Abstract
Burkholderia pseudomallei (Bpm) is a bacterial pathogen that causes Melioidosis, a disease with up to 40% mortality and an infection relapse of 15–23% despite antibiotic treatment. Ineffective clearance of Bpm by antibiotics is believed to be due to persistence, a hibernation-like survival mechanism modulated, in part, by toxin–antitoxin systems (TAS). Several organisms possess a repertoire of TASs but defining environmental cues eliciting their activity is hindered by laborious in vitro experiments, especially when there are many toxins with redundant function. Here, we identified which of 103 proteins in Bpm that share features found in toxins of the TAS and repurposed transcriptional data to identify which ones play a role in surviving intracellular host defenses. Putative toxins with the strongest transcriptional response were found to have low conservation between Bpm strains, while toxins that were constitutively expressed were highly conserved. Further examination of highly conserved toxins BPSS0899, BPSS1321, and BPSL1494 showed that they were functional, and their mutation led to reduce survival within macrophages and reduced in vivo persistence-associated pathology (abscesses) during treatment, but did not affect macrophages persistence. These findings highlight the utility of a data-driven approach to select putative toxins and suggests a selective role for some TAS in host survival.
Similar content being viewed by others
Introduction
Burkholderia pseudomallei (Bpm) is an opportunistic bacterial pathogen that can infect both animals and humans causing the disease Melioidosis. The estimated yearly predicted disease and mortality burden for Bpm infections is 169,000 cases and 89,000 (54%) deaths worldwide1,2. Without an existing vaccine, the only way to combat a Bpm infection is through a partially effective multi-antibiotic treatment that still results in high mortality rates and relapse in 15–23% of convalescent patients2,3,4. Although Bpm is intrinsically resistant to many antibiotics, emergence of resistance to clinically relevant antibiotics is rare, suggesting an alternative mode of bacterial survival2,5,6,7. Bacterial persistence has been associated with treatment failure of chronic infections caused by pathogens such as Escherichia coli, Mycobacterium tuberculosis, Staphylococcus aureus, Acinetobacter baumannii, Pseudomonas aeruginosa, and the fungal pathogen Candida albicans8,9,10,11. The mechanism of survival used in persistence differs from antibiotic resistance in that the bacteria are susceptible to antibiotics, but become transiently refractive to killing by reducing cellular functions and growth12,13. Previous publications have shown that Bpm has higher rates of persister cell formation than several other bacteria, reaching up to 10–64% when stationary growth phase bacteria are exposed to antibiotics14,15,16,17,18. Due to the high rate of Bpm persistence during infection, identification and targeting of persistence mechanisms offers a means to improve treatment efficacy.
Bacterial toxin-antitoxin systems (TAS) have been identified as one of the molecular switches that sense stress and in many cases activate a persistence phenotype. This phenomenon occurs using a variety of mechanisms to induce growth senescence. In most cases, TAS are composed of two cistronic genes, one encoding an antitoxin and another a toxin. During a favorable environment, the toxin is bound and neutralized by the antitoxin. Upon encountering a stressful stimulus, the toxin is freed from the antitoxin’s repression and becomes able to induce persistence by selectively inhibiting cellular processes19,20. Not all toxins are reversible, but those activated in persistence are, allowing for a dynamic population that responds to environmental stresses and recovers when the stress is removed.
Although the role of TAS in persistence induction has been controversial in non-pathogenic E. coli, there is strong evidence that these systems are playing a role in more complex organisms such as Salmonella enterica serovar Typhimurium, M. tuberculosis, and Bpm14,21,22. The TAS can respond to any number of environmental cues, including dehydration, nutrient limitation, bacteriophage infection, or nitric oxide release by macrophages23,24. The traditional approach to studying TAS is by over-expression of each toxin, often in a surrogate organism, and testing for induction of persistence via growth senescence in nutrient rich or minimal laboratory media15,25. As more toxins are being identified, this approach has become more laborious. Currently, there is no in silico predictive model to determine which environmental conditions might mediate toxin induction. To address this, we used a data mining approach and repurposed existing transcriptional data across 82 environmental conditions to determine which Bpm putative toxins are associated with survival in the host26. The overall goal was to generate a data-driven approach that directs selection of TAS for further investigation as mediators of persistence and their association with chronic infection and treatment failure.
Results
Identification of putative toxins
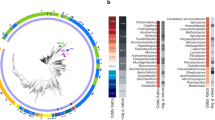
To predict whether toxins have a role in host persistence, it is important to first determine how many putative toxins are encoded in the Bpm genome. Employing the same methodology used to find PIN domain-containing toxins, we PSI-Blasted all bacterial sequences contained in the Pfam for each of the 21 previously identified type II toxin families (Supplemental Table S1; Supplemental Figure S1). Using Bpm K96243 as the query, 103 predicted proteins containing toxin features were identified with an expect value of ≤ 0.001 (Fig. 1A) and we selected only those smaller than 500 amino acids15,27,28,29. The size cut off was derived from our knowledge that Bpm BPSS1584 is a functional toxin of 450 amino acids15. Two-thirds (60.2%) of the Bpm putative toxins were larger than standard toxin size of 200 amino acids (Fig. 1B)27,28 and the predominant classes identified were PIN (also known as VapC), and Doc toxins, representing 17% (n = 18) and 22% (n = 23) of all toxins, respectively (Fig. 1C). The toxins identified were asymmetrically distributed in the genome, with a heavier concentration in chromosome 1 (Fig. 1D), which generally encodes for core genes associated with central metabolism and growth, while chromosome 2 encodes accessory functions linked to adaptation and survival in different niches and virulence factors30. Although our goal was to streamline selection of toxins for experimental investigation, we also wanted to validate the functionality of the toxins identified. Using the traditional over-expression assay, we assessed functionality of eight toxins described in the next section and not previously investigated. Of the toxins BPSL0034, BPSL1494, BPSL2775, BPSL2851, BPSS0899, BPSS0698, BPSS0899, BPSS1321, and BPSS2196 that were tested, 75% of the over-expressed genes led to a reduction in growth, indicating their functionality (Supplemental Figure S2).
