Abstract
Gastrointestinal stromal tumours that are wild type for KIT and PDGFRA are referred to as WT GISTs. Of these tumours, SDH-deficient (characterized by the loss of SDHB) and quadruple WT GIST (KIT/PDGFRA/SDH/RAS-P WT) subgroups were reported to display a marked overexpression of FGF4, identifying a putative common therapeutic target for the first time. In SDH-deficient GISTs, methylation of an FGF insulator region was found to be responsible for the induction of FGF4 expression. In quadruple WT, recurrent focal duplication of FGF3/FGF4 was reported; however, how it induced FGF4 expression was not investigated. To assess whether overexpression of FGF4 in quadruple WT could be driven by similar epigenetic mechanisms as in SDH-deficient GISTs, we performed global and locus-specific (on FGF4 and FGF insulator) methylation analyses. However, no epigenetic alterations were detected. Conversely, we demonstrated that in quadruple WT GISTs, FGF4 expression and the structure of the duplication were intimately connected, with the copy of FGF4 closer to the ANO1 super-enhancer being preferentially expressed. In conclusion, we demonstrated that in quadruple WT GISTs, FGF4 overexpression is not due to an epigenetic mechanism but rather to the specific genomic structure of the duplication. Even if FGF4 overexpression is driven by different molecular mechanisms, these findings support an increasing biologic relevance of the FGFR pathway in WT GISTs, both in SDH-deficient and quadruple WT GISTs, suggesting that it may be a common therapeutic target.
Similar content being viewed by others
Introduction
Approximately 10–15% of gastrointestinal stromal tumours (GISTs) do not harbour mutations in KIT or PDGFRA and are referred to as KIT/PDGFRA wild-type (WT)1. In general, WT GISTs are more resistant to imatinib, and a proper therapeutic strategy is still lacking. Of these cases, 20–40% are characterized by the loss of SDHB (SDH-deficient GIST)2,3,4, approximately 15% carry BRAF/RAS or NF1 mutations, and the remaining cases are designated as KIT/PDGFRA/SDH/RAS-P WT GISTs (or quadruple WT GISTs) in which the molecular driver alterations are still unknown.
In recent years, many efforts have been made to characterize quadruple WT GISTs; however, whole-exome sequencing uncovered considerable molecular heterogeneity in this subgroup of patients despite a homogeneous gene expression profile5. Nonetheless, alterations in FGFR1 (point mutations and fusion genes) and FGF4 (duplication and overexpression) have been detected in several cases, supporting the idea that the FGFR pathway could play a relevant role in the biology of quadruple WT GISTs5,6,7. Interestingly, overexpression of FGF4 and activation of its signalling were recently reported in SDH-deficient GISTs8. In these cases, the hypermethylator phenotype of SDH-deficient GISTs9 was associated with the methylation of an insulator region (referred to as the “FGF insulator”) located in the upstream region of FGF4 (between FGF3 and AP003555.2). This epigenetic alteration caused genome topology changes, allowing ANO1 (which has super-enhancer activity) to induce the expression of the FGF4 oncogene8. FGFR1 was found to be highly expressed in GISTs7,8,10,11, and activation of the downstream signalling of FGFR, through AKT or MAPK, was demonstrated both in quadruple WT and SDH-deficient GISTs, supporting the presence of an autocrine loop between FGF4 and FGFR17,8.
Together, these data highlight the involvement of FGF4 in the biology of GISTs that do not rely on KIT or PDGFRA. However, how the FGF4 duplication detected in quadruple WT could induce FGF4 expression was not investigated. Thus, in the present work, we studied the methylation status and the structure of FGF4 duplication in quadruple WT GISTs to assess whether epigenetic changes occurred and whether they could be involved in the regulation of FGF4 expression.
