Abstract
Most land plants form beneficial associations with arbuscular mycorrhizal (AM) fungi which improves mineral nutrition, mainly phosphorus, in the host plant in exchange for photosynthetically fixed carbon. Most of our knowledge on the AM symbiosis derives from dicotyledonous species. We show that inoculation with the AM fungus Funneliformis mosseae stimulates growth and increases Pi content in leaves of rice plants (O. sativa, cv Loto, ssp japonica). Although rice is a host for AM fungi, the systemic transcriptional responses to AM inoculation, and molecular mechanisms underlying AM symbiosis in rice remain largely elusive. Transcriptomic analysis identified genes systemically regulated in leaves of mycorrhizal rice plants, including genes with functions associated with the biosynthesis of phospholipids and non-phosphorus lipids (up-regulated and down-regulated, respectively). A coordinated regulation of genes involved in the biosynthesis of phospholipids and inositol polyphosphates, and genes involved in hormone biosynthesis and signaling (jasmonic acid, ethylene) occurs in leaves of mycorrhizal rice. Members of gene families playing a role in phosphate starvation responses and remobilization of Pi were down-regulated in leaves of mycorrhizal rice. These results demonstrated that the AM symbiosis is accompanied by systemic transcriptional responses, which are potentially important to maintain a stable symbiotic relationship in rice plants.
Similar content being viewed by others
Introduction
Arbuscular mycorrhizal fungi (AMF or AM fungi) are soil-borne obligate biotrophs that form mutualistic associations with most terrestrial plants1,2,3,4,5,6. The fungus provides the host plant with nutrients, mainly phosphorus, which generally improves plant growth and fitness. In return, the host plant provides the AM fungus with sugars and lipids, both representing a major source of organic carbon in the fungus7. In this symbiosis, the fungus penetrates and colonizes the root cortex, forming differentiated symbiotic structures, known as arbuscules, which are the sites for nutrient exchange between the two organisms. Besides plant nutrition improvement, AM fungi often confer tolerance to biotic and abiotic stresses8,9,10,11,12,13,14. The molecular mechanisms of interactions between plants and AM fungi have mainly been investigated using the model legumes of Medicago truncatula and Lotus japonicus15.
Rice is one of the most important cereal crops in the world and a staple for more than half of the global population. Rice is also a host for AM fungi, and several studies have found positive effects of AM fungi on growth, pathogen resistance, and drought tolerance in rice plants16. Even though rice can establish symbiotic relationships with AM fungi, the molecular and physiological responses to inoculation with an AM fungus in rice plants remain less explored. The main reason for this backwardness is that rice is mostly grown under flooded conditions (paddy fields), and plants growing in aquatic environments have been traditionally considered to be non-mycorrhizal. However, natural colonization with AM fungi has been reported in rice plants under aerobic and flooded conditions17,18.
Root colonization by AM fungi involves drastic reprogramming of gene expression for the control AM symbiosis. Most studies focused on transcriptional responses in roots of mycorrhizal plants19,20,21,22, whereas transcriptional responses in leaves of mycorrhizal plants have not yet been systematically investigated. Most AM-responsive genes in roots are involved in transcriptional regulation (e.g., transcription factors) and signal transduction (e.g., components of the “common symbiosis signaling” pathway), as well as in nutrient transport, or sugar and lipid transport15,23. Phytohormones are also important regulators of AM symbiosis6,24,25.
This study aimed to investigate systemic transcriptional alterations in leaves of mycorrhizal rice plants. These studies were carried out on the rice cultivar Loto (Oryza sativa, ssp. japonica) interacting with the AM fungus Funneliformis mosseae. Evidence is presented that the AM symbiosis triggers coordinated regulations in the expression of genes involved in lipid signaling (phospholipids and non-phosphorus lipids), hormone biosynthesis and signaling (jasmonic acid, ethylene), and genes involved in Pi homeostasis. These results provided a global view of genes, and members of gene families, that will advance our understanding on mechanisms underlying the AM symbiosis in rice.
Results
Effect of F. mosseae inoculation on growth of rice plants
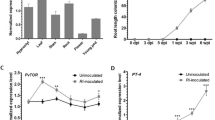
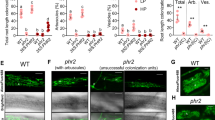
It is known that rice cultivars differ widely for colonization with the AM fungus R. irregularis26. In this work, we observed that inoculation with the AM fungus F. mosseae has an important effect on growth of Loto rice plants (Fig. 1A). F. mosseae-inoculated plants grew faster than non-inoculated plants, the stimulatory effect on plant growth being observed as early as 17 days after F. mosseae inoculation (Fig. 1A,B). Chlorophyll content, and indicator of the physiological status of plants, also increased in mycorrhizal plants (Fig. 1C). Inoculation with F. mosseae significantly increased Pi content in leaves of Loto plants (Fig. 1D), thus, supporting the formation of a functional AM symbiosis. Microscopic observations of cotton blue-stained roots confirmed root colonization by F. mosseae. Typical AM fungal structures, such as hyphae, vesicles and arbuscules were clearly observed in roots of F. mosseae-inoculated Loto plants (Fig. 2).
Phenotype of rice (Oryza sativa spp. japonica, cv Loto) inoculated with the AM fungus F. mosseae. (A) Rice plants were inoculated with F. mosseae (+ F.m; red line), or non-inoculated (-F.m; black line). Height of Loto rice plants was determined at the indicated times after inoculation with F. mosseae. Data are mean ± SE (n = 10). Asterisks denote statistical differences (*p < 0.05, ANOVA test; F. mosseae-inoculated vs non-inoculated). (B) Appearance of Loto rice plants at 23 days post-inoculation (dpi). (C) Chlorophyll content in leaves of mycorrhizal and non-mycorrizal plants at 23 dpi. Data are mean ± SE (n = 10). (D) Pi content in leaves of mycorrhizal and non-mycorrhizal Loto plants at 23 dpi. Data are mean ± SE (n = 5, each biological replicate is a pool of 2 individual leaves). FW fresh weight.
Colonization by F. mosseae in roots of rice (cv Loto) plants. (A) Representative cotton blue staining of roots colonized by F. mosseae at 8 wpi. Light micrographs show intracellular hyphae (ih), arbuscules (a) and vesicles (v). Scale bar = 100 μm. (B) Estimation of root colonization was determined by the Trouvelot method at 8 weeks post-inoculation. M%, Intensity of the mycorrhizal colonization in the root system; A%, Arbuscule abundance in the root system.
Transcriptome analysis reveals the impact of AM symbiosis in rice leaves
To investigate the molecular mechanisms underlying AM symbiosis in rice, we performed transcriptome analysis of leaves from mycorrhizal and non-mycorrhizal plants. Differentially expressed genes (DEGs) were identified based on significance level (FDR ≤ 0.01) and log2 fold change (FC) as the threshold for up-regulated (FC ≥ + 0.5) and down-regulated (FC ≤—0.5) genes. Using these criteria, 4993 and 5221 genes were found to be up-regulated and down-regulated, respectively, in leaves of mycorrhizal plants relative to non-inoculated plants (Fig. 3A; Supplementary Fig. S1A,B; Supplementary Table S1). Principal component analysis revealed a clear separation between non-inoculated and F. mosseae-inoculated plants (Supplementary Fig. S1C). These results support that the AM symbiosis has a strong impact on the leaf transcriptome of Loto rice plants. Similarly, massive transcriptional changes were reported in rice roots during AM symbiosis19.
Differentially expressed genes (DEGs) in leaves of F. mosseae-inoculated rice plants relative to non-inoculated plants by RNA-seq analysis. Leaves of 30-day old rice plants (cv Loto) were collected after 23 dpi of root inoculation with the AM fungus F. mosseae (+ F. mosseae). No inoculum was added to non-inoculated plants (-F. mosseae). (A) Number of DEGs (+ F. mosseae vs -F. mosseae). Up-regulated (log2 fold change [FC] ≥ 0.5; purple) and down-regulated (log2FC ≤ -0.5; green) genes (p < 0.05, false discovery rate [FDR] < 0.01, n = 3). (B) GO terms enriched in up- (left panel) and down- (right panel) regulated DEGs. The most enriched GO terms were identified (number of DEG ≥ 10; Enrichment Score ≥ 2.5) and visualized using REVIGO after reducing redundancy and clustering of similar GO terms (https://revigo.irb.hr/). Disc color (blue to red) represent the degree of GO enrichment (Enrichment Score) and disc size is proportional to the frequency of the GO term in the GO Oryza sativa database (larger and smaller discs represent more general and more specific terms, respectively). Full list of genes in enriched GO terms is presented in Supplementary Table S2.
Gene Ontology (GO) enrichment analysis of Biological Process (Supplementary Table S2) revealed that GO annotations related to “Phosphatidylinositol phosphorylation” were over-represented in the set of up-regulated genes in leaves of mycorrhizal plants (Fig. 3B, left panel). The GO categories of “Gene silencing”, “RNA modification and RNA processing”, and “Regulation of translation”, were also enriched among the up-regulated genes (Fig. 3B, left panel).
A significant enrichment in the GO term of “Cellular response to phosphate starvation” occurred in the set of down-regulated DEGs (Fig. 3B, right panel). Other GO categories enriched among the down-regulated genes were “Proteasomal protein catabolism”, or “Catabolic processes”. Results obtained by RNA-Seq analysis were validated by qRT-PCR (Supplementary Fig. S2A, B). A good correlation between qRT-PCR and RNA-seq data was observed (Supplementary Fig. S2C).
Transcriptional regulation of phosphoinositide and inositol polyphosphate biosynthesis genes in mycorrhizal rice plants
As previously mentioned, GO analysis revealed that genes involved in “Phosphatidylinositol phosphorylation” were significantly enriched in leaves of mycorrhizal rice. In plants, Phosphatidylinositol (PtdIns) (or phosphoinositide, PI) is an essential phospholipid that is required for the production of both membrane-bound phosphoinositides and soluble inositol polyphosphates (InsPs, or IPs). Besides their roles as membrane building blocks, phosphoinositides also exert regulatory functions through interaction with a wide range of cellular proteins27. As for inositol polyphosphates, they function as cofactors, regulators, and second messengers in plants. At present, however, there are major gaps in our knowledge about the possible contribution of phosphoinositides and inositol polyphosphates in interactions between plants and AM fungi. This fact prompted us to further investigate the expression of genes involved in inositol polyphosphate and phosphoinositide metabolism in mycorrhizal rice plants. To note, we found that the expression of phosphoinositide and inositol polyphosphate biosynthesis was highly regulated in leaves of mycorrhizal rice plants (Fig. 4A; Table 1; Supplementary Table S3). The mycorrhiza-regulated genes in the phosphoinositide and inositol polyphosphate biosynthesis pathway identified in this study are presented in Fig. 4B.
Differential expression of genes involved in inositol polyphosphates (InsP) and phosphoinositides (PtdIns) metabolism in leaves of rice (cv Loto) plants. RNA-Seq was used to identify DEGs in leaves of F. mosseae-inoculated vs non-inoculated (− F.m vs − F.m) rice at 23 dpi. (A) Heatmaps showing expression level (left panel; log10 [FPKM + 1]) and Fold Change (right panel; log2FC) of DEGs. Gene expression is represented from pale yellow (less expressed) to orange (more expressed). Upregulated (log2FC ≥ 0.5; purple) and downregulated (log2FC ≤ − 0.5; green) DEGs. Data represented are individual replicates (n = 3, each biological replicate (R) consisting in a pool of 6 leaves from individual plants). The full gene names and ID list are in listed in Table 1 and Supplementary Table S3. (B) Biochemical pathways for the synthesis of InsP and PtdInsP. Interactions with JA signaling and Pi homeostasis are indicated. DEGs in leaves of AM-inoculated vs leaves of non-inoculated rice plants are indicated in purple (up-regulated) and green (down-regulated). Phosphoinositides species identified in plants include PtdIns monophosphates (PtdIns3P, PtdIns4P, PtdIns5P), and PtdIns biphosphates (PtdIns(3,5)P2, PtdIns(4,5)P2). The presence of PtdIns(3,4)P2 or PtdIns(3,4,5)P3 in plants has not been reliable confirmed. Enzyme abbreviations are as follows: 5PTase Phosphatidylinositol 5-phosphatase, CDS Cytidine diphosphate diacylglycerol synthase, D50 DWARF50, CDP-DAG Cytidine diphosphate diacylglycerol, DAG diacylglycerol, DGK Diacylglycerol kinase, FAB Phosphatidylinositol 3phosphate 5-kinase, GH1 Grain number and plant Height1, IMP Inositol monophosphatase, Ins myo-inositol (the positions on the inositol ring are designated 1–6), IPKK Inositol-pentakisphosphate 2-kinase, IPMK Inositol polyphosphate multikinase, IPTK Inositol 1,3,4-trisphosphate 5/6-kinase, MIK Myo-inositol kinase, MTM Phosphatidylinositol 3-phosphatase myotubularin, NPC Non-specific phospholipase C, PA phytic acid, PAP Phosphatidic acid phosphatase, PATL Patellin Sec14-domain, PI3K Phosphatidylinositol 3-kinase, PI4K Phosphatidylinositol 4-kinase, PIP5K Phosphatidylinositol 4-phosphate 5-kinase, PI-PLC Phosphatidylinositol-specific phospholipase, PIS Phosphatidylinositol synthase, PITP Phosphatidylinositol-transfer proteins, PtdChol Phosphatidylcholine, PtdIns Phosphatidylinositol, PtdOH Phosphatidic acid, VIP VIP/PPIP5K diphosphoinositol pentakisphosphate kinase, SAC Phosphatidylinositol 3,5-bisphosphate 5-phosphatase and Phosphatidylinositol 4-phosphate phosphatase, SNDP Sec14-nodulin domain-containing protein.
Among the genes up-regulated in leaves of mycorrhizal plants there were genes involved in: (i) the synthesis of PtdIns, namely cytidine diphosphate-diacylglycerol (CDP-DAG) synthase (OsCDS2), and PtdIns synthase (OsPIS); (ii) kinase genes acting in the production of phosphorylated derivatives of PtdIns, either PtdIns monophosphates [PtdIns3P, PtdIns4P] (PI3K and PI4K family proteins), or PtdIns bisphosphates [PtdIns(3,5)P2, PtdIns(4,5)P2] (FAB1/PIKFYVE and PIP5K family proteins) (Fig. 4A,B; Table 1). Genes encoding PtdIns3P and PtdIns4P phosphatases, and PtdIns(3,5)P2 (Myotubularin, SAC1, and 5´phosphatases family proteins) were also up-regulated. The production of Ins3P from Glucose-6P appears to be repressed in leaves of mycorrhizal plants, as revealed by the down-regulation of myo-inositol-1-phosphate synthase (INO1 or MIPS) (Fig. 4B). Altogether these findings point to fine-tuning of phosphoinositide levels in mycorrhizal rice.
In plants, InsPs are synthesized via two pathways: the lipid-independent (or soluble) and lipid–dependent pathways28. In the lipid-independent pathway, soluble inositol is phosphorylated through the action of kinases, whereas in the lipid-dependent pathway, different InsP intermediates originate through degradation of PtdIns by the action of phosphoinositide-specific phospholipases28. Here, the unique pathway that appears to be activated in leaves of mycorrhizal plants is the soluble pathway, as revealed by the up-regulation the myo-inositol kinase (MIK1) and down-regulation of the phosphoinositide-specific phospholipase C (PI-PLC4) (Fig. 4A,B; Table 1). Also, the expression of genes encoding kinases responsible for the production of the different inositol polyphosphates was found to be regulated in leaves of mycorrhizal plants (Fig. 4A,B; Table 1). They were: OsIPK1 (Inositol-pentakisphosphate 2-kinase, IPPK), involved in the phosphorylation of Ins(1,3,4,5,6)P5 to produce Ins(1,2,3,4,5,6)P6 (phytic acid, or InsP6), OsIPK2 (Inositolpolyphosphate multikinase, IPMK), involved in sequential phosphorylations from inositol triphosphates Ins(1,4,5)P3] to inositol-pentakisphosphate Ins(1,3,4,5,6)P5] and OsIPTK1, OsIPTK3, OsIPTK5, OsIPTK6 (Inositol 1,3,4-trisphosphate 5/6-kinase, IPTK), involved in phosphorylation of the 5/6-position of inositol triphosphates Ins(1,3,4)P3] to produce inositol-tetrakisphosphate Ins(1,3,4,5)P4 and Ins(1,3,4,6)P4 and Ins(1,3,4)P3. OsIPTK1, OsIPTK3, OsIPTK5 have InsP6 kinase activity in vitro, and produce InsP7 and InsP8 from phytic acid29. Also, OsVIP1 and OsVIP2, encoding inositol hexakisphosphate and diphosphoinositol-pentakisphosphate kinase Vip1/PPIP5K-like proteins, produce InsP7 and InsP8. VIP2, the rice homolog for the Arabidopsis VHI2 protein, critical for InsP8 production30, was upregulated in leaves of mycorrhizal rice plants.
Collectively, results here presented support that the inoculation of rice roots with the AM fungus F. mosseae results in systemic alterations in the expression of genes involved in the biosynthesis of inositol polyphosphates and phosphoinositides. It is worth mentioning that although several studies described a correlation between inositol phosphate metabolites and Pi homeostasis in the model dicotyledonous Arabidopsis30,31,32,33,34, at present, no systematic study of phosphoinositide or inositolpolyphosphate signaling has been performed in AM symbiosis in cereal species, as rice is. Similarly, cross-talk between inositol polyphosphates and JA signaling has been described in Arabidopsis35,36,37. Taking into account that inositol phosphate metabolites act as important signaling molecules, the observed mycorrhiza-induced alterations in the expression of genes involved in inositol phosphate metabolic pathways is expected to have an impact on multiple signaling processes in mycorrhizal rice plants, including Pi homeostasis and hormonal regulation. Along with this, genes involved in hormone signaling were also found to be regulated in leaves of mycorrhizal rice plants, as described below.
Expression of genes related to hormone signaling in leaves of mycorrhizal rice plants
In addition to developmental regulation, plant hormones also play important roles in symbiotic interactions between plants and AM fungi38. Each of the plant hormones possesses specific functions and they also interact with each other either in an antagonistic or cooperative way. However, contradictory results have been reported in studies related to hormone signaling in AM symbiosis, such as Jasmonic acid (JA), ethylene (ET), Salicylic acid (SA) and auxins, which led to the notion that these regulatory mechanisms are dependent on the plant species and growth conditions (e.g., Pi supply conditions)38. Contrary to what was previously described in Medicago truncatula and tomato, the establishment of AM symbiosis appears to occur independently of Jasmonic Acid in rice39.
Transcriptome analysis revealed a down-regulation of many genes involved in JA biosynthesis and signaling in leaves of mycorrhizal plants (Fig. 5A; Supplementary Table S4). The mycorrhiza-regulated genes in the JA biosynthesis pathway identified in this study are presented in Fig. 5B. Although most of our knowledge in JA biosynthesis and signaling comes from studies in Arabidopsis, all homologs of Arabidopsis genes have been identified in rice40. JA (and JA-derivatives) originates from lipids, preferentially α–linolenