Abstract
Grapevine (Vitis vinifera L.) is a crop of major economic importance. However, grapevine yield is guaranteed by the massive use of pesticides to counteract pathogen infections. Under temperate-humid climate conditions, downy mildew is a primary threat for viticulture. Downy mildew is caused by the biotrophic oomycete Plasmopara viticola Berl. & de Toni, which can attack grapevine green tissues. In lack of treatments and with favourable weather conditions, downy mildew can devastate up to 75% of grape cultivation in one season and weaken newly born shoots, causing serious economic losses. Nevertheless, the repeated and massive use of some fungicides can lead to environmental pollution, negative impact on non-targeted organisms, development of resistance, residual toxicity and can foster human health concerns. In this manuscript, we provide an innovative approach to obtain specific pathogen protection for plants. By using the yeast two-hybrid approach and the P. viticola cellulose synthase 2 (PvCesA2), as target enzyme, we screened a combinatorial 8 amino acid peptide library with the aim to identify interacting peptides, potentially able to inhibit PvCesa2. Here, we demonstrate that the NoPv1 peptide aptamer prevents P. viticola germ tube formation and grapevine leaf infection without affecting the growth of non-target organisms and without being toxic for human cells. Furthermore, NoPv1 is also able to counteract Phytophthora infestans growth, the causal agent of late blight in potato and tomato, possibly as a consequence of the high amino acid sequence similarity between P. viticola and P. infestans cellulose synthase enzymes.
Similar content being viewed by others
Introduction
Pesticide use is at the basis of intensive agriculture, as it guarantees protection from pathogens, pests and weeds, which otherwise would count for up to 40% of production losses1,2,3. In the European Union (EU), almost 500 active substances, most of them able to inhibit key metabolic pathways of plant pathogens4, are approved as pesticides (European commission, EU pesticides database, 2018; ec.europa.eu/food/plant/pesticides/eu-pesticides-database), with annual sales of 374,000 tons (Eurostat. Pesticide sales, 2018; ec.europa.eu/eurostat/web/products-datasets/-/aei_fm_salpest09), whereas the global use of pesticides accounted for 4,113,591 tonnes in 2017 (FAOSTAT, fao.org/faostat/en/#data/RP/visualize). However, despite the beneficial effects on agricultural production, some pesticides can have a harmful impact on the environment and on the health of humans and animals depending on their concentration, bioavailability and rate of transport through the soil5,6,7,8,9,10,11. Dermatological, gastrointestinal, carcinogenic, respiratory and endocrine pathologies are among the negative health effects that have been associated with the massive use of chemical pesticides12. Residues of pesticides can be found in foods and beverages13,14, although in most of the cases the concentrations do not exceed the safe levels15. However, the simultaneous exposure to different compounds may have negative synergistic effects16. Consequently, stricter regulations have been enacted both in the EU [Sustainable Use Directive 2009/128/EC and Plant Protection Products Regulation (EC) 1107/2009] and in the USA (Insecticide, Fungicide, and Rodenticide Act). Several compounds have been banned or included in a list of candidates for substitution (ec.europa.eu/food/plant/pesticides/approval_active_substances_en) because of problems of environmental toxicity, effects on human health or development of resistance in the target pathogen, similarly to the case of antibiotics17.
Taken these considerations together, it appears clear that new biotechnological sustainable solutions need to be explored to find safe and reliable alternatives to conventional pesticides that specifically inhibit the activity of pathogen key enzymes. In recent years, peptide aptamers—i.e. short synthetic peptides able to specifically bind and inhibit a protein target—have emerged as novel molecular tools that have attracted the attention of different research groups interested in developing antimicrobial compounds18,19, presumably with a better environmental fate, lower off-target effects and possibly a low-risk alternative to conventional pesticides, although this technology requires experimental validation.
Among crops, grapevine represents a great agricultural and economic value worldwide, with 7.534,00 Mha invested in viticulture20. However, the grapevine industry relies predominantly on Vitis vinifera, which is susceptible to a large spectrum of pathogens and requires a frequent application of chemical antimicrobials to avoid yield and quality loss21. In particular, under temperate-humid climate conditions downy mildew, a worldwide destructive disease caused by the biotrophic oomycete Plasmopara viticola Berl. & De Toni22, is considered of primary importance for viticulture20. In lack of treatments and with favourable weather conditions, downy mildew can devastate up to 75% of the crop in one season and weaken newly emerging shoots, causing serious economic losses23,24. P. viticola attacks grapevine green organs, such as leaves and young fruits, by releasing flagellate zoospores at high humidity and warm temperatures25. When the zoospores encounter a stoma, they shed the flagella and encyst. Subsequently, a germ tube emerges from each spore and reaches the substomatal cavity, where it dilates into a vesicle that initiates the infection26.
Data available for the oomycete Phytophthora infestans, the causal agent of late blight in potato and tomato23, provide useful information on the molecular mechanisms at the basis of oomycete plant infection. The main component of P. infestans cell wall is cellulose, accounting for the 33.6% over the total glucan composition (85.6%)27. Cellulose appears to be essential for appressorium formation and effective potato infection by P. infestans, since inhibition of its biosynthesis leads to a dramatic reduction in the number of normal germ tubes with appressoria, severe disruption of the cell wall in the pre-infection structures, and a complete loss of pathogenicity28. Furthermore, cellulose synthase gene expression is up-regulated during pre- and early-infection stages of P. infestans in potato28. Additionally, P. infestans and P. viticola cellulose synthase enzymes have been identified as the target of the fungicide mandipropamid, highlighting the relevance of cell wall biosynthesis in oomycete disease development29,30.
In this manuscript, we describe the identification of a novel peptide aptamer of 8 amino acid residues, named NoPv1 (No Plasmopora viticola 1), able to counteract P. viticola infection ex vivo (leaf disks) and in vivo (potted plants). In particular, the PvCesA2 (P. viticola cellulose synthase 2) enzyme was used as a bait in the yeast two-hybrid assay to isolate the NoPv1 peptide, from a combinatorial peptide library31. NoPv1 prevents P. viticola germ tube formation and grapevine infection without affecting the growth of off-target organisms, and without being toxic for plant and human cells. Furthermore, NoPv1 is also able to arrest the growth of P. infestans probably due to the high amino acid sequence similarity of its cellulose synthase enzymes with the ones of P. viticola.
Results
Identification of peptide aptamers interacting with the Plasmopara viticola cellulose synthase 2, PvCesA2
The PvCesA2 cellulose synthase enzyme has been selected as target enzyme for our study since cellulose biosynthesis plays a pivotal role during pathogen infection, as demonstrated by Grenville-Briggs and co-workers28 in the closely related oomycete species, P. infestans. Furthermore, phylogenetic analysis performed by comparing cellulose synthase amino acid sequences from different organisms (see Fig. 1, Table S1 and Data S1), highlighted that oomycete cellulose synthase proteins group together as a distinct clade, separate from bacterial, cyanobacterial, viral, fungal and plant cellulose synthases28,30. In addition, it is also interesting to observe that among the four Oomycota CesAs, CesA3 enzymes cluster in a slightly different clade from CesA1, CesA2 and CesA4, probably due to the lack of a Plekstrin Homology domain (PH domain), involved in regulation, trafficking and/or targeting of polysaccharide synthases to the correct site for cell wall biosynthesis32.
Dendrogram of cellulose synthases derived from Oomycota, Prokarya and Eukarya sequences. Phylogenetic analyses were conducted using MEGA version X66 (see Data S1 for Clustal omega multiple sequence alignment). The generated dendrogram demonstrates that Oomycota CesAs group together in a separate clade from all the other considered cellulose synthase and cellulose synthase-like sequences. The analysis is based on complete amino acid sequences reported in Table S1. CesA, cellulose synthase; CslA-H, cellulose synthase-like; Pv, Plasmopara viticola; Pi, Phytophthora infestans; Ns, Nostoc sp.; Tv, Trichormus variabilis; At, Arabidopsis thaliana; Vv, Vitis vinifera; Pb, Physcomitrella patens; Pt, Populus tremula; Sl, Solanum tuberosum; Zm, Zea mays; Os, Oryza sativa; Atm, Agrobacterium tumefaciens; Ec, Escherichia coli; Kx, Komagataeibacter xylinus; Rl, Rhizobium leguminosarum; Ps, Pseudomonas syringae; Pf, Pseudomonas fluorescens; Ba, Bacillus amyloliquefaciens; PbC, Paramecium bursaria Chlorella virus 1; Cr, Ceratopteris richardii; Pp, Physcomitrella patens; Tl, Trichoderma longibrachiatum; Th, Trichoderma harzianum; Jf, Jimgerdemannia flammicorona; Gp, Gonapodya prolifera; Am, Allomyces macrogynus.
These notions, together with the fact that CesA2 is the most abundant among the PH domain-containing CesA enzymes28, and that cellulose synthase enzymes are absent in many beneficial organisms, such as Saccharomyces cerevisiae, and in animal cells, including human cells, make PvCesA2 the ideal target for developing antimicrobial active compounds.
