Abstract
Flooding periods, as one probable consequence of climate change, will lead more frequently to plant hypoxic stress. Hypoxia sensing and signaling in the root, as the first organ encountering low oxygen, is therefore crucial for plant survival under flooding. Nitric oxide has been shown to be one of the main players involved in hypoxia signaling through the regulation of ERFVII transcription factors stability. Using SNP as NO donor, we investigated the NO-responsive genes, which showed a significant response to hypoxia. We identified 395 genes being differentially regulated under both hypoxia and SNP-treatment. Among them, 251 genes showed up- or down-regulation under both conditions which were used for further biological analysis. Functional classification of these genes showed that they belong to different biological categories such as primary carbon and nitrogen metabolism (e.g. glycolysis, fermentation, protein and amino acid metabolism), nutrient and metabolites transport, redox homeostasis, hormone metabolism, regulation of transcription as well as response to biotic and abiotic stresses. Our data shed light on the NO-mediated gene expression modulation under hypoxia and provides potential targets playing a role in hypoxia tolerance. These genes are interesting candidates for further investigating their role in hypoxia signaling and survival.
Similar content being viewed by others
Introduction
Nitric oxide (NO) is an important signaling molecule involved in a wide range of physiological processes during plant development. NO has been reported to play a role in photomorphogenesis and flowering1, development of lateral root2, organ growth3 and senescence4. Moreover, NO has been shown to be involved in response to different abiotic and biotic stresses such as heat5, drought6,7,8, salinity9,10, ozon11, heavy metal12,13, pathogen attack14 as well as flooding and hypoxia15,16,17.
NO is a highly reactive molecule and therefore its conversion to a non-toxic molecule is crucial to avoid cell toxicity18. NO level in plant cell is mainly regulated via S-nitrosoglutathione reductase (GSNOR) activity19. Interestingly, S-nitrosylation of GSNOR1 itself at Cys10, resulted in conformational changes and direct interaction with ATG8 leading to the autophagy of GSNOR1 in Arabidopsis20,21. Furthermore, phytoglobins (PGBs) have been reported to be involved in NO scavenging and its conversion to nitrate22,23.
S-Nitrosoglutathione (GSNO) is one of the important NO-bioactive molecules inside the cell involved in NO-mediated post-translational modifications (PTMs) such as cysteine S-nitrosylation, metal nitrosylation and tyrosine nitration of target proteins18,24,25. Moreover, NO can lead to fatty acid nitration and nitro fatty acid production, which might further contribute to plant NO signaling26,27,28.
NO can be synthesized via oxidative or reductive pathways, either enzymatic or nonenzymatic. NO synthase (NOS) is the enzyme involved in NO biosynthesis in mammals29. Although in higher plants, NO synthase inhibitors decrease the production of NO and citrulline from L-arginine, the NOS homologe in the plant cells has not yet been identified, except in Ostreococcustauri, a single cell alga24,30. In plants, Xanthine oxidoreductase (XOR), a key enzyme involved in purine catabolism, has been shown to convert nitrite to NO in vitro in the presence of reducing substrates NADH or xanthine in an oxygen dependent manner31,32,33. The cytosolic nitrate reductase (NR), the key enzyme involved in nitrate assimilation which converts nitrate to nitrite, may also catalyze the conversion of nitrite to NO34. Cytochrome c oxidase, located in the inner membrane of mitochondria, is involved in NO synthesis from nitrite35,36. Loss-of-function mutant of Arabidopsis NOA1 (noa1) showed reduced level of NO37. NOA1 is a plastid targeted GTPase, which is associated with ribosome function. Therefore, NO-deficient phenotype of noa1 mutant is possibly an indirect effect due to the hindered chloroplast function37. Oxygen dependent enzymatic extracellular NO production has been suggested to be important for sensing the availability of nitrate. In this direction, a Nitrite-NO reductase (NiNOR) activity has been discovered in root plasma membrane (PM), which uses the nitrite provided by PM bound NR as substrate under low oxygen condition16,38. Recently, a new enzymatic NO producing route has been reported in Chlamydomonas reinhardtii which produces NO via activity of two molybdenum cofactor enzymes, NOFNiR (nitric oxide-forming nitrite reductase) and NR39,40, however, its role in NO production in higher plants is still not clear.
Conversion of nitrite to NO can also occur non-enzymatically under low pH in apoplast and plastids37,41,42. This complexity of plant NO production in different parts of the cell via various enzymatic and non-enzymatic reactions, complicates investigating the NO effect using mutant lines. This is even more problematic in crops compared to Arabidopsis due to their genome complexity. Hence, NO donors are used as an alternative tool for NO studies. Among different NO donors, sodium nitroprusside (SNP) is widely used to study the effect of NO. Beside its lower cost compared to other NO donors, SNP releases NO continuously and for longer period of the time, which might be of advantage for biological studies43,44,45.
The effect of NO during flooding stress has been intensively addressed in a recent review46. Former studies have shown that NO-mediated S-nitrosylation of several proteins such as ERFVIIs, phytoglobins, cytochrome c oxidase (COX), aconitase, and ascorbate peroxidase (APX1) might be associated with flooding signaling and tolerance46,47,48. Moreover, it has been reported that under hypoxia, NO plays a pivotal role by regulation of COX and alternative oxidase (AOX) activity and therefore, mitochondrial oxygen consumption49.
Endogenous NO level plays a role in regulation of transcription under hypoxia through the N-degron pathway48,50. It has been shown that degradation of MC-ERFVII transcription factors (TFs) occurs under NO availability via the N-degron pathway. Adversely, ERFVII TFs are stabilized in the absence of NO48.
Tomato is an economically important crop plant which is sensitive to flooding induced hypoxia51,52. However, our understanding of the precise mechanism of hypoxia tolerance and in particular the role of NO during this process is not complete. Moreover, our knowledge about NO effect on gene expression regulation in tomato during hypoxia is still scarce. Therefore, identification of NO-responsive genes under hypoxia is essential for improving flooding stress tolerance in tomato.
In our former study, we identified short-(6 h) and long-term (48 h) hypoxia-responsive genes in tomato (cv. Moneymaker) roots53. Those data suggest a distinct temporal transcriptional response. While short term hypoxia resulted in transcriptional acclimation, hypoxia progression resulted in a transcriptional reprogramming to support an escape mechanism probably through aerenchyma and adventitious root formation53. This indicates the ability of a cultivated crop such as tomato to temporally adjust its response mechanism to hypoxia, both metabolically and anatomically53.In the current study, using RNA-seq approach, tomato root gene regulation changes after long-term (48 h) application of NO donor SNP was studied. In order to find NO-responsive genes under hypoxia, common differentially regulated genes with similar regulation changes (being up- or down-regulated) under both hypoxia and SNP-treatment were identified. This study provides the potential targets of hypoxia-induced NO. The identified genes can serve as candidates for investigating their role in NO-mediated hypoxia tolerance in tomato as well as other crops.
Result
SNP-treatment showed distinct response in physiological parameters as well as in modulation of gene expression
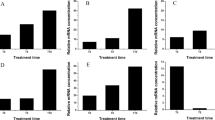
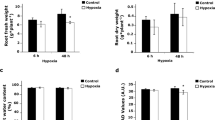
In contrast to SNP-treatment, only the 48 h hypoxia resulted in a significantly lower root fresh weight (Fig. 1a). Moreover, neither hypoxia nor SNP-treatment resulted in statistically significant differences in root dry weight and root water content (%) (Fig. 1b,c). Relative chlorophyll level (SPAD values) in leaf #3showed a significant (P < 0.05) reduction in response to 48 h hypoxia, but not to SNP-treatment (Fig. 1d).
Fresh weight, dry weight and relative chlorophyll content of tomato plants in response to hypoxia and SNP-treatment. (a) Fresh weight; (b) dry weight and (c) water content (%) of 5 weeks old tomato roots after 48 h hypoxia and 48 h SNP-treatment were compared to their representative control. (d) Relative chlorophyll contents in leaf #3 of plants under hypoxia and SNP-treatment in comparison to their respective control are shown as SPAD values. Data represent means ± SD; n = 3; *significant differences (Student’s t test, *P < 0.05).
High throughput sequencing of root RNA samples
High throughput sequencing results of 6 samples, including SNP-treated and untreated control are summarized in Table 1. Preprocessing steps, adapter clipping and low base quality filtering, resulted in about 98 and 107 million high quality reads in total (ca. 33 and 36 million reads per sample) for SNP-treated and untreated control, respectively. In total, ca. 90 and 100 million reads (91% and 93% for each sample) for untreated control and SNP-treated samples were mapped on to tomato reference genome (ITAG2.4), using Genomics workbench V7.5.5.
Differentially expressed genes (DEGs) in response to hypoxia and SNP-treatment (functional classification of NO and hypoxia-responsive genes)
The number of 1144 genes were differentially regulated (Padj < 0.05) in response to 48 h SNP-treatment (792 down- and 352 up-regulated genes). After comparison with 1421 differentially regulated genes (897 up- and 524 down-regulated) under 48 h hypoxia (Padj < 0.05)53, it was observed that 395 DEGs, were concertedly regulated under both, hypoxia and SNP-treatment (Fig. 2). Among above-mentioned genes, 144 genes showed the opposite- while 251 genes showed similar regulation changes (154 up- and 97 down-regulated) (Fig. 2). For further analysis, only those 251 genes with similar regulation changes under NO and hypoxia were chosen for downstream biological pathway analysis. The list of all common differentially regulated genes between hypoxia and SNP-treatment is provided in Supplementary Table S1.
Venn diagram of common differentially expressed genes (DEGs) (Padj < 0.05) under both hypoxia and SNP-treatment. Red and green arrows represent down- and up-regulated genes, respectively. Boxes containing double green and double red arrows indicates number of down (154)- and up-regulated (97) DEGs under both hypoxia and SNP-treatments. The box containing both red and green arrows indicates the number of DEGs (144) with opposite regulation in response to hypoxia and SNP-treatment.
Gene Ontology (GO) analysis
To visualize enriched regulated GO terms, all significantly enriched (Padj < 0.05) Gene Ontology (GO) annotations, according to cellular component, biological processes and molecular function are presented (Fig. 3). It must be noted that GO terms refer to the proteins encoded by the genes and therefore in some cases, the word activity is used in GO term results.
GO terms associated with transcriptome modulation of tomato roots in response to hypoxia and SNP-treatment. Enriched GO terms (Padj < 0.05), describing molecular function, biological process and cellular compartment. The regulated genes in all samples were analyzed for enriched GO terms using online tool PANTHER 14.0 and Solanum lycopersicum as a reference organism. The bars represent all significantly enriched GO terms associated with regulated genes in response to hypoxia and SNP-treatment.
