Abstract
The gut microbiome plays an important role in early life, protecting newborns from enteric pathogens, promoting immune system development and providing key functions to the infant host. Currently, there are limited data to broadly assess the status of the US healthy infant gut microbiome. To address this gap, we performed a multi-state metagenomic survey and found high levels of bacteria associated with enteric inflammation (e.g. Escherichia, Klebsiella), antibiotic resistance genes, and signatures of dysbiosis, independent of location, age, and diet. Bifidobacterium were less abundant than generally expected and the species identified, including B. breve, B. longum and B. bifidum, had limited genetic capacity to metabolize human milk oligosaccharides (HMOs), while B. infantis strains with a complete capacity for HMOs utilization were found to be exceptionally rare. Considering microbiome composition and functional capacity, this survey revealed a previously unappreciated dysbiosis that is widespread in the contemporary US infant gut microbiome.
Similar content being viewed by others
Introduction
The neonatal period represents a unique stage of life when critical foundations of lifelong health are established1,2. Throughout this period, the gut microbiome can provide protection against enteric infections and is crucial for the proper development of the immune system3,4. In infants, it is well established that gut microbiome perturbations characterized by the overrepresentation of potentially pathogenic taxa are implicated in mediating persistent pathophysiological and immune abnormalities, including heightened risk for immunological disorders later in life and acute chronic inflammation3,5,6,7. These gut microbial perturbations, conceptually referred to as dysbiosis, are strongly associated with the absence of infant-associated Bifidobacterium that encode key functions required for the metabolization of human milk oligosaccharides (HMOs) in the infant8.
Disruptions to the optimum infant intestinal microbiome are thought to result from changes to the infant’s diet, antibiotic exposure, and cesarean section delivery, as well as other interventions or exposures that might alter the transmission of microbes from mother to infant or the communities themselves9,10,11,12. However, there is mounting evidence suggesting that disparities between microbiome compositions observed today in different human populations may also echo historical changes including migrations13, historical infant feeding practices14, or the sum of all changes associated with modern lifestyles15. Despite the importance of the microbiome for infant health, nutrition, and development, most reports on the infant gut microbiome in developed countries have described communities in preterm infants16,17, and only few studies have utilized metagenomics to characterize microbiome function at distinct sites from large cohorts of infants18,19. Recent metagenomic surveys focused on genetically similar, but socioeconomically distinct populations in three European countries identifying microbiome compositions in early life with clear implications for long term health and development regarding the development of autoimmune diseases including Type 1 Diabetes19. Other studies have identified health status-specific differences among infants in low-income countries that later impacted vaccine responses, further signaling the important role that the microbiome in early life plays in immune development20,21. Similarly, recent large-scale studies have identified parallel differences in the microbiomes among infants even within high-income countries in Europe, which were associated with the development of autoimmune diseases such as asthma22. Mechanistic support for the critical role Bifidobacterium play in early life immune development has also been recently described23. Thus, it appears increasingly important to describe population-level norms among otherwise healthy infants within and across countries, especially large, ethnically, and socioeconomically diverse countries such as the United States.
In order to begin to address this need for population level surveys of the healthy infant gut microbiome in the United States, fecal samples from 227 infants (0–6 months of age) were collected from pediatric research sites located in five states, representing a cross-section of infants from across the United States. Stool samples underwent shotgun metagenomics sequencing. Specifically, we applied shotgun metagenomics to characterize: (1) gut bacterial communities of healthy US infants in the first 6 months of life; (2) ecosystem functions by determining the metabolic potential of gut microbiomes in different enterotypes to metabolize human milk oligosaccharides (HMOs) from breast milk; and (3) the carriage of antibiotic resistant genes (ARGs) in infants across different US states. The above criteria were then used to classify microbiomes as dysbiotic or not, based on the concept of ecosystem services24 adapted to evaluate the benefits infants receive from functions provided by their gut microbiomes8.
Overall, this survey revealed that, on average, infants in the US have a low abundance of Bifidobacterium, a high abundance of potentially pathogenic bacteria carrying high levels of ARGs, as well as limited capacity of metabolizing HMOs from breast milk. These findings were widespread in infants in the US, independent of location, age, and diet. This survey offers a new perspective when considering infants in the context of a healthy microbiome and the acute and long-term consequences it implies.
Results
Infant gut microbiome compositions vary independently of age, diet and location
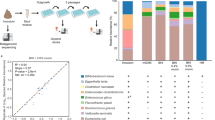
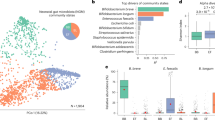
Our cohort of 227 samples (one sample per subject) from 7 research sites in 5 states (California, Georgia, Oregon, Pennsylvania, South Carolina), was assessed for microbiome composition and function via shotgun metagenomic sequencing. Demographic data for every subject is reported in Supplementary Table 1. After quality filtering, Illumina sequencing led to an average of 28 million PE reads per sample (± SD 6.4 million), of which 4.6% (± SD 12%) per sample on average were discarded as human reads. High-quality, human-filtered reads were subjected to taxonomic and functional profiling (see “Methods”). A total of 367 classified bacterial species belonging to 119 genera, 53 families, 25 orders, 15 classes, and 7 phyla were identified across the samples (Supplementary Table 2). Potential pathogenic bacteria were present among the top ten bacterial families, which combined accounted for 93% of the infant’s microbiome. The most abundant on average were Enterobacteriaceae (35%), Streptococcaceae (5.6%), Clostridiaceae (3.6%), and Staphylococcaceae (1.23%) (Fig. 1A). The bacterial family of Bifidobacteriaceae was low on average in infants from all five states (19.9%) with the lowest abundance in infants from Georgia (12%) and the highest (37%) in Pennsylvania. When considering only samples collected within the first 100 days of life (0–3 months), a critical window of time for immune system development25, we did not see a difference by state in terms of bacterial families composing the microbiome, with a composition that appears to be independent of location25 (Supplementary Fig. 1). Splitting the data by both age (0–3 vs. 4–6 months) and diet (exclusively breast or formula fed and mixed), revealed that the 0–3 months samples had a higher individual variation in bacterial family composition compared to the 4–6 months samples, probably due to the natural acquisition of bifidobacteria (Fig. 1B).
