Abstract
CHARGE syndrome, a rare multiple congenital anomaly condition, is caused by haploinsufficiency of the chromatin remodeling protein gene CHD7 (Chromodomain helicase DNA binding protein 7). Brain abnormalities and intellectual disability are commonly observed in individuals with CHARGE, and neuronal differentiation is reduced in CHARGE patient-derived iPSCs and conditional knockout mouse brains. However, the mechanisms of CHD7 function in nervous system development are not well understood. In this study, we asked whether CHD7 promotes gene transcription in neural progenitor cells via changes in chromatin accessibility. We used Chd7 null embryonic stem cells (ESCs) derived from Chd7 mutant mouse blastocysts as a tool to investigate roles of CHD7 in neuronal and glial differentiation. Loss of Chd7 significantly reduced neuronal and glial differentiation. Sholl analysis showed that loss of Chd7 impaired neuronal complexity and neurite length in differentiated neurons. Genome-wide studies demonstrated that loss of Chd7 leads to modified chromatin accessibility (ATAC-seq) and differential nascent expression (Bru-Seq) of neural-specific genes. These results suggest that CHD7 acts preferentially to alter chromatin accessibility of key genes during the transition of NPCs to neurons to promote differentiation. Our results form a basis for understanding the cell stage-specific roles for CHD7-mediated chromatin remodeling during cell lineage acquisition.
Similar content being viewed by others
Introduction
CHARGE syndrome is a multisystem developmental disorder characterized by Coloboma, Heart defects, Atresia of the choanae, Retardation of growth and development, Genital hypoplasia, and Ear malformations including deafness and vestibular disorders1. Nervous system abnormalities are also common among individuals with CHARGE, including brain malformations (i.e., hindbrain anomalies, brainstem anomalies, and ventricular enlargement), abnormal sensory organ structure and function (i.e., ocular coloboma and sensorineural hearing loss), and impaired cranial nerves (hypoplasia or aberrant innervation)2,3,4,5. Together, these malformations contribute to a wide range of cognitive defects and behavioral challenges for individuals with CHARGE, including learning disabilities, speech and communication difficulties, motor incoordination, and autistic behaviors6,7.
Pathogenic variants in CHD7 are a major diagnostic criterion for CHARGE syndrome and are present in over 71% of cases8. CHD7 encodes Chromodomain Helicase DNA binding protein 7 (CHD7), an ATP-dependent chromatin remodeling enzyme and member of the Group III CHD family of chromatin remodelers9. Chd7 is expressed highly in the adult mouse subventricular zone (SVZ) and subgranular zone (SGZ) where neural progenitors contribute to formation of adult born neurons10. CHD7 has been broadly implicated in neurogenesis in multiple areas of the nervous system, during development and beyond. In mice, heterozygous loss of Chd7 mimics human CHARGE syndrome, with reduced volume of the brain, cortex, and cerebellum compared to controls3. In adult Chd7 conditional knockout mice, SVZ neural stem/progenitor cells exhibit impaired neurogenesis11, and loss of Chd7 leads to reduced neurogenesis and abnormal dendritic development of newly born neurons in the adult mouse SVZ and SGZ10. Adult hippocampal loss of Chd7 causes premature neuron production and depletion of neural stem cells12. In the telencephalon, midbrain and spinal cord, neuroepithelial thickness is reduced in E10.5 heterozygous gene trap (Chd7Gt/+) embryos13. Together, these observations implicate CHD7 in a wide variety of central nervous system and neuron-specific processes.
In addition to central nervous system defects, Chd7 deficiency has been implicated in peripheral nervous system neurogenesis, including in the olfactory and auditory epithelia14,15. In the ear, germline and conditional loss of Chd7 leads to a severe reduction in neuroblasts in the developing otic epithelium and the cochleovestibular ganglion15. Taken together, these observations demonstrate that CHD7 promotes neurogenesis and gliogenesis in early embryonic central and peripheral nervous system development.
CHD7, like other chromatin remodelers, is known to regulate chromatin accessibility using the energy of ATP to reposition nucleosomes16,17. CHD7 is required in cerebellar granule neural progenitor cells for maintenance of open chromatin differentiation18, regulation of neuronal differentiation10,15, and fate determination by influencing enhancer activity18,19. In mouse embryonic stem cells (ESCs), CHD7 co-localizes with transcription factors including OCT4, SOX2, and NANOG at active enhancers to promote expression of pluripotency markers in ESC-specific genes19. In addition, CHD7 is a reader of H3K4me1, a canonical histone mark of enhancers, and its binding pattern varies with developmental stage from ESCs to neural progenitor cells (NPCs)19,20. CHD7 dosage reduction leads to a fate switch from neural epithelial to neural crest cell lineages in induced pluripotent stem cells (iPSCs) derived from individuals with CHARGE and CHD7 pathogenic variants21. CHD7 also regulates central nervous system lineages in patient derived iPSCs via SOX21 and BRN2 activation at super enhancers22.Together, these observations suggest that CHD7 regulates enhancer activity and expression of nervous system genes in iPSCs. However, the temporal and genome-region specific requirements for CHD7 in developing neurons.
To better understand CHD7 function in mammalian neurogenesis, we derived Chd7 null ESCs from Chd7 mutant blastocysts. Neuronal lineage differentiation was induced in vitro and the effects of CHD7 on gene expression (using Bru-Seq) and chromatin accessibility (using ATAC-seq) were analyzed. We found that loss of Chd7 in ESCs leads to decreased neuronal and glial cell differentiation and impaired neuronal complexity and neurite length in differentiated neurons. We also observed modified chromatin accessibility and altered expression of neural-specific genes with loss of Chd7. Our results suggest developmental stage-specific roles for CHD7 in neuronal development via changes in chromatin accessibility and gene transcription.
Results
Loss of Chd7 does not disrupt development or proliferation of ESCs and NPCs
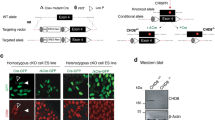
Chd7 is highly expressed in developing neurons in vivo14; thus, we hypothesized that CHD7 may play an active role in the differentiation of embryonic stem cells (ESCs) to neural progenitor cells (NPCs) and subsequently to neurons and glia (Fig. 1a). To test this hypothesis, we generated ESCs from mouse Chd7+/+ and Chd7Gt/Gt blastocysts. Western-blotting (Fig. 1b, Supplementary Fig. 1) and immunostaining (Fig. 2a) showed an absence of CHD7 protein, and qRT-PCR showed significantly reduced Chd7 mRNA (Fig. 1c), in Chd7Gt/Gt vs. Chd7+/+ ESCs. These observations validate prior studies showing that the Chd7Gt allele gives rise to a β-gal-Chd7 fusion transcript but no detectable CHD7 protein, confirming the null status of Chd7Gt/Gt ESCs13.
CHD7 promotes differentiation of neurons from ESCs. (a) Flow chart of ESC differentiation over 14 days into primary and secondary EBs, neurospheres, NPCs, neurons and glia. The protocol involves sequential treatment of ESCs with specified media. After secondary EB formation, EBs are cultured in suspension to form neurospheres to generate NPC cell lines or plated on Poly-l-lysine coated coverslips for direct induction into PAX6+ NPCs and subsequent differentiation into TUJ1+ neurons or GFAP+ glia. (b) Western blot showing absence of CHD7 protein in Chd7Gt/Gt ESCs. (c) ESC pluripotency genes are expressed based on RT-qPCR of steady state RNA in both Chd7+/+ and Chd7Gt/Gt ESCs (N = 3). Significant differences were found in all of the genes. *P < 0.05, **P < 0.01, ***P < 0.001. (d) Chd7Gt/Gt and Chd7+/+ ESCs grow at similar rates over 6 days in culture (N = 3 for each time point). (e) Chd7+/+ and Chd7Gt/Gt NPCs grow at similar rates over 6 days in culture (N = 3 for each time point). (f) Chd7+/+ and Chd7Gt/Gt ESCs, embryoid bodies (EBs) and NPCs exhibit similar morphologies. Scale bars = 50 µm. (g) Immunostaining for PAX6 (left) in NPCs derived from Chd7+/+ and Chd7Gt/Gt ESCs. Quantification of immunostaining is shown on the right. NPCs show no difference between Chd7+/+ and Chd7Gt/Gt in the proportion of PAX6+ cells (P = 0.94). Data of three independent experiments are represented as mean ± SEM. Unpaired student’s t-test was used for statistical analysis. Scale bars = 50 µm.