Functional annotation of the regulated genes showed that their encoded proteins are mainly involved in catalytic activity (GO:0003824) (> 61%), transporter activity (GO:0005215) (17%) and binding (GO:0005488) (14%). Biological processes such as metabolic process (GO:0008152) (40%), response to stimulus (GO:0050896) (22%) and cellular process (GO:0009987) (22%) showed the highest percentage of regulated genes. The cellular component categories with the highest percentage of regulated genes were cell (GO:0005623) (67%), organelle (GO:0043226) (14%) and cell junction (GO:0030054) (13%) (Fig. 3). For a more detailed analysis of the biological pathways, we used a plant based database, MapMan54. MapMan categories54 are based on ITAG2.3 annotations (Supplementary Table S1).
Validation of differentially expressed genes in response to SNP-treatment using qPCR
To validate RNA-Seq data, qPCR was performed on 17 regulated genes in response to SNP-treatment (Supplementary Table S2). We observed similar gene regulation changes (SNP-treated/control) between RNA-Seq and qPCR (Fig. 4). These data confirm the validity of the RNA-Seq results used in this study. The confirmation of RNA-Seq data under hypoxia is provided elsewhere53.
Hypoxia and SNP-associated phytohormone related genes
It was observed that 23 hypoxia-induced phytohormone related genes responded to SNP-treatment. These genes were related to different phytohormone categories such as abscisic acid (Solyc04g008960), auxin (ATB2, Solyc03g006490, IAA14, AILP1 and PIN2), brassinosteroid (Solyc11g006270), ethylene (Solyc12g006380, DLO1, two transcripts annotated as DLO2, DMR6, Solyc06g066830, Solyc03g116260, Solyc09g089680, Solyc06g073580 as well as two nitrilase encoded transcripts MES3 and Solyc09g011140), gibberellin (GASA5, two transcripts annotated as GASA6 and Solyc06g067860), and jasmonate (two transcripts annotated as LOX1) (Fig. 5).
Transcriptional changes of genes related to different phytohormone categories. The heat map displays the up-regulated (red bars) or down-regulated (green bars) tomato genes and their Arabidopsis thaliana homologs in response to hypoxia and SNP-treatment. Depicted are differentially expressed genes (Padj < 0.05) (n = 3). ABA, abscisic acid; AUX, Auxin; BR, brassinosteroid; ETH, ethylene; GA: gibberellin; JA, jasmonate; 2OG, 2-oxoglutarate; ATB2, NAD(P)-LINKED OXIDUREDUCTASE-LIKE PROTEIN; IAA14, INDOLE-3-ACETIC ACID INDUCIBLE 14; AILP1, ALUMINIUM INDUCED PROTEIN WITH YGL AND LRDR MOTIFS; PIN2, PIN-FORMED 2; DLO1, DMR6-LIKE OXYGENASE 1; DLO2, DMR6-LIKE OXYGENASE 2; DMR6, DOWNY MILDEW RESISTANT 6; GASA5, GAST1 PROTEIN HOMOLOG 5; GASA6, GAST1 PROTEIN HOMOLOG 5; LOX1, LIPOXYGENASE 1.
Hypoxia and SNP-responsive genes encoding proteins involved in PTM and regulation of transcription (TFs)
Two genes encoding proteins involved in PTM (CIPK11 and Solyc02g086360) showed regulation changes in response to hypoxia and SNP-treatment. Moreover, 16 genes encoding members of different transcription factor (TF) families were observed, as follow: NAC family (two transcripts annotated as ATAF2/NAC081 and one transcript as NAC083), AP2/EREBP (RAP2.2), Aux/IAA family (IAA14), bHLH (Solyc09g098110), C2C2 (Solyco9g074560, JAZ9 and CDF1), C2H2 zinc finger family(REIL1), G2-like (Solyc05g009720), MYB (MYB62), MYB-like (Solyc11g006720), WRKY (WRKY7), one transcript annotated as unclassified (Solyc12g087940) and two transcripts encoding a putative transcription factor (NFXL1) (Fig. 6).
Hypoxia and SNP-responsive genes encoding proteins involved in post-translational modification (PTM) and regulation of transcription (TFs). Heat map represents the up-regulated (red bars) or down-regulated (green bars) tomato genes and their Arabidopsis thaliana homologs in response to hypoxia and SNP-treatment. Depicted are differentially expressed genes (Padj < 0.05), (n = 3). CIPK11, CBL-INTERACTING PROTEIN KINASE 11; ATAF2 (NAC081), ARABIDOPSIS NAC DOMAIN CONTAINING PROTEIN 81; NOC083, ARABIDOPSIS NAC DOMAIN CONTAINING PROTEIN 83; RAP2.2, RELATED TO AP2 2; JAZ9, RELATED TO AP2 2; CDF1, CYCLING DOF FACTOR 1; REIL1, REI1-LIKE 1; MYB62, MYB DOMAIN PROTEIN 62; NFXL1, NF-X-LIKE 1; WRKY7, WRKY DNA-BINDING PROTEIN 7.
Common DEGs involved in primary metabolism
Among the responsive genes to both hypoxia and NO, were the genes encoding enzymes belonging to the primary metabolism such as glycolysis (ENO2 and Solyc04g008740), fermentation(ADH1, Solyc02g077240 and solyc10g076510), carbon metabolism (carbonic anhydrase: CA1 and CA2), starch degradation (Solyc06g073190, Solyc08g079080 and SUS4), photosynthesis: Calvin cycle (FBA4), photorespiration (GOX1) and electron transport chain (Solyc04g074740 and ENODL16), amino acid synthesis (MTO3, ASP3, Solyc02g087740), protein synthesis (BT1) and protein degradation (KRS-1 and UBP17) (Fig. 7).
Hypoxia- and SNP-induced and repressed genes associated with primary metabolism. Heat maps display the up-regulated (red bars) or down-regulated (green bars) tomato genes and their Arabidopsis thaliana homologs. Depicted are differentially expressed genes (Padj < 0.05) (n = 3). ENO2, ENOLASE2; ADH1, ALCOHOL DEHYDROGENASE 1; CA1, CARBONIC ANHYDRASE 1; CA2, CARBONIC ANHYDRASE 2; SUS4, SUCROSE SYNTHASE 4; FB4, AUXIN SIGNALING F-BOX 4; GOX1, GLYCOLATE OXIDASE 1; ENODL16, EARLY NODULIN-LIKE PROTEIN 16; MTO3, METHIONINE OVER-ACCUMULATOR 3; ASP3, ASPARTATE AMINOTRANSFERASE 3; KRS.1, LYSYL-TRNA SYNTHETASE 1; BT1, BTB AND TAZ DOMAIN PROTEIN 1; UBP17, UBIQUITIN-SPECIFIC PROTEASE 17.
Redox associated genes responsive to hypoxia and SNP-treatment
Among regulated genes under hypoxia and SNP-treatment, 35 genes were observed encoding redox-associated proteins belonging to different categories such as catalases (CAT2), reductases (SDR5), peroxidases (PRX71, two transcripts annotated as PRX72, five transcripts annotated as PRX52, PRX2, two transcripts annotated as PRX9, PRX64, five transcripts annotated as RCI3, Solyc02g090450, Solyc02g090470, Solyc07g017880 and Solyc08g075830, oxidases (Solyc12g013690 and SKU5), glutathione S-transferases (GSTU1, three transcripts annotated as GSTU8, GSTU19 and GSTL3), cytochrome P450 (CYP716A1, CYP76C2, CYP72A14 and CYP707A3) (Fig. 8). These data indicate the importance of redox regulation under hypoxia-induced nitrosative stress.
Expression pattern of genes encoding members of different antioxidant classes. Heat map represents the up-regulated (red bars) or down-regulated (green bars) tomato genes and their Arabidopsis thaliana homologs in response to hypoxia and SNP-treatment. Depicted are differentially expressed genes (Padj < 0.05), (n = 3). GST, GLUTATHIONE S-TRANSFERASE; CAT2, CATALASE 2; SDR5, SHORT-CHAIN DEHYDROGENASE REDUCTASE 5; PRX71, PEROXIDASE 71; PRX72, PEROXIDASE 72; PRX52, PEROXIDASE 52; PRX2, PEROXIDASE 2; PRX9, PEROXIDASE 9; PRX64, PEROXIDASE 64; RCI3, RARE COLD INDUCIBLE GENE 3; SKU5; encodes SKU5 protein; GSTU1 (GST19), GLUTATHIONE S-TRANSFERASE TAU 1 (GLUTATHIONE S-TRANSFERASE 19); GSTU8, GLUTATHIONE S-TRANSFERASE TAU 8; GSTU19 (GST8), GLUTATHIONE S-TRANSFERASE TAU 19 (GLUTATHIONE S-TRANSFERASE 8); GSTL3, encoding a member of glutathione S-transferase family protein; CYP716A1, CYTOCHROME P450, FAMILY 716, SUBFAMILY A, POLYPEPTIDE 1; CYP76C2, CYTOCHROME P450, FAMILY 76, SUBFAMILY C, POLYPEPTIDE 2; CYP72A14, CYTOCHROME P450, FAMILY 72, SUBFAMILY A, POLYPEPTIDE 14; CYP707A3, CYTOCHROME P450, FAMILY 707, SUBFAMILY A, POLYPEPTIDE 3.
Concertedly regulated DEGs associated with cellular transport
Among common DEGs under hypoxia and SNP-treatment, 27 genes were observed encoding proteins related to transport (3 up- and 17 down-regulated). These genes encode proteins involved in the transport of sulfate (SULTR1;3), peptides and oligopeptides (NRT1.1) and nitrate (two transcripts encoding NRT2.4). Furthermore, a member of Sec14p-like phosphatidylinositol transfer family(solyc02g070210), an ATP-binding cassette(ABC) transporter (ALS1), mitochondria membrane metabolite transporter (SLAH1), a sugar transporter (Solyc01g080680) and major intrinsic proteins, aquaporins ( AQP), such as 12 plasma membrane intrinsic proteins (PIPs: PIP1;3, three transcripts annotated as PIP1;4 (three transcripts), PIP2;1 (two transcripts), PIP2;2, PIP2;5 (two transcripts) and PIP2;7 (two transcripts). Among the regulated genes were also 7 tonoplast intrinsic proteins (TIPs: TIP1;1 (two transcripts), TIP1;3, TIP2;1 (two transcripts), TIP2;2 and major facilitator superfamily (MSF: Solyc01080680) were identified (Fig. 9).