Furthermore, we performed a microbiome multivariable association analysis with linear models, using MaAsLin2 (see “Methods”), by including all the clinical variables available in this study to test for microbiome differences at the family level according to metadata. Interestingly, while we did find significant taxa associated with few metadata variables, most of the bacterial families reported, were very low in relative abundance overall (< 2%) and thus we do not believe that their biological impact is of a major concern or clinical interest. However, two top families, namely Bifidobacteriaceae (19.9%) and Bacteroidaceae (10.4%) were all significantly associated with age but not with any other variable (Supplementary Table 6), confirming again the natural acquisition of these families from the environment as the infants grow older.
At the species level, we identified known potentially pathogenic species of particular relevance, especially as it pertains to the neonatal population, accounting for 26% of the microbiome on average in all samples (Supplementary Table 2). In order of abundance we identified, Escherichia coli (12.6%) the most abundant among all species overall, Klebsiella pneumoniae (7%), Klebsiella oxytoca (2.5%), Enterobacter cloaceae (2.7%), Clostridium perfringens (1.1%), Clostridium difficile (0.1%), Staphylococcus aureus (0.03%) and Streptococcus agalactiae (0.01%). We also identified a taxon classified as Escherichia (5%) for which the species level assignment was not classified.
Bifidobacteria levels are low on average in the US independent of diet
As bifidobacteria are considered a fundamental component of a healthy infant gut microbiome, we performed targeted genus-, species- and strain-level metagenomic analysis using unique clade-specific marker genes (see “Methods”). Overall, the average abundance of Bifidobacterium was 20%. When accounting for diet, within the first 100 days of life (0–3 months) or later (4–6 months), there were no significant differences in the abundance of Bifidobacterium, independent of whether the infants were exclusively fed breast milk, formula or were fed a mixed diet of breast milk and formula (Fig. 2A). However, breastfed infants in the 4–6 months age group, had a higher abundance of Bifidobacterium (75% of the microbiome on average), though not significantly different (P > 0.05) amongst feeding types, likely due to high individual variation from the formula fed infants in that age group, which had an average abundance of Bifidobacterium below 25%. At the species level, we identified 10 Bifidobacterium species, with the most abundant being Bifidobacterium longum (8.5%), Bifidobacterium breve (5.8%), and Bifidobacterium bifidum (3.75%) (Fig. 2B). All three species were significantly lower in the 0–3 months age group compared to the older infants (4–6 months) and had a greater degree of intra-group variability with samples ranging from 0% to more than 90% of relative abundance of B. breve, B. longum, or B. bifidum.
Relative abundance of Bifidobacterium at the genus and species level. (A) Bifidobacterium relative abundance (%) grouped by age and diet. (B) Relative abundance (%) of all species identified within the Bifidobacterium genus and grouped by patient age. The lower and upper hinges of the box plots correspond to the first and third quartiles (the 25th and 75th percentiles), while internal hinge is the 50th percentile (median). Data beyond the end of the whiskers are "outlying" points and are plotted individually.
B. infantis, a key infant gut symbiont, is missing in 90% of infants
We assessed the bifidobacteria composition at the strain level by investigating the presence of genes specifically involved in the consumption of human milk oligosaccharides (HMOs)26,27. We performed a nucleotide-level analysis of six gene clusters previously identified to be relatively unique to the genome of Bifidobacterium longum subsp. infantis (B. infantis)27. Notably, the 43 kb H1 gene cluster that encodes glycosyl hydrolases active on four key HMO glycosidic linkages, has so far only been identified in B. infantis26 and is broadly conserved among the subspecies8,28 while other HMO-related gene loci are less conserved and found among different bacteria beyond bifidobacteria27. To account for genomic variation within the subspecies, we used the detection of at least 90% of the genes in these six genomic loci (H1, H2, H3, H4, H5, and a urease gene cluster) to detect B. infantis among infant samples by metagenomic sequencing. Using this criterion, we detected B. infantis in only 10% (23 samples) of all 227 samples. However, of these samples, only 3% (6 samples) had a relative abundance of B. longum species (inclusive of B. infantis) greater than 40%, which collectively represented a mean abundance of 65.10% (± SD 19.74%). Among the remaining 17 samples (7% of samples) where B. infantis was detected, the overall B. longum species only achieved a mean relative abundance of 7.38% (± SD 6.95%).
To confirm our findings, we also applied a subspecies-specific qPCR assay (see “Methods”) to detect B. infantis among these samples. Using qPCR, we detected B. infantis in 11% of samples (n = 24), in strong agreement with our metagenomic detection approach (Supplementary Table 1). One sample not detected by our metagenomics approach, but positive by qPCR, demonstrated a low relative abundance of B. longum species (%) and was missing more than 33% and 42% of the genes in the H1 and urease gene clusters, respectively.
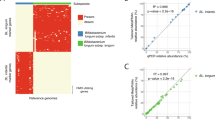
Thus, we used the detection of the H1 gene cluster as a proxy to determine the presence of B. infantis in the gut microbiome. All the genes belonging to the H1-cluster were detected in only 7 out of the 227 microbiome samples analyzed, meaning 97% of infants were likely missing B. infantis. Additionally, all 220 infants with incomplete or missing genes in the H1-cluster were also lacking crucial genes from other H-clusters non uniquely found in B. infantis, which are also involved in the capture, transport and metabolism of HMOs27 (Fig. 3). Furthermore, a comparison of the relative abundance of the genus Bifidobacterium on a per sample basis against the presence of the H-clusters showed that even with high abundance of Bifidobacterium (> 90%) in some of the infant microbiomes, key HMO genes were largely undetected (Fig. 3). This suggests that even when high levels of B. breve, B. longum, or B. bifidum are found in infants, it does not translate into a complete functional capacity of HMO utilization. Together, these analyses revealed that genes crucial to the utilization of HMOs, and therefore the main species that encodes them (i.e., B. infantis), are largely absent from the microbiome of infants in the US, independent of age, location, and diet (Fig. 3).
Functional HMO-cluster gene profiling by samples. (A) Heatmap showing presence/absence of the 56 HMO-cluster genes in all samples in the survey. Every column represents an individual sample and ordered by age block. Within a age block samples are ordered by decreasing presence of genes from left to right. Bottom of heatmap shows relative abundance (%) of Bifidobacterium by corresponding individual samples.