Next, we examined the overlap between our study and existing results from other experimental reports, computational programs, and databases. The most recent platform, TASmania, relies on 369 Hidden Markov Models (HMM; generalized protein sequence) to identify toxins31. TASmania identified 30 TAS with an expect value below 0.001, of which 8 (27.5%) were found in our current study (Supplemental Figure S3A and B). Re-running the RASTA computational program, we were unable to obtain the same results previously published15, and when compared the published results, we only had an overlap of 8 toxins (BPSL0549A, BPSL0562, BPSL1564, BPSL2333, BPSS0390, BPSS0845a, BPSS2142, and BPSS1226), which may be due to the stringent size and gene structure filters used by the RASTA analysis (Supplemental Figure S3A and B)28. Further, The overlap of data between RASTA and TASmania was also examined and showed low convergence. The Toxin-Antitoxin Database (TADB) is a compilation of all previously published computational and experimental results from which we found all toxins.
Prediction of toxins important in the host using existing datasets
Historically, TAS have been investigated for their impact in antibiotic-mediated persistence but are also induced by environmental stress, such as low nutrients, high temperature, extracellular death factors, and immune system attacks23,24. Here, we predicted which toxins were host-induced by using an existing RNA expression dataset, exposing Bpm to 82 different conditions ranging from general laboratory growth, desiccation, survival in water, to in vivo infection (Supplemental Table S2)26. It is important to note that the dataset includes three wild type Bpm strains K96234, Bp22, and Bp008, and the latter two were included in the original publication as isogenic controls. Bpm K96243 and Bp22 share 94% genome with a 98% identity; however, the genome for Bp008 is not available.
Hierarchical clustering identified seven sets of genes and seven clusters of conditions (Fig. 2). Of the putative toxins, 54% of the toxins are associated with most of the conditions tested and this finding could suggest redundancy in the toxin functions. Anaerobic conditions (strain K96243), in vivo lung infection (K96243), macrophage infection (Bp22) and pH 7 (K96243) are clustered together and represent conditions the bacteria would encounter in the host. Three gene clusters representing 26 genes were identified as being host-associated. Cluster one includes the novel putative toxins BPSS1321, BPSL1494, BPSS0899, BPSS2775, and BPSS2196. Seven of the genes in clusters 2 and 3 (BPS1584, BPSS0395, BPSL0559, BPSL2333, BPSS1060, BPSS0390, and BPSL0175) have been previously tested by either our lab or in the study by Butt et al. and some were shown to induce antibiotic-mediated persistence and influence persistence in a murine model of infection14,15. As controls, the type 6 secretion system (T6SS) subunit gene hcp1 and the multi-nucleated giant cells-inducing gene bimA were included. Consistent with the literature, bimA and hcp1 were induced during macrophage and in vivo infection32,33,34.
Predicting toxin-condition association. The dataset is presented as a heat map. Toxin genes are presented on the horizontal axis and conditions on the vertical axis. Both genes and conditions are clustered as shown on the top and left. Red boxes highlight antibiotic exposure (top) and exposure to host-like conditions (bottom), such as infection of RAW 264.7 macrophages, bacteria recovered from BALB/c mouse lung, pH 7, and exposure to anaerobic conditions. For ease, genes previously investigated are denoted with blue arrows and genes investigated in the current study are denoted in green arrows.
Conservation and expression of the predicted host-associated toxins
We further evaluated toxins predicted to be important in the host-induced persistence phenotype and with a potential role in chronic infection. We hypothesize that for the gene product to play a role in a defined phenotype, it might be highly conserved within the bacterial species or isolates; therefore, we evaluated the conservation (presence/absence) of putative toxin genes in clusters 1–3 of Bpm and closely related Burkholderia strains (Table 1). Among 677 Bpm strains available in the database, putative toxins in clusters 1 and 2 were highly conserved. B. mallei (Bm) is the closest related species to Bpm and is believed to have originated following reductive evolution from Bpm to become a host restricted pathogen. We speculated that conserved toxins in Bm should be important for survival in the host. B. mallei displayed similar trends as Bpm, with higher toxin conservation in cluster 1 and the least conservation in cluster 3 (Table 1). Conversely, B. thailandensis shares 85% of the toxin genes with Bpm, and it is an environmentally restricted bacteria that has only caused a handful of reported cases of human illness35. As such, B. thailandensis had even fewer of the toxins conserved than Bm. To evaluate the conservation in a more distantly related species, we selected B. cenocepacia, which is notorious for causing chronic, life threatening infections in patients with lung abnormalities such as cystic fibrosis36. Interestingly, B. cenocepacia strains share only three of the putative toxins evaluated, BPSS0899, BPSL2527, and BPSS1584.
In the processing steps of preparing the data for analysis and visualization, gene expression was min–max normalized to place all of them on the same scale. As with any data normalization, there are flaws and for min–max, it inflates the importance of constitutively active genes. Due to this, the standard deviation across all conditions was investigated to evaluate the overall gene expression (Table 2). Toxin expression was lowest in cluster 1 and highest in cluster 3. A closer look uncovered that cluster 1 was overly inflated with respect to their constitutive expression. When comparing expression to conservation, the data showed an inverse correlation (Table 2). Previously, we had investigated three of the toxin genes that had high expression and low-to-moderate conservation (BPSS0390, BPSS0395, and BPSL1584) and found moderate to no impact in host persistence18. Constitutively expressed toxin genes have not been investigated but owing to their high conservation, we chose to further evaluate for potential function of the putative toxins BPSS1321, BPSL1494, BPSS0899, BPSS2775, and BPSS2196, found in cluster 1.
Over-expression of toxins in surrogate organisms
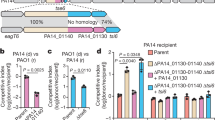
Next, we over-expressed the genes in E. coli DH10B (Fig. 3A). When toxin expression was induced, BPSS0899, BPSS1321, and BPSL1494 reduced the growth rate of E. coli as compared to the control strain carrying the empty vector. Reduced growth indicated some form of growth arrest. We further tested whether BPSS0899, BPSS1321, and BPSL1494 could cause growth arrest in the closer related bacterial surrogate, B. thailandensis. When over-expressed, only BPSL1494 induced a reduction in growth compared to the strain carrying the empty plasmid (Fig. 3B). Interestingly, B. thailandensis carries orthologous toxin-antitoxin operons for BPSS1321 and BPSL1494, and over-expressed toxins’ activity may be abrogated by binding suppression of the orthologue antitoxin. As such, the bacteria were exposed to levofloxacin to promote toxin activation via release from antitoxin repression. After 24 h of antibiotic exposure, only expression of BPSS1321 conferred a survival advantage compared to the control strain (Fig. 3C). Although we found different results between E. coli and B. thailandensis, we chose to further investigate BPSS0899 (putative doc toxin), BPSS1321 (PemK/MazF-like), and BPSL1494 (doc toxin) in Bpm by allelic exchange mutagenesis (Supplemental Table S3).
