Abstract
Phytophthora nicotianae is highly pathogenic to Solanaceous crops and is a major problem in tobacco production. The tobacco cultivar Beihart1000-1 (BH) is resistant, whereas the Xiaohuangjin 1025 (XHJ) cultivar is susceptible to infection. Here, BH and XHJ were used as models to identify resistant and susceptible genes using RNA sequencing (RNA-seq). Roots were sampled at 0, 6, 12, 24, and 60 h post infection. In total, 23,753 and 25,187 differentially expressed genes (DEGs) were identified in BH and XHJ, respectively. By mapping upregulated DEGs to the KEGG database, changes of the rich factor of “plant pathogen interaction pathway” were corresponded to the infection process. Of all the DEGs in this pathway, 38 were specifically regulated in BH. These genes included 11 disease-resistance proteins, 3 pathogenesis-related proteins, 4 RLP/RLKs, 2 CNGCs, 7 calcium-dependent protein kinases, 4 calcium-binding proteins, 1 mitogen-activated protein kinase kinase, 1 protein EDS1L, 2 WRKY transcription factors, 1 mannosyltransferase, and 1 calmodulin-like protein. By combining the analysis of reported susceptible (S) gene homologs and DEGs in XHJ, 9 S gene homologs were identified, which included 1 calmodulin-binding transcription activator, 1 cyclic nucleotide-gated ion channel, 1 protein trichome birefringence-like protein, 1 plant UBX domain-containing protein, 1 ADP-ribosylation factor GTPase-activating protein, 2 callose synthases, and 2 cellulose synthase A catalytic subunits. qRT-PCR was used to validate the RNA-seq data. The comprehensive transcriptome dataset described here, including candidate resistant and susceptible genes, will provide a valuable resource for breeding tobacco plants resistant to P. nicotianae infections.
Similar content being viewed by others
Introduction
As a typical oomycete, Phytophthora nicotianae has a broad host range; this pathogen causes root rot, crown rot, fruit rot, and leaf and stem infections 1,2,3,4. P. nicotianae can attack all parts of Nicotiana tabacum, including the roots, stems, and leaves at any stage of its growth, and the most common symptom of infection is a black base or shank of the stalk 5. The disease can be devastating to tobacco in the greenhouse, as well as in the field, leading to severe yield losses every year worldwide 6. In a recent ranking of oomycete species based on scientific and economic importance, P. nicotianae was listed eighth 7. Additionally, P. nicotianae is well adapted to high temperatures; therefore, it is gaining importance in agriculture and plant health worldwide as the trend of global warming increases8.
Multiple sources have been used to improve resistance in cultivated tobacco for P. nicotianae. For example, the Php and Phl genes were introgressed from the closely related species N. plumbaginifolia and N. longiflora, providing immunity to race 0 of P. nicotianae 4. In addition to dominant resistance, polygenic resistance, which occurs in commercial flue-cured and burley tobacco cultivars, was likely derived from the cigar tobacco cultivar Florida 301 9. This cultivar was produced from crossing the cultivars Big Cuba and Little Cuba by Tisdale in the 1930s, and it is resistant to all known strains of P. nicotianae 9. The cultivar Beinhart 1000 (BH), which originated from selection of the cultivar Quin Diaz, has the highest reported level of quantitative resistance to P. nicotianae, and resistance in this line may be effective against all races 10. Another alien gene, Wz, introgression from N. rustica, has been found to confer a high level of resistance to race 0 and 1 11. Although multiple resistance sources are used in breeding against P. nicotianae, none of these resistant genes have been cloned, and the mechanism of resistance has not been elucidated.
The coevolution of plants and pathogens has resulted in the development of a multifaceted and sophisticated plant immune system. In addition to barriers at the surface of plant cells, plants have developed two layers of induced defense responses that rely on the recognition of pathogen-, microbe-, or damage-associated molecular patterns (PAMPs, MAMPs, or DAMPs, respectively) and pathogen effectors, known as PAMP-triggered immunity (PTI) and effector-triggered immunity (ETI) 12. This type of immune system has been termed a “zigzag” model 12. Conversely, pathogens require host cooperation to establish a compatible interaction. Plant genes that facilitate infections can be considered as susceptibility (S) genes 13, and disrupting these S genes may interfere with the compatibility between the host and the pathogen 13. A breeding strategy that involved disabling plant S genes was proposed in 2010 14. Recently, more attention has been paid to the study and exploitation of plant S genes as a source of broad-spectrum and durable resistance 15. Functional screens in Arabidopsis have yielded many S gene candidates 13. Based on available genome and transcriptome sequencing data, homologous gene sequences from other crops can be identified.
Thus, systematic determination of key resistance and susceptibility genes in response to infection by P. nicotianae would help to accelerate the breeding of new strains. RNA sequencing (RNA-seq) is a powerful tool for studying disease resistance in plants. In this study, we used the resistant cultivar Beinhart 1000–1 (hereafter, BH), a selection of Beinhart 1000, and the susceptible cultivar Xiaohuangjin 1025 (hereafter, XHJ) 16 as models. RNA-seq was used to analyze the gene expression profiles of roots from both cultivars at 6, 12, 24, and 60 h post-inoculation (hpi) with P. nicotianae. We identified DEGs in both cultivars during the infection process and screened tobacco resistant and susceptible genes to P. nicotianae. Our results provide a basis for understanding the mechanisms of the responses of tobacco to P. nicotianae and provide a potentially valuable resource for the future development of resistant plants.
Results
Transcriptome sequencing of resistant and susceptible tobacco cultivars infected by P. nicotianae
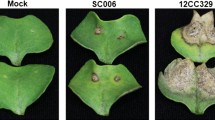
The primary symptoms induced by P. nicotianae infection, such as slight leaf wilting, appeared in XHJ at 60 hpi. At 5 days post-inoculation (dpi), differences in symptoms were observed between the two cultivars. Leaf wilting and severe stem necrosis occurred in XHJ, whereas these symptoms were not apparent in BH (Fig. 1).
Disease symptoms in the cultivars BH (resistant) and XHJ (susceptible) at 5 dpi by Phytophthora nicotianae. The basal parts of stems are magnified and shown in the circles. In BH (Left), no symptoms were apparent. In XHJ (right), leaves were withered and the basal part of stem was severely necrosed and became black.
To investigate the transcriptional differences between BH and XHJ in response to P. nicotianae, we exposed 6-week-old seedlings of BH and XHJ to this pathogen. Five sequencing libraries were generated for each cultivar from the total RNA of healthy root tissues and infected root tissues at 6, 12, 24, and 60 hpi. A total of 1,475,991,460 clean reads (221.4 GB) were generated using Illumina RNA-Seq deep sequencing. Clean data were submitted to the NCBI Sequence Reads Archive (SRA) database (Accession number: PRJNA679433). The reads of all samples (inoculated BH, non-inoculated BH, inoculated XHJ, and non-inoculated XHJ) were used for transcriptome assembly (Table 1). On average, 91.09% (BH at 0 hpi), 92.19% (BH at 6 hpi), 92.08% (BH at 12 hpi), 89.33% (BH at 24 hpi), 77.94% (BH at 60 hpi), 90.60% (XHJ at 0 hpi), 92.65% (XHJ at 6 hpi), 92.26% (XHJ at 12 hpi), 71.26% (XHJ at 24 hpi), and 7.26% (XHJ at 60 hpi ) were mapped to the reference transcriptome (Supplementary Table S1).
Identification of DEGs in resistant and susceptible tobacco cultivars during the infection process and GO enrichment analysis
The fragments per kilobase per million (FPKM) values for each unigene in all 30 libraries were computed and are displayed in Supplementary Table S2. Gene expression profiles in healthy BH and XHJ roots were used as the baselines. If there was a two-fold (or more) difference in gene expression in infected roots relative to the baseline (p < 0.001), the gene was regarded as a DEG. As shown in Fig. 2, for BH inoculated with P. nicotianae, there were 11,696 DEGs at 6 hpi, and the number of DEGs increased gradually between 12 hpi (14,448) and 24 hpi (16,669) and finally increased to 17,179 at 60 hpi. In comparison, in P. nicotianae-infected XHJ roots, there were 16,050 DEGs at 6 hpi and only 9,200 DEGs at 12 hpi, which then increased to 19,618 at 24 hpi and again decreased to 9,725 at 60 hpi. Venn diagrams were generated from the DEGs identified at 6, 12, 24, and 60 hpi, corresponding to each of the cultivar-pathogen combinations. At all-time points, 23,753 DEGs were identified in the BH group, whereas there were 7,060 DEGs shared at all-time points. A total of 25,187 DEGs were identified in the XHJ group, with a total of 4,009 shared DEGs at all-time points.
To identify differences in Gene Ontology (GO) term enrichment analysis between the two cultivars after inoculation, all DEGs at different time points in BH (23,753) and XHJ (25,187) were analyzed using GOseq, with a cut-off corrected p value of 10−10. 29 and 15 GO terms were identified in BH and XHJ respectively (Table 2). The most significantly enriched GO term in BH and XHJ was the structural constituent of the ribosome. In biological process (BP) terms, expression of genes related to metabolic processes, cellular protein metabolic processes, protein phosphorylation, and phosphorylation were remarkably more significant in BH than in XHJ. In cellular component (CC) terms, intracellular non-membrane-bound organelles and non-membrane-bounded organelles were more significant in BH than in XHJ. In molecular function (MF) terms, structural molecule activity, antioxidant activity, oxidoreductase activity, acting on peroxide as an acceptor, protein kinase activity, peroxidase activity, heme binding, and tetrapyrrole binding were remarkably more significant in BH than in XHJ. In contrast, transferase activity was more significant in XHJ than in BH.