Antibiotic resistance in the US infant microbiome
Analysis of the total load of antibiotic resistance genes (ARGs) in the infants’ microbiomes across the US (i.e., the resistome), revealed a high abundance of ARGs in all five states analyzed in this survey, with a total of 325 unique ARGs identified from the CARD database (Supplementary Table 3). Particularly, South Carolina, California and Georgia had the highest abundance of ARGs (0.10%/0.014%/0.013%, average ARGs relative abundance compared to overall microbiome), followed by Pennsylvania (0.01%) and Oregon (0.006%) (Fig. 4A). No significant differences were observed by state in terms of overall ARG % microbiome composition. A total of 155 ARGs were common to the five states and despite the total abundance levels, more unique ARGs were identified in California (n = 20) (Fig. 4B). After accounting for sequencing depth, we confirmed that both the type and load of ARGs identified were not related to the number of sequences per sample grouped by state (Oregon = 32.5 M; Georgia = 28.9 M; California = 28.1 M; Pennsylvania = 27.5 M; South Carolina = 26.2 M), but rather possibly related to the number of individual patient samples collected (Fig. 4B). However, there appears to be a virtual plateau in ARG diversity assessed in this study considering that California had the highest diversity with almost half the patient samples compared to Georgia. A complete list of the identified ARGs by state is reported in Supplementary Table 4. Interestingly, the 20 unique ARGs found in California in this study have been also previously found in another cohort of 60 healthy, exclusively breastfed infants lacking B. infantis at day 21 of life29. Amongst the core ARGs found in every state, the most abundant on average were “ARO_3003369” (Escherichia coli EF-Tu mutants conferring resistance to Pulvomycin); followed by “ARO:3003317” (Salmonella serovars parE conferring resistance to fluoroquinolones); and “ARO:3003890” (Escherichia coli UhpT with mutation conferring resistance to fosfomycin). When considering age, there was a higher load of ARGs in the microbiome of infants between 0 and 3 months, though due to the high degree of intra-individual variation, not significantly different from the 4–6 months age window (P > 0.05) (Supplementary Fig. 2). We analyzed the ARG abundance on both the gene type and antimicrobial class levels using principal coordinate analyses (PCoA). With regard to the sample resistome dissimilarities at both genes and drug class levels, there was no clear separation when considering all the metadata collected, though geography and age were significant, but these factors had a very low effect size (adonis) on the overall resistome (Supplementary Table 5). Collapsing the ARGs into their respective drug class and by state, we found that on average 54% of the resistome was associated with genes conferring multi drug resistance, followed by fosfomycin, elfamycin and fluoroquinolone antibiotics resistance (Supplementary Fig. 3).
Antibiotic resistance genes characterization by state. (A) Relative abundance (%) of antibiotic resistance genes (ARGs) compared to overall microbiome composition identified by state. (B) Venn diagram showing shared ARG types by state. 155 ARGs types were core to all states (center of diagram), with unique ARG types identified in single state reported at the edge of the diagram. N’s represent number of patient samples by state.
Enterotypes of the US infant gut microbiome and functional analysis of HMO-metabolism potential
Three enterotype clusters were identified; 99 subjects were assigned to Bifidobacteriaceae, 81 to Enterobacteriaceae, and 47 to Bacteroidaceae enterotype (Fig. 5A). Other bacterial taxa identified contributed less significantly to each enterotype cluster compared to the three main bacterial families highlighted (Fig. 5B). Relative abundances of the three bacterial taxa that are principally responsible for the separation of samples confirmed expected predominance of Bifidobacteriaceae, Enterobacteriaceae and Bacteroidaceae, respectively (Supplementary Fig. 4). We confirmed the enterotype classification results using other well-known clustering approaches, including principal component analysis (PAM) as well as Bray–Curtis, which are commonly used to compare microbiome compositions across a given variable; here, our enterotype cluster assignments. We used the permutational multivariate analysis of variance (adonis) to test whether the clustering by our Enterotype classification was supported. The PAM clustering reported a P = 0.001; R2 = 0.17, while Bray–Curtis reported a P = 0.001; R2 = 0.32, confirming that the Enterotypes are indeed different in terms of the composition and represent distinct compositional states. Furthermore, absolute abundances as measured via quantitative PCR targeting the Bifidobacterium genus, also confirmed a significantly higher abundance of Bifidobacterium within the Bifidobacteriaceae-driven enterotype compared to the Enterobacteriaceae (P < 0.0001, Holm-adjusted Dunn’s) or Bacteroidaceae (P < 0.0001, Holm-adjusted Dunn’s). Subjects in each enterotype did not significantly differ according to diet, location, or sex. However, when considering age, there was a significant difference between the Bifidobacteriaceae and Enterobacteriaceae enterotypes (P < 0.0001, Bonferroni).
Enterotype grouping, Bifidobacterium abundance and functional gene annotation. (A) Enterotype analysis of samples at the family level. Samples clustered in three different enterotypes reported by different colors. In the center of the cluster is shown the main bacterial family responsible for the enterotype formation. (B) Bacterial families driving enterotype formation. Longer arrows indicate a stronger contribution. (C) Quantification of total Bifidobacterium spp. by enterotype. (D) Abundance of ARGs in all samples grouped by enterotype. (E) Heatmap showing relative abundance and functional annotation of KEGG orthologous genes related to HMO-utilization across enterotypes.
Analysis of the resistome composition measured by the total amount of ARGs identified amongst enterotypes, showed a significant difference in reductions of the total abundance of ARGs in the samples clustering within the Bifidobacteriaceae enterotype compared to Enterobacteriaceae (P = 0.0004; Bonferroni) and Bacteroidaceae (P = 0.006; Bonferroni) enterotypes. No significant differences in the abundance of ARGs were observed between the Enterobacteriaceae and Bacteroidaceae enterotypes (P = 0.3; Bonferroni).
