Abstract
Maize is China’s largest grain crop. Mechanical grain harvesting is the key technology in maize production, and the kernel moisture concentration (KMC) is the main controlling factor in mechanical maize harvesting in China. The kernel dehydration rate (KDR) is closely related to the KMC. Thus, it is important to conduct genome-wide association studies (GWAS) of the KMC and KDR in maize, detect relevant quantitative trait nucleotides (QTNs), and mine relevant candidate genes. Here, 132 maize inbred lines were used to measure the KMC every 5 days from 10 to 40 days after pollination (DAP) in order to calculate the KDR. These lines were genotyped using a maize 55K single-nucleotide polymorphism array. QTNs for the KMC and KDR were detected based on five methods (mrMLM, FASTmrMLM, FASTmrEMMA, pLARmEB, and ISIS EM-BLASSO) in the package mrMLM. A total of 334 significant QTNs were found for both the KMC and KDR, including 175 QTNs unique to the KMC and 178 QTNs unique to the KDR; 116 and 58 QTNs were detected among the 334 QTNs by two and more than two methods, respectively; and 9 and 5 QTNs among 58 QTNs were detected in 2 and 3 years, respectively. A significant enrichment in cellular component was revealed by Gene Ontology enrichment analysis of candidate genes in the intervals adjacent to the 14 QTNs and this category contained five genes. The information provided in this study may be useful for further mining of genes associated with the KMC and KDR in maize.
Similar content being viewed by others
Introduction
Maize (Zea mays L.) is the largest grain crop in China. The planting area for maize and maize production were 41.3 million ha and 2.6 billion tons in 2019 in China, respectively1. Mechanical grain harvesting is the key technology in maize production, and the kernel moisture concentration (KMC) is the main controlling factor in mechanical maize harvesting in China2. A low KMC at harvest may facilitate mechanical harvesting, shelling efficiency, and grain quality and reduce additional drying cost and shrinkage penalties3,4,5. When the KMC at harvest is greater than 25 percent, the breakage rate quickly increases, which significantly reduces farmers' incomes6. The kernel dehydration rate (KDR) is defined as the rate of moisture loss between two adjacent periods after pollination, which is closely related to the KMC. It is the key measurements to select maize hybrids with a low KMC and a high KDR at mature period for achieving mechanical maize harvesting.
Changes in the KMC and KDR occur in two distinct phases7,8. The first phase spans the time from pollination to physiological maturity (PM) and is defined as physiological dehydration. The second phase spans the time from PM to harvest and is defined as a natural drying process. During the first phase, kernel dehydration is an internal process under the control of growth and development regulatory processes. Environmental factors have no significant effects on such dehydration9. Beginning as early as the 1960s, the KMC and KDR have been controlled by multi-genes and could be stably inherited10,11,12. Previous research has also shown that selection based on a low KMC and a high KDR before PM was an effective strategy to achieve a low KMC at harvest13,14,15. Therefore, measuring the KMC and KDR in various maize cultivars before PM and conducting quantitative trait locus (QTL) mapping, genome-wide association studies (GWAS), and candidate gene mining for these two traits in maize are crucial tasks.
Extensive research has been conducted to map QTLs for the KMC and KDR in maize3,4,16,17,18,19. Some intervals or genes associated with the KDR in maize have been obtained3,4, but only Dai et al.20 and Zhang et al.21 have conducted GWAS for these two traits in maize.
The most popular method for GWAS is the mixed linear model (MLM)22,23. In the past decade, several MLM algorithms, such as compressed MLM24 and enriched compressed MLM25, were developed to improve the computational efficiency. However, all these models perform one-dimensional genome scans and require multiple corrections. Generally, the above traditional methods had significant limitations in mapping QTLs with relatively small effects. Therefore, Wang et al.26 proposed a new model. Then, GWAS methods based on a multi-locus random-SNP-effect MLM (mrMLM) were proposed. The methods included iterative modified-sure independence screening expectation–maximization (EM)-Bayesian LASSO (ISIS EM-BLASSO), polygenic-background-control-based least angle regression plus empirical Bayes (pLARmEB), fast multi-locus random-SNP-effect efficient mixed model association (FASTmrEMMA), and fast multi-locus random-SNP-effect mixed linear model (FASTmrMLM)26,27,28,29,30. These methods could effectively detect small-effect quantitative trait nucleotides (QTNs) and improve the efficiency and accuracy of GWAS.
In this study, the mrMLM was used to perform a GWAS based on Axiom Maize 55K SNP Array data from 132 maize inbred lines. QTNs associated with the KMC and KDR were detected using five methods in the package mrMLM (mrMLM, FASTmrMLM, FASTmrEMMA, pLARmEB, and ISIS EM-BLASSO). Associated candidate genes were predicted and submitted to Gene Ontology (GO) enrichment analysis. The results have theoretical significance and application value for improving maize kernel dehydration characteristics using marker-assisted selection (MAS).