Expression pattern of genes related to cellular transport. Heat map represents the up-regulated (red bars) or down-regulated (green bars) tomato genes and their Arabidopsis thaliana homologs in response to hypoxia and SNP-treatment. Depicted are differentially expressed genes (Padj < 0.05), (n = 3). ABC, ATP-binding cassette transporter; PIP, major intrinsic proteins; TIP, tonoplast intrinsic protein; MT MET, metabolite transporters at the mitochondrial membrane, SEC14, Sec14p-like phosphatidylinositol transfer family; NRT, nitrate transporter; MFS, major facilitator superfamily; SULTR, sulfur transporter. ALS1, ALUMINUM SENSITIVE 1; PIP1;3, PLASMA MEMBRANE INTRINSIC PROTEIN 1;3; PIP1;4, PLASMA MEMBRANE INTRINSIC PROTEIN 1;4; PIP2;1, PLASMA MEMBRANE INTRINSIC PROTEIN 2;1; PIP2;2, PLASMA MEMBRANE INTRINSIC PROTEIN 2;2; PIP2;5, PLASMA MEMBRANE INTRINSIC PROTEIN 2;5; PIP2;7, PLASMA MEMBRANE INTRINSIC PROTEIN 2;7; TIP1;1, TONOPLAST INTRINSIC PROTEIN 1;1; TIP1;3, TONOPLAST INTRINSIC PROTEIN 1;3; TIP2;1, TONOPLAST INTRINSIC PROTEIN 2;1; TIP2;2, TONOPLAST INTRINSIC PROTEIN 2;2; SLAH1, SLAC1 HOMOLOGUE 1; NRT2.4, NITRATE TRANSPORTER 2.4; NRT1.1, NITRATE TRANSPORTER 1.1; SULTR1.3, SULFATE TRANSPORTER 1.3.
DEGs associated with biotic and abiotic stress
Out of 19 stress related genes, which were responsive to hypoxia and SNP-treatment, six were related to biotic stress, encoding PR genes (Solyc08g080650, Solyc04g015220, Solyc09g091210, Solyc06g075630, Solyc10g055190 and Solyc06g075630). 13 genes were identified being related to abiotic stresses such as drought/salt (two transcripts annotated as SRO5), heat (HSP23.6, Solyc06g068500 and ATJ3) and an unspecified class of stress (Solyc09g014530, Solyc09g005400, MLP34, Solyc04g007750, Solyc10g080190 and Solyc01g100370). The rest of the abiotic stress related genes (OSM34, Solyc04g015220, Solyc09g091210, Solyc06g075630, Solyc10g055190 and Solyc06g075630) were unspecified (Fig. 10).
Expression pattern of biotic and abiotic stress related genes. Heat map represents the up-regulated (red bars) or down-regulated (green bars) tomato genes and their Arabidopsis thaliana homologs in response to hypoxia and SNP-treatment. Depicted are differentially expressed genes (Padj < 0.05), (n = 3). SRO5, SIMILAR TO RCD ONE 5; HSP23.6, MITOCHONDRION-LOCALIZED SMALL HEAT SHOCK PROTEIN 23.6; ATJ3, DNAJ HOMOLOGUE 3; MLP34, MLP-LIKE PROTEIN 34; OSM34, OSMOTIN 34.
Cell wall related genes regulated in response to hypoxia and SNP-treatment
Several cell wall related genes showed regulation changes in response to hypoxia and SNP-treatment. These transcripts were related to cell wall synthesis (CSLD3, UGT85A2, UGT71B1, UGT73C1 and UGT73B3), Cell wall modification (XTH8, XTH5 and XTH24, EXPA6 and EXPA3 ) and cell wall degradation (RD22) (Fig. 11).
Expression pattern of regulated cell wall related genes. Heat map represents the up-regulated (red bars) or down-regulated (green bars) tomato genes and their Arabidopsis thaliana homologs in response to hypoxia and SNP-treatment. Depicted are differentially expressed genes (Padj < 0.05), (n = 3). UGT85A2, UDP-GLUCOSYL TRANSFERASE 85A2; UGT71B1, UDP-GLUCOSYL TRANSFERASE 71B1; UGT73C1, UDP-GLUCOSYL TRANSFERASE 73C1; UGT73B3, UDP-GLUCOSYL TRANSFERASE 73B3; CSLD3, CELLULOSE SYNTHASE LIKE D3; RD22, RESPONSIVE TO DESICCATION 22; EXPA6, EXPANSIN 6; XTH8, XYLOGLUCAN ENDOTRANSGLUCOSYLASE/HYDROLASE 8; XTH5, XYLOGLUCAN ENDOTRANSGLUCOSYLASE/HYDROLASE 5; XTH24, XYLOGLUCAN ENDOTRANSGLUCOSYLASE/HYDROLASE 24; EXPA3, EXPANSIN 3; MES3, METHYL ESTERASE 3.
Other biological pathways
Differentially regulated genes responsive to hypoxia and SNP-treatment encoding proteins belonging to additional functional categories were identified: lipid metabolism (LP1, ACBP6, FAD2 and Solyc10g083720) cell cycle (solyc01g111170, Solyc01g111170), cell organization (WVD2, PP2-A12, TUB1 and TUA6), cell vesicle transport (SNAP33), co-factor and vitamin metabolism (PDX1), development/storage proteins (PLP1 and solyc01g104110), DNA synthesis (Solyc01g080600), metal handling (MT2B and FP3), S-assimilation (two transcripts annotated as APR3, APR2 and SIR), secondary metabolism such as lignin (solyc04g054690 and LAC7) and phenol (three transcripts annotated as PAL1), Signaling (GRF2, IQD1 and Solyc07g006830), nucleotide metabolism (APY1), beta-galactosidase (BGAL1) and protease inhibitor (solyc03g079880) (Fig. 12).
Expression pattern of genes belonging to different biological categories. Heat map represents the up-regulated (red bars) or down-regulated (green bars) tomato genes and their Arabidopsis thaliana homologs in response to hypoxia and SNP-treatment. Depicted are differentially expressed genes (Padj < 0.05), (n = 3). WVD2, WAVE-DAMPENED 2; PP2-A12, PHLOEM PROTEIN 2-A12, TUB1, TUBULIN BETA-1 CHAIN; TUA6, TUBULIN ALPHA-6; SNAP33, SOLUBLE N-ETHYLMALEIMIDE-SENSITIVE FACTOR ADAPTOR PROTEIN 33; PDX1, PYRIDOXINE BIOSYNTHESIS 1; PLP1, PHOSPHOLIPASE 1; FAD2, FATTY ACID DESATURASE 2; ACBP6, ACYL-COA-BINDING PROTEIN; LP1, LIPID TRANSFER PROTEIN 1; MT2B, METALLOTHIONEIN 2B; FP3, FARNESYLATED PROTEIN 3; APR3, APS REDUCTASE 3; APR2, 5′ADENYLYLPHOSPHOSULFATE REDUCTASE 2; SIR, SULFITE REDUCTASE; PAL1, PHENYLALANINE AMMONIA-LYASE; LAC7, LACCASE 7; GRF2, GENERAL REGULATORY FACTOR 2; IQD13, IQ-DOMAIN 13; APY1, APYRASE 1; BGAL1, BETA GALACTOSIDASE 1.
Discussion
Tomato is a commercially important edible crop10. Improving tomato fruit size during the domestication process has been achieved by compensating its stress tolerance55. Former studies showed that SNP application on tomato root enhances salt stress tolerance, exhibited in improved growth and higher chlorophyll content. In the same study, SNP-treatment resulted in lower lipid oxidation, higher activity of antioxidant enzymes (SOD, APX, GR and POD) as well as an increase in ascorbate and proline content10. These data suggest that exogenous NO application via SNP-treatment is a reliable system for investigating the NO effect during stress response in the plant.
Several former studies investigated gene expression regulation in response to exogenous NO application using different NO donors in various plant species such as Arabidopsis11,18,56, cotton57 and birch58. However, our knowledge about the role of hypoxia-induced NO on gene expression modulation in tomato root is still scarce.
The current study represents the comparative transcriptome modulation of tomato (cv. Moneymaker) root in response to long-term (48 h) hypoxia53 and SNP-application. Long-term hypoxia, but not SNP-treatment, resulted in significantly (P < 0.05) lower root fresh weight and chlorophyll content (SPAD values) (Fig. 1a,d).
Under hypoxia as well as under SNP-treatment, 395 genes (28% of hypoxia-regulated genes), were concertedly regulated. Among them, 251 genes, corresponding to. 64% of common regulated genes, were similarly up- or down-regulated under both conditions (Fig. 2). It was also noticeable that the number of common down-regulated genes (154) in response to hypoxia and SNP-treatment was higher than up-regulated genes (97).
To validate the RNA-Seq. data, the expression changes of 17 differentially regulated genes were confirmed using qPCR. A high level of consistency between the qPCR and RNA-Seq expression data indicates the reliability of transcriptome data (Fig. 4).
Phytohormone associated genes showed regulation changes response to hypoxia and SNP-treatment
Genes belonging to different phytohormonal categories such as auxin, ethylene, jasmonate and gibberellin revealed regulation changes in response to NO application and hypoxia (Fig. 5). NO modulates auxin effect on root architecture. Transcriptional regulation of auxin related genes in response to exogenous NO application has already been reported59. Among auxin related DEGs in response to hypoxia and SNP-treatment in our study, polar auxin transporter PIN2 showed the highest up-regulation (12-fold) under hypoxia (Fig. 5). In Arabidopsis, ERFVII mediated repression of PIN2, has been shown to be associated with root bending under hypoxia in soil grown roots60. Since in this study tomato were cultivated in a hydroponic system, hypoxia might not cause the same response as in soil grown roots. These data suggest that polar auxin transport seems to be involved in root growth response under hypoxia. However, species-specific response, effects of growth condition as well as NO-dependent regulation of PIN2 expression remain unclear.
IAA14 showed down-regulation in response to hypoxia and SNP-treatment (Fig. 5). IAA14 is a negative regulator of auxin response factors ARF7 and ARF19, which are involved in lateral root initiation via induction of multiple LBD/ASLs such as LBD16/ASL18 and LBD29/ASL1661. It has been shown that overexpression of IAA14 leads to inhibition of lateral root formation in Arabidopsis thaliana62. Moreover, this indicates that IAA14 negatively regulates lateral root formation. This is in accordance with its downregulation in response to hypoxia, when adventive root formation is beneficial for low oxygen tolerance. Down-regulation of IAA14 in response to SNP in this study indicates that its regulation might be NO dependent. Recently, it has been shown that ERF VII TFs are involved in the regulation of lateral root formation through repression of auxin-induced genes (LBD16, LBD18, and PUCHI and IAA19)63. To what extent this process is associated with NO signaling, requires further investigation.