CHD7 promotes neuronal and glial differentiation in established NPC lines. (a,b) Immunostaining identifies CHD7 in Chd7+/+ but not in Chd7Gt/Gt ESCs (a) or NPCs. (b). Scale bars = 50 µm. (c) Chd7 mRNA levels significantly increase as Chd7+/+ cells transition from ESCs to EBs and NPCs (N = 3). Chd7 mRNA levels also increase significantly between Chd7+/+ ESCs and NPCs (68.68 ± 13.08, P = 0.006) and between Chd7+/+ EBs and NPCs (65.77 ± 13.08, P = 0.007). ***P < 0.001. (d) RT-qPCR of neural progenitor genes in Chd7Gt/Gt vs Chd7+/+ ESC-derived NPCs. (e) Immunostaining for the glial marker GFAP in cells derived from Chd7+/+ and Chd7Gt/Gt NPCs, and for the neuronal marker TUJ1 in cells derived from Chd7+/+ and Chd7Gt/Gt NPCs. Scale bars = 50 µm. (f) Quantification of the immunostaining in (e). There is a significant reduction in the proportion of GFAP+ (− 0.14 ± 0.02, P = 5.8 × 10–9) and TUJ1+ cells (− 0.14 ± 0.04, P = 9.7 × 10–4) in Chd7Gt/Gt compared to Chd7+/+. Unpaired student’s t-test was used for statistical analysis. ***P < 0.001.
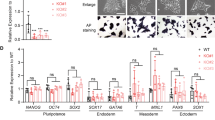
As a first step toward characterizing Chd7 null ESCs, we measured expression of the pluripotency markers Oct4, Sox2, Nanog, and c-Myc. All markers showed modest, albeit significant, differences in expression in Chd7Gt/Gt ESCs compared to Chd7+/+ cells (P < 0.05), with fold changes ranging from 0.69 and 1.35 (Fig. 1c). Neither Chd7+/+ or Chd7Gt/Gt ESCs showed defects in proliferation at passages up to P21 when measured over 6 days in standard ESC culture medium (Fig. 1d). In addition, there were very few (~ 2%) apoptotic cells as identified by immunostaining for cleaved Caspase 3 in Chd7+/+ and Chd7Gt/Gt ESCs (Supplementary Fig. 2a,b), suggesting that Chd7 deficiency does not alter ESC apoptosis. Moreover, no obvious differences in morphology were observed between Chd7+/+ and Chd7Gt/Gt ESCs (Fig. 1f). These data indicate that loss of Chd7 does not disrupt ESC morphology, proliferation, or cell death, but may have mild effects on pluripotency markers, which is consistent with other observations19.
ESCs were differentiated into NPCs via directed differentiation without Leukemia Inhibitory Factor (LIF) (Fig. 1f). Chd7+/+ and Chd7Gt/Gt ESCs were successfully differentiated to embryoid bodies (EBs) and NPCs, indicating that CHD7 is not essential for differentiation to neuroectoderm and progenitor fates. The percentage of Chd7+/+ and Chd7Gt/Gt NPCs expressing the marker PAX6 were similar (Fig. 1g), indicating that CHD7 is not necessary for the transition from ESCs to NPCs. Other NPC markers showed similar results (Supplementary Fig. 3). Over a 6-day period, Chd7+/+ and Chd7Gt/Gt NPC cell counts were comparable (Fig. 1e), suggesting no major defects in cell proliferation or survival with Chd7 loss. There were also no differences in the number of apoptotic NPCs (based on Caspase 3 immunostaining) in Chd7+/+ vs. Chd7Gt/Gt NPCs (Supplementary Fig. 2c,d). These observations demonstrate that CHD7 is dispensable for the generation and survival of NPCs from ESCs.
CHD7 is required for differentiation of NPC into neurons and glia
CHD7 is expressed at the protein and mRNA levels in Chd7+/+ NPCs, as confirmed by immunostaining (Fig. 2b) and qRT-PCR (Fig. 2d). As expected, CHD7 protein was absent in Chd7Gt/Gt NPCs (Fig. 2b) and Chd7 mRNA (reflecting a chimeric β-gal-Chd7 transcript from the Chd7Gt allele) was reduced in Chd7Gt/Gt NPCs (Fig. 2d). Interestingly, Chd7 mRNA levels gradually increased during neural differentiation of Chd7+/+ ESCs to EBs and NPCs (Fig. 2c). Expression of multipotency factors Brn2, Ncan, Sox1, and Sox11 was significantly different between Chd7+/+ and Chd7Gt/Gt NPCs (P < 0.05) (Fig. 2d). Notably, Sox1 mRNA levels were reduced to 4% of wild type levels in Chd7Gt/Gt NPCs (P = 1.1 × 10–6), consistent with previous studies showing that Sox1 promotes progenitor cell cycle exit and establishment of neuroectodermal lineages in mouse cells23.
Neurons and glia arise from common progenitors, with neurogenesis preceding gliogenesis during development24. In order to test the function of Chd7 in NPC transition to neural and glial lineages, we established NPC cell lines from Chd7+/+ and Chd7Gt/Gt ESCs. There were no differences in the proportion of Chd7+/+ and Chd7Gt/Gt cells expressing the neural markers NEUN, MAP2, Calretinin, or Neurofilament-200 (Supplementary Fig. 4). However, the proportion of TUJ1+ cells decreased from 24% in Chd7+/+ to 10% in Chd7Gt/Gt (SD = 4%, P = 9.7 × 10–4) (Fig. 2e,f). The proportion of GFAP+ cells was also reduced from 34% in Chd7+/+ to 20% in Chd7Gt/Gt (SD = 2%, P = 5.8 × 10–9) (Fig. 2e,f). These results suggest that CHD7 promotes NPC differentiation into both glial and neuronal lineages in mouse ESC-derived NPCs, instead of selectively regulating one lineage at the expense of the other.
Loss of Chd7 impairs the number, length and complexity of neurites in NPC-derived neurons
Reduced percentages of cells expressing the neural marker TUJ1 and the glial marker GFAP in Chd7Gt/Gt cultures suggests that CHD7 has broad functions in ESC-derived cell differentiation. To address this, we assayed the morphology of ESC-derived neurites using Sholl analysis on MAP2-positive cultures. Use of MAP2 instead of TUJ1 allowed us to label proportionately similar numbers of cells in cultures from both genotypes. Typical tracings of MAP2-positive neurons from DIV5 Chd7+/+ and Chd7Gt/Gt neurons are shown in Fig. 3a. Sholl analysis of neuritic complexity shows a reduction in the number of intersections for neurites in Chd7Gt/Gt compared to Chd7+/+ (Fig. 3b). The average individual neurite length at DIV5 was significantly reduced from 111.8 (± 27.1) μm in Chd7+/+ neurons to 72.2 (± 16.8) μm in Chd7Gt/Gt neurons (Fig. 3c). Total neurite length was also reduced from 229.4 (± 46.2) μm in Chd7+/+ neurons to 126.2 (± 37.8) μm in Chd7Gt/Gt neurons (Fig. 3d). The average number of primary neurites extending from the soma was reduced from 2.8 (± 0.6) in Chd7+/+ neurons to 2.3 (± 0.5) in Chd7Gt/Gt neurons (Fig. 3e). The total number of neurite branches was also reduced from 3.8 (± 1.1) in Chd7+/+ neurons to 2.6 (± 0.6) in Chd7Gt/Gt neurons (Fig. 3f). Together, these data show that CHD7 is necessary for normal neurite length, complexity, and branching in ESC-derived neurons.
