Abstract
Several species of crustose coralline algae (CCA) and their associated microbial biofilms play important roles in determining the settlement location of scleractinian corals on tropical reefs. In recent decades, peyssonnelid algal crusts (PAC) have become spatial dominants across large areas of shallow Caribbean reefs, where they appear to deter the recruitment of scleractinians. Our genetic investigations of PAC in St. John, US Virgin Islands, amplifying the large-subunit ribosomal RNA and psbA protein D1 marker genes, revealed them to be identical to Ramicrusta textilis previously reported overgrowing corals in Jamaica. Specimens of PAC sampled from the Honduras were likewise identical, confirming that this crustose alga inhabits the easternmost and westernmost regions of the Caribbean. We also analysed 16S rDNA tag amplicon libraries of the biofilms associated with PAC and sympatric CCA, which is favoured for coral settlement. Our results show that the microbial communities on PAC (vs. CCA) are characterized by significantly lower numbers of the epibiotic bacterial genus Pseudoalteromonas, which facilitates the recruitment and settlement of marine invertebrates. From these data, we infer that PAC are therefore unlikely to be attractive as settlement sites for coral larvae. Given the significant ecological change anticipated on these reefs due to increasing cover of PAC, there is an urgent need to further investigate competitive interactions between PAC and scleractinian corals, and elucidate the role of PAC and their associated microbiomes in accentuating phase shifts from coral to algae on tropical reefs.
Similar content being viewed by others
Introduction
As tropical reefs become increasingly degraded, many are transitioning from spatial dominance by stony corals to occupation by macroalgae1. Through competition with macroalgae for space on hard substrata, the distribution and abundance of coral recruits is restricted, as is their post-settlement success, thus favouring profound changes in coral reef community structure2,3,4. Since the 1980s, Caribbean reefs have been exposed to multiple agents driving large-scale declines in coral cover, including diseases causing mass mortality of the echinoid Diadema antillarum5 and the coral Acropora spp.6, seawater warming7 leading to global coral bleaching in 1987, 1998, 2005, and 20168,9,10,11, and major hurricanes12. In a region (i.e., the Caribbean) classically characterised by low rates of coral larval recruitment (i.e., circa 100 corals m−2)13,14,15, compared to the Indo-Pacific16,17, and challenged by overfishing18,19, many present day reefs have undergone changes in benthic community structure to favour taxa that grow faster than scleractinian corals. These include macroalgae4,20, sponges21, and octocorals22 to the detriment of scleractinian coral cover.
Recruitment of pelagic larvae to benthic surfaces by scleractinians is one of a number of processes necessary for populations of scleractinians to recover following severe disturbances14. Freshly released brooded coral larvae, as well as those developing from mass-spawned gametes, rely upon a complex sequence of habitat-specific cues at increasingly smaller scales23 to select a suitable settlement location, and then initiate attachment to, and metamorphosis on, this surface24,25. Several decisive factors are involved in settlement choices by coral larvae, the best documented of which are the physical properties of the substratum (e.g., rugosity and light microenvironment) and the presence of select species of crustose coralline algae (CCA)26,27 and their associated microbial biofilms3,28,29,30,31,32. Within these microbial biofilms, members of the bacterial genus Pseudoalteromonas, that have been isolated from tissues of CCA (including Hydrolithon onkodes and Neogoniolithon fosliei from the Great Barrier Reef and Paragoniolithon solubile from the Caribbean), are potent inducers of settlement and metamorphosis of coral larvae24,33,34. In these cases, larval settlement is induced by the secondary metabolite tetrabromopyrrole (TBP) produced by the bacteria34,35. Recent work36 has also reaffirmed the earlier view that certain CCA can directly promote coral settlement without a participatory role of bacteria37,38.
In 2010, a new threat to Caribbean reefs emerged as aggressively spreading peyssonnelid algal crusts (PAC), which were first documented in Florida and Belize39,40 and later in Jamaica41, Bonaire, Netherlands Antilles42,43, Puerto Rico44 and now in St. John, US Virgin Islands45. PAC have been reported as common on shallow coral reefs (\(<3\) m depth) in these locations, and in Bonaire for example (Lac Bay in 2010), PAC aggressively grew over vacant space, corals (e.g., example from St. John, Fig. 1), and sponges to occupy as much as 18.7 ± 20.9% of the benthos43. Rhodophyte crusts, including CCA and PAC, are common on shallow continental shelves throughout the world46, and while peyssonnelids have long been components of the algal flora of Caribbean reefs47,48,49,50, their recent appearance as spatially dominant algae on shallow reefs is novel for these locations41. It is likely that PAC are a biodiverse group composed of endemics, for example, Peyssonnelia stoechas in St. John, US Virgin Islands (R. Steneck, unpublished data), Metapeyssonnelia milleporoides in Puerto Rico44, and Ramicrusta sp. from Bonaire and Puerto Rico51,52. The genus Ramicrusta was established in 1981, based on samples described from the Paracel Islands, South China Sea53, and prior to its detection in Jamaica41, it was unknown in the Caribbean. Since 2013, PAC have encroached upon large areas of space on some shallow reefs (\(< 7\) m depth) in St. John54, with surveys at 3 m depth showing an increase in mean cover of PAC from 8.5 ± 3.6% (mean ± SE, n = 5 sites) in 201554 to 22.0 ± 9.1% (mean ± SE, n = 5 sites) in August 201745, with up to 34.5 ± 3.2% (mean ± SE, n = 39) cover at an exposed headland. Two months later, high cover of PAC persisted after two Category Five hurricanes impacted St. John in September 201745. To date, 14 weeks of fieldwork over 3 years in St. John have revealed only two coral recruits on PAC, and rare signs of fish herbivory on these crusts. The reasons underlying the novel ecological success of PAC on present-day Caribbean reefs remain unknown.
Crusts of CCA (family Corallinaceae, Rhodophyta) can promote coral recruitment through microbial flora associated with their surface33,34,35,55. As PAC have been reported to deter larval settlement and metamorphosis45,56, and different species of macroalgae harbour unique and species-specific bacterial communities32,57, here we asked whether PAC might harbour microbial consortia differing from those associated with CCA. We compared microbial communities in surface mucous (SM) from PAC (collected from igneous rocks) and also from CCA (on both igneous rocks and carbonate substrata) sampled from shallow reefs of St. John. Using high-throughput sequencing (HTS), we analysed amplicon libraries of biofilms associated with each algal crust. Using additional molecular and microscopic tools, we identified the algal species comprising PAC aggressively overgrowing shallow reefs in St. John.
Results
The samples of PAC removed from the reefs of St. John in August 2018 supported a definitive identification of the alga involved. BLASTn comparision with the NCBI database for both LSU (590bp) and psbA (410bp) gene fragments amplified from PAC genomic DNA, resulted in 100% sequence identity with voucher specimens of Ramicrusta textilis (Accession Numbers FJ848970 and KM360015, respectively) sampled from the coastal waters of Jamaica in 200341. Low resolution (\(\times\) 20) microscopy of radial vertical sections of the PAC (Fig. 2), showing the well-defined tabular layer of basal cells and abundant rhizoids on the alga’s lower surface, also confirmed that the specimens were identical to those previously described by Pueschel and Saunders41.