Screening and analysis of resistant genes against P. nicotianae in tobacco
To further elucidate the functions of DEGs and to analyze the resistance mechanism of BH, KEGG pathway enrichment analysis involving upregulated DEGs at different infection time points was conducted by mapping to the KEGG database. The top 20 metabolic pathways associated with these DEGs in BH are shown in Fig. 3. Among these, changes in the rich factor of the “plant-pathogen interaction” pathway corresponded to the infection process and were much higher in BH than in XHJ (Supplementary Fig. 1). As shown in Fig. 3, the rich factor of the “plant pathogen interaction pathway” in BH was nearly 0.45 at 12 hpi, which increased to almost 0.58 at 24 hpi and declined to 0.52 thereafter.
Scatterplot of KEGG pathways enrichment analysis for upregulated DEGs at (a) 6, (b) 12, (c) 24, and (d) 60 hpi in BH. The rich factor is the ratio of the number of DEGs annotated in a given pathway term to the number of all genes annotated in the pathway term. A higher rich factor indicates greater intensity. The Q value is the corrected P value and ranges from 0 to 1, with a lower Q value indicating greater intensity. The sizes of the circles indicate the number of genes. The top 20 enriched pathway terms in the KEGG database are listed. The blue arrows indicates the plant-pathogen interaction pathway.
To investigate the differences in resistance mechanisms between BH and XHJ, a Venn diagram was generated from the DEGs identified in the BH and XHJ groups and all DEGs in the “plant pathogen interaction pathway” (Fig. 4). In total, 5,489 DEGs were specifically identified in BH, 6,923 DEGs were specifically identified in XHJ, and 18,264 DEGs were common to both BH and XHJ. Of all the DEGs in the “plant pathogen interaction pathway”, 38 were specifically regulated in BH.
The expression data of the 38 DEGs are shown in Fig. 5. Remarkably, there were 11 disease resistance proteins upregulated in BH, especially at 24 hpi and 4 pathogenesis-related proteins upregulated in BH, especially at 60 hpi. In addition, 4 RLP/RLKs, 2 CNGCs, 7 calcium-dependent protein kinases, 4 calcium-binding proteins, 1 mitogen-activated protein kinase kinase, 1 protein EDS1L, 2 WRKY transcription factors, and 1 mannosyltransferase were upregulated in BH, whereas 1 calmodulin-like protein was downregulated in BH.
By further focusing on the disease resistance proteins and pathogenesis-related protein, we found that, 1 disease resistance protein was a homolog of At4g10780, 3 were homologs of At4g27190, 4 were homologs of RPM1 and 1 was a homolog of RPP8. Pathogenesis-related proteins were homologs of pathogenesis-related protein 1B and 1C. The fold change versus the mock-infected control is shown in Table 3.
Screening and analysis of P. nicotianae susceptible genes in tobacco
To screen for S genes in tobacco, 28 S genes reported to interact with oomycetes or fungi in other crops were chosen, and their 56 homologs in tobacco were identified (Supplementary Table S3). Venn diagrams were generated from the DEGs identified in the BH group, XHJ group, and 56 S gene homologs (Fig. 6); 9 S gene homologs were specifically identified in XHJ.
Expression data of the 9 S gene homologs specifically identified in XHJ are shown in Fig. 7. These included 1 calmodulin-binding transcription activator, 1 cyclic nucleotide-gated ion channel, 1 protein trichome birefringence-like protein, 1 plant UBX domain-containing protein, 1 probable ADP-ribosylation factor GTPase-activating protein, and 2 callose synthases, all of which were specifically upregulated in XHJ, especially at 24 hpi. Two cellulose synthase A catalytic subunits were specifically downregulated in XHJ, especially at 60 hpi. Pathogens require the cooperation of host S genes to establish a compatible interaction. During the infection process, pathogens deploy effectors to inhibit defense networks, thereby activating S genes. Therefore, gene upregulation may be a criterion for screening candidate S genes.
In XHJ, at 24 hpi, expression of homologs of calmodulin-binding transcription activator 3, cyclic nucleotide-gated ion channel 4, protein trichome birefringence-like 37, plant UBX domain-containing protein 2, ADP-ribosylation factor GTPase-activating protein, and 2 callose synthases increased more than threefold compared to the mock control (Table 4).
Identification of P. nicotianae genes during the early infection stage
To identify DEGs in P. nicotianae during the early infection stage in the susceptible variety, reads from 9 sequencing libraries (XHJ at 6 hpi, XHJ at 12 hpi, and XHJ at 24 hpi) were aligned to the reference genome of P. nicotianae race 0 (NCBI: PRJNA294216). On average, 0.08% (XHJ at 6 hpi), 0.09% (XHJ at 12 hpi), and 6.17% (XHJ at 24 hpi) were mapped to the reference genome. The fragments per kilobase per million (FPKM) values for each unigene in 9 libraries were computed and are shown in Supplementary Table S4. Within these genes, some RxLR effectors 17, AM587_10007145, AM587_10001643, and AM587_10002874 were sharply expressed at 6 hpi and 24 hpi in XHJ.