Overexpression of cluster one toxins. (A,B) Toxins were cloned into pBAD-myc-his or pSCRba2 plasmids and then transformed into either E. coli DH10b or B. thailandensis, respectively. After bacteria were grown in the corresponding media, cultures were divided into 6 tubes and toxin expression induced with (A) 0.2% arabinose (pBAD), (B) 0.2% rhamnose (pSCRba2), or mock treated with PBS and growth monitored by optical density. (C) To assay if toxin overexpression affected persistence, induced, and repressed B. thailandensis strains were treated with 5–10 × MIC of levofloxacin and represented as a fold-change between induced and repressed for each toxin. ****P < 0.0001.
Non-host persistence phenotypes: swarming and antibiotic-induced persistence
Isogenic, unmarked mutants were constructed for the toxin genes BPSS0899, BPSS1321 and BPSL1494 to evaluate their function in the Bpm K96243 strain background. Typical assays to test persistence include swarming and antibiotic-induced persistence. Reduced swarming has been correlated to persistence in several pathogens37,38,39,40,41,42,43. All three toxin mutant strains had reduced swarming motility even when supplemented with a higher concentration of glucose (Supplemental Figure S4A). Antibiotic stress is typically the focus of persistence research; therefore, we carried out antibiotic-induced persistence assays and found reduced persistence of ΔBPSS1321 and ΔBPSL1494 in LB, but not RPMI (Supplemental Figure S4B and C). Conversely, ΔBPSS0899 resulted in increased persistence in RPMI, but not LB.
Macrophage uptake, survival, and persistence
Next, we evaluated the role of the toxins for persistence in host cells. When Bpm is in the host, macrophages are thought to be the preferred cell infected prior to establishment of a chronic infection44. Evidence shows that macrophages assist in dissemination as early as 24 h post infection and therefore, they are encountered early during infection44. U937-derived macrophages were infected and assayed for bacterial uptake, survival, and persistence. Significantly lower uptake was seen with only ΔBPSL1494 as compared to the wild type strain (Fig. 4A). All three mutants had lower ability to survive in macrophages, suggesting a defect in intracellular adaptation (Fig. 4B). However, when cell permeable antibiotic levofloxacin was added to kill actively growing intracellular bacteria (Fig. 4B), there was no significant difference in intracellular persistence between the mutants and the wild type strain. Compared to levofloxacin only treatment, macrophage exposure increased persistence as we have previously shown (Fig. 4B and18).
Invasion, survival, and persistence in macrophages. Macrophages were derived by incubating the U937 pleural monocytic cell line with PMA for 48 h. (A) Invasion, (B) survival, and persistence were assayed after 30 min of uptake followed by a 24 h incubation with or without 20 × MIC of levofloxacin. (C) To further investigate the role of macrophages on survival, U937 derived macrophages were polarized to a M1 phenotype using IFNγ for 16 h prior to infection and incubated for 24 h. One-way ANOVA was used to evaluate the difference within a treatment group. *P < 0.05, ****P < 0.0001.
The reduced macrophage survival of ΔBPSS0899, ΔBPSS1321, and ΔBPSL1494 indicated a potential increase in susceptibility to innate antimicrobial killing found in these cells. Previous studies have shown that M1 polarized macrophages kill Bpm more efficiently than M0 differentiated macrophages due to NO production45. To evaluate the susceptibility of the mutants to M1 macrophage killing, U937 macrophages were differentiated with PMA then stimulated with IFNγ prior to infection. Activated cells were more readily able to kill Bpm; however, there was no significant difference between wild type and mutant strains (Fig. 4C).
Simulating in vitro host-induced persistence
As we have shown, all three toxin mutants had a reduced capacity to survive in macrophages but behaved like wild type when levofloxacin was added in the macrophage-induced persistence assay. Two possibilities are feasible: (1) persistence is not affected and non-persister bacteria are being killed by the macrophage or (2) the toxins respond to host-stresses, but the levofloxacin treatment activates other TAS that compensate for the absence of the mutated toxins. To clarify this dichotomy, the effect of macrophage-like stress stimuli was assessed. Nutrient deprivation can induce persistence and occurs in some microenvironments within the macrophage which is a preferred niche for Bpm18,44,46,47. Nutrient deprivation also contributes to the reduction of bacterial growth in late growth stages. We hypothesize that if the toxins induce persistence during nutrient deprivation, loss of function could lead to higher bacterial titers at late growth stages. As hypothesized, all mutants tested grew to higher titers in both nutrient rich medium (LB) and a host-cell supportive, nutrient-defined, medium (RPMI) (Fig. 5A,B) suggesting a lack of nutrient deprivation recognition. We next tested survival in presence of antibiotics after nutrient starvation. Bacteria were grown in RPMI and then diluted and incubated in M9 minimal media supplemented with glycerol and casamino acids for 30 min prior to antibiotic exposure (Fig. 5C). Loss of BPSS0899 and BPSL1494 resulted in a significant defect in survival, while loss of BPSS1321 trended toward lower survival but was not statistically different than wild type.
The effect of nutrient deprivation and nitric oxide stress on persistence. (A,B) The response to natural nutrient deprivation in late growth state was assayed by growing bacteria in either LB or RPMI and monitoring optical density and growth over 48 h. To test the impact of nutrient depravation further, bacteria were grown overnight in RPMI then exposed to (C) nutrient limitation for 30 min then treated with 20 × MIC of levofloxacin for 2 h. Survival was enumerated by normalizing to bacterial counts before antibiotics were added. In (D) a similar experiment was carried out with nitric oxide as the stress for 30 min prior to exposure to 20 × MIC of levofloxacin. One-way ANOVA was done with in each conditional group or between conditions for one strain. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
During infection of macrophages Bpm is also exposed to nitric oxide (NO), which has been shown to increase Bpm persistence to the antibiotics imipenem and levofloxacin18,45. Exposure to NO for 30 min prior to levofloxacin treatment led to reduced survival when comparing ΔBPSS1321 and ΔBPSL1494 to wildtype (Fig. 5D). It is important to note that with antibiotics alone, ΔBPSS0899 survived better than wild type (Supplemental Figure S4C), but it did not when pre-treated with NO (Fig. 5D).