Alpha Diversity indices (Observed, Shannon and Inverse Shannon) of CCA and PAC SM prokaryote richness and diversity (based on OTUs derived from HTS of 16S rDNA amplicons). The ends of the box represent the 25th and 75th percentiles, the whiskers represent minimum and maximum range, the band inside each box represents the median and the black filled circles represent the samples. iCCA = igneous CCA (n = 3), cCCA = carbonate CCA (n = 3), PAC = peyssonnelid algal crusts (n = 3).
The microbiome diversity (in terms of operational taxonomic units [OTUs], defined here by unique 16S prokaryote ribosomal RNA gene sequences) associated with PAC differed from that of CCA, with richness (total number of observed OTUs) and diversity (Shannon and Inverse Simpson) significantly higher in microbes associated with PAC (Fig. 3). Of the 7955 OTUs recovered, 73.6% were in at least one sample from PAC, compared with 47.7% and 59.6% in samples from carbonate-associated CCA (hereafter cCCA) and igneous-associated CCA (hereafter iCCA), respectively. The numbers of unique OTUs in each sample type were similarly proportioned, with 23.1% associated with PAC, 3.0% with cCCA, and 5.8% with iCCA. There was a small core microbiome (sensu58) consisting of 203 OTUs (2.6%) in samples regardless of source, whilst within samples from algal crusts, the core microbiome from PAC (1705 OTUs) was more diverse than from cCCA (583 OTUs) or iCCA (941 OTUs). A one-way Analysis of Similarity (ANOSIM) revealed a significant difference in OTU community structure between the two crustose algal types (Global R = 1.0, \(p \le\) 0.01); however, ANOSIM of the CCA samples only, showed that there was no significant difference between microbial communities associated with CCA on the two different substrate types (Global R = 0.148, \(p \le\) 0.3). Prokaryote diversity in samples from PAC was greater than in samples from CCA, and yet the diversity between samples from PAC was more conserved than in CCA (Fig. 3).
Constrained canonical analysis (ConCA) of the OTU data (Fig. 4) confirmed that the microbial communities on PAC tightly clustered on the first axis (ConCA1), and were distinct from the more divergent microbial samples from CCA. Microbial communities from cCCA and iCCA separated similarly from PAC but on the second axis (ConCA2), indicating that crustose algal type strongly influenced the prokaryote community, and that communities associated with CCA on different substrata were more similar to each other than to PAC. The weighting of individual OTUs in describing variation among samples was investigated by Principal Component Analysis (PCA), with a scatterplot of square-root transformed PCA data for OTUs highlighting the predominant contribution of a group of OTUs along the first axis (Dim1) comprising almost entirely of sequences affiliated with Pseudoalteromonas spp. (Fig. 5).
The relative abundance of Pseudoalteromonas spp. OTUs from algal crusts, revealed a large difference between samples harvested from PAC and CCA (Fig. 6). Sequences related to Pseudoalteromonas spp. were present at low relative abundance (0.14 ± 0.04% of reads [n = 307,225 ± 15,415]) in samples (n = 3) from PAC, but had over a hundredfold greater abundances (21.33 ± 8.63% of OTUs [n = 131,100 ± 14,824]) in samples (n = 6) from CCA.
Discussion
Surveys of shallow reefs (\(\le\) 14 m depth) along 5 km of the south shore of St. John revealed that rocks coated in CCA and algal turf are common settlement substrata for scleractinian recruits (reported in more detail in45). Densities of small scleractinians (\(\le\) 4 cm diameter) routinely are circa 12 colonies m\(^{-2}\) on these surfaces59, yet only two small scleractinians that putatively settled on PAC were found during the surveys conducted for densities of small corals over 2017–2019. Although the abundance of PAC in July 2017 varied strongly among quadrats, some of the quadrats were 83% occupied by PAC. Thus, it is reasonable to infer that scleractinian larvae were repelled from settling on benthic surfaces dominated by PAC, or alternatively, that they settled but were overgrown or subsequently removed by grazing or detachment. Our study provides the first evidence that contemporary sheets of PAC in the Caribbean harbour very low population densities of bacterial species previously implicated in promoting coral larval recruitment. Using HTS of 16S rDNA amplicons derived from the SM of two types of crustose algae, sequences affiliated with Pseudoalteromonas spp. bacteria were found in significantly lower numbers on PAC versus CCA, where they dominated the microbial consortia (Fig. 6). Given the dramatic and recent increase in abundance of PAC on shallow reefs in the Caribbean39,40,41,42,43,44, the absence of scleractinians recruiting to its surfaces45, and the strong role of Pseudoalteromonas spp. in the settlement of some marine invertebrate larvae31, we hypothesise that the paucity of this microbial genus on PAC may contribute to the recent ecological success of this functional group of algae in the Caribbean.
The initial impetus for this present study was to follow-up on previous research60, which investigated the factors influencing coral recruitment to, and post-settlement success on, different types of rock (i.e. igneous versus carbonate) on the reefs of St. John. This previous work suggested that larvae did not discriminate between rock types at settlement60. The present study was conducted to test the hypothesis that the microbial consortia differed between putative settlement surfaces in St. John (i.e. igneous versus carbonate rock) and, therefore, could provide a means to mediate larval settlement behaviour and distribution of juvenile corals. PAC microbiomes were concurrently sampled from the same site to provide an ecological context to the CCA, as a crustose alga which shared the substrata with CCA, but on which very few coral recruits had been observed. Whilst the OTU richness was higher in microbial communities associated with cCCA than iCCA (Fig. 3), substratum type had no significant effect on epibiotic microbial community composition; cCCA and iCCA were associated with similar relative abundances of Pseudoalteromonas spp. (18.1% ± 9.0 s.d and 24.6% ± 8.5 s.d, for iCCA and cCCA respectively). Interestingly, collection site (i.e. sampling locations along 4 km of the south shore of St. John) had a greater effect on CCA microbial composition than substratum type, with a PCA of prokaryote OTUS showing samples of both iCCA and cCCA clustering closely together by site, but with sites clearly separated (see Supplementary Fig. S1 online). These findings therefore complement the empirical data in the previous study60, by revealing that rock surfaces to which corals recruited in proportionately equal numbers were not distinguishable in terms of the microbial communities on their surfaces.
The majority of obligate marine Pseudoalteromonas spp. share intimate associations with diverse eukaryotic hosts61, where they are constituents of the microbial community occupying the biofilms coating their exposed outer surfaces. Pseudoalteromonas spp. are well suited to this niche, because many of its species produce anti-bacterial compounds62 and employ quorum sensing63 to aid their competitive colonisation of surfaces. Additionally, many secrete extracellular polymeric substances (EPS) essential for biofilm structural integrity64. However, it is their involvement in the larval metamorphosis and settlement stages of marine invertebrates30,55,63,65,66, and in particular with several species of CCA24,33, which has garnered most recent interest. The primary means by which Pseudoalteromonas spp. contribute to the settlement of scleractinian larvae appears to be through the production of the secondary metabolite, tetrabromopyrrole (TBP), which is a potent inducer of larval settlement and metamorphosis in several species of coral34,35. More recently, a second mechanism for the induction of invertebrate metamorphosis by Pseudoalteromonas spp. has been suggested in larvae of the tubeworm Hydroides elegans, where ordered arrays of phage tail-like metamorphosis-associated contractile structures (MACs), mediate bacterial-host interactions66. MACs in Pseudoalteromonas luteoviolacea induce H. elegans larval settlement, initiate loss of host cilia and activate host metamorphosis. As of yet, this mechanism has only been investigated in H. elegans, but genomic comparisons of P. luteoviolacea with other bacterial strains known to induce settlement in the tubeworm revealed it to be unique in producing these structures67, reinforcing the importance of this bacterial genus in the life cycles of various marine invertebrates.
Whilst the disproportionate numbers of sequences related to Pseudoalteromonas spp. on CCA and PAC provide compelling circumstantial evidence of differential capacity to induce larval settlement, the contrasts in overall microbiome diversity between the two algal types are also of great interest. The richness of prokaryotes in both the total (Fig. 3) and core microbiomes of PAC was significantly higher than those on the CCA, yet there was less variability between independently sampled microbiomes from PAC versus CCA, suggesting a selective pressure of PAC on their core microbiome. The identification of core OTUs is oftentimes important in microbial ecology, as it has been suggested that organisms that occur in all host species (or indeed habitats) are likely critical to the functioning of that particular community58. Metagenomic analysis of the microbiome of PAC will likely offer a greater insight into the functional roles of the core microbiome (and in particular, Pseudoalteromonas spp.) and the maintenance of what appears to be a stable symbiotic relationship between the PAC and their epifauna.
Present-day large scale crusts of PAC in the Caribbean, which are likely to include at least two speciose genera (Peyssonnelia spp. and Ramicrusta spp.), have not been extensively studied, and until now, little information is known regarding the taxonomic identity of the species composing the crust, their ecology, or their range distribution43,68,69. A recent study in which PAC were observed to be overgrowing corals in the South China Sea revealed that crusts frequently co-occurred in a complex which comprised Ramicrusta textilis, members of the family Peyssonneliaceae and also the genus Lobophora70. Peyssonnelid algae often are also only identified to genus, as species share a simple gross morphology (as in69). However, the recent increased abundance of PAC in the Caribbean41, has motivated augmentation of traditional morphological taxonomy with molecular taxonomy, the application of which has identified several new taxa including Ramicrusta sp. in Bonaire52. Prior to the present study, only one sample from the US Virgin Islands had been identified morphologically as Peyssonnelia stoechas (R. Steneck, Pers. Comm.). However, now that the PAC from one site on the south shore of St. John have been identified as Ramicrusta textilis, future research efforts can begin to address the critical need for a widespread comparative analysis of taxonomic composition of the PAC spreading rapidly on shallow Caribbean reefs. A full-length CDS for the psbA gene (Accession Number MT215153) from Ramicrusta textilis recently sampled on reefs in St Thomas (also in USVI) shows 100% sequence identity to the partial psbA gene fragments elucidated in this study (P. Gabrielsen, Pers. Comm). A preliminary survey of reefs in the Honduras in the Western Caribbean has also revealed that PAC found there also share 100% sequence identity with the specimens described in the present analysis of PAC samples from St. John (ENA Accession Number LR828131). An important conclusion supported by these results, is that Ramicrusta-dominated PAC may now have a cosmopolitan range in the Caribbean. A draft genome assembly is also currently in preparation by one of us (B. Wilson) which will likely increase our knowledge of PAC biology.
Whilst much of the biology of tropical PAC remains to be more completely described, and most studies which have addressed the ecology of benthic reef communities accord scant mention to PAC, a few details regarding their biology are unequivocal. First, their high abundance in shallow water on present-day Caribbean reefs is a recent development that first gained attention around 199839, and the trend has been noted along a 3000-km arc extending from Honduras (B. Wilson, unpublished data) to Jamaica41, Puerto Rico44, St. John54, and Bonaire42. Most records of this new phenomenon have come from shallow reefs (i.e., \(\le\) 3 m), but in St. John it is spreading to at least 14 m depth, where in July 2017, it covered a mean of 3.3 ± 0.6% (mean ± SE, range 0–13%, n = 30) of the benthos. South of St. Thomas, PAC has been found in \(>75\%\) of non-overlapping video clips from 30 m depth at College Shoals East71. Whilst PAC have been recorded on Caribbean reefs since at least the 1920’s47,72,73, and across multiple habitats, its recent increase in abundance in select locations probably reflects the combined effects of multiple factors. Of these, whilst it has been suggested that Ramicrusta found in PAC is invasive to the Caribbean42, the ongoing trend for phase-shifts from coral- to macroalgal-domination of Caribbean reefs18,74 may have also have favoured increased abundances of endemic PAC (e.g. Peyssonnelia spp.) along with fleshy macroalgae.
However, one interesting aspect of its biology that might be contributing to the current ecological success of PAC in the Caribbean may be its ability to deter settlement by invertebrates upon its surface. As the present study highlights, this likely reflects the effects of a combination of potent chemical, microbial and mechanical defenses. Many marine algae are protected from herbivory by either secondary metabolites75, or high concentrations of calcium carbonate76, with different groups of fish affected by each mode of deterence77. Peyssonnelia spp. in Fiji produce novel sesquiterpene hydroquinones called peyssonneic acids78,79 that inhibit the growth of Pseudoalteromonas bacteriolytica80, a pathogen of the commercially important macroalga Laminaria japonica81. The exclusion of Pseudoalteromonas spp. by PAC may therefore potentially be a pathogen defense mechanism, an indirect result of which is the inhibition of coral larval settlement. PAC may accentuate their competitive advantage on present day reefs by employing both defensive mechanisms (secondary metabolite production and high calcification) in concert, therefore effectively repelling a broad range of grazing fish and potentially utilising the same mechanisms in their aggressive overgrowth of live corals. However, as the spread of high-cover growths of PAC throughout the Caribbean seems to be a relatively recent phenomenon, it is likely that multiple factors have contributed to its success on present day reefs.
This study highlighted the significant difference in microbial communities associated with CCA versus PAC, in particular illuminating the disparity in the relative abundances of sequences related to the ubiquitous and ecologically-important marine bacterium Pseudoalteromonas. Given the ongoing and severe decline in coral abundance in the Caribbean, further investigations into the role of Pseudoalteromonas in coral settlement and the indirect means by which its populations may be affected by PAC are necessary.
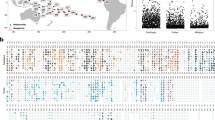
Map showing study site at Lameshur Bay, St. John, US Virgin Islands. Map created using QGIS (v2.18.17; https://qgis.org/en/site/) under GNU General Public License (Version 2)98, using Natural Earth vector and raster map data (http://naturalearthdata.com/). WP = White Point, CH = Cabritte Horn, VIERS = Virgin Islands Environmental Resource Station.
Methods
Sampling
Mucous swabs of crustose algae (PAC and CCA) were collected for microbiome studies at 7–10 m depth across Great Lameshur Bay, St. John, US Virgin Islands (\(18{^{\circ }} 18^{\prime} 25.1448^{\prime \prime }\) N \(64{^{\circ }} 43^{\prime } 14.4248^{\prime \prime }\) W), in August 2016 (Fig. 7). PAC was defined as a crust of peyssonnelid algae, thought to be in the genera Peyssonnelia and Ramicrusta41,51,52,54, with pooling of taxa into a single functional group (PAC) rationalized by their convergence in function82. Patches of CCA and PAC were haphazardly selected for sampling at various locations on the shallow fringing reefs between White Point (WP) and Cabritte Horn (CH) (Fig. 7) where scleractinians, octocorals sponges, and macroalgae are common, but where coral cover has remained \(< 4.5\%\) for decades59,83. PAC were sampled from igneous substrata (n = 3), whilst CCA were sampled from both igneous (n = 3; see Supplementary Fig. S2 online) and carbonate rock (n = 3; see Supplementary Fig. S3 online) surfaces, with this sampling regime reflecting both the profusion of igneous boulders and submerged cliffs on the shallow reefs of the Virgin Islands, and the distribution of PAC and CCA on these surfaces. CCA was sampled from two rock types in order to determine whether substratum type was associated with surface mucous microbiome composition. Sterile cotton swabs were stored prior to sampling in sealed, sterile 15 mL polypropylene tubes. Inverted tubes were opened underwater immediately above the crustose algae and their SM was collected by rolling the swab over the algal surface, and returning the swabs to the inverted tube such that it was sealed with minimal seawater ingress and the swab maintained in an air pocket. Upon return to the laboratory (\(< 60\) min after sampling), swabs were trimmed to 2 cm length and added to cryogenic vials (2 mL; Corning, Part No. 430488) containing 500 \(\upmu \hbox {L}\) RNAlater (ThermoFisher Scientific), with the swab fully immersed. Cryogenic vials were stored at − 20 °C in St. John for five days, but were thawed to room temperature for shipment to Europe where they were stored at 4 °C prior to analysis. The collection of mucous samples in St. John was permitted by the Virgin Islands National Park (VIIS-2016-SCI-0031).
For microscopic examination and phylogenetic identification of the PAC, tissue samples were collected in August 2018 from an area of a few square metres at approximately 4 m depth at Cabritte Horn. Upon immediate return to the laboratory, tissue samples were carefully examined and brushed clean to remove as much contaminating debris (e.g. macroalgae and invertebrates) as possible. Prepared samples were immersed in RNALater and stored at 4 °C, prior to shipment at room temperature to the Department of Embryology, Carnegie Institution for Science, Baltimore, USA. The collection of tissue samples in St. John and their export was permitted by the Virgin Islands National Park (VIIS-2018-SCI-0012) and Department of Planning and Natural Resources (DFW 18087J).
Microscopy
Specimens for microscopy were fixed for 4 h in a solution containing 3% glutaraldehyde and 1% formaldehyde in a 0.1 M sodium cacolyte buffer (containing 2 mM each calcium and magnesium salts) at pH 7.4. Samples were rinsed for 10 min (four times each) in distilled water before being decalcified, firstly in 10% formic acid at \(4 {^{\circ }}\hbox {C}\) for 3 d and then in fresh 10% formic acid (at \(4 {^{\circ }}\hbox {C}\) again) overnight. After two further rinses (10 min each) in distilled water, specimens were dehydrated for 15 min in each of a graded ethanol series (35%, 50%, 75% and 95%), before two final 100% ethanol treatments for 15 min. Specimens were then treated twice with a solution of 100% ethanol and resin (either LR White or Micro-Bed), for 1 h with ethanol:resin (2:1), then 1 h with ethanol:resin (1:2). Specimens were infiltrated with 100% resin overnight at \(4 {^{\circ }}\hbox {C}\), before two further treatments of 100% resin for 1 h each. The final embedding in 100% resin was carried out at \(55 {^{\circ }}\hbox {C}\) overnight. \(1\upmu \hbox {m}\) sections were cut and mounted on slides and stained with 1% Methylene Blue (for visualisation of cell walls) and 1% Azure B (cytoplasm), warm, for light microscopy.
PAC Microbiome DNA Extraction, PCR and HTS
Genomic DNA was extracted from mucous swabs using a modified protocol for the Fast DNA SPIN Kit for Soil (MP Biomedicals). Briefly, the contents of the Lysing Matrix E tube were transferred to the sample vial containing the cotton swab. Sodium Phosphate Buffer (300 \(\upmu \hbox {L}\)) was added to the sample and vortexed for 5 s. MT Buffer (122 \(\upmu \hbox {L}\)) was added to the cryogenic vial and the sample homogenised on a Pulsing Vortex Mixer (VWR) at maximum speed for 40 s. The sample was centrifuged at 10,000 \(\times\) g for 10 min and the supernatant transferred to a 2 mL tube. Solution PPS (250 \(\upmu \hbox {L}\)) was added to the supernatant and mixed by inversion ten times. The sample was centrifuged again at 14,000 \(\times\) g for 5 min and the supernatant transferred to 15 mL tube. The Binding Matrix suspension was resuspended by vortexing for 5 s and 1 mL immediately added to the supernatant, before mixing by inversion for 2 min and settling in a rack for 3 min. Avoiding the Binding Matrix, 500 \(\upmu \hbox {L}\) supernatant was removed and discarded and the remaining supernatant and Binding Matrix resuspended by inversion, before 600 \(\upmu \hbox {L}\) suspension was transferred to a SPIN filter. The SPIN filter was centrifuged at 14,000 \(\times\) g for 1 min and the filtrate discarded. The remaining suspension was added to the SPIN filter and centrifuged again at 14,000 \(\times\) g for 1 min. The filtrate was again discarded and 500 \(\upmu \hbox {L}\) SEWS-M added to the pellet on the SPIN filter and the pellet gently resuspended using the pipette. The SPIN filter was centrifuged at 14,000 \(\times\) g for 1 min, the filtrate discarded and the SPIN filter centrifuged again at 14,000 \(\times\) g for 2 min. The catch tube containing the filtrate was discarded and the SPIN filter placed in a clean catch tube and air-dried for 5 min. The pellet on the SPIN filter was gently resuspended in ddH2O (100 \(\upmu \hbox {L}\)) using a pipette and the SPIN filter incubated for 5 min at \(55 {^{\circ }}\hbox {C}\), before a final centrifugation at 14,000 \(\times\) g for 1 min. The SPIN filter was discarded and the eluate stored at \(-20 {^{\circ }}\hbox {C}\).
For analysis of the microbiome, the V4 region of the small-subunit (SSU) 16S rRNA gene was amplified from genomic DNA using a two-step nested PCR approach (after84). Briefly, samples were amplified using primers 519F (5\(^{\prime }\)-CAG CMG CCG CGG TAA-3\(^{\prime }\))85 and 806R (5\(^{\prime }\)-GGA CTA CHV GGG TWT CTA AT-3\(^{\prime }\))86. The reaction mixture consisted of 10 \(\upmu \hbox {L}\) HotStarTaq Master Mix (Qiagen), 500 nM of each primer, 1 \(\upmu \hbox {L}\) template DNA, \(200\hbox { ng mL}^{-1}\) non-acetylated BSA and nuclease-free water to bring the total volume to 20 \(\upmu \hbox {L}\). Reactions were initially denatured for 15 min at \(95 {^{\circ }}\hbox {C}\), followed by 25 cycles of denaturation at \(95 {^{\circ }}\hbox {C}\) for 20 s, primer annealing at \(55 {^{\circ }}\hbox {C}\) for 30 s and extension at \(72 {^{\circ }}\hbox {C}\) for 30 s, followed by a final extension step of \(72 {^{\circ }}\hbox {C}\) for 7 min. In the second step, pooled amplicons were amplified and barcoded by nested PCR using MID-tagged primers 519F and 806R in a reaction mixture comprising 25 \(\upmu \hbox {L}\) HotStarTaq Master Mix, 500 nM of each primer, 1 \(\upmu \hbox {L}\) amplicon DNA, and nuclease-free water to bring the total volume to 50 \(\upmu \hbox {L}\). Reactions were initially denatured for 15 min at \(95 {^{\circ }}\hbox {C}\), followed by 15 cycles of denaturation at \(95 {^{\circ }}\hbox {C}\) for 20 s, primer annealing at \(62 {^{\circ }}\hbox {C}\) for 30 s and extension at \(72 {^{\circ }}\hbox {C}\) for 30 s. This was followed by a final extension step of \(72 {^{\circ }}\hbox {C}\) for 7 min. The quantity and quality of the second-step PCR amplicons was assessed by agarose gel electrophoresis. The second-step PCR amplicons were purified using Agencourt AMPure XP Beads (Beckman Coulter Inc., CA, USA) and quantified using a Qubit 3.0 Fluorometer and by agarose gel electrophoresis. MID-tagged amplicons were then pooled in equimolar amounts for library construction. Libraries were sent to the Norwegian Sequencing Centre (Oslo, Norway) for paired-end (2 \(\times\) 300bp) HTS on a MiSeq platform (Illumina, CA, USA) using the MiSeq Reagent Kit v3 (Illumina) kit.
PAC DNA extraction, PCR and sanger sequencing
Tissue (100 mg wwt) was ground with a pestle and mortar and genomic DNA extracted using the DNeasy Plant Mini Kit (Qiagen), as per the manufacturer’s instructions and eluted in a final volume of 200 \(\upmu \hbox {L}\).
The phylogenetic identification of the PAC was ascertained by amplification of fragments each of the nuclear-encoded, large-subunit (LSU) ribosomal RNA gene and the psbA protein D1 gene (which comprises part of the reaction center of photosystem II) by PCR. Samples were amplified using primers X (5\(^{\prime }\)-GCA GGA CGG TGG CCA TGG AAG T-3\(^{\prime }\)) and 28F (5\(^{\prime }\)-CAG AGC ACT GGG CAG AAA TCA C-3\(^{\prime }\))87 and primers F40 (5\(^{\prime }\)-CGT TAT GAG TCA GGT GTA ATT CC-3\(^{\prime }\)) and R851.1 (5\(^{\prime }\)-GCC ATT GTT TGA ATA GC-3\(^{\prime }\))88 for the LSU gene and primers psbA-F (5\(^{\prime }\)-ATG ACT GCT ACT TTA GAA AGA CG-3\(^{\prime }\)) and psbA-R2 (5\(^{\prime }\)-TCA TGC ATW ACT TCC ATA CCT A-3\(^{\prime }\))89 for the psbA gene. The reaction mixture comprised 25 \(\upmu \hbox {L}\) GoTaq GT HotStart Green Master Mix (Promega), 200 nM of each primer, 1 \(\upmu \hbox {L}\) of template DNA (tenfold-diluted in nuclease-free water), 200 ng mL\(^{-1}\) non-acetylated BSA and nuclease-free water to bring the total volume to 50 \(\upmu \hbox {L}\). Reactions were denatured for 15 min at \(95 {^{\circ }}\hbox {C}\), followed by 35 cycles of denaturation at \(95 {^{\circ }}\hbox {C}\) for 60 s, primer annealing at 57 and \(48 {^{\circ }}\hbox {C}\) for 60 s (for LSU and psbA genes, respectively) and extension at \(72 {^{\circ }}\hbox {C}\) for 2 min, followed by a final extension step of \(72 {^{\circ }}\hbox {C}\) for 10 min. The quantity and quality of the PCR amplicons was assessed by agarose gel electrophoresis, prior to excision of bands for sequencing by Genewiz (MD, USA) with the same primers used for PCR.
Bioinformatic analyses
HTS reads were processed using BBTools (https://jgi.doe.gov/data-and-tools/bbtools/). Briefly: (1) phiX contaminants (calibration controls used during sequencing) were removed; (2) paired end reads were merged; (3) forward and reverse primer sequences and linkers were removed; (4) reads were quality end-trimmed at a phred quality score \(\le\) 27; and (5) reads \(< 200\) bp in length were removed. The remaining sequence reads were checked for chimeras with the identify chimeric seqs and filter fasta scripts in QIIME (Quantitative Insights into Microbial Ecology, v1.9.090), using usearch6191 and the ChimeraSlayer92 16S rRNA reference database (Gold.fa) found in the Broad Microbiome Utilities suite (http://microbiomeutil.sourceforge.net/). The pick de novo otus script in QIIME (using default parameters) was used for de novo OTU picking (using uclust91) and a sequence similarity threshold of 97%), taxonomy assignment (using PyNAST93) at 90% sequence similarity against the Greengenes core reference alignment database (Release 13_894), and finally, the assembly of a table of OTU abundances with taxonomic identifiers for each OTU.
Sanger FASTQ files were quality end-trimmed at a phred quality score \(\le\) 27 using Sickle (Version 1.33)95 and processed using EMBOSS96. LSU and psbA sequences were compared with those in the NCBI database using the Nucleotide Basic Local Alignment Search Tool (BLASTn) algorithm97.
Code availability
HTS data were submitted to the European Nucleotide Archive (ENA) under Study PRJEB39143, whilst PAC marker gene sequences were submitted to ENA under Accession Numbers LR828130-LR828131 (LSU) and LR861111 (psbA).
References
Jackson, J., Donovan, M., Cramer, K. & Lam, V. Status and Trends of Caribbean Coral Reefs: 1970–2012. Glob. Coral Reef Monit. Network, Int. Union for Conserv. Nat. Glob. Mar. Polar Program, Washington, DC (2014).
Baird, A. H., Babcock, R. C. & Mundy, C. P. Habitat selection by larvae influences the depth distribution of six common coral species. Mar. Ecol. Prog. Ser. 252, 289–293 (2003).