Regulation changes in ethylene related genes were observed in the current study (Fig. 5). Ethylene, one of the key regulators of hypoxia response, is involved in aerenchyma formation under flooding stress and has been shown to have cross talk with NO64. Recently, it has been reported that early ethylene-induced PHYTOGLOBIN1 (PGB1) acts as NO scavenger leading to the ERFVII stability and adaption to upcoming hypoxia65. Moreover, investigation of ein2-1 nos1/noa1 double mutant in A. thaliana revealed cross talk between ethylene, through EIN2, and NO signaling in the regulation of dark-induced leaf senescence66. These data indicate the importance of NO signaling and ethylene response during hypoxia.
Defense related phytohormones such as JA and SA have been shown to have an interplay with NO. In the current study, two transcripts being annotated as LOX1 (LIPOXYGENASE 1) showed regulation changes (Fig. 5). Mutation of LOX1 gene in Arabidopsis has been shown to modify both signaling and redox related response under cadmium stress67. Former studies in A. thaliana confirmed the expression induction of AOS and LOX2 in response to SNP application68. It has been shown that NO increased the expression of several JA biosynthetic genes among them LOX3, encoding the enzyme involved in the conversion of linolenic acid to 13(s) hydro peroxy octadecatrieonic acid69. Our result is in accordance with the above-mentioned studies and suggests a possible role for hypoxia-induced NO on regulation of some of the JA biosynthesis genes.
NO has been shown to play a role in regulating GA biosynthesis and signal transduction70. Both antagonistic and synergic interaction between NO and GA has been reported71. Five GA related genes showed down-regulation in our study (Fig. 3), among them, GASA5 has been reported to negatively regulate GA-induced flowering and stem growth72. However, its role during hypoxia in the root and its interaction with NO has not yet been investigated. Our results imply a cross talk between NO, hypoxia and different phytohormone related genes.
NO responsive TFs play role in diverse physiological processes and stress response
Group VII ERFs (ERFVII) has been shown to play a key role in hypoxia sensing and signaling via oxygen and NO dependent N-end rule pathway50,73,74. NO sensing has been shown to be executed via oxidation of Cysteine residue of ERFVII TFs, followed by arginylation and ubiquitination for proteasomal degradation (N-degron pathway)48,75. An ERFVII member, RAP2.2, was significantly up-regulated in response to hypoxia and SNP-treatment (Fig. 6). It has been reported that RAP2.2 overexpressing lines exhibited an improved hypoxia survival response while knockout lines showed weaker survival rate compared to the wild type. Moreover, RAP2.2 regulates the expression of hypoxia-responsive genes which their encoded enzymes are involved in sugar metabolism as well as fermentation pathways76. These data indicate the significant role of RAP2.2 in root response to hypoxia.
Moreover, among regulated genes were several transcripts (DLO1, DLO2 and DMR6) encoding proteins belonging to 2OG and Fe(II) dependent oxygenase superfamily. In a recent study in Arabidopsis, it was demonstrated that loss of function of PRT1, involved in N-degron pathways of ubiquitin-mediated proteolysis, improves the plant immune system. DMR6 and DLO1 proteins were accumulated in the former study indicating their importance in regulating the basal defense system77.
Several members of the NAC TF family such as ATAF2, NAC102 and NAC032 have been shown to be regulated by NO78,79. In accordance with the former studies, two transcripts (Solyc04g009440 and Solyc11g017470) annotated as ATAF2/NAC081, showed up-regulation in the current study in response to hypoxia and SNP-treatment (Fig. 6). NAC TFs are involved in plant development and response to different abiotic stresses80. The expression of ATAF2/NAC081 TF in maize has been shown to be positively regulated by ZmPTF1. ZmPTF1 is a member of the basic helix-loop-helix (bHLH) family involved in phosphate starvation and drought tolerance as well as root development in maize. ZmPTF1 binds to the G-box element within the promoter of several TFs such as ATAF2/NAC081 and NAC3081. Up-regulation of the above-mentioned genes has been reported to be involved in root development and stress response81. These data are in line with the up-regulation of ATAF2/NAC081 TF in our study indicating that it might be involved in root hypoxia response. However, whether NO is involved in ATAF2/NAC081 regulation requires further investigation.
NAC083, also known as VND-INTERACTING2 (VNI2), showed down-regulation in response to hypoxia and SNP-treatment in the current study. NAC083/VNI2 is an ABA responsive TF which has been shown to be involved in plant stress response. High salinity has been shown to increase the expression of NAC083/VNI2 in an ABA-dependent manner. Moreover, NAC083 negatively regulates stress-induced leaf senescence through regulation of COLD REGULATED (COR) and RESPONSIVE TO DEHYDRATION (RD) genes82,83. Two abiotic stress marker genes, COR15A/B and RD29A/B have been shown to be regulated by direct NAC083/VNI2 binding to their promoter. In the current study, only RD22 showed down-regulation which is in line with NAC083/VNI2 down-regulation83. Our data suggest that the down-regulation of NAC083/VNI2 under hypoxia might be NO-dependent. However, how hypoxia and NO results in repression of NAC083/VNI2 in the root, requires more investigations and might provide more insight into the role of NAC083/VNI2 in the regulation of hypoxia response.
WRKY7 showed up-regulation in the current study (Fig. 6). Up-regulation of WRKY encoding transcription factors, such as WRKY22, in response to submergence, has been reported in Arabidopsis, resulting in induction of immunity related marker genes84. wrky22 mutant showed the lower expression level of defense related genes after submergence84. These data indicate that there is a link between submergence induced hypoxia and defense in plants. However, the underlying signaling network, particularly the role of NO, is not yet unraveled.
REIL1, a member of the C2H2 zinc finger family, showed the highest up-regulation in our study in response to hypoxia (17.3-fold) and SNP-treatment (12.1-fold) (Fig. 6). REIL1 and REIL2 have been shown to be involved in A. thaliana leaf growth in the cold but not in normal temperature85. REIL1 provides an interesting candidate for further investigating its role in response to NO and hypoxia tolerance.
Genes related with ROS metabolism were regulated in response to hypoxia and SNP-treatment
Genes encoding different categories of ROS associated proteins such as peroxidases, oxidases, nitrilases, glutathione S-transferases as well as cytochrome P450 showed up- and down-regulation in the current study (Fig. 8).
Accumulation of ROS and reactive nitrogen species (RNS) is associated with low oxygen stress53,86,87. NO and its derivatives such as peroxynitrite (OONO-), dinitrogen trioxide (N2O3), and nitrous acid (HNO2) have been reported to be involved in the modification of cellular redox statues88,89.
Among the genes, regulated under hypoxia and SNP-treatment, two Cytochrome P450 encoding genes (CYP72A14 and CYP707A3), showed the highest up-regulation, > 30 and > 6-fold, respectively, in response to hypoxia (Fig. 8). Cytochrome P450 (CYP450) can convert toxic metabolites (e.g. superoxide anion, hydrogen peroxide and hydroxyl radical) to H2O2 to prevent harmful effects on the cell. Therefore, CYP450s are considered as markers for oxidative stress.
High level of ROS has to be scavenged by the cellular antioxidant system consisting of enzymatic antioxidants (e.g. superoxide dismutase (SOD), catalase (CAT) and glutathione peroxidases (GPXs), thioredoxin (Trx)) and non-enzymatic antioxidants (e.g. ascorbic acid, glutathione (GSH), carotenoids)90. Accordantly, up-regulation of genes encoding different classes of ROS scavenging enzymes were observed. Among six regulated GST encoding genes in our study, GSTU8 showed the highest up-regulation (ca. 6-fold) under hypoxia (Fig. 8). It has been shown that Arabidopsis gstu8 mutant line does not demonstrate any phenotypic changes nor modifications in the glutathione profile. However, it became evident that interaction between different dehydroascorbate reductases (DHARs) mediates the link between ascorbate and glutathione pools to ensure glutathione associated signaling under excessive H2O291.
MT2B, encoding metallothionein-like protein 2B, a ROS scavenger, showed down-regulation in our study (Fig. 12). It has been shown that MT2B down-regulation is associated with ROS accumulation and subsequent aerenchyma formation in rice64. This indicates that MT2B might be involved in hypoxia tolerance in tomato root. However, the role of NO during this process has not yet been addressed.
Hypoxia and SNP-treatment led to the down-regulation of AQP encoding genes
Members of two families of aquaporin (AQP) encoding genes (PIPs: 12 genes and TIPs: 7 genes) were down-regulated in our study in response to hypoxia and SNP-treatment (Fig. 9).
AQPs are involved in the transport of water and other small molecules such as ammonia, boron, CO2, H2O2 and urea across membranes92. Moreover, AQPs have been shown to be involved in plant biotic and abiotic stress response93. Expression reduction of NtAQP1, a member of the PIP1 family, in tobacco, led to a decrease in hydraulic conductivity of the root and eventually reduced drought stress resistance94.
In accordance with our results, microarray analysis showed that O2 deficiency resulted in the down-regulation of AQPs in Arabidopsis (Liu et al., 2005) and Avocado95. It has been shown that beside modifications in cytosolic Ca2+ and H2O2 level, low cytosolic pH during anoxia results in inhibition of hydraulic conductivity through a mechanism of pH dependent AQP gating96. During flooding, AQPs remain phosphorylated but closed due to the protonation of His193 in PIP2;1 of spinach plants97.
Tobacco PIP1;3 has been shown to be potentially involved in the O2 transmembrane transport98. Hypoxia stress in hydroponically grown tobacco, resulted in the up-regulation of PIP1;3 in the whole root98 and its down-regulation in lateral (LR) but not adventitious (AR) roots. The latter study demonstrated that beside PIP1;3, PIP1;1 was also down-regulated after 2 days of hypoxia treatment. Moreover, down-regulation of other AQP encoding genes such as PIP1;2, PIP1;4, PIP2;1 in LR, was observed after one week of hypoxia treatment99. The authors did not observe any difference in hydraulic conductance (Kr) between hypoxic and aerated plants, indicating the efficiency of AR in root water transport in tobacco. The down-regulation of PIP genes in our study is in accordance with AR response in tobacco. However, our study was focused on the whole root (Fig. 9). It has been reported that hypoxia is associated with a lower Kr in some species but an unchanged Kr in the others100,101. Tomato plants grown in soil, showed an early negative root hydraulic conductivity in response to flooding without affecting stomata closure102. It remains to determine whether expression changes in PIPs encoding genes is correlated with the functionality of PIPs and Kr in tomato root in response to hypoxia.