In vivo persistence
In our prior publication, we showed that toxin mutants ΔBPSS0390 and ΔBPSS0395, found in clusters 2 and 3, showed reduced bacterial colonization upon infection of BALB/c mice and treatment with levofloxacin18. Testing colonization of ΔBPSS0899, ΔBPSS1321, and ΔBPSL1494 after infection and levofloxacin treatment resulted in transiently (days 14–18) higher weights of mice infected with ΔBPSS0899 and ΔBPSL1494 as compared to wild type (Fig. 6A). At the end of the experiment, the spleens of mice infected with mutants had lower splenic weight (Fig. 6B) which corresponded to reduced gross pathology in all spleens infected with ΔBPSS0899 or ΔBPSL1494 and in half of ΔBPSS1321-infected spleens (Fig. 6C). Interestingly, this finding did not correlate with the difference in splenic bacterial burden. Additionally, the lung and liver bacterial burden were not significantly altered by toxin loss (Fig. 6D–F).
In vivo Persistence of ΔBPSS0899, ΔBPSS1321, and ΔBPSL1494 Following Treatment. Mice were intranasally infected with 3.5 CFU equivalent of each mutant and treated for 5 days with 25 mg/kg/day of levofloxacin. (A) Weight was monitored for the 20-day infection period after which organs of surviving mice were collected, (B) weighed, and (C) visualized for gross pathology. (C) Representative images are displayed, and two images were selected from ΔBPSS1321 due to variability in pathology. (D) Lungs, (E) livers, and (F) spleens were homogenized and bacteria cultured on agar to assess bacterial burden. Statistics were completed using Two-Way ANOVA or One-Way ANOVA. *P < 0.05 and **P < 0.01.
In our in vitro macrophage assays, mutants showed reduced intracellular survival in the absence of antibiotics (Fig. 4B), but not when antibiotics were added (Fig. 4B). To test if colonization was altered in the absence of antibiotics, mice were infected with a low dose (1.5 LD50) of each bacterial strain. After challenge, all mice became ill, had weight loss, and some succumbed to infection (Fig. 7A,B). There was no significant difference in survival between mice infected with the respective toxin mutants and compared to those infected with the wild type strain (Fig. 7A). In contrast to the study carried out with antibiotics, infection with ΔBPSL1494 led to large splenic abscesses (Fig. 7C). Quantification of bacterial burden showed no significant difference in the spleens, lungs, or livers when comparing mice (n = 3) infected with wild type and ΔBPSS0899 (Fig. 7D–F). Because we had a single survivor (n = 1) infected with ΔBPSS1321 and ΔBPSL1494, we could not make similar conclusions, but noted that ΔBPSS1321 and ΔBPSL1494 had a splenic burden of 2 and 6 logs higher than the average burden observed with the wild type mice, respectively (Fig. 7D–F).
In vivo Persistence of ΔBPSS0899, ΔBPSS1321, and ΔBPSL1494 in the Absence of Treatment. Mice were intranasally infected with 1.5 LD50 of wild type or CFU equivalent of each mutant and (A) survival and (B) weight was monitored for 14 days after which organs of surviving mice were collected, (C) visualized for gross pathology, and CFU burden enumerated. (D) Lungs, (E) livers, and (F) spleens were homogenized and bacteria cultured on agar to assess bacterial burden. Statistics were completed using Two-Way ANOVA or One-Way ANOVA. *P < 0.05 and **P < 0.01.
Discussion
Due to the likelihood that an intracellular bacteria such as Bpm might enter a persistence state, targeting the mechanisms associated with this phenotype offers an opportunity to reduce non-antibiotic resistance treatment failures, infection relapses, and potentially prevents the emergence of new antibiotic resistant strains12,48. Although targeting TASs is an ideal approach because it is a bacterial specific mechanism, many pathogens harbor numerous TAS and in many cases they have redundancy in their functions21,49. It is also unclear how many toxins are elicited by any given condition and therefore, it is difficult to determine how many TAS play a role in persistence during infection in the host. The approach presented here used known data on toxin families and existing Bpm gene expression data to identify and down-select putative toxins based on their association with environmental cues. We found that approximately half of the putative toxins were associated with a wide variety of stress conditions, including antibiotic treatment, supporting the idea that multiple TAS might be required and their functions overlapped to achieve a persistent phenotype. This data supports previous findings indicating that free-living organisms have more toxins, likely to promote adaptation to changing surroundings49. This observation is also true for Bpm because: (a) this pathogen is a soil and water dwelling organism and many of the conditions evaluated are likely found during environment changes and, (b) antibiotics are a strong selective pressure, which are encountered in the environment during interspecies bacterial warfare. In contrast, fewer genes were associated with host-like conditions. These findings are expected because bacteria have been adapting to their environment and competing with other bacteria for a billion years, while bacterial-host interaction has occurred more recently.
The prediction method used here proved to be an efficient method of down-selecting toxins to study. As with any bioinformatic platform there are strengths and weaknesses. Here, min–max normalization was used and resulted in over representation of constitutively active genes. This brought into light a reciprocal correlation between gene expression activity and conservation. It is also important to mention that identification of antitoxins and their association with their respective toxins was not performed in this study because we want to first provide a wide view of the Bpm genes sharing features with known toxins. Further, constitutively expressed genes have not been the focus in TAS research. Continuous expression of a gene product is energetically costly to the bacteria and coupled with their high conservation, might suggest that the protein product is important for metabolic/survival purposes. In terms of TAS, being constitutively expressed has the potential to have a protein that can be activated faster and which might induce persistence more readily, therefore the effect of these gene products was the focus of the rest of our study.