Material and methods
Tumour samples
Tissue samples from GIST patients (fresh frozen or formalin-fixed paraffin-embedded already centralized at “Giorgio Prodi” Cancer Research Center (CIRC), University of Bologna) were studied. Diagnosis of GIST was based on histologic evaluation and immunohistochemistry of CD117 and DOG1. SDH deficiency was checked using IHC of SDHB. Quadruple WT cases were defined as being negative for mutations in the KIT, PDGFRA, SDHA-B-C-D, and RAS-P genes, which was confirmed by Sanger sequencing and next generation sequencing (NGS)7. Six quadruple WT cases positive for FGF4 duplication were identified7 and used in the present study. Clinically, tumours were localized in the duodenum in four cases and in the ileum in two cases: two females and four males, with a median age of 60 years (range 44–73). From a molecular point of view, the presence of relevant gene fusions or somatic mutations was excluded through whole-exome and whole-transcriptome sequencing, with the exception of two patients in which mutations on tumour suppressor genes were identified: GIST127 carried a mutation in CTNND2, and GIST320 had MEN1 and TP53 mutations5.
As a cohort for comparison, we used GIST samples already present in our tissue bank and for which molecular data useful for this study were available: 26 SDH-deficient samples (15 mutated in SDHA, 4 in SDHB and 7 in SDHC) and 17 KIT mutant samples (10 mutated in exon 11 and 7 mutated in exon 9) and two quadruple WT GIST samples negative for FGF4 duplication (described in7). The tumour samples analysed were all from primary tumours. This study was approved by the local institutional ethical committee of Azienda Ospedaliero-Universitaria Policlinico S. Orsola-Malpighi (number 113/2008/U/Tess). All experiments were performed in accordance with relevant guidelines/regulations, and informed consent was obtained from all participants.
Global methylation profile
The global methylation profile was evaluated in 4 quadruple WT, 4 SDH-deficient (3 SDHA mutant and 1 SDHC epimutant) and 4 KIT mutant GISTs (exon 11 mutated). To correct for the different tumour localization of SDH-deficient and quadruple WT cases, KIT mutant cohort included both stomach and intestinal tumours. Genomic DNA was analysed using the Infinium Human Methylation 450 BeadChip (Illumina, San Diego, CA) following the manufacturer’s instructions. Data analysis was performed by adopting the approach described by Bacalini et al. (2015)12. Briefly, the Infinium 450k annotation was used to group the CpG probes on the same island/shelf/shore associated with the genes. For each gene, we evaluated the methylation changes by applying a multivariate analysis of variance (MANOVA) correcting for multiple hypothesis comparisons with Benjamini–Hochberg correction.
Target methylation analysis
CpG methylation at specific loci was assessed through bisulfite conversion and enzymatic digestion on the six quadruple WT samples positive for FGF4 duplication and on an expanded cohort of other molecular subgroups of GIST: 26 SDH-deficient cases, 17 KIT mutant cases, and two quadruple WT GISTs negative for FGF4 duplication. For bisulfite conversion, 200 ng of DNA was converted using the Methylamp DNA Modification Kit (Epigentek, Farmingdale, NY) following the manufacturer’s instructions. Then, 3 µl of converted DNA was amplified with primers specific for the FGF insulator region located at chr11:69,918,992–69,919,031 (kindly provided by Dr William Flavahan, Harvard Medical School, Boston, MA, USA). Amplicons were then sequenced using the BigDye Terminator v1.1 Cycle Sequencing Kit for Sanger sequencing (Thermo Fisher Scientific).
For enzymatic digestion, 100 ng of DNA was digested using the EpiJET DNA Methylation Analysis Kit (MspI/HpaII) (Thermo Fisher Scientific, Monza, Italy). In brief, DNA was digested with the MspI and HpaII restriction enzymes, two isoschizomers with differing sensitivities to CpG methylation. When CpG regions are methylated, cleavage with Epi HpaII is blocked, but cleavage with Epi MspI is not affected. HpaII is able to cleave only unmethylated DNA, while MspI cleaves both methylated and unmethylated DNA. qPCR using primers specific for the FGF insulator was performed on undigested (Cq1), HpaII digested (Cq2) or MspI digested (Cq3) DNA. The primers used were FW 5′-AATGTCCCCTGCACATGGAG-3′ and RV 5′-GCCGCGTCTCTCACATTTTC-3′. All samples were completely digested with the control enzyme MspI (threshold cycles Cq3 – Cq1 > 4.5) and were used for the analysis. The level of DNA methylation (%) was estimated using the formula: % of 5-mC = 100/(1 + E) Cq2 – Cq1, where E was the PCR efficiency value.
Copy number
Copy number assessments were performed using TaqMan copy number assays (Thermo Fisher Scientific). Evaluation of the FGF insulator copy number was performed using Hs06280902_cn and Hs06332446_cn FAM‐labelled probes, covering the upstream and downstream regions of the insulator, respectively. The FGF4 copy number was evaluated as previously described7. Hs03800758_cn and Hs03782780_cn FAM‐labelled probes targeting ENDOD1 and XXRA1 located outside FGF4 duplication margins, respectively, were used for copy number normalization. TaqMan RNaseP Control Reagent (VIC‐labelled; Thermo Fisher Scientific) was used as an internal reference control. The copy number was estimated using the DDCt method and using a normal diploid sample as the calibrator. Moreover, high-density copy number array data (CytoScan HD or Oncoscan CNV Plus, Thermo Fisher Scientific) of quadruple WT GISTs were obtained from our previously published paper7.
Sanger sequencing
Ten nanograms of DNA was used for amplification and sequencing of regions of interest: the FGF insulator locus and two SNPs located inside the UTR of FGF4 (rs3168175 in the 3′UTR and rs9666584 in the 5′UTR). For fusion gene validation, 500 ng of RNA was reverse-transcribed with SuperScript IV Reverse Transcriptase (Thermo Fisher Scientific) using random primers. Then, cDNA was used for amplification of fusion gene breakpoint regions or for the evaluation of FGF4-UTR SNP expression. The primers used are listed in Suppl. Table 1. Amplified regions were purified and sequenced using the BigDye Terminator 1.1 Cycle Sequencing Kit on the ABI 3730 Genetic Analyzer (Applied Biosystems).
Gene expression
Raw RNA sequencing data were obtained from our previously published manuscripts5,7. Reads were aligned using the TopHat/BowTie pipeline. TPM normalization was used to evaluate gene expression levels. Fusion gene prediction was performed on RNA-seq data of six quadruple WT GISTs using Defuse, ChimeraScan and FusionMap software.
Results
Methylation analysis of quadruple WT GISTs
Overexpression of FGF4 was reported in both quadruple WT and SDH-deficient GISTs7,8. Since it was shown that the hypermethylator phenotype of SDH-deficient GISTs was responsible for FGF4 transcriptional activation8, we investigated for the first time the global DNA methylation profile of quadruple WT GISTs to uncover whether overexpression of FGF4 could be regulated by the same epigenetic mechanism as SDH-deficient cases.
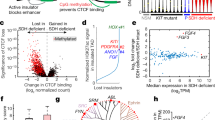
Global methylation analysis was performed on quadruple WT GISTs in comparison with SDH-deficient and KIT mutant cases. Quadruple WT cases showed a global methylation level similar to that of the KIT mutant, while SDH-deficient GISTs had significantly higher genomic methylation. This result indicated that the hypermethylator phenotype characteristic of SDH-deficient GISTs was not present in quadruple WT cases (Fig. 1A). Moreover, focusing on the FGF4 locus, we showed that neither gene methylation was responsible for FGF4 upregulation in quadruple WT GIST. In fact, CpG dinucleotides located inside the FGF4 gene were hypomethylated in all GIST subgroups independent of FGF4 gene expression, which was higher in SDH-deficient and quadruple WT GISTs and was absent in KIT mutant GISTs (Fig. 1B). This result indicated that in these tumours, the CpG methylation status of the FGF4 locus was not responsible for its transcriptional activation. Moreover, as previously demonstrated7, FGF4 expression seemed to be higher in quadruple WT cases than in SDH-deficient cases, suggesting that different activation mechanisms subtended FGF4 induction in quadruple WT GIST.