TIPs are involved in water transport between the vacuole and cytoplasm and therefore play a role in the regulation of cellular turgor pressure in plants93,103. One of the down-regulated TIP encoding genes in our study was TIP2;2 (Fig. 9). Overexpression of TIP2;2, in tomato, improved drought stress tolerance of transgenic plants through regulation of transpiration rate. This indicates that the water permeability rate across tonoplast is involved in drought stress tolerance104. Our data suggest that studying expression changes as well as activity of TIP2;2 during flooding induced hypoxia gives more insight into the importance of vacuole-cytoplasm water relation in flooding stress tolerance. It is noteworthy that protein storage vacuoles (PSVs) in stem cell niche and lytic vacuoles (LVs) in mature cell, contain distinct TIPs proteins in their membran105. Further studies are required to unravel the response of cell type and vacuole specific TIPs to nitric oxide and flooding.
NO transport through AQPs, as well as its potential role in expression regulation of AQPs, is not yet clear. In a human cell line expressing AQP1, NO permeability across the cell membrane was correlated with water permeability106. In the latter study, NO transport was significantly reduced after the addition of HgCl2, an aquaporin inhibitor. The authors concluded that NO transport by AQP1 controls intracellular NO levels and its consequences106. However, whether NO transport and controlling its cellular level during hypoxia can be executed via aquaporins in plant cells, needs further investigations.
This result indicates that little is known about the role of AQPs in response to hypoxia and the role of NO in this process is still unknown. Gene regulation does not necessarily correlate with protein amount and function. Transcriptional, post-transcriptional and posttranslational regulation can eventually affect the amount and the activity of AQPs. Moreover, the clear link between NO and AQPs under hypoxic conditions is not yet fully known. Our result indicates the necessity of future investigations to unravel the functional significance of AQPs and NO in root hypoxia tolerance.
Stress associated genes showed expression changes under hypoxia and SNP-treatment
Among stress related genes, SRO5 (SIMILAR TO RCD ONE 5) showed the highest up-regulation level in response to hypoxia (22.2-fold) in our study (Fig. 10). SRO5 has already been reported to be a common hypoxia-responsive gene, particularly in the root, throughout the plant kingdom65,107,108. The role of SRO5 in salt stress has been addressed in Arabidopsis. SRO5 overexpression led to a reduction in root H2O2 content in response to salt stress, compared to WT and sro5 plants109. Our data suggest that SRO5 might play a role in alleviating H2O2 level during hypoxia stress. Further investigations are required to unravel the link between hypoxia-induced NO and SRO5 in tomato root.
Among the stress related genes, Heat shock protein 23.6 (HSP23.6) showed the highest up-regulation under hypoxia. SNP application in Arabidopsis has shown that different members of the heat shock TF family are responsive to NO110. Moreover, HSPs and HSFs are also responsive to anoxia111,112. These data indicate that hypoxia-induced NO production might be involved in the regulation of HSPs. The possible cross talk between hypoxia and NO-mediated thermo-tolerance and HSPs chaperon function remains to be determined.
Comparison to former hypoxia and NO studies
Common regulated genes in response to hypoxia and SNP-treatment (FDR < 0.05) in our study were compared with the data obtained from a well-designed former study on Arabidopsis seedlings48. In the former study, 357 genes showed regulation changes in response to hypoxia in WT. Out of the above-mentioned genes, only four genes were shared with our study (Supplementary Table S3). This might be related to the fact that in our study only roots were exposed to hypoxia and SNP in a hydroponic system but in the other study, the whole seedlings were under submergence. Moreover, the plant age, 5 weeks old tomato vs. Arabidopsis seedlings, might partially explain the low overlap of the regulated genes between both studies. Moreover, regulated genes in our study were compared with the genes regulated in a triple NO mutant (nia1nia2noa1-2)under normoxia relative to WT48. 70 out of 251 (28%) genes in the current study were also regulated in the triple NO mutant under normoxia. Concomitantly, 51 out of 70 genes (73%) were regulated similarly in both studies (Supplementary Table S4).
The Arabidopsis N-degron pathway mutants (prt6 and ate1ate2) have been reported to exhibit constitutive expression of several core hypoxia genes (e.g. ADH1, SUS4 and PDC1) under normoxia leading to better resistance against hypoxia48. To identify the possible targets of the N-degron pathway in our study, regulated tomato genes were compared with the above-mentioned mutants in Arabidopsis. The result showed 13 common genes between tomato and Arabidopsis prt6 mutant. 10 genes, among them SUS4, were up-regulated (Supplementary Table S5). Comparison of our results with Arabidopsis ate1ate2 mutant revealed that 12 genes were common between two data sets, with 8 genes showing similar regulation changes (Supplementary Table S6).
In summary, our data suggest an overlap in gene expression response to long-term hypoxia and SNP-treatment. The concertedly regulated genes belong to different biological categories such as phytohormone signalling and transcription factor related genes as well as genes which their encoded proteins are involved in various metabolic pathways such as redox regulation, transport across membrane, glycolysis and fermentation. It can be proposed that the identified genes in our study could be considered as targets of hypoxia-induced NO and requires more investigation to unravel their role in the anatomic and metabolic adjustment of long-term hypoxia response and tolerance in tomato root. Beside TFs and phytohormones, the emphasis of the future studies needs to be placed on investigating the function of redox regulated proteins and their interaction with NO under hypoxia. These findings are essential to understand the cellular control of stress induced ROS/RNS as signal molecules as well as harmful radicals for the cell during hypoxia. A schematic model is illustrated to summarize the genes addressed in the discussion section (Fig. 13). It is noteworthy that this study was conducted on the RNA extracted from the whole root. Future investigations on phenotyping with cell type map and Single-cell RNA-seq approach can provide a more precise view on the cell type specific response and gene expression changes in response to nitric oxide and flooding stress.
A schematic model of the molecular response of tomato root to the yin and yang of oxygen and nitric oxide. Depicted are the genes belonging to different biological categories with identified or proposed roles in response to hypoxia and/or NO. Red and green font colors or arrows represent up- and down-regulation, respectively. ADH1, ALCOHOL DEHYDROGENASE 1; ENO2, ENOLASE 2; CA1, CARBONIC ANHYDRASE 1; CA2, CARBONIC ANHYDRASE 2; Kr: hydraulic conductivity, TIP, tonoplast intrinsic proteins; PIP, plasma membrane intrinsic proteins; PRX, peroxidase; GST, glutathione S-transferase; MT2B, METALLOTHIONEIN 2B;CYP450,cytochrome P-450; SRO5, SIMILAR TO RCD ONE 5; HSP23.6, HEAT SHOCK PROTEIN 23.6; ATAF2 (NAC081), ARABIDOPSIS NAC DOMAIN CONTAINING PROTEIN 81; WRKY7, WRKY DNA-BINDING PROTEIN 7; VNI2 (NAC083), VND-INTERACTING 2; RAP2.2, RELATED TO AP2 2; GASA5, GAST1 PROTEIN HOMOLOG 5; PIN2, PIN-FORMED 2; IAA14, INDOLE-3-ACETIC ACID INDUCIBLE 14; LOX1, LIPOXYGENASE 1; GA, gibberellic acid; AUX, auxin; JA, jasmonic acid; ETH, ethylene.
Methods
Plant material and growth conditions
Plant age and growth condition was the same for both hypoxia and SNP studies. The hypoxia treatment has been previously described in details53. Tomato plants (Solanum lycopersicum L. cv. Moneymaker) were grown on sand in the greenhouse (500 μmol photons/m2/s and 25 °C under a 14/10‐h light/dark) for three weeks. During this time, a modified Hoagland nutrient solution containing 5 mM nitrate (NO3-), as described previously113, was used for treatment. Three-week-old plants were transferred to hydroponic plastic boxes containing 6 L of nutrient solution (pH 5.5). Roots were submerged in the nutrient solution and aerated by mild bubbling using aquarist air pumps (Hailea ACO-9620, Raoping, Guangdong, China) and air outlets (Tetratec, Osnabrück, Germany). The hydroponic boxes were covered with dark covers, so that the roots were kept in dark. Despite dark, the low pH of the nutrient solution around the root will facilitate the release of NO from SNP114. The hypoxia treatment was conducted on five weeks old plant roots using N2 gas (≥ 99.99 Vol. %) (Air Liquide, Germany) for 48 h53. For SNP-treatment, the root of five weeks old plants was treated with final concentration of 500 μM SNP. The roots were harvested 48 h after the initiation of SNP-treatment in the solution for this study. Hypoxia was induced via root exposure to N2 gas53. Both treatments were started at 8:00 a.m. and continued 48 h until the harvest time at 8:00 a.m.
A Konica Minolta SPAD-502 chlorophyll meter was used to measure the relative chlorophyll levels of leaf #3 (the third leaf above cotyledones)53.
Read mapping and identification of differentially expressed genes
All the processes involved in data mapping and analysis has been described in a former study53. Adaptor clipped reads obtained from the NextSeq500 Illumina platform (LGC Biosearch Technologies, Berlin, Germany) were used for the following processes. After omitting short fragments and low quality reads, rRNA sequences were filtered. Remaining sequences were mapped to tomato reference genome (ITAG 2.4) (The Tomato Genome Consortium, 2012) using CLC Genomics Workbench (Qiagen, V. 7.5.5). Adaptor clipped reads obtained from the NextSeq500 Illumina platform (LGC Biosearch Technologies, Berlin, Germany. Sequencing data are deposited in the Sequence Read Archive (SRA) database (bioproject accession PRJNA553994) at the National Centre for Biotechnology Information (NCBI). The bioproject’s metadata are available at https://dataview.ncbi.nlm.nih.gov/object/PRJNA553994 reviewer = 684sto9a948tin240f0tlt1o1h.
The TMM (trimmed means of M values)115 and the edgeR algorithm116 were used for normalization and estimation of P-values, respectively. The algorithm edgeR was used for log fold change values. The P-values were adjusted for multiple testing117. All calculations were performed with the CLC Genomics Workbench software (Qiagen, V. 7.5.5). Differentially expressed genes (DEGs) (Padj < 0.05) (Supplementary Table S1) were selected for subsequent analysis. The FDR threshold was used for the P-value in multiple tests (Padj). GO term enrichment analysis (Padj < 0.05), was performed using the Panther database118,119. For biological pathway analysis of differentially regulated genes, MapMan categories based on ITAG 2.3 annotations54 were used. Heat maps of differentially regulated genes were created using MultiExperiment Viewer (MeV 4.9.0)120.