Highly conserved, constitutively expressed, toxins BPSS0899, BPSS1321, and BPSL1494 were further investigated in Bpm using both over-expression and unmarked mutant strains. All three toxin mutants were defective in responding to one or more host stress stimuli (nutrient deprivation or NO) and had a reduced ability to survive in macrophages, but the survival defect was lost when macrophage persistence was induced using levofloxacin. These findings indicate that the BPSS0899, BPS1321, and BPSL1494 toxins might played a role in surviving the harsh environment of the host cell, but their absence may be compensated by activation of other toxins when an antibiotic is added to the medium early during the experiment. The selective activation of persistence by host stress alone is not discussed in the persistence field but may explain why people in endemic areas can be found harboring Bpm in the form of abscesses without being symptomatic50,51,52. Additionally, it is speculated that asymptomatic persistent infections provide a continuous immune protection from other Bpm infections, which has been corroborated by murine vaccine studies showing that the strongest protection was achieved with the vaccine strains that asymptomatically persists in the host53,54. To further examine the role of host-mediated persistence, there is a need for the development of novel methods to examine persistence in the absence of antibiotics.
When persistence was tested in an in vivo murine model of inhalational melioidosis, there was no evident effect on colonization with or without levofloxacin treatment. A surprising observation was that after treatment ΔBPSS0899 and ΔBPS1321 resulted in a reduced gross pathology compared to wild type strain. Conversely, without levofloxacin treatment ΔBPSL1494 had larger splenic abscesses. In the absence of CFU changes, we conclude that the change in gross pathology might be immune-mediated. For a bacterial pathogen such as Bpm, whose preferred niche in the host is the macrophage, persistence not only relies on the bacterial but also the host cell states. Previous work has shown that Salmonella persisters can modulate macrophage polarization toward a M2 phenotype55. For Bpm, there are no studies investigating the role of persistence on immune state modulation. Epidemiological and clinical data show that immune dysregulation seen in diabetes is the largest risk factor associated with melioidosis recurrent infections2. Future direction should aim to further understand the role of Bpm persistence in macrophages, the effect on cellular immune state, and whether bona fide TAS are playing a specific role within this environment. Until then, the findings here are intended to shift the paradigm of toxin–antitoxin research by using a data-driven approach to facilitate their identification and supports the utility of big data analysis to investigate constitutively expressed toxins as possible regulators of the persistence phenotype.
Materials and methods
Ethics statement
All manipulations with Bpm were conducted in CDC/USDA-approved and registered biosafety level 3 (BSL3) facilities at UTMB. Experiments with select agents were performed in accordance with BSL3 standard operating practices. The animal studies were carried out humanely in strict accordance with the recommendations in the Guide for the Care and Use of Laboratory Animals by the National Institutes of Health. The protocol (IACUC #0503014D) was approved by the Institutional Animal Care and Use Committee of the University of Texas Medical Branch.
Bacterial strains and growth conditions
The bacterial strains used in this study are listed in Supplemental Table S4. B. pseudomallei strain K96243 was obtained from BEI Resources, Manassas, VA, USA. E. coli and Burkholderia strains were stored in LB with 5% glycerol (LBG) at − 80 °C and prior to use strains were streaked onto Luria–Bertani (LB) agar and grown for 36–48 h at 37 °C. For liquid cultures, 3–5 colonies were inoculated into LB broth or RMPI with HEPES and L-glutamine and grown for 16 h at 37 °C with agitation (200 rpm). LBG agar was used in the process of generating isogenic mutants.
Toxin identification
All known toxins used to identify TAS in M. tuberculosis were considered in our identification21 (presented in Supplemental Table S1). All bacterial and archaeal sequences were downloaded from the EMBL-EBI Pfam database (https://pfam.xfam.org) for each toxin family. Sequences were PSI-Blast against Bpm K96243 (taxid:272560) with an expect cut-off 0.002. Expect value is a parameter that describes the number of hits one can “expect” to see by chance when searching a database, like a p-value. The results were sorted to remove redundancies, expect value further reduced to 0.001, and only toxins below 500 amino acid included28. Identified toxins were compared to toxins identified were compared to exiting results from:
RATSA (https://hdl.handle.net/10871/9303),
TASMania (https://shiny.bioinformatics.unibe.ch/apps/tasmania/),
TADB for chromosome one and two (https://bioinfo-mml.sjtu.edu.cn/TADB2/browse_org_result.php?type=genome&org_id=NC_006351) and (https://bioinfo-mml.sjtu.edu.cn/TADB2/browse_org_result.php?type=genome&org_id=NC_006350).
Toxin conservation
Presence or absence of toxin identified in cluster 1–3 was done using the ortholog data available on the Burkholderia Genome Database (during June 2018) (https://burkholderia.com). The analysis is carried out using DIAMOND high throughput protein alignment via translated DNA. For the analysis, the ortholog data for putative toxins was downloaded, sorted by species, and the percent of total strains on the databased used to calculated conservation across Bpm (n = 677), B. mallei (65), B. thailandensis (28), and B. cenocepacia (243) strains. All genomes were included regardless of their completion. Genomes with homologous genes with an expect value of 0.0001 were considered positives orthologs.
Repurposing existing data
Gene expression of Bpm following exposure to 82 independent conditions was obtained from a previous publication26. Median log10 raw reads of putative toxins and selected control genes were isolated from the data. The values were normalized on the expression of bacteria in LB at stationary phase (LBS) and min–max normalized presented as a heat map (https://www1.heatmapper.ca/expression/) with Manhattan distance metric clustering.
Over-expression assays
Selected toxin genes of interest were amplified with primers specified in Supplemental Table S5, cleaned with Qiagen PCR purification kit (Qiagen, Hilden, Germany) then ligated into either pBad-myc-his (Thermo Fisher Scientific, Massachusetts, USA) or pSCRba2 (Addgene.org), using Gibson kit (NEB, Massachusetts, USA). All constructs were transformed into E. coli DH5α competent cells (NEB, Massachusetts, USA). Constructs were confirmed with PCR and Sanger sequencing (UTMB Sequencing core). pBAD-myc-his constructs were transferred into E. coli DH10B cells which have a disrupted arabinose metabolism pathway (araD139; NEB, Massachusetts, USA). pSCRba2 constructs were transferred into B. thailandensis E264. Bacteria cultures grown in M9, supplemented with 2% glycerol and 1% casamino acids (BD), were diluted and then grown at 37 °C with agitation (200 rpm) until they reached an OD600 of 0.2. E. coli were induced with 0.2% arabinose or not stimulated, and growth monitored by optical density over time. B. thailandensis induced with 0.2% rhamnose or not stimulated and growth monitored at 37 °C with agitation (200 rpm). All experiments were carried out with antibiotics to maintain the plasmids.