Loss of Chd7 impairs the number, length, and complexity of neurites in NPC-derived neurons. (a–f) Neurons at DIV5. (g–l) Neurons at DIV9. (a) Representative images of Chd7+/+ neurons (left) and Chd7Gt/Gt neurons (right). For Sholl analysis, the total number of neurite crossings was counted at each circle with a radius increasing in increments of 10 μm. Scale bar = 20 μm. (b) Sholl analysis of the neuritic complexity of MAP2+ neurons. The data are shown as mean ± SEM (N = 20 per genotype). Two-way ANOVA was used for data analysis. ****P < 0.0001. (c,d) Loss of Chd7 results in a significant decrease in the maximum and total neurite length of differentiated neurons. Each symbol represents a single MAP2+ neuron (N = 20 per genotype). Unpaired student’s t-test was used for statistical analysis. ****P < 0.0001. (e,f) Loss of Chd7 causes a significant reduction in the number of primary neurites from the soma and total number of neurite branches of differentiated neurons. Each symbol represents a single MAP2+ neuron (N = 20 per genotype). Unpaired student’s t-test was used for statistical analysis. *P < 0.05; ***P < 0.001. (g) Representative images of Chd7+/+ neurons (left) and Chd7Gt/Gt neurons (right) at DIV9. Radius increments are 10 μm for Sholl analysis, Scale bar = 20 μm. (h) Sholl analysis of MAP2+ neurons. The data are shown as mean ± SEM (N = 20 per genotype). ****P < 0.0001 by two-way ANOVA. (i,j) The maximum and total neurite length of differentiated neurons were significantly reduced in Chd7Gt/Gt neurons compared to Chd7+/+ neurons. Each symbol represents a single MAP2+ neuron (N = 20 per genotype). Unpaired student’s t-test was used for statistical analysis. ***P < 0.001; ****P < 0.0001. (k,l) The number of primary neurites from the soma and total number of neurite branches were decreased in Chd7Gt/Gt neurons compared to Chd7+/+ neurons. Each symbol represents a single MAP2+ neuron (N = 20 per genotype). Unpaired student’s t-test was used for statistical analysis. ****P < 0.0001.
To further assess the role of Chd7 in neuronal differentiation, we extended the culture time to DIV9. Typical tracings of MAP2-positive Chd7+/+ and Chd7Gt/Gt neurons are shown in Fig. 4g. We observed similar and more severe impairments in neurite number, branching, and complexity at DIV9 compared to DIV5. Sholl analysis of neuritic complexity shows a reduction in the number of intersections for Chd7Gt/Gt compared to Chd7+/+ (Fig. 3h).The average maximum neurite length was reduced from 135.1 (± 23.2) μm in Chd7+/+ neurons to 103.1 (± 28.2) μm in Chd7Gt/Gt neurons (Fig. 3i). Total neurite length was reduced from 581.5 (± 132.3) μm in Chd7+/+ neurons to 224.1 (± 55.6) μm in Chd7Gt/Gt neurons (Fig. 3j). Additionally, the number of primary neurites extending from the soma ranged from 2 to 4 in Chd7Gt/Gt neurons and from 3 to 7 in Chd7+/+ neurons (Fig. 3k). The total number of neurite branches was reduced on average from 10.2 (± 4.4) in Chd7+/+ neurons to 3.55 (± 1.2) in Chd7Gt/Gt neurons (Fig. 3l). Taken together, these results suggest that Chd7 is required over a nine-day period for multiple aspects of neuronal morphogenesis including neurite extension, branching, and complexity.
Genes with high CHD7 occupancy display low RNAPII pausing and increased RNAPII throughout the gene body. (a–d) Pipeline used to identify expressed genes occupied by RNAPII and CHD7 in mouse ESCs. (e) Meta-analysis profile of RNAPII occupancy along gene territories in ESCs as determined from previously published RNAPII ChIP-Seq data. Normalized RNAPII binding averages in 150 bins are shown to illustrate the general RNAPII distribution along genes with high (red) and low (orange) CHD7 occupancy as determined from ChIP-seq. (f) Boxplots comparing RNAPII promoter pausing indices from the three different RNAPII ChIP-Seq datasets in relationship to two CHD7-associated gene sets (high vs low). P value was determined using two-sample Wilcoxon test.
CHD7 is enriched at genes with high RNA polymerase occupancy
The Drosophila orthologue of Chd7, kismet, has also been shown to positively regulate neural differentiation and promote transcriptional elongation on polytene chromosomes25. Given these observations, we asked whether mammalian CHD7 may also affect rates of transcriptional elongation, in addition to its known roles in activation or repression of target genes. A key control point for transcriptional elongation is RNA polymerase II (RNAPII) release from promoter-proximal pause sites26, whereby RNAPII pausing generates stable DNA secondary structures upon unwinding of double-stranded DNA. In mouse cerebellar granule cells, CHD7 interacts with Topoisomerase to facilitate chromatin unwinding18. Thus, CHD7 could regulate RNAPII pause release to help resolve DNA structures that stall transcriptional elongation.
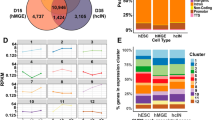
To determine whether CHD7 localization correlates with RNAPII occupancy and gene expression, we accessed publicly available ChIP-Seq data for CHD7 and RNAPII as well as RNA-seq data, all derived from mouse ESCs. We used these data to calculate pausing index, defined as the ratio of Reads Per Kilobase of transcript, per Million mapped reads (RPKM) at the transcription start site (TSS), to RPKM at the gene body, for genes with high vs low CHD7 occupancy over the whole gene (Fig. 4). Mouse ESC RNA-Seq data were obtained from GEO dataset GSM881355 (Fig. 4a). From these data, we selected genes expressed with FPKM ≥ 1.0 and length > 750 bp. Next, genes bound by RNAPII were identified by analysis of three RNAPII ChIP-Seq data sets (GSM699166, GSM1534642, GSM1566094) for RPKM at TSS > 1.6 and RPKM at the gene body > 0.5 (Fig. 4b). Genes expressed in ESCs and enriched for RNAPII at both the TSS and the gene body were identified by comparison of these datasets. This analysis revealed 3915 genes in GSM1534642, 6318 genes in GSM1566094, and 5133 genes in GSM699166 which were expressed in ESCs and also enriched for RNAPII at the TSS and gene body. These genes were then ranked in order from highest to lowest CHD7 occupancy levels throughout the gene body based on ESC Chd7 ChIP-Seq dataset GSM558674 (Fig. 4c). The top 500 and bottom 500 CHD7-bound genes were then selected for further calculation of pausing indices (i.e. ratio of RNAPII RPKMs at the TSS (− 50 bp to + 200 bp) to RPKM at the gene body (+ 500 bp to transcription termination site)) (Fig. 4d).
Meta-analysis of all three datasets showed a low pausing index for genes with high CHD7 occupancy in the gene body and a high pausing index for genes with low CHD7 occupancy in the gene body (Fig. 4e). For all three RNAPII ChIP-seq datasets, gene body RPKM for RNAPII were higher for those genes bound by CHD7 than for genes not bound by CHD7 (Fig. 4f). These results suggest a correlation between high CHD7 occupancy and slowed RNAPII processivity in gene bodies. Thus, CHD7 may participate in general chromatin unwinding and remodeling in ESC-derived cells and/or control one or more aspects of transcriptional elongation18.
CHD7 regulates genes expression
CHD7 enrichment in the gene body of ESCs suggests it may alter rates of RNA transcriptional elongation in addition to nascent RNA transcription. Because prior studies of gene expression in Chd7 mutant cells and tissues focused on steady state RNA, which does not distinguish newly transcribed from older RNA molecules, we asked whether CHD7 may function in ESC-derived NPCs to regulate transcriptional activation, elongation, or both. Bru-Seq is a recently developed large scale sequencing method that uses Bromouridine (Bru) to label newly transcribed RNA36. Thus, we used Bru-Seq in Chd7+/+ and Chd7Gt/Gt NPCs (N = 3) to test the effects of chronic Chd7 loss on nascent transcription. Principal components plots showing sample clustering, along with summaries of replicates, are presented in Supplementary Fig. 3 and Table 2.
Bru-Seq confirmed higher nascent Chd7 expression in Chd7+/+ NPCs compared to Chd7Gt/Gt NPCs, consistent with the overall reduction of Chd7 observed in Chd7Gt/Gt ESCs (Fig. 2d). PCA of the Bru-Seq data show a clear separation of genotypes along principal component one and principal component two, indicating consistency across replicates (Supplementary Fig. 5). The total number of genes analyzed in Chd7+/+ and Chd7Gt/Gt NPCs was 17,346. There were 6,122 differentially expressed genes between Chd7+/+ and Chd7Gt/Gt NPCs (35%) (FDR < 0.05), with 2,716 (44%) being down-regulated in Chd7Gt/Gt and 3,406 (56%) being up-regulated in Chd7Gt/Gt NPCs (Fig. 5a). This result is consistent with previous studies showing that CHD7 both positively and negatively regulates expression of other genes3,18,27.
CHD7 loss affects accessibility and expression of key NPC genes. (a) Volcano plot showing log2 fold changes and FDRs for genes in the Bru-Seq analysis. (b) Pie chart showing the distribution of peak locations from ATAC-seq peaks that were associated with genes expressed in NPCs by Bru-Seq. (c) Scatter plot showing peaks from the combined analysis of ATAC-seq and Bru-Seq data. The y-axis is the log2 fold change in peaks. The x-axis is the log2 fold change in gene expression. Peaks are labeled with different colors based on significance thresholds of FDR < 0.05. The upper-right quadrant shows genes with higher accessibility in their promoter region and higher expression in Chd7Gt/Gt compared to Chd7+/+. The lower left quadrant shows genes with lower accessibility in their promoter region and lower expression in Chd7Gt/Gt compared to Chd7+/+. (d) GO Term analysis of “doubly significant” genes. The p-values were corrected with a Bonferroni adjustment. The black-dashed line shows the cutoff of a corrected p-value of 0.05. Blue bars represent GO terms associated with genes that were up-regulated in Chd7Gt/Gt; red bars represent GO terms associated with down-regulated genes. (e) Plot of ATAC-seq and Bru-Seq read tracks along the gene body of Tfap2b using the UCSC Genome Browser. The promoter region is highlighted in yellow. The image shows greater accessibility of the promoter region and higher expression in the Chd7Gt/Gt cell line.
CHD7 loss disrupts chromatin accessibility at differentially expressed genes
CHD7 uses the energy of ATP hydrolysis to evict, slide, or rearrange nucleosomes on chromatin16 . In previous studies, CHD7 binding sites were found in transcriptional start sites as well as intergenic sites and intragenic regions20. Most CHD7 binding sites in mouse NPCs have been confirmed as CHD7 binding sites in human cells20. To test whether CHD7 modifies the chromatin landscape in mouse NPCs, we used ATAC-seq, an assay for genome-wide mapping of chromatin accessibility, on both Chd7+/+ and Chd7Gt/Gt NPCs (N = 3) and compared peaks across genotypes. Each peak was associated with a nearby gene and annotated by its location relative to that gene. There were 121,161 combined peaks identified in Chd7+/+ and Chd7Gt/Gt NPCs. Of these, 99,770 peaks were associated with genes expressed by Bru-Seq assay. Of these peaks, 45% (45,228) were located in intronic regions, 29% (28,876) were in distal intergenic regions, and 16% (16,130) were in promoter regions (Fig. 5b).
To identify misregulated genes whose promoters were also differentially accessible in Chd7Gt/Gt cells, we further analyzed 12,299 genes that were expressed by Bru-Seq analysis and were associated with at least one ATAC-seq peak in a promoter region. Among these genes, some contained multiple promoter peaks, typically reflecting different transcription start sites for isoforms. Altogether, we observed a total of 16,130 ATAC-seq peaks associated with these 12,299 expressed genes. A scatterplot was created using fold changes for all peak/gene combinations from this combined analysis (Fig. 5c). A Kendall’s tau correlation was performed to determine the relationship between fold change in ATAC-seq peak accessibility and Bru-Seq gene expression between Chd7+/+ and Chd7Gt/Gt NPCs. There was a significant, positive correlation between ATAC-seq and Bru-Seq fold changes (Kendall’s tau = 0.11, P < 2.2 × 10–16), consistent with increased accessibility at the promoter and higher levels of gene expression. Of particular interest were 2,742 differentially expressed genes which were also associated with 3,119 differentially accessible peaks. We call these genes “doubly significant”. GO term analysis of these doubly significant genes was performed for both up- and down-regulated genes. The down-regulated genes (N = 1094, 40%) showed enrichment in many different processes including nervous system development, neurogenesis and cell differentiation. The up-regulated genes (N = 1,648, 60%) showed an enrichment in mitotic cell cycle process, DNA repair and mitotic nuclear division (Fig. 5d).
Review of the top doubly significant genes and significant GO terms (i.e. neurogenesis) revealed several genes of particular interest (Table 1), five of which encode transcription factors (Tfap2b, Pax3, Zic5, Tbx3 and Nkx6-1) that control neuronal or glial cell fate28,29,30,31,32,33,34. Another gene, Stmn2, was downregulated in Chd7Gt/Gt cells and encodes a stathmin protein family member that promotes neuronal growth35. These six genes were further analyzed by examination of chromatin accessibility and gene expression using the UCSC Genome Browser. Representative read tracks for Tfap2b highlight differences in chromatin accessibility and gene expression (Fig. 5e). All read tracks for Tfap2b are shown in Supplementary Fig. 6. Several other doubly significant genes also showed differences in expression and accessibility (Supplementary Fig. 7). Based on these combined genome level data, we speculate that CHD7 alters the chromatin landscape in NPCs by modifying chromatin accessibility at genes that regulate neuron-specific and glial-specific processes.
Discussion
This study demonstrates that CHD7 is dispensable for maintenance/survival of mouse ESC pluripotency and differentiation to neural progenitor cells yet promotes differentiation of neurons and glia from embryonic stem cell-derived NPCs. CHD7 also promotes neurite length, branching, and complexity in ESC-derived neurons, and regulates gene expression and chromatin accessibility at key neural and glial developmental genes. This developmental stage-specific requirement for CHD7 may reflect the need for a changing chromatin landscape as cells transition from pluripotent to lineage restricted cells, highlighting unique functions for CHD7 in later stages of the neuronal/glial differentiation process.
We found no evidence for defects in growth (cell number) in Chd7Gt/Gt ESCs or NPCs, suggesting that CHD7 is not necessary for ESC or NPC proliferation. Interestingly, individuals with CHARGE syndrome and mouse models of CHD7 deficiency show cerebellar hypoplasia but not microcephaly or macrocephaly, and CHD7 promotes granule cell, but not cortical neural progenitor cell, proliferation3,18,36. In the mouse cerebellum, loss of Chd7 leads to decreases in the number of dividing granule cell precursors3,18. In contrast, conditional knockout of Chd7 in hippocampal glia and neural progenitors is associated with a transient increase in cell division and genesis of immature neurons, with depletion of the stem cell pool in the sub-granular zone12. Germline and conditional Chd7 knockout mice also show decreased neurogenesis in the adult sub-ventricular zone and hippocampus10,11, dentate gyrus12, and inner ear15. In non- neuronal cell types like ESCs, however, loss of Chd7 has no impact on cell growth19. These observations together support the conclusion that CHD7 can promote, inhibit, or have no effects on cell proliferation, depending on the context and cell type.