To determine if the enterotype classification was related to the HMO-metabolism potential in the microbiomes at the functional level, we determined the abundance of ortholog genes (KOs) involved in the uptake, degradation and metabolism of HMO in each of the enterotypes. We found that, on average, microbiomes belonging to the Bifidobacteriaceae enterotype had a significantly increased capacity for utilization of HMO (119 CPM, mean) compared to the Bacteroidaceae (43 CPM, mean; P < 0.0001; Holm-adjusted Dunn’s) and Enterobacteriaceae enterotypes (47 CPM, mean; P < 0.0001; Holm-adjusted Dunn’s) enterotypes. The HMO-metabolism potential was also significantly different between Bacteroidaceae and Enterobacteriaceae enterotypes (P = 0.04; Holm-adjusted Dunn’s). When grouping KOs by function, on average, those related to the uptake and degradation of sialylated HMOs were significantly more abundant in the Bifidobacteriaceae (69.7 CMP, mean) and the Bacteroidaceae (51.4 CMP, mean) enterotype compared to the Enterobacteriaceae (1.23 CMP, mean) enterotypes (P < 0.0001 and P < 0.0001, respectively), but not significantly different between the Bifidobacteriaceae and Bacteroidaceae enterotypes (P = 0.32; Wilcoxon-FDR). Functions related to lacto-N-biose metabolism were significantly different in all three enterotypes with the Bifidobacteriacea enteroype encoding the most functions (50.3 CMP, mean) and the Enterobacteriaceae enterotype encoding the least (0.62 CMP, mean). The ability to degrade fucosylated HMO was also differentially significant between enterotypes but was higher in the Bacteroidaceae enterotypes (159.9 CMP, mean), followed by the Bifidobacteriacea (97.7 CMP, mean) and the Enterobacteriaceae (13.8 CMP, mean). Orthologous genes encoding functions related to the transport and catabolism of mono and polysaccharides were not significantly different between the Bacteroidaceae (49.5 CPM, mean) and Enterobacteriaceae (52.0 CPM, mean) enterotypes (P = 0.48; Wilcoxon-FDR), but were significantly higher in the Bifidobacteriaceae enteroype (162 CPM, mean) compared to the Bacteroidaceae (P < 0.0001; Wilcoxon-FDR) and the Enterobacteriaceae (P < 0.0001; Wilcoxon-FDR) enterotypes. Urea-related KOs were not statistically different between Bifidobacteriaceae (26.1 CPM, mean) and Enterobacteriaceae (28.6 CPM, mean) enterotypes (P = 0.84; Wilcoxon-FDR), but were significantly lower in the Bacteroidaceae (13.5 CPM, mean) compared to the Bifidobacteriaceae (P < 0.0001; Wilcoxon-FDR) and the Enterobacteriaceae (P < 0.0001; Wilcoxon-FDR) enterotypes (Fig. 5). Finally, we determined the number of samples that had a complete set of HMO-utilization orthologous genes. We found that only 16 out of 227 had the complete set of these KO functions. Amongst them, more than half (n = 9) where classified in the Bifidobacteriaceae enterotype, while 5 were in Bacteroidaceae and 2 in the Enterobacteriaceae enterotype. By location, 5 infants were in California, 4 in Oregon, and 7 in South Carolina. By age, 5 infants were in the 4–6-month age block and the rest were 3 months or less. Individual family microbiome composition of the top bacterial families (≥ 1% relative abundance, on average), and sorted by enterotype classification, is also reported in Supplementary Fig. 5.
Discussion
In the past few years, reliable evidence has emerged on the status of the US infant gut microbiome showing a general trend toward dysbiosis and associated negative acute- and long-term health consequences23,30,31,32. However, to date, researchers have derived conclusions mainly from single-site microbiome studies, which are often limited to a geographical area where samples were collected, or small association studies that carry inherent limits in terms of reproducibility of methodologies from sample collection to analysis. Furthermore, most infant microbiome studies in the US have focused on preterm infants, who are a very distinct population, experiencing epithelial gut immaturity and with a high degree of microbiome instability and a more severe dysbiosis compared to what is observed in term infants31,33.
Here, we aimed to broadly sample infants in the US across multiple geographies, in order to establish an understanding of the current status of the infant gut microbiome. Even though only five states were analyzed, our results suggest strong similarities across the research site regions, which appears to be independent of location, age and feeding modality (Fig. 1). Predominantly, 10 bacterial families accounted for 93% of the overall microbiome, while 40 families accounted for the remaining 7% on average. Amongst the 10 most abundant families, the majority were composed of potentially pathogenic species. Of the most abundant, the highest overall was E. coli (12.6% mean abundance), followed by Klebsiella pneumoniae (7% mean abundance). While the pathogenicity of these taxa is not universal34,35, in infants, the overrepresentation of γ-Proteobacteria, which includes the genera Escherichia and Klebsiella, are implicated in mediating pro-inflammatory responses associated with negative pediatric health outcomes23,36,37.
Several other bacterial species implicated with negative pediatric health outcomes, including sepsis and necrotizing enterocolitis (NEC) were identified38,39. For example, we detected relatively high levels of Klebsiella oxytoca (2.5% mean abundance) and Enterobacter cloaceae (2.7% mean abundance), both well described species related to NEC and Streptococcus agalactiae (Group B Strep; 0.01% mean abundance), which remains the primary cause of neonatal sepsis40,41. Infant microbiomes from across the US, irrespective of location, diet, or age could be classified into three distinct enterotypes, driven by abundances of Bifidobacteriaceae (43.6% mean abundance, n = 99), Enterobacteriaceae (35.7% mean abundance, n = 81), or Bacteroidaceae (20.7% mean abundance, n = 47). Infants classified as Enterobacteriaceae enterotype were typified by the highest relative abundance of Enterobacteriaceae, and the highest abundance of antibiotic resistance genes (ARGs) (Fig. 5D). We detected a high amount of ARGs in a small number of species, with more than half of the genes associated with multi-drug resistance (Supplementary Fig. 3). Infants classified within the Bifidobacteriaceae enterotype had a lower abundance of pathogenic bacteria and ARGs compared to the infants in the other two enterotypes. These results are in accordance with previous reports showing microbiomes rich in Bifidobacterium carry fewer ARGs29,42 and have a lower abundance of Enterobacteriaceae. The antagonistic effects exerted by bifidobacteria against Enterobacteriaceae, which are known to carry a wide array of ARGs, may contribute to this association43,44,45. Interestingly, we did find specific ARG signatures by state. Particularly, 20 ARGs were found in this study to be uniquely present in Southern California and were the same genes previously reported in a Northern California infant microbiome study46. This suggests that there might be distinct resistomes by different geographical area, which may coincide with regional disparities in antibiotic use and stewardship47 and may warrant further investigation.
The delivery of ecosystem services in the infant gut such as impeding colonization of infectious bacteria are dependent on the efficient utilization of human milk oligosaccharides (HMOs) for the production of organic acids8,44. As HMO consumption is pivotal to the establishment of a bifidobacteria-rich microbiota in breastfed infants, the abundance of genes related to the uptake of HMO correlates strongly with the abundance of Bifidobacterium in infants from different nations, including the US48. Therefore, we screened our data for the presence of HMO-related orthologous genes known to be fundamental for the complete metabolism of HMOs27. The analysis of such orthologous (HMO-KO) by enterotype revealed that there was a significantly higher presence of these crucial genes in the Bifidobacteriaceae-driven enterotype compared to the other enterotypes. However, we found that although 43.6% of infants (n = 99) were classified as having a Bifidobacteriaceae-driven enterotype, the microbiomes of only a small subset (n = 16) of infants carried all the KO-HMO functions. Notably, HMO assimilation abilities significantly vary among different species and strains of Bifidobacterium48,49. Therefore, it is likely that species within the Bifidobacteriaceae-enterotypes were limited in the ability to access HMOs. Some Bifidobacterium (e.g., B. bifidum50) and Bacteroides possess enzymes for the purpose of degrading mucin glycans that hold structural similarities with HMO51,52. This may explain why functions related to sialic acid, which decorates both mucin and HMOs, were not differentially abundant between the Bifidobacteriaceae and Bacteroidaceae enterotypes. This idea is further supported by the fact that only 10% of the samples (n = 23) showed detectable B. infantis, which is the only known species fully equipped and ecologically adept at capturing, transporting and metabolizing HMOs26. Further, only a small number of these samples where B. infantis was detected (3% of total samples, n = 6) exhibited a microbiome composition characterized by a high abundance of B. longum species (inclusive of B. infantis) demonstrative of the efficient provision of ecosystem services8.
The absence or substantial decrease of B. infantis during the first trimester postnatal has been associated with (1) higher susceptibility to invasion of allochthonous bacteria with pathogenic potential31,44; (2) higher levels of ARGs29 and virulence factors53; (3) increased enteric inflammation23; (4) progressive mucus erosion compromising the intestinal mucosal barrier 54; (5) limited conversion of HMOs into organic acids31; (6) limited HMO-derived nutritional absorption8; and (7) altered response to vaccines21. Together, data generated in this study demonstrate that B. infantis, which is considered a key infant gut symbiont that has co-evolved with the host to provide important ecosystem services, is exceptionally rare among infants in the US8,26,27,55.
Our study carries the limitations of a cross-sectional study and infants analyzed were mostly located in urban and suburban areas and may not be representative of rural populations56,57. Also, the use of medications (e.g. antibiotics) and delivery mode are known to alter the microbiome58,59, but the impact of these interventions was not examined in the context of this study. However, we note that the aim of this study was to gain insight into infant gut microbiome compositions as they exist among the population and characterize the functional capacities of these communities, rather than to identify the relative impact of medically necessary interventions on the gut microbiome of infants. Further, associations between these interventions and the gut microbiome may be context-dependent, and connections between cesarean section delivery on the infant gut microbiome in the US do not match observations in Bangladesh11,20.
In conclusion, our survey highlights the microbiome composition and its metabolic capacity among infants in the US. Given recent findings linking the microbiome in early life to key elements of infant health and the understanding of this community has improved, our findings reveal that infants in the US have microbiomes that may fail to provide functions necessary in early life including shaping the immune system, protecting against pathogen colonization, and maximizing nutrition from breastmilk (e.g. HMOs). These diminished ecosystem services suggest that these communities could thus be classified as dysbiotic8. Future efforts should be directed to determine the cause and extent of these results and confirm relevance more broadly and in specific populations of concern. No matter the cause, these findings demonstrate a need to develop interventions that address the dysbiosis reported here, considering microbiome functional deficiencies and restoring the necessary ecosystem services to infants.
Methods
Sample collection
The study was conducted at five pediatrician offices and two breastfeeding support centers across five geographically diverse states (California, Georgia, Oregon, Pennsylvania and South Carolina). The experimental protocol was approved by Advarra Central IRB (IRB Registration number: 00000971). All methods were carried out in accordance with relevant guidelines and regulations set forth by the Declaration of Helsinki. Informed consent was obtained from a parent or legal guardian of all participants in the study. Generally healthy aged 0–6 months infants were enrolled. Participants were excluded if they presented with jaundice at the time of enrollment, were undergoing current antibiotic treatment, or were diagnosed with carbohydrate malabsorption syndrome. Site personnel processed fresh diapers within eight hours of receipt. If they could not be processed within 8 h, diapers were frozen at − 20 °C for up to 72 h and thawed for 1 h prior to processing. Site personnel collected stool swabs from each diaper and placed them in a barcoded collection tube containing DNA/RNA Shield stabilization buffer (Zymo Research, Irvine CA) and lysis beads. Stool swab samples in stabilization buffer were stored at room temperature. Additionally, staffs were instructed to aliquot at least a bean-sized amount of stool into an empty, barcoded specimen tube. This aliquot was stored at − 20 °C prior to shipment. All samples were shipped on dry ice to Evolve BioSystems, Inc. Upon receipt, all samples were stored at − 80 °C until further processing. 229 stool samples were collected during the enrollment period but only 227 passed final quality filtering.
DNA extraction and metagenomics sequencing
gDNA was extracted from 229 stool swab samples stored in DNA/RNA shield lysis tubes (Zymo Research, Irvine CA) using the ZymoBIOMICS 96 MagBead DNA kit (Zymo Research, Irvine CA). Extracted DNA was quantified using QuantIT dsDNA Assay kit, high sensitivity (ThermoFisher Scientific, Waltham, MA) according to the manufacturer’s protocol. 2 Samples did not meet library preparation input specifications and were omitted from downstream analysis. Libraries were prepared for each sample using the Illumina Nextera DNA Flex library kit (Illumina, San Diego, CA) with unique dual indexes according to manufacturer guidelines. Libraries were pooled and submitted to UC Davis DNA Technologies core for sequencing on the Illumina NovaSeq S4 flow cell. (Illumina, San Diego, CA). Each lane of the S4 flow cell contained 96 libraries.
