Abstract
Colistin is considered a last-resort reserved drug for the treatment of critical human infections by Gram-negative bacteria. Phenotypic colistin-resistance is strongly associated with plasmid-mediated mobile colistin resistance (mcr) genes. The mcr-bearing Enterobacteriaceae have been detected in many countries from environments, animals, and humans. This study investigated phenotypic colistin-resistance and the distribution of mcr-1, mcr-2, mcr-3, mcr-4, and mcr-5 genes in chicken-gut bacteria in Bangladesh. Bacteria were isolated from poultry- and native-chicken droppings, and their susceptibilities to colistin were determined by agar dilution and E-test minimal inhibitory concentration (MIC) measurements. Multiplex polymerase chain reactions detected mcr-1 to mcr-5 genes. Overall, 61.7% (92/149) of the isolates showed colistin resistance by agar dilution assessment (MIC > 2.0 μg/mL). The phenotypic resistance was observed considerably higher in poultry-chicken isolates (64.6%, 64/99) than in native-chicken isolates (56%, 28/50; p = 0.373). All the resistant isolates showed MIC levels between > 2 and > 128 μg/mL. The mcr-genes (mcr-1and mcr-2 combined) were detected more in poultry gut bacteria (36.4%) than native-chicken isolates (20%, p = 0.06). Despite bacteria sources, mcr-genes appeared to be significantly associated with phenotypic colistin-resistance phenomena (p < 0.001). Prior colistin usage led to a substantial increase in the proportion of bacteria with mcr-genes and phenotypic resistance (p < 0.001).
Similar content being viewed by others
Introduction
Emerging antimicrobial resistance (AMR) is a leading public health concern worldwide1. Multiple studies have reported a strong association of antimicrobial usage (AMU) in animals with the extensive AMR burden2,3. Colistin (polymyxin E) has been used worldwide in food animals for decades for therapeutic, prophylactic, and growth promotion purposes4,5,6. China and Brazil were two major users of colistin as a growth promoter in agriculture and livestock7,8, until a recent ban due to AMR as a health hazards9. However, the ban in China includes the prohibition of using colistin as a feed additive only, and its use as a therapeutic agent for sick animals is not prohibited10. In most Asian countries, including India, Japan, Korea, and Vietnam, colistin has been used extensively as an feed-administered animal growth promoter to increase the body weights of farm animals11,12. The indiscriminate presence of therapeutic or prophylactic antimicrobials in either humans or animal husbandry generates selection pressure and influences the plausible development of AMR in pathogens and other diverse commensal microbial populations13,14. The presence of residual antimicrobials in farmed animal foods has been evidenced by multiple studies15,16, and is a major human health concern. Acquisition of the plasmid-mediated mobilized colistin resistance gene variant-1 (mcr-1) was first described in Enterobacteriaceae from both farm-animal products and humans7. After that, multiple mcr-1-carrying species of Enterobacteriaceae were detected in many countries in the environment, animals, and humans17,18,19. Subsequently, more variants of transferable colistin resistance mcr gene (mcr-1 to mcr-9) have been described in Enterobacteriaceae20,21. As colistin was extensively used in animal production in China, increased colistin resistance has been observed there in recent years18,22. The outcomes further attest to the relationship between colistin usage and the acquisition of antimicrobial-resistant bacteria from food-producing animals13,14. Moreover, colistin heteroresistance (wherein traditionally susceptible pathogens eventually appear resistant) has also been reported for many pathogens23,24,25. The mcr-1 gene was found in human pathogens with and without a history of colistin treatment26,27,28. A recent outbreak with colistin-resistant pathogens ended a very high case-fatality rate in humans29. Thus, identification of the root cause, transmission, and trajectories of colistin-resistant infection has drawn worldwide attention. Colistin has been approved as a veterinary medicine and as a feed additive in Bangladesh. To our knowledge, no known study has reported either the prevalence of colistin usage in Bangladeshi food animals or the association of colistin-usage in food animals with evolution of corresponding drug-resistant pathogens. This study investigated the prevalence of the mcr-1 to mcr-5 genes linked to phenotypic colistin resistance in farm-origin poultry chicken bacteria and household-dwelling native chicken isolates in Bangladesh.
Results
Study farms and samples
A total of 105 poultry dropping samples from 21 farms were collected for this study. After routine cross-checking, five samples from one farm were discarded due to incompleteness of the exposure history data. Thus, 100 faeces from 20 poultry farms located in five districts of Bangladesh were screened for distinct types of enteric bacteria (Fig. 1). The selected farms were a combination of small (capacity 50–2000 chickens), medium (2001–5000 chickens), and large (5000+ chickens) farms. More than half of the farmed chickens were intended for meat production, namely, broiler chickens (6/20, 30%) and Pakistani cocks (30%). The remaining 40% (8/20) of the chickens were layer type, and were farmed primarily for egg production. Layer chickens are also used as meat sources when egg production rates are low. Colistin usage was reported in 66.7% of the meat-producing chickens (8/12) and 37.5% of the layer chickens (3/8) in the PFs. Over 80% of the farm managers in this study had no professional training regarding the dosing of different veterinary medicines and their potential side-effects. However, the managers were familiar with different brand names of ‘over the counter’ (OTC) antimicrobials and growth-promoting drugs used for chickens. Our data collector captured the brand names as original data and subsequently translated to generic names before entry into the database. Other 25 native-chicken faeces were collected from five individual houses and investigated in the study. None of the farmers who provided droppings of native chickens had profession training for animal husbandry farming.