Apoptosis is key to proper organismal development and tissue maintenance37 and involves similar cellular processes as differentiation10. We observed very few apoptotic ESCs or NPCs in either Chd7+/+ or Chd7Gt/Gt cells, and no difference in the number of apoptotic cells between genotypes. Contrary to this finding, loss of CHD7 is associated with increased cell death in mouse cerebella as measured by the number of Caspase-positive cells3,18. Differences in apoptosis rates across studies may reflect differing sensitivities in neuronal cell type to Chd7 loss.
We observed changes in expression of ESC-related genes like Oct4, Sox2, and Nanog between Chd7 null and wild-type ESCs. It is unclear why these pluripotency markers differ in expression between Chd7+/+ and Chd7Gt/Gt cells, but this could reflect direct regulation of CHD7 on the promoters of these genes, or other subtle forms of regulation. Schnetz et al. reported normal mRNA levels for Oct4, Sox2, and Nanog in Chd7 null ESCs, and showed that loss of Chd7 does not preclude ESC pluripotency and differentiation into embryoid bodies19. Differences between their study and ours may be related to the underlying genetic defect (nonsense vs gene-trapped alleles), genetic background, or other experimental conditions. Interestingly, CHD7 co-localizes with OCT4, SOX2, NANOG and P300 in mouse ESCs, and Oct4 is required for CHD7 binding to genomic segments that are bound by multiple transcription factors13,19,38. Moreover, loss of Chd7 does not impair advancement of embryonic development to the blastocyst stage from which ESCs are generated, since null embryos die after E10.538. Together, these observations support the conclusion that Chd7 is dispensable for ESC pluripotency and self-renewal.
In contrast to these early stages of development where Chd7 is not absolutely required, later stages of lineage formation and gene expression have been shown to depend on CHD7. For example, downregulation of CHD7 disrupts EB formation and expression of endoderm and mesoderm genes39. In our study, Chd7Gt/Gt ESCs were able to differentiate normally into PAX6-positive NPCs, but Chd7Gt/Gt NPCs showed reduced ability to differentiate into TUJ1-positive neurons. Similarly, in previous studies, neural stem and progenitor cells from the sub-ventricular zone of Chd7+/− and Chd7−/− mice showed reduced potential to differentiate into TUJ1-positive neurons11. Additionally, loss of Chd7 in mice leads to fewer neurons in the olfactory bulb11. Thus, normal levels of CHD7 are required to maintain multiple neuronal lineages in a variety of peripheral and central nervous system sites22. Our results showed that the number of ESC-derived neurons expressing NEUN, MAP2, Calretinin or Neurofilament-200 was unchanged with Chd7 loss, suggesting that only certain aspects of neuronal differentiation require Chd7. We cannot exclude the possibility that certain neuronal subtypes (e.g. TUJ1 +) or developmental stages (e.g. early differentiation) are more sensitive to Chd7 loss than others. Indeed, we observed defective neurite branching number, length, and complexity in Chd7Gt/Gt neurons in vitro, both at 5 and 9 days in culture (Fig. 4), and similar defects in neurite length and branching were reported in newly generated hippocampal neurons with loss of Chd710, consistent with our findings. It will be interesting to determine in future studies whether specific neuronal subtypes are preferentially affected by Chd7 loss. Our analysis of existing RNAPII and CHD7 ChIP-Seq data in ESCs showed enrichment of RNAPII in genes that are also occupied by CHD7. This analysis also uncovered lower RNAPII pausing indices for genes with high CHD7 occupancy. While correlative, these data suggest a model whereby CHD7 promotes release of RNAPII from the transcription start site and regulates the rate at which RNAPII travels through the gene body during transcription. Additional studies using controlled, inducible, CHD7 and RNAPII expression will help explore this possibility. It will also be helpful to determine whether enrichment of CHD7 in the gene body of highly expressed genes also occurs in differentiated cells like neurons and glia.
Our combined analysis using Bru-Seq and ATAC-seq on Chd7+/+ and Chd7Gt/Gt NPCs identified many genes whose expression and accessibility were significantly altered by Chd7 loss. Several of these genes are implicated in neural progenitor cell differentiation, and the biological processes (based on GO terms) associated with these gene expression changes include nervous system development and differentiation. One gene, Pax3, exhibited a striking loss of open chromatin near the promoter, and reduced expression in Chd7Gt/Gt NPCs. Pax3 has been shown to directly bind and regulate Hes1 and Neurog2, thereby promoting differentiation of sensory neurons40. Uncovering the extent to which Pax3 and other CHD7-sensitive genes mediate the effects of Chd7 in neural progenitors will require new methods that allow for simultaneous up- and down-regulation of multiple genes and genetic pathways. Such approaches will help clarify not only which genes and pathways are critical for CHD7 function but will also enable the identification of putative therapeutic targets.
Our studies also identified Stmn2, the gene encoding phosphoprotein stathmin family member 2, as a putative target of CHD7. STMN2 regulates microtubules and neuronal growth, is highly expressed in the developing brain, and is downregulated in postmortem brains from individuals with Parkinson’s Disease41. Stmn2 has been shown to function in synaptic trafficking and is necessary for axonal outgrowth and regeneration41, akin to the changes in complexity we observed in neurons lacking Chd7. Reduced Stmn2 expression may help explain the defects in neuronal complexity and neurite outgrowth observed with loss of Chd7 in ESC-derived neural and hippocampal cells.
We also observed reduced expression of Tfap2b, encoding transcription factor AP2-β in Chd7Gt/Gt ESC-derived neural cells. Pathogenic variants in TFAP2B cause human CHAR syndrome, an autosomal dominant disorder that presents in individuals with patent ductus arteriosus, facial dysmorphisms, and abnormalities of the 5th digits of the hands42. Some features of CHAR overlap with CHARGE, include developmental delay, broad forehead, short philtrum, low-set ears, and congenital heart disease. Of note, TFAP2B has been shown to promote differentiation of neural crest cells via heterodimerization with the pioneer transcription factor and family member TFAP2A43. Since neural crest cells contribute to craniofacial, cardiac, and skeletal structures, this cell population is implicated in both CHAR and CHARGE syndromes and could be considered as a candidate target population for therapeutics44,45.
In conclusion, this study provides the first evidence for CHD7 regulation of nascent transcription and chromatin accessibility in ESC-derived neural progenitors. We present several candidate genes as putative genetics targets for CHD7 that merit further analysis, including Tfap2b, Stmn2, Sox1, Pax3, Zic5, Tbx3, and Nkx6-1. These genes may mediate CHD7 effects in neuronal and glial differentiation, and a genome-wide, combinatorial approach will be helpful to further address this possibility. Our results suggest that therapies directed toward correction of human phenotypes of CHD7 deficiency, particularly those affecting the central and peripheral nervous systems, may also need to target multiple genetic pathways during early stages of cellular differentiation, rather than focusing on stem or progenitor cell stages.
Materials and methods
All reagents and culture media were purchased from Life Technologies/Invitrogen (Carlsbad) unless otherwise indicated. Methods were carried out in accordance with relevant guidelines and regulations, and experimental protocols involving mice (generation of embryonic stem cells from blastocysts) were approved by the University of Michigan Institutional Animal Care & Use Committee (IACUC).
Western blotting
Nuclear protein was extracted and quantified using the BCA Protein Assay Kit (Life Technologies). Protein samples (10–20 µg) were electrophoresed in 4–12% Bis–Tris gels and transferred overnight to a nitrocellulose membrane at 4 °C. Primary antibodies used were anti-HDAC2 (Santa Cruz) and anti-CHD7 (#6505, Cell Signaling). Secondary antibodies were goat anti-rabbit IgG HRP (#31461, Thermo Fisher) and rabbit anti-goat IgG HRP (#31403, Thermo Fisher).
