Abstract
The capacity to assimilate carbon and nitrogen, to transport the resultant sugars and amino acids to sink tissues, and to convert the incoming sugars and amino acids into storage compounds in the sink tissues, are key determinants of crop yield. Given that all of these processes have the potential to co-limit growth, multiple genetic interventions in source and sink tissues, plus transport processes may be necessary to reach the full yield potential of a crop. We used biolistic combinatorial co-transformation (up to 20 transgenes) for increasing C and N flows with the purpose of increasing tomato fruit yield. We observed an increased fruit yield of up to 23%. To better explore the reconfiguration of metabolic networks in these transformants, we generated a dataset encompassing physiological parameters, gene expression and metabolite profiling on plants grown under glasshouse or polytunnel conditions. A Sparse Partial Least Squares regression model was able to explain the combination of genes that contributed to increased fruit yield. This combinatorial study of multiple transgenes targeting primary metabolism thus offers opportunities to probe the genetic basis of metabolic and phenotypic variation, providing insight into the difficulties in choosing the correct combination of targets for engineering increased fruit yield.
Similar content being viewed by others
Introduction
The yield of the harvested organs of crop plants is influenced by both developmental and metabolic processes1,2,3,4. While the green revolution was underpinned by the former5, major international projects to generate future high yielding crops such as the C4 rice project6,7, RIPE8,9,10, project and CASS11,12 are increasingly focused on the latter. Indeed, there is ample evidence that the net capacity for assimilation of carbon (C) and nitrogen (N) and their subsequent metabolism into the main cellular biomass polymers is a major determinant of crop yield13,14,15,16. For example, an analysis of the historical yield gains achieved in wheat demonstrate that recent yield increases were related to increased photosynthesis and enhanced production of stem CHO reserves17. Furthermore, transgenic interventions have demonstrated that plant growth and yield can be improved by enhancing the catalytic activity of specific enzymes18,19,20,21,22,23,24.
Given the strong need for crop yield improvement there is a substantial interest in the engineering of key metabolic processes for increased source-to-sink C and N flows. There are several major challenges in such engineering projects: first it must be decided which are the key metabolic processes; second, an engineering strategy to increase flux of those processes must be designed; and third the necessary genetic changes to implement this strategy must be made. In choosing the key metabolic processes, researchers have tended to focus either on source processes (e.g. the metabolic assimilation of inorganic C into organic precursors25,26,27,28,29,30); or on sink processes (e.g., the synthesis of starch, lipid or protein in tubers, fruits or seeds31,32,33,34,35). This choice is usually a pragmatic one: there is a limit to the number of genetic interventions that can be made and therefore it makes sense to focus on the process that is thought to impose the greatest limitation on the overall source-to-sink flow. Essentially, this reduces to an argument as to whether a particular crop is source- or sink-limited. Many of the recent consortium projects to increase crop yield are predicated on the argument that crops are source limited36,37, and are thus focusing on source processes such as photosynthesis and N assimilation.
Considerable experimental data is in support of theoretical assessments that both source and sink metabolisms co-limit whole plant fluxes. That said, modulation of net C flow by simultaneous modification of source and sink processes38,39, or alternatively genetically modification of C fluxes via manipulation of individual processes of either source or sink tissues such as photosynthesis40,41 or carbohydrate synthesis31,32,33,42,43,44, respectively, have led to increases in plant growth and yield30,45. Moreover, Nunes-Nesi et al.2 showed that regulation of source-sink interactions is also depending on developmental stage and environmental conditions. Most importantly, there is a strong argument to be made that simultaneous manipulation of source and sink processes lead to a considerable yield increases39,46,47. This is mainly due to signals that communicate and regulate the mechanisms of shifting C flow between source and sink tissues. The potential of this strategy is demonstrated by the only experiments to date to make targeted manipulations of both source and sink39,48. First, expression of transgenes in potato leaves to increase the partitioning of photoassimilates towards sucrose and away from starch was combined with over expression of two transporters to increase the capacity for starch storage in the tuber39. This led to an impressive doubling of potato tuber yield and starch content per plant. Secondly, these studies were achieved with minimal genetic intervention (combined expression of three and one gene – albeit in two specific cell types, respectively). However, the same argument about redistribution of metabolic control applies equally to the local metabolic network as it does to source and sink. For example, it has been suggested that the failure of overexpression of glutamine synthase to consistently increase N assimilation in transgenic crops is due to the lack of simultaneous manipulation of downstream enzymes and transporters49,50. The aim of the current study was therefore to use genetic engineering to relieve potential flux bottlenecks at multiple points in the metabolic networks of both tomato leaves, phloem and fruits with the purpose of substantially increasing fruit yield. To do so we took the emergent combinatorial biolistic transformation approach which promises to revolutionize plant metabolic engineering51. This approach relies on two unique features of biolistic transformation: (1) the regular integration of multiple copies of transgenes, and (2) their usual integration into a single chromosomal locus51,52, with in principle no limit to the number of transgenes that can be integrated simultaneously. Indeed, this route has been taken to achieve increases in three vitamins in maize through the simultaneous integration of five transgenes53. Although impressive, the pathways targeted were easy to engineer because of their position at the periphery of the metabolic network and because of known enzyme deficiencies in each of these pathways in maize54. We aimed to considerably advance the state-of-the-art by systematically manipulating the core of the metabolic network, a substantially greater challenge because of the larger number of targets that we envisage (up to 20 transgenes) and the distributed control of flux in central metabolism. We assessed the transgenic plants that we created with regard to the expression levels of the introduced genes, their photosynthetic parameters and their metabolite composition. The results are discussed in terms of the overall success of the approach and the implications they have for similar scale metabolic engineering approaches in the future.
Results
Generation of tomato plants modifying source and sink metabolisms
Sugar and amino acid accumulation in sink organs is impacted by multiple metabolic and transport processes, ranging from CO2 and NO3 assimilation to the storage and consumption of the products of these assimilation in sink tissues. We here engineered both source and sink tissues by creating transgenic tomato plants containing up to 20 genes involving in different metabolic and transport processes. These target genes were selected based on the characterization of their effects in single-gene transgenic plants and demonstrated to have positive effects on source or sink carbon or nitrogen flows (Table 1).
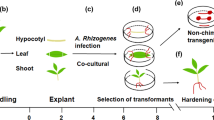
We performed stable co-transformation of tomato plants (cv. MoneyMaker) to simultaneously introduce multiple genes under control of different promoters to confer appropriate tissue specificity (Fig. 1, Supplementary Table S1; Supplementary Note). Using an established combinatorial biolistic co-transformation protocol we were able to generate a total of 18 primary transformant lines (T0), which were grown in the greenhouse to produce seeds (T1). The T1 seeds were germinated on kanamycin-containing media to select for hetero- and homozygous plants. Additionally, the T1 plants were fully genotyped by polymerase chain reaction (PCR) assays using transgene-specific primers that do not amplify the endogenous gene. As result, a different combination of transgenes was inserted in each independent transgenic line is shown in Supplementary Table S1.
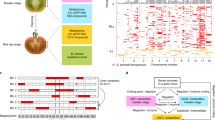
Schematic overview of stable combinatorial-transformation of tomato plants to simultaneously introduce multiple genes under different promoters to confer appropriate tissue specificity. Transgenes are involved in three different processes of carbon and nitrogen fluxes. (i) assimilation ([1] SlmMDH, Solanum lycopersicum mitochondrial malate dehydrogenase; [2] AtSBP, Arabidopsis thaliana sedoheptulose 1,7-bisphosphatase; [3] SlSPA, Solanum lycopersicum sugar partitioning affected; [4] EcPP, Escherichia coli pyrophosphatase; [5] NtGS2, Nicotiana tabacum chloroplast glutamine synthetase 2; [6] FpGLDH, Flaveria pringlei H-protein of glycine decarboxylase); (ii) transport ([7] AtSWEET11, Arabidopsis thaliana sugar efflux transporter 11; [8] AtSUC2, Arabidopsis thaliana sucrose transporter 2; [9] AtAAP1, Arabidopsis thaliana amino acid permease 1); and (iii) sink metabolism ([10,11] AtSUC2/9, Arabidopsis thaliana sucrose transporter 2/9; [12, 13] AtSTP3/6, Arabidopsis thaliana sugar transporter 3/6; [14] SpLIN5, Solanum pennellii tomato apoplastic invertase 5; [15] AtSUS1, Arabidopsis thaliana sucrose synthase 1; [16] ShAgpL1, Solanum habrochaites Large subunit of ADPglucose pyrophosphorylase 1; [17] AtTMT1, Arabidopsis thaliana tonoplast monosaccharide transporter 1; [18] AtAAP6, Arabidopsis thaliana amino acid permease 6; [19] SlINVINH, Solanum lycopersicum apoplastic invertase inhibitor; [20] SlCAT9, Solanum lycopersicum cationic amino acid transporter 9). Overexpression (showed as red color) or silencing (showed as blue color) of these genes were achieved using different tissue-specific promoters; (i) leaf- and mesophyll-specific, ribulose-bisphosphate carboxylase (RbcS), and fructose-1,6-bisphosphate (cyFBP); (ii) constitutive, 35S-cauliflower mosaic virus (35S); (iii) companion cell-specific, commelina yellow mottle virus (CoYMV); (iv) fruit specific, patatin B33 (B33), and ripening-specific ethylene-inducible E8 (E8); and (v) native promoter of S. habrochaites Large subunit of ADPglucose pyrophosphorylase 1 (ShAgpL1). Transgenic lines were grown under glasshouse and polytunnel conditions. SlSPA resides in the plastid but is not known to catalyze an enzymatic reaction, GLDH is associated to the inner mitochondrial membrane where it catalyzes the terminal reaction of ascorbate biosynthesis.