Sampling areas. Spatial locations of poultry farms and houses were shown from where samples of poultry-chicken droppings (n = 100) and native-chicken droppings (n = 50) were collected. Sampling locations covering five districts in Bangladesh are indicated in the map. We used geographic information mapping software, ArcGIS version 10 for Windows to draw the sampling spot-location map.
Isolation and identification of chicken faecal bacteria
The poultry faeces yielded 375 different isolates, and native faeces generated 50 isolates (Supplementary Fig. 1A). The poultry faeces carried more varieties of bacteria than the native chicken faeces. Of the total isolates, 99 were randomly selected from poultry bacteria, and all the native bacteria were subjected to assessment of colistin susceptibility and mcr-1 to mcr-5 carriage. Overall, all the poultry- and native-chicken faeces yielded at least one type of bacteria. Very few culture plates exhibited no growth. The 99 isolates from poultry faeces were classified as, 21 E. coli (21.2%), 36 Proteus spp. (36.4%), 20 Klebsiella spp. (20.2%), 10 Salmonella spp. (10.1%), 3 Shigella spp. (3.0%) and 9 Enterobacter spp. (9.1%). The native chicken faeces yielded 13 E. coli (26.0%), 16 Proteus spp. (32.0%), 10 Klebsiella spp. (20.0%), 6 Salmonella spp. (12.0%), and 5 Enterobacter spp. (10.0%), but no Shigella spp.
Phenotypic colistin susceptibility assessment
Following EUCAST guidelines, isolates were considered susceptible (S) when the MIC values exhibited ≤ 2 μg/mL and resistant (R) when MICs appeared > 2 μg/mL. (Supplementary Fig. 1B). Experiments were repeated when some single colonies or a thin haze growth was observed within the inoculated spot. Concordant results were found in independent MIC assessment by E-test (Supplementary Fig. 1C). A total of 92 (61.7%) isolates showed colistin-resistance by the agar dilution test. Of these, 64 (64.6%) were poultry chicken isolates compared to 28 (56%) from the native group (Table 1, p = 0.373). Isolates from poultry- and native-chicken faeces exhibited overall indifferent phenotypic colistin resistance (Fig. 2A). Intraspecies bacteria from the two sources demonstrated similar phenotypic resistance levels (Supplementary Table 1).
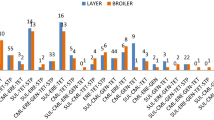
Isolate-wise phenotypic colistin resistance and mcr-gene carriage. (A) The agar dilution methods evaluated phenotypic colistin resistance. The bacteria were considered resistant when their minimum inhibitory concentrations (MICs) appeared > 2 μg/mL. Bacterial species examined for the phenotypic resistance were presented on the X-axis. The white-coloured bars in the Y-axis indicate the percentage of colistin resistance exhibited by poultry isolates. Likely, the black-coloured bars illustrate the percentages of resistant isolates from native chicken faeces. Percentile distributions of the poultry isolates show similar levels of colistin phenotypic resistance compared to native chicken isolates. (B) Carriage of the mcr genes (mcr-1 to mcr-5) was examined in chicken gut bacteria by polymerase chain reaction (PCR). The percentage of the identified mcr-genes in poultry-chicken bacteria was compared to that of native chicken bacteria. White bars to the Y-axis represent the proportions of the mcr-genes in poultry isolates, and black bars represent the native-chicken bacteria. Percentile distributions show the poultry-chicken isolates carried more mcr genes than that of the native chicken isolates.
Detection of the mcr gene variants
The colistin resistance genes, mcr-1 to mcr-5 were assessed in 149 chicken isolates using PCR analysis. The mcr-1 and mcr-2 genes were detected in 43 (28.9%) and 5 (3.4%) of isolates, respectively, from a combination of poultry- and native-chicken faeces (Supplementary Fig. 1D). Of them, two isolates carried mcr-1 and mcr-2 concurrently, and the remaining were mutually exclusive. None of the isolates was detected with mcr-3, mcr-4, and mcr-5. Separately, mcr-1 was found in 36.4% of the poultry bacteria (36/99) and 20% of the native chicken isolates (10/50). A statistically significant and moderately strong correlation was observed between mcr-genes and poultry faeces-origin bacteria (r = 0.167, p = 0.041). In contrast, no marked differences were observed in the carriage of mcr-2: only two isolates (2%) from poultry-chicken faeces and three (6%) from native-chicken faeces carried the allele (p = 0.335). The mcr-bearing isolates were detected in all five districts located in two divisions, namely, Savar (n = 20), Manikganj (n = 9), Tangail (n = 6), Mymensingh (n = 4), and Gazipur (n = 2) (Fig. 1). In this study, 17 Proteus spp. 10 Klebsiella spp. nine E. coli, seven Salmonella spp. one Shigella spp. and two Enterobacter isolate harboured the mcr-1 gene. The mcr trait was found in overall higher proportion in poultry-chicken isolates than in native chicken isolates (Fig. 2B). However, Chi-square analysis showed a statistically non-significance difference in the overall mcr-carriages in isolates from the two different origins (p = 0.06). The separate intraspecies presence of mcr was not significantly different between the two groups. (Supplementary Table 1).
Sequence analyses of the amplified mcr-1 gene showed high identity with global mcr-1 gene sequences in the NCBI database. A majority of the nucleotide sequences among the three clones were identical, except for a four-nucleotide mismatch with one clone (Supplementary Fig. 2A). The four-nucleotide substitutions appeared to be non-synonymous, which changed four amino acid substitutions in one bacterial mcr-1 (Supplementary Fig. 2B).
