Abstract
Pediatric community-acquired bloodstream infections (CA-BSIs) in sub Saharan African humanitarian contexts are rarely documented. Effective treatment of these infections is additionally complicated by increasing rates of antimicrobial resistance. We describe the findings from epidemiological and microbiological surveillance implemented in pediatric patients with suspected CA-BSIs presenting for care at a secondary hospital in the conflict affected area of Zamfara state, Nigeria. Any child (> 2 months of age) presenting to Anka General Hospital from November 2018 to August 2020 with clinical severe sepsis at admission had clinical and epidemiological information and a blood culture collected at admission. Bacterial isolates were tested for antibiotic susceptibility. We calculated frequencies of epidemiological, microbiological and clinical parameters. We explored risk factors for death amongst severe sepsis cases using univariable and multivariable Poisson regression, adjusting for time between admission and hospital exit. We included 234 severe sepsis patients with 195 blood culture results. There were 39 positive blood cultures. Of the bacterial isolates, 14 were Gram positive and 18 were Gram negative; 5 were resistant to empiric antibiotics: methicillin-resistant Staphylococcus aureus (MRSA; n = 2) and Extended Spectrum Beta-Lactamase positive enterobacterales (n = 3). We identified no significant association between sex, age-group, ward, CA-BSI, appropriate intravenous antibiotic, malaria positivity at admission, suspected focus of sepsis, clinical severity and death in the multivariable regression. There is an urgent need for access to good clinical microbiological services, including point of care methods, and awareness and practice around rational antibiotic in healthcare staff in humanitarian settings to reduce morbidity and mortality from sepsis in children.
Similar content being viewed by others
Introduction
Sepsis, a life threatening multi-organ dysfunction caused by a dysregulated host response to infection1, is considered one of the main causes of mortality in children globally. In 2017, there were an estimated 20.3 million sepsis cases globally among children younger than 5 years of age2. Mortality from sepsis in children is also high, and up to 25% hospital mortality has been reported in a multi-center study for children admitted for intensive care with severe sepsis3.
Approximately 70% of all sepsis cases are thought to be community-acquired infections4. Public health prevention of these infections can be achieved by high vaccination coverage and improved access to adequate sanitation and water (both quality and quantity)5. However, early diagnosis and appropriate clinical management (antimicrobials and fluid resuscitation) reduce mortality and long-term morbidity5. The majority of pediatric community-acquired sepsis are from pneumonia or meningitis, but unknown sources of infection are also common3. Identifying a causative pathogen in patients with sepsis is also not guaranteed, even in high resource settings6,7. Two recent studies in pediatric intensive care units (PICUs) in seven European countries and the United States, isolated a bacteria in 54% and 48% of pediatric sepsis cases, respectively6,7.
The incidence and clinical outcomes of bloodstream infections (BSIs) in pediatric patients in sub Saharan Africa are poorly documented8. Information from humanitarian settings in Africa is even scarcer. A recent systematic review on community-acquired BSIs (CA-BSIs) in low- and middle-income countries identified only nine studies on the topic in four African countries (Kenya, Mozambique, Nigeria and Zimbabwe)9. A recent study in South Africa showed that 35% of BSIs identified in hospitalized children were community-acquired infections (the remainder healthcare associated or hospital acquired), with 47% due to Gram-negative bacteria (GNB) infections (mostly E. coli) and 50% due to Gram-positive bacterial (GPB) infections (mostly Staphylococcus aureus and Streptococcus pneumoniae)8. Another study in South Africa with CA-BSI surveillance in pediatric inpatients, showed that 40% of infections were due to GNB (54% were E. coli) and 60% due to GPB (45% from S. aureus)10.
Surveillance for antibiotic resistance is improving across the continent, but information on bacterial susceptibility to commonly used antibiotics for the treatment of BSIs in pediatric patients remains limited and reported in non-standardized ways11. Even so, reported resistance to commonly prescribed antibiotics across Africa is high12 and it is assumed to be similar in humanitarian contexts. A systematic review on resistance in bacteria isolated in children with sepsis in Africa showed that E. coli isolates were resistant to ampicillin, gentamicin and ceftriaxone of 93%, 29% and 16% respectively12. Similarly, S. aureus isolates showed resistance of 90%, 29% and 20% for ampicillin, gentamicin and cloxacillin respectively12.