Total Bifidobacterium was quantified using a standard curve Taqman-based quantitative PCR assay as previously described using the Bif F, Bif R and Bif P primer/probe set31,60. The primer final concentrations were adjusted to 900 nM. Quantification of the total B. infantis was performed by quantitative real-time PCR using Blon_2348 sialidase gene primers Inf2348F (5′-ATA CAG CAG AAC CTT GGC CT-3′), Inf2348_R (5′-GCG ATC ACA TGG ACG AGA AC-3′), and Inf2348_P (5′-/56-FAM/TTT CAC GGA /ZEN/TCA CCG GAC CAT ACG/3lABkFQ/-3′)61. Each reaction contained 10 μL of 2 × TaqMan Universal Master Mix II with UNG master mix (ThermoFisher Scientific, Waltham, MA), 0.9 µM of each primer, 0.25 µM probe and 5 μL of template DNA. Thermal cycling was performed on a QuantStudio 3 Real-Time PCR System and consisted of an initial UNG activation step of 2 min at 50 °C followed by a 10 min denaturation at 95 °C, succeeded by 40 cycles of 15 s at 95 °C and 1 min at 60 °C. Standard curves for absolute quantification were generated using genomic DNA extracted from a pure culture of B. infantis EVC001.
Quality filtering and removal of human sequences
Quality filtering and removal of human sequences were performed following a previously described pipeline29. Demultiplexed Fastq files from NovaSeq were quality filtered using Trimmomatic v0.3662 with default parameters. Quality-filtered sequences were screened to remove human sequences using GenCoF v1.063 against a non-redundant version of the Genome Reference Consortium Human Build 38, patch release 7 (GRCh38_p7; www.ncbi.nlm.nih.gov). Human sequence-filtered raw reads were deposited in the Sequence Read Archive (SRA; https://www.ncbi.nlm.nih.gov/sra) under the reference number PRJNA633576.
Taxonomic and strain profiling
Taxonomic and strain profiling was performed as previously described29. Taxonomic profiling of the metagenomic samples was performed using MetaPhlAn264, which uses a library of clade-specific markers to provide pan microbial (bacterial, archaeal, viral, and eukaryotic) profiling (http://hutten-hower.sph.harvard.edu/metaphlan2), in combination with HUMAnN265 to profile functional metagenomics against Uniref90 following the updated global profiling of the Human Microbiome Project (2017)66. Strain-specific gene markers were used to determine the presence of B. longum subs. infantis as previously described29. Cross database annotations (e.g., UniProt to KEGG) were performed within Humman2 using the “utility_mapping” conversion tool package.
Antibiotic resistance gene analysis
Antibiotic resistance gene analysis was computed as previously described29. We applied ShortBRED67,68 to profile antibiotic resistance (AR) abundance and composition in the infant gut microbiome. We first produced a set of new AR marker sequences by applying ShortBRED-Identify to the Comprehensive Antibiotic Resistance Database database (CARD)69. We then used ShortBRED-Quantify to profile the relative abundance of corresponding antibiotic resistance genes (ARGs). Final values were normalized in Reads Per Kilobase per Million mapped reads (RPKM) to account for sequencing depth as well as gene length. We used custom scripts to collapse CARD individual antibiotic resistance gene entries in their corresponding drug class. Conversion rules are offered within every CARD package update.
Statistical analysis
All statistical analyses were performed in R v3.6.2. A Kruksal-Wallis one-way analysis of variance coupled with an FDR or Bonferroni correction was used for statistical comparisons between individual genes and taxa among groups. Statistical analysis to assess total resistome or enterotype composition by group was performed using a Mann–Whitney or Holm–adjusted Dunn's test. Rarefaction curves were computed to estimate the diversity of the identified ARGs across samples. A nonparametric two-sample t-test was used to compare rarefaction curves using Monte Carlo permutations (n = 999). The microbiome multivariable association analysis with linear models was perfomed using MaAsLin2 (https://huttenhower.sph.harvard.edu/maaslin/), by including all the clinical variables presented in the Supplementary Table 1 to test for microbiome differences at the family level according to metadata. All variables were run as fixed effects, except for the variable “State”, which was run as a random effect for the model (Supplementary Table 1). Enterotype analysis was performed as previously described70. Principal component analysis and Bray–Curtis beta diversity to confirm enterotype classification was performed as previously decribed29, and confirmed with a permutational multivariate analysis of variance (adonis) to test whether the clustering by our Enterotype classification was supported. The P-values throughout the manuscript are represented by asterisks (*P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001).
Change history
20 May 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41598-021-90391-4
References
Derrien, M., Alvarez, A.-S. & de Vos, W. M. The gut microbiota in the first decade of life. Trends Microbiol. 27, 997–1010 (2019).
Gollwitzer, E. S. & Marsland, B. J. Impact of early-life exposures on immune maturation and susceptibility to disease. Trends Immunol. 36, 684–696 (2015).
Olin, A. et al. Stereotypic immune system development in newborn children. Cell 174, 1277-1292.e1214. https://doi.org/10.1016/j.cell.2018.06.045 (2018).
Gensollen, T., Iyer, S. S., Kasper, D. L. & Blumberg, R. S. How colonization by microbiota in early life shapes the immune system. Science 352, 539–544 (2016).
Cox, L. M. et al. Altering the intestinal microbiota during a critical developmental window has lasting metabolic consequences. Cell 158, 705–721 (2014).
Prescott, S. L. Early-life environmental determinants of allergic diseases and the wider pandemic of inflammatory noncommunicable diseases. J. Allergy Clin. Immunol. 131, 23–30 (2013).
Stiemsma, L. T. & Turvey, S. E. Asthma and the microbiome: Defining the critical window in early life. Allergy Asthma Clin. Immunol. 13, 3 (2017).
Duar, R. M., Henrick, B. M., Casaburi, G. & Frese, S. A. Integrating the ecosystem services framework to define dysbiosis of the breastfed infant gut: the role of B. infantis and human milk oligosaccharides. Front. Nutr. 7, 33 (2020).