CpG methylation in quadruple WT and SDH-deficient GISTs. (A) Violin plot showing the B-value of global CpG methylation evaluated using the Infinium Human Methylation 450 BeadChip in GISTs. (B) Bar plot showing the absence of correlation between FGF4 CpG methylation and FGF4 mRNA expression. The B-value of the CpG island located at the FGF4 locus is shown in the left panel, and the gene expression level (log2TPM) of FGF4 is shown on the right. The t-test statistic was used (* < 0.05; *0.01). (C) Methylation level of the FGF insulator region evaluated through comparative enzymatic digestion in an extended cohort of GISTs (6 FGF4-positive quadruple WT, 26 SDH-deficient and 17 KIT mutant). (D) Box plot showing the copy number of the FGF insulator region (the two TaqMan assays, covering upstream and downstream regions of the insulator, were performed) of quadruple WT in comparison with KIT mutant and SDH-deficient GISTs. In quadruple WT cases, the FGF insulator was duplicated (three copies), which is similar to FGF4.
Then, we explored the epigenetic status of the so-called FGF insulator described by Flavahan et al., whose hypermethylation in SDH-deficient GISTs causes the loss of functional chromatin boundaries, thus allowing the adjacent ANO1 super-enhancer to contact and activate FGF48. We confirmed in an extended cohort that the FGF insulator was methylated specifically in SDH-deficient with respect to other GIST subgroups (Fig. 1C), thus reinforcing the role of SDH loss in the alteration of the epigenetic status of FGF4 in GIST. In contrast, it was fully unmethylated in both FGF4-positive quadruple WT and KIT mutant GISTs (Fig. 1C and Suppl. Figure 1), indicating that the methylation status of the FGF insulator was not the driver of the remarkably high expression of FGF4 in these quadruple WT cases. Of note, the analysis was also performed on two additional quadruple WT GISTs negative for FGF4 duplication and for FGF4 expression (described in7), confirming the absence of methylation of the FGF insulator in this setting (data not shown). Since it was also demonstrated that disruption of this insulator by gene editing induces overexpression of FGF48, we further assessed the integrity of the FGF insulator in quadruple WT cases in addition to methylation status. Through direct sequencing, we excluded the presence of mutations inside this region (not shown), and through copy number assay, we confirmed that the region was duplicated along with FGF4 (Fig. 1D).
Altogether, these data reinforced the role of SDH loss in the alteration of the epigenetic status of FGF4 in GISTs, while they excluded the possibility that FGF4 overexpression in quadruple WT cases could be due to an epigenetic loss of the FGF insulator region. This result raised the question as to whether the genomic structure of the duplication could be responsible for FGF4 transcriptional activation.
Correlation between FGF4 duplication structure and FGF4 expression
To understand the molecular mechanism underlying the high transcriptional level of FGF4 in quadruple WT GISTs, we first dissected the genomic structure and location of the duplicated FGF4 allele.