Results
Phenotypic data analysis of 132 inbred lines
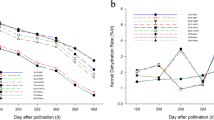
The trends of both the KMC and KDR were very similar in 2014, 2015 and 2016 (Fig. 1). The KMC gradually decreased over time, while the KDR gradually increased over time (Table 1). The different phenotypic traits also showed different levels of variation. The KMC had relatively low coefficients of variation (CV), and the lowest CVs were found at 15 days after pollination (DAP), with values of 1.34%, 1.48%, and 1.44% in 2014, 2015, and 2016, respectively. CVs for the KMC gradually increased over time, with values of 12.68%, 13.56% and 13.71% in 2014, 2015, and 2016 at 40 DAP, respectively. By contrast, the KDR had relatively high CV, with the maximum values found at 35 DAP, 42.09%, 39.4% and 37.06% in 2014, 2015, and 2016, respectively. This result suggests that the optimal time for maize KDR measurement may be after 35 DAP. The heritability ranged from 68.54 to 76.42% for the KMC, and ranged from 61.75 to 74.16% for the KDR. The highest heritability of both traits appeared at 40 DAP. The results from analysis of variance for maize KMC and KDR revealed highly significant differences across cultivars and across periods, except KDR15.
Population structure and linkage disequilibrium (LD) analysis
The Axiom Maize 55K SNP Array (CapitalBio Corp., Beijing, China) had 55,229 SNPs31, and the 132 inbred lines were genotyped based on the Affy AXIOM Array 2.0 platform. The raw genotyping data were filtered using PLINK based on a minor allele frequency (MAF) > 0.05 and a missing genotype rate (GENO) < 0.1. After filtering, the remaining 41,357 SNPs were used for LD analysis and GWAS.
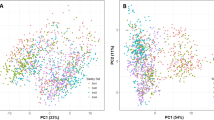
When r2 = 0.1, the attenuation distance of LD was approximately 200 kb (Fig. 2A), and genes within 200 kb upstream or downstream of the marker exceeding the threshold were taken as candidates. The number of subpopulations (k) was plotted based on the delta k calculated using STRUCTURE software (Fig. 2B). The line chart showed a peak at k = 6, indicating that the natural population could be divided into six subpopulations (Fig. 2C).
The quality and accuracy of the SNP markers and lines
Measurement of the KDR is relatively difficult, and natural populations have rarely been used in GWAS. In this study, we conducted the GWAS using cob colour, a qualitative trait in maize, to verify the validity and accuracy of the experimental populations and markers. Pericarp colour 1 (P1) regulates red pigmentation in cob, pericarp, tassel glumes, and husks, and its gene is located at mk187 on chromosome (chr.) 1 at position 48.1 Mb. A total of seven significant QTNs were detected on chr.1 using the package mrMLM (Table 2). Among them, AX-86284808 and AX-91425354 were detected by two and four methods, respectively, which explained up to 32.12% of the phenotypic variation. When r2 = 0.1 and the attenuation distance of LD was approximately 200 kb, AX-86284808 and AX-91425354 were mapped onto B73 Ref Gen_V4, and both QTNs were internally scanned to Zm00001d028850 (P1), which is consistent with the results of a previous study screening genes involved in cob colour32. These results suggest that we should identify candidate genes within the attenuation distance of LD around the QTNs shared in common by multiple detection methods to achieve accurate candidate gene mining (Table 2).
QTNs associated with the KMC and KDR
A total of 334 significant QTNs were identified for the KMC and KDR in maize, using five multi-locus GWAS methods in the package mrMLM, with a critical LOD (logarithm of odds) score of 3 (Supplementary Table S1, Fig. 3). A total of 175 significant QTNs were found for the KMC, including 28, 35, 23, 34, 35, 23, and 30 significant QTNs associated with KMC10, KMC15, KMC20, KMC25, KMC30, KMC35, and KMC40, respectively, in which their proportions of phenotypic variation explained (PVE) by each QTN ranged from 1.22 to 25.34%. Of these QTNs, 41 and 21 QTNs were detected in 2 and 3 years, respectively. For the KDR, 178 significant QTNs were detected, including 30 28, 37, 39, 22, and 23 significant QTNs associated with KDR15, KDR20, KDR25, KDR30, KDR35, and KDR40, respectively, in which their PVEs ranged from 0.54 to 24.62%. Among these QTNs, 19 and 5 were detected in 2 and 3 years, respectively. Moreover, 1, 3, 5, and 32 QTNs were found to be simultaneously associated with the above 5, 4, 3 and 2 traits, respectively.
Genome-wide distribution of SNPs throughout the 132 lines’ genomes. The outermost box with the scale represents the 10 maize chromosomes. The red histogram represents the density of SNPs. The yellow, red and blue scatters represent significant QTNs for the KMC and KDR traits in 2014, 2015 and 2016, respectively.