Persister assays with B. thailandensis E264 were carried out on bacteria collected after 2 h of induction. Bacteria were normalized to the same concentration, exposed to either 5 × or 10 × MIC (MIC = 1 μg) of levofloxacin for 24 h, then quantified and normalized to the input concentration. Data is represented as a fold-change in persistence compared to non-induced bacteria. One-Way-ANOVA statistical analysis was done to examine significant difference between bacteria with empty vector and those expressing a toxin.
Mutagenesis scheme
The Bpm ΔBPSS0899, ΔBPSS1321, and ΔBPSL1494 mutants were constructed using a parental mating approach using pMo130 as previously described18,56. Two fragments were amplified (1) 400–600 base pair upstream and (2) the first 21 base pairs of the toxin gene, and the last 9 base pairs of the toxin gene accompanied by 400–600 bases pairs downstream (primers listed in Supplemental Table S5). The fragments were inserted into pMo130 using Gibson Assembly (NEB, Massachusetts, USA) and transformed into E. coli S17-1 λpir and introduce to Bpm K96243 via biparental mating. Merodiploids were selected using LBG agar with kanamycin and then plasmid loss was promoted by counter selecting on YT agar supplemented with 15% sucrose to obtain single deletion mutants. The mutations were then confirmed via PCR and Sanger sequencing (UTMB Sequencing core).
Growth curves
Isolates were grown overnight in LB or RPMI with HEPES (Gibco, Thermo Fisher Scientific, Massachusetts, USA) and the following day, diluted to an OD600 of 0.1. Cultures were incubated at 37 °C with agitation (200 rpm) and optical density read over 48 h.
Swarming assays
Bacterial cultures were diluted 1:10 and 2 μl was spotted in the center of semisolid nutrient agar plate (5 g/l bacto agar, 8 g/l nutrient broth N°2, +/− 0.5% [w/v] glucose). Plates were incubated for 24 h and the diameter of bacterial growth measured and presented as a percentage of the plates’ total diameter which was analyzed with One-Way ANOVA with Kruskal–Wallis correction.
Traditional persistence assays
Bacterial cultures were adjusted to 1 × 108 CFU/ml in medium containing 100 × MIC of levofloxacin. Cultures were incubated for 24 h at 37 °C without shaking, after which the surviving bacteria were quantified by serial dilution and plating on LB agar. Surviving bacteria were normalizing to the input and analyzed using One-Way ANOVA with Kruskal–Wallis correction.
Macrophage survival assays
Human U937 pleural monocytic cells (CRL-1593.2) were maintained in Modified RPMI-1640 Medium (ATCC 30–2001, Virginia, USA) supplemented with 10% FBS (Gibco), 1 mM sodium pyruvate (Invitrogen, California, USA), and 1 mM penicillin–streptomycin (Invitrogen). For infection assays, 2 × 105 cells/well were seeded in 24-well cell culture plates and supplemented with 25 ng/ml phorbol myristate acetate (PMA; Sigma, Missouri, USA) to induce macrophage differentiation 72 h prior to infection. Twenty-four h prior to infection, U937 cells were washed once and media replenished without PMA. For M1 differentiation, 16 h prior to infection media was replaced with 10 ng/ml of human IFNγ (Peprotech) supplemented medium. Cells were infected with bacteria at a MOI of 10 and synchronized by centrifugation. After a 30 min invasion, cells were washed with PBS and fresh medium containing 100 μg/ml kanamycin (extracellular restricted) or 80 μg/ml of levofloxacin (20 × MIC; cell permeable). At 24 h post infection, cells were washed with PBS and lysed with 0.1% Triton-X100 (Sigma), serial diluted in PBS, and plated on LB agar. After incubation, colonies were counted, normalized for invasion, and analyzed with One-Way ANOVA with Kruskal–Wallis correction.
Pre-induced persistence assays
Bacteria were grown in RPMI then adjusted to 5 × 107 CFU/ml in RPMI with 100 µM of spermine NONOate as a NO donor (sNO, Sigma Missouri, USA). Alternatively, bacteria were adjusted to approximately 5 × 107 CFU/ml in minimal medium supplemented with 2% glycerol and 1% casamino acids (M9; Becton Dickinson, New Jersey, USA). After 30 min of exposure at 37 °C without shaking, the samples were diluted 1:1 with medium containing levofloxacin to yield a final concentration of 20 × MIC (80 µg/ml) and in the case of the NO study, additional spermine NONOate to maintain their respective concentration. Bacteria were then incubated for 2 h at 37 °C without shaking and then survival enumerated. Percent survival was determined by normalizing the input after the initial 30 min incubation. Statistical significance was determined using One-Way ANOVA with Kruskal–Wallis correction.
In vivo bacterial infection model
Female, 6 to 8-week-old BALB/c mice were obtained from Jackson Laboratories (Bar Harbor, ME, USA). Mice were housed in microisolator cages under pathogen-free conditions and provided with rodent feed and water ad libitum and maintained on a 12 h light cycle. Before experiments, mice were given 5 days of acclimation. Anesthetized BALB/c mice (n = 5 per group) were inoculated intranasally (I.N.) with equivalent to 3.5 LD50 Bpm K96243 wild type (1 LD50 = 312 CFU), and CFU equivalent of ΔBPSS0899, ΔBPSS1321, or ΔBPSL1494 diluted with PBS in a total volume of 50 µL (25 µL/ nare). Mice received daily intraperitoneal injections of levofloxacin (25 mg/kg/day in PBS) starting at 24 h post-infection and continuing for five days. Mice were monitored and weighed daily throughout the 20-day study. For the low-dose infection BALB/c mice were anesthetized (n = 5 per group) and then inoculated intranasally (I.N.) with equivalent to 1.5 LD50 Bpm K96243 wild type, ΔBPSS0899, ΔBPSS1321, or ΔBPSL1494 and monitored and weighed daily throughout the 14-day study. Humane endpoints were strictly observed, and time of death was recorded upon animals succumbing to infection or at the study’s endpoint. Survival curves were analyzed by using the Kaplan–Meier method with log-rank test. For CFU enumeration, animals were euthanized, their lungs, liver, and spleen collected, and homogenized using Covidien Precision tissue grinders (Thermo Fisher Scientific, Massachusetts, USA). Tissue homogenates were serially diluted in PBS, plated, and incubated for 48–72 h at 37 °C. Colonies were counted and normalized to organ weight (g) and significance determined using T-test with a Mann–Whitney correction or One-way ANOVA depending on the number of groups.