Neuronal lineage differentiation
Chd7+/+ and Chd7Gt/Gt ESCs were derived from sibling mouse blastocysts in the Transgenic Animal Model Core (TAMC) of The University of Michigan. Chd7+/+ (passage 10) and Chd7Gt/Gt (passage 10) cells were sent to Cell Line Genetics (Madison, Wisconsin, US) for karyotyping based on cytogenetic analysis on G-banded metaphase cells. Both cell lines were > 85% euploid and were used only up to passage 20–21.
ESCs were cultured in knockout DMEM, 20% FBS (Fetal bovine serum, Atlas Biologicals, Fort Collins, CO), Glutamax, MEM Nonessential amino acids, 2-Mercaptoethanol, and Recombinant Mouse Leukemia Inhibitory Factor (LIF) without feeder cells. ESCs were differentiated to neuronal cells following an established protocol46. Mouse ESCs were dissociated to single cells using 0.05% Trypsin–EDTA, then seeded at a density of 3 × 105 cells /ml in embryoid body (EB) medium (withdrawal of LIF) onto bacteriological dishes. Cells were cultured as aggregates in suspension. EBs formed after 4 days, then 5 μM ATRA was added to the medium and cultures allowed to grow for 4 more days. EBs were dissociated and plated onto poly-L-Lysine (Sigma) coated coverslips in 6-well-plates at 1 × 105 cells per well in NPC medium (10 ng/ml basic fibroblast growth factor, 1% N2, 1% Penicillin/Streptomycin, supplemented by DMEM/F12). Neurospheres were maintained in DMEM/F12 supplemented with B27, 20 ng/ml EGF, 20 ng/ml bFGF, Glutamax and Penicillin/ Streptomycin for cell line establishment. NPCs were harvested for immunofluorescence for PAX6, NESTIN, VIMENTIN and SOX2.
Cells were cultured in NPC medium for 3 days, then switched to neuronal media (1 mM Glutamax, 1% Penicillin/ Streptomycin, 1% N2, 2% B27, supplemented with DMEM/F12) and differentiated into neurons and glia46. Cells were incubated at 37 °C with 5% CO2 and cultured on plates coated with poly-L-lysine. The medium was changed every other day. Neural cells were harvested on day in vitro (DIV) 5 for immunostaining with anti-TUJ1, GFAP, NEUN, MAP2, Calretinin and Neurofilament-200 (NF200).
Quantitative RT-PCR
Cells (at least 106) were collected by centrifugation, lysed by cold Trizol and incubated for 5 min to ensure complete lysis. An equal volume of Chloroform (Thermo Fisher Scientific) was added to the lysis product and vortexed for 15 s. After centrifugation, the supernatant was removed and added to the same volume of Isopropanol (Thermo Fisher Scientific) to precipitate RNA. After centrifugation, the supernatant was discarded, and the pellet washed with 70% DNase/RNase free ethanol solution. After drying for 5 min, RNA was dissolved in 20 µl nuclease free H2O.
cDNA was synthesized using the SuperScript First-Strand Synthesis System kit (Thermo Fisher Scientific) and stored at − 20 °C. DNA samples were quantified using a Nanodrop 2000c (Thermo Fisher Scientific). qRT-PCR experiments were performed using SYBR Green Master Mix (Applied Biosystems, Foster City, CA) on an ABI 7500 qPCR machine (Applied Biosystems).
Primers are shown in Supplementary Table 1. Gene expression was normalized to Gapdh. Statistical significance was determined using unpaired, two-tailed Student’s t-tests. P-values were corrected using the Holm-Sidak method47.
Cell growth assays
For cell growth assays, 50,000 ESCs or 100,000 NPCs were plated per well in a 6-well plate. Wells were coated with 0.1% gelatin for ESCs or with poly-L-lysine (Invitrogen, Carlsbad, CA) for NPCs. Cells were maintained in culture media for the length of the assay. Cell numbers were assayed every two days. Colonies were digested to single cells with 0.05% trypsin–EDTA for ESCs and Accutase for NPCs. Cells were diluted in culture media and counted with a hemocytometer. Statistical significance was determined using unpaired, two-tailed Student’s t-tests. P-values were corrected using the Holm-Sidak method47.
Immunostaining
Cells were plated on poly-l-lysine coated coverslips, washed three times with PBS, then fixed in 4% paraformaldehyde (PFA, Sigma) in phosphate buffer (pH 7.4) for 30 min at room temperature. Cells were stored at 4 °C in 4% paraformaldehyde or immediately used for immunostaining. Coverslips were washed with PBS three times, then treated with 0.1% Triton-X-100 (Sigma) for 15 min. Coverslips were blocked with 2% bovine serum albumin (BSA) for one hour and incubated with primary antibodies: rabbit anti-PAX6 antibody (1:300, #901301, Biolegend, San Diego, CA), rat anti-Nestin antibody (1: 200, #rat-401, Developmental Studies Hybridoma Bank, Iowa City, IA), goat anti-Vimentin, (1:100, #V4630, Sigma-Aldrich, St. Louis, MO), goat anti-SOX2 (1:200, #GT15098, Neuromics, Edina, MN), mouse anti-Tuj1 (1:200, #801202, Biolegend, San Diego, CA), mouse anti-NEUN (1:100, #MAB3777, Millipore Billerica, MA), mouse anti-MAP2 (1:200, #M4403, Sigma Aldrich St. Louis, MO), rabbit anti-Neurofilament 200 (1:100, #N4142, Sigma Aldrich St. Louis, MO), rabbit anti-Calretinin (1:100, #AB5054, Sigma Aldrich St. Louis, MO), rabbit anti-cleaved Caspase 3 (1:200, Cell Signaling, Danvers, MA), and mouse anti-GFAP (1:200, #G6171, Sigma Aldrich St. Louis, MO) overnight at 4 °C. After incubation with primary antibody, cells were washed three times with PBS, then incubated with secondary antibody for 2 h at room temperature. Secondary antibodies used were as follows: goat anti-rabbit IgG H&L Alexa Fluor 555 (1:300, #ab150078, Abcam), goat anti-mouse IgG H&L Alexa Fluor 555 (1:500, #ab150118, Abcam), donkey anti-goat Alexa Fluor 488 (1:200, #ab150129, Abcam), and donkey anti-mouse Alexa Fluor 488 (1:200, #ab150105, Abcam). Coverslips were washed with PBS (3 × 10 min) and incubated with 1:10,000 4′,6-diamidino-2-phenylindole (DAPI; Thermo Fisher Scientific, Waltham, MA) for 5 min. Samples were washed with PBS and mounted with Prolong Gold Antifade reagent. Cells were imaged using a Leica DMB upright fluorescence microscope. Statistical significance was determined using paired, two-tailed Student’s t-tests.
Sholl analysis
Neural cells were harvest at DIV5 and DIV9 and immunostained with anti-MAP2 as described above. The longest neurite for each MAP2 positive cell was measured using Sholl analysis, as previously described48 with Fiji software (National Institutes of Health, Bethesda, MD, USA). Sholl analysis of single neuron branching was performed manually using Simple Neurite Tracer.
Bru-Seq
Chd7+/+ and Chd7Gt/Gt mouse ESC and NPC cells were collected and at least 6 × 106 cells for genotype were used for the Bru-Seq analysis as previously reported42. Briefly, cells in the logarithmic growth phase were treated with bromouridine to a final concentration of 2 mM for 30 min to label newly synthesized nascent RNA. Cells were then lysed in TRIzol (Invitrogen) and total RNA was isolated. Bru-labeled, nascent RNA was then immunocaptured using anti-BrdU antibodies (BD Biosciences, San Jose, CA, USA) conjugated to magnetic beads (Invitrogen) and then converted into cDNA libraries and sequenced at the University of Michigan Sequencing Core using an Illumina HiSeq 2000 sequencer (San Diego, CA). Sequencing reads were mapped to the mm10 reference sequence and the relative rates of nascent transcription genome-wide were calculated as previously described42.