Common QTNs by multiple methods and years
Among the above-mentioned 334 QTNs, 116 and 58 were detected by two and more than two methods, respectively (Supplementary Tables S2, S3). Nine of the 58 QTNs were detected in 2 years, which were associated with KMC10, KMC20, KMC20, KMC30, KMC35, KMC40, KDR15, KDR30, and KDR40, respectively. Five of the 58 QTNs were detected in 3 years, which were associated with KMC10, KMC30, KMC35, KDR20 and KDR30, respectively. Among the 14 QTNs across multiple environments (Table 3), AX-91654966 on chr.5 was found by five methods in 2015 and 2016 to be associated with KDR40; AX-91629217 on chr.4 was found by four methods in 3 years to be associated with KMC30; and their PVEs ranged up to 19.76% and 14.50%, respectively.
Validation of fourteen common QTNs
Based on the alleles of 14 QTNs, the 132 inbred lines were divided into two groups, and we tested for significant differences in mean phenotypes between the two groups (Fig. 4). The results showed that the mean values of the groups with favourable alleles were larger than those of the other groups with alternative alleles, indicating significant or extremely significant differences.
GO enrichment analysis
According to the genomic information of B73 Ref Gen_V4, a total of 2,970 genes were present in the intervals adjacent to the 334 QTNs detected for the KMC and KDR in maize (Supplementary Tables S1, S4). A total of 142 genes were detected in the intervals adjacent to the 14 QTNs in two and 3 years. Significance was observed for cellular component in the GO enrichment analysis (P ≤ 0.05; Supplementary Table S5), and this category contained five genes (Supplementary Table S6). This information may be useful for further mining of genes associated with the KMC and KDR in maize.
Discussion
Measurements and physiological mechanisms of KMC and KDR
According to Hart and Golumbic33, methods to measure seed KMC can be divided into direct and indirect methods. With direct methods, the KMC in a seed sample is measured directly. With indirect methods, the KMC is measured according to the relationship between the chemical and physical characteristics of seed water and its KMC, with calibration to a reference. Direct methods have the advantage of high repeatability, but require a substantial workload in the field and are therefore unsuitable for multiple repeated measurements of many materials. In this study, we directly measured the KMC in maize following the experimental method of Reid et al.34. This method can not only reduce the workload in the field but also decrease experimental errors during shelling, thus saving time and labour.
The KMC in maize kernels is closely linked to their dry weight35. In the early stage of kernel development, water accumulates more than dry matter. As the dry weight increases, the KMC gradually decreases. Therefore, kernel dehydration before PM is an internal process controlled by growth and development, and environmental factors have no significant effects on dehydration in this process9.
Genotypic differences lead to variations in the KDR, and the KDR shows a marked variation between various cultivars before PM, which is still inheritable. This study showed that in the early stage of kernel development, the KMC decreased slowly, while the KDR was relatively low; and with maturation of the kernels, the KMC and KDR increased sharply after 35 DAP (the kernels were close to PM); these trends were the same for 3 years. Although the slope of the KDR or KMC is a parameter used for evaluation of the dehydration process, the index calculated by the fitting curve has no biological meaning. The purpose of the paper is to ensure the key periods and detect the key genes. Meanwhile, the specific moment is also the key point of phenotype identification.
The feasibility of multi-locus GWAS for detection of QTNs associated with the KMC and KDR
The multi-locus GWAS model has a higher discrimination ability and a lower false-positive rate for detection of animal and plant genes compared with the single-locus GWAS model36,37. Geneticists have introduced pleiotropy and population structure into the single-locus GWAS model to reduce the errors in effect estimation by controlling the genetic background24,38,39. Although the modification of the single-locus GWAS model improves its detection accuracy to a certain degree, the multiple-testing correction (e.g., Bonferroni correction) of significance thresholds for the single-locus model is too strict. This strict correction leads to the exclusion of important loci, especially when large experimental errors occur in field trials of crop genetics. To solve this problem, the application of multi-locus mrMLM is essential.
Previous studies have applied the multi-locus GWAS model to detect QTNs associated with important traits in various crops. Based on this model, Guan et al. detected 149 QTNs associated with the fatty acid content and composition of rape40, while Misra et al. detected 224 significant QTNs for the grain quality of rice, 97 of which were detected by at least two methods41. Additionally, Li et al. identified 42 SNPs and 5 QTNs for quality traits of forage sorghum42. Peng et al. detected 328 significant QTNs associated with free amino acid (FAA) levels in wheat, including 66 QTNs that were detected by more than two models, which revealed the complexity of metabolic and genetic regulation43. Lü et al. detected 159 QTNs for the photosynthetic response of soybean under low-phosphorus stress and discovered 52 candidate genes44. Hou et al.45 detected 20 QTNs that were significantly associated with drought resistance traits in cotton and further identified 1326 relevant genes, 205 of which were induced after drought stress treatment. Hu et al.46 detected 913 QTNs for agronomic traits of barley, including 39 QTNs that were repeatedly detected in various environments and by different methods, with 10 candidate genes identified through gene annotation In maize, Ma et al. 47 detected 63 QTNs for the regenerative capacity of embryonic callus and found 40 candidate genes based on these common QTNs. Moreover, Xu et al.48 identified a total of 60 QTNs for the gelatinization properties of maize starch. Zhang et al.49 detected 423 QTNs for maize stalk traits related to lodging resistance, 29, 34, and 48 of which were associated with stem diameter, stalk bending strength, and rind penetrometer resistance, respectively, as detected by multiple methods or across multiple environments. Based on these studies, conducting multi-locus GWAS for KMC- and KDR-associated traits in maize is feasible.