Since 2014 Zamfara state in northwest Nigeria is affected by an intensified internal conflict and population displacement13. Since 2018, the Ministry of Health and Médecins Sans Frontières (MSF) implement antimicrobial resistance (AMR) control activities (antibiotic stewardship, infection prevention and control and microbiological surveillance) in the pediatric inpatient, isolation and inpatient therapeutic feeding centre (ITFC; dedicated to children with severe acute malnutrition) of Anka General Hospital (AGH) (Fig. 1). We report the clinical, epidemiological and microbiological findings from pediatric admissions to the hospital between November 2018 and August 2020 with suspected severe sepsis.
Map indicating the location of Anka (Zamfara State) where AGH is and Sokoto (Sokoto State) where the microbiology laboratory is based with the road that connects them. Note: Contains information from OpenStreetMap and OpenStreetMap Foundation, which is made available under the Open Database License.
Methods
Study setting
Access to the hospital is hampered by the lack of primary healthcare referral options, distance and security to travel. A hospital emergency triage functions as the sole point for hospital admission to the isolation, inpatient or ITFC wards (150 beds total). The hospital discharged an average of 769 pediatric patients per month during 2019 (range: 419–1066 per month; unpublished data, MSF).
Severe sepsis upon admission management
From October 2018, we established surveillance for CA-BSIs in pediatric patients that presented to the emergency triage with signs and symptoms of severe sepsis. A case of suspected severe sepsis was any pediatric patient over the age of 2 months that presented with at least two of the following clinical criteria: axillary temperature > 38 °C or < 35.5 °C, tachycardia or bradycardia for their age, tachypnea for age and/or oxygen saturation < 92% or leukocytosis or leukopenia or > 10% immature neutrophils and one of the following criteria: decreased level of consciousness (pain or unresponsive on the alert/verbal/pain/unresponsive [AVPU] scale) or circulatory insufficiency (defined as tachycardia with weak pulse or capillary refill time > 3 s or oligo-anuria).
Any severe sepsis patient had a blood culture taken and was treated using standard clinical protocols14. Empirical treatment for severe sepsis is with intravenous (IV) ceftriaxone monotherapy with the addition of: cloxacillin (suspected sepsis source: skin/soft tissue); metronidazole (suspected sepsis source: aspiration pneumonia); clindamycin (suspected sepsis source: bone/joint); metronidazole (suspected sepsis source: intestinal/biliary or abdominal).
We collected information on: age, sex, weight, date of admission, date of and reason for exit, primary diagnosis at exit, admission ward, suspected source of sepsis at admission, signs of sepsis at admission (respiratory rate, heartrate, skin color, urine output etc.), malaria positivity from rapid diagnostic tests, use of antibiotics prior to admission to hospital, start and end dates of IV antibiotics received during hospitalization and clinical outcome.
Microbiological testing
Blood cultures were collected from severe sepsis patients identified at admission using an aseptic technique. Not all patients presenting with severe sepsis had blood cultures performed due to transport problems, staff availability and logistical constraints (i.e. security restrictions for transport, temporary rupture of materials etc.). For infants or malnourished children, a single blood volume of 1-2 mL was collected and for pediatric patients > 1 years of age a single blood volume of 2.5–5 mL was collected using neonatal and pediatric aerobic blood collection bottles respectively (Liofilchem, Italy). Following collection, culture bottles were incubated at 35 °C. Culture bottles were transported at ambient temperature in triple packaging by a private taxi service to the microbiology laboratory (established in July 2018) in Noma Children’s Hospital (NCH) in Sokoto (Sokoto State) (3 h drive from Anka). Positive blood cultures were Gram stained to determine subsequent choice of enriched and/or selective media for subculture (Polyvitex (PVX), Columbia Colistin Nalidixic Acid (CNA), MacConkey (MAC) or CHROMagar Orientation (CRO) agar plates). Colonies obtained from the media were identified using standard biochemical tests and the API® system (bioMérieux, France). Antibiotic susceptibility was assessed using the Kirby–Bauer disk diffusion method on Mueller–Hinton agar and interpreted following the European Committee on Antimicrobial Susceptibility Testing (EUCAST) recommended breakpoints from 201915. The control organisms used for quality control were American type culture collection strains (ATCC) based on the EUCAST recommendation.