We selected three to ten T1 plants per line to be grown under two different growth conditions; (1) glasshouse under low light (< 450 µmol (photons) m−2 s−1 of Photosynthetically Active Radiation—PAR) and limited soil (i.e. pots contained approx. 0,004 m3 of substrate), and (2) polytunnel (semi-commercial conditions) under high light (> 1200 µmol (photons) m−2 s−1 of PAR) and non-limited soil. Initially, we set up an extra experiment under glasshouse conditions in which tomato plants were allowed to develop naturally (i.e. only side shoots were removed), however we observed that some fruits did not reach ripe stage in all transgenic plants and the two controls. Therefore, we decided to work with pruned plants to standardize and directly compare both grown conditions. Thus, all plants were pruned one week after fruit set to five fruits/truss and three trusses per plant. In addition, due to the normal early fruit-set of the first fruit of each truss, this fruit was removed in order to synchronize growth of fruits in the same truss.
Overview of the changes in carbon- and nitrogen-related genes under low and high light and limited and non-limited soil growth conditions and in different organs
In order to explore the changes in the level of transcription of all transferred genes related to carbon and nitrogen fluxes, we evaluated the relative abundance of all studied transcripts by qRT-PCR in fully expanded leaves from 4 week-old plants and mature red fruits from plants grown in the greenhouse and the polytunnel (Fig. 2). From these analyses, we confirmed that there was a reduction or overexpression of the target gene transcript restricted to tissue specificity expected for the promoter used. It is, however, important to note that a few lines showed changes in gene expression not related to the transgene (for example SBP3 expression was increased in lines 23, 34, 42, 102, and 117 in comparison to control), although these lines were not transformed with this target gene (Supplementary Table S1). In both tissues, gene transcript levels displayed similar patterns of changes in both glasshouse and polytunnel grown conditions (Fig. 2). Effect of growth conditions and genotypes (lines) on gene expression is shown as Supplementary Data and Supplementary Table S2.
Gene expression of genes involved in carbon and nitrogen fluxes. Expression by quantitative real-time PCR (qRT-PCR) of AAP1, SBP, SUC2, PP, GLDH, GS2, CAT9, INVINH, mMDH, SPA, AAP6, SBP3, STP6, LIN5, SUC9, SUS1, TMT1, and AgpL1 genes in transgenic lines under glasshouse and polytunnel conditions in fully expanded leaves and mature red fruits. The increase or decrease in expression of each gene is shown relative to the control value. Error bars indicate means ± SD. Asterisks indicate the values that were determined by the t-test to be significantly different (P < 0.05) from control. Note the different axes scale in the independent plots. This data is plotted with the individual data points visible in Supplementary Table S8.
Detailed phenotypic analysis of transgenic lines under low light, limited soil and high light, non-limited soil grown conditions
To further characterize these lines, we first performed a detailed phenotypic analysis of the plants grown either in glasshouse or polytunnel conditions. Phenotypic variation in terms of photosynthesis, dark respiration, stomatal conductance, chloroplast electron transport rate (ETR) was measured prior to flowering. In general, variation of these traits were largely similar between the growth conditions. However, large variability was evident in some lines for some traits when comparing the growth conditions (Supplementary Figure S1). In particular, we observed a decrease in (1) photosynthesis in lines 42 and 116; (2) dark respiration in lines 14, 23, 102, and 121; (3) ETR in lines 8, 42, 116, and 128 when comparing with control plants (Supplementary Fig. S1).
When analyzing fruit ripening-related traits, five lines (in particular, lines 8, 30, 111, 117 and 121) flowered significantly earlier than their respective controls in the glasshouse or polytunnel, respectively (Supplementary Figure S2A,B). Moreover, as would perhaps be anticipated, the same lines produced red fruit earlier than controls. By contrast, some lines displayed later flowering time in comparison to controls (Supplementary Figure S2C,D). Namely, when plants were grown in the polytunnel, the late flowering of lines 2 and 42 correlated with a later appearance of the first red fruit. Similarly, lines 128 and 140 showed the same behavior in the greenhouse (Supplementary Figure S2D). We next determined yield parameters of mature fruit. In glasshouse, two transformants (lines 111 and 116) displayed mild reductions in fruit yield, however it is important to note that four lines (lines 14, 36, 102, and 121) showed a significantly increased fruit yield ranging from 13.5 to 23% (Table 2). Interestingly, when transformants were grown in the polytunnel the same behavior was observed for these lines but also for lines 117 and 133 (Table 2). Moreover, the lines showing higher yield also exhibited a clear increase in the total soluble solids (Brix) content of their fruits (Table 2). By contrast, the same lines displayed unaltered or even mild decreases in Brix content when grown in the glasshouse.
Metabolite profiling reveals differential metabolic responses to light and soil growth conditions
In order to gain a deeper understanding of the metabolic changes underlying the above-mentioned increased yield in the transgenic lines (glasshouse [experiment 1], lines 14, 36, 102 and 121; polytunnel [experiment 2], lines 14, 36, 102, 117, 121, 133), we next determined metabolite levels in the pericarp tissue of mature fruit harvested from plants grown under both growth conditions using a gas chromatography-time of flight-mass spectrometry (GC–TOF–MS)-based metabolite profiling method. A total of 47 primary metabolites were annotated after this analysis and their relative levels were normalized of each sample for each grown condition (Supplementary Tables S3 and S4). In addition, metabolite levels were analyzed on a dry weight basis to avoid the effect of differential water contents.
Each dataset was examined by principal component analysis (PCA) (Supplementary Figure S3). For fruits from plants grown in the glasshouse (experiment 1), clear differences were evident between the analyzed genotypes. However, for fruits of the high light, non-limited soil growth conditions (polytunnel; experiment 2) PCA clearly separated the genotypes along PC2, with the exception of line 121 that was separated along PC1. Overall the global composition changes induced in mature fruit in experiment 2, high light and non-limited soil grown conditions (polytunnel), seem lower than those recorded in experiment 1 (glasshouse).
The effects of the genetic intervention on the levels of individual metabolites are summarized in Supplementary Tables S3 and S4. Of the compounds analyzed, approximately 50% were significantly altered in experiment 1 (glasshouse) while more than 80% were significantly altered in experiment 2 (p < 0.05) (Fig. 3). Some metabolites showed a clear tendency of differential accumulation across both experiments. For example, glutamine, methionine, alanine, and putrescine accumulated in both experiments while others such as malic acid, lysine, and valine decreased (Figs. 3 and 4). Under low light and limited soil conditions (experiment 1, glasshouse), sucrose, glucose, fructose, rhamnose, galactonic acid, and proline were reduced in the high yielding transgenics in comparison to the control line. By contrast, these metabolites accumulated under in high light and non-limited soil conditions (experiment 2) in the high yielding transgenics in comparison to the control line. Decreased contents of phenylalanine and glycine were observed under both conditions, whereas ß-alanine was decreased only in polytunnel grown transgenics. Moreover, increased contents of aspartic acid, citric acid, tryptophan and isoleucine were observed solely in transgenic plants grown in polytunnel conditions.