Detection of QTLs and QTNs associated with the KMC and KDR
The KMC and KDR of maize are complex quantitative traits with different impact factors in different periods. Physiological dehydration is controlled by genotype and could be stably inherited11,22,50. Natural drying is jointly determined by the KMC at PM, drying time, and the KDR and KMC at harvest, which are susceptible to environmental conditions9. Both the KMC and KDR have high heritability and can be stably inherited. It is feasible to map major-effect QTLs for the KMC and KDR, to mine associated candidate genes, and to develop practical functional markers for marker-assisted selection, which will be valuable for the breeding of maize cultivars for rapid dehydration. Many studies have mapped QTLs for the KMC and the KDR in maize3,4,16,17,18,19, but GWAS for these two traits have rarely been reported. Dai et al.20 selected 80 maize inbred lines to conduct a field survey for two consecutive years, performed a GWAS using 1,490,007 high-quality SNPs, and eventually detected 19 SNPs associated with natural KDR in the field. Zhang et al. conducted GWAS on kernel dehydration traits in maize using 290 inbred lines in combination with 201 simple sequence repeat (SSR) molecular markers, and 17 SSRs associated with natural KDR in the field were finally detected21. This study conducted a multi-locus GWAS for KMC- and KDR-associated traits in maize based on the Axiom Maize 55K Array data of 132 maize inbred lines. A total of 116 and 58 QTNs were detected by two and more than two methods, respectively. Among the 58 QTNs detected by three or more methods, 9 and 5 were detected in 2 or 3 years.
The QTNs identified in this study were correlated with QTLs reported earlier than the present report. Among these 14 QTNs, AX-123944713 and AX-90786057 were located in the qKdr-2-1 interval detected by Wang et al.51 and the intervals of q9GDR13-2-1 and qcGDR23-2-1 by Li et al.18; AX-90614081 were located in the q8GDR14-2-1 and qcGDR14-2-2 interval detected by Li et al.18 and the qKdr-2-2 interval by Liu et al.19; AX-90843001 was located in the intervals of q8GWC20-5-1, q9GWC10-5-2, qcGWC10-5-1, qcGWC20-5-1, q8GDR12-5-1 and qcGDR12-5-1 detected by Li et al.18; AX-90848774 was located in the qKdr-3-6 interval detected by Wang et al.51; AX-91652225 was located in the q9GWC10-5-11 interval detected by Li et al.18; AX-91654966 and AX-91442789 were located in the intervals of q8GWC10-5-1, q8GWC30-5-1, q8GWC40-5-1, q9GWC20-5-1, q9GWC30-5-1, q9GWC40-5-1, qcGWC30-5-1, qcGWC40-5-1, q8GDR13-5-1, q8GDR14-5-1 and qcGDR13-5-1 detected by Li et al.18; AX-91760347 was located in qKdr-8-2 interval detected by Wang et al.51; AX-91785266 was located in intervals of q9GWC30-9-1、q9GDR13-9-1、q9GDR34-9-1 and qcGDR13-9-1 detected by Li et al.18; AX-86286026, AX-91629217 and AX-86291963 have not been reported. Furthermore, a significant enrichment in cellular component was obtained through GO enrichment analysis of candidate genes in the intervals adjacent to the 14 QTNs detected in 2 or 3 years, and this category contained five genes.
Conclusions
The 132 inbred lines were genotyped using a maize 55K single-nucleotide polymorphism array. QTNs for the KMC and KDR in maize were detected based on five methods (mrMLM, FASTmrMLM, FASTmrEMMA, pLARmEB, and ISIS EM-BLASSO) in the package mrMLM. A total of 334 significant QTNs were identified for the KMC and the KDR, 116 and 58 of which were detected by two and more than two methods, respectively, while 9 and 5 QTNs were detected in 2 and 3 years, respectively. A significant enrichment in cellular component was revealed by GO enrichment analysis of candidate genes in the intervals adjacent to the fourteen QTNs, and this category contained five genes. The information provided in this study may be useful for further mining of genes associated with the KMC and the KDR in maize.
Materials and methods
Plant materials
A total of 132 maize inbred lines were used in this study, which were provided by the Institute of Crop Resources, Jilin Academy of Agricultural Sciences (JAAS) (Supplementary Table S7). These lines represented six subpopulations: BSSS (American BSSS including Reid), PA (group A germplasm derived from modern U.S. hybrids in China), PB (group B germplasm derived from modern U.S. hybrids in China), Lan (Lancaster Surecrop), LRC (derivative lines from Lvda Red Cob, a Chinese landrace), and SPT (derivative lines from Si-ping-tou, a Chinese landrace).