Verification of DEGs using quantitative reverse-transcription PCR (qRT-PCR)
To confirm the results obtained using RNA-seq, we chose 10 sharply upregulated homologs of 6 reported S genes (ADH1, WRKY48, bHLH25, PLP2, KMD3, PUB24) 13 and performed qRT-PCR. The results of the qRT-PCR correlated with the RNA-seq data (evaluated by FPKM), and the gene expression increased more significantly in XHJ than in BH (Fig. 8). Expression of homologs of PUB24 and bHLH25 rose sharply at 12 hpi, homologs of ADH1 and KMD3 rose sharply at 24 hpi, and the homologs of WRKY48 and PLP2 rose sharply at 60 hpi.
The relative expression levels of identified susceptible genes at each time point after P. nicotianae inoculation in BH and XHJ. Blue bars indicate BH, whereas red bars indicate XHJ. Actin was used as an internal control, and the transcript level in non-infected plants was set as 1.0. Error bars represent standard error (n = 3).
Discussion
The transcriptome has been widely used to study the defense response in plants and to identify specific genes that interact with pathogens. In Lomandra longifolia roots, callose synthase genes, MAPK 15, 2 PR genes, and 5 receptor-like protein genes were found to be significantly expressed when infected by Phytophthora cinnamomi 18. In N. benthamiana leaves, expression of 13 b-1,3-glucanases from the PR-2 family, and 16 chitinases from the PR-3, PR-4, PR-8, and PR-11 families were induced following infection with P. parasitica 19. In sugarcane stalks, cytochrome P450, chitinase, NBS-LRR domain-containing proteins, and leucine zipper domain proteins were identified when infected by Sporisorium scitamineum 20. Furthermore, pathogens deploy effector proteins to inhibit defense networks; therefore, susceptibility factors encoded by S genes can be activated during infection 13. In pepper, the expression of CaMlo2 is upregulated at an earlier time point following Leveillula taurica infection, and complementation experiments confirmed the role of CaMlo2 as a susceptibility factor to different powdery mildews 21. SWEET sugar transporters, susceptible factors, can be upregulated during pathogen attack to export sugars from cells into the extracellular spaces 22. The transcription factors bHLH25 and bHLH27 positively influence cyst nematode parasitism and were upregulated at 1 dpi 23. Thus, it is feasible to identify resistant and susceptible genes using RNA-seq.
By profiling genes specifically regulated in resistant tobacco before and after infection with P. nicotianae, 11 disease resistance proteins, 3 pathogenesis-related proteins, 4 RLP/RLKs, 2 CNGCs, 7 calcium-dependent protein kinases, 4 calcium-binding protein, 1 mitogen-activated protein kinase kinase, 1 protein EDS1L, 2 WRKY transcription factor, 1 mannosyltransferase and 1 calmodulin-like protein were identified. In Arabidopsis, RPM1 confers resistance against Pseudomonas syringae expressing AvrRpm1 or AvrB 24, whereas RPP8 confers resistance to P. parasitica 25. PR-1 proteins are produced abundantly during defense responses, and have been shown to possess sterol-binding activity 26. Cyclic nucleotide-gated channels (CNGCs) are nonselective cation channels that permit the diffusion of divalent and monovalent cations. CNGCs are involved in both basal and R gene-mediated plant immunity 27. In plants, Ca2+-stimulated protein kinase activities occur via activation of calcium-dependent protein kinases (CDPKs) 28. NtCDPK2 was initially identified in the Cf-9/Avr9 pathosystem and is activated in response to race-specific elicitation 29. The Arabidopsis calmodulin-like protein CML36 is a Ca2+ sensor that interacts with ACA8 and stimulates its activity 30. The wheat calmodulin-like protein TaCML36 positively participates in the immune response to Rhizoctonia cerealis 31. EDS1 family members control plant basal immunity and ETI 32. Remarkably, the homologs of RPM1 (LOC107804213, LOC107804212, LOC107802041 and LOC107775646) increased sharply in BH at 24 hpi, whereas the homologs of pathogenesis-related protein 1 (LOC107807833 and LOC107808770) increased sharply in BH at 60 hpi. By profiling P. nicotianae genes during the early infection stage, some RxLR effectors were upregulated at 24 hpi. Phtophthora RxLR effectors affect various aspects of plant immune systems. Some of them inhibit the positive regulation of plant immunity 33,34,35, whereas some of them promote negative regulators of plant immunity 36,37,38. The specific recognition of RxLR effectors by one of the NB-LRR proteins will activate plant immunity 11. In N. benthamiana leaves infected with P. nicotianae, biotrophic growth was dominant before 24 hpi, followed by a rapid switch to necrotrophic growth 39. It is speculated that 24 hpi is a key time point of interaction between tobacco roots and P. nicotianae. With the spread of abundant hyphae invasion at 24 hpi, some effectors were recognized by LOC107804213, LOC107804212, LOC107802041, and LOC107775646 to trigger plant immune response. The pathogenesis-related PR-1 proteins were activated during defense responses around 60 hpi and inhibited the growth of P. nicotianae by sequestrating sterol from this pathogen.