References
Limmathurotsakul, D. et al. Predicted global distribution of Burkholderia pseudomallei and burden of melioidosis. Nat. Microbiol. 1, 15008 (2016).
Wiersinga, W. et al. Melioidosis. Nat. Rev. Dis. Primers 4, 17107 (2018).
Whitmore, A. An account of a glanders-like disease occurring in Rangoon. J. Hyg. (Lond.) 13, 1–34 (1913).
Chaowagul, W. et al. Relapse in melioidosis: Incidence and risk factors. J. Infect. Dis. 168, 1181–1185 (1993).
Sarovich, D. S. et al. Development of ceftazidime resistance in an acute Burkholderia pseudomallei infection. Infect. Drug Resist. 5, 129–132 (2012).
Schweizer, H. P. Mechanisms of antibiotic resistance in Burkholderia pseudomallei: Implications for treatment of melioidosis. Fut. Microbiol. 7, 1389–1399 (2012).
Wuthiekanun, V. et al. Trimethoprim/sulfamethoxazole resistance in clinical isolates of Burkholderia pseudomallei. J. Antimicrob. Chemother. 55, 1029–1031 (2005).
Gunn, J. S. et al. Salmonella chronic carriage: Epidemiology, diagnosis, and gallbladder persistence. Trends Microbiol. 22, 648–655 (2014).
Mulcahy, L. R., Burns, J. L., Lory, S. & Lewis, K. Emergence of Pseudomonas aeruginosa strains producing high levels of persister cells in patients with cystic fibrosis. J. Bacteriol. 192, 6191–6199 (2010).
Fauvart, M., De Groote, V. N. & Michiels, J. Role of persister cells in chronic infections: Clinical relevance and perspectives on anti-persister therapies. J. Med. Microbiol. 60, 699–709 (2011).
Alkasir, R. et al. Characterization and transcriptome analysis of Acinetobacter baumannii persister cells. Microb. Drug Resist. 24, 1466–1474. https://doi.org/10.1089/mdr.2017.0341 (2018).
Balaban, N. Q. et al. Definitions and guidelines for research on antibiotic persistence. Nat. Rev. Microbiol. 17, 441–448 (2019).
Blair, J. M., Webber, M. A., Baylay, A. J., Ogbolu, D. O. & Piddock, L. J. Molecular mechanisms of antibiotic resistance. Nat. Rev. Microbiol. 13, 42–51 (2015).
Butt, A. et al. The HicA toxin from Burkholderia pseudomallei has a role in persister cell formation. Biochem. J. 459, 333–344 (2014).
Butt, A., Müller, C., Harmer, N. & Titball, R. W. Identification of type II toxin–antitoxin modules in Burkholderia pseudomallei. FEMS Microbiol. Lett. 338, 86–94 (2013).
Nierman, W. C., Yu, Y. & Losada, L. The in vitro antibiotic tolerant persister population in Burkholderia pseudomallei is altered by environmental factors. Front. Microbiol. 6, 1338 (2015).
Ross, B. N., Myers, J. N., Muruato, L. A., Tapia, D. & Torres, A. G. Evaluating new compounds to treat Burkholderia pseudomallei infections. Front. Cell Infect. Microbiol. 8, 210 (2018).
Ross, B. N., Micheva-Viteva, S., Hong-Geller, E. & Torres, A. G. Evaluating the role of Burkholderia pseudomallei K96243 toxins BPSS0390, BPSS0395, and BPSS1584 in persistent Infection. Cell Microbiol. 21, e13096 (2019).
Unterholzner, S. J., Poppenberger, B. & Rozhon, W. Toxin–antitoxin systems. Bioengineered 5, 1–13 (2014).
Wen, Y., Behiels, E. & Devreese, B. Toxin–antitoxin systems: Their role in persistence, biofilm formation, and pathogenicity. Pathog. Dis. 70, 240–249 (2014).
Ramage, H. R., Connolly, L. E. & Cox, J. S. Comprehensive functional analysis of Mycobacterium tuberculosis toxin-antitoxin systems: Implications for pathogenesis, stress responses, and evolution. PLoS Genet. 5, e1000767 (2009).
Helaine, S. et al. Internalization of Salmonella by macrophages induces formation of nonreplicating persisters. Science 343, 204–208 (2014).
Fraikin, N., Goormaghtigh, F. & Van Melderen, L. Type II toxin–antitoxin systems: Evolution and revolutions. J. Bacteriol. https://doi.org/10.1128/JB.00763-19 (2020).
Lee, K. Y. & Lee, B. J. Structure, biology, and therapeutic application of toxin–antitoxin systems in pathogenic bacteria. Toxins (Basel). https://doi.org/10.3390/toxins8100305 (2016).
Ramage, H. R., Connolly, L. E. & Cox, J. S. Comprehensive functional analysis of Mycobacterium tuberculosis toxin–antitoxin systems: Implications for pathogenesis, stress responses, and evolution. PLoS Genet 5, e1000767. https://doi.org/10.1371/journal.pgen.1000767 (2009).
Ooi, W. F. et al. The condition-dependent transcriptional landscape of Burkholderia pseudomallei. PLoS Genet. 9, e1003795 (2013).
Unterholzner, S. J., Poppenberger, B. & Rozhon, W. Toxin–antitoxin systems: Biology, identification, and application. Mob. Genet. Elements 3, e26219 (2013).
Sevin, E. W. & Barloy-Hubler, F. RASTA-Bacteria: A web-based tool for identifying toxin-antitoxin loci in prokaryotes. Genome Biol. 8, R155 (2007).
Makarova, K. S., Wolf, Y. I. & Koonin, E. V. Comprehensive comparative-genomic analysis of type 2 toxin-antitoxin systems and related mobile stress response systems in prokaryotes. Biol. Direct 4, 19. https://doi.org/10.1186/1745-6150-4-19 (2009).
Holden, M. T. et al. Genomic plasticity of the causative agent of melioidosis, Burkholderia pseudomallei. Proc. Natl. Acad. Sci. USA 101, 14240–14245. https://doi.org/10.1073/pnas.0403302101 (2004).
Akarsu, H. et al. TASmania: A bacterial toxin–antitoxin systems database. PLoS Comput. Biol. 15, e1006946 (2019).