ATAC-seq
Chd7+/+ and Chd7Gt/Gt neural progenitor cells (differentiated from ESCs) were collected according to protocol49. Briefly, 50,000 cells for each genotype were collected and lysed in 50 µl cold ATAC-Resuspension Buffer containing 0.1% NP40 (Sigma), 0.1% Tween-20 (Sigma), and 0.01% Digitonin (Promega). Lysis product was incubated with Tn5 transposase (Illumina, Nextera XT DNA Library Prep Kit) for 30 min at 37 °C. cDNA from Tn5-treated cells was purified and used as a template to amplify a DNA library by PCR with 2 × NEB master mix. Libraries were deep-sequenced with Next-seq 500 and high output kit (75 bp) with paired-end 36 bp reads. Three replicates for each cell line were collected and processed.
ATAC libraries were prepared as described previously50. In brief, 50,000 cells per sample were processed for the transposase reaction. ATAC-seq libraries were PCR amplified with barcoded primers. ATAC-seq libraries were sequenced (paired-end 36 bp on a HiSeq4000at the UM Sequencing Core) and raw reads trimmed using cta (https://github.com/ParkerLab/cta)50 to eliminate barcodes and aligned to the mm9 reference genome using BWA-MEM (v. 0.7.15)51. Picard was used to remove duplicates, and reads were subsequently filtered for high mapping quality (mapq ≥ 30) and properly paired alignments, as described52. In order to minimize potential confounding by sequencing depth, libraries were uniformly down sampled to the minimum sequencing depth (31.9 M reads) for downstream analysis steps.
Peaks were called using MACS253 with options –nomodel –shift -100 –extsize 200 –broad -B –keep-dup all on the down sampled BAM files. Peak calls that met a 5% False Discovery Rate and did not intersect genomic regions with known mappability artifacts (blacklisted regions from https://sites.google.com/site/anshulkundaje/projects/blacklists) were retained for further analyses. A virtual session of the quality control metrics for the ATAC-seq samples is provided at the following URL: https://theparkerlab.med.umich.edu/data/albanus/martin/ataqv_npc/.
There were 88,641 peaks in the Chd7+/+ NPCs and 81,773 peaks in the Chd7Gt/Gt NPCs. The peak regions from Chd7+/+ and Chd7Gt/Gt cells were merged together to obtain a common set of peaks for NPCs, in order to compare peak reads across genotypes. The coverage from each replicate was then determined for the combined peaks. This analysis was done on Python (v. 2.7.14 Anaconda) with Bedtools (v. 2.25.0)54. The raw counts were then merged into a matrix which was analyzed in R (v. 3.5) using DESeq2 (v. 1.20.0)55. Peaks were matched to genes with ChIPseeker (v. 1.18.0)56 with a promoter definition of 1000 bp upstream to 1 bp downstream from the transcription start site57.
UCSC genome browser
Sequencing tracks were generated for the ATAC-seq and the Bru-Seq in bigwig format. These files were then uploaded and viewed in the UCSC Genome Browser, using the mm10 assembly58.
GO term analysis
The Generic Gene Ontology (GO) Term Finder developed at the Lewis-Sigler Institute at Princeton University59 was used to analyze the data. All enrichment calculations were done using only expressed genes from the Bru-Seq analysis as the background.
Statistical analysis
Data were analyzed using GraphPad Prism and R. Unless otherwise indicated, graphs show mean and standard deviations. Unpaired t-tests were used and were adjusted for multiple comparisons with the Holm-Sidak method, unless otherwise indicated. P-value adjustment with the Benjamini–Hochberg method was used to calculate the upper bound for the expected false discovery rates (FDR). Kendall’s tau coefficient was used to measure the ordinal association between measured quantities due to a lack of normality in the data. All statistical tests were two-sided.
Data availability
The datasets generated and/or analyzed during the current study are available from the corresponding author upon request.
References
Pagon, R. A., Graham, J. M. Jr., Zonana, J. & Yong, S. L. Coloboma, congenital heart disease, and choanal atresia with multiple anomalies: CHARGE association. J. Pediatr. 99, 223–227 (1981).
Legendre, M. et al. Phenotype and genotype analysis of a French cohort of 119 patients with CHARGE syndrome. Am. J. Med. Genet. C 175, 417–430 (2017).
Whittaker, D. E. et al. The chromatin remodeling factor CHD7 controls cerebellar development by regulating reelin expression. J. Clin. Invest. 127, 874–887 (2017).
de Geus, C. M. et al. Guidelines in CHARGE syndrome and the missing link: Cranial imaging. Am. J. Med. Genet. C 175, 450–464 (2017).
van Ravenswaaij-Arts, C. & Martin, D. M. New insights and advances in CHARGE syndrome: diagnosis, etiologies, treatments, and research discoveries. Am. J. Med. Genet. C 175, 397–406 (2017).
Hartshorne, T. S., Hefner, M. A. & Davenport, S. L. Behavior in CHARGE syndrome: introduction to the special topic. Am. J. Med. Genet. A 133A, 228–231 (2005).
Hartshorne, T. S., Stratton, K. K., Brown, D., Madhavan-Brown, S. & Schmittel, M. C. Behavior in CHARGE syndrome. Am. J. Med. Genet. C 175, 431–438 (2017).
Hale, C. L., Niederriter, A. N., Green, G. E. & Martin, D. M. Atypical phenotypes associated with pathogenic CHD7 variants and a proposal for broadening CHARGE syndrome clinical diagnostic criteria. Am. J. Med. Genet. A 170, 344–354 (2016).
Vissers, L. E. et al. Mutations in a new member of the chromodomain gene family cause CHARGE syndrome. Nat. Genet. 36, 955–957 (2004).
Feng, W. et al. The chromatin remodeler CHD7 regulates adult neurogenesis via activation of SoxC transcription factors. Cell Stem Cell 13, 62–72 (2013).
Micucci, J. A. et al. CHD7 and retinoic acid signaling cooperate to regulate neural stem cell and inner ear development in mouse models of CHARGE syndrome. Hum. Mol. Genet. 23, 434–448 (2014).
Jones, K. M. et al. CHD7 maintains neural stem cell quiescence and prevents premature stem cell depletion in the adult hippocampus. Stem cells 33, 196–210 (2015).
Hurd, E. A. et al. Loss of Chd7 function in gene-trapped reporter mice is embryonic lethal and associated with severe defects in multiple developing tissues. Mamm. Genome 18, 94–104 (2007).
Layman, W. S. et al. Defects in neural stem cell proliferation and olfaction in Chd7 deficient mice indicate a mechanism for hyposmia in human CHARGE syndrome. Hum Mol Genet 18, 1909–1923 (2009).
Hurd, E. A., Poucher, H. K., Cheng, K., Raphael, Y. & Martin, D. M. The ATP-dependent chromatin remodeling enzyme CHD7 regulates pro-neural gene expression and neurogenesis in the inner ear. Development 137, 3139–3150 (2010).
Bouazoune, K. & Kingston, R. E. Chromatin remodeling by the CHD7 protein is impaired by mutations that cause human developmental disorders. Proc. Natl. Acad. Sci. USA 109, 19238–19243 (2012).
Clapier, C. R. & Cairns, B. R. The biology of chromatin remodeling complexes. Annu Rev Biochem 78, 273–304 (2009).
Feng, W. et al. Chd7 is indispensable for mammalian brain development through activation of a neuronal differentiation programme. Nat. Commun. 8, 14758 (2017).
Schnetz, M. P. et al. CHD7 targets active gene enhancer elements to modulate ES cell-specific gene expression. PLoS Genet. 6, e1001023 (2010).
Schnetz, M. P. et al. Genomic distribution of CHD7 on chromatin tracks H3K4 methylation patterns. Genome Res 19, 590–601 (2009).
Okuno, H. et al. CHARGE syndrome modeling using patient-iPSCs reveals defective migration of neural crest cells harboring CHD7 mutations. Elife 6, e21114 (2017).
Chai, M. et al. Chromatin remodeler CHD7 regulates the stem cell identity of human neural progenitors. Genes Dev. 32, 165–180 (2018).