First, by exploiting previously published RNA-seq and copy number variation (CNV) information7, we investigated the genomic position of the duplicated copy of the FGF4 gene. Fusion transcripts were predicted starting from RNA-seq data of the whole series of FGF4-positive quadruple WT GISTs, and RT-PCR was used for validation. Supplementary Table 2 shows all the predicted fusion transcripts involving chromosome 11. In two samples, GIST133 and GIST219, no fusions were detected. In the remaining samples, several fusions were identified, among which intrachromosomal rearrangements involving q13 cytobands of chromosome 11 were predominant, further indicating the high instability of the genomic region surrounding FGF4 in quadruple WT GIST. GIST127 and GIST320 showed several consecutive structural alterations on chromosome 11 that were responsible for the majority of fusion genes identified in these samples (Suppl. Table 3). However, due to the complexity of the region and the absence of fusion genes directly affecting FGF4, it was not possible to properly dissect the structure of FGF4 in these samples. Conversely, GIST400 and GIST401 harboured only one duplicated segment in chr11, the one encompassing FGF4. Moreover, two fusion genes involving the terminal untranslated region (3′UTR) of FGF4 were detected in these two tumours: FGF4-3UTR was fused with the intergenic region upstream of CCND1 in GIST401 and with the upstream region of AP003555.2 in GIST400. Breakpoints of these fusions corresponded approximatively to the boundaries of the duplication events detected (Suppl. Table 3, Fig. 2A), indicating that these fusion genes were produced by the insertion of the duplicated copy of FGF4. Thus, it was possible to demonstrate that the duplicated allele was integrated in the same chromosome and in close proximity to the parental copy of FGF4. Therefore, in these two GISTs, by combining information from RNA sequencing and CNV analysis, we can describe the structure of the duplication, in which FGF4 is either inserted in tandem in the intergenic region between FGF3 and AP003555.2 (GIST_400) or is fused with the inverted upstream region of CCND1 (GIST_401) (Fig. 2A).
Structure of FGF4 duplication and monoallelic expression of FGF4 mRNA. (A) Schematic representation of the structure of FGF4 duplication in two quadruple WT cases. In GIST400 (left panel), the genomic region between FGF4-3UTR and the upstream region of AP003555.2 was duplicated in tandem, producing the fusion gene FGF4-3UTR/AP003555.2-upstr. In GIST401 (right panel), the genomic region between CCND1 and AP003555.2 was duplicated, inverted and inserted into the 3′UTR of FGF4, producing the fusion gene FGF4-3UTR/CCND1-upstr. Duplicated regions are shown in violet. Genes are indicated by yellow arrows, depending on their transcriptional direction. The FGF4 copy involved in the fusion is shown in red. Breakpoint sequence and position on the genome are shown for each fusion: sequences corresponding to FGF4-3UTR are shown in red (+ or − marks indicated if the sequence maps on the positive or negative strand). (B) Plot (in the left) showing the coverage per base of FGF4 mRNA in the two quadruple WT cases carrying FGF4-3UTR fusion. The FGF4 region retained in the fusion gene is highlighted in yellow. As a comparison, coverage of a quadruple WT without fusion (GIST133) is shown (lower panel). A relevant drop in coverage was detected in the cases carrying FGF4-3UTR fusion after the breakpoint of each case (indicated by a black bar). On the right, a box plot shows the coverage depth of exon 3 in FGF4 before and after the fusion breakpoint. (C) Bar plot showing the allelic fraction (%) of two SNPs located at the FGF4 locus (specifically rs3168175 in the 3′UTR and rs9666584 in the 5′UTR) at the DNA and mRNA levels in two quadruple WT GISTs (GIST127 and GIST133), indicating that FGF4 expression is monoallelic.
Finally, we investigated whether FGF4 expression was uniformly upregulated or whether its overexpression was due to a single rearranged allele. Interestingly, we discovered that only one of the three copies of FGF4 was predominantly expressed. In the two quadruple WT GISTs harbouring FGF4-3UTR fusion genes, we observed that the depth of RNA-seq reads over FGF4 had a relevant drop immediately after the breakpoint region of each case, indicating that the copy involved in the fusion was preferentially transcriptionally activated (Fig. 2B). Interestingly, in both cases, the copy of FGF4 expressed and involved in the fusion was the one closer to ANO1. Similarly, among the other quadruple WT samples, it was possible to identify two informative heterozygous SNPs at the DNA level located in UTR regions of FGF4 that were monoallelically expressed in two cases (GIST127 and GIST133) (Fig. 2C), indicating that FGF4 expression was derived from only one copy of FGF4 also in these tumours. Globally, these findings indicated that FGF4 overexpression detected in quadruple WT is a distinctive and recurrent feature that was not due to an epigenetic mechanism but rather to the specific genomic structure of the 11q13 duplication that induces the transcriptional activation of one allele of FGF4.