Association of the mcr and phenotypic colistin resistance
A strongly significant association was observed between mcr-1 and colistin-resistant phenotypes by the agar dilution test (p < 0.001). All the five isolates carrying mcr-2 were phenotypically colistin-resistance; however, no statistically significant association between mcr-2 and phenotypically colistin-resistance was observed (p = 0.064). The lost statistical significance could be attributable to the small sample sizes. By combining the two variants' acquisition, isolates possessing the mcr demonstrated a very strong harmony to the phenotypic resistance attribute (p < 0.001). All of the isolates carrying mcr trait showed colistin MIC levels between16 and > 128 μg/mL (Fig. 3A). Unexpectedly, some mcr-negative isolates (44.7%) also showed phenotypic colistin-resistance by the agar dilution test (Supplementary Table 2). The finding indicates the presence of some other factors besides the identified mcr variants to contribute colistin resistance to the isolates.
Minimum inhibitory concentrations (MICs) of isolates based on the mcr-gene acquisition and sources of origin. All the bacteria in this study were subjected to measure minimum inhibitory concentrations (MICs) of colistin between 1.0 and 128.0 μg/mL by agar dilution tests. Percentages of bacteria exhibiting each MIC level were calculated. The X-axis shows each of the MIC points, and Y-axis represents the percentages of isolates at a particular MIC. (A) The percentages of mcr-positive and mcr-negative isolates at MIC endpoints were compared. Red-coloured bars show the magnitudes of mcr-positive isolates, and the green-coloured bars are for mcr-negative. Percentile distributions show that mcr-positive isolates exhibited higher MICs compared to mcr-negative isolates. (B) The proportions of poultry- and native-chicken isolates at particular MIC points were compared. Red-coloured bars show the percentages of poultry-chicken origin isolates, and the green-coloured bars are for native-chicken bacteria. No marked differences of MIC were observed in isolates categorized in two source groups.
Association of prior colistin usage with its phenotypic resistance and mcr acquisition
Twelve poultry farms (60%, 12/20) reported using colistin sulphate for the purposes of treatment or growth promotion. One lot of the household chickens had a history of colistin usage (20%, 1/50). The 12 farms with colistin exposure yielded 71 isolates, of which 33 were mcr-positive (46.5%). In contrast, the droppings from the remaining non-exposed chickens yielded 16.7% mcr harbouring isolates (13/78). Therefore, a substantially higher proportion of mcr was observed in colistin-exposed bacteria (p < 0.001). Similar higher proportion of phenotypic colistin resistance was noted in the colistin-exposed group of isolates (p < 0.001). Statistical significance became attenuated for phenotypic resistance and mcr-acquisition in separate analyses for poultry- and native-chicken isolates separately (Table 2). It is likely that the reduced sample sizes in the separate analyses led to loss of statistical significance.
Minimum inhibitory concentration determination
Entire isolates were subjected to MIC measurements by agar dilution tests, in which > 2 μg/mL is considered a clinical breakpoint for colistin resistance. An overall consistent result was observed between the agar dilution and the E-test methods for colistin MIC determination. Each time, the control plate without colistin sulfate showed adequate growth of both test-isolate and susceptible control Escherichia coli ATCC25922 strains. For several isolates, mismatching of two- to four-fold MIC values were observed between Etest and dilution methods. Consistent MIC patterns were observed in mcr-1/mcr-2 carrying isolates: the bacteria's growth was found from the lowest concentration of 1.0 μg/ml colistin to their MICs. All the mcr-bearing isolates showed higher MIC ranging between > 16 and > 128 μg/mL (Fig. 3A). However, some inconsistent ‘skipped’ (no growth) patterns were observed for several mcr-naive isolates: no growths were observed at lower colistin concentrations, such as 1–4 μg/mL, but growth appeared at immediate higher concentration (8 μg/mL). We repeated the experiments and considered the MIC values based on the growth observed at the highest concentrations. Although most mcr-naïve isolates demonstrated lower colistin MICs, a portion had shown seriously high MIC levels (Fig. 3A). With the acquisition of mcr-gene, each type of bacteria, such as Proteus spp., Klebsiella spp., E. coli, Salmonella spp., and Enterobacter spp., revealed higher MIC endpoints (Table 3). On the other hand, isolates from poultry-chicken faeces exhibited an almost indifferent MIC levels with the isolates from native-chicken droppings (Fig. 3B). Further, independent intraspecies MIC manifestations from the two sources appeared reasonably similar (Table 4).