RNA isolation and cDNA synthesis
The whole root was snap-frozen and grounded in liquid N2. 250 mg of the homogenized grounded powder was used for total RNA extraction following phenol–chloroform extraction method121. The integrity of RNA was checked on 1.2% agarose gel. RNA concentration was quantified photometrically using NanoDrop (ND-1000, Thermo Scientific, Wilmington, DE, USA).
cDNA synthesis was performed with two μg DNase I-digested total RNA as template and using oligo-(dT)18 and RevertAid H Minus First Strand kit (Thermo Scientific, Waltham, USA).
qPCR primer design and assay
Primers for quantitative real-time PCR (qPCR) were calculated by QuantPrime122 (Supplementary Table S2). qPCR reactions were performed in total volume of 5 μl including 2.5 μl Power SYBR Green Master Mix (Thermo Fisher Scientific), 0.5 μM forward and reverse primers and 0.5 μl cDNA. ACTIN was used as reference gene123. The thermal profile used for all qPCRs was: 2 min 50 °C > 10 min 95 °C > (15 s 95 °C > 1 min 60 °C)40x. Data were analyzed by the 2−ΔΔCt method124.
Data availability
All materials and data sets represented in the current study are available in the main text or the supplementary materials. RNA-Seq. data are deposited to the Sequence Read Archive (SRA) database (bioproject accession PRJNA553994) at the National Center for Biotechnology Information (NCBI). The bioproject’s metadata are available at https://dataview.ncbi.nlm.nih.gov/object/PRJNA553994 reviewer = 684sto9a948tin240f0tlt1o1h.
References
Seligman, K., Saviani, E. E., Oliveira, H. C., Pinto-Maglio, C. A. & Salgado, I. Floral transition and nitric oxide emission during flower development in Arabidopsis thaliana is affected in nitrate reductase-deficient plants. Plant Cell Physiol.. 49, 1112–1121. https://doi.org/10.1093/pcp/pcn089 (2008).
Kolbert, Z., Bartha, B. & Erdei, L. Exogenous auxin-induced NO synthesis is nitrate reductase-associated in Arabidopsis thaliana root primordia. J. Plant. Physiol. 165, 967–975. https://doi.org/10.1016/j.jplph.2007.07.019 (2008).
Takahashi, M. et al. Nitrogen dioxide regulates organ growth by controlling cell proliferation and enlargement in Arabidopsis. New Phytol. 201, 1304–1315. https://doi.org/10.1111/nph.12609 (2014).
Mishina, T. E., Lamb, C. & Zeier, J. Expression of a nitric oxide degrading enzyme induces a senescence programme in Arabidopsis. Plant Cell Environ. 30, 39–52. https://doi.org/10.1111/j.1365-3040.2006.01604.x (2007).
Song, L., Ding, W., Zhao, M., Sun, B. & Zhang, L. Nitric oxide protects against oxidative stress under heat stress in the calluses from two ecotypes of reed. Plant Sci. 171, 449–458. https://doi.org/10.1016/j.plantsci.2006.05.002 (2006).
Montilla-Bascon, G. et al. Reduced nitric oxide levels during drought stress promote drought tolerance in barley and is associated with elevated polyamine biosynthesis. Sci. Rep. 7, 13311. https://doi.org/10.1038/s41598-017-13458-1 (2017).
Hasanuzzaman, M. et al. Nitric oxide pretreatment enhances antioxidant defense and glyoxalase systems to confer PEG-induced oxidative stress in rapeseed. J. Plant Interact. 12, 323–331. https://doi.org/10.1080/17429145.2017.1362052 (2017).
Xie, Y. et al. Reactive oxygen species-dependent nitric oxide production contributes to hydrogen-promoted stomatal closure in arabidopsis. Plant Physiol. 165, 759–773. https://doi.org/10.1104/pp.114.237925 (2014).
Ahmad, P. et al. Nitric oxide mitigates salt stress by regulating levels of osmolytes and antioxidant enzymes in chickpea. Front. Plant Sci. 7, 347. https://doi.org/10.3389/fpls.2016.00347 (2016).
Manai, J., Kalai, T., Gouia, H. & Corpas, F. J. Exogenous nitric oxide (NO) ameliorates salinity-induced oxidative stress in tomato (Solanum lycopersicum) plants. J. Soil Sci. Plant Nutr. 14, 433–446 (2014).
Ahlfors, R., Brosche, M., Kollist, H. & Kangasjarvi, J. Nitric oxide modulates ozone-induced cell death, hormone biosynthesis and gene expression in Arabidopsis thaliana. Plant J. 58, 1–12. https://doi.org/10.1111/j.1365-313X.2008.03756.x (2009).
Nabi, R. B. S. et al. Nitric oxide regulates plant responses to drought, salinity, and heavy metal stress. Environ. Exp. Bot. 161, 120–133. https://doi.org/10.1016/j.envexpbot.2019.02.003 (2019).
Dong, Y., Chen, W., Bai, X., Liu, F. & Wan, Y. Effects of exogenous nitric oxide and 24-epibrassinolide on the physiological characteristics of peanut seedlings under cadmium stress. Pedosphere 29, 45–59. https://doi.org/10.1016/s1002-0160(17)60376-x (2019).
Ding, Y., Gardiner, D. M., Xiao, D. & Kazan, K. https://doi.org/10.1101/779173 (2019)
Limami, A. M., Diab, H. & Lothier, J. Nitrogen metabolism in plants under low oxygen stress. Planta 239, 531–541. https://doi.org/10.1007/s00425-013-2015-9 (2014).
Stöhr, C. & Stremlau, S. Formation and possible roles of nitric oxide in plant roots. J. Exp. Bot. 57, 463–470. https://doi.org/10.1093/jxb/erj058 (2006).
van Dongen, J. T. & Licausi, F. Oxygen sensing and signaling. Annu. Rev. Plant Biol. 66, 345–367. https://doi.org/10.1146/annurev-arplant-043014-114813 (2015).
Hussain, A. et al. Nitric oxide mediated transcriptome profiling reveals activation of multiple regulatory pathways in Arabidopsis thaliana. Front. Plant Sci. 7, 975. https://doi.org/10.3389/fpls.2016.00975 (2016).
Jahnova, J., Luhova, L. & Petrivalsky, M. S-Nitrosoglutathione reductase-the master regulator of protein S-nitrosation in plant NO signaling. Plants (Basel) https://doi.org/10.3390/plants8020048 (2019).
Sun, X., Zhuang, Y., Lin, H. & Zhou, H. Patellin1 negatively regulates plant salt tolerance by attenuating nitric oxide accumulation in Arabidopsis. Plant Signal Behav. 14, 1675472. https://doi.org/10.1080/15592324.2019.1675472 (2019).
Zhan, N. et al. S-Nitrosylation targets GSNO reductase for selective autophagy during hypoxia responses in plants. Mol. Cell 71, 142–154. https://doi.org/10.1016/j.molcel.2018.05.024 (2018).
Gupta, K. J., Hebelstrup, K. H., Mur, L. A. & Igamberdiev, A. U. Plant hemoglobins: Important players at the crossroads between oxygen and nitric oxide. FEBS Lett. 585, 3843–3849. https://doi.org/10.1016/j.febslet.2011.10.036 (2011).
Hebelstrup, K. H. et al. Haemoglobin modulates NO emission and hyponasty under hypoxia-related stress in Arabidopsis thaliana. J. Exp. Bot. 63, 5581–5591. https://doi.org/10.1093/jxb/ers210 (2012).
Feng, J., Chen, L. & Zuo, J. Protein S-Nitrosylation in plants: Current progresses and challenges. J. Integr. Plant Biol. 61, 1206–1223. https://doi.org/10.1111/jipb.12780 (2019).
Corpas, F. J., Palma, J. M., del Rio, L. A. & Barroso, J. B. Evidence supporting the existence of L-arginine-dependent nitric oxide synthase activity in plants. New Phytol. 184, 9–14. https://doi.org/10.1111/j.1469-8137.2009.02989.x (2009).
Schopfer, F. J. et al. Fatty acid transduction of nitric oxide signalling. Nitrolinoleic acid is a hydrophobically stabilized nitric oxide donor. J. Biol. Chem. 280, 19289–19297. https://doi.org/10.1074/jbc.M414689200 (2005).
Padilla, M. N., Mata-Perez, C., Melguizo, M. & Barroso, J. B. In vitro nitro-fatty acid release from Cys-NO2-fatty acid adducts under nitro-oxidative conditions. Nitric Oxide 68, 14–22. https://doi.org/10.1016/j.niox.2016.12.009 (2017).
Astier, J., Gross, I. & Durner, J. Nitric oxide production in plants: An update. J. Exp. Bot. 69, 3401–3411. https://doi.org/10.1093/jxb/erx420 (2018).
Forstermann, U. & Sessa, W. C. Nitric oxide synthases: Regulation and function. Eur. Heart J. 33, 829–837, 837a–837d. https://doi.org/10.1093/eurheartj/ehr304 (2012).
Foresi, N. et al. Characterization of a nitric oxide synthase from the plant kingdom: NO generation from the green alga Ostreococcus tauri is light irradiance and growth phase dependent. Plant Cell 22, 3816–3830. https://doi.org/10.1105/tpc.109.073510 (2010).
Godber, B. L. J., Doel, J. J., Durgan, J., Eisenthal, R. & Harrison, R. A new route to peroxynitrite: A role for xanthine oxidoreductase. FEBS Lett. 475, 93–96 (2000).
Godber, B. L. J. et al. Reduction of nitrite to nitric oxide catalyzed by xanthine oxidoreductase. J. Biol. Chem. 275(11), 7757–7763 (2000).
Cantu-Medellin, N. & Kelley, E. E. Xanthine oxidoreductase-catalyzed reduction of nitrite to nitric oxide: Insights regarding where, when and how. Nitric Oxide 34, 19–26. https://doi.org/10.1016/j.niox.2013.02.081 (2013).
Yamamoto-Katou, A., Katou, S., Yoshioka, H., Doke, N. & Kawakita, K. Nitrate reductase is responsible for elicitin-induced nitric oxide production in Nicotiana benthamiana. Plant Cell Physiol. 47, 726–735. https://doi.org/10.1093/pcp/pcj044 (2006).
Planchet, E., Jagadis Gupta, K., Sonoda, M. & Kaiser, W. M. Nitric oxide emission from tobacco leaves and cell suspensions: Rate limiting factors and evidence for the involvement of mitochondrial electron transport. Plant J. 41, 732–743. https://doi.org/10.1111/j.1365-313X.2005.02335.x (2005).
Igamberdiev, A. U., Ratcliffe, R. G. & Gupta, K. J. Plant mitochondria: Source and target for nitric oxide. Mitochondrion 19(Pt B), 329–333. https://doi.org/10.1016/j.mito.2014.02.003 (2014).
Gas, E., Flores-Perez, U., Sauret-Gueto, S. & Rodriguez-Concepcion, M. Hunting for plant nitric oxide synthase provides new evidence of a central role for plastids in nitric oxide metabolism. Plant Cell 21, 18–23. https://doi.org/10.1105/tpc.108.065243 (2009).