Hierarchical clustering of the primary metabolite data from selected transgenic lines under glasshouse (A) and polytunnel (B) conditions. Relative metabolite levels were normalized (Z-Score) of each sample for each grown condition and to dry weight. Each biological replicate is shown independently. For negative controls, WT and PH200 were used (PH200 was originated from an independent transformation, containing only the nptII gene under 35S promoter). Full documentation of metabolite profiling data acquisition is provided in Supplementary Table S3 and S4. Data analysis and graphical representation were performed using R Software (https://www.R-project.org/).
Schematic representation of metabolite changes occurring in selected transgenic lines. The heat maps represent the Log2 of the fold change level of metabolites with respect to the control in plants under glasshouse (violet-green) and in polytunnel (red-blue) conditions. Changes that were significant in the statistical analysis are denoted with an asterisk. The lines have been ordered by yield increase (Table 2).
We next investigated the strength of correlations (based on Pearson correlation coefficients at the threshold of p < 0.05) between the levels of each metabolite and fruit yield in either the glasshouse or polytunnel experiment. We postulate that this would allow us to identify metabolites closely related to fruit yield under the different growth conditions. In the polytunnel grown plants levels of aspartic acid displayed a positive correlation while raffinose displayed a negative correlation to fruit yield (Fig. 5, Supplementary Table S5). Under glasshouse condition, levels of rhamnose and galactonic acid displayed negative correlation with fruit yield (Fig. 5, Supplementary Table S5). This finding suggests that these metabolites are possible candidate metabolite biomarkers related to fruit yield and highlights that the key points of regulation vary depending on the environmental conditions.
Correlation between metabolite levels and fruit yield under (A) glasshouse and (B) polytunnel conditions. Levels of selected metabolites showing significant correlation (p < 0.05) were plotted (B) and (D) against fruit yield. Correlation coefficient and p-value were calculated based on Pearson correlation analysis. Data analysis and graphical representation were performed using R Software (https://www.R-project.org/).
Sparse partial least squares (sPLS) regression modeling can predict fruit yield from a combination of transcript levels
We next constructed a sparse Partial Least Squares (sPLS) regression model in order to ascertain if we could identify genes that could highly affect fruit yield in each growth condition (glasshouse and polytunnel) and also distinguish leaf and fruit tissues55. The model is creating variable importance in the projection (VIP) coefficients of the relative importance of each independent variable (in this instance the gene expression levels measured in this study Fig. 2, Supplementary Fig. S4), for each dependent variable (yield) of every single combinatorial experiment. In other words, the greater the VIP coefficient the greater the explanatory power with regard to yield. The model was applied to data coming from each growth condition (experiment 1; glasshouse and experiment 2; polytunnel) as well as to distinguish variables from different tissues (leaves and fruits). We ran three independent simulations for the leaves, fruits and the combination of leaves and fruits, respectively (Table 3).
In leaves, we identified that the SPA protein contributed most significantly to variation of fruit yield under low light and limited soil (glasshouse). In addition to this protein, we also observed that pyrophosphatase and the invertase inhibitor were highly significant contributors for describing the variation in yield under high light and non-limited soil conditions (polytunnel) (Table 3).
When estimating the VIP coefficients in fruit, a total of three (under glasshouse conditions) and six (under polytunnel conditions) proteins displayed high VIP values, suggesting the significant contribution of these proteins to explain fruit yield variation under the two different grown conditions, respectively (Table 3). These proteins are: sugar partitioning affecting protein (SPA), sucrose transporter 2 (SUC2), and amino acid permease 6 (AAP6) for glasshouse conditions, and mitochondrial malate dehydrogenase (mMDH), H-protein of glycine decarboxylase (GLDH), sucrose transporter 2 (SUC2), amino acid permease 6 (AAP6), apoplastic invertase 5 (LIN5), and cationic amino acid transporter 9 (CAT9) for polytunnel (Table 3).
Furthermore, when calculating the VIP coefficients in the joint dataset (leaves and fruits combined), we observed that a large proportion of the enzymes contributing to the variation of fruit yield could be explained by the additive effects of the individual analysis for each tissue (Table 3). This confirms the importance of the expression of SPA, pyrophosphatase and the invertase inhibitor in leaves and LIN5 and AA6 in fruits Moreover, the modeling of the combined data set highlighted two transporters, amino acid permease 1 (AAP1), and sucrose transport 9 (SUC9), that also exhibited significant contribution to explain fruit yield variation only under glasshouse condition (Table 3). Whilst on the basis of the current study we cannot formally state if the variation in gene expression and enzyme activity lies in the genetic diversity or in the genotype-environment interaction, it is evident that the three processes of assimilation, transport, and sink metabolism are important in determining the fruit yield.
Discussion
Current agriculture faces a considerable challenge with respect to securing food for the growing population on the planet, a fact that is exacerbated by the deteriorating environment and increasing pressure for land use. It is, therefore, becoming imperative to develop strategies which enable us to substantially increase crop yields on existing farmland56. Numerous studies have shown that partitioning and allocation of C and N assimilates play an essential role in crop yield. Considering that source-sink partitioning is determined by the synchronization of a highly complex signaling network that also embraces developmental processes12, there is a substantial interest in the engineering of key metabolic processes for increased C and N flow. Several published studies have determined that high availability of C sources leads to higher C accumulation on the sink57,58. However, there are also a number of previous studies of sink-dependent alteration of photosynthesis of source leaves by using single-transgene transformation59,60,61,62,63. This suggests that the photosynthetic activity of source tissues is controlled either by the metabolism of photoassimilates within source tissue, insufficient sink strength or inhibition of their transport64. This hypothesis is further supported by experiments in potato and pea which indicate that transgenic manipulation of both source and sink is a highly effective route for enhancing the harvest index of a crop species39,48. Recently, a multi-transgenic approach has been used that targeted both C and N metabolism was proven to be effective in enhancing Arabidopsis growth65. Our study expands on the basis of those above by generating multi-transgenic tomato plants that are affected in both source and sink metabolism to simultaneously increase the flow of C and N from leaves to fruit with a view to altering yield. The aim of this work was to determine the importance of twenty proteins previously implicated (see the summary in Table 1), in diverse processes of source-sink partitioning, in the reconfiguration of plant metabolism required to increase fruit yield.
In search of the combination with the greatest impact on yield, we expressed different genes under diverse promoters in order to achieve a range of protein overexpression or silencing. For overexpression, to achieve high expression levels, we used the CaMV 35S viral promoter which has been widely and successfully used in the past to drive high expression of transgenes66. In addition, RbcS, cyFBP, CoYMV, Patatin B33, and E8 promoters allowed us to achieve intermediate level expression and leaf-, mesophyll-, companion cell-, fruit- and fruit ripening- specific expression, respectively. For gene silencing, either the RbcS or the CoYMV promoter was used. We subsequently evaluated the physiological and metabolic effects of these genetic interventions under two different grown conditions, (1) glasshouse under relative low light (< 450 PAR) and limited soil (pots contained approx. 0.004 m3 of substrate), and (2) polytunnel (semi-commercial conditions) under high light (> 1200 PAR) and non-limited soil.
We observed common transgenic lines (namely L14, L36, L102, L121) exhibiting significantly increased fruit yield in our experiments under both low light, limited soil conditions and high light, non-limited soil growth conditions. In addition, two more transgenic lines (L117 and L133) displayed elevated fruit yield in comparison to control plants under high light, non-limited soil conditions. That said, the rest of transgenic lines did not display consistent differences across the experiments rendering it difficult to associate phenotypic and metabolic characteristics of these plants with fruit yield. Focusing exclusively on the transgenic lines displaying increased fruit yield, we observed that these plants produced heavier fruits although the number of fruits were identical since the plants had previously been pruned. Moreover, neither morphological not developmental alterations appeared under both grown conditions (greenhouse and polytunnel). Given the lack of significant alteration in photosynthetic parameters our results indicate a more efficient transfer of photoassimilate between source and sink. This hypothesis was supported by the analysis relating gene expression and fruit yield by applying a sparse Partial Least Squares (sPLS) regression model on leaves and fruits separately. When the transcript levels relation was tested in leaves under low light, limited soil grown conditions, we found that only the expression of the Sugar Partitioning-Affecting (SPA) gene, exhibited a high VIP value with fruit yield. Our analysis is in line with the observation that deficiency of this protein, which is encoded by a single gene in tomato67, leads to a pronounced phenotype, with increased harvest index and reduction in the level of sucrose, glucose and fructose in leaves68. These changes indicate that SPA activity promotes carbon export from leaves to sink organs. Interestingly, under the same grown condition, when we tested the regression model on fruit, expression of SUC2 and AAP6 genes appeared to be important, in addition to SPA, to explain higher fruit yield under low light, limited soil grown conditions. APP6 has been described to play a role in xylem-phloem transfer69. This hypothesis is supported by showing a reduction in amino acid contents of sieve elements in aap6 mutant in Arabidopsis70. Moreover, this mutant did not display a strong phenotype, only a slight increase in leaf width and seed size. Interestingly, the third candidate gene highlighted from the model was SUC2, an apoplastic loader, stressing the importance of sugar movement system across the plasma membrane for phloem loading to increase fruit yield. In particular, sucrose is loaded into the sieve element-companion cell complex in the phloem by the sucrose-H+ co-transporter SUC2 from the apoplasm (cell wall space)71. Interestingly, potato plants that expressed reduced levels of this sucrose transporter showed a dramatic reduction in tuber yield, supporting the importance of transport capacity for growth and development of the plant71.