By profiling genes specifically regulated in susceptible tobacco before and after infection with P. nicotianae, 9 S gene homologs were identified. These genes included 1 calmodulin-binding transcription activator (CAMTA3), 1 cyclic nucleotide-gated ion channel (CNGC4), 1 protein trichome birefringence-like protein (PMR5), 1 plant UBX domain-containing protein (PUX2), 1 probable ADP-ribosylation factor GTPase-activating protein (AGD5), 2 callose synthases (PMR4) and 2 cellulose synthase A catalytic subunits (CESA3). By focusing on the upregulated genes, CAMTA3 negatively regulates SA accumulation and plant defenses through calmodulin binding 40. Mutations in the pmr5 gene can render Arabidopsis resistant to the powdery mildew species Erysiphe cichoracearum and E. orontii 41. Mutants of PUX2, a plant ubiquitin regulatory X domain-containing protein 2, results in significantly enhanced resistance to powdery mildew Golovinomyces orontii in Arabidopsis 42. Arabidopsis ARF-GAP protein, AGD5, is a susceptibility factor for Hyaloperonospora arabidopsidis 43. CNGC4 is a cyclic nucleotide-gated ion channel gene, the mutation of which can enhance resistance to Pseudomonas syringae 44. In Arabidopsis, mutation in CNGC4 lead to high expression of PR-1, elevated levels of SA, and elevation “SAR-like” resistance in response to virulent pathogens 44. PMR4 (also known as GSL5) is a callose synthase gene. Silencing of the ortholog in tomato resulted in increased resistance to the adapted powdery mildew pathogen 45, and silencing of potato orthologs increased the resistance to late blight 46. In addition, we identified several S gene homologs that were sharply upregulated in both BH and XHJ cultivars. They were homologs of ADH1 (LOC107772756, LOC107830380), WRKY48 (LOC107832061), bHLH25 (LOC107804684), PLP2 (LOC107797353), KMD3 (LOC107775293, LOC107797281), PUB24 (LOC107808916, LOC107831963, LOC107791734). Alcohol dehydrogenase 1 (ADH1) of barley modulates susceptibility to the fungus Blumeria graminis f.sp. hordei 47. WRKY48 negatively regulates PR gene expression and basal resistance to the bacterial pathogen P. syringae 48. Arabidopsis bHLH25 and bHLH27 transcription factors positively influence the susceptibility to the cyst nematode Heterodera schachtii 23. Patatin-like protein 2 (PLP2) promotes cell death and negatively regulates Arabidopsis resistance to the fungus Botrytis cinerea 49. Expression of the F-box/Kelch-repeat protein At2g44130 (KMD3) promotes susceptibility to the root-knot nematode Meloidogyne incognita 50. A homologous triplet of U-box type E3 ubiquitin ligases (PUBs), PUB22, PUB23, and PUB24 in Arabidopsis, negatively regulates PTI in response to several distinct PAMPs51. These genes were not specifically regulated in XHJ, suggesting that they may not be key S genes determining different resistance in BH and XHJ. As pathogens can regulate S gene expression, further study of these sharply upregulated S genes in both BH and XHJ will contribute to revealing the interaction between tobacco and P. nicotianae. Further research on the S genes in XHJ will contribute to uncovering the differences in resistance mechanisms between BH and XHJ. Remarkably, all upregulated genes increased sharply in XHJ at 24 hpi, which confirmed the hypothesis that 24 hpi was a key time point of interaction between tobacco roots and P. nicotianae.
In this study, we identified candidate genes related to resistance and susceptibility to P. nicotianae in tobacco-resistant and -susceptible materials. Our results provide further insights into the molecular mechanisms underlying the interaction between P. nicotianae and tobacco, which will be useful in organizing resistant breeding practices. However, further reliable evidence is required to validate our results. For example, the use of VIGS to transiently silence genes to elucidate their function, and the application of CRISPR/Cas9 to fully knockout candidate genes would help to verify our results.
In this study, we provided insight into the P. nicotianae infection process in tobacco cultivars and investigated resistant and susceptible genes using the Illumina HiSeq platform. The resistant tobacco cultivar BH and susceptible tobacco cultivar XHJ were used as research objects, and the samples were collected at 0, 6, 12, 24, and 60 hpi. Thirty-eight defense-related genes were identified in BH, whereas nine susceptible genes were identified in XHJ. Our results provide a valuable resource for resistant breeding to P. nicotianae although further research is needed to explore the function of the identified resistant and susceptible genes.