Stöhr, C., Strube, F., Marx, G., Ullrich, W. R. & Rockel, P. A plasma membrane-bound enzyme of tobacco roots catalyses the formation of nitric oxide from nitrite. Planta 212, 835–841. https://doi.org/10.1007/s004250000447 (2001).
Chamizo-Ampudia, A. et al. A dual system formed by the ARC and NR molybdoenzymes mediates nitrite-dependent NO production in Chlamydomonas. Plant Cell Environ. 39, 2097–2107. https://doi.org/10.1111/pce.12739 (2016).
Chamizo-Ampudia, A., Sanz-Luque, E., Llamas, A., Galvan, A. & Fernandez, E. Nitrate reductase regulates plant nitric oxide homeostasis. Trends Plant Sci. 22, 163–174. https://doi.org/10.1016/j.tplants.2016.12.001 (2017).
Bethke, P. C., Badger, M. R. & Jones, R. L. Apoplastic synthesis of nitric oxide by plant tissues. Plant Cell 16, 332–341. https://doi.org/10.1105/tpc.017822 (2004).
Cooney, R. V., Harwood, P. J., Custer, L. J. & Franke, A. A. Light-mediated conversion of nitrogen dioxide to nitric oxide by carotenoids. Environ. Health Perspect. 102(5), 460–462 (1994).
Antoniou, C. et al. Developmental stage- and concentration-specific sodium nitroprusside application results in nitrate reductase regulation and the modification of nitrate metabolism in leaves of Medicago truncatula plants. Plant Signal Behav. https://doi.org/10.4161/psb.25479 (2013).
Zandonadi, D. B. et al. Nitric oxide mediates humic acids-induced root development and plasma membrane H+-ATPase activation. Planta 231, 1025–1036. https://doi.org/10.1007/s00425-010-1106-0 (2010).
Floryszak-Wieczorek, J., Milczarek, G., Arasimowicz, M. & Ciszewski, A. Do nitric oxide donors mimic endogenous NO-related response in plants?. Planta 224, 1363–1372. https://doi.org/10.1007/s00425-006-0321-1 (2006).
Sasidharan, R. et al. Signal dynamics and interactions during flooding stress. Plant Physiol. 176, 1106–1117. https://doi.org/10.1104/pp.17.01232 (2018).
Millar, A. H. & Day, D. A. Nitric oxide inhibits the cytochrome oxidase but not the alternative oxidase of plant mitochondria. FEBS Lett. 398, 155–158. https://doi.org/10.1016/s0014-5793(96)01230-6 (1996).
Gibbs, D. J. et al. Nitric oxide sensing in plants is mediated by proteolytic control of group VII ERF transcription factors. Mol. Cell 53, 369–379. https://doi.org/10.1016/j.molcel.2013.12.020 (2014).
Gupta, K. J., Igamberdiev, A. U. & Mur, L. A. NO and ROS homeostasis in mitochondria: A central role for alternative oxidase. New Phytol. 195, 1–3. https://doi.org/10.1111/j.1469-8137.2012.04189.x (2012).
Gibbs, D. J. et al. Homeostatic response to hypoxia is regulated by the N-end rule pathway in plants. Nature 479, 415–418. https://doi.org/10.1038/nature10534 (2011).
Kuo, C. G. & Chen, B. W. Physical responses of tomato cultivars to flooding. J. Am. Soc. Hortic. Sci 105, 751–755 (1980).
Kuo, C. G., Tsay, J. S., Chen, B. W. & Lin, P. Y. Screening for flooding tolerance in the genus Lycopersicon. HortScience 17(1), 76–78 (1982).
Safavi-Rizi, V., Hede, M. & Stöhr, C. RNA-Seq reveals novel genes and pathways associated with hypoxia duration and tolerance in tomato root. Sci. Rep. 10, 1692. https://doi.org/10.1038/s41598-020-57884-0 (2020).
Thimm, O. et al. MAPMAN: A user-driven tool to display genomics data sets onto diagrams of metabolic pathways and other biological processes. Plant J. 37, 914–939 (2004).
Tuteja, N. Mechanisms of high salinity tolerance in plants. Methods Enzymol. 428, 419–438. https://doi.org/10.1016/S0076-6879(07)28024-3 (2007).
Imran, Q. M. et al. Transcriptome profile of NO-induced Arabidopsis transcription factor genes suggests their putative regulatory role in multiple biological processes. Sci. Rep. 8, 771. https://doi.org/10.1038/s41598-017-18850-5 (2018).
Huang, J., Wei, H., Li, L. & Yu, S. Transcriptome analysis of nitric oxide-responsive genes in upland cotton (Gossypium hirsutum). PLoS One 13, e0192367. https://doi.org/10.1371/journal.pone.0192367 (2018).
Zeng, F., Sun, F., Li, L., Liu, K. & Zhan, Y. Genome-scale transcriptome analysis in response to nitric oxide in birch cells: Implications of the triterpene biosynthetic pathway. PLoS One 9, e116157. https://doi.org/10.1371/journal.pone.0116157 (2014).
Terrile, M. C. et al. Nitric oxide influences auxin signaling through S-nitrosylation of the Arabidopsis TRANSPORT INHIBITOR RESPONSE 1 auxin receptor. Plant J 70, 492–500. https://doi.org/10.1111/j.1365-313X.2011.04885.x (2012).
Eysholdt-Derzso, E. & Sauter, M. Root Bending is antagonistically affected by hypoxia and ERF-mediated transcription via auxin signaling. Plant Physiol. 175, 412–423. https://doi.org/10.1104/pp.17.00555 (2017).
Okushima, Y., Fukaki, H., Onoda, M., Theologis, A. & Tasaka, M. ARF7 and ARF19 regulate lateral root formation via direct activation of LBD/ASL genes in Arabidopsis. Plant Cell 19, 118–130. https://doi.org/10.1105/tpc.106.047761 (2007).
Fukaki, H., Tameda, S., Masuda, H. & Tasaka, M. Lateral root formation is blocked by a gain-of-function mutation in the SOLITARY-ROOT/IAA14 gene of Arabidopsis. Plant J. 29(2), 153–168 (2002).
Shukla, V. et al. Endogenous hypoxia in lateral root Primordia controls root architecture by antagonizing auxin signaling in arabidopsis. Mol. Plant 12, 538–551. https://doi.org/10.1016/j.molp.2019.01.007 (2019).
Steffens, B., Geske, T. & Sauter, M. Aerenchyma formation in the rice stem and its promotion by H2O2. New Phytol. 190, 369–378. https://doi.org/10.1111/j.1469-8137.2010.03496.x (2011).
Hartman, S. et al. Ethylene-mediated nitric oxide depletion pre-adapts plants to hypoxia stress. Nat. Commun. 10, 4020. https://doi.org/10.1038/s41467-019-12045-4 (2019).
Niu, Y. H. & Guo, F. Q. Nitric oxide regulates dark-induced leaf senescence through EIN2 in Arabidopsis. J. Integr. Plant Biol. 54, 516–525. https://doi.org/10.1111/j.1744-7909.2012.01140.x (2012).
Keunen, E. et al. A mutant of the Arabidopsis thaliana LIPOXYGENASE1 gene shows altered signalling and oxidative stress related responses after cadmium exposure. Plant Physiol. Biochem. 63, 272–280. https://doi.org/10.1016/j.plaphy.2012.12.005 (2013).
Huang, X. et al. Nitric oxide is induced by wounding and influences jasmonic acid signaling in Arabidopsis thaliana. Planta 218, 938–946. https://doi.org/10.1007/s00425-003-1178-1 (2004).
Mur, L. A., Prats, E., Pierre, S., Hall, M. A. & Hebelstrup, K. H. Integrating nitric oxide into salicylic acid and jasmonic acid/ethylene plant defense pathways. Front. Plant Sci. 4, 215. https://doi.org/10.3389/fpls.2013.00215 (2013).
Lozano-Juste, J. & Leon, J. Nitric oxide regulates DELLA content and PIF expression to promote photomorphogenesis in Arabidopsis. Plant Physiol. 156, 1410–1423. https://doi.org/10.1104/pp.111.177741 (2011).
Freschi, L. Nitric oxide and phytohormone interactions: Current status and perspectives. Front. Plant Sci. 4, 398. https://doi.org/10.3389/fpls.2013.00398 (2013).
Zhang, S., Yang, C., Peng, J., Sun, S. & Wang, X. GASA5, a regulator of flowering time and stem growth in Arabidopsis thaliana. Plant Mol. Biol. 69, 745–759. https://doi.org/10.1007/s11103-009-9452-7 (2009).
Bailey-Serres, J. et al. Making sense of low oxygen sensing. Trends Plant Sci. 17, 129–138. https://doi.org/10.1016/j.tplants.2011.12.004 (2012).
Licausi, F. et al. Oxygen sensing in plants is mediated by an N-end rule pathway for protein destabilization. Nature 479, 419–422. https://doi.org/10.1038/nature10536 (2011).
Castillo, M. C., Coego, A., Costa-Broseta, A. & Leon, J. Nitric oxide responses in Arabidopsis hypocotyls are mediated by diverse phytohormone pathways. J. Exp. Bot. 69, 5265–5278. https://doi.org/10.1093/jxb/ery286 (2018).
Hinz, M. et al. Arabidopsis RAP2.2: An ethylene response transcription factor that is important for hypoxia survival. Plant Physiol. 153, 757–772. https://doi.org/10.1104/pp.110.155077 (2010).
Till, C. J. et al. The Arabidopsis thaliana N-recognin E3 ligase PROTEOLYSIS1 influences the immune response. Plant Direct. 3, e00194. https://doi.org/10.1002/pld3.194 (2019).
Christianson, J. A., Dennis, E. S., Llewellyn, D. J. & Wilson, I. W. ATAF NAC transcription factors: Regulators of plant stress signaling. Plant Signal Behav. 5, 428–432. https://doi.org/10.4161/psb.5.4.10847 (2010).
Parani, M. et al. Microarray analysis of nitric oxide responsive transcripts in Arabidopsis. Plant Biotechnol. J. 2, 359–366. https://doi.org/10.1111/j.1467-7652.2004.00085.x (2004).
Jin, C. et al. A novel NAC transcription factor, PbeNAC1, of Pyrus betulifolia confers cold and drought tolerance via interacting with PbeDREBs and activating the expression of stress-responsive genes. Front. Plant Sci. 8, 1049. https://doi.org/10.3389/fpls.2017.01049 (2017).
Li, Z. et al. The bHLH family member ZmPTF1 regulates drought tolerance in maize by promoting root development and abscisic acid synthesis. J. Exp. Bot. 70, 5471–5486. https://doi.org/10.1093/jxb/erz307 (2019).
Yang, S. D., Seo, P. J., Yoon, H. K. & Park, C. M. The Arabidopsis NAC transcription factor VNI2 integrates abscisic acid signals into leaf senescence via the COR/RD genes. Plant Cell 23, 2155–2168. https://doi.org/10.1105/tpc.111.084913 (2011).
Seo, P. J. & Park, C. M. Signaling linkage between environmental stress resistance and leaf senescence in Arabidopsis. Plant Signal Behav. 6, 1564–1566. https://doi.org/10.4161/psb.6.10.17003 (2011).
Hsu, F. C. et al. Submergence confers immunity mediated by the WRKY22 transcription factor in Arabidopsis. Plant Cell 25, 2699–2713. https://doi.org/10.1105/tpc.113.114447 (2013).
Schmidt, S., Dethloff, F., Beine-Golovchuk, O. & Kopka, J. The REIL1 and REIL2 proteins of Arabidopsis thaliana are required for leaf growth in the cold. Plant Physiol. 163, 1623–1639. https://doi.org/10.1104/pp.113.223925 (2013).
Blokhina, O. & Fagerstedt, K. V. Oxidative metabolism, ROS and NO under oxygen deprivation. Plant. Physiol. Biochem. 48, 359–373. https://doi.org/10.1016/j.plaphy.2010.01.007 (2010).
Paradiso, A. et al. ROS production and scavenging under anoxia and re-oxygenation in Arabidopsis cells: A balance between redox signaling and impairment. Front. Plant Sci. 7, 1803. https://doi.org/10.3389/fpls.2016.01803 (2016).
Schafer, F. Q. & Buettner, G. R. Redox environment of the cell as viewed through the redox state of the glutathione disulfide/glutathione couple. Free Radic. Biol. Med. 30, 1191–1212. https://doi.org/10.1016/s0891-5849(01)00480-4 (2001).
Blanco, S., Hernandez, R., Franchelli, G., Ramos-Alvarez, M. M. & Peinado, M. A. Melatonin influences NO/NOS pathway and reduces oxidative and nitrosative stress in a model of hypoxic-ischemic brain damage. Nitric Oxide 62, 32–43. https://doi.org/10.1016/j.niox.2016.12.001 (2017).
He, L. et al. Antioxidants maintain cellular redox homeostasis by elimination of reactive oxygen species. Cell Physiol. Biochem. 44, 532–553. https://doi.org/10.1159/000485089 (2017).
Rahantaniaina, M. S. et al. Glutathione oxidation in response to intracellular H2O2: Key but overlapping roles for dehydroascorbate reductases. Plant Signal Behav. 12, e1356531. https://doi.org/10.1080/15592324.2017.1356531 (2017).
Sabir, F. et al. Exploring three PIPs and three TIPs of grapevine for transport of water and atypical substrates through heterologous expression in aqy-null yeast. PLoS One 9, e102087. https://doi.org/10.1371/journal.pone.0102087 (2014).
Afzal, Z., Howton, T. C., Sun, Y. & Mukhtar, M. S. The roles of aquaporins in plant stress responses. J. Dev. Biol. https://doi.org/10.3390/jdb4010009 (2016).
Kaldenhoff, R., Bertl, A., Otto, B., Moshelion, M. & Uehlein, N. Characterization of plant aquaporins. Methods Enzymol. 428, 505–531. https://doi.org/10.1016/S0076-6879(07)28028-0 (2007).
Reeksting, B. J., Olivier, N. A. & van den Berg, N. Transcriptome responses of an ungrafted Phytophthora root rot tolerant avocado (Persea americana) rootstock to flooding and Phytophthora cinnamomi. BMC Plant Biol. 16, 205. https://doi.org/10.1186/s12870-016-0893-2 (2016).
Tournaire-Roux, C. et al. Cytosolic pH regulates root water transport during anoxic stress through gating of aquaporins. Lett. Nat. 120, 393–397 (2003).
Tornroth-Horsefield, S. et al. Structural mechanism of plant aquaporin gating. Nature 439, 688–694. https://doi.org/10.1038/nature04316 (2006).
Zwiazek, J. J., Xu, H., Tan, X., Navarro-Rodenas, A. & Morte, A. Significance of oxygen transport through aquaporins. Sci. Rep. 7, 40411. https://doi.org/10.1038/srep40411 (2017).
Tan, X. & Zwiazek, J. J. Stable expression of aquaporins and hypoxia-responsive genes in adventitious roots are linked to maintaining hydraulic conductance in tobacco (Nicotiana tabacum) exposed to root hypoxia. PLoS One 14, e0212059. https://doi.org/10.1371/journal.pone.0212059 (2019).
Araki, H. Water uptake of soybean (Glycine max L. Merr.) during exposure to O2 deficiency and field level CO2 concentration in the root zone. Field Crops .earch 96, 98–105. https://doi.org/10.1016/j.fcr.2005.05.007 (2006).
Islam, M. A. & Macdonald, S. E. Ecophysiological adaptations of black spruce (Picea mariana) and tamarack (Larix laricina) seedlings to flooding. Trees Struct. Funct. 18, 35–42. https://doi.org/10.1007/s00468-003-0276-9 (2004).
Else, M. A., Davies, W. J., Malone, M. & Jackson, M. B. A negative hydraulic message from oxygen-deficient roots of tomato plants? (Influence of soil flooding on leaf water potential, leaf expansion, and synchrony between stomatal conductance and root hydraulic conductivity). Plant Physiol. 109, 1017–1024. https://doi.org/10.1104/pp.109.3.1017 (1995).
Kapilan, R., Vaziri, M. & Zwiazek, J. J. Regulation of aquaporins in plants under stress. Biol. Res. 51, 4. https://doi.org/10.1186/s40659-018-0152-0 (2018).
Sade, N. et al. Improving plant stress tolerance and yield production: Is the tonoplast aquaporin SlTIP2;2 a key to isohydric to anisohydric conversion?. New Phytol. 181, 651–661. https://doi.org/10.1111/j.1469-8137.2008.02689.x (2009).
Jiang, L. P. et al. The protein storage vacuole. JCB 156, 991–1002 (2001).
Herrera, M., Hong, N. J. & Garvin, J. L. Aquaporin-1 transports NO across cell membranes. Hypertension 48, 157–164. https://doi.org/10.1161/01.HYP.0000223652.29338.77 (2006).
Mustroph, A. et al. Cross-kingdom comparison of transcriptomic adjustments to low-oxygen stress highlights conserved and plant-specific responses. Plant Physiol. 152, 1484–1500. https://doi.org/10.1104/pp.109.151845 (2010).
van Veen, H. et al. Transcriptomes of eight Arabidopsis thaliana accessions reveal core conserved, genotype- and organ-specific responses to flooding stress. Plant Physiol. 172, 668–689. https://doi.org/10.1104/pp.16.00472 (2016).
Babajani, G., Effendy, J. & Plant, A. L. Sl-SROl1 increases salt tolerance and is a member of the radical-induced cell death 1—similar to RCD1 gene family of tomato. Plant Sci. 176, 214–222. https://doi.org/10.1016/j.plantsci.2008.10.012 (2009).
Singh, P. K. et al. Nitric oxide mediated transcriptional modulation enhances plant adaptive responses to arsenic stress. Sci. Rep. 7, 3592. https://doi.org/10.1038/s41598-017-03923-2 (2017).
Banti, V. et al. Heat acclimation and cross-tolerance against anoxia in Arabidopsis. Plant Cell Environ. 31, 1029–1037. https://doi.org/10.1111/j.1365-3040.2008.01816.x (2008).
Loreti, E., Poggi, A., Novi, G., Alpi, A. & Perata, P. A genome-wide analysis of the effects of sucrose on gene expression in Arabidopsis seedlings under anoxia. Plant Physiol. 137, 1130–1138. https://doi.org/10.1104/pp.104.057299 (2005).
Wendlandt, T., Moche, M., Becher, D. & Stöhr, C. A SDD1-like subtilase is exuded by tobacco roots. Funct. Plant Biol. 43, 141. https://doi.org/10.1071/fp15211 (2016).
Smith, N. & Dasgupta, T. P. Mechanism of nitric oxide release. I. Two-electron reduction of sodium nitroprusside by l-cysteine in aqueous solution inorganic reaction mechanisms. Inorg. React. Mech. 3, 181–195. https://doi.org/10.1080/10286620210352 (2002).
Robinson, M. D. & Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 11, R25. https://doi.org/10.1186/gb-2010-11-3-r25 (2010).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: A bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140. https://doi.org/10.1093/bioinformatics/btp616 (2010).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. R. Statist. Soc. B 57(1), 289–300 (1995).
Thomas, P. D. et al. PANTHER: A library of protein families and subfamilies indexed by function. Genome Res. 13, 2129–2141. https://doi.org/10.1101/gr.772403 (2003).
Thomas, P. D. et al. Applications for protein sequence-function evolution data: mRNA/protein expression analysis and coding SNP scoring tools. Nucleic Acids Res. 34, W645-650. https://doi.org/10.1093/nar/gkl229 (2006).
Saeed, A. I. et al. TM4 microarray software suite. Methods Enzymol. 411, 134–193. https://doi.org/10.1016/S0076-6879(06)11009-5 (2006).
Chomczynski, P. & Sacchi, N. The single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction: Twenty-something years on. Nat. Protoc. 1, 581–585. https://doi.org/10.1038/nprot.2006.83 (2006).
Arvidsson, S., Kwasniewski, M., Riano-Pachon, D. M. & Mueller-Roeber, B. QuantPrime—a flexible tool for reliable high-throughput primer design for quantitative PCR. BMC Bioinform. 9, 465. https://doi.org/10.1186/1471-2105-9-465 (2008).
Ren, Z. et al. The auxin receptor homologue in Solanum lycopersicum stimulates tomato fruit set and leaf morphogenesis. J. Exp. Bot. 62, 2815–2826. https://doi.org/10.1093/jxb/erq455 (2011).
Schmittgen, T. D. & Livak, K. J. Analyzing real-time PCR data by the comparative C(T) method. Nat. Protoc. 3, 1101–1108 (2008).
Acknowledgements
We thank Seyedhamid Safavi Rizi, PNU University, for his computational assistance in management and analysing the large-scale data set. We acknowledge financial support for the Article Processing Charge from the DFG (German Research Foundation, 393148499) and the Open Access Publication Fund of the University of Greifswald.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
V.S.R. conceived, designed, and performed experiments, analysed the data and wrote the paper; M.H. contributed to RNA-Seq analysis, C.S. conceived the experiment and edited the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Safavi-Rizi, V., Herde, M. & Stöhr, C. Identification of nitric oxide (NO)-responsive genes under hypoxia in tomato (Solanum lycopersicum L.) root. Sci Rep 10, 16509 (2020). https://doi.org/10.1038/s41598-020-73613-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-73613-z
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.