Srinon, V., Chaiwattanarungruengpaisan, S., Korbsrisate, S. & Stevens, J. M. Burkholderia pseudomallei BimC is required for actin-based motility, intracellular survival, and virulence. Front. Cell Infect. Microbiol. 9, 63 (2019).
Burtnick, M. N. et al. The cluster 1 type VI secretion system is a major virulence determinant in Burkholderia pseudomallei. Infect. Immun. 79, 1512–1525 (2011).
Burtnick, M. N. & Brett, P. J. Burkholderia mallei and Burkholderia pseudomallei cluster 1 type VI secretion system gene expression is negatively regulated by iron and zinc. PLoS ONE 8, e76767 (2013).
Majerczyk, C. D. et al. Cross-species comparison of the Burkholderia pseudomallei, Burkholderia thailandensis, and Burkholderia mallei quorum-sensing regulons. J. Bacteriol. 196, 3862–3871 (2014).
Scoffone, V. C. et al. Burkholderia cenocepacia infections in cystic fibrosis patients: Drug resistance and therapeutic approaches. Front. Microbiol. 8, 1592 (2017).
Curtis, T. D., Gram, L. & Knudsen, G. M. The small colony variant of Listeria monocytogenes is more tolerant to antibiotics and has altered survival in RAW 264.7 Murine macrophages. Front. Microbiol. 7, 1056 (2016).
Kim, N. H. et al. Small-colony variants in persistent and recurrent Staphylococcus aureus bacteremia. Microb. Drug Resist. 22, 538–544 (2016).
Tuchscherr, L. et al. Staphylococcus aureus develops increased resistance to antibiotics by forming dynamic small colony variants during chronic osteomyelitis. J. Antimicrob. Chemother. 71, 438–448 (2016).
See, J. X. et al. Experimental persistent infection of BALB/c mice with small-colony variants of Burkholderia pseudomallei leads to concurrent upregulation of PD-1 on T cells and skewed Th1 and Th17 responses. PLoS Negl. Trop. Dis. 10, e0004503 (2016).
Butt, A. et al. Burkholderia pseudomallei kynB plays a role in AQ production, biofilm formation, bacterial swarming and persistence. Res. Microbiol. 167, 159–167 (2016).
Lai, S., Tremblay, J. & Déziel, E. Swarming motility: A multicellular behaviour conferring antimicrobial resistance. Environ. Microbiol. 11, 126–136 (2009).
Trastoy, R. et al. Mechanisms of bacterial tolerance and persistence in the gastrointestinal and respiratory environments. Clin. Microbiol. Rev. 31, e00023-18 (2018).
Liu, P. J. et al. Induction of mouse melioidosis with meningitis by CD11b+ phagocytic cells harboring intracellular B. pseudomallei as a Trojan horse. PLoS Negl Trop Dis 7, e2363 (2013).
Jones-Carson, J. et al. Nitric oxide from IFNγ-primed macrophages modulates the antimicrobial activity of β-lactams against the intracellular pathogens Burkholderia pseudomallei and Nontyphoidal Salmonella. PLoS Negl. Trop. Dis. 8, e3079 (2014).
Anutrakunchai, C. et al. Impact of nutritional stress on drug susceptibility and biofilm structures of Burkholderia pseudomallei and Burkholderia thailandensis grown in static and microfluidic systems. PLoS ONE 13, e0194946 (2018).
Cabral, D. J., Wurster, J. I. & Belenky, P. Antibiotic persistence as a metabolic adaptation: Stress, metabolism, the host, and new directions. Pharmaceuticals 11, E14 (2018).
Windels, E. M. et al. Bacterial persistence promotes the evolution of antibiotic resistance by increasing survival and mutation rates. ISME J. 13, 1239–1251 (2019).
Pandey, D. P. & Gerdes, K. Toxin-antitoxin loci are highly abundant in free-living but lost from host-associated prokaryotes. Nucleic Acids Res. 33, 966–976 (2005).
Price, E. P. et al. Within-host evolution of Burkholderia pseudomallei over a twelve-year chronic carriage infection. mBio 4, e00388-313 (2013).
Cheng, A. C. & Currie, B. J. Melioidosis: Epidemiology, pathophysiology, and management. Clin. Microbiol. Rev. 18, 383–416 (2005).
Chow, T. K. et al. Incidental splenic granuloma due to Burkholderia pseudomallei: a case of asymptomatic latent melioidosis?. Am. J. Trop. Med. Hyg. 94, 522–524 (2016).
Mott, T. M., Vijayakumar, S., Sbrana, E., Endsley, J. J. & Torres, A. G. Characterization of the Burkholderia mallei tonB mutant and its potential as a backbone strain for vaccine development. PLoS Negl. Trop. Dis. 9, e0003863 (2015).
Norris, M. H. et al. The Burkholderia pseudomallei Δasd mutant exhibits attenuated intracellular infectivity and imparts protection against acute inhalation melioidosis in mice. Infect. Immun. 79, 4010–4018 (2011).
Stapels, D. A. C. et al. Salmonella persisters undermine host immune defenses during antibiotic treatment. Science 362, 1156–1160 (2018).
Hamad, M. A., Zajdowicz, S. L., Holmes, R. K. & Voskuil, M. I. An allelic exchange system for compliant genetic manipulation of the select agents Burkholderia pseudomallei and Burkholderia mallei. Gene 430, 123–131 (2009).
Acknowledgements
This work was partially supported by NIH NIAID Grant AI148913, the Defense Threat Reduction Agency (DTRA) Grant HDTRA1-12-17-FRCWMD and UTMB funds awarded to AGT. BNR received a NIH Biodefense T32 fellowship. The contents are solely the responsibility of the authors and do not necessarily represent the official views of NIH, DTRA or the Department of Defense (DoD).
Author information
Authors and Affiliations
Contributions
B.N.R. and A.G.T. Designed the experiments, B.N.R., J.D.T., S.M.W. performed the experiments, B.N.R. and A.G.T. wrote the manuscript, B.N.R., J.D.T., S.M.W., I.C.G., and A.G.T. edited and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ross, B.N., Thiriot, J.D., Wilson, S.M. et al. Predicting toxins found in toxin–antitoxin systems with a role in host-induced Burkholderia pseudomallei persistence. Sci Rep 10, 16923 (2020). https://doi.org/10.1038/s41598-020-73887-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-73887-3
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.