Suter, D. M., Tirefort, D., Julien, S. & Krause, K. H. A Sox1 to Pax6 switch drives neuroectoderm to radial glia progression during differentiation of mouse embryonic stem cells. Stem Cells 27, 49–58 (2009).
Guerout, N., Li, X. & Barnabe-Heider, F. Cell fate control in the developing central nervous system. Exp. Cell Res. 321, 77–83 (2014).
Srinivasan, S. et al. The Drosophila trithorax group protein Kismet facilitates an early step in transcriptional elongation by RNA Polymerase II. Development 132, 1623–1635 (2005).
Adelman, K. & Lis, J. T. Promoter-proximal pausing of RNA polymerase II: emerging roles in metazoans. Nat. Rev. Genet. 13, 720–731 (2012).
Yao, H. et al. CHD7 represses the retinoic acid synthesis enzyme ALDH1A3 durng inner ear development. JCI Insight 3, e97440 (2018).
Li, Y., Tzatzalos, E., Kwan, K. Y., Grumet, M. & Cai, L. Transcriptional regulation of Notch1 expression by Nkx6.1 in neural stem/progenitor cells during ventral spinal cord development. Sci. Rep. 6, 38665 (2016).
Sander, M. et al. Ventral neural patterning by Nkx homeobox genes: Nkx6.1 controls somatic motor neuron and ventral interneuron fates. Genes Dev. 14, 2134–2139 (2000).
Esmailpour, T. & Huang, T. TBX3 promotes human embryonic stem cell proliferation and neuroepithelial differentiation in a differentiation stage-dependent manner. Stem Cells 30, 2152–2163 (2012).
Ohnishi, T. et al. A spontaneous and novel Pax3 mutant mouse that models Waardenburg syndrome and neural tube defects. Gene 607, 16–22 (2017).
Wei, C., Ren, L., Li, K. & Lu, Z. The regulation of survival and differentiation of neural stem cells by miR-124 via modulating PAX3. Neurosci. Lett. 683, 19–26 (2018).
Furushima, K., Murata, T., Kiyonari, H. & Aizawa, S. Characterization of Opr deficiency in mouse brain: subtle defects in dorsomedial telencephalon and medioventral forebrain. Dev. Dyn. 232, 1056–1061 (2005).
Inoue, T. et al. Mouse Zic5 deficiency results in neural tube defects and hypoplasia of cephalic neural crest derivatives. Dev. Biol. 270, 146–162 (2004).
Yue, F. et al. A comparative encyclopedia of DNA elements in the mouse genome. Nature 515, 355–364 (2014).
Whittaker, D. E. et al. Distinct cerebellar foliation anomalies in a CHD7 haploinsufficient mouse model of CHARGE syndrome. Am. J. Med. Genet. C Seminars in Med. Genet. 175, 465–477 (2017).
Potten, C. S. & Booth, C. The role of radiation-induced and spontaneous apoptosis in the homeostasis of the gastrointestinal epithelium: a brief review. Comp. Biochem. Physiol. B 118, 473–478 (1997).
Bosman, E. A. et al. Multiple mutations in mouse Chd7 provide models for CHARGE syndrome. Hum. Mol. Genet. 14, 3463–3476 (2005).
Yamamoto, T. et al. Differentiation potential of Pluripotent Stem Cells correlates to the level of CHD7. Sci. Rep. 8, 241 (2018).
Ichi, S. et al. Role of Pax3 acetylation in the regulation of Hes1 and Neurog2. Mol. Biol. Cell 22, 503–512 (2011).
Klim, J. R. et al. ALS-implicated protein TDP-43 sustains levels of STMN2, a mediator of motor neuron growth and repair. Nat. Neurosci. 22, 167–179 (2019).
Satoda, M. et al. Mutations in TFAP2B cause Char syndrome, a familial form of patent ductus arteriosus. Nat. Genet. 25, 42–46 (2000).
Rothstein, M. & Simoes-Costa, M. Heterodimerization of TFAP2 pioneer factors drives epigenomic remodeling during neural crest specification. Genome Res. 30, 35–48 (2020).
Bajpai, R. et al. CHD7 cooperates with PBAF to control multipotent neural crest formation. Nature 463, 958–962 (2010).
Schulz, Y. et al. CHD7, the gene mutated in CHARGE syndrome, regulates genes involved in neural crest cell guidance. Hum. Genet. 133, 997–1009 (2014).
Klincumhom, N. et al. Generation of neuronal progenitor cells and neurons from mouse sleeping beauty transposon-generated induced pluripotent stem cells. Cell Reprogram 14, 390–397 (2012).
Holm, S. A simple sequentially rejective multiple test procedure. Scand. J. Stat. 6, 65–70 (1979).
Marraudino, M. et al. Sexually dimorphic effect of genistein on hypothalamic neuronal differentiation in vitro. Int. J. Mol. Sci. 20, 2465 (2019).
Corces, M. R. et al. An improved ATAC-seq protocol reduces background and enables interrogation of frozen tissues. Nat. Methods 14, 959–962 (2017).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Scott, L. J. et al. The genetic regulatory signature of type 2 diabetes in human skeletal muscle. Nat. Commun. 7, 11764 (2016).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Yu, G., Wang, L. G. & He, Q. Y. ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31, 2382–2383 (2015).
Konermann, S. et al. Genome-scale transcriptional activation by an engineered CRISPR-Cas9 complex. Nature 517, 583–588 (2015).
Kent, W. J. et al. The human genome browser at UCSC. Genome Res. 12, 996–1006 (2002).
Boyle, E. I. et al. GO::TermFinder–open source software for accessing Gene Ontology information and finding significantly enriched Gene Ontology terms associated with a list of genes. Bioinformatics 20, 3710–3715 (2004).
Acknowledgements
Involvement of Yiwen Zhai in this work was supported by Henan Province Health and Family Planning Technology Talents Overseas Training Project (No. 2018060).
Funding
This work was funded by Henan Province Health and Family Planning Technology Talents Overseas Training Project (No.2018060), U.S. Department of Health & Human Services | National Institutes of Health (NIH) (Grant No. R01DC009410), U.S. Department of Health & Human Services | NIH | National Institute on Deafness and Other Communication Disorders (NIDCD) (Grant Nos. R01DC014456, R01DC0184014).
Author information
Authors and Affiliations
Contributions
H.Y., Y.Z., D.M.M., P.S., M.L. and S.C.J.P. conceived experiments. J.M.S. and Bielas provided technical assistance. H.Y., Y.Z., and S.F.H. performed experiments and, along with D.F.H., prepared figures and analyzed data for Figs. 1, 2, 3, and 5. G.S., A.S. and P.S. generated and analyzed data for Fig. 4. H.Y., R.D.A., M.L., S.C.J.P. and D.F.H. produced and analyzed data for Fig. 5. W.L. assisted with manuscript editing. H.Y., Y.Z., S.F.H., D.F.H. and D.M.M. wrote the manuscript. All authors reviewed and edited the manuscript and approved the content.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yao, H., Hannum, D.F., Zhai, Y. et al. CHD7 promotes neural progenitor differentiation in embryonic stem cells via altered chromatin accessibility and nascent gene expression. Sci Rep 10, 17445 (2020). https://doi.org/10.1038/s41598-020-74537-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-74537-4
This article is cited by
-
Improving the differentiation potential of pluripotent stem cells by optimizing culture conditions
Scientific Reports (2022)
-
Hybrid tumors with perineurioma components: a systematic review of the literature and illustrative case
Acta Neurochirurgica (2022)
-
CHD8 safeguards early neuroectoderm differentiation in human ESCs and protects from apoptosis during neurogenesis
Cell Death & Disease (2021)
-
Hybrid schwannoma–perineurioma frequently harbors VGLL3 rearrangement
Modern Pathology (2021)
-
The emerging role of chromatin remodelers in neurodevelopmental disorders: a developmental perspective
Cellular and Molecular Life Sciences (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.