Discussion
FGFR1 is one of the most highly expressed tyrosine kinase receptors in GISTs after KIT and PDGFRA7,8,10,11. It has been reported that quadruple WT GISTs, which do not rely on KIT or PDGFRA, may present FGFR1 activating mutations and fusion events5,6. Recently, in two different papers, overexpression of an FGFR1 ligand, FGF4, has been reported as a novel molecular event in two subgroups of GISTs without KIT and PDGFRA mutations, both SDH-deficient and quadruple WT GISTs. Moreover, activation of FGFR downstream signalling pathways through AKT and MAPK was demonstrated7,8.
In particular, Flavahan et al. reported that in SDH-deficient GISTs, the hypermethylator phenotype induced by SDH loss9 disrupts the binding of CTCF in regions located in proximity to the FGF3/FGF4 locus, causing FGF4 overexpression8. Generally, CTCF binding sites demarcate the boundaries of topologically associated domains (TADs), which are contiguous genomic intervals in which the majority of loci interact more frequently with each other than with loci outside the TAD. Thus, the disruption of these CTCF binding sites inactivates their insulator function, allowing contact with the adjacent TAD13. The CTCF binding site identified by Flavahan et al. was demonstrated to have an insulator function on FGF3 and FGF4, and its hypermethylation allowed contact between the adjacent ANO1 super-enhancer and FGF genes, causing the overexpression of FGF48.
Conversely, we had previously reported that FGF4 was recurrently overexpressed in quadruple WT GISTs in association with a focal duplication encompassing the gene. Duplication and overexpression of FGF4 were mutually exclusive with other relevant alterations, and they were not detected in GISTs harbouring FGFR1 mutations, supporting the pathogenic role of the event7. In our case series of quadruple WT GISTs, we investigated the epigenetic mechanisms modulating FGF4 expression identified in SDH-deficient GISTs8, and we demonstrated that aberrant methylation was not responsible for FGF4 overexpression in quadruple WT cases. Moreover, we found that FGF4 expression was derived from the duplicated allele, preferentially from the copy nearest to ANO1. This suggests that the mechanism of upregulation of FGF4 in quadruple WT GISTs is related to the genomic structure alteration caused by the duplication. One hypothesis is that the insertion of the duplicated allele in the proximity of FGF4 could alter the chromatin conformation, allowing FGF4 transcription to be induced by the super-enhancer activity of ANO18. It has been shown that TAD boundaries are altered in a cancer-specific manner for each tumour type, presumably reflecting the distribution of cancer driver genes14. In cancer, duplication events frequently extend over the neighbouring TAD (inter-TAD), resulting in the formation of new chromatin domains (neo-TAD) that could alter the expression level of the involved genes14,15,16. Since the duplicated region identified in quadruple WT GISTs is involved at least two neighbouring TADs8, it is possible that the resulting neo-TAD formation allowed for the interaction with the ANO1 super-enhancer inducing the aberrant expression of FGF4. Further studies are needed to confirm this hypothesis; however, the extreme rarity of these tumours, the extensive molecular characterization already performed and the lack of appropriate cellular models hamper the development of deeper analyses on this aspect.