Discussion
We investigated the gut flora of both farm-based poultry chickens and home-based cage-free chickens for the prevalence of the five mcr gene variants, mcr-1 to mcr-5, which is linked to phenotypic colistin resistance in Bangladesh. The study showed that approximately one-third of the isolates from poultry chicken droppings carried the hazardous mcr-1 gene. Carriage of mcr-2 variants was found much lower (3–4%) in bacteria from both poultry- and native-chicken droppings. Some co-occurrences of mcr-1 and mcr-2 were also observed. None of the isolates were detected with the presence of mcr-3, mcr-4, and mcr-5. The findings were consistent with earlier reports of the mcr gene in some GNB isolated from food animals in several countries7,30. Furthermore, we detected the mcr-gene in native chicken gut isolates as well, although at marginally lower proportions. The mcr-habouring bacteria, regardless of specimen source, were also found to be resistant to colistin by the agar dilution method. Moreover, the determined MIC values for all mcr-positive isolates were between > 16 and ≥ 128 μg/mL, which confirmed the association of mcr genes with colistin resistance. To the best of our knowledge, there is no scientific information available about colistin-resistant bacteria identified from either livestock or the clinical sector in Bangladesh. One brief report described mcr-1 from sewage samples in Bangladesh31. Our study is the first of its kind, describing carriage of the colistin-resistance mcr gene in both farmed poultry and cage-free native chicken bacteria in Bangladeshi settings. As anticipated, occurrences of the mcr genes, singly or a co-existence of the two (mcr-1 and mcr-2) were associated with an increased phenotypic colistin resistance (higher MIC values). A combination of the two genes may enable bacteria to accumulate multiple genes simultaneously to emerge as high-risk enteric bacterial clones easily. Multiple studies have reported that the co-occurrences of multiple clinically relevant antibiotic-resistant genes with mcr into the same bacteria possess increased resistance to different antimicrobials32,33. On the other hand, a significant portion of isolates without carrying any detectable mcr gene, showed phenotypic resistance to colistin. The inconsistency of the genotype–phenotype association could be explained by other variants of mcr genes or factors that have not been investigated in this study21. Mutational inactivation in the essential chromosomal gene, mgrB could further explain the colistin-resistant isolates without the carriage of mcr genes34. More studies will be required to find some mechanistic explanations for the colistin phenotypic discrepancy to mcr genes. This study identified mcr-bearing colistin resistant enteric bacteria in five different districts in Bangladesh, and the obtained results are expected to be generalizable for the remaining territory of the country. The study findings carry public health importance in the establishment of potential surveillance for enhanced antimicrobial resistance and stewardship in human-veterinary interfaces.
Antimicrobials commonly used in animal husbandry and human health are more concerning in the context of zoonotic transmission of resistant bacteria. The discovery of mcr in chicken gut bacteria is of both national and global importance because of colistin's clinical applications35. Considerably large proportions of E.coli in the study were found to carry mcr-1, consistent with the multiple previous studies7,36. This study identified mcr at a higher frequency in Salmonella spp. than in E. coli, although colistin is not an antibiotic of choice for treating Salmonella infections. These salmonellae can be particular serovars that intrinsically remain resistant to colistin. Some earlier studies have identified group D serovars of Salmonella possessing O:9 somatic antigens confer natural colistin resistance37,38. However, this study did not confirm the serotyping of Salmonella interlinked to mcr-gene and colistin resistance. The natural reservoirs of colistin resistance may contribute to horizontal transmission of the resistant traits to surrounding susceptible isolates39. Furthermore, mcr presence was observed in Proteus spp. and Klebsiella spp. with a clear disparity between caged and cage-free chicken sources. The feeding types and nature of poultry chickens may contribute to differential gut flora isolates and their resistance attributes. Moreover, colistin use for treatment of poultry chickens may have contributed to the increased acquisition of mcr by the two types of bacteria. The other studies also showed that prior exposure was associated with high positive detection of mcr in the poultry isolates11.
The procedure used to collect data on antimicrobial usage history in this study had some inherent limitations. The previous colistin usage data provided by the chicken owners were self-reported and have not been evaluated in-depth. The drug-usage information collected from unskilled and less-educated workers may contain some information bias towards underreporting. A majority of the owners, managers, and workers in the PFs were untrained and uneducated enough to not understand the chemical types of antimicrobials. Sometimes they were unresponsive and unable to provide the generic names of the antimicrobial drugs that they used. Furthermore, neither the poultry farms nor the household chicken owners had an inventory and recording system for their applied therapeutic antimicrobials. Some chicken owners presented the commercial brand names of the antimicrobials used, or alternatively provided drug sachets or bottles directly to the field workers. The data enumerator entered the brand names of the antibiotics used as original data and generic names in parallel. Refusal and self-censoring were encountered occasionally. The factors together may have led to underreporting of and self-selection bias in the colistin usage data. There is a minute possibility of overreporting as our data enumerator had collected colistin treatment history only after checking validated evidence of the content. Therefore, the identified association between colistin exposure and mcr acquisition should definitely be valid. We anticipate an even stronger association if further cohort studies are designed with wide coverage of samples.
Unlike PFs chickens, the native chickens had less reported usage of colistin for treatment or growth promoter. Hypothetically, there should have considerably low phenotypic colistin resistance or mcr acquisition in the native chicken isolates. However, we detected a steady phenotypic resistance and mcr-carriage in the isolates of the native chicken group. There are several hypothetical explanations for these unanticipated outcomes. First, colistin usage history may be unreported by the home-based native chicken owners. Second, the native chickens may receive some feeds that were previously mixed with colistin as a growth-promoting agent, which was not known to the users. Similar evidence of extensive colistin use for growth promotion has been found in many countries4. Third, horizontal gene transfer (HGT) could explain the transmission of the mcr gene and associated phenotypic resistance from the poultry gut microbiota to the native chicken enteric flora39.
This study had a few more limitations. The investigation was conducted following a cross-sectional study-design, and no follow up was carried out due to resource limitations. This study analyzed five mcr gene variants such as mcr-1 to mcr-5; some more recently reported variants, like, mcr-6, mcr-7, mcr-8, and mcr-9 have not been investigated. Serological identification of Salmonella spp. has not been carried out. Salmonella serotyping, particularly against somatic O-antigen, could explain whether the identified bacteria were natural reservoirs of colistin resistance or carrying newly acquired resistance traits. Without the serotyping of Salmonella, the reporting of overall colistin resistance may become slightly overestimated. However, total Salmonellae identified in this study was around 10% only, which is unable to shift the whole study findings significantly. The small sample size was also a constraint to conducting fully powered statistical analyses. However, our results were generated from a resource-limited setting and maintained internal validity by repeating independent experiments where necessary. The findings have shed some light on colistin usage for animal husbandry, varieties of chicken-gut bacteria, their carriage of mcr genes, co-occurrences, and the genotypic-phenotypic association of colistin resistance. Further studies will be worthwhile to assess the external validity of our findings.