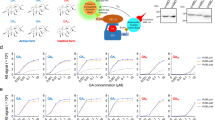
Peptide aptamers able to specifically interact with PvCesA2 were identified by using the yeast two-hybrid strategy. The cytoplasmic soluble portion of PvCesA2 (from aa 331 to 790; see also Data S1), containing all typical signatures of most processive glycosyltransferases, including three aspartic acid residues and the QXXRW motif28,33, was fused to the GAL4 DNA Binding Domain (BD) and used as bait to screen a combinatorial library of linear peptide aptamers (8 a.a. in length; see also “Materials and methods” for further details). Ten small scale transformations were performed (⁓ 106 transformants obtained) and only 30 colonies were able to grow on selective media, lacking either adenine or histidine, and supplemented with 10 mM of 3-amino-1,2,4-triazole (3-AT), a competitive inhibitor for the HIS3 reporter gene. To identify the peptide aptamers interacting with PvCesA2, plasmids were purified and sequenced, and priority was given to peptides whose sequences appeared at least three times among the 30 colonies isolated. In total three peptides, named “No Plasmopora viticola 1”, − 2 and − 3 (NoPv1, NoPv2 and NoPv3; see also Table 1), were prepared via Microwave-assisted Solid Phase Peptide Synthesis (see “Materials and methods” for further details) and analysed for their capability to counteract P. viticola infection of grapevine leaf disks through co-inoculation assays.
In details, leaf disks of V. vinifera cv. Pinot noir, susceptible to P. viticola infection, were treated with sporangia suspension in either absence or presence of 200 μM NoPv1, NoPv2 and NoPv3 (Fig. 2a–f). P. viticola sporulation was clearly visible on grapevine leaf disks in the absence of NoPv peptides (Fig. 2a,c,g; Control) at 5 and 7 days post-inoculation (dpi), and NoPv2 and NoPv3 peptide aptamers showed intermediate and no effect on P. viticola severity, respectively (Fig. 2e,f,g). On the contrary, 200 μM NoPv1 peptide was able to completely inhibit P. viticola severity on leaf disks (Fig. 2b,d,g), without causing any damage to the leaf tissues, as demonstrated by very similar values in the maximum quantum yield of photosystem II (FV/FM) and effective quantum yield of photosystem II [Y(II)] of young developing grapevine leaves, treated or not with 400 μM and 1 mM NoPv1, measured using the IMAGING-PAM (Walz, Germany) to evaluate their photosynthetic performance (Fig. 3 and Table S2).
Images of Pinot noir leaf disks co-inoculated ex vivo with Plasmopara viticola sporangia in the presence/absence of NoPv peptide aptamers. In the rectangle (a-b) is displayed the experimental setup: 5 leaf disks for each Petri dish and 5 droplets for each leaf disk of P. viticola sporangia suspension in the presence/absence of NoPv peptides. (a) P. viticola sporangia suspension was used to infect grapevine leaf disks, as control. (b) P. viticola sporangia suspension mixed with NoPv1 peptide (200 μM) was used to inoculate grapevine leaf disks as in (a). (c) Detail of a single leaf disk shown in (a) infected with P. viticola sporangia suspension, and used as control. P. viticola sporulation can be observed as white spots on leaf surface. (d) Detail of a single leaf disk shown in (b) infected with P. viticola sporangia in the presence of NoPv1 peptide (200 μM). No P. viticola sporulation can be observed on leaf disks, i.e. no white spots, pinpointing the anti-oomycete properties of NoPv1. (e) Single leaf disk infected with P. viticola sporangia in the presence of NoPv2 peptide (200 μM). White spots due to P. viticola sporulation are visible, indicating the poor anti-oomycete activity of NoPv2. (f) Single leaf disk infected with P. viticola sporangia in the presence of NoPv3 peptide (200 μM). White sectors due to P. viticola sporulation are visible, indicating the poor anti-oomycete activity of NoPv3. Images were taken at 7 dpi. (g) Quantitative evaluation of sporulation of P. viticola isolates on leaf disks. Disease severity was assessed at 5 and 7 days post-inoculation (dpi) as percentage of leaf disk area covered by P. viticola sporulation, calculated as sum of the disease severity of the five drops for each disk74. Each inoculated area was scored as surface with no sporulation (0%), scarce sporulation (10%) or fully covered by sporulation (20%). Each experiment was carried out twice. For each treatment, average and standard error values of ten replicates from the two experiments are shown. Different lowercase and uppercase letters indicate significant differences among treatments at 5 and 7 dpi respectively, according to a Kruskal–Wallis test (P ≤ 0.05).
Photosynthetic performance of grapevine leaves exposed ex vivo to 400 µM and 1 mM NoPv1 for 7 days to evaluate NoPv1 cytotoxicity. The maximum quantum yield (FV/FM) and the effective quantum yield [Y(II)] of photosystem II were measured with an imaging Chl fluorometer (Imaging PAM) after 7 days exposure to 400 µM (P1) and 1 mM NoPv1 (P2). As positive control, the same leaves were treated with 0.1% v/v (BH1) and 0.2% v/v (BH2) of BASTA herbicide. Treatment with water (M1 and M2) was used as negative control. Representative images from 10 biological replicates are shown. Herbicide treated spots showed a clear reduction of photosynthetic performance, even though no clear symptoms were visible in bright-field leaf images, highlighting the high sensitivity of the assay. Furthermore, NoPv1- and water-treated spots showed comparable photosynthetic values (see Table S2), indicating that the peptide aptamer is not cytotoxic for plant cells. See Table S2 for average and standard deviation values of photosynthetic parameters. Scale bar = 1 cm.
Properties of NoPv1
As shown in Table 1, NoPv1 is a peptide of 8 l-amino acid residues with the following sequence: RLTAQCRL. Its molecular weight is of 960.16 Da, the isoelectric point is at pH 10.43, it has a net charge of + 1.9 at pH 7 and a good water solubility (pepcalc.com). Interestingly, the net positive charge has been proposed to be optimal for antimicrobial peptides, since cationic peptides are thought to undertake electrostatic interactions with the negatively charged phospholipid membranes of bacteria, fungi and other microorganisms34,35,36,37.
To verify the importance of the two Arginine (R) residues for the biological activity of NoPv1, three alanine point mutated derivatives were prepared: NoPv1-R1A (ALTAQCRL), NoPv1-R7A (RLTAQCAL), and NoPv1-R1A-R7A (ALTAQCAL) (see also Table 1). Interestingly, all three NoPv1 derivatives were able to interact physically with PvCesA2, as shown by yeast two-hybrid assay (Fig. 4a), but unable to counteract P. viticola infection (Fig. 4b), pointing to the fundamental role of the net positive charge of NoPv1 for its biological activity.
Yeast two-hybrid interaction assay and antimicrobial activity of NoPv1 modified versions. (a) NoPv1-R1A, NoPv1-R7A and NoPv1-R1A-R7A were tested for their physical interaction with PvCesA2 enzyme. The unmodified version of NoPv1 was used as positive control in the assay. Interestingly, all the modified versions of NoPv1 interact with the PvCesA2. -W -L, permissive medium devoid of Tryptophan and Leucine; -W -L -H -A, selective medium devoid of Tryptophan, Leucine, Histidine and Alanine supplemented with 5 mM 3-AT (3-Amino-1,2,4-triazole, a histidine biosynthesis inhibitor). (b) The anti-oomycete activities of NoPv1-R1A, NoPv1R7A and NoPv1-R1A-R7A (200 μM) were tested by co-inoculating them with P. viticola sporangia on grapevine leaf disks and NoPv1 was used as positive control. Disease severity was evaluated at 5 and 7 days post inoculation (dpi). The experiment was carried out twice. For each treatment, average and standard error values of ten replicates from the two experiments are presented. Different lowercase and uppercase letters indicate significant differences among treatments at 5 and 7 dpi respectively, according to a Kruskal–Wallis test (P ≤ 0.05).
Antimicrobial activity and specificity of NoPv1 peptide aptamer
Properties of NoPv1 against P. viticola were further investigated on leaf disks and on potted plants grown in greenhouse. In particular, NoPv1 displayed a good inhibitory activity in the co-inoculation assay on leaf disks at concentrations equal or higher than 100 μM, where the percentage of leaf disk area covered by P. viticola sporulation was far below 10% at 5 and 7 dpi (Fig. 5a). On the contrary, lower concentrations were able to contrast only partially P. viticola on leaf disks, as disease severity was almost half of the control sample at 5 and 7 dpi in the case of 50 μM NoPv1 and no anti-oomycete activity was observed at 20 μM NoPv1 (Fig. 5a). In addition, nebulization of 400 μM NoPv1 on leaf disks performed using the Potter Precision Spray Tower, recognised as the standard of reference for chemical spraying techniques in the laboratory38, indicated a reduction of P. viticola severity at 7 dpi (Fig. 5b). Similarly, a solution of 800 μM NoPv1 resulted to be as effective as the copper-based fungicide Kocide 2000 (1.42 g l−1 active compound; see also Fig. 5b). Preliminary nebulization tests were also conducted on leaf disks to assess both the preventive and curative action of NoPv1. In particular, 400 μM NoPv1 was the lowest concentration that maximises the efficacy against P. viticola (see Fig. 5b) and it was sprayed on leaf disks at different times before [from 7 days (− 7 d) to 2 hours (− 2 h)] and after [from 1 hour (+ 1 h) to 2 days (2 d)] pathogen inoculation (Fig. 6a). The treatment efficacy was evaluated as a measure of the difference of disease severity between control (43.2 ± 4.4%; average ± standard error values) and treated (400 μM NoPv1) samples (see “Materials and methods” for further details) for each time point (Table S3). NoPv1 was effective in suppressing P. viticola infection even when applied 7 days before inoculation (-7 d), whereas its efficacy decreased very rapidly when applied after P. viticola inoculation, i.e. in post-inoculation treatments (Fig. 6a).