Even if the mechanism underlying FGF4 upregulation seems to be different between SDH-deficient and quadruple WT GISTs, a common altered signalling pathway could be identified, and the therapeutic consequences for patients could be similar. Taken together, our findings indicated that FGFR pathway alteration is a common oncogenic event in these two groups of KIT/PDGFRA WT GISTs, and targeted therapeutic approaches should be considered10. In vitro models of KIT/PDGFRA WT GISTs are still lacking, but Flavahan et al. developed an SDH-deficient GIST PDX model carrying epigenetic alterations and consequent overexpression of FGF4 and demonstrated that it was responsive to BGJ-398, a selective inhibitor of FGFR1–4. This study further strengthened the biological relevance of FGFR signalling in the tumorigenesis of GISTs not dependent on KIT and PDGFRA, indicating a potential therapeutic option for these tumours8. Currently, no specific FGFR inhibitors are approved for GISTs. Regorafenib, which was approved in GISTs resistant to imatinib and sunitinib, is the only available drug that may also have a weak potential inhibition of FGFR. In fact, in vitro studies demonstrated that regorafenib also inhibited FGFR signalling in MCF‐7 breast cancer by inhibiting phosphorylated FGFR substrate 2 (pFRS2) and the downstream signalling kinase pERK1/217. SDH-deficient GISTs were shown to obtain durable benefits from regofarenib treatment, but this positive result probably depends more on the inhibition of the KIT pathway and angiogenic kinases (VEGFR1/3)18. The ongoing clinical trial REGISTRI (NCT02638766) is evaluating whether WT KIT/PDGFRA may benefit from regorafenib upfront in first-line therapy; however, at present, no definitive conclusion can be drawn due to the slow accrual. In addition to regorafenib, in the last decade, other multi-TKIs active against FGFR (e.g., dovitinib, masitinib, ponatinib and pazopanib) have also been tested in GISTs, and some of them have produced encouraging results10,19,20. However, all these trials were predominantly conducted on mutant KIT GISTs, and no conclusion can be drawn about the efficacy of these multi-TKIs in rarer mutational subsets of GISTs.
In conclusion, we demonstrated the presence of a novel mechanism of FGF4 regulation in quadruple WT GISTs. Even if the genetic alteration is different, overexpression of the FGF4 oncogene is a common event in both SDH-deficient and quadruple WT GISTs, indicating for the first time a putative common therapeutic target for GISTs that do not rely on KIT/PDGFRA/RAS-P activation. Clinical experience or trials specific for KIT/PDGFRA/RAS-P WT GISTs using FGFR inhibitors should be encouraged.
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- CNV:
-
Copy number variation
- GIST:
-
Gastrointestinal stromal tumours
- TAD:
-
Topologically associated domains
- WT:
-
Wild-type
References
Corless, C. L., Fletcher, J. A. & Heinrich, M. C. Biology of gastrointestinal stromal tumors. J. Clin. Oncol. 22, 3813–3825 (2004).
Miettinen, M. et al. Succinate dehydrogenase-deficient GISTs. Am. J. Surg. Pathol. 35, 1712–1721 (2011).
Gill, A. J. et al. Immunohistochemistry for SDHB divides gastrointestinal stromal tumors (GISTs) into 2 distinct types. Am. J. Surg. Pathol. 34, 636–644 (2010).
Boikos, S. A. et al. Molecular subtypes of KIT/PDGFRA wild-type gastrointestinal stromal tumors. JAMA Oncol. 2, 922–928 (2016).
Pantaleo, M. A. et al. Genome-wide analysis identifies MEN1 and MAX mutations and a neuroendocrine-like molecular heterogeneity in Quadruple WT GIST. Mol. Cancer Res. 15, 553–562 (2017).
Shi, E. et al. FGFR1 and NTRK3 actionable alterations in “wild-type” gastrointestinal stromal tumors. J. Transl. Med. 14, 339 (2016).
Urbini, M. et al. Gain of FGF4 is a frequent event in KIT/PDGFRA/SDH/RAS-P WT GIST. Genes Chromosomes Cancer 58, 636–642 (2019).
Flavahan, W. A. et al. Altered chromosomal topology drives oncogenic programs in SDH-deficient GISTs. Nature 575, 229–233 (2019).
Killian, J. K. et al. Succinate dehydrogenase mutation underlies global epigenomic divergence in gastrointestinal stromal tumor. Cancer Discov. 3(6), 648–657 (2013).
Astolfi, A., Pantaleo, M. A., Indio, V., Urbini, M. & Nannini, M. The emerging role of the FGF/FGFR pathway in gastrointestinal stromal tumor. Int. J. Mol. Sci. 21, 3313 (2020).