Conclusions
The mobile colistin resistance genes circulate in bacteria isolated from both poultry chicken- and native chicken-droppings. mcr-1 variant predominates in the Bangladeshi chicken gut bacteria over other variants, such as mcr-2, mcr-3, mcr-4, and mcr-5. The distribution of the mcr genes and phenotypic colistin resistance was moderately higher in poultry-chicken isolates. The prior use of colistin in chickens leads to increased acquisition of higher phenotypic resistance and mcr genes by gut-bacteria.
Methods
Study design and specimen collection
A cross-sectional study was conducted to examine mcr-1 to mcr-5 genes prevalence in bacterial isolates from chicken droppings. Poultry farms (PFs) were conveniently selected for chicken-faeces collection between July 2017 and June 2018. We selected 20 PFs from areas of the Dhaka, Manikgang, Gazipur, Tangail, and Mymensing districts, where major PFs are located in Bangladesh. Geographic information mapping software, ArcGIS version 10 for Windows, was used to draw a sampling spot-location map (Fig. 1). A structured questionnaire was administered to farm-owners to investigate the types of poultry chickens, their recent disease history, and records of antibiotics applied for treatment and prophylaxis. The study also made enquiries regarding the farm-owners’ levels of education and whether they had attained animal husbandry training. Native chickens are also a substantial and high demand meat source for the Bangladeshi population. People living in villages and sub-urban areas of the country raise native chickens on a non-commercial basis. The native chickens are maintained cage-free in the day time and are allowed to explore around the house to find natural foods. Sometimes commercial foods are provided. This study also examined the faeces of native chickens for bacterial isolates and for their carriage of the mcr-1 to mcr-5 genes and associated colistin susceptibility. The same structured questionnaire was administered to home-owners to seek information regarding the antibiotic exposure history of the native chickens. Data enumerators were trained beforehand on the use of the questionnaire by minimizing selection bias and interviewer bias.
Bacterial isolation and identification
Chicken faecal samples were directly collected in specific specimen collection tubes following all safety precautions and aseptic techniques. For long distance transport, faeces samples were dipped into the Cary Blair transport medium (Oxoid, UK) before being transported to the laboratory. For bacterial isolation, approximately one gram of chicken-faeces was mixed in 400 μL of phosphate-buffered saline (PBS), and one loopful of diluted sample was streaked on MacConkey agar (Oxoid, UK) for growth of Gram-negative enteric bacilli. For the detection of Salmonella and Shigella, the diluted chick-droppings were enriched in Rappaport Vassiliadis soya broth (RVS Broth, Oxoid, UK) overnight and streaked separately on Salmonella-Shigella (SS) agar (Oxoid, UK) medium. After overnight incubation at 37 °C, distinct single colonies were picked and cultured again on tryptone soya agar (Lyophilchem, Italy) to prepare a pure-culture repository, which was stored at -80ºC. The purified bacterial colonies were identified by conventional biochemical procedures followed by a rapid biochemical-test kit (API 20E, BioMérieux, Durham, NC). Bacterial identification was validated further by 16S rDNA analyses.
Antibiogram and minimum inhibitory concentration (MIC) assessment
The isolates' phenotypic antimicrobial susceptibilities were tested by the agar dilution method on Mueller–Hinton agar (MHA, Oxoid, Basingstoke, UK). The method further determined the lowest concentration of colistin (minimal inhibitory concentration, MIC) capable of inhibiting the visible growth of isolates40,41. Agar dilution was performed by incorporating different concentrations of colistin-sulfate powder (Santa Cruz Biotechnology Inc, TX) into an MHA medium from 1.0 to 128.0 μg/mL in a two-fold dilution series. Approximate twenty-milliliter volumes of MHA were used in 90-mm Petri dishes for agar dilution MICs. Each test inoculum was prepared by inoculating one pure culture colony into Mueller–Hinton broth and allowed incubation for three-hour at 37 °C that generates a density of inoculum about 104 colony-forming units (CFU) per spot on the MHA. A 0.5 McFarland standard was used to visually compare a density equivalent to approximately 108 CFU/mL. A micropipette was be used to inoculate plates and allowed the inoculum to dry on spots at room temperature. The plates were then inverted for incubation at 37 °C in air for 18–20 h. Agar dilution MICs were performed in duplicates. Several earlier studies conducted the epsilometer test (E-test) parallel to the agar dilution method to validate colistin MIC determination42,43. E-test was performed using a commercial strip containing a predefined gradient of colistin concentrations (Liofilchem Inc, Italy) to compare MIC with the agar dilution method44. Escherichia coli ATCC25922 was used as the susceptible-control reference strain for each batch of MIC tests. Also, a control plate without colistin-sulfate was tested for the growth of both test and control strains. The clinical breakpoints for colistin resistance were interpreted according to the European Committee on Antimicrobial Susceptibility Testing (EUCAST) guidelines when the MIC value was > 2 μg/mL45.