Tapiainen, T. et al. Impact of intrapartum and postnatal antibiotics on the gut microbiome and emergence of antimicrobial resistance in infants. Sci. Rep. 9, 1–11 (2019).
Lewis, Z. T. et al. Validating bifidobacterial species and subspecies identity in commercial probiotic products. Pediatr. Res. 79, 445 (2016).
Bokulich, N. A. et al. Antibiotics, birth mode, and diet shape microbiome maturation during early life. Sci. Transl. Med. 8, 343ra382-343ra382 (2016).
Davis, J. C. et al. Growth and morbidity of gambian infants are influenced by maternal milk oligosaccharides and infant gut microbiota. Sci. Rep. 7, 40466 (2017).
Vangay, P. et al. US immigration westernizes the human gut microbiome. Cell 175, 962–972 (2018).
Sonnenburg, E. D. et al. Diet-induced extinctions in the gut microbiota compound over generations. Nature 529, 212–215 (2016).
Blaser, M. J. The theory of disappearing microbiota and the epidemics of chronic diseases. Nat. Rev. Immunol. 17, 461 (2017).
Groer, M. et al. (Epub 2014/10/22. https://doi.org/10.1186/2049-2618-2-38 PMID: 25332768).
Gritz, E. C. & Bhandari, V. The human neonatal gut microbiome: A brief review. Front. Pediatr. 3, 17 (2015).
Stewart, C. J. et al. Temporal development of the gut microbiome in early childhood from the TEDDY study. Nature 562, 583–588 (2018).
Vatanen, T. et al. The human gut microbiome in early-onset type 1 diabetes from the TEDDY study. Nature 562, 589–594 (2018).
Huda, M. N. et al. Stool microbiota and vaccine responses of infants. Pediatrics 134, e362–e372 (2014).
Huda, M. N. et al. Bifidobacterium abundance in early infancy and vaccine response at 2 years of age. Pediatrics 143, e20181489 (2019).
Depner, M. et al. Maturation of the gut microbiome during the first year of life contributes to the protective farm effect on childhood asthma. Nat. Med. 26, 1766–1775 (2020).
Henrick, B. M. et al. Colonization by B. infantis EVC001 modulates enteric inflammation in exclusively breastfed infants. Pediatr. Res. 86, 749–757 (2019).
McKenney, E., Koelle, K., Dunn, R. & Yoder, A. The ecosystem services of animal microbiomes. Mol. Ecol. 27, 2164–2172 (2018).
Fujimura, K. E. et al. Neonatal gut microbiota associates with childhood multisensitized atopy and T cell differentiation. Nat. Med. 22, 1187 (2016).
Sela, D. et al. The genome sequence of Bifidobacterium longum subsp. infantis reveals adaptations for milk utilization within the infant microbiome. Proc. Natl. Acad. Sci. 105, 18964–18969 (2008).
LoCascio, R. G., Desai, P., Sela, D. A., Weimer, B. & Mills, D. A. Broad conservation of milk utilization genes in Bifidobacterium longum subsp. infantis as revealed by comparative genomic hybridization. Appl. Environ. Microbiol. 76, 7373–7381 (2010).
Albert, K., Rani, A. & Sela, D. A. Comparative pangenomics of the mammalian gut commensal Bifidobacterium longum. Microorganisms 8, 7 (2020).
Casaburi, G. et al. Early-life gut microbiome modulation reduces the abundance of antibiotic-resistant bacteria. Antimicrob. Resist. Infect. Control 8, 131 (2019).
Tanaka, M. & Nakayama, J. Development of the gut microbiota in infancy and its impact on health in later life. Allergol. Int. 66, 515–522 (2017).
Frese, S. A. et al. Persistence of supplemented Bifidobacterium longum subsp. infantisEVC001 in breastfed infants. mSphere https://doi.org/10.1128/mSphere.00501-17 (2017).
Kostic, A. D. et al. The dynamics of the human infant gut microbiome in development and in progression toward type 1 diabetes. Cell Host Microbe 17, 260–273 (2015).
Raju, T. N. et al. Long-term healthcare outcomes of preterm birth: an executive summary of a conference sponsored by the National Institutes of Health. J. Pediatr. 181, 309–318 (2017).
Lau, H. Y., Huffnagle, G. B. & Moore, T. A. Host and microbiota factors that control Klebsiella pneumoniae mucosal colonization in mice. Microbes Infect. 10, 1283–1290 (2008).
Katouli, M. Population structure of gut Escherichia coli and its role in development of extra-intestinal infections. Iran. J. Microbiol. 2, 59 (2010).
Pammi, M. et al. Intestinal dysbiosis in preterm infants preceding necrotizing enterocolitis: A systematic review and meta-analysis. Microbiome 5, 31 (2017).
Hackam, D. J. & Sodhi, C. P. Toll-like receptor-mediated intestinal inflammatory imbalance in the pathogenesis of necrotizing enterocolitis. Cell. Mol. Gastroenterol. Hepatol. 6, 229–238 (2018).
Coggins, S. A., Wynn, J. L. & Weitkamp, J.-H. Infectious causes of necrotizing enterocolitis. Clin. Perinatol. 42, 133–154 (2015).
van Acker, J. et al. Outbreak of necrotizing enterocolitis associated with Enterobacter sakazakii in powdered milk formula. J. Clin. Microbiol. 39, 293–297 (2001).
Puopolo, K. M., Baker, C. J. & Edwards, M. Group B streptococcal infection in neonates and young infants (Waltham, Massachusetts, 2013).
Hanna, M. & Noor, A. In StatPearls [Internet] (StatPearls Publishing, 2020).
Taft, D. H. et al. Bifidobacterial dominance of the gut in early life and acquisition of antimicrobial resistance. mSphere 3, e00441-e418 (2018).
Fukuda, S. et al. Bifidobacteria can protect from enteropathogenic infection through production of acetate. Nature 469, 543–547 (2011).
Duar, R. M., Kyle, D. & Casaburi, G. Colonization resistance in the infant gut: The role of B. Infantis in reducing pH and preventing pathogen growth. High-Throughput 9, 7 (2020).
Iredell, J., Brown, J. & Tagg, K. Antibiotic resistance in Enterobacteriaceae: Mechanisms and clinical implications. BMJ 352, h6420 (2016).
Casaburi, G. et al. Early-life gut microbiome modulation reduces the abundance of antibiotic-resistant bacteria. Antimicrob. Resist. Infect. Control 8, 1–18 (2019).
Llor, C. & Bjerrum, L. Antimicrobial resistance: Risk associated with antibiotic overuse and initiatives to reduce the problem. Ther Adv. Drug Saf. 5, 229–241 (2014).
Sakanaka, M. et al. Evolutionary adaptation in fucosyllactose uptake systems supports bifidobacteria-infant symbiosis. Sci. Adv. 5, eaaw7696 (2019).
Thomson, P., Medina, D. A. & Garrido, D. Human milk oligosaccharides and infant gut bifidobacteria: Molecular strategies for their utilization. Food Microbiol. 75, 37–46 (2018).
Turroni, F. et al. Genome analysis of Bifidobacterium bifidumPRL2010 reveals metabolic pathways for host-derived glycan foraging. Proc. Natl. Acad. Sci. 107, 19514–19519 (2010).
Charbonneau, M. R. et al. Sialylated milk oligosaccharides promote microbiota-dependent growth in models of infant undernutrition. Cell 164, 859–871 (2016).
Marcobal, A. et al. Bacteroides in the infant gut consume milk oligosaccharides via mucus-utilization pathways. Cell Host Microbe 10, 507–514 (2011).
Casaburi, G. & Frese, S. A. Colonization of breastfed infants by Bifidobacterium longum subsp. infantisEVC001 reduces virulence gene abundance. Hum. Microbiome J. 9, 7–10 (2018).
Karav, S., Casaburi, G. & Frese, S. A. Reduced colonic mucin degradation in breastfed infants colonized by Bifidobacterium longum subsp. infantisEVC001. FEBS Open Bio 8, 1649–1657 (2018).
Underwood, M. A., German, J. B., Lebrilla, C. B. & Mills, D. A. Bifidobacterium longum subspecies infantis: Champion colonizer of the infant gut. Pediatr. Res. 77, 229 (2015).
Martínez, I. et al. The gut microbiota of rural papua new guineans: Composition, diversity patterns, and ecological processes. Cell Rep. 11, 527–538 (2015).
Yatsunenko, T. et al. Human gut microbiome viewed across age and geography. Nature 486, 222–227 (2012).
Salminen, S., Gibson, G. R., McCartney, A. L. & Isolauri, E. Influence of mode of delivery on gut microbiota composition in seven year old children. Gut 53, 1388–1389 (2004).
Vangay, P., Ward, T., Gerber, J. S. & Knights, D. Antibiotics, pediatricdysbiosis, and disease. Cell Host Microbe 17, 553–564 (2015).
Penders, J. et al. Quantification of Bifidobacterium spp., Escherichia coli and Clostridium difficile in faecal samples of breast-fed and formula-fed infants by real-time PCR. FEMS Microbiol. Lett. 243, 141–147 (2005).
Lawley, B. et al. Differentiation of Bifidobacterium longum subspecies longum and infantis by quantitative PCR using functional gene targets. PeerJ 5, e3375 (2017).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Czajkowski, M. D., Vance, D. P., Frese, S. A. & Casaburi, G. GenCoF: A graphical user interface to rapidly remove human genome contaminants from metagenomic datasets. Bioinformatics 35, 2318–2319 (2018).
Truong, D. T. et al. MetaPhlAn2 for enhanced metagenomic taxonomic profiling. Nat. Methods 12, 902 (2015).
Franzosa, E. A. et al. Species-level functional profiling of metagenomes and metatranscriptomes. Nat. Methods 15, 962 (2018).
Lloyd-Price, J. et al. Strains, functions and dynamics in the expanded Human Microbiome Project. Nature (2017).
Jakobsson, H. E. et al. Short-term antibiotic treatment has differing long-term impacts on the human throat and gut microbiome. PLoS ONE 5, e9836 (2010).
Kaminski, J. et al. High-specificity targeted functional profiling in microbial communities with ShortBRED. PLoS Comput. Biol. 11, e1004557 (2015).
Alcock, B. P. et al. CARD 2020: Antibiotic resistome surveillance with the comprehensive antibiotic resistance database. Nucleic Acids Res. 48, D517–D525 (2020).
Arumugam, M. et al. Enterotypes of the human gut microbiome. Nature 473, 174–180 (2011).
Acknowledgements
Sequencing in this study was carried by the DNA Technologies and Expression Analysis Cores at the UC Davis Genome Center, supported by NIH Shared Instrumentation Grant 1S10OD010786-01.
Funding
This work was supported by Evolve BioSystems, Inc.
Author information
Authors and Affiliations
Contributions
G.C., R.M.D., H.B., S.K., R.D.M., S.C. and S.A.F. processed and analyzed the data. O.C., R.L.F., B.M.H. and S.L.F. organized the sample collection. All authors contributed to writing and editing and final approval of the manuscript.
Corresponding author
Ethics declarations
Competing interests
KGS is a professor at Stanford University. GC, RMD, HB, RDM, SK, SC, OC, RLF, SAF, BMH and SLF are employees of Evolve BioSystems, a company focused on restoring the infant microbiome. We are committed to making our raw data, materials, and analysis methods publicly available.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Casaburi, G., Duar, R.M., Brown, H. et al. Metagenomic insights of the infant microbiome community structure and function across multiple sites in the United States. Sci Rep 11, 1472 (2021). https://doi.org/10.1038/s41598-020-80583-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-80583-9
This article is cited by
-
CRISPR-Cas-based identification of a sialylated human milk oligosaccharides utilization cluster in the infant gut commensal Bacteroides dorei
Nature Communications (2024)
-
Primary succession of Bifidobacteria drives pathogen resistance in neonatal microbiota assembly
Nature Microbiology (2024)
-
Faecal microbiota and cytokine profiles of rural Cambodian infants linked to diet and diarrhoeal episodes
npj Biofilms and Microbiomes (2024)
-
Longitudinal quantification of Bifidobacterium longum subsp. infantis reveals late colonization in the infant gut independent of maternal milk HMO composition
Nature Communications (2024)
-
Human milk oligosaccharides modify the strength of priority effects in the Bifidobacterium community assembly during infancy
The ISME Journal (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.