Materials and methods
Plant growth condition and inoculation treatments
The resistant tobacco cultivar BH and the susceptible tobacco cultivar XHJ were cultivated in Hogland nutrient solution in a growth chamber under a 16 h light/8 h dark photoperiod, at 22 °C in the Tobacco Research Institute of Chinese Academy of Agricultural Sciences. A field-isolate of P. nicotianae race 0 was used for all inoculations throughout the study. Mycelial cultures of P. nicotianae were grown on oatmeal agar medium at 25 °C for 14 days. The roots of 8-week-old tobacco plantlets were laid on oatmeal agar medium and inoculated at 25 °C in the dark. The infected roots were harvested at five time points: 0, 6, 12, 24, and 60 hpi. Three independent experiments were performed for each treatment condition. Roots from all groups were sampled and immediately stored at − 80 °C.
RNA-seq
Total RNA was isolated from samples using TRIzol reagent according to the method of kit instructions. After quality confirmation, RNA samples were sent to Novogene (Beijing, China) for RNA sequencing (Illumina Novaseq platform with 150-bp paired-end reads). According to the manufacturer’s instructions, cDNA library construction and Illumina sequencing were performed with three technical replicates performed per sample.
Transcriptome data processing
The sequencing data were filtered with SOAPnuke (v1.5.2) by: (1) Removing reads containing a sequencing adapter; (2) Removing reads with a low-quality base ratio more than 20% (base quality less than or equal to 5); (3) Removing reads whose unknown base (‘N’ base) ratio is more than 5%. Thus, clean reads were obtained and stored in FASTQ format. De novo assembly of the transcriptome was performed with the short reads assembling program, Trinity v2.0.6 52. The candidates that had the probable longest open reading frame were generated from the Trinity assembly result. A set of candidates was used as the reference transcriptome. If multiple transcripts belonged to one unigene, the coding sequences of a transcript were extracted and used for functional annotations of the unigene. Tgicl (v2.0.6) was used to perform clustering and to eliminate redundant data in the assembled transcripts to obtain unique genes. A transdecoder was used to identify coding region sequences of the unigene. All assembled unigenes were compared using the public protein databases, including the NCBI non-redundant database, Swiss-Prot, and KEGG databases, using the software BLAST (v2.2.23) with a cut-off E-value of 10−5. GO annotation was performed using Blast2GO (v2.5.0) with NR annotations.
Screening of DEGs
The expression levels of the unigenes were calculated using the FPKM methods 53,54. The FPKM values of each unigene were calculated based on the length of the gene and the mapped reads count. DEGs were detected using the edgeR program package (3. 22. 5). The p value (< 0.05) threshold in multiple tests and analyses was determined using the false discovery rate (FDR). The DEGs were deemed significant according to the following criteria: p < 0.001, and the absolute value of log2 (ratio) ≥ 1. All DEGs were mapped to each term of the KEGG (KOBAS v2.0 procedure) or GO (GOSeq release2.12 procedure) databases, and significant pathways were defined based on a corrected p-value ≤ 0.05.
qRT-PCR validation
With the isolated total RNA, first-strand cDNA was synthesized using a cDNA Synthesis Kit (Takara, Japan). The actin gene (GenBank no. X63603) served as an internal control. Primer sets were designed using Primer Premier 6.0 software, and the primers are listed in Supplementary Table S4. qRT-PCR was performed using the SYBR Green Quantitative RT-qPCR kit (New England Biolab). All independent samples were analyzed in triplicates.
References
Alvarez, M. E. et al. Reactive oxygen intermediates mediate a systemic signal network in the establishment of plant immunity. Cell 92, 773–784 (1998).
Moralejo, E. et al. Multiple alien Phytophthora taxa discovered on diseased ornamental plants in Spain. Plant Pathol. 58, 100–110 (2009).
Mammella, M. A. et al. Analyses of the population structure in a global collection of Phytophthora nicotianae isolates inferred from mitochondrial and nuclear DNA sequences. Phytopathology 103, 610–622 (2013).
Johnson, E. S. et al. Origin of the black shank resistance gene, Ph, in tobacco cultivar coker 371-Gold. Plant Dis. 86, 1080–1084 (2002).
Sullivan, M. J., Melton, T. A. & Shew, H. D. Managing the race structure of Phytophthora parasitica var. nicotianae with cultivar rotation. Plant Dis. 89, 1285–1294 (2005).
Biasi, A. et al. Genetic analysis of Phytophthora nicotianae populations from different hosts using microsatellite markers. Phytopathology 106, 1006–1014 (2016).
Kamoun, S. et al. The top 10 oomycete pathogens in molecular plant pathology. Mol. Plant Pathol. 16, 413–434 (2015).
Franck, P. et al. Phytophthora nicotianae diseases worldwide: new knowledge of a long-recognized pathogen. Phytopathol. Mediterr. 55, 20–40 (2016).
Csinos, A. S. Stem and root resistance to tobacco black shank. Plant Dis. 83, 777–780 (1999).
Vontimitta, V. & Lewis, R. S. Mapping of quantitative trait loci affecting resistance to Phytophthora nicotianae in tobacco (Nicotiana tabacum L.) line Beinhart-1000. Mol. Breed. 29, 89–98 (2012).
McCorkle, K. L. et al. Characterization of Phytophthora nicotianae resistance conferred by the introgressed Nicotiana rustica Region, Wz, in flue-cured tobacco. Plant Dis. 102, 309–317 (2018).
Jones, J. D. G. & Dangl, J. L. The plant immune system. Nature 444, 323–329 (2006).
Chris, C. N. S. & Frank, L. W. T. Susceptibility genes 101: how to be a good host. Annu. Rev. Phytopathol. 52, 551–581 (2014).
Pavan, S. et al. Loss of susceptibility as a novel breeding strategy for durable and broad-spectrum resistance. Mol. Breed. 25, 1–12 (2010).
Zaidi, S. S. E. A., Mukhtar, M. S. & Mansoor, S. Genome editing: targeting susceptibility genes for plant disease resistance. Trends Biotechnol. 36, 898–906 (2018).
Yusheng, Z. et al. Identification of stably expressed QTL for resistance to black shank disease in tobacco (Nicotiana tabacum L.) line Beinhart 1000-1. Crop J. 6, 282–290 (2018).
Hui, L. et al. Genomes and virulence difference between two physiological races of Phytophthora nicotianae. GigaScience 5, 1–8 (2016).
Islam, M. T. et al. Transcriptome analysis, using RNA-Seq of Lomandra longifolia roots infected with Phytophthora cinnamomi reveals the complexity of the resistance response. Plant Biol. 20, 130–142 (2018).
Shen, D. Y. et al. Comparative RNA-Seq analysis of Nicotiana benthamiana in response to Phytophthora parasitica infection. Plant Growth Regul. 80, 59–67 (2016).
McNeil, M. D. et al. Analysis of the resistance mechanisms in sugarcane during Sporisorium scitamineum infection using RNA-seq and microscopy. PLoS ONE 13, 1–28 (2018).
Zheng, Z. et al. Loss of function in Mlo orthologs reduces susceptibility of pepper and tomato to powdery mildew disease caused by Leveillula taurica. PLoS ONE 8, 1–14 (2013).
Bezrutczyk, M. et al. Sugar flux and signaling in plant-microbe interactions. Plant J. 93, 675–685 (2018).
Jin, J., Hewezi, T. & Baum, T. J. The Arabidopsis bHLH25 and bHLH27 transcription factors contribute to susceptibility to the cyst nematode Heterodera schachtii. Plant J. 65, 319–328 (2011).
Mackey, D. et al. RIN4 interacts with Pseudomonas syringae type III effector molecules and is required for RPM1-mediated resistance in Arabidopsis. Cell 108, 743–754 (2002).
Cooley, M. B. et al. Members of the Arabidopsis HRT/RPP8 family of resistance genes confer resistance to both viral and oomycete pathogens. Plant Cell. 12, 663–676 (2000).
Breen, S. et al. Emerging insights into the functions of pathogenesis-related protein 1. Trends Plant Sci. 22, 871–879 (2017).
Du, L. et al. Ca(2+)/calmodulin regulates salicylic-acid-mediated plant immunity. Nature 457, 1154–1158 (2009).
Cheng, S. H. et al. Calcium signaling through protein kinases. The Arabidopsis calcium-dependent protein kinase gene family. Plant Physiol. 129, 469–485 (2002).
Romeis, T., Piedras, P. & Jones, J. D. Resistance gene-dependent activation of a calcium-dependent protein kinase in the plant defense response. Plant Cell. 12, 803–816 (2000).
Astegno, A. et al. Arabidopsis calmodulin-like protein CML36 is a calcium (Ca2+) sensor that interacts with the plasma membrane Ca2+-ATPase isoform ACA8 and stimulates its activity. J. Biol. Chem. 292, 15049–15061 (2017).
Lu, L. et al. TaCML36, a wheat calmodulin-like protein, positively participates in an immune response to Rhizoctonia cerealis. Crop J. 7, 608–618 (2019).
Lapin, D., Bhandari, D. D. & Parker, J. E. Origins and immunity networking functions of EDS1 family proteins. Annu. Rev. Phytopathol. 58, 1–24 (2020).
Du, Y. et al. Immune activation mediated by the late blight resistance protein R1 requires nuclear localization of R1 and the effector AVR1. New Phytol. 207, 735–747 (2015).
Du, Y. et al. Phytophthora infestans RXLR effector AVR1 interacts with exocyst component sec5 to manipulate plant immunity. Plant Physiol. 169, 1975–1990 (2015).
Gilroy, E. M. et al. CMPG1-dependent cell death follows perception of diverse pathogen elicitors at the host plasma membrane and is suppressed by Phytophthora infestans RXLR effector AVR3a. New Phytol. 190, 653–666 (2011).