Detection of the colistin resistance mcr gene variants
Multiplex polymerase chain reaction ((PCR) was conducted to detect the mcr1 to mcr-5 genes in the isolates. The primer pair, CLR-F, (5′ CGGTCAGTCCGTTTGTTC 3′) and CLR-R, (5′ CTTGGTCGGTCTGTAGGG 3′), was used to yield a 309 bp DNA band for mcr-1, as described elsewhere7. The other four primer pairs to detect mcr-2, mcr-3, mcr-4, and mcr-5 gene amplicons were obtained from another original study20. In brief, the modified protocol was as follows: prepared bacterial DNA (2.0 μL) was added to a 2× PCR premixture, (15 μL, GeneON, Germany) and five pmol of each primer (1 μL), and deionized water was added to obtain a final volume of 30 μL. Reactions underwent an initial denaturation at 94 °C for 15 min followed by 25 cycles of amplification (Applied Biosystems 2720 Thermal Cycler, Singapore), consisting of denaturation for 30 s at 94 °C, annealing for 90 s at 55 °C, and extension for 1 min at 72 °C, and a final 10 min elongation at 72 °C. Expected amplicons for mcr-1 (309 bp), mcr-2 (715 bp), mcr-3 (929 bp), mcr-4 (1116 bp), and mcr-5 (1644 bp) were visualized under UV light after 1.2% agarose gel electrophoresis followed by staining with ethidium bromide. The obtained results were validated by separate singleplex PCR analyses of the mcr-genes.
Sequencing of the mcr-1 gene
Three randomly amplified mcr-1 genes obtained by PCR were sequenced and blasted against the NCBI nucleotide database (https://blast.ncbi.nlm.nih.gov) to confirm their identity with other reported mcr-1 genes. Sequence alignments were performed using the BioEdit program (version 7.2) for Windows. The revealed mcr-1 sequences were submitted to GenBank, and the accession numbers obtained were MK615113, MK615114, and MK615116.
Statistical analysis
A validated Bengali version of the questionnaire was used for data collection. Data collection supervisors rechecked completed data collection forms. In addition, 10% of the laboratory analyses were randomly selected twice by different researchers to assess inter-researcher biases. Verified data were entered and subsequently analysed using the IBM SPSS Statistics data editor (version 21). Missing data were excluded from the bivariate analysis. Descriptive and inferential statistical procedures were used to ascertain the types of chicken gut bacteria and their carriage of the mcr genes. Pearson's chi-square test was used to test the significance of association between categorical data, and Yate's correction for continuity was applied where appropriate. A two-tailed p-value small than 0.05 was considered to measure statistical significance.
Ethics statement
This study was approved by the Ethics and Research Review Committee of the Jahangirnagar University Faculty of Biological Sciences [No. BBEC, JU/M 2017 12(4), dated December 27, 2017]. All methods were performed in accordance with the relevant guidelines and regulations. Informed consent was obtained from the respective farm owners or managers for collecting poultry droppings and information from their farms. Farm identities and information were kept strictly anonymous to protect their commercial, personal, and private information. The respective sample identification code was assigned duly for each sample collected.
Conference presentation
ASM Biothreats on “Prophylactic Uses of Antimicrobials in Bangladeshi Poultry Drive Microbial Resistance (129)”, Baltimore, Maryland, February 12–14, 2018.
Data availability
A dataset was generated and analysed during the study that stored at the databank repository of One Health Laboratory, Department of Microbiology, Jahangirnagar University. Data can be shared upon request from the corresponding author following universal data-sharing.
rules.
References
WHO. Global Action Plan on Antimicrobial Resistance. World Health Organization (2015).
Marshall, B. M. & Levy, S. B. Food animals and antimicrobials: Impacts on human health. Clin. Microbiol. Rev. 24, 718–733. https://doi.org/10.1128/CMR.00002-11 (2011).
Aidara-Kane, A. Containment of antimicrobial resistance due to use of antimicrobial agents in animals intended for food: WHO perspective. Rev. Sci. Technol. 31, 277–287 (2012).
Rhouma, M., Beaudry, F. & Letellier, A. Resistance to colistin: What is the fate for this antibiotic in pig production?. Int. J. Antimicrob. Agents 48, 119–126. https://doi.org/10.1016/j.ijantimicag.2016.04.008 (2016).
Poirel, L., Jayol, A. & Nordmann, P. Polymyxins: Antibacterial activity, susceptibility testing, and resistance mechanisms encoded by plasmids or chromosomes. Clin. Microbiol. Rev. 30, 557–596. https://doi.org/10.1128/CMR.00064-16 (2017).
Catry, B. et al. Use of colistin-containing products within the European Union and European Economic Area (EU/EEA): Development of resistance in animals and possible impact on human and animal health. Int. J. Antimicrob. Agents 46, 297–306. https://doi.org/10.1016/j.ijantimicag.2015.06.005 (2015).
Liu, Y. Y. et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: A microbiological and molecular biological study. Lancet. Infect. Dis. 16, 161–168. https://doi.org/10.1016/S1473-3099(15)00424-7 (2016).
Fernandes, M. R. et al. Silent dissemination of colistin-resistant Escherichia coli in South America could contribute to the global spread of the mcr-1 gene. Euro Surveill. https://doi.org/10.2807/1560-7917.ES.2016.21.17.30214 (2016).
Wang, R. et al. The global distribution and spread of the mobilized colistin resistance gene mcr-1. Nat. Commun. 9, 1179. https://doi.org/10.1038/s41467-018-03205-z (2018).
Walsh, T. R. & Wu, Y. China bans colistin as a feed additive for animals. Lancet. Infect. Dis. 16, 1102–1103. https://doi.org/10.1016/S1473-3099(16)30329-2 (2016).
Kempf, I., Jouy, E. & Chauvin, C. Colistin use and colistin resistance in bacteria from animals. Int. J. Antimicrob. Agents 48, 598–606. https://doi.org/10.1016/j.ijantimicag.2016.09.016 (2016).