Fang, L. et al. FGFR-mediated reactivation of MAPK signaling attenuates antitumor effects of imatinib in gastrointestinal stromal tumors. Cancer Discov. 5, 438–451 (2015).
Bacalini, M. G. et al. A meta-analysis on age-associated changes in blood DNA methylation: Results from an original analysis pipeline for Infinium 450k data. Aging (Albany NY). 7, 97–109 (2015).
Dixon, J. R., Gorkin, D. U. & Ren, B. Chromatin domains: The unit of chromosome organization. Mol. Cell 62, 668–680 (2016).
Akdemir, K. C. et al. Disruption of chromatin folding domains by somatic genomic rearrangements in human cancer. Nat. Genet. 52, 294–305 (2020).
Franke, M. et al. Formation of new chromatin domains determines pathogenicity of genomic duplications. Nature 538, 265–269 (2016).
Lupiáñez, D. G. et al. Disruptions of topological chromatin domains cause pathogenic rewiring of gene-enhancer interactions. Cell 161, 1012–1025 (2015).
Wilhelm, S. M. et al. Regorafenib (BAY 73-4506): A new oral multikinase inhibitor of angiogenic, stromal and oncogenic receptor tyrosine kinases with potent preclinical antitumor activity. Int. J. Cancer. 129, 245–255 (2011).
Ben-Ami, E. et al. Long-term follow-up results of the multicenter phase II trial of regorafenib in patients with metastatic and/or unresectable GI stromal tumor after failure of standard tyrosine kinase inhibitor therapy. Ann Oncol. 27, 1794–1799 (2016).
Joensuu, H. et al. Dovitinib in patients with gastrointestinal stromal tumour refractory and/or intolerant to Imatinib. Br. J. Cancer 117, 1278–1285 (2017).
Adenis, A. et al. Masitinib in advanced gastrointestinal stromal tumor (GIST) after failure of imatinib: A randomized controlled open-label trial. Ann. Oncol. 25, 1762–1769 (2014).
Acknowledgements
Special thanks to Annalisa Altimari, Francesco Buia, Paolo Castellucci, Massimo Cervellera, Claudio Ceccarelli, Matteo Cescon, Antonietta D’Errico, Antonio De Leo, Valerio Di Scioscio, Stefano Fanti, Michelangelo Fiorentino, Elena Fumagalli, Giovanni Grignani, Ludovica Murrone, Fabio Niro, Maria Giulia Pirini, Nico Pagano, Donatella Santini, Maristella Saponara, Valeria Tonini, and Bruno Vincenzi.
Funding
This study was possible thanks to the research contribution received in memory of Alberto Arenghi, Caravaggio (BG), Italy, and was supported by Petra S.r.L.
Author information
Authors and Affiliations
Contributions
M.U., M.A.P. and A.A. conceived and designed the work and drafted the manuscript. M.N., M.D.G., P.S., P.U., G.R. and S.A. participated in data analysis and interpretation. V.I. performed bioinformatics analysis. G.C. and M.G.B. performed global methylation analysis. M.U., E.G., A.S. and G.R. performed molecular analyses and validations. M.N., V.I., S.A., M.D.G., P.S. and P.U. helped with drafting the manuscript. All authors revised and approved the final version of the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Urbini, M., Astolfi, A., Indio, V. et al. Gene duplication, rather than epigenetic changes, drives FGF4 overexpression in KIT/PDGFRA/SDH/RAS-P WT GIST. Sci Rep 10, 19829 (2020). https://doi.org/10.1038/s41598-020-76519-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-76519-y
This article is cited by
-
New treatment strategies for advanced-stage gastrointestinal stromal tumours
Nature Reviews Clinical Oncology (2022)
-
Concurrent inhibition of CDK2 adds to the anti-tumour activity of CDK4/6 inhibition in GIST
British Journal of Cancer (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.