A confirmed CA-BSI was defined as any patient for whom a positive blood culture (bacterial or yeast) was obtained that was not a contaminant. Coagulase-negative Staphylococci, Bacillus sp., Corynebacterium sp. or Propionibacterium sp. were considered contaminants. For other bacteria or yeast, the laboratory team discussed the result with the antibiotic stewardship focal point in AGH to determine if the organism was the likely cause of symptoms. If the microorganism was determined not to fit the clinical presentation, the bacteria were considered a probable contaminant.
Data sources
We combined data from three separate sources for this current analysis: the health information data (District Health Information Software [DHIS2]), a severe sepsis database (Epi Data Manager; http://www.epidata.dk/download.php; which stored the additional clinical and epidemiological data collected for all pediatric patients with severe sepsis) and the WHONET microbiological database (http://www.whonet.org).
Data analysis
We merged all datasets and retained data from DHIS2 and WHONET if the patient identification number was available in the severe sepsis dataset. Patient characteristics at admission were described for all severe sepsis cases by age group, sex, ward of admission, duration of hospitalization and clinical outcome. We were unable to calculate a Pediatric Early Warning Score (PEWS) from the data available, thus we created a modified severity score. We assigned scores of 0, 1, 2 or 3 (3 being the most severe) to each category of presentation for capillary refill time (< 3 s = 0; CRT 3–4 s = 1 and CRT > = 5 s = 2), oxygen saturation (95–100% = 0; 90–95% = 1; < 90% = 2; < 80% = 3), AVPU (Alert, Verbal, Pain, Unresponsive: A = 1, V = 2, P = 3, U = 3), respiratory rate (normal = 0, tachypnoea = 2, bradypnea = 3), respiratory distress (1), tachycardia (1) and bradycardia (1). We calculated the 25th quartile and the median for the total severity scores in this patient population. Low severity was defined as being below the 25% quartile, moderate severity was from the 25th–50th quartile and high severity was greater than the 50% quartile.
Due to resource constraints, we limited data collection related to treatment received during hospitalization to IV antibiotics only. Appropriateness of antibiotic therapy was independently reviewed by FC and KC using compliance to the MSF Pediatric Guidelines14 for diagnosis and treatment of sepsis and fever and evaluating whether antibiotics were switched when results of blood cultures were available. The clinical presentation of severe sepsis, clinical course and outcomes was compared between patients admitted to the ITFC or the pediatric/isolation wards. We assessed the differences between the proportions of outcomes in both hospital wards using Pearson’s chi-squared test. The time of IV antibiotic administration was calculated as the time from IV insertion to removal (for reasons of death, de-escalation, discharge etc.). We performed Kaplan–Meier survival analysis to assess the time to death following IV antibiotic administration in the ITFC and pediatric/isolation wards. The log rank test was used to determine if the difference between the survival curves was statistically significant.
We compared epidemiological and clinical risk factors upon admission for patients with confirmed bloodstream infections with those who had blood culture contamination or a negative blood culture using descriptive statistics and testing for significant differences using Chi-square and Fisher exact test (p-values under 0.05 were considered statistically significant). Only one clinical variable showed evidence of a statistically significant difference between the two groups, thus we did not conduct further regression analysis.
To understand the risk factors that contributed to death in the severe sepsis patient group, we calculated person-time as the days from date of admission until date of death or other outcome (discharged, Left Against Medical Advice [LAMA], Lost to Follow Up [LTFU] and referred). We were unable to correctly ascertain treatment switch for each patient (as we only collected data on IV antibiotic treatment and not oral antibiotic treatment). Time to treatment switch to appropriate antibiotics was not taken into account in this analysis. Crude and adjusted rate ratios (iRR) were calculated using Poisson regression models. All variables with p ≤ 0.2 in the univariate analysis were included in the multivariable model.