Lim, S. K. et al. First detection of the mcr-1 gene in Escherichia coli isolated from livestock between 2013 and 2015 in South Korea. Antimicrob. Agents Chemother. 60, 6991–6993. https://doi.org/10.1128/AAC.01472-16 (2016).
Canton, R. & Morosini, M. I. Emergence and spread of antibiotic resistance following exposure to antibiotics. FEMS Microbiol. Rev. 35, 977–991. https://doi.org/10.1111/j.1574-6976.2011.00295.x (2011).
Mezhoud, H. et al. Presence of antimicrobial resistance in coliform bacteria from hatching broiler eggs with emphasis on ESBL/AmpC-producing bacteria. Avian Pathol. 45, 493–500. https://doi.org/10.1080/03079457.2016.1167837 (2016).
Reig, M. & Toldra, F. Veterinary drug residues in meat: Concerns and rapid methods for detection. Meat. Sci. 78, 60–67. https://doi.org/10.1016/j.meatsci.2007.07.029 (2008).
Goetting, V., Lee, K. A. & Tell, L. A. Pharmacokinetics of veterinary drugs in laying hens and residues in eggs: A review of the literature. J. Vet. Pharmacol. Ther. 34, 521–556. https://doi.org/10.1111/j.1365-2885.2011.01287.x (2011).
Hembach, N. et al. Occurrence of the mcr-1 colistin resistance gene and other clinically relevant antibiotic resistance genes in microbial populations at different municipal wastewater treatment plants in Germany. Front. Microbiol. 8, 1282. https://doi.org/10.3389/fmicb.2017.01282 (2017).
Huang, X. et al. High prevalence of colistin resistance and mcr-1 gene in Escherichia coli isolated from food animals in China. Front. Microbiol. 8, 562. https://doi.org/10.3389/fmicb.2017.00562 (2017).
Zhou, H. W. et al. Occurrence of plasmid- and chromosome-carried mcr-1 in waterborne Enterobacteriaceae in China. Antimicrob. Agents Chemother. 61, 562. https://doi.org/10.1128/AAC.00017-17 (2017).
Rebelo, A. R. et al. Multiplex PCR for detection of plasmid-mediated colistin resistance determinants, mcr-1, mcr-2, mcr-3, mcr-4 and mcr-5 for surveillance purposes. Euro Surveill. https://doi.org/10.2807/1560-7917.ES.2018.23.6.17-00672 (2018).
Gharaibeh, M. H. & Shatnawi, S. Q. An overview of colistin resistance, mobilized colistin resistance genes dissemination, global responses, and the alternatives to colistin: A review. Vet. World 12, 1735–1746. https://doi.org/10.14202/vetworld.2019.1735-1746 (2019).
Wang, Y. et al. Prevalence, risk factors, outcomes, and molecular epidemiology of mcr-1-positive Enterobacteriaceae in patients and healthy adults from China: An epidemiological and clinical study. Lancet. Infect. Dis. 17, 390–399. https://doi.org/10.1016/S1473-3099(16)30527-8 (2017).
Nicoloff, H., Hjort, K., Levin, B. R. & Andersson, D. I. The high prevalence of antibiotic heteroresistance in pathogenic bacteria is mainly caused by gene amplification. Nat. Microbiol. 4, 504–514. https://doi.org/10.1038/s41564-018-0342-0 (2019).
Meletis, G., Tzampaz, E., Sianou, E., Tzavaras, I. & Sofianou, D. Colistin heteroresistance in carbapenemase-producing Klebsiella pneumoniae. J. Antimicrob. Chemother. 66, 946–947. https://doi.org/10.1093/jac/dkr007 (2011).
Jayol, A., Nordmann, P., Brink, A. & Poirel, L. Heteroresistance to colistin in Klebsiella pneumoniae associated with alterations in the PhoPQ regulatory system. Antimicrob. Agents Chemother. 59, 2780–2784. https://doi.org/10.1128/AAC.05055-14 (2015).
Hasman, H. et al. Detection of mcr-1 encoding plasmid-mediated colistin-resistant Escherichia coli isolates from human bloodstream infection and imported chicken meat, Denmark 2015. Euro Surveill. https://doi.org/10.2807/1560-7917.ES.2015.20.49.30085 (2015).
Wang, Q. et al. Genomic insights into mcr-1-positive plasmids carried by colistin-resistant Escherichia coli isolates from inpatients. Antimicrob. Agents Chemother. https://doi.org/10.1128/AAC.00361-17 (2017).
Beyrouthy, R. et al. MCR-1 and OXA-48 in vivo acquisition in KPC-producing Escherichia coli after colistin treatment. Antimicrob. Agents Chemother. https://doi.org/10.1128/AAC.02540-16 (2017).
Tian, G. B. et al. MCR-1-producing Klebsiella pneumoniae outbreak in China. Lancet. Infect. Dis. 17, 577. https://doi.org/10.1016/S1473-3099(17)30266-9 (2017).
Rhouma, M., Beaudry, F., Theriault, W. & Letellier, A. Colistin in pig production: Chemistry, mechanism of antibacterial action, microbial resistance emergence, and one health perspectives. Front. Microbiol. 7, 1789. https://doi.org/10.3389/fmicb.2016.01789 (2016).
Islam, A. et al. Colistin resistant Escherichia coli carrying mcr-1 in urban sludge samples: Dhaka, Bangladesh. Gut Pathog. 9, 77. https://doi.org/10.1186/s13099-017-0227-4 (2017).