Field design and phenotypic measurement
The 132 lines were grown at the Gongzhuling (124°47′ N and 43°27′ E, Jilin Province, China) Experimental Base, JAAS in 2014, 2015 and 2016, with a final plant density of 75,000 plants ha-1. A randomized block experimental design with three replications was adopted. In 3 years, each plot had 5 rows, with a row length of 5 m, row spacing of 0.65 m, plant spacing of 0.20 m and plot area of 16.25 m2. Normal agronomic practices for maize were used in field management. To avoid border effects, for each plot, two border rows and the first two plants at each end of the middle three rows were not used for future trait determination.
The ears were bagged before silking (50% of plants in the row with extruded silks). Then the bagged ears were pollinated by hand. One week later, the bags were removed and five tested ears were randomly selected, tagged and labelled in each plot. The moisture concentration was recorded from 10 to 40 DAP, with one measurement of every five days. At 9:00 a.m., per the method published by Reid et al.34, for each ear, a SK-300 probe for moisture concentration measurement (manufactured by Harbin Yuda Electronic Technology Co., Ltd., China) was used to pierce into kernels after penetrating the bract leaves in the middle of the ear.
KMCs at 10, 15, 20, 25, 30, 35, and 40 DAP were measured, which were designated as KMC10, KMC15, KMC20, KMC25, KMC30, KMC35, and KMC40, respectively. KDRs were then calculated based on two consecutive KMC measurements. KDR = (KMC at a given time—KMC at the next time)/number of days during the time span. The KDRs for the six time spans (namely, 10–15, 15–20, 20–25, 25–30, 30–35 and 35–40 DAP) were denoted as KDR15, KDR20, KDR25, KDR30, KDR35, and KDR40, respectively.
Statistical analysis of phenotypic data
The F-values for genotypes (FG) and environments phenotypic data (FE) were analysed by SPSS 22 (IBM Corp., Armonk, NY, USA). Broad-sense heritability was calculated using the formula proposed by Knapp et al.52: \({H}^{2}={\delta }_{g}^{2}/\left[{\delta }_{g}^{2}+\left({\delta }_{gl}^{2}/n\right)+{\delta }_{e}^{2}/nr\right]\), where \({\delta }_{g}^{2}\) is the genetic variance, \({\delta }_{gl}^{2}\) is the variance of the genotype-by-environment interaction, \({\delta }_{e}^{2}\) is the error variance, n is the number of sites, and r is the number of replicates.\(CV=SD/Mean\).
Genotyping and filtering
Genomic DNA was extracted from young leaf samples of the 132 maize inbred lines using the modified cetyltrimethylammonium bromide (CTAB) method53. The quality of DNA was assessed using 0.8% agarose gel electrophoresis and a NanoDrop 2000 spectrophotometer (NanoDrop, Wilmington, DE, USA). Genotyping was performed using the Axiom Maize 55K SNP Array (CapitalBio Corp., Beijing, China) 31, which contained a total of 55,229 SNPs. The 132 inbred lines were genotyped based on the Affy AXIOM Array 2.0 platform. Genotyping data of the 132 inbred lines were filtered using the software PLINK54 with the settings of MAF > 0.05, GENO < 0.1, and genotype heterozygous loci missing.
Linkage disequilibrium and population structure analysis
LD among markers was calculated using PLINK software. The window size for LD calculation was set based on the number of SNPs located in the genome. Pair-wise linkage disequilibrium was measured using the squared allele frequency correlations, according to Weir55, and assessed by calculating r2 for pairs of SNP loci.
The population structure of 132 lines was assessed using STRUCTURE software 2.3.456. A burn-in of 5000 iterations followed by 50 000 Markov Chain Monte Carlo (MCMC) replicates was implemented to estimate the number of subpopulations (k) in a putative range of 2–10. To estimate the robustness of the inferred population structure, five replications were conducted for each k. The subpopulation number was estimated using an ad hoc statistic delta k based on the rate of change in the log probability of data between successive values57.
Genome-wide association studies
All the phenotypic and genotypic information in the above mapping population was used to detect QTNs using the mrMLM26, FASTmrEMMA29, FASTmrMLM30, pLARmEB28, and ISIS EM-BLASSO27 approaches, implemented by the software programme mrMLM v4.0 (https://cran.r-project.org/web/packages/mrMLM.GUI/index.html). The unified parameter settings for the five methods were as follows: (1) the Q + K model was used, where the population structure value Q was calculated by Structure 2.3.4 software56, and the kinship value K was analyzed by the “mrMLM” software package; and (2) the significance threshold p value was set as 0.0002 (limit of detection (LOD) = 3.0). In addition, while using mrMLM and FASTmrEMMA, the search radius of candidate genes was specified as 20 kb; using pLARmEB, 50 potential association loci were selected on each chromosome.