When the above approach was used to identify genes that highly affect fruit yield in leaves from plants grown under high light and non-limited soil condition, we found that two proteins having a role in assimilation of carbon, soluble pyrophosphatase (PP) and in sink metabolism, apoplastic invertase inhibitor (INVINH), were identified to have high contributions to explain increased fruit yield on plants grown in polytunnel. These results pointed to the importance of increase the gradient of translocation from source to sink and hence the net import into the fruit under high light grown condition. Consistent with this hypothesis, overexpression of E. coli PP previously described in tobacco and potato resulted in sugar-storing leaves72,73—a feature which could subsequently be exploited by re-routing these photoassimilates to the sink organs39. In particular, transgenic lines of tobacco and potato showed perturbed sink growth but different responses. In tobacco, plant growth was inhibited, while potato plants produced a larger number of smaller tubers in comparison to controls72,73. In addition, Jin et al.74 showed that decreasing the INVINH activity in tomato correlated with an increased fruit sugar level and seed size without a negative impact on fruit yield.
Finally, a tight co-regulation of C-N metabolism was observed in fruits from plants grown under high light and non-limited soil conditions, since the combination of six protein activities (named as mMDH, GLDH, SUC2, AAP6, LIN5, and CAT9) were needed to significantly explain the increased fruit yield. In particular, these results illustrate the intertwined crosstalk of metabolic pathways through assimilation, transport, and sink metabolism of photoassimilates for the maintenance of carbon and nitrogen metabolism to increase fruit yield. In this sense, our data support the hypothesis of enhance fruit yield under high light grown condition only through a tightly coordinated increase in carbon assimilation, export, and utilization. This scenario is in agreement with previous studies in which reduced activity of mMDH detected in source leaves correlated with an induction of photosynthetic metabolism in leaves, resulting in increased fruit yield75; however, fruit-specific antisense suppression of this enzyme resulted in a relatively small effect on total fruit yield76. Moreover, using an in vitro assay, Hasse et al.77, demonstrated that increased glycine decarboxylase (GLD) H-protein supply enhances the activity of GLD P-protein, an essential protein for the interconversion of glycine and serine in photorespiration78. Furthermore, overexpression of GLDH resulted in an increase in photosynthesis and yield24,79. The present data suggest that the principal tomato phloem unloading under high light grown condition to favor an increase in fruit yield may be apoplastic through the activity of LIN5 protein as previously described80,81. This hypothesis is supported by the facts that reduction of LIN5 activity in tomato plants resulted in a compromised fruit yield, approximately 40% reduction of that showed for wild type81. CAT9 activity was also significantly identified to explain the variation of fruit yield under high light and non-limited grown conditions. CAT9 has been identified as tonoplast-localized transporter that facilities the exchange of glutamic acid, aspartic acid and GABA. This may result from the importance of GABA metabolism in signaling, redox regulation, energy production and the maintenance of carbon/nitrogen balance82, however, further studies are required in order to elucidate the role of this protein in the elevation of tomato yield. Another aim of this study was to identify whether there were metabolic features that rendered the transgenic lines that displayed higher yield. In this regard, we made some interesting observations further discussed in Supplementary Discussion that lead to a more complete understanding of the metabolic process in tomato to improve source-to-sink partitioning and thereby yield.
Conclusion
The primary aim of this work was to test if a multi-step metabolic engineering of primary metabolism could be utilized to improve source-to-sink partitioning and thereby yield. For this purpose we introduced up to 20 transgenes targeted at step in source and sink metabolism as well as at the transport process itself. Under two different growth regimes we were able to identify a subset of the 20 obtained transgenic lines which had a similar magnitude of effect on yield as was achieved by single-transgene transformations but were not able to isolate lines in which the increase in yield was in excess of that previously achieved. Several possible reasons can be postulated for this however we find two of these to be most likely. Firstly, it is highly possible that we did not screen enough transgenic lines in this study to ensure that the optimal expression level of the transgenes was achieved. Secondly, it is additionally possible that our understanding of metabolism is not quite at the level whereby we can rationally “pick and mix” the best combinations of genes. It is important to note that one possible reason that we did not observe genotypes exhibiting higher yield than that achieved following single transgene manipulation was the growth space constraints in a research laboratory setting (although the growth space we utilized was considerably). As such, industrial-scale testing of this approach may allow isolate of such successful genotypes given that testing all the combinations of expression would need a vast amount of independent transformants. Since the initiation of this project a handful of elegant papers boosting tomato yield by affecting development associated genes have been published83,84. It seems likely that, as was recently postulated83, approaches incorporating both metabolic and developmental genes would be more likely to result in larger yield increases than reported here. Despite the biolistic combinatorial co-transformation approach taken here not being highly successful from a biotechnological standpoint it did provide considerable insight into source-sink partitioning. Indeed, both the physiological and metabolic measurements support the conclusion that the phloem transport step is highly important in determining source-sink relations in tomato whilst the importance of source and sink metabolism per se is more context dependent. That said under commercial growth conditions it would seem likely that all three processes co-limit tomato fruit yield.
Methods
Plant material
Tomato plants (Solanum lycopersicum cv. Moneymaker) were grown under sterile conditions on agar-solidified MS medium85 supplemented with 20 g/L sucrose. Genetically modified plants were propagated and rooted in the same medium additionally containing 35 mg/L kanamycin. For sampling and seed production, plants were transferred to soil and grown under experimental growth conditions.
Experimental growth conditions
Three to ten T1 plants per line were cultivated under two types of semi-controlled conditions. (1) In “experiment 1”, plants were grown in a glasshouse as previously reported86. Plants in the “experiment 1” were exposed to low light (< 450 µmol photons m−2 s−1 of Photosynthetically active radiation-PAR) and limited soil (i.e. pots contained approx. 0.004 m3 of substrate) at controlled temperature 24 °C/16 °C day/night. The plants were irradiated with supplemental light to maintain an irradiance close to 400 μmol photons m−2 s−1. (2) In “experiment 2”, plants were cultivated in polytunnel conditions (similar to semi-commercial conditions), with high light (> 1200 µmol photons m−2 s−1) and non-limited soil. Plants were pruned one week after fruit set to five fruits per truss and three trusses per plant. In addition, due to the normal early fruit-set of the first fruit in each truss, this fruit also was removed in order to avoid unbalanced growth between fruits of the same truss. Systematically, every week side shoots and new flowers were removed. Young fully expanded leaves were harvested from 4 week-old-plants. The stage of fruit development was followed by tagging the truss upon appearance of the flower. Pericarp samples were harvested from mature red fruit. Harvested fruits were weighed, and pericarp was separated from the placental tissue, weighed, and then immediately frozen in liquid nitrogen before being stored at − 80 °C until further analysis.
Construction of transformation vectors
Transformation vectors (pSKJ1, 2, 3, 6, 8, 10, 12, 15, 16, 18, 20, 22, 24, 26, 28, 30 ad 32) were constructed based on the pUC18 plasmid, containing the cauliflower mosaic virus (CaMV 35S) promoter region upstream of the multiple cloning site (MCS) and the nopaline synthase nos terminator sequence downstream of the MCS. Full coding sequences of genes of interest (GOI) were amplified using a standard PCR protocol from donated plasmids, amplified from cDNA as a template or synthesized commercially (GeneCust, France). GOI sequences were subcloned into the pUC18 backbone via standard restriction enzyme type IIS and ligation-based protocol. Where needed the 35S promoter sequence was exchanged for a number of tissue-specific promoters such as Commelina yellow mottle virus (CoYMV) promoter region, B33 Patatin promoter region, Solanum tuberosum cytosolic fructose-1,6-bisphosphatase (StcyFBP) promoter region, Solanum lycopersicum small subunit of Rubisco (SlRbcS) promoter region, ethylene-inducible, ripening-specific (E8) promoter region and a native promoter region of the Solanum habrochaites ADP-glucose pyrophosphorylase Large subunit 1.Silencing vectors (pSKJ33 and pSKJ35) were constructed based on the pK7GWIWG2(I) destination vector according to the Gateway cloning protocol (Supplementary Table S6). Prior to transformation all constructs were validated by sequencing and GOI sequences were confirmed.