García, V. et al. Co-occurrence of mcr-1, mcr-4 and mcr-5 genes in multidrug-resistant ST10 enterotoxigenic and Shiga toxin-producing Escherichia coli in Spain (2006–2017). Int. J. Antimicrob. Agents 52, 104–108 (2018).
dos Santos, L. D. R. et al. Co-occurrence of mcr-1, mcr-3, mcr-7 and clinically relevant antimicrobial resistance genes in environmental and fecal samples. Arch. Microbiol. 17, 1–6 (2020).
Poirel, L. et al. The mgrB gene as a key target for acquired resistance to colistin in Klebsiella pneumoniae. J. Antimicrob. Chemother. 70, 75–80 (2015).
Marchaim, D., Kaye, D. & Kaye, K. S. Use of colistin in critically Ill patients. Adv. Exp. Med. Biol. 1145, 155–179. https://doi.org/10.1007/978-3-030-16373-0_12 (2019).
Olaitan, A. O., Morand, S. & Rolain, J. M. Mechanisms of polymyxin resistance: Acquired and intrinsic resistance in bacteria. Front. Microbiol. 5, 643. https://doi.org/10.3389/fmicb.2014.00643 (2014).
European Food Safety Authority, and European Centre for Disease Prevention and Control.. The European Union summary report on antimicrobial resistance in zoonotic and indicator bacteria from humans, animals and food in 2014. EFSA J.https://doi.org/10.2903/j.efsa.2016.4380 (2016).
Ricci, V., Zhang, D., Teale, C. & Piddock, L. J. V. The O-Antigen Epitope Governs Susceptibility to colistin in Salmonella enterica. mBio 11, 5625. https://doi.org/10.1128/mBio.02831-19 (2020).
Tse, H. & Yuen, K. Y. Dissemination of the mcr-1 colistin resistance gene. Lancet. Infect. Dis. 16, 145–146. https://doi.org/10.1016/S1473-3099(15)00532-0 (2016).
Wiegand, I., Hilpert, K. & Hancock, R. E. Agar and broth dilution methods to determine the minimal inhibitory concentration (MIC) of antimicrobial substances. Nat. Protoc. 3, 163–175. https://doi.org/10.1038/nprot.2007.521 (2008).
E. D. D. E. D. Determination of minimum inhibitory concentrations (MICs) of antibacterial agents by agar dilution. European Committee for Antimicrobial SusceptibilityTesting (EUCAST) of the European Society of Clinical Microbiology and Infectious Diseases (ESCMID)6, 509–515 (2000).
Behera, B. et al. Evaluation of susceptibility testing methods for polymyxin. Int. J. Infect. Dis. 14, e596-601. https://doi.org/10.1016/j.ijid.2009.09.001 (2010).
Maalej, S., Meziou, M., Rhimi, F. & Hammami, A. Comparison of disc diffusion, Etest and agar dilution for susceptibility testing of colistin against Enterobacteriaceae. Lett. Appl. Microbiol. 53, 546–551 (2011).
Balouiri, M., Sadiki, M. & Ibnsouda, S. K. Methods for in vitro evaluating antimicrobial activity: A review. J. Pharm. Anal. 6, 71–79. https://doi.org/10.1016/j.jpha.2015.11.005 (2016).
Testing, E. C. o. A. S. Breakpoint tables for interpretation of MICs and zone diameters. Version 10.0 (2020).
Acknowledgements
Dr. Salequl Islam received a research grant from the Grants for Advanced Research in Education (GARE), the Ministry of Education, Bangladesh (Award ID: LS 201685). The above grant provided support in study design, data collection, and laboratory investigation. It also supported a stipend for a co-author, ULU, but did not provide any part in the publication of the manuscript. This study was further supported by a research grant from the World Academy of Sciences (TWAS) awarded to Dr. Shamsun Nahar (Award ID: RG/BIO/AS_I–FR3240297771). The latter grant provided support in laboratory investigation and played a partial role in manuscript processing and publication. The authors would like to thank the chicken-farm owners, managers, and the household chicken owners for their active support. They are grateful to Mahbubul H. Siddiqee for his critical review and suggestions for developing the manuscript.
Author information
Authors and Affiliations
Contributions
S.I. conceived and designed the study, analyzed the data, and wrote the manuscript. U.L.U. conducted major laboratory experiments, prepared the results, and assisted with manuscript drafting. M.R., F.S., N.J., S.Iq., and B.H. helped with sample collection, laboratory experiments, data acquisition, and sequence analysis. A.S.M.M. prepared questionnaires and consent forms, assisted with translation into verified Bengali formats, and reviewed the manuscript. M.M.H. helped study design, conducted the statistical analysis, and reviewed the manuscript. S.N. coordinated the study and assisted with manuscript drafting.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Islam, S., Urmi, U.L., Rana, M. et al. High abundance of the colistin resistance gene mcr-1 in chicken gut-bacteria in Bangladesh. Sci Rep 10, 17292 (2020). https://doi.org/10.1038/s41598-020-74402-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-74402-4
This article is cited by
-
Phenotypic and genotypic characterization of colistin-resistant Escherichia Coli with mcr-4, mcr-5, mcr-6, and mcr-9 genes from broiler chicken and farm environment
BMC Microbiology (2023)
-
Farm to table: colistin resistance hitchhiking through food
Archives of Microbiology (2023)
-
Electrochemical immunosensor for the detection of colistin in chicken liver
3 Biotech (2022)
-
The varying effects of antibiotics on gut microbiota
AMB Express (2021)
-
Surveillance of antimicrobial resistance in low- and middle-income countries: a scattered picture
Antimicrobial Resistance & Infection Control (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.