Biological activity of NoPv1 peptide aptamer against Plasmopara viticola on leaf disks. (a) The activity of NoPv1 peptide solubilised at different concentrations in water (20 μM, 50 μM, 100 μM and 200 μM) and co-inoculated with P. viticola sporangia suspension on grapevine leaf disks, as droplets, was monitored at 5 and 7 days post-inoculation (dpi), as percentage of leaf disk area covered by P. viticola sporulation, according to the protocol reported in Lazazzara et al.74. NoPv1 displayed a good anti-oomycete activity at concentrations equal or higher than 100 μM. (b) The anti-oomycete activity of NoPv1 was subsequently evaluated using a Potter Precision Spray Tower, employed to mimic in field spray conditions. Leaf disks were treated with NoPv1 at different concentrations, ranging from 100 to 800 μM, and its efficacy, measured as percentage of disease severity, was compared with a copper-based commercial fungicide, (Kocide 2000, copper hydroxide). Each experiment was carried out twice. For each treatment, average and standard error values of ten replicates from the two experiments are presented. Different lowercase and uppercase letters indicate significant differences among treatments at 5 and 7 dpi, respectively, according to a Kruskal–Wallis test (P ≤ 0.05).
Preventive and curative NoPv1 properties against Plasmopara viticola on leaf disks and greenhouse grown plants. (a) The anti-oomycete activity of NoPv1 (400 μM) was tested, using the Potter Precision Spray Tower, on leaf disks at different times [days (d)/hours (h)] before (from − 7 d to − 2 h) and after (+ 1 h to + 2 d) P. viticola inoculation. The co-inoculation assay (0 h) is also reported in the histogram. Treatment efficacy (see also “Materials and methods”) was evaluated at 7 dpi. Efficacy of NoPv1 was found in preventive treatments, but not in the case of curative treatments. (b) Biological activity of NoPv1 peptide on grapevine plants grown under greenhouse conditions. The NoPv1 peptide at 1 mM concentration was sprayed at 7 and 2 days before P. viticola inoculation (− 7 d and − 2 d) and co-inoculated (0 h) and the treatment efficacy was assessed at 7 dpi. Preventive treatments with the NoPv1 peptide control P. viticola infection under greenhouse conditions. Each experiment was carried out twice. For each treatment, average and standard error values of ten replicates from the two experiments are presented. Different lower case letters indicate significant differences among treatments, according to a Kruskal–Wallis test (P ≤ 0.05).
On the basis of the ex vivo results, the activity of NoPv1 against P. viticola was tested in vivo on grapevine plants grown under greenhouse conditions. Different NoPv1 concentrations were sprayed on grapevine plants immediately before P. viticola inoculation and the disease severity of plants treated with 400 μM (27.3 ± 5.2%), 800 μM (26.2 ± 4.2%), 1 mM (20.8 ± 2.4%) and 2 mM (21.1 ± 4.2) NoPv1 was lower than that of control plants (58.7 ± 6.5%; P < 0.05, Kruskal–Wallis test). In particular, 1 mM NoPv1 was the lowest concentration showing a P. viticola severity comparable to the copper treatment (5.4 ± 3.9%; P > 0.05, Kruskal–Wallis test) and it was further used in the following experiments. Therefore, 1 mM NoPv1 was applied at 7 and 2 days before inoculation (− 7 d and − 2 d) and immediately before (0 h) P. viticola inoculation to assess the persistence on grapevine plants. As shown in Fig. 6b, the NoPv1 efficacy was always higher than 60% in plants treated with NoPv1 compared to control plants (Fig. 6b), which showed a mean disease severity of 58.15 ± 6.6% (see Table S4).
NoPv1 specificity was further tested to verify potential harmful effects on non-target organisms. Coherently with the very low homology of CesA enzymes from Escherichia coli, Agrobacterium tumefaciens and Bacillus amyloliquefaciens with PvCesA2 (see also Fig. 1 and Data S1), NoPv1 peptide aptamer, provided directly in the growth medium at 100 μM and 200 μM, was not able to inhibit the growth of these bacteria, as shown by the very similar growth curves (Fig. 7a–c). Furthermore, the co-inoculation of NoPv1 at 200 μM and 400 μM with Erysiphe necator, an Ascomycota responsible for grapevine powdery mildew, was not able to reduce the disease severity on leaves (Fig. 7d), in agreement with the fact that the cell wall of E. necator, as other Ascomycota, is mainly formed by chitin and other types of glucans39, rather than cellulose. On the contrary, comparison of the PvCesA2 cytoplasmic domain used in the yeast two-hybrid screening with the orthologous P. infestans CesA2 protein (PiCesA2) showed 97% of amino acid residue identity (Data S1). As it could be hypothesised, NoPv1 was able to inhibit P. infestans growth in vitro at the same concentrations used against P. viticola, i.e. 100 μM and 200 μM (Fig. 8), however further experiments are needed before concluding that NoPv1 interacts with PiCesA2 and it is able to inhibit cellulose biosynthesis.
Biological activity of NoPv1 peptide on Escherichia coli, Agrobacterium tumefaciens, Bacillus amyloliquefaciens and Erysiphe necator. The NoPv1 peptide was employed to test its effects, at 100 μM and 200 μM, on the growth of E. coli (a), Agrobacterium tumefaciens (b) and Bacillus amyloliquefaciens (c) for 5 h. The growth curves are the average of three biological replicates. (d) NoPv1 (200–400 μM) was also co-inoculated with Erysiphe necator spores on young grapevine leaves to monitor its antifungal activity. The percentage of disease severity was evaluated at 14 dpi, using the protocol reported in “Materials and methods”. Overall, no inhibitory effects were found. The experiment in (d) was carried out twice, average and standard error values of ten replicates from the two experiments are shown. No significant differences were detected (P > 0.05), according to the Kruskal–Wallis test.
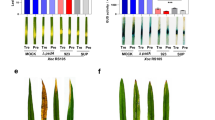
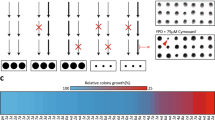
Anti-oomycete activity of NoPv1 on Phytophthora infestans. (a) The ability of NoPv1 to inhibit the mycelial growth of P. infestans at 20 °C was evaluated in vitro by adding 100 μM and 200 μM of peptide to the pea agar medium and measuring the radial growth at 4, 5 and 7 days post inoculation (dpi). (b) In representative images of P. infestans growth on Petri dishes at 7 dpi, the diameter of P. infestans mycelia is smaller in the presence of NoPv1 than under control conditions, indicating that NoPv1 is able to inhibit P. infestans growth. Each treatment was carried out on five petri-dishes (replicates), and experiments were carried out twice. Different lowercase, uppercase and italics letters indicate significant differences among treatments at 4, 5 and 7 dpi, respectively, according to a Kruskal–Wallis test (P ≤ 0.05).
Absence of potential cytotoxicity of NoPv1
To investigate further the potential cytotoxicity of NoPv1, human cells were used to perform an in vitro cell viability test, or MTT assay. Immortalized human cell cultures (HKC8) at different densities (1 K, 3 K and 6 K/100 μl) were grown in DMEM-F12 medium in the presence of 400 μM NoPv1 and their viability was measured after 24 and 48 h from the treatment. No significant differences between control and treated samples were observed after 24 h with respect to the level of absorbance at 570 nm, and cytotoxic effects could only be observed at 1 K cell density after 48 h, indicating that NoPv1 has no major deleterious effects on human cells (see Fig. 9).
Cell-viability assay to assess the potential cytotoxicity of NoPv1. Immortalized human cells (HKC8), grown in the DMEM-F12 medium at different densities (1 K, 3 K and 6 K cells/100 μl), were grown in the presence of 400 μM NoPv1 and in its absence (Control) for 24 and 48 h. The results demonstrate the absence of any major effect of NoPv1 on the viability of the cells, as shown by the almost identical absorbance levels of MTT-formazan at 570 nm, with the exception of 1 K cell density at 48 h post-treatment (hpt). For each treatment, average and standard error values of four biological replicates are shown. Statistical significance was determined by two-tailed paired Student’s t-test. Each treatment was considered individually and compared with the relative control. p-values are represented as *P ≤ 0.05.
NoPv1 blocks Plasmopara viticola germ tube formation
Leaf disks of V. vinifera cv. Pinot noir were inoculated with P. viticola zoospores in the absence (control, Fig. 10a–c) and in the presence (Fig. 10d–f) of 200 μM NoPv1. In control leaf disks, the biflagellate zoospores (n = 90, Fig. 10a) were able to reach the stomata localised on the abaxial leaf surface (Fig. 10b). There, they encysted either alone or in groups (up to 4 germinating cysts/stomata) and emitted a single germ tube (see arrow in Fig. 10b) for each zoospore that penetrated the stomata (Fig. 10b,c). In presence of 200 μM NoPv1, P. viticola zoospores were unable to develop any germ tube essential to penetrate the stomata (n = 160, Fig. 10d,e). In the very few cases (5 zoospores out of 160, 3.12%) where a germ tube primordium could be detected (see arrow in Fig. 10f), its orientation was opposite to the stomata localization (see asterisk in Fig. 10f), indicating that NoPv1 prevents germ tube formation and proper stomatal recognition (Fig. 10f).
Scanning Electron Microscopy (SEM) images of leaf disks infected with Plasmopara viticola zoospores in presence/absence of NoPv1. (a–c) Grapevine leaf disks infected with a P. viticola sporangial suspension (1 × 105 sporangia ml−1) were incubated for 6 h in the dark at 22 °C, (f, flagellum). (d–f) Identical grapevine leaf disks were infected with a P. viticola sporangial suspension (1 × 105 sporangia ml−1) in presence of 200 μM NoPv1 and incubated for 6 h in the dark at 22 °C. All leaf disks were then fixed, covered with gold and observed with a LEO 1430 scanning electron microscope. NoPv1-treated zoospores were unable to develop a proper germ tube and to penetrate the stomatal cavity of grapevine leaf disks. In (f), the black arrow indicates a germ tube primordium that points in the opposite direction with respect to the nearest stomata (asterisk), suggesting that NoPv1-treated zoospores are also unable to sense stomata proximity. In total, about 250 zoospores with and without NoPv1 were observed. The analysis was carried out twice, using 12 independent leaf disks.
Discussion
The need of alternatives to traditional pesticides
Plasmopara viticola is a relevant plant pathogen which requires control measures to avoid severe yield losses. Researchers have dedicated many energies to identify host resistance genes to the pathogen attacks, and a number of breeding programs have introgressed resistance loci from wild North American and Asian Vitis species into V. vinifera resulting in new downy mildew resistant grapevine cultivars40,41. However, most of the European cultivars carry a single major resistance locus named Rpv3 (Resistance to P. viticola), and P. viticola isolates able to overcome this resistance have arisen in Europe42,43,44,45. Because of that and due to the long time required for conventional breeding46, wine production is heavily dependent on the use of pesticides to control this disease. Nevertheless, strong selection pressure following repeated pesticide applications has led to the development of resistant pathogen populations, which has limited the success of chemical pesticides in grapevine and other crops47,48,49,50.
In this scenario, it is important to develop new tools that allow to easily and quickly isolate molecules, intrinsically less dangerous for consumers, farmers and the environment, able to counteract emerging pathogens. Here, we reported the development of an in vivo strategy aimed to isolate synthetic antimicrobial peptide aptamers from a combinatorial library. These synthetic peptide aptamers share several features, including size, level of hydrophobicity and net charge, with the antimicrobial peptides (AMPs), synthesised by the innate immune system of various species including human, animals and plants, which are the first-line defence against foreign attacks51,52. This strategy allowed the isolation of the 8 a.a. NoPv1 peptide aptamer able to counteract P. viticola and P. infestans infections, but it might have a much broader impact in the long term by being applicable to other species and type of crop pathogens, i.e. viruses and bacteria.
The mechanism of action of NoPv1
NoPv1 inhibits zoospore germ tube formation (see Fig. 10) that requires active cellulose biosynthesis, as reported in P. infestans where the cellulose synthases were shown to be localised and play a major role in the growing tip of the germ tube28, suggesting a possible inhibitory role of NoPv1 on PvCesA2. This notion seems to be supported by the capability of NoPv1 to affect severely P. infestans growth, given that its CesA2 enzyme shares high amino acid sequence identity (97%) with the PvCesA2 (see Fig. 8) and by the failure of NoPv1 to inhibit the growth of bacteria, such as E. coli, A. tumefaciens and B. amyloliquefaciens and to counteract the leaf infection caused by the Ascomycota E. necator (see Fig. 7). However, further interaction analyses and in vitro enzymatic assays are needed before concluding that NoPv1 is a specific inhibitor of PvCesA2 and PiCesA2 enzymatic activity. The selectivity of NoPv1 for certain microorganisms might suggest that NoPv1, with two positively charged “R” residues, does not behave as a typical broad spectrum cationic AMP able to alter membrane permeability by simply inserting into the lipid membranes and forming ion channels or pores that eventually result in leakage of cell contents and cell death53,54,55,56. However, the role of NoPv1 on cell membrane permeabilization remains to be investigated. The replacement of “R” residues with “A” in NoPv1-R1A, NoPv1-R7A and NoPv1-R1A-R7A (see Table 1) abolished the antimicrobial activity (Fig. 4b), but not the physical interaction with PvCesA2 (see Fig. 4a), suggesting that the two cationic residues might be essential for NoPv1 cellular uptake by P. viticola, thus reaching the PvCesA2 catalytically active portion, which localizes intracellularly. This is in agreement with the mechanism of action of some cationic AMPs, reported to penetrate cells and affect the cellular physiological processes, without altering the permeability of microbial membranes57,58. For instance, the natural echinocadin AMP family targets specifically 1,3 β glucan synthase, an enzyme essential for cell wall integrity of fungi59. Furthermore, chitin biosynthesis is blocked by nikkomycins which are the most widely studied peptidyl nucleoside inhibitors of chitin synthases60,61. Alternatively, NoPv1 biological activity could strictly rely on the presence of “R” residues without necessarily interacting with the PvCesA2 enzyme.
The safety and sustainability of NoPv1 potential application under field conditions
Beside target specificity, novel drugs must have a broad-spectrum activity, low toxicity and ideally no off-target organisms. These properties are generally used as guidelines in drug discovery and might give the impression that such molecules are unattainable. The selection of PvCesA2 as a target for peptide aptamers makes NoPv1 both specific for the target and, concomitantly, able to act on a broad spectrum of oomycete pathogens since their cellulose synthases group together in a clade separated from bacteria, viral and plant cellulose synthases (see Fig. 1). As a consequence, NoPv1 is able to inhibit the growth of P. infestans (see Fig. 8), an oomycete that causes the serious potato and tomato disease known as late blight or potato blight, but at the same time it does not affect any off-target organism among the ones tested, including plants (see Figs. 3 and 7). Furthermore, the absence of any major cytotoxicity on mammalian cells (see Fig. 9), likely due to the lack of cellulose synthase enzymes and the neutral net charge of cell membranes35,62 confers to NoPv1 almost all the desirable features required for an anti-oomycete active compound. Nevertheless, the relatively high concentration of NoPv1 peptide, around 1 mM, shown to have efficient antimicrobial activity under greenhouse conditions seriously challenges its employment under field conditions, mainly as a consequence of the elevated costs of production that are certainly not competitive with the conventional pesticides. However, lower NoPv1 concentrations showed high efficacy on leaf disk assays (e.g. 100 µM and 400 µM in the co-inoculation and nebulization experiment, respectively), suggesting that appropriated formulations could improve NoPv1 stability under field conditions and possibly reduce the risk of degradation by UV light and/or leaf-associated microorganisms. Moreover, a lot can be learnt from the biomedical sector, where recent publications report the successful employment of antifungal peptides in combination with conventional antifungal drugs63. Similarly, NoPv1 could be sprayed in combination with conventional active compounds under field conditions, resulting in the reduction of peptide dose, i.e. reduced treatment costs, and in the prevention of drug resistance development as a direct consequence of the mechanistic polyfunctionality improvement of the active compounds. Furthermore, the design of novel delivery systems aimed to improve safety and bioavailability64, in combination with precision agriculture tools, may contribute to make the peptide-based pesticides sustainable for the agricultural sector in future.
Conclusions
We have described a novel approach to identify active compounds to counteract P. viticola infection and proposed that this strategy could be applicable to other crop pathogens. In this context, NoPv1 is reported as a highly effective, relatively safe peptide to be used as potential anti-oomycete agent. Despite these features, it must be observed that only a very limited number of antimicrobial peptide-based drugs, around 9, have been approved by the Food and Drug Administration, so far56. Based on that, it is reasonable to think that NoPv1 must be subjected to major improvements before its use for agricultural purposes. Obviously, the stability of NoPv1 must be verified at high temperature, in the presence of proteases or at different pH in order to be used under field conditions. However, unfavourable properties limiting peptide usage could be overcome by chemical optimization. For instance, synthetic peptides can be stabilised once chemically modified. Some of them have been reported to keep biological activity at high temperature, and some can resist to hydrolysis mediated by proteases, like tripsin and pepsin54. Moreover, the wide variety of structural and functional features identified for various natural antimicrobial peptides represent an invaluable source of ideas for adapting peptides to our needs, providing promising perspectives to the identification of safer alternatives to conventional pesticides. Finally, a cost-efficient peptide manufacturing technology needs to be set-up to compete with the production costs of conventional pesticides. Indeed, the peptide chemical synthesis is not applicable to agriculture purposes due to the high costs of production, however several companies in recent years offer cost-efficient fermentative bioprocesses, up to ton scales, for the production of recombinant peptides.
Materials and methods
Phylogenetic and amino acid sequence analysis
CesA and Csl amino acid sequences were obtained from UniprotKB database (uniprot.org), except for KJD55249.1 (BaCesA) which was obtained from the National Center for Biotechnology Information database (ncbi.nlm.nih.gov). Multiple sequence alignment was generated with Clustal omega (ebi.ac.uk/Tools/msa/clustalo)65. The dendrogram was constructed with a Minimum-Evolution algorithm and 1000 bootstrap repetitions using MEGA X (megasoftware.net)66. Transmembrane regions were predicted with SOSUI (harrier.nagahama-i-bio.ac.jp/sosui)67 and domain prediction was performed with CD-BLAST (ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi)68.
Plant material and phytopathogens
Two year-old plants of the susceptible V. vinifera cv. Pinot noir were grown under greenhouse conditions at 25 ± 1 °C, with a photoperiod of 16 h light and a relative humidity of 70 ± 10%, as described by Perazzolli et al.69. A P. viticola population was collected in an untreated vineyard sited in the Trentino region (Northern Italy) and maintained by weekly inoculations on potted Pinot noir plants grown in greenhouse as described by Perazzolli et al.69. To obtain P. viticola inoculum, plants with disease symptoms were incubated overnight in the dark at 99–100% RH and 25 ± 1 °C to promote pathogen sporulation. Sporangia were collected by washing the abaxial surfaces bearing freshly sporulating lesions with cold (4 °C) distilled water and the concentration of the inoculum suspension was adjusted to 1 × 105 sporangia ml−1 by counting with a haemocytometer under a light microscope. The P. infestans isolate (kindly provided by M. Finckh and A. Butz, University of Kassel, Germany) was grown on pea agar medium (PAM, 12.5% w/v frozen peas in distilled water and 1.2% w/v bacteriological agar) at 20 °C, as reported by Tomada et al.70. Infected leaves of greenhouse-grown plants were used as a source of the E. necator inoculum.
RNA extraction and cDNA synthesis
P. viticola sporangia were collected by washing the abaxial surfaces of grapevine leaves bearing freshly sporulating lesions with cold (4 °C) distilled water. After centrifugation at 4000×g and 4 °C, the pellet (0.05 g) was immediately frozen in N2-liquid and stored at − 80 °C. Total RNA was extracted using the Spectrum Plant total RNA kit (Sigma-Aldrich, St. Louis, MO), quantified using the Nanodrop 8000 (Thermo Fisher Scientific, Wilmington, DE) and its quality was checked by agarose gel electrophoresis. RNA was treated with DNase I (Invitrogen, Thermo Fisher Scientific), and first-strand cDNA was synthesised from 500 ng of total RNA using ImProm-II Reverse Transcription System Kit (Promega Corporation, Madison, WI, USA) with a combination of oligo (dT) primers and random hexamers.
Gene amplification and cloning
The gene fragment encoding the PvCesA2 cytoplasmic portion was amplified from the cDNA template with Phusion High-Fidelity DNA Polymerase (Thermo Fisher Scientific), according to manufacturer’s instruction, using primers GrA007 (5′-GGGGACAAGTTTGTACAAAAAAGCAGGCTTCGACGAGTTTGAGCCGCC-3′) and GrA008 (5′-GGGGACCACTTTGTACAAGAAAGCTGGGTCTACCACTCGGGGTCAAAATATTGG -3′), which contain attB sites, and cloned in the Gateway-compatible variant of pGBKT7 (kindly provided by Brendan Davies) passing through pDONR207 (Thermo Fisher Scientific).
NoPv1 mutated derivatives (R residues replaced with A residues) for the two-hybrid experiments were cloned in the following way: the primers GrA_045 (5′-AGTGGATCCAAGCGCTGACGGCGCAGTGTCGTCTTAAGGGGCCCAAAATG-3′) and GrA_046 (5′-CATTTTGGGCCCCTTAAGACGACACTGCGCCGTCAGCGCTTGGATCCACT- 3′) for NoPv1-R1A; GrA_047 (5′-AGTGGATCCAACGTCTGACGGCGCAGTGTGCGCTTAAGGGGCCCAAAATG-3′) and GrA_048 (5′-CATTTTGGGCCCCTTAAGCGCACACTGCGCCGTCAGACGTTGGATCCACT-3′) for NoPv1-R7A; GrA_049 (5′-AGTGGATCCAAGCGCTGACGGCGCAGTGTGCGCTTAAGGGGCCCAAAATG-3′) and GrA_050 (5′-CATTTTGGGCCCCTTAAGCGCACACTGCGCCGTCAGCGCTTGGATCCACT-3′) for NoPv1-R1A-R7A. All primers contain complementary regions and flanking 5′-BamHI and 3′-ApaI restriction sites. After self-annealing, the DNA fragments were digested with BamHI and ApaI restriction enzymes (Fastdigest, Thermo Fisher Scientific), dephosphorylated with FastAP Thermosensitive Alkaline Phosphatase (Thermo Fisher Scientific) and finally ligated into the plasmid pLib2, previously digested with BamHI and ApaI.
Construction of the peptide aptamer library
The peptide aptamer library was constructed according to Reverdatto et al.31 with modifications. The oligonucleotide 5′-GGCAGAGTGGATCCAA(NNK)8AAGGGGCCCCTT-3′, which contains complementary regions and flanking 5′-BamHI and 3′-ApaI restriction sites, was self-annealed and the 5′ and 3′ single-stranded ends were filled-in by using the Klenow Fragment (Thermo Fisher Scientific). Subsequently, the DNA fragment was digested with BamHI and ApaI restriction enzymes (Fastdigest, Thermo Fisher Scientific), dephosphorylated with FastAP Thermosensitive Alkaline Phosphatase (Thermo Fisher Scientific) and finally ligated into the plasmid pLib2 (kindly provided by Alexander Shekhtman). The library was then amplified in E. coli DH5α competent cells. About 2.5 × 106 colonies were collected with LB medium containing ampicillin and let grow for about 4 h. Plasmids were then purified by using the Qiagen Maxi prep kit.
Yeast two-hybrid (Y2H) library screening and peptide aptamer identification
The aptamer library was transformed into the yeast strain Saccharomyces cerevisiae AH109 (Clontech Laboratories, Palo Alto, CA, USA) harbouring the bait-containing plasmid pGBKT7-PvCesA271. Transformants were selected for growth on selective media without tryptophan, leucine, and either adenine or histidine supplemented with 10 mM 3-AT (3-Amino-1,2,4-triazole, a histidine biosynthesis inhibitor). Yeast plasmid DNA was purified from the positive colonies, electroporated into E. coli cells, and the plasmids isolated and sequenced using the GrA_019 primer (5′-TCCAAGCTTTGCAAAGATGG-3′).
Peptide synthesis and purification
The peptides identified by the Y2H strategy and the NoPv1 mutated derivatives were initially prepared by microwave assisted solid phase synthesis, based on Fmoc chemistry on pre-loaded Wang resin (0.4 meq/g substitution) using a fivefold molar excess of 0.2 M Fmoc-protected amino acids dissolved in NMP and using HOBT/HBTU/DIEA (5:5:10) as activators72,73). Coupling reactions were performed for 5 min at 40 W with a maximum temperature of 75 °C. De-protection was performed in two stages using 20% v/v piperidine in DMF (5 min and 10 min each). The cleavage from the resin was performed using 10 ml of Reagent K (TFA/phenol/water/thioanisole/EDT; 82.5/5/5/5/2.5) for 180 min. After cleavage, peptides were precipitated out and washed using ice-cold anhydrous ethyl ether. All peptides were purified by RP-HPLC using a gradient elution of 5–70% v/v solvent B (solvent A: water/acetonitrile/TFA 95/5/0.1; solvent B: water/acetonitrile/TFA 5/95/0.1) over 20 min at a flow rate of 20 ml/min. The purified peptides were freeze-dried and stored at 0 °C. The peptides purity was > 95% and was determined using analytical HPLC (95% v/v A for 5 min; 95–30% v/v A over 20 min) high resolution mass spectrometry (HRMS) and NMR (see Data S2). Peptide identity was also confirmed by electrospray ionization mass spectrometry (ESI–MS). NoPv1 peptide was later purchased at > 95% purity from Bio-Fab Research (Rome, Italy). All the lyophilized peptides and mocks were prepared using the same solvent, either distilled water or 5% v/v DMSO, according to the overall solubility of used peptides.
Activity of peptide aptamers against Plasmopara viticola on leaf disks
Leaves (from the fourth-sixth node) were collected from V. vinifera cv. Pinot noir plants grown under greenhouse conditions and surface sterilised by immerging them for 2 min in 1% v/v sodium hypochlorite solution. After that, they were rinsed three times with water and dried on paper. Leaf disks (18 mm diameter) were cut using a cork-borer and placed, abaxial side up, in 9 cm diameter plastic Petri dishes (five disks for each dish), containing four filter papers moistened with 4 ml of sterilised distilled water42. Each leaf disk was inoculated with five drops (10 μl) of a P. viticola sporangia suspension (1 × 105 sporangia ml−1) mixed with the respective peptide aptamer at the appropriate concentration. Dishes were sealed with a plastic film and incubated overnight (16 h) in a growth chamber at 22 ± 1° C in the dark, then dried with filter paper and incubated for 7 days in a growth chamber at 22 ± 1° C and 16 h photoperiod. Disease severity was assessed at 5 and 7 days post inoculation (dpi) as percentage of leaf disk area covered by P. viticola sporulation, calculated as sum of the disease severity of the five drops for each disk74. Each inoculum drop was scored as surface with no sporulation (0%), scarce sporulation (10%) or fully covered by sporulation (20%). Five replicates (dishes with five disks each) were assessed for each treatment and the experiment was carried out twice.
To analyse the peptide aptamer effects by spray nebulization, Petri dishes (with five leaf disks each) were sprayed using a Potter Precision Spray Tower (Burkard Scientific Co., Uxbridge, UK) with 1.67 ml of the peptide aptamer solution (corresponding to the standard dosage of 10 hl ha−1 in a vineyard with a ‘Pergola trentina’ training system) at a pressure of 55 kPa. As control treatments, Petri dishes were sprayed with water (mock) or with copper hydroxide (1.42 g l−1 Kocide 2000; Du Pont, Wilmington, DE, USA) as a reference fungicide. Petri dishes were incubated under a laminar flow hood for 5–10 min in order to dry the leaf disks, sprayed with a fresh P. viticola sporangia suspension using a small hand sprayer device (0.6 ml per dish). Afterwards, Petri dishes were incubated overnight in the dark at 22 ± 1 °C in a growth chamber, then dried under a laminar flow hood and incubated for 7 days in a growth chamber, as described above. Disease severity was assessed visually as percentage of leaf disk area covered by P. viticola sporulation at 5 and 7 dpi75. The disease reduction (efficacy) was calculated according to the following formula76: (disease severity of mock disks—disease severity of treated disks) / (disease severity of mock disks) × 100. Five replicates (Petri dishes with five leaf disks each) were assessed for each treatment and the experiment was carried out twice. To assess the persistence, as well as the preventive and curative action of the peptide aptamers, treatments were applied on the leaf disks at different times (from 7 days to 2 h) before pathogen inoculation. After each application, leaf disks were dried under a laminar flow hood for 5–10 min and then kept in the growth chamber. In the curative treatment the peptide aptamer was applied after pathogen inoculation (0, 1, 24, 48 h post infection).
Activity of peptide aptamers against Plasmopara viticola on greenhouse-grown plants
Vitis vinifera cv. Pinot noir plants were grown in a greenhouse for 2 months as described by Perazzolli et al.69. Peptide aptamer was applied with a compressed-air hand sprayer to the abaxial and adaxial surfaces of all leaves (20 to 30 ml, depending on the number of leaves) at 7 days before inoculation, 2 and 0 (co-inoculation), whereas not treated plants were used as control. A fresh sporangia suspension (1 × 105 sporangia ml−1) was applied to the abaxial leaf surface using a compressed air hand sprayer (20–30 mL per plant). Inoculated plants were incubated overnight in the dark at 25 ± 1 °C with 99–100% RH. Six days after inoculation, plants were incubated overnight at 25 ± 1 °C with 99–100% RH to promote P. viticola sporulation and the disease severity was assessed visually as percentage of abaxial leaf area covered by sporulation in relation to the total leaf area according to EPPO standard guidelines75. Five replicates (plants) were assessed for each treatment and time point, and the experiment was carried out twice.
Antimicrobial activity of peptide aptamers on bacteria
The antibacterial activity of NoPv1 was evaluated in liquid culture using three different bacterial species, Escherichia coli, Agrobacterium tumefaciens and Bacillus amyloliquefaciens (subsp. plantarum) strain D 747. Overnight pre-cultures were diluted to OD600 of 0.1 (E. coli; B. amyloliquefaciens) or 0.2 (A. tumefaciens) into sterile LB medium. NoPv1 (100 and 200 µM) was then added to the liquid medium of three biological replicates. Bacteria were grown at 28 °C (A. tumefaciens) or 37 °C (E. coli; B. amyloliquefaciens). The density of the cell population was measured spectrophotometrically at OD600 every hour, for 5 h.
Activity of peptide aptamers against Phytophthora infestans
The activity of NoPv1 against P. infestans was evaluated on Petri dishes with Pea Agar Medium (PAM) supplemented with 100 µM and 200 µM NoPv1. Mycelium plugs (7 mm in diameter) were cut from 7-day old colonies and placed upside down at the centre of the Petri dish. Petri dishes containing only PAM, without NoPv1, were used as control. Dishes were sealed with a plastic film and incubated in the dark at 20 ± 1 °C. The diameter of P. infestans colonies was assessed at 4, 5 and 7 dpi. Five replicates (petri dishes) were analysed for each treatment and the experiment was carried out twice.
Antifungal activity of peptide aptamers against Erysiphe necator
Evaluation of the ability of NoPv1 to protect grapevine plants against E. necator was carried out using the method described by Miclot et al.77 with minor modifications. The second and third fully expanded leaves from the apex were collected from different V. vinifera cv. Pinot noir plants grown under greenhouse condition and surface sterilised by washing for 2 min in a 1% v/v sodium hypochlorite solution. Leaves were rinsed three times for 2 min with distilled water and briefly dried on paper. The base of the petiole was cut off and leaves were placed, adaxial side up, in square culture dishes (120 × 120 mm), containing 1% w/v agar covered with sterile filter paper (1 layer, moistened with 2 ml of sterile distilled water). A small cut was performed with a sterile scalpel on the paper and the petiole was inserted. Four leaves were placed in each culture dish. NoPv1 solution or water (mock) were sprayed on the leaves inside a laminar flow (1–2 ml for each dish) using a small air hand sprayer. Leaves were left to air-dry for 20 min. Pathogen inoculation was performed inside the laminar flow using the dry inoculation method. The open dishes containing the leaves were placed inside a settling tower, a home-made plastic box of 26 × 26 cm base and 42 cm high. Conidia were blown from heavily infected grapevine leaves at the top of the settling tower using an aquarium air pump (Newa Wind NW2 Air Pump, NEWA Tecno Industria, Loreggia, Italy) connected to a Pasteur pipette. Three infected leaves were used as source of inoculum for each round of inoculation. The dishes were then placed in a growth chamber at 22 ± 1 °C, 70% RH and 16 h photoperiod. At 14 dpi, the percentage of disk area covered (% of disease severity) by E. necator sporulation was visually assessed, according to the EPPO standard guidelines78. Sixteen replicates (leaves) were used for each treatment and the experiment was carried out twice.
Cell viability and phytotoxicity assays
NoPv1 potential cytotoxicity was tested on SV40-Immortalised human fibroblasts, HKC8, using the MTT assay. This colorimetric assay is dependent on mitochondrial respiration and serves to assess the metabolic activity of a cell. In particular, it measures the activity of the mitochondrial succinate dehydrogenase, active only in living cells, that is capable of reducing MTT (3-[4,5-dimethylthiazol-2-yl]-2,5-diphenyltetrazolium bromide, yellow) to MTT-formazan, a blue/purple substance, the formation of which can be followed by measuring absorption at 570 nm. Therefore, the levels of MTT-formazan and the absorbance at 570 nm are directly proportional to the amount of viable cells present in the sample. Concerning NoPv1 cytotoxicity, HKC8 fibroblasts cultured in DMEM-F12 medium at different cell densities (1 K, 3 K and 6 K cells/100 μl) were exposed to 400 μM NoPv1 for 24 and 48 h.
Potential phytotoxicity of NoPv1 was monitored by estimating the photosynthetic performance of Pinot noir leaves [Maximum quantum yield—FV/FM—and effective quantum yield—Y(II)—of photosystem II] incubated for 7 days with 400 μM and 1 mM NoPv1 at room temperature, using the Imaging PAM (WALZ), as similarly reported in Perreault et al.79 and Schreiber et al.80. As positive control, 0.1% v/v and 0.2% v/v of BASTA herbicide were used81. Mock treatments were performed using water. In total, ten independent leaves were treated and analysed.
Scanning electron microscopy (SEM) analysis
Leaf disks infected with a P. viticola sporangia suspension (1 × 105 sporangia ml−1) with and without NoPv1 (200 µM) were fixed for 6 h in 50% v/v ethanol, 5% v/v acetic acid and 3.7% v/v formaldehyde in 0.025 M phosphate buffer, pH 7.0. Samples were collected soon after the infection and at 6 h after the infection. Samples were subsequently washed twice (about 20 min) in 70% v/v ethanol in 0.025 M phosphate buffer, pH 7.0. The material was dehydrated gradually in ethanol series to 100% v/v ethanol, dried in liquid carbon dioxide at the critical point. Samples were subsequently covered with gold using a sputter coater (SEMPREP2; Nanotech, Manchester, UK) and observed with a LEO 1430 scanning electron microscope (LEO Electron Microscopy, leo-usa.com).
Statistical analysis
Each experiment was carried out at least twice using independent biological samples. Statistical significance of cell viability assay was determined by two-tailed paired Student’s t-test. Different experimental setups were analysed singularly and compared with the relative control. For all the other experiments, data were analysed using the Statistica 13.1 software (TIBCO Software Inc., Tulsa, OK, USA) and a Kruskal–Wallis test was used to demonstrate equivalent results in the two experiments (P > 0.05, non-significant differences between experimental repetitions). Data from the two experimental repetitions were pooled and a Kruskal–Wallis test was then used to detect significant differences among treatments (P ≤ 0.05).
Patent PCT/IB2018/059834
The amino acid sequence of the NoPv1 peptide and its antimicrobial properties against Plasmopora viticola are described in the International Application No. PCT/IB2018/059834; Publication Number WO/2019/116203; Publication Date 20.06.2019.
References
Flood, J. The importance of plant health to food security. Food Secur. 2, 215–231 (2010).
Oerke, E. C. Crop losses to pests. J. Agric. Sci. 144, 31–43 (2006).
Savary, S. et al. The global burden of pathogens and pests on major food crops. Nat. Ecol. Evol. 3, 430–439 (2019).
FRAC Code List ©* 2018: Fungicides sorted by mode of action (including FRAC Code numbering). (2018).
Aktar, M. W., Sengupta, D. & Chowdhury, A. Impact of pesticides use in agriculture: Their benefits and hazards. Interdiscip. Toxicol. 2, 1–12 (2009).
Geiger, F. et al. Persistent negative effects of pesticides on biodiversity and biological control potential on European farmland. Basic Appl. Ecol. 11, 97–105 (2010).
Roberts, J. R. & Karr, C. J. Pesticide exposure in children. Pediatrics 130, e1765–e1788 (2012).
Mascarelli, A. Growing up with pesticides. Science 341, 740–741 (2013).
Carvalho, F. P. Pesticides, environment, and food safety. Food Energy Secur. 6, 48–60 (2017).
Kim, K.-H., Kabir, E. & Jahan, S. A. Exposure to pesticides and the associated human health effects. Sci. Total Environ. 575, 525–535 (2017).
Mostafalou, S. & Abdollahi, M. Pesticides: An update of human exposure and toxicity. Arch. Toxicol. 91, 549–599 (2017).
Nicolopoulou-Stamati, P., Maipas, S., Kotampasi, C., Stamatis, P. & Hens, L. Chemical pesticides and human health: The urgent need for a new concept in agriculture. Front. Public Health 4, 148 (2016).
Cabras, P. & Angioni, A. Pesticide residues in grapes, wine, and their processing products. J. Agric. Food Chem. 48, 967–973 (2000).
Nag, S. K. & Raikwar, M. K. Persistent organochlorine pesticide residues in animal feed. Environ. Monit. Assess. 174, 327–335 (2011).
Nougadère, A. et al. Total diet study on pesticide residues in France: Levels in food as consumed and chronic dietary risk to consumers. Environ. Int. 45, 135–150 (2012).
Kortenkamp, A. T. years of mixing cocktails: A review of combination effects of endocrine-disrupting chemicals. Environ. Health Perspect. 115, 98–105 (2007).
Vidaver, A. K. Uses of antimicrobials in plant agriculture. Clin. Infect. Dis. 34, S107–S110 (2002).
Colombo, M., Mizzotti, C., Masiero, S., Kater, M. M. & Pesaresi, P. Peptide aptamers: The versatile role of specific protein function inhibitors in plant biotechnology. J. Integr. Plant Biol. 57, 892–901 (2015).
Xu, Q., Yan, X., Zhang, Y. & Wu, J. Current understanding of transport and bioavailability of bioactive peptides derived from dairy proteins: A review. Int. J. Food Sci. Technol. 54, 1930–1941 (2019).
Buonassisi, D. et al. Breeding for grapevine downy mildew resistance: A review of “omics” approaches. Euphytica 213, 1–21 (2017).
Armijo, G. et al. Grapevine pathogenic microorganisms: Understanding infection strategies and host response scenarios. Front. Plant Sci. 7, 382 (2016).
Gessler, C., Pertot, I. & Perazzolli, M. Plasmopara viticola: A review of knowledge on downy mildew of grapevine and effective disease management. Phytopathol. Mediterr. 50, 3–44 (2011).
Agrios, G. Plant pathology: Fifth edition. 9780080473, (Academic Press, 2005).
Toffolatti, S. L. et al. A time-course investigation of resistance to the carboxylic acid amide mandipropamid in field populations of Plasmopara viticola treated with anti-resistance strategies. Pest Manag. Sci. 74, 2822–2834 (2018).
Unger, S., Büche, C., Boso, S. & Kassemeyer, H. H. The course of colonization of two different Vitis genotypes by Plasmopara viticola indicates compatible and incompatible host-pathogen interactions. Phytopathology 97, 780–786 (2007).
Kiefer, B., Riemann, M., Büche, C., Kassemeyer, H. H. & Nick, P. The host guides morphogenesis and stomatal targeting in the grapevine pathogen Plasmopara viticola. Planta 215, 387–393 (2002).
Mélida, H., Sandoval-Sierra, J. V., Diéguez-Uribeondo, J. & Bulone, V. Analyses of extracellular carbohydrates in oomycetes unveil the existence of three different cell wall types. Eukaryot. Cell 12, 194–203 (2013).
Grenville-Briggs, L. J. et al. Cellulose synthesis in Phytophthora infestans is required for normal appressorium formation and successful infection of potato. Plant Cell 20, 720–738 (2008).
Blum, M. et al. Mandipropamid targets the cellulose synthase-like PiCesA3 to inhibit cell wall biosynthesis in the oomycete plant pathogen, Phytophthora infestans. Mol. Plant Pathol. 11, 227–243 (2010).
Blum, M., Waldner, M. & Gisi, U. A single point mutation in the novel PvCesA3 gene confers resistance to the carboxylic acid amide fungicide mandipropamid in Plasmopara viticola. Fungal Genet. Biol. 47, 499–510 (2010).
Reverdatto, S. et al. Combinatorial library of improved peptide aptamers, CLIPs to inhibit RAGE signal transduction in mammalian cells. PLoS ONE 8, e65180 (2013).
Fugelstad, J. et al. Functional characterization of the pleckstrin homology domain of a cellulose synthase from the oomycete Saprolegnia monoica. Biochem. Biophys. Res. Commun. 417, 1248–1253 (2012).
Fugelstad, J. et al. Identification of the cellulose synthase genes from the oomycete Saprolegnia monoica and effect of cellulose synthesis inhibitors on gene expression and enzyme activity. Fungal Genet. Biol. 46, 759–767 (2009).
Montesinos, E. & Bardají, E. Synthetic antimicrobial peptides as agricultural pesticides for plant-disease control. Chem. Biodivers. 5, 1225–1237 (2008).
Bahar, A. A. & Ren, D. Antimicrobial peptides. Pharmaceuticals 6, 1543–1575 (2013).
Yeaman, M. R. & Yount, N. Y. Mechanisms of antimicrobial peptide action and resistance. Pharmacol. Rev. 55, 27–55 (2003).
Brown, K. L. & Hancock, R. E. W. Cationic host defense (antimicrobial) peptides. Curr. Opin. Immunol. 18, 24–30 (2006).
Paramasivam, M. & Selvi, C. Laboratory bioassay methods to assess the insecticide toxicity against insect pests. J. Entomol. Zool. Stud. 5, 1441–1445 (2017).
Barr, M. E. Ascomycota. in Systematics and Evolution 161–177 (Springer, Berlin, 2001). https://doi.org/10.1007/978-3-662-10376-0_8
Eibach, R., Zyprian, E., Welter, L. & Töpfer, R. The use of molecular markers for pyramiding resistance genes in grapevine breeding. VITIS J. Grapevine Res. 46, 120–120 (2007).
Merdinoglu, D. et al. Breeding for durable resistance to downy and powdery mildew in grapevine. Acta Hortic. 1046, 65–72 (2014).
Peressotti, E. et al. Breakdown of resistance to grapevine downy mildew upon limited deployment of a resistant variety. BMC Plant Biol. 10, 1–11 (2010).
Venuti, S. et al. Historical introgression of the downy mildew resistance gene Rpv12 from the Asian species Vitis amurensis into grapevine varieties. PLoS ONE 8, e61228 (2013).
Delmotte, F. et al. Rapid and multiregional adaptation to host partial resistance in a plant pathogenic oomycete: Evidence from European populations of Plasmopara viticola, the causal agent of grapevine downy mildew. Infect. Genet. Evol. 27, 500–508 (2014).
Toffolatti, S. L., Venturini, G., Maffi, D. & Vercesi, A. Phenotypic and histochemical traits of the interaction between Plasmopara viticola and resistant or susceptible grapevine varieties. BMC Plant Biol. 12, 124 (2012).
Morales, F. J. Conventional breeding for resistance to bemisia tabaci-transmitted geminiviruses. in Crop Protection, Vol. 20, 825–834 (Elsevier, Amsterdam, 2001).
Gisi, U., Sierotzki, H., Cook, A. & McCaffery, A. Mechanisms influencing the evolution of resistance to Qo inhibitor fungicides. in Pest Management Science, Vol. 58, 859–867 (Wiley, New York, 2002).
Grasso, V., Palermo, S., Sierotzki, H., Garibaldi, A. & Gisi, U. Cytochrome b gene structure and consequences for resistance to Qo inhibitor fungicides in plant pathogens. Pest Manag. Sci. 62, 465–472 (2006).
Heaney, S. P., Hall, A. A., Davies, S. A. & Olaya, G. Resistance to fungicides in the Qol-STAR cross-resistance group: current perspectives. BCPC Conf. Pests Dis. Vol. 2. Proc. an Int. Conf. held Bright. Hilt. Metrop. Hotel. Bright. UK, 13–16 Novemb. 2000 755–762 (2000).
Baudoin, A., Olaya, G., Delmotte, F., Colcol, J. F. & Sierotzki, H. QoI Resistance of Plasmopara viticola and Erysiphe necator in the mid-Atlantic United States. Plant Health Prog. 9, 25 (2008).
Kościuczuk, E. M. et al. Cathelicidins: Family of antimicrobial peptides. A review. Mol. Biol. Rep. 39, 10957–10970 (2012).
Starr, C. G., Maderdrut, J. L., He, J., Coy, D. H. & Wimley, W. C. Pituitary adenylate cyclase-activating polypeptide is a potent broad-spectrum antimicrobial peptide: Structure–activity relationships. Peptides 104, 35–40 (2018).
Jäkel, C. E., Meschenmoser, K., Kim, Y., Weiher, H. & Schmidt-Wolf, I. G. H. Efficacy of a proapoptotic peptide towards cancer cells. Vivo 26, 419–426 (2012).
Mahlapuu, M., Håkansson, J., Ringstad, L. & Björn, C. Antimicrobial peptides: An emerging category of therapeutic agents. Front. Cell. Infect. Microbiol. 6, 194 (2016).
Wang, K. et al. Antitumor effects, cell selectivity and structure–activity relationship of a novel antimicrobial peptide polybia-MPI. Peptides 29, 963–968 (2008).
Lei, J. et al. The antimicrobial peptides and their potential clinical applications. Am. J. Transl. Res. 11, 3919–3931 (2019).
Moravej, H. et al. Antimicrobial peptides: Features, action, and their resistance mechanisms in bacteria. Microb. Drug Resist. 24, 747–767 (2018).
Aisenbrey, C., Marquette, A. & Bechinger, B. The mechanisms of action of cationic antimicrobial peptides refined by novel concepts from biophysical investigations. in Advances in Experimental Medicine and Biology Vol. 1117, 33–64 (Springer New York LLC, 2019).
Schmatz, D. M. et al. Pneumocandins from Zalerion arboricola iv. Biological evaluation of natural and semisynthetic pneumocandins for activity against Pneumocystis carinii and Candida species. J. Antibiot. 45, 1886–1891 (1992).
Chen, W., Zeng, H. & Tan, H. Cloning, sequencing, and function of sanF: A gene involved in nikkomycin biosynthesis of Streptomyces ansochromogenes. Curr. Microbiol. 41, 312–316 (2000).
Matejuk, A. et al. Peptide-based antifungal therapies against emerging infections. Drugs Future 35, 197–217 (2010).
Taniguchi, M. et al. Cationic peptides from enzymatic hydrolysates of soybean proteins exhibit LPS-neutralizing and angiogenic activities. J. Biosci. Bioeng. 127, 176–182 (2019).
Bondaryk, M., Staniszewska, M., Zielińska, P. & Urbańczyk-Lipkowska, Z. Natural antimicrobial peptides as inspiration for design of a new generation antifungal compounds. J. Fungi 3, 46 (2017).
Vega-Vásquez, P., Mosier, N. S. & Irudayaraj, J. Nanoscale drug delivery systems: From medicine to agriculture. Front. Bioeng. Biotechnol. 8, 79–79 (2020).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
Kumar, S., Stecher, G., Li, M., Knyaz, C. & Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 35, 1547–1549 (2018).
Mitaku, S., Hirokawa, T. & Tsuji, T. Amphiphilicity index of polar amino acids as an aid in the characterization of amino acid preference at membrane-water interfaces. Bioinformatics 18, 608–616 (2002).
Lu, S. et al. CDD/SPARCLE: The conserved domain database in 2020. Nucleic Acids Res. 48, D265–D268 (2020).
Perazzolli, M., Roatti, B., Bozza, E. & Pertot, I. Trichoderma harzianum T39 induces resistance against downy mildew by priming for defense without costs for grapevine. Biol. Control 58, 74–82 (2011).
Tomada, S. et al. Dual RNA-Seq of Lysobacter capsici AZ78—Phytophthora infestans interaction shows the implementation of attack strategies by the bacterium and unsuccessful oomycete defense responses. Environ. Microbiol. 19, 4113–4125 (2017).
Schiestl, R. H. & Gietz, R. D. High efficiency transformation of intact yeast cells using single stranded nucleic acids as a carrier. Curr. Genet. 16, 339–346 (1989).
Macut, H. et al. Tuning PFKFB3 bisphosphatase activity through allosteric interference. Sci. Rep. 9, 1–10 (2019).
Pellegrino, S., Annoni, C., Contini, A., Clerici, F. & Gelmi, M. L. Expedient chemical synthesis of 75mer DNA binding domain of MafA: An insight on its binding to insulin enhancer. Amino Acids 43, 1995–2003 (2012).
Lazazzara, V. et al. Downy mildew symptoms on grapevines can be reduced by volatile organic compounds of resistant genotypes. Sci. Rep. 8, 1–14 (2018).
Plasmopara viticola. EPPO Bull.31, 313–317 (2001).
Abbott, W. S. A method of computing the effectiveness of an insecticide. J. Econ. Entomol. 18, 265–267 (1925).
Miclot, A. S., Wiedemann-Merdinoglu, S., Duchêne, E., Merdinoglu, D. & Mestre, P. A standardised method for the quantitative analysis of resistance to grapevine powdery mildew. Eur. J. Plant Pathol. 133, 483–495 (2012).
Uncinula necator. EPPO Bull.32, 315–318 (2002).
Perreault, F. et al. Evaluation of copper oxide nanoparticles toxicity using chlorophyll a fluorescence imaging in Lemna gibba. J. Bot. 2010, 1–9 (2010).
Schreiber, U., Quayle, P., Schmidt, S., Escher, B. I. & Mueller, J. F. Methodology and evaluation of a highly sensitive algae toxicity test based on multiwell chlorophyll fluorescence imaging. Biosens. Bioelectron. 22, 2554–2563 (2007).
Logemann, E., Birkenbihl, R. P., Ülker, B. & Somssich, I. E. An improved method for preparing Agrobacterium cells that simplifies the Arabidopsis transformation protocol. Plant Methods 2, 1–5 (2006).
Acknowledgements
We thank Fondazione Cariplo for the financial support of this research to P. P. (GraptaResistance, Rif. 2015-0430). We are also grateful to Sergey Reverdatto and Alexander Shekhtman for providing the pLib2 vector. Valerio Parravicini and Mario Beretta are acknowledged for their invaluable support with plant growth. Our deep thanks also go to Delia Tarantino for her availability and kindness. Delia passed away a year ago, this manuscript is dedicated to her memory. Part of this work was carried out at the Model Plants Platform, established by the Department of Biosciences at University of Milan.
Author information
Authors and Affiliations
Contributions
M.C., S.M., S.R., R.V., M.P., S.V. and P.P. conceived experiments; M.C., S.M., S.R., E.C., S.L.T., C.M., L.T., F.R., S.P., R.M., M.P., S.V. carried out the experimental work; M.C., S.M., S.R., M.P., S.V. and P.P. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Colombo, M., Masiero, S., Rosa, S. et al. NoPv1: a synthetic antimicrobial peptide aptamer targeting the causal agents of grapevine downy mildew and potato late blight. Sci Rep 10, 17574 (2020). https://doi.org/10.1038/s41598-020-73027-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-73027-x
This article is cited by
-
Unravelling molecular mechanisms involved in resistance priming against downy mildew (Plasmopara viticola) in grapevine (Vitis vinifera L.)
Scientific Reports (2023)
-
Combating fungal phytopathogens with human salivary antimicrobial peptide histatin 5 through a multi-target mechanism
World Journal of Microbiology and Biotechnology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.