All data analysis was done using R (Version 4.0.2) and Rstudio (Version 1.3.1056). Microbiological data was analyzed using the AMR for R package which applies EUCAST clinical breakpoints and dosing of antibiotic guidance 202016,17.
Ethics approval and consent to participate
This study used routinely collected clinical and microbiological data from patients to inform their clinical management and monitor program implementation. As the data was not collected for research purposes, no explicit informed consent for the collection of data was required. The Nigerian Federal Ethical Review Board and the Zamfara Ethical Review board exempted the study from ethical review. This research fulfilled the exemption criteria set by the Médecins Sans Frontières Ethics Review Board for a posteriori analyses of routinely collected clinical data and thus did not require MSF ERB review. It was conducted with permission from Melissa McRae, Medical Director, Operational Centre Amsterdam (OCA), Médecins Sans Frontières.
Results
Overall characteristics of severe sepsis patients
We analysed information from 234 patients of severe sepsis who were admitted to AGH between 1 November 2018 and 31 August 2020. The sex distribution between patients was similar, most patients were below the age of 2 years and most were admitted to the pediatric ward (Table 1). Thirty-five percent (n = 82) of severe sepsis patients died during their hospitalization with almost half (49%) dying within 24 h of admission to hospital (Table 1). Most ITFC patient (91%) were younger than 2 years of age, compared to 64% in the pediatric/isolation ward (p < 0.001). ITFC patients also had a significantly higher mortality rate compared to those in the pediatric and isolation ward (49% vs. 28%) (Table 1).
Most patients with severe sepsis at admission had a suspected respiratory focus (38%) or an unknown focus (21%) (Table 2). Patients in the ITFC were more likely to have respiratory and gastrointestinal focus of sepsis whereas those admitted to the pediatric or isolation wards were more likely to have an unknown or suspected CNS focus (Table 2). The proportion of patients with a suspected source of sepsis from skin and soft tissue infection were similar in both wards. The malaria positivity rate in those presenting with suspected CNS focus was significantly higher compared to all the other categories of suspected focus of sepsis (82% vs. 58%, p = 0.003). Only two patients were recorded as having received antibiotics before admission to AGH, but this variable was missing for 165 severe sepsis patients (71%).
Patients in ITFC received IV antibiotics for a median of four days compared with three days in the pediatric/isolation ward (Table 3). The main reason for cessation of IV antibiotics was death in the ITFC and discharge in the pediatric/isolation ward (Table 3). The Kaplan–Meier survival analysis showed little evidence of a difference between the time to death following IV insertion (p = 0.08; supplemental information). Most patients received IV ceftriaxone (in line with clinical algorithms). Receipt of two or more antibiotics was more common in the ITFC patients (77% vs 46.5%) as was cloxacillin use (Table 4). Fewer children in ITFC received appropriate antibiotics compared to those in admitted to pediatric/isolation wards (61% vs 78%, p = 0.01) (Table 3).
Confirmed bacterial bloodstream infections
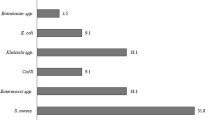
195 patients with severe sepsis (83.3%) had blood cultures taken. They were negative in 116 patients (59.5%) and positive in 79 patients (40.5%). Forty isolates (50.6%) were considered contaminants and 39 isolates (49.3%) were considered confirmed CA-BSIs. Fifty percent of contaminants were coagulase-negative Staphylococcus and unknown GPB (Table 4). We also classified six Aeromonas sp. isolates, three Burkholderia cepacia and one Achromobacter sp. isolates as contaminants. Of the confirmed CA-BSIs, most were GNB (n = 18), followed by GPB (n = 14) and yeasts (n = 6). Most of the GNB were Pseudomonas sp. (n = 8), three of which were P. aeruginosa (all sensitive to gentamicin). We identified three ESBL positive isolates of Escherichia coli, Klebsiella pneumoniae and Serratia liquefaciens. Most of the GPB were Staphylococcus aureus (n = 11) of which two were methicillin resistant (MRSA). Six blood cultures contained yeast of which one was Candida albicans and the remainder were unspecified (Table 4).
Fifty-one percent of patients with confirmed BSIs were alert (A) and verbal (V) at admission compared to 37% of patients without confirmed BSIs. Patients with confirmed BSIs were less likely to be unresponsive at admission (2.6% vs 19%, p = 0.008). All other clinical criteria at admission were similar between these two patient groups.
Information on the time to communication of preliminary and final results was available for 151 blood cultures (77%) and 158 blood cultures (81%) respectively. Preliminary results were communicated with a mean of 3.2 days (SD: 1.16; median = 3 days) from the date of admission and final results were communicated with a mean of 7.1 days (SD: 2.4 median = 7 days) from the date of admission. For 116 (76%, n = 153) patients, the final blood culture results arrived an average 4.9 days (median: 4) after their exit from hospital (for all clinical outcomes). For 36 patients (24%), the final blood culture results arrived an average of 6.3 days (median: 5 days) before their exit from hospital. For 157 patients, for whom the information was available, 93% (n = 146) received their blood culture results after the last dose of IV antibiotics was administered.
Risk factors for death
The univariable Poisson regression only identified male patients as having a lower risk of death compared to female patients (p = 0.04; Table 5). In the multivariable model, no further epidemiological or clinical characteristics showed a statistically significant association with death (Table 5).
Discussion
In this acute humanitarian setting in Zamfara state, pediatric patients that present for hospital care with signs and symptoms of severe sepsis have high mortality rates (35%) and approximately 20% have confirmed CA-BSIs. We were unable to determine any clear epidemiological or clinical risk factors for death in this patient group. We also showed that confirmed CA-BSIs are not associated with higher mortality rates compared with other causes. This might be from the high proportion of patients receiving the correct empirical antibiotic treatment upon admission.
The mortality in pediatric patients with suspected bloodstream infections in Anka is higher than the average pediatric mortality reported from secondary hospitals in Nigeria in Aba (5.7%)18 and Asaba (5.8%)19. It is also higher than the overall average monthly pediatric hospital mortality from AGH (8.3%, MSF unpublished data). This speaks to the severity of disease in patients presenting with severe sepsis at AGH. Even though global estimates for hospital mortality from sepsis in children is around 25%3, the higher rates in Anka might also be linked to difficulties in accessing the hospital as suggested by the high proportion of patients that die within 24 h of being admitted to the hospital20.
The 20% blood culture positivity rate identified in our study is similar to those obtained in the few available studies from hospitalized children < 5 years of age in Nigeria, ranging between 11 and 30%21,22,23. Half of the bacteria implicated in confirmed CA-BSI in this study were GNB. This is similar to the results from a recent meta-analysis of pediatric CA-BSI that included six studies from African hospitals where 54.8% (CI95% 45.1–64.4) of infections were due to GNB. ESBL producing bacteria were identified in 22% of the GNB isolates which is higher than the 6% of ESBL producing Enterobacterales isolated in Enugu (Nigeria) in 201924 but closer to the 39% of ESBL positive GNB reported from central and northwest Nigeria in 2020 in similar patient groups22. It has been shown that malnourished children have an increased risk for acquisition of ESBL producing Enterobacterales when exposed to amoxicillin25. The high prevalence of ESBL-GNB at admission in Anka should also be a reminder for antibiotic stewardship and infection prevention and control in the hospital, due to the high numbers of severe acute malnutrition patients treated and risk for hospital-acquired infections with ESBL-GNB in this patient group. We were unable to differentiate the Salmonella isolates, but it has been previously described that the majority of these isolates in children in Nigeria are from S. typhi21,23. We are unable to comment on the high rate of yeast positive blood cultures in our patient population due to the limited additional clinical information available on these patients.
The widespread use of antibiotics in Nigeria without doctors prescriptions is well documented26. We had difficulty identifying which patients had received antibiotics and what these antibiotics were prior to their admission to hospital. This may have influenced the treatment choice for the individual patient and it could have facilitated the interpretation of negative blood cultures. We currently do not test for HIV status in the pediatric population, and even though the overall prevalence is assumed to be low, this information would allow for better interpretation of blood culture results (particularly yeast). Approximately half of the positive blood cultures were contaminated which is an issue that we are actively working to reduce through training staff in aseptic blood collection. Due to resource constraints, we have only been using a single blood culture bottle, but are working towards using two bottles as standard practice. Our ability to analyze the clinical progression of severe sepsis patients for their duration of hospitalization was limited due to the restricted amount of data we were able to collect thus we were unable to evaluate fluid resuscitation and de-escalation to oral antibiotics. This could further improve the current clinical management and AMR activities if resources allow.
Conclusions
Pediatric sepsis accounts for approximately 8% of all critically ill children27. In 2020 WHO called for global action on sepsis with a focus on improving data from low-income countries with better definitions for sepsis and more information on microbiological aspects and antibiotic resistance28,29. Our study sheds much needed light on the incidence and causative organisms implicated in pediatric sepsis in a humanitarian context in Nigeria. Interventions around antibiotic stewardship require context-specific approaches30 and humanitarian settings are no different. The current study reinforces the urgent need for improved clinical bacteriology to improve awareness and practice around optimal antibiotic use in clinical care providers in these locations31. In the absence of microbiology, new, affordable and validated point of care diagnostic tools might also reduce inappropriate antibiotic use32. In addition, the mainstay activities of humanitarian response to increase vaccination coverage, improve water and sanitation conditions, increase access to early diagnostics and healthcare and improve the nutritional status of affected populations will serve to prevent CA-BSIs. These preventative measures should be included as part of context specific stewardship activities in humanitarian settings.
Data availability
MSF has a managed access system for data sharing. Data are available on request in accordance with MSF’s data sharing policy. Requests for access to data should be made to data.sharing@msf.org. For more information please see: (1) MSF’s Data Sharing Policy: http://fieldresearch.msf.org/msf/handle/10144/306501, (2) MSF’s Data Sharing Policy PLOS Medicine article: http://journals.plos.org/plosmedicine/article?id=10.1371/journal.pmed.1001562.
Abbreviations
- AGH:
-
Anka General Hospital
- AMR:
-
Antimicrobial resistance
- AVPU:
-
Alert, Verbal, Pain, Unresponsive
- CA-BSI:
-
Community acquired bloodstream infection
- CAN:
-
Columbia Colistin Nalidixic Acid
- CNS:
-
Central nervous system
- CRO:
-
CHROMagar Orientation
- ESBL:
-
Extended Spectrum Beta-Lactamase
- EUCAST:
-
European Committee on Antimicrobial Susceptibility Testing
- GNB:
-
Gram negative bacteria
- GPB:
-
Gram positive bacteria
- HIV:
-
Human immunodeficiency virus
- IQR:
-
Inter quartile range
- ITFC:
-
Inpatient therapeutic feeding centre
- IV:
-
Intravenous
- LAMA:
-
Left against medical advice
- LTFU:
-
Lost To Follow up
- MAC:
-
MacConkey
- MRSA:
-
Methicillin Resistance Staphylococcus aureus
- MSF:
-
Médecins Sans Frontières
- NCH:
-
Noma Children’s Hospital
- PEWS:
-
Pediatric Early Warning Score
- PICU:
-
Pediatric Intensive Care Unit
- PVX:
-
Polyvitex
- RDT:
-
Rapid Diagnostic Test
- SD:
-
Standard deviation
- WHO:
-
World Health Organisation
References
Singer, M. et al. The third international consensus definitions for sepsis and septic shock (sepsis-3). JAMA J. Am. Med. Assoc. 315, 801–810 (2016).
Rudd, K. E. et al. Global, regional, and national sepsis incidence and mortality, 1990–2017: Analysis for the Global Burden of Disease Study. Lancet 395(10219), 200–211 (2020).
Weiss, S. L. et al. Global epidemiology of pediatric severe sepsis: The sepsis prevalence, outcomes, and therapies study. Am. J. Respir. Crit. Care Med. 191(10), 1147–1157 (2015).
Reinhart, K. et al. Recognizing sepsis as a global health priority—A WHO resolution. N. Engl. J. Med. 377(5), 414–417 (2017).
World Health Organisation (WHO). Improving the prevention, diagnosis and clinical management of sepsis [Internet] (2020). https://www.who.int/servicedeliverysafety/areas/sepsis/en/
Boeddha, N. P. et al. Mortality and morbidity in community-acquired sepsis in European pediatric intensive care units: A prospective cohort study from the European Childhood Life-threatening Infectious Disease Study (EUCLIDS). Crit. Care 22(1), 143 (2018).
Ames, S. G. et al. Infectious etiologies and patient outcomes in pediatric septic shock. J. Pediatr. Infect. Dis. Soc. 6(1), 80–86 (2017).
Lochan, H., Pillay, V., Bamford, C., Nuttall, J. & Eley, B. Bloodstream infections at a tertiary level paediatric hospital in South Africa. BMC Infect. Dis. 17(1), 750 (2017).
Droz, N. et al. Bacterial pathogens and resistance causing community acquired paediatric bloodstream infections in low- and middle-income countries: A systematic review and meta-analysis. Antimicrob. Resist. Infect. Control 8, 207 (2019).
Crichton, H., O’Connell, N., Rabie, H., Whitelaw, A. C. & Dramowski, A. Neonatal and paediatric bloodstream infections: Pathogens, antimicrobial resistance patterns and prescribing practice at Khayelitsha District Hospital, Cape Town, South Africa. South Afr. Med. J. 108(2), 99 (2018).
Tadesse, B. T. et al. Antimicrobial resistance in Africa: A systematic review. BMC Infect. Dis. 17(1), 1–17 (2017).
Williams, P. C. M., Isaacs, D. & Berkley, J. A. Antimicrobial resistance among children in sub-Saharan Africa. Lancet Infect. Dis 18, e33–e44 (2018).
Anyadike, O. The longshot bid to end rampant banditry in Nigeria’s northwest. The New Humanitarian (2021).
Medecins Sans Frontieres (MSF). Pediatric Guidelines: Adapted for Operational Centre Amsterdam (March 2018), 107–113 (2018).
EUCAST Breakpoint tables for interpretation of MICs and zone diameters. Version 9.0, 2019 (2019).
European Committee on Antimicrobial Susceptibility Testing (EUCAST). EUCAST Breakpoint tables for the interpretation of MICs and zone diameters. Version 10.0 (2020).
Berends, M. et al. AMR—An R package for working with antimicrobial resistance data. BioRxiv. https://doi.org/10.1101/810622 (2019).
Okoronkwo, N. C., Onyearugha, C. N. & Ohanenye, C. A. Pattern and outcomes of paediatric medical admissions at the Living Word Mission Hospital, Aba, South East Nigeria. Pan. Afr. Med. J. 30, 202 (2018).
Ezeonwu, B., Chima, O., Oguonu, T., Ikefuna, A. & Nwafor, I. Morbidity and mortality pattern of childhood illnesses seen at the children emergency unit of federal medical center, Asaba, Nigeria. Ann. Med. Health Sci. Res. 4(9), 239 (2014).
Maisa, A., Lawal, A. M., Islam, T., Nwankwo, C., Oluyide, B. et al. Impact of healthcare awareness and access on patient mortality and loss to follow-up in two paediatric hospital wards in Zamfara, North-West Nigeria, 2016–2018 (submitted).
Obaro, S. K. et al. Salmonella bacteremia among children in central and Northwest Nigeria, 2008–2015. Clin. Infect. Dis. 61(Suppl 4), S325–S331 (2015).
Duru, C. et al. Molecular characterization of invasive Enterobacteriaceae from pediatric patients in Central and Northwestern Nigeria. PLoS One 15, e0230037 (2020).
Popoola, O. et al. Bacteremia among febrile patients attending selected healthcare facilities in Ibadan, Nigeria. Clin. Infect. Dis. 69(Suppl 6), S466–S473 (2019).
Oli, O. et al. Multi-antibiotic resistance and factors affecting carriage of extended spectrum β-lactamase-producing enterobacteriaceae in pediatric population of Enugu Metropolis, Nigeria. Med. Sci. 7(11), 104 (2019).
Maataoui, N. et al. Increased risk of acquisition and transmission of ESBL-producing Enterobacteriaceae in malnourished children exposed to amoxicillin. J. Antimicrob. Chemother. 75(3), 709–717 (2020).
Awosan, K. J., Ibitoye, P. K. & Abubakar, A. K. Knowledge, risk perception and practices related to antibiotic resistance among patent medicine vendors in Sokoto metropolis, Nigeria. Niger. J. Clin. Pract. 21(11), 1476–1483 (2018).
Harder, T., Seidel, J., Eckmanns, T., Weiss, B. & Haller, S. Predicting late-onset sepsis by routine neonatal screening for colonisation by gram-negative bacteria in neonates at intensive care units: A protocol for a systematic review. BMJ Open 7(3), e014986 (2017).
World Health Organization (WHO). WHO calls for global action on sepsis—cause of 1 in 5 deaths worldwide [Internet] (2020). https://www.who.int/news/item/08-09-2020-who-calls-for-global-action-on-sepsis---cause-of-1-in-5-deaths-worldwide
World Health Organization (WHO). Global report on the epidemiology and burden of sepsis: current evidence, identifying gaps and future directions (2020).
Punpuing, S. et al. Community-based antibiotic access and use in six low-income and middle-income countries: A mixed-method approach. Lancet Glob. Health. https://doi.org/10.1016/S2214-109X(21)00024-3 (2021).
Ombelet, S. et al. Clinical bacteriology in low-resource settings: Today’s solutions. Lancet Infect. Dis. 18, e248–e258 (2018).
Do, N. T. T. et al. Point-of-care C-reactive protein testing to reduce inappropriate use of antibiotics for non-severe acute respiratory infections in Vietnamese primary health care: A randomised controlled trial. Lancet Glob. Health 4(9), e633–e641 (2016).
Acknowledgements
We would like to thank all laboratory, clinical staff and data coding staff in AGH and the microbiology laboratory in Noma Children’s hospital for the hard work that made this surveillance possible. We would like to thank Omar Contigiani for facilitating meaningful data quality checks in the existing surveillance data in AGH. We would also like to thank Zhian Kamvar for his support in coding scripts in R. We would like to thank Nienke Meeuwissen for the map.
Funding
This research received no specific grant from any funding agency in the public, commercial or not-for-profit sectors. MSF staff carried out the research as part of their routine roles. The corresponding and final authors had full access to all data and final responsibility for the decision to submit for publication.
Author information
Authors and Affiliations
Contributions
F.C., A.L., H.W., J.H. and K.C. conceived the study design. F.C., A.L., R.O., A.M.L., B.O., G.O., D.G., H.R. and K.C. contributed to the acquisition of data. F.C., A.L., C.A. and K.C. contributed to the analysis of data. F.C. and A.L. drafted the manuscript. All authors contributed to the interpretation of data and revised the manuscript. All authors have approved the submitted version of this manuscript and have agreed to be personally accountable for their own contributions.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chukwumeze, F., Lenglet, A., Olubiyo, R. et al. Multi-drug resistance and high mortality associated with community-acquired bloodstream infections in children in conflict-affected northwest Nigeria. Sci Rep 11, 20814 (2021). https://doi.org/10.1038/s41598-021-00149-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-00149-1
This article is cited by
-
Epidemiology and clinical presentation of community-acquired Staphylococcus aureus bacteraemia in children under 5 years of age admitted to the Manhiça District Hospital, Mozambique, 2001–2019
European Journal of Clinical Microbiology & Infectious Diseases (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.