Annotation of candidate genes analysis
QTNs for the KMC and the KDR in maize detected by multiple methods were mapped to the maize reference genome B73 RefGen_V4 to identify associated candidate genes. The obtained candidate genes were subjected to GO enrichment analysis using AgriGO v2.058, and the final set of genes associated with the KMC and the KDR were identified.
References
National Bureau of Statistics of the People’s Republic of China. China Statistical Yearbook 1–12 (China Statistical Press, Beijing, 2019).
Chai, Z. H. et al. Current status of maize mechanical grain harvesting and its relationship with grain moisture content. Sci. Agric. Sin. 50, 2036–2043. https://doi.org/10.3864/j.issn.0578-1752.2017.11.009 (2017).
Capelle, V. et al. QTLs and candidate genes for Desiccation and Abscisic Acid content in maize kernels. BMC Plant Biol. 10, 2. https://doi.org/10.1186/1471-2229-10-2 (2010).
Xiang, K., Reid, L. M., Zhang, Z. M., Zhu, X. Y. & Pan, G. T. Characterization of correlation between grain moisture and ear rot resistance in maize by QTL meta-analysis. Euphytica 183, 185–195. https://doi.org/10.1007/s10681-011-0440-z (2012).
Sweeney, P. M., St Martin, S. K. & Clucas, C. P. Indirect inbred selection to reduce grain moisture in maize hybrids. Crop Sci. 34, 391–396. https://doi.org/10.2135/cropsci1994.0011183X003400020016x (1994).
Li, Z. Rapid Determination Method and Genome-wide Association Study of Maize Kernel Moisture Content in Mature Period. MAS Dissertation. Hebei Agricultural University, China (2019).
Kang, M. S. & Zhang, S. Narrow-sense heritability for and relationship between seed imbibition and grain moisture loss rate in maize. J. New Seeds. 3, 1–16. https://doi.org/10.1300/J153v03n02_01 (2001).
Shaw, R. H. & Loomis, W. E. Bases for the prediction of corn yields. Plant Physiol. 25, 225–244. https://doi.org/10.1104/pp.25.2.225 (1950).
Wang, K. R. & Li, S. K. Analysis of influencing factors on kernel dehydration rate of maize hybrids. Sci. Agric. Sin. 11, 27–35. https://doi.org/10.3864/j.issn.0578-1752.2017.11.008 (2017).
Hillson, M. T. & Penny, L. H. Dry matter accumulation and moisture loss during maturation of corn grain. Agron. J. 57, 150–153 (1965).
Purdy, J. L. & Crane, P. L. Inheritance of drying rate in “mature” corn (Zea mays L.). Crop Sci. 7, 294–297. https://doi.org/10.2135/cropsci1967.0011183X000700040003x (1967).
Nass, H. G. & Crane, P. L. Effect of endosperm mutants on drying rate in corn (Zea mays L.). Crop Sci. 10, 141–144. https://doi.org/10.2135/cropsci1970.0011183X001000020005x (1970).
Cross, H. Z. A selection procedure for ear drying-rate in maize. Euphytica 34, 409–418. https://doi.org/10.1007/BF00022936 (1985).
Cross, H. Z., Chyle, J. R. & Hammond, J. J. Divergent selection for ear moisture in early maize. Crop Sci. 27, 914–918. https://doi.org/10.2135/cropsci1987.0011183X002700050016x (1987).
Freppon, J. T., Martin, S. K. S., Pratt, R. C. & Henderlong, P. R. Selection for low ear moisture in corn, using a hand-held Meter. Crop Sci. 32, 1062–1064. https://doi.org/10.2135/cropsci1992.0011183X003200040046x (1992).
Zhang, L. The QTL Analysis of Kernel Dehydration Rate in Maize. MAS Dissertation of Yangzhou University, Jiangsu, Yangzhou, (2016).
Song, W. et al. Molecular mapping of quantitative trait loci for grain moisture at harvest in maize. Plant Breed. 136, 28–32. https://doi.org/10.1111/pbr.12430 (2016).
Li, Y. L. et al. QTL detection for grain water relations and genetic correlations with grain matter accumulation at four stages after pollination in maize. Plant Biochem. Physiol 2, 1–9. https://doi.org/10.4172/2329-9029.1000121 (2014).
Liu, X. J., Wang, Z. H., Wang, X., Li, T. F. & Zhang, L. Primary mapping of QTL for dehydration rate of maize kernel after physiological maturing. Acta Agron. Sin. 36, 47–52. https://doi.org/10.3724/SP.J.1006.2010.00047 (2010).
Dai, L. Q. et al. Genome-wide association study of field grain drying rate after physiological maturity based on a resequencing approach in elite maize germplasm. Euphytica 213, 182. https://doi.org/10.1007/s10681-017-1970-9 (2017).
Zhang, J. et al. Genome-wide association study identifies genetic factors for grain filling rate and grain drying rate in maize. Euphytica 212, 201–212. https://doi.org/10.1007/s10681-016-1756-5 (2016).
Zhang, L., Wang, Z. H., Jin, Y. & Yu, T. J. Combining ability analysis of water content in harvest stage in corn. Southwest China J. Agric. Sci. 05, 32–35 (2005).
Yu, J. & Buckler, E. S. Genetic association mapping and genome organization of maize. Curr. Opin. Biotechnol 17, 155–160 (2006).
Zhang, Z. et al. Mixed linear model approach adapted for genome-wide association studies. Nat. Genet. 42, 355–360. https://doi.org/10.1038/ng.546 (2010).
Li, M. et al. Enrichment of statistical power for genome-wide association studies. BMC Biol. 12, 73. https://doi.org/10.1186/s12915-014-0073-5 (2014).
Wang, S. B. et al. Improving power and accuracy of genome-wide association studies via a multi-locus mixed linear model methodology. Sci. Rep. 6, 19444. https://doi.org/10.1038/srep19444 (2016).
Tamba, C. L., Ni, Y. L. & Zhang, Y. M. Iterative sure independence screening EMBayesian LASSO algorithm for multi-locus genome-wide association studies. PLoS Comput. Biol. 13, e1005357. https://doi.org/10.1371/journal.pcbi.1005357 (2017).
Zhang, J. et al. pLARmEB: integration of least angle regression with empirical Bayes for multilocus genome-wide association studies. Heredity 118, 517–524. https://doi.org/10.1038/hdy.2017.8 (2017).
Wen, Y. J. et al. Methodological implementation of mixed linear models in multi-locus genome-wide association studies. Brief. Bioinform. 19, 700–712. https://doi.org/10.1093/bib/bbw145 (2018).
Tamba, C. L. & Zhang, Y. M. A fast mrMLM algorithm for multi-locus genome-wide association studies. bioRxiv https://doi.org/10.1101/341784 (2018).
Xu, C. et al. Development of a Maize 55 K SNP array with improved genome coverage for molecular breeding. Mol Breed. 37, 20. https://doi.org/10.1007/s11032-017-0622-z (2017).
Xie, C. et al. Zea mays (L.) P1 locus for cob glume color identified as a post-domestication selection target with an effect on temperate maize genomes. Crop J. 1, 15–24. https://doi.org/10.1016/j.cj.2013.07.002 (2013).
Hart, L. P., Gendloff, E. & Rossman, E. C. Effect of corn genetypes on ear rot infection by Gibberella zeae. Plant Dis. 68, 296–298. https://doi.org/10.1094/PD-69-296 (1984).
Reid, L. M. et al. A non-destructive method for measuring maize kernel moisture in a breeding program. Maydica 55, 163–171. https://doi.org/10.3198/jpr2009.06.0350crmp (2010).
Gambín, B. L., Borrás, L. & Otegui, M. E. Kernel water relations and duration of grain filling in maize temperate hybrids. Field Crops Res. 101, 1–9. https://doi.org/10.1016/j.fcr.2006.09.001 (2007).
Segura, V. et al. An efficient multi-locus mixed-model approach for genome-wide association studies in structured populations. Nat. Genet. 44, 825–830. https://doi.org/10.1038/ng.2314 (2012).
Wang, S. B. et al. Mapping small-effect and linked quantitative trait loci for complex traits in backcross or DH populations via a multi-locus GWAS methodology. Sci. Rep. 6, 29951. https://doi.org/10.1038/srep29951 (2016).
Zhang, Y. M. et al. Mapping quantitative trait loci using naturally occurring genetic variance among commercial inbred lines of maize. Genetics 169, 2267–2275. https://doi.org/10.1371/journal.pone.0029350 (2005).
Yu, J. et al. A unified mixed-model method for association mapping that accounts for multiple levels of relatedness. Nat. Genet. 38, 203–208. https://doi.org/10.1038/ng1702 (2006).
Guan, M. W. et al. Association mapping analysis of fatty acid content in different ecotypic rapeseed using mrMLM. Front. Plant Sci. 9, 1872. https://doi.org/10.3389/fpls.2018.01872 (2019).
Misra, G. et al. Deciphering the genetic architecture of cooked rice texture. Front. Plant Sci. 9, 1405. https://doi.org/10.3389/fpls.2018.01405 (2018).
Li, J. Q. et al. Genome-wide association studies for five forage quality-related traitsin sorghum (Sorghum bicolor L.). Front. Plant Sci. 9, 1146. https://doi.org/10.3389/fpls.2018.01146 (2018).
Peng, Y. C. et al. Genome-wide association studies of free amino acid levels by six multi-locus models in bread wheat. Front. Plant Sci. 9, 1196. https://doi.org/10.3389/fpls.2018.01196 (2018).
Lü, H. Y. et al. Genome-wide association studies of photosynthetic traits related to phosphorus efficiency in soybean. Front. Plant Sci. 9, 1226. https://doi.org/10.3389/fpls.2018.01226 (2018).
Hou, S. et al. Genome-wide association studies reveal genetic variation and candidate genes of drought stress related traits in cotton (Gossypium hirsutum L.). Front. Plant Sci. 9, 1276. https://doi.org/10.3389/fpls.2018.01276 (2018).
Hu, X. et al. Multi-locus genome-wide association studies for 14 main agronomic traits in barley. Front. Plant Sci. 9, 1683. https://doi.org/10.3389/fpls.2018.01683 (2018).
Ma, L. L. et al. Genetic dissection of maize embryonic callus regenerative capacity using multi-locus genome-wide association studies. Front. Plant Sci. 9, 561. https://doi.org/10.3389/fpls.2018.00561 (2018).
Xu, Y. et al. Genome-wide association mapping of starch pasting properties in maize using single-locus and multi-locus models. Front. Plant Sci. 9, 1311. https://doi.org/10.3389/fpls.2018.01311 (2018).
Zhang, Y. L. et al. Multi-locus genome-wide association study reveals the genetic architecture of stalk lodging resistance-related traits in maize. Front. Plant Sci. 9, 611. https://doi.org/10.3389/fpls.2018.00611 (2018).
Shi, Y. Q., Meng, Q. L., Yang, S. W. & Zhang, Y. W. Research development of kernel dehydration rate in maize. China Seed Ind. 278, 33–35. https://doi.org/10.19462/j.cnki.1671-895x.20180404.015 (2018).
Wang, Z. H. et al. QTL underlying field grain drying rate after physiological maturity in maize (Zea mays L.). Euphytica 185, 521–528. https://doi.org/10.1007/s10681-012-0676-2 (2012).
Knapp, S. J., Stroup, W. W. & Ross, W. M. Exact confidence intervals for heritability on a progeny mean basis. Crop Sci. 25, 192–194. https://doi.org/10.2135/cropsci1985.0011183X002500010046x (1985).
Saghai-Maroof, M. A., Soliman, K., Jorgensen, R. A. & Allard, R. W. Ribosomal DNA spacer length polymorphism in barley: Endelian inheritance, chromosomal location and population dynamics. Proc. Natl. Acad. Sci. USA 81, 8014–8018 (1984).
Purcell, S. et al. Plink: A tool set for whole-genome association and population based linkage analyses. Am. J. Hum. Genet 81, 559–575. https://doi.org/10.1086/519795 (2007).
Williams, J. T. Genetic data analysis II: Methods for discrete population genetic data by Bruce S. Weir. Hum. Biol. 4, 583–586 (1996).
Pritchard, J. K., Stephens, M., Rosenberg, N. A. & Donnelly, p. Association mapping in structured populations. Am. J. Hum. Genet. 67, 170–181. https://doi.org/10.1086/302959 (2000).
Evanno, G., Regnaut, S. & Goudet, J. Detecting the number of clusters of individuals using the software structure: A simulation study. Mol. Ecol. 14, 2611–2620. https://doi.org/10.1111/j.1365-294x.2005.02553.x (2005).
Tian, T. et al. AgriGO v2.0: A GO analysis toolkit for the agricultural community. Nucleic Acids Res. 45, W122–W129. https://doi.org/10.1093/nar/gkx382 (2017).
Acknowledgements
The authors thank Professor Yuan-Ming Zhang from College of Plant Science & Technology of Huazhong Agricultural University for providing the mrMLM software package and helping us in GWAS, and thank Researcher Han Zhao from Provincial Key Laboratory of Agrobiology, Institute of Crop Germplasm and Biotechnology of Jiangsu Academy of Agricultural Sciences for providing assistance.
Funding
The work was supported by the Key Research Project of Science and Technology Department of Jilin Province (20200402024NC).
Author information
Authors and Affiliations
Contributions
X.L. and W.L. initiated the research. X.L., D.Y., S.L. and W.L. designed the experiments. C.Z., Y.Q. and F.J. performed the molecular experiments and data analysis. M.L., X.L. and S.L. conducted the phenotypic evaluations and collected the data from the field. S.L. and C.Z. drafted the manuscript. X.L. and W.L. finalized the manuscript. All authors contributed in the interpretation of results and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, S., Zhang, C., Yang, D. et al. Detection of QTNs for kernel moisture concentration and kernel dehydration rate before physiological maturity in maize using multi-locus GWAS. Sci Rep 11, 1764 (2021). https://doi.org/10.1038/s41598-020-80391-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-80391-1
This article is cited by
-
A combination of QTL mapping and genome-wide association study revealed the key gene for husk number in maize
Theoretical and Applied Genetics (2024)
-
Genome-wide association analysis for grain moisture content and dehydration rate on maize hybrids
Molecular Breeding (2023)
-
Evaluation of local wheat (Triticum aestivum L.) germplasm resources and mining of excellent drought resistance loci in Shanxi province
Genetic Resources and Crop Evolution (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.