The plasmid cocktail (pSKJcombi1) for combinatorial transformation was prepared by mixing equal quantities of pSKJ1, 2, 3, 6, 8, 10, 12, 15, 16, 18, 20, 22, 24, 26, 28, 30, 32, 33, 35 and pK7GWIWG2(I)_SlSPA68 (each at a concentration of 2 µg/µL) and plasmid pPH200 that contains the nptII gene for kanamycin resistance between the 35S promoter and terminator (Supplementary Table S6).
Combinatorial nuclear transformation and selection of transgenic tomato plants
Young leaves from plants grown under aseptic conditions were harvested and bombarded with gold particles coated with a plasmid DNA mixture pSKJ-combi1 (Supplementary Table S6) using the DuPont PDS1000He biolistic gun as previously described by Elghabi et al.87. Kanamycin-resistant shoots were selected on plant regeneration medium containing 2.0 mg/L Zeatin, 0.1 mg/L IAA, 0.5 g/L MES and 35 mg/L kanamycin. Resistant shoots were rooted in agar-solidified MS medium, then transferred to soil and grown to maturity under standard greenhouse conditions. As negative controls wild type (WT) plants were used, as well as PH200 line, which contained only the nptII gene controlled by 35S promoter. The PH200 line, was originated from an independent transformation. Material from T0 plants was harvested and used for initial molecular analysis.
Isolation of nucleic acids
Tomato leaf genomic DNA was isolated using a CTAB-based protocol88 and used for genotyping. For total tomato leaf RNA extraction, samples of 100 mg of frozen leaf powder material were extracted with the NucleoSpin RNA Plant kit following the manufacturer’s instructions (Macherey–Nagel, Düren, Germany. The RNA was eluted in 60 µl of RNase-free water and stored at − 80 °C until used for the cDNA synthesis. Tomato pericarp RNA was obtained using the TRIZOL reagent according to the manufacturer’s instructions. Obtained RNA was additionally purified using the NucleoSpin RNA Plant kit.
cDNA synthesis
Isolated RNA was tested for the presence of DNA contamination by a standard PCR using 1 ng of RNA as template. cDNA was synthesized using the SuperScript III Reverse Transcriptase kit according to the manufacturer’s instructions (Invitrogen, Carlsbad, CA). The quality of the cDNA was tested by a standard PCR reaction.
Genotyping
Genotyping of transgenic lines was performed using genomic DNA isolated from 2-week old seedlings germinated on kanamycin-containing media. Gene-specific primers were used for genotyping. Genotyping was performed using a standard PCR protocol.
Gene expression analysis by quantitative real-time PCR (qRT-PCR)
Quantitative RT-PCR was performed in a LightCycler 480 (Roche, Mannheim, Germany) using cDNA as template in 5 µL reactions containing 1 µL of each gene-specific primer (1.25 µM; Supplementary Table S7), 2.5 µL of the LightCycler 480 SYBR green I Master mix and 0.5 µL of a 1:50 cDNA dilution. Two biological replicates (independent plants) and three technical replicates per line were analyzed. The relative transcript levels were determined using the formula (1 + E)−ΔΔCp where E is the binding efficiency of the primers89. Expression data were normalized to the reference gene SlFRG03 (Solyc02g063070) according to Cheng et al., 201790.
Metabolite analysis
Metabolite extraction, derivatization, and sample injection for gas chromatography coupled to electron impact ionization-time of flight-mass spectrometry (GC-EI-TOF/MS) were performed according to Osorio et al.91. Chromatograms and mass spectra were evaluated using ChromaTOF 1.0 (Leco, www.leco.com) and TagFinder v.4.092, respectively Cross-referencing of mass spectra was performed with the Golm Metabolome database93. Data is reported following the standards suggested in Fernie et al.94.
Measurement of fruit °Brix and yield
Ripe fruit tissue was homogenized with a razor blade, and the soluble solids (Brix) content of the resulting juice measured on a portable refractometer (Digitales Refrktometer DR6000; Krüss Optronic GmbH, Hamburg, Germany). Fruit yield was determined in red fruit considering each biological replicate the weight of 15 fruits per individual plant.
Measurements of photosynthetic parameters
Leaf gas exchange and chlorophyll a fluorescence were measured simultaneously with an open infrared gas‐exchange analyser system equipped with a leaf chamber fluorometer (Li‐6400XT, Li‐Cor Inc., Lincoln, NE, USA). The measurements were performed during mornings (9:00–11:00 h) in full expanded leaves at growth light (i.e. Glasshouse: 450 µmol (photons) m−2 s−1 of PAR, and Polytunnel 1200 µmol (photons) m−2 s−1 of PAR) while the amount of blue light was set to 10% photosynthetically active photon flux density to optimize stomatal aperture. The reference CO2 concentration was set at 400 µmol CO2 mol–1 air. All measurements were performed using the 2 cm2 leaf chamber maintaining the block temperature at 25 °C and flow rate 300 mmol air min–1. Dark respiration and maximum quantum efficiency of PSII (Fv/Fm) were measured during mornings in leaflets after 2 h of dark adaptation. Relative electron transport rate (rETR) was calculated according to Krall and Edwards95. The photorespiration rate was calculated following the model based on gas exchange and Chl fluorescence measurements proposed by Valentini et al.96.
Data analysis
Data mining, normalization, clustering and graphical representation were performed using R Software (https://www.R-project.org/) and pheatmap: Pretty Heatmaps. R package version 1.0.12. (https://CRAN.R-project.org/package=pheatmap). Sparse Partial Least Squares (sPLS) regression model was performed using quantitative data. In particular, the levels of transcripts as independent variables and fruit yield under glasshouse and polytunnel conditions as dependent variables. Six different matrixes were used to feed the model; i.e.: in glasshouse (experiment 1) (i) leaf, (ii) fruit gene expression and (iii) the mixed matrix considering both datasets. Same manner, the matrixes (iv), (v) and (vi) with data coming from polytunnel (experiment 2). To determine the optimal number of components and variables of a given model, we searched the parameter space spanned all possible component combinations. For each such component/variable combination, 100 iterations of fivefold cross-validation rounds were tested. One an optimal number of components and variables was determined for each response variable, we obtained the variable importance in projection (VIP) coefficients reported. This analysis was performed using the package mixOmics97.
References
Hanson, A. D. & Roje, S. One-carbon metabolism in higher plants. Annu. Rev. Plant Phys. 52, 119–137 (2001).
Nunes-Nesi, A., Fernie, A. R. & Stitt, M. Metabolic and signaling aspects underpinning the regulation of plant carbon nitrogen interactions. Mol. Plant. 3, 973–996 (2010).
Redestig, H. et al. Exploring molecular backgrounds of quality traits in rice by predictive models based on high-coverage metabolomics. BMC Syst. Biol. 5, 176 (2011).
Genard, M., Baldazzi, V. & Gibon, Y. Metabolic studies in plant organs: don’t forget dilution by growth. Front Plant Sci 5, 85 (2014).
Borlaug, N. Nobel Lecture. The Green Revolution, Peace, and Humanity. NobelPrize.org: Nobel Media AB (1970).
Schuler, M. L., Mantegazza, O. & Weber, A. P. M. Engineering C-4 photosynthesis into C-3 chassis in the synthetic biology age. Plant J 87, 51–65 (2016).
Ermakova, M., Danila, F. R., Furbank, R. T. & von Caemmerer, S. On the road to C-4 rice: advances and perspectives. Plant J. 101(4), 940–950 (2020).
Kromdijk, J. et al. Improving photosynthesis and crop productivity by accelerating recovery from photoprotection. Science 354, 857–861 (2016).
Glowacka, K. et al. Photosystem II Subunit S overexpression increases the efficiency of water use in a field-grown crop. Nat. Commun. 9(1), 868 (2018).
South, P.F., Cavanagh, A.P., Liu, H.W. & Ort, D.R. Synthetic glycolate metabolism pathways stimulate crop growth and productivity in the field. Science. 363 (6422), eaat9077 (2019).
Obata, T. et al. Metabolic profiles of six African cultivars of cassava (Manihot esculenta Crantz) highlight bottlenecks of root yield. Plant J. https://doi.org/10.1111/tpj.14693 (2020).
Fernie, A. R. et al. (2020) Synchronization of developmental, molecular and metabolic aspects of source–sink interactions. Nat. Plants. 6(2), 55–66 (2020).
Paul, M. J. & Foyer, C. H. Sink regulation of photosynthesis. J. Exp. Bot. 52, 1383–1400 (2001).
Gregersen, P. L., Culetic, A., Boschian, L. & Krupinska, K. Plant senescence and crop productivity. Plant Mol. Biol. 82, 603–622 (2013).
Sulpice, R. et al. Impact of the carbon and nitrogen supply on relationships and connectivity between metabolism and biomass in a broad panel of Arabidopsis Accessions. Plant Physiol. 162, 347–363 (2013).
Ambavaram, M. M. R. et al. Coordinated regulation of photosynthesis in rice increases yield and tolerance to environmental stress. Nat. Commun. 5, 5302 (2014).
Driever, S. M., Lawson, T., Andralojc, P. J., Raines, C. A. & Parry, M. A. J. Natural variation in photosynthetic capacity, growth, and yield in 64 field-grown wheat genotypes. J. Exp. Bot. 65(17), 4959–4973 (2014).
Nunes-Nesi, A., Carrari, F., Lytovchenko, A. & Fernie, A. R. Enhancing crop yield in Solanaceous species through the genetic manipulation of energy metabolism. Biochem Soc. T. 33, 1430–1434 (2005).
Fujita, D. et al. NAL1 allele from a rice landrace greatly increases yield in modern indica cultivars. Proc. Natl. Acad. Sci. USA 110, 20431–20436 (2013).
Park, S. J. et al. Optimization of crop productivity in tomato using induced mutations in the florigen pathway. Nat. Genet. 46, 1337–1342 (2014).
Driever, S. M. et al. Increased SBPase activity improves photosynthesis and grain yield in wheat grown in greenhouse conditions. Philos. T. R. Soc. B 372(1730), 20160384 (2017).
Leach, K. A., Tran, T. M., Slewinski, T. L., Meeley, R. B. & Braun, D. M. Sucrose transporter2 contributes to maize growth, development, and crop yield. J. Integr. Plant Biol. 59, 390–408 (2017).
Bezrutczyk, M. et al. Impaired phloem loading in zmsweet13a, b, c sucrose transporter triple knock-out mutants in Zea mays. New Phytol. 218, 594–603 (2018).
Lopez-Calcagno, P. E. et al. Overexpressing the H-protein of the glycine cleavage system increases biomass yield in glasshouse and field-grown transgenic tobacco plants. Plant Biotechnol. J. 17, 141–151 (2019).
Smith, A. M. & Stitt, M. Coordination of carbon supply and plant growth. Plant Cell Environ. 30, 1126–1149 (2007).
Sulpice, R. et al. Starch as a major integrator in the regulation of plant growth. Proc. Natl. Acad. Sci. USA 106, 10348–10353 (2009).
Nunes-Nesi, A., Araujo, W. L. & Fernie, A. R. Targeting mitochondrial metabolism and machinery as a means to enhance photosynthesis. Plant Physiol. 155, 101–107 (2011).
Braun, D. M., Wang, L. & Ruan, Y. L. Understanding and manipulating sucrose phloem loading, unloading, metabolism, and signalling to enhance crop yield and food security. J. Exp. Bot. 65, 1713–1735 (2014).
Yang, L.Y., Wang, L.T., Ma, J.H., Ma, E.D., Li, J.Y. & Gong, M. Effects of light quality on growth and development, photosynthetic characteristics and content of carbohydrates in tobacco (Nicotiana tabacum L.) plants. Photosynthetica. 55, 467–477 (2017).
Tegeder, M. & Masclaux-Daubresse, C. Source and sink mechanisms of nitrogen transport and use. New Phytol. 217, 35–53 (2018).
McCormick, A. J., Cramer, M. D. & Watt, D. A. Sink strength regulates photosynthesis in sugarcane. New Phytol. 171, 759–770 (2006).
Weichert, N. et al. Increasing sucrose uptake capacity of wheat grains stimulates storage protein synthesis. Plant Physiol. 152, 698–710 (2010).
Ainsworth, E. A. & Bush, D. R. Carbohydrate export from the leaf: a highly regulated process and target to enhance photosynthesis and productivity. Plant Physiol. 155, 64–69 (2011).
Muller, B. et al. Water deficits uncouple growth from photosynthesis, increase C content, and modify the relationships between C and growth in sink organs. J. Exp. Bot. 62, 1715–1729 (2011).
Sekhon, R. S. et al. Integrated genome-scale analysis identifies novel genes and networks underlying senescence in maize. Plant Cell. 31, 1968–1989 (2019).
Zhu, X. G., Long, S. P. & Ort, D. R. Improving Photosynthetic Efficiency for Greater Yield. Annu. Rev. Plant Biol. 61, 235–261 (2010).
Ray, D. K., Ramankutty, N., Mueller, N. D., West, P. C. & Foley, J. A. Recent patterns of crop yield growth and stagnation. Nat. Commun. 3, 1293 (2012).
Ferreira, S. J. & Sonnewald, U. The mode of sucrose degradation in potato tubers determines the fate of assimilate utilization. Front. Plant Sci. 3, 23 (2012).
Jonik, C., Sonnewald, U., Hajirezaei, M. R., Flugge, U. I. & Ludewig, F. Simultaneous boosting of source and sink capacities doubles tuber starch yield of potato plants. Plant Biotechnol. J. 10, 1088–1098 (2012).
Long, S. P., Zhu, X. G., Naidu, S. L. & Ort, D. R. Can improvement in photosynthesis increase crop yields?. Plant Cell Environ. 29, 315–330 (2006).
Kirschbaum, M. U. F. Does enhanced photosynthesis enhance growth? Lessons Learned from CO2 Enrichment Studies. Plant Physiol. 155, 117–124 (2011).
Rosche, E. et al. Seed-specific overexpression of a potato sucrose transporter increases sucrose uptake and growth rates of developing pea cotyledons. Plant J. 30, 165–175 (2002).
Li, B. et al. Constitutive expression of cell wall invertase genes increases grain yield and starch content in maize. Plant Biotechnol. J. 11, 1080–1091 (2013).
Nuccio, M. L. et al. Expression of trehalose-6-phosphate Phosphatase in maize ears improves yield in well-watered and drought conditions. Nat. Biotechnol. 8, 862–869 (2015).
Bennett, E., Roberts, J. A. & Wagstaff, C. Manipulating resource allocation in plants. J. Exp. Bot. 63, 3391–3400 (2012).
Reynolds, M., et al. Raising yield potential of wheat. I. Overview of a consortium approach and breeding strategies. J. Exp. Bot. 62, 439–452 (2011).
Lawlor, D. W. & Paul, M. J. Source/sink interactions underpin crop yield: the case for trehalose 6-phosphate/SnRK1 in improvement of wheat. Front. Plant Sci. 5, 418 (2014).
Lu, M. Z., Snyder, R., Grant, J. & Tegeder, M. Manipulation of sucrose phloem and embryo loading affects pea leaf metabolism, carbon and nitrogen partitioning to sinks as well as seed storage pools. Plant J. 101, 217–236 (2020).
Brauer, E.K., Rochon, A., Bi, Y.M., Bozzo, G.G., Rothstein, S.J. & Shelp, B.J. Reappraisal of nitrogen use efficiency in rice overexpressing glutamine synthetase1. Physio.l Plantarum. 141, 361–372 (2011).
Urriola, J. & Rathore, K. S. Overexpression of a glutamine synthetase gene affects growth and development in sorghum. Transgenic Res. 24, 397–407 (2015).
Bock, R. Strategies for metabolic pathway engineering with multiple transgenes. Plant Mol. Biol. 83, 21–31 (2013).
Fuentes, P. et al. A new synthetic biology approach allows transfer of an entire metabolic pathway from a medicinal plant to a biomass crop. Elife. 5, e13664 (2016).
Naqvi, S. et al. Transgenic multivitamin corn through biofortification of endosperm with three vitamins representing three distinct metabolic pathways. Proc. Natl. Acad. Sci USA 106, 7762–7767 (2009).
Fitzpatrick, T. B. et al. Vitamin Deficiencies in Humans: Can Plant Science Help?. Plant Cell. 24, 395–414 (2012).
Lê Cao, K.-A., Martin, P.G.P., Robert-Granié, C. & Besse, P. Sparse canonical methods for biological data integration: application to a cross-platform study. BMC Bioinformatics. 10, 34 (2009).
Zhang, W. F. et al. Closing yield gaps in China by empowering smallholder farmers. Nature 537, 671–674 (2016).
Van Roekel, R. J. & Coulter, J. A. Agronomic Responses of Corn to Planting Date and Plant Density. Agron J. 103, 1414–1422 (2011).
Antonietta, M., Fanello, D. D., Acciaresi, H. A. & Guiamet, J. J. Senescence and yield responses to plant density in stay green and earlier-senescing maize hybrids from Argentina. Field Crop Res. 155, 111–119 (2014).
Micallef, B. J. et al. Altered photosynthesis, flowering, and fruiting in transgenic tomato plants that have an increased capacity for sucrose synthesis. Planta 196, 327–334 (1995).
D’Aoust, M. A., Yelle, S. & Nguyen-Quoc, B. Antisense inhibition of tomato fruit sucrose synthase decreases fruit setting and the sucrose unloading capacity of young fruit. Plant Cell. 11, 2407–2418 (1999).
Dai, N. et al. Overexpression of Arabidopsis hexokinase in tomato plants inhibits growth, reduces photosynthesis, and induces rapid senescence. Plant Cell. 11, 1253–1266 (1999).
Menu, T. et al. High hexokinase activity in tomato fruit perturbs carbon and energy metabolism and reduces fruit and seed size. Plant Cell Environ. 27, 89–98 (2004).
Ruan, Y. L. et al. Expression of sucrose synthase in the developing endosperm is essential for early seed development in cotton. Funct. Plant Biol. 35, 382–393 (2008).
Adams, W. W., Muller, O., Cohu, C. M. & Demmig-Adams, B. May photoinhibition be a consequence, rather than a cause, of limited plant productivity?. Photosynth Res. 117, 31–44 (2013).
Kaachra, A., Vats, S. K. & Kumar, S. Heterologous expression of key C and N metabolic enzymes improves re-assimilation of photorespired CO2 and NH3, and growth. Plant Physiol. 177, 1396–1409 (2018).
Kohler, I. H. et al. Expression of cyanobacterial FBP/SBPase in soybean prevents yield depression under future climate conditions. J. Exp. Bot. 68, 715–726 (2017).
Bermudez, L. et al. A candidate gene survey of quantitative trait loci affecting chemical composition in tomato fruit. J. Exp. Bot. 59, 2875–2890 (2008).
Bermudez, L. et al. Silencing of the tomato Sugar Partitioning Affecting protein (SPA) modifies sink strength through a shift in leaf sugar metabolism. Plant J. 77, 676–687 (2014).
Okumoto, S. et al. High affinity amino acid transporters specifically expressed in xylem parenchyma and developing seeds of Arabidopsis. J. Biol. Chem. 277, 45338–45346 (2002).
Hunt, E. et al. A mutation in amino acid permease AAP6 reduces the amino acid content of the Arabidopsis sieve elements but leaves aphid herbivores unaffected. J. Exp. Bot. 61, 55–64 (2010).
Riesmeier, J. W., Willmitzer, L. & Frommer, W. B. Evidence for an essential role of the sucrose transporter in phloem loading and assimilate partitioning. EMBO J. 13, 1–7 (1994).
Jelitto, T., Sonnewald, U., Willmitzer, L., Hajirezeai, M. & Stitt, M. Inorganic pyrophosphate content and metabolites in potato and tobacco plants expressing E. coli pyrophosphatase in their cytosol. Planta. 188(2), 238–244 (1992).
Sonnewald, U. Expression of E. coli inorganic pyrophosphatase in transgenic plants alters photoassimilate partitioning in leaves of transgenic plants. Plant J. 2(4), 571–581 (1992).
Jin, Y., Ni, D. A. & Ruan, Y. L. Posttranslational elevation of cell wall invertase activity by silencing its inhibitor in tomato delays leaf senescence and increases seed weight and fruit hexose level. Plant Cell. 21, 2072–2089 (2009).
Nunes-Nesi, A. et al. Enhanced photosynthetic performance and growth as a consequence of decreasing mitochondrial malate dehydrogenase activity in transgenic tomato plants. Plant Physiol. 137, 611–622 (2005).
Centeno, D. C. et al. Malate plays a crucial role in starch metabolism, ripening, and soluble solid content of tomato fruit and affects postharvest softening. Plant Cell. 23, 162–184 (2011).
Hasse, D., Mikkat, S., Hagemann, M. & Bauwe, H. Alternative splicing produces an H-protein with better substrate properties for the P-protein of glycine decarboxylase. FEBS J. 276, 6985–6991 (2009).
Bauwe, H. & Kolukisaoglu, U. Genetic manipulation of glycine decarboxylation. J. Exp. Bot. 54, 1523–1535 (2003).
Timm, S. et al. Glycine decarboxylase controls photosynthesis and plant growth. FEBS Lett. 586, 3692–3697 (2012).
Fridman, E., Carrari, F., Liu, Y. S., Fernie, A. R. & Zamir, D. Zooming in on a quantitative trait for tomato yield using interspecific introgressions. Science 305, 1786–1789 (2004).
Zanor, M. I. et al. RNA interference of LIN5 in tomato confirms its role in controlling brix content, uncovers the influence of sugars on the levels of fruit hormones, and demonstrates the importance of sucrose cleavage for normal fruit development and fertility. Plant Physiol. 150, 1204–1218 (2009).
Bouche, N. & Fromm, H. GABA in plants: just a metabolite?. Trends Plant Sci. 9, 110–115 (2004).
Soyk, S. et al. Duplication of a domestication locus neutralized a cryptic variant that caused a breeding barrier in tomato. Nat. Plants. 5, 471–479 (2019).
Kwon, C.-T. et al. Rapid customization of Solanaceae fruit crops for urban agriculture. Nat. Biotechnol. 38, 182–188 (2020).
Murashige, T. & Skoog, F. A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol. Plant. 15, 473–497 (1962).
Devkar, V. et al. Multifaceted regulatory function of tomato SlTAF1 in the response to salinity stress. New Phytol. 225(4), 1681–1898 (2019).
Elghabu, Z., Ruf, S. & Bock, R. Biolistic co-transformation of the nuclear and plastid genomes. Plant J. 67(5), 941–948 (2011).
Doyle, J. & Doyle, J. A rapid DNA isolation procedure for small quantities of leaf tissue. Focus. 12, 13–15 (1990).
Pfaffl, M. W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 29(9), e45 (2001).
Cheng, Y. et al. Genome-Wide identification and evaluation of reference genes for quantitative RT-PCR analysis during tomato fruit development. Front. Plant Sci. 8, 1440 (2017).
Osorio, S. et al. Integrative comparative analyses of transcript and metabolite profiles from pepper and tomato ripening and development stages uncovers species-specific patterns of network regulatory behavior. Plant Physiol 159, 1713–1729 (2012).
Luedemann, A., Strassburg, K., Erban, A. & Kopka, J. TagFinder for the quantitative analysis of gas chromatography–mass spectrometry (GC–MS)-based metabolite profiling experiments. Bioinformatics 24, 732–737 (2008).
the Golm Metabolome Database. Kopka, J., et al. GMD@CSB.DB. Bioinformatics 21, 1635–1638 (2005).
Fernie, A. R. et al. Recommendations for reporting metabolite data. Plant Cell. 23, 2477–2482 (2011).
Krall, J. P. & Edwards, G. E. Relationship between Photosystem-Ii activity and CO2 fixation in leaves. Physiol. Plantarum. 86, 180–187 (1992).
Valentini, R., Epron, D., Deangelis, P., Matteucci, G. & Dreyer, E. In-Situ Estimation of net CO2 assimilation, photosynthetic electron flow and photorespiration in turkey oak (Q-Cerris L) leaves: diurnal cycles under different levels of water-supply. Plant Cell Environ. 18, 631–640 (1995).
Rohart, F., Gautier, B., Singh, A. & Le Cao, K. A. mixOmics: An R package for ‘omics feature selection and multiple data integration. PLoS Comput. Biol. 13, e1005752 (2017).
Rosenthal, D. M. et al. Over-expressing the C-3 photosynthesis cycle enzyme Sedoheptulose-1-7 Bisphosphatase improves photosynthetic carbon gain and yield under fully open air CO2 fumigation (FACE). BMC Plant Biol. 11, 123 (2011).
Ding, F., Wang, M., Zhang, S. & Ai, X. Changes in SBPase activity influence photosynthetic capacity, growth, and tolerance to chilling stress in transgenic tomato plants. Sci. Rep. 6, 32741 (2016).
Migge, A., Carrayol, E., Hirel, B. & Becker, T. W. Leaf-specific overexpression of plastidic glutamine synthetase stimulates the growth of transgenic tobacco seedlings. Planta 210, 252–260 (2000).
Chen, L. Q. et al. Sucrose efflux mediated by SWEET proteins as a key step for phloem transport. Science 335, 207–211 (2012).
Sauer, N. & Stolz, J. Suc1 and Suc2 - 2 Sucrose Transporters from Arabidopsis-thaliana - expression and characterization in bakers-yeast and identification of the histidine-tagged protein. Plant J. 6, 67–77 (1994).
Dasgupta, K. et al. Expression of sucrose transporter complementary DNAs specifically in companion cells enhances phloem loading and long-distance transport of sucrose but leads to an inhibition of growth and the perception of a phosphate limitation. Plant Physiol. 165, 715–731 (2014).
Fischer, W. N. et al. Low and high affinity amino acid H+-cotransporters for cellular import of neutral and charged amino acids. Plant J. 29, 717–731 (2002).
Yang, H. Y., Stierhof, Y. D. & Ludewig, U. The putative Cationic Amino Acid Transporter 9 is targeted to vesicles and may be involved in plant amino acid homeostasis. Front Plant Sci. 6, 212 (2015).
Xu, S.M., Brill, E., Llewellyn, D.J., Furbank, R.T. & Ruan, Y.L. Overexpression of a potato sucrose synthase gene in cotton accelerates leaf expansion, reduces seed abortion, and enhances fiber production. Mol. Plant. 5, 430–441 (2012).
Petreikov, M. et al. Temporally extended gene expression of the ADP-Glc pyrophosphorylase large subunit (AgpL1) leads to increased enzyme activity in developing tomato fruit. Planta 224, 1465–1479 (2006).
Wingenter, K. et al. Increased activity of the vacuolar monosaccharide transporter TMT1 alters cellular sugar partitioning, sugar signaling, and seed yield in Arabidopsis. Plant Physiol. 154, 665–677 (2010).
Scholz-Starke, J., Büttner, M. & Sauer, N. AtSTP6, a new pollen-specific H+-Monosaccharide symporter from Arabidopsis. Plant Physiol. 131, 70–77 (2003).
Büttner, M. et al. AtSTP3, a green leaf-specific, low affinity monosaccharide-H+ symporter of Arabidopsis thaliana. Plant Cell Environ 23, 175–184 (2000).
Nguyen-Quoc, B. & Foyer, C. H. A role for “futile cycles” involving invertase and sucrose synthase in sucrose metabolism of tomato fruit. J. Exp. Bot. 52, 881–889 (2001).
Nguyen-Quoc, B., N’Tchobo, H., Foyer, C. H. & Yelle, S. Overexpression of sucrose phosphate synthase increases sucrose unloading in transformed tomato fruit. J. Exp. Bot. 50, 785–791 (1999).
Ho, L. C. Partitioning of assimilates in fruiting tomato plants. Plant Growth Regul. 2, 277–285 (1984).
Lytovchenko, A. et al. Tomato fruit photosynthesis is seemingly unimportant in primary metabolism and ripening but plays a considerable role in seed development. Plant Physiol. 157, 1650–1663 (2011).
McCollum, J. P. & Skok, J. Radiocarbon studies on the translocation of organic constituents into ripening tomato fruits. Proc. Am. Soc. Hortic. Sci. 75, 611–616 (1960).
Balibrea, M. E., Martinez-Andujar, C., Cuartero, J., Bolarin, M. C. & Perez-Alfocea, F. The high fruit soluble sugar content in wild Lycopersicon species and their hybrids with cultivars depends on sucrose import during ripening rather than on sucrose metabolism. Funct. Plant Biol. 33, 279–288 (2006).
Yativ, M., Harary, I. & Wolf, S. Sucrose accumulation in watermelon fruits: genetic variation and biochemical analysis. J. Plant Physiol. 167, 589–596 (2010).
Taji, T. et al. Important roles of drought- and cold-inducible genes for galactinol synthase in stress tolerance in Arabidopsis thaliana. Plant J. 29, 417–426 (2002).
Van den Ende, W. Multifunctional fructans and raffinose family oligosaccharides. Front. Plant Sci. 4, 247 (2013).
Hochberg, U., Batushansky, A., Degu, A., Rachmilevitch, S. & Fait, A. Metabolic and physiological responses of Shiraz and Cabernet Sauvignon (Vitis vinifera L.) to near optimal temperatures of 25 and 35 degrees C. Int. J. Mol. Sci. 16, 24276–24294 (2015).
ElSayed, A. I., Rafudeen, M. S. & Golldack, D. Physiological aspects of raffinose family oligosaccharides in plants: protection against abiotic stress. Plant Biol. 16, 1–8 (2014).
Kaur, H. et al. Ectopic expression of the ABA-inducible dehydration-responsive chickpea l-myo-inositol 1-phosphate synthase 2 (CaMIPS2) in Arabidopsis enhances tolerance to salinity and dehydration stress. Planta 237, 321–335 (2013).
Carrari, F. et al. Integrated analysis of metabolite and transcript levels reveals the metabolic shifts that underlie tomato fruit development and highlight regulatory aspects of metabolic network behavior. Plant Physiol. 142, 1380–1396 (2006).
Sauter, M., Moffatt, B., Saechao, M. C., Hell, R. & Wirtz, M. Methionine salvage and S-adenosylmethionine: essential links between sulfur, ethylene and polyamine biosynthesis. Biochem J. 451, 145–154 (2013).
Sicher, R. C. & Barnaby, J. Y. Impact of carbon dioxide enrichment on the responses of maize leaf transcripts and metabolites to water stress. Physiol. Plant. 144, 238–253 (2012).
Barnaby, J. Y. et al. Drought responses of foliar metabolites in three maize hybrids differing in water stress tolerance. PLoS ONE 8(10), e77145 (2013).
Hare, P. D. & Cress, W. A. Metabolic implications of stress-induced proline accumulation in plants. Plant Growth Regul. 21, 79–102 (1997).
Ishizaki, K. et al. The critical role of Arabidopsis electron-transfer flavoprotein: Ubiquinone oxidoreductase during dark-induced starvation. Plant Cell. 17, 2587–2600 (2005).
Matas, A. J. et al. Tissue- and cell-type specific transcriptome profiling of expanding tomato fruit provides insights into metabolic and regulatory specialization and cuticle formation. Plant Cell. 23, 3893–3910 (2011).
Shinozaki, Y. et al. High-resolution spatiotemporal transcriptome mapping of tomato fruit development and ripening. Nat. Commun. 9, 364 (2018).
Tohge, T. & Fernie, A. R. Leveraging natural variance towards enhanced understanding of phytochemical sunscreens. Trends Plant Sci. 22, 308–315 (2017).
Acknowledgements
We would like to thank Dr. Jacopo Martinis for providing vector pUC18-SlRcbS-MCS-NosT. Also, we would like to thank Dr. John M. Ward for providing plasmids with SUC2 and SUC9 sequences. This work was supported with an ERA-CAPS—European Commission’s 7th framework programme grant and by the Max Planck Society (Germany). We thank the MPI-MP Green Team for plant cultivation.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
J.V., S.K.-J., R.B., L.J.S. and A.R.F. conceptualized the study. S.R., M.R., S.T., H.B., F.C. and D.R. carried out cloning and transformation. J.V., L.J.S. and A.R.F. wrote the manuscript which was proofread and approved of all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Vallarino, J.G., Kubiszewski-Jakubiak, S., Ruf, S. et al. Multi-gene metabolic engineering of tomato plants results in increased fruit yield up to 23%. Sci Rep 10, 17219 (2020). https://doi.org/10.1038/s41598-020-73709-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-73709-6
This article is cited by
-
Genetic architecture of fresh-market tomato yield
BMC Plant Biology (2023)
-
Development of a facile genetic transformation system for the Spanish elite rice paella genotype Bomba
Transgenic Research (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.