Turnbull, D. et al. RXLR effector AVR2 up-regulates a brassinosteroid-responsive bHLH transcription factor to suppress immunity. Plant Physiol. 174, 356–369 (2017).
Wang, X. et al. A host KH RNA-binding protein is a susceptibility factor targeted by an RXLR effector to promote late blight disease. Mol. Plant. 8, 1385–1395 (2015).
Boevink, P. C. et al. A Phytophthora infestans RXLR effector targets plant PP1c isoforms that promote late blight disease. Nat. Commun. 7, 1–14 (2016).
Huang, G. et al. An RXLR Effector secreted by Phytophthora parasitica is a virulence factor and triggers cell death in various plants. Mol. Plant Pathol. 1, 1–16 (2018).
Nie, H. et al. SR1, a Calmodulin-binding transcription factor, modulates plant defense and ethylene-induced senescence by directly regulating NDR1 and EIN3. Plant Physiol. 4, 1847–1859 (2012).
Vogel, J. P. et al. Mutations in PMR5 result in powdery mildew resistance and altered cell wall composition. Plant J. 40, 968–978 (2004).
Divya, C. et al. Host cell ploidy underlying the fungal feeding site is a determinant of powdery mildew growth and reproduction. Mol. Plant Microbe Interact. 26, 537–545 (2013).
Schmidt, S. M. et al. Interaction of a Blumeria graminis f. sp. hordei effector candidate with a barley ARF-GAP suggests that host vesicle trafficking is a fungal pathogenicity target. Mol. Plant Pathol. 15, 535–549 (2014).
Jurkowski, G. I. et al. Arabidopsis DND2, a second cyclic nucleotide-gated ion channel gene for which mutation causes the “defense, no death” phenotype. Mol. Plant Microbe Interact. 17, 511–520 (2004).
Huibers, R. P. et al. Powdery mildew resistance in tomato by impairment of SlPMR4 and SlDMR1. PLoS ONE 8, 1–8 (2013).
Sun, K. et al. Silencing of six susceptibility genes results in potato late blight resistance. Transgenic Res. 25, 731–742 (2016).
Pathuri, I. P. et al. Alcohol dehydrogenase 1 of barley modulates susceptibility to the parasitic fungus Blumeria graminis f. sp. hordei. J. Exp. Bot. 62, 449–457 (2011).
Xing, D. H. et al. Stress- and pathogen-induced Arabidopsis WRKY48 is a transcriptional activator that represses plant basal defense. Mol Plant. 1, 459–470 (2008).
La, C. S. et al. The Arabidopsis patatin-like protein 2 (PLP2) plays an essential role in cell death execution and differentially affects biosynthesis of oxylipins and resistance to pathogens. Mol. Plant Microbe Interact. 22, 469–481 (2009).
Curtis, R. H. et al. The Arabidopsis F-box/Kelch-repeat protein At2g44130 is upregulated in giant cells and promotes nematode susceptibility. Mol. Plant Microbe Interact. 26, 36–43 (2013).
Trujillo, M. et al. Negative regulation of PAMP-triggered immunity by an E3 ubiquitin ligase triplet in Arabidopsis. Curr Biol. 18, 1396–1401 (2008).
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 29, 644–652 (2011).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 12, 323 (2011).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Acknowledgements
This research was funded by the National Natural Science Foundation of China (31571738) and the Agricultural Science and Technology Innovation Program of China (ASTIP-TRIC01).
Author information
Authors and Affiliations
Contributions
H.M. and L.C. designed the experiments. Y.L., Y.S., G.Y., W.Y., M.S., and Z.J. performed the experiments. C.J., M.R., D.L., Q.F., and Y.W. analyzed the data. H.M., A.Y. and L.C. drafted the manuscript. All authors discussed the results and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Meng, H., Sun, M., Jiang, Z. et al. Comparative transcriptome analysis reveals resistant and susceptible genes in tobacco cultivars in response to infection by Phytophthora nicotianae. Sci Rep 11, 809 (2021). https://doi.org/10.1038/s41598-020-80280-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-80280-7
This article is cited by
-
An improved bacterial mRNA enrichment strategy in dual RNA sequencing to unveil the dynamics of plant-bacterial interactions
Plant Methods (2024)
-
Rhizosphere microbiomes derived from vermicompost alter gene expression and regulatory pathways in tomato (Solanum lycopersicum, L.)
Scientific Reports (2024)
-
Deciphering fungicide resistance in Phytophthora: mechanisms, prevalence, and sustainable management approaches
World Journal of Microbiology and Biotechnology (2024)
-
CRISPR-Cas technology secures sustainability through its applications: a review in green biotechnology
3 Biotech (2023)
-
Comparative transcriptome profiling of resistant and susceptible foxtail millet responses to Sclerospora graminicola infection
BMC Plant Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.