Harrington, L., Fabricius, K., De’ath, G. & Negri, A. Recognition and selection of settlement substrata determine post-settlement survival in corals. Ecol. 85, 3428–3437 (2004).
Roff, G. & Mumby, P. J. Global disparity in the resilience of coral reefs. Trends Ecol. & Evol. 27, 404–413 (2012).
Lessios, H. A. Mass mortality of Diadema antillarum in the Caribbean: What Have We Learned? Annu. Rev. Ecol. Syst. 19, 371–393 (1988).
Patterson, K. L. et al. The etiology of white pox, a lethal disease of the Caribbean elkhorn coral, Acropora palmata. Proc. Natl. Acad. Sci. 99, 8725–8730 (2002).
Eakin, C. M. et al. Caribbean Corals in Crisis: Record Thermal Stress, Bleaching, and Mortality in 2005. PLoS ONE 5, 1–9 (2010).
Wilkinson, C. R. In Status of Coral Reefs of the World: 2004 (Townsville: Australian Institute of Marine Science, 2004).
Miller, J., Waara, R., Muller, E. & Rogers, C. Coral bleaching and disease combine to cause extensive mortality on reefs in US Virgin Islands. Coral Reefs 25, 418 (2006).
Donner, S. D., Knutson, T. R. & Oppenheimer, M. Model-based assessment of the role of human-induced climate change in the 2005 Caribbean coral bleaching event. Proc. Natl. Acad. Sci. 104, 5483–5488 (2007).
Hughes, T. P. et al. Global warming and recurrent mass bleaching of corals. Nat. 543, 373–377 (2017).
Gardner, T. A., Côté, I. M., Gill, J. A., Grant, A. & Watkinson, A. R. Hurricanes and Caribbean coral reefs: impacts, recovery patterns, and role in long-term decline. Ecol. 86, 174–184 (2005).
Smith, S. R. Patterns of Coral Recruitment and Post-Settlement Mortality on Bermuda’s Reefs: Comparisons to Caribbean and Pacific Reefs. Am. Zool. 32, 663–673 (1992).
Arnold, S. N. & Steneck, R. S. Settling into an Increasingly Hostile World: The Rapidly Closing “Recruitment Window” for Corals. PLoS ONE 6, e28681 (2011).
Edmunds, P. J. et al. Geographic variation in long-term trajectories of change in coral recruitment: a global-to-local perspective. Mar. Freshw. Res. 66, 609–622 (2015).
Glassom, D., Zakai, D. & Chadwick-Furman, N. Coral recruitment: a spatio-temporal analysis along the coastline of Eilat, northern Red Sea. Mar. Biol. 144, 641–651 (2004).
Edmunds, P. J. et al. Why more comparative approaches are required in time-series analyses of coral reef ecosystems. Mar. Ecol. Prog. Ser. 608, 297–306 (2019).
Hughes, T. P. Catastrophes, Phase Shifts, and Large-Scale Degradation of a Caribbean Coral Reef. Sci. 265, 1547–1551 (1994).
Jackson, J. B. C. et al. Historical Overfishing and the Recent Collapse of Coastal Ecosystems. Sci. 293, 629–637 (2001).
McCook, L. Macroalgae, nutrients and phase shifts on coral reefs: scientific issues and management consequences for the Great Barrier Reef. Coral Reefs 18, 357–367 (1999).
Loh, T., McMurray, S. E., Henkel, T. P., Vicente, J. & Pawlik, J. R. Indirect effects of overfishing on Caribbean reefs: sponges overgrow reef-building corals. PeerJ 3, e901 (2015).
Lenz, E., Bramanti, L., Lasker, H. & Edmunds, P. J. Long-term variation of octocoral populations in St. John, US Virgin Islands. Coral Reefs 34, 1099–1109 (2015).
Gleason, D. F. & Hofmann, D. K. Coral larvae: From gametes to recruits. J. Exp. Mar. Biol. Ecol. 408, 42–57 (2011).
Siboni, N. et al. Using Bacterial Extract along with Differential Gene Expression in Acropora millepora Larvae to Decouple the Processes of Attachment and Metamorphosis. PLoS ONE 7, e37774 (2012).
Doropoulos, C. et al. Characterizing the ecological trade-offs throughout the early ontogeny of coral recruitment. Ecol. Monogr. 86, 20–44 (2016).
Ritson-Williams, R., Arnold, S. N., Paul, V. J. & Steneck, R. S. Larval settlement preferences of Acropora palmata and Montastraea faveolata in response to diverse red algae. Coral Reefs 33, 59–66 (2014).
Ritson-Williams, R., Paul, V. J., Arnold, S. N. & Steneck, R. S. Larval settlement preferences and post-settlement survival of the threatened caribbean corals Acropora palmata and A. cervicornis. Coral Reefs 29, 71–81 (2010).
Johnson, C. R., Muir, D. G. & Reysenbach, A. L. Characteristic bacteria associated with surfaces of coralline algae: a hypothesis for bacterial induction of marine invertebrate larvae. Mar. Ecol. Prog. Ser. 74, 281–294 (1991).
Heyward, A. J. & Negri, A. P. Natural inducers for coral larval metamorphosis. Coral Reefs 18, 273–279 (1999).
Webster, N. S. et al. Metamorphosis of a Scleractinian Coral in Response to Microbial Biofilms. Appl. Environ. Microbiol. 70, 1213–1221 (2004).
Hadfield, M. G. Biofilms and Marine Invertebrate Larvae: What Bacteria Produce that Larvae Use to Choose Settlement Sites. Annu. Rev. Mar. Sci. 3, 453–470 (2011).
Sneed, J. M., Ritson-Williams, R. & Paul, V. J. Crustose coralline algal species host distinct bacterial assemblages on their surfaces. ISME J. 9, 2527–2536 (2015).
Negri, A. P., Webster, N. S., Hill, R. T. & Heyward, A. J. Metamorphosis of broadcast spawning corals in response to bacteria isolated from crustose algae. Mar. Ecol. Prog. Ser. 223, 121–131 (2001).
Sneed, J. M., Sharp, K. H., Ritchie, K. B. & Paul, V. J. The chemical cue tetrabromopyrrole from a biofilm bacterium induces settlement of multiple Caribbean corals. Proc. Royal Soc. B 281, 20133086 (2014).
Tebben, J. et al. Induction of Larval Metamorphosis of the Coral Acropora millepora by Tetrabromopyrrole Isolated from a Pseudoalteromonas Bacterium. PLoS ONE 6, e19082 (2011).
Tebben, J. et al. Chemical mediation of coral larval settlement by crustose coralline algae. Sci. Reports 5, 10803 (2015).
Morse, D. E., Morse, A. N. C., Raimondi, P. T. & Hooker, N. Morphogen-Based Chemical Flypaper for Agaricia humilis Coral Larvae. The Biol. Bull. 186, 172–181 (1994).
Raimondi, P. T. & Morse, A. N. C. The consequences of complex larval behavior in a coral. Ecol. 81, 3193–3211 (2000).
Antonius, A. & Ballesteros, E. Epizoism: a new threat to coral health in Caribbean reefs. Revista de Biol. Trop. 46, 145–156 (1998).
Verlaque, M., Ballesteros, E. & Antonius, A. Metapeyssonnelia corallepida sp. nov. (Peyssonneliaceae, Rhodophyta), an Atlantic Encrusting Red Alga Overgrowing Corals. Bot. Mar. 43, 191–200 (2000).
Pueschel, C. M. & Saunders, G. W. Ramicrusta textilis sp. nov. (Peyssonneliaceae, Rhodophyta), an anatomically complex Caribbean alga that overgrows corals. Phycol. 48, 480–491 (2009).
Eckrich, C. E., Engel, M. S. & Peachey, R. B. J. Crustose, calcareous algal bloom (Ramicrusta sp.) overgrowing scleractinian corals, gorgonians, a hydrocoral, sponges, and other algae in Lac Bay, Bonaire, Dutch Caribbean. Coral Reefs 30, 131–131 (2011)
Eckrich, C. E. & Engel, M. S. Coral overgrowth by an encrusting red alga (Ramicrusta sp.): a threat to Caribbean reefs?. Coral Reefs 32, 81–84 (2013).
Ballantine, D. & Ruiz, H. Metapeyssonnelia milleporoides, a new species of coral-killing red alga (Peyssonneliaceae) from Puerto Rico, Caribbean Sea. Bot. Mar. 54, 47–51 (2011).
Edmunds, P. J., Zimmerman, S. A. & Bramanti, L. A spatially aggressive peyssonnelid algal crust (PAC) threatens shallow coral reefs in St. John, US Virgin Islands. Coral Reefs 38, 1329–1341 (2019).
Basso, D. Carbonate production by calcareous red algae and global change. Geodiversitas 34, 13–33 (2012).
Taylor, W. & Arndt, C. The marine algae of the southwestern Peninsula of Hispaniola. Am. J. Bot. 16, 651–662 (1929).
Van Den Hoek, C., Cortel-Breeman, A. & Wanders, J. Algal zonation in the fringing coral reef of Curaçao, Netherlands Antilles, in relation to zonation of corals and gorgonians. Aquatic. Bot. 1, 269–308 (1975).
Sammarco, P. W. Effects of grazing by Diadema antillarum Philippi (Echinodermata: Echinoidea) on algal diversity and community structure. J. Exp. Mar. Biol. Ecol. 65, 83–105 (1982).
Littler, M. M., Taylor, P. R., Littler, D. S., Sims, R. H. & Norris, J. N. Dominant macrophyte standing stocks, productivity and community structure on a Belizean barrier-reef. Atoll Res. Bull. 302, 1–24 (1987).
Ballantine, D. & Ruiz, H. A unique red algal reef formation in Puerto Rico. Coral Reefs 32, 411–411 (2013).
Ballantine, D., Ruiz, H., Lozada-Troche, C. & Norris, J. N. The genus Ramicrusta (Peyssonneliales, Rhodophyta) in the Caribbean Sea, including Ramicrusta bonairensis sp. nov. and Ramicrusta monensis sp. nov. Bot. Mar. 59, 417–431 (2016).
Zhang, D. & Zhou, J. Ramicrusta, a new genus of Peyssonneliaceae (Cryptonemiales, Rhodophyta). Oceanol. et Limnol. Sinica 12, 538–544 (1981).
Bramanti, L., Lasker, H. R. & Edmunds, P. J. An encrusting peyssonnellid preempts vacant space and overgrows corals in St. John, US Virgin Islands. Reef Encount. 32, 68–70 (2017).
Tran, C. & Hadfield, M. G. Larvae of Pocillopora damicornis (Anthozoa) settle and metamorphose in response to surface-biofilm bacteria. Mar. Ecol. Prog. Ser. 433, 85–96 (2011).
Golbuu, Y. & Richmond, R. H. Substratum preferences in planula larvae of two species of scleractinian corals, Goniastrea retiformis and Stylaraea punctata. Mar. Biol. 152, 639–644 (2007).
Longford, S. R. et al. Comparisons of diversity of bacterial communities associated with three sessile marine eukaryotes. Aquatic. Microb. Ecol. 48, 217–229 (2007).
Shade, A. & Handelsman, J. Beyond the Venn diagram: the hunt for a core microbiome. Environ. Microbiol. 14, 4–12 (2011).
Edmunds, P. J. The hidden dynamics of low coral cover communities. Hydrobiol. 818, 193–209 (2018).
Green, D. H., Edmunds, P. J., Pochon, X. & Gates, R. D. The effects of substratum type on the growth, mortality, and photophysiology of juvenile corals in St. John, US Virgin Islands. J. Exp. Mar. Biol. Ecol. 384, 18–29 (2010).
Holmström, C. & Kjelleberg, S. Marine Pseudoalteromonas species are associated with higher organisms and produce biologically active extracellular agents. FEMS Microbiol. Ecol. 30, 285–293 (1999).
Offret, C. et al. Spotlight on Antimicrobial Metabolites from the Marine Bacteria Pseudoalteromonas: Chemodiversity and Ecological Significance. Mar. Drugs 14, 129 (2016).
Huang, Y.-L., Dobretsov, S., Xiong, H. & Qian, P.-Y. Effect of Biofilm Formation by Pseudoalteromonas spongiae on Induction of Larval Settlement of the Polychaete Hydroides elegans. Appl. Environ. Microbiol. 73, 6284–6288 (2007).
Wolfaardt, G. M., Lawrence, J. R. & Korber, D. R. Function of EPS. In Wingender, J., Neu, T. R. & Flemming, H. (eds.) Microbial extracellular polymeric substances: characterization, structure, and function, 186–190 (Springer, New York, NY, 1999).
Huggett, M. J., Williamson, J. E., de Nys, R., Kjelleberg, S. & Steinberg, P. D. Larval settlement of the common Australian sea urchin Heliocidaris erythrogramma in response to bacteria from the surface of coralline algae. Oecologia 149, 604–619 (2006).
Shikuma, N. J., Antoshechkin, I., Medeiros, J. M., Pilhofer, M. & Newman, D. K. Stepwise metamorphosis of the tubeworm Hydroides elegans is mediated by a bacterial inducer and MAPK signaling. Proc. Natl. Acad. Sci. 113, 10097–10102 (2016).
Freckleton, M. L., Nedved, B. T. & Hadfield, M. G. Induction of Invertebrate Larval Settlement; Different Bacteria, Different Mechanisms? Sci. Reports 7, 42557 (2017).
Kato, A., Baba, M., Kawai, H. & Masuda, M. Reassessment of the little-known crustose red algal genus Polystrata (Gigartinales), based on morphology and SSU rDNA sequences. J. Phycol. 42, 922–933 (2006).
Dixon, K. R. & Saunders, G. W. DNA barcoding and phylogenetics of Ramicrusta and Incendia gen. nov., two early diverging lineages of the Peyssonneliaceae (Rhodophyta). Phycol. 52, 82–108 (2013).
Nieder, C., Chen, P.-C., Chen, C. A. & Liu, S.-L. New record of the encrusting alga Ramicrusta textilis overgrowing corals in the lagoon of Dongsha Atoll, South China Sea. Bull. Mar. Sci. 95, 459–462 (2019).
Smith, T. B., Ennis, R., Kadison, E., Nemeth, R. S. & Henderson, L. The United States Virgin Islands Territorial Coral Reef Monitoring Program. 2016 Annu. Report, Univ. Virgin Islands, United States Virgin Islands 1–286 (2016).
Adey, W. H. A survey of red algal biology and ecology with reference to carbonate geology, and the role of reds in algal ridge and reef construction. In Recent Advances in Carbonate Studies, vol. 6, 3–6 (West Indies Laboratory, St. Croix, USVI, 1974).
Woelkerling, W. J. South Florida benthic marine algae. In Sedimenta V (University of Miami, 1976).
Hoegh-Guldberg, O. et al. Coral reefs under rapid climate change and ocean acidification. Sci. 318, 1737–1742 (2007).
Hay, M. E. Marine Chemical Ecology: Chemical Signals and Cues Structure Marine Populations, Communities, and Ecosystems. Annu. Rev. Mar. Sci. 1, 193–212 (2009).
Schupp, P. J. & Paul, V. J. Calcium Carbonate and Secondary Metabolites in Tropical Seaweeds: Variable Effects on Herbivorous Fishes. Ecol. 75, 1172–1185 (1994).
Pennings, S. C., Puglisi, M. P., Pitlik, T. J., Himaya, A. C. & Paul, V. J. Effects of secondary metabolites and CaCO\(_{3}\) on feeding by surgeonfishes and parrotfishes: within-plant comparisons. Mar. Ecol. Prog. Ser. 134, 49–58 (1996).
Blunt, J. W. et al. Marine natural products. Nat. Prod. Reports 24, 31–86 (2007).
Blunt, J. W. et al. Marine natural products. Nat. Prod. Reports 25, 35–94 (2008).
Lane, A. L. et al. Ecological leads for natural product discovery: novel sesquiterpene hydroquinones from the red macroalga Peyssonnelia sp. Tetrahedron 66, 455–461 (2010).
Sawabe, T. et al. Pseudoalteromonas bacteriolytica sp. nov., a marine bacterium that is the causative agent of red spot disease of Laminaria japonica. Int. J. Syst. Evol. Microbiol. 48, 769–774 (1998).
Steneck, R. & Dethier, M. A Functional Group Approach to the Structure of Algal-Dominated Communities. Oikos 69, 476–498 (1994).
Edmunds, P. J. Decadal-scale changes in the community structure of coral reefs of St. John, US Virgin Islands. Mar. Ecol. Prog. Ser. 489, 107–123 (2013).
Wilson, B. et al. Changes in Marine Prokaryote Composition with Season and Depth Over an Arctic Polar Year. Front. Mar. Sci. 4, 95 (2017).
Øvreås, L., Forney, L., Daae, F. L. & Torsvik, V. Distribution of bacterioplankton in meromictic Lake Saelenvannet, as determined by denaturing gradient gel electrophoresis of PCR-amplified gene fragments coding for 16S rRNA. Appl. Environ. Microbiol. 63, 3367–3373 (1997).
Caporaso, J. G. et al. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc. Natl. Acad. Sci. 108, 4516–4522 (2011).
Freshwater, D., Fredericq, S. & Bailey, J. Characteristics and utility of nuclear-encoded large-subunit ribosomal gene sequences in phylogenetic studies of red algae. Phycol. Res. 47, 33–38 (1999).
Krayesky, D. M., Norris, J. N., Gabrielson, P. W., Gabriel, D. & Fredericq, S. A new order of red algae based on the Peyssonneliaceae, with an evaluation of the ordinal classification of the Florideophyceae (Rhodophyta). Proc. Biol. Soc. Wash. 122, 364–391 (2009).
Yoon, H. S., Hackett, J. D. & Bhattacharya, D. A single origin of the peridinin- and fucoxanthin-containing plastids in dinoflagellates through tertiary endosymbiosis. Proc. Natl. Acad. Sci. 99, 11724–11729 (2002).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–6 (2010).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinforma. 26, 2460–2461 (2010).
Haas, B. J. et al. Chimeric 16S rRNA sequence formation and detection in Sanger and 454-pyrosequenced PCR amplicons. Genome Res. 21, 494–504 (2011).
Caporaso, J. G. et al. PyNAST: a flexible tool for aligning sequences to a template alignment. Bioinforma. 26, 266–267 (2009).
DeSantis, T. Z. et al. Greengenes, a Chimera-Checked 16S rRNA Gene Database and Workbench Compatible with ARB. Appl. Environ. Microbiol. 72, 5069–5072 (2006).
Joshi, N. A. & Fass, J. N. Sickle: A sliding-window, adaptive, quality-based trimming tool for FastQ files (Version 1.33) (2011). Available at https://github.com/najoshi/sickle. Accessed 6 Nov 2020.
Rice, P., Longden, I. & Bleasby, A. EMBOSS: The European Molecular Biology Open Software Suite. Trends Genet. 16, 276–277 (2000).
Altschul, S., Gish, W., Miller, W., Myers, E. & Lipman, D. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
GNU General Public License. URL http://www.gnu.org/licenses/gpl.html. Accessed 6 Nov 2020.
Acknowledgements
This work was funded by a US National Science Foundation Long Term Research in Environmental Biology (LTREB) (Grant Number DEB 13-50146) and Rapid Response Research (RAPID) (Grant Number RAPID 18-01335) grants awarded to PJE and was completed under research permits issued by the Virgin Islands National Park (VIIS-2016-SCI-0031 and VIIS-2018-SCI-0012). We are extremely grateful to the CSUN graduate students who assisted with this work, in particular Ashley Potter, Arien Widrick, Sigfredo Zimmerman and Megan Williams and also the facilities at the Virgin Islands Environmental Resource Station (VIERS). CMF is supported by the Carnegie Institution for Science and thanks also go to Michael Sepanski there for his technical assistance with microscopy. We thank Dan Exton and Operation Wallacea for support and assistance with collection and export of PAC samples from Honduras, under a research permit (DE-MP-81-2017) and export permit (ICF-DVS-17-2018) issued to Dan Exton by the Honduran Government’s Instituto de Conservación Forestal (ICF). We also thank Stephen Harris at the Department of Plant Science, University of Oxford and Robert Steneck, University of Maine for their advice regarding algal taxonomy, and David Bourne at James Cook University and the Australian Institute of Marine Science for comments that improved an earlier draft of this paper. Finally, BW is indebted to PJE and CMF for their generosity and support with this work.
Author information
Authors and Affiliations
Contributions
B.W. and P.J.E. designed the research; B.W., P.J.E and C.M.F. performed experiments; B.W. analysed the data; B.W. and P.J.E. wrote the manuscript. All authors reviewed and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wilson, B., Fan, CM. & Edmunds, P.J. An unusual microbiome characterises a spatially-aggressive crustose alga rapidly overgrowing shallow Caribbean reefs. Sci Rep 10, 20949 (2020). https://doi.org/10.1038/s41598-020-76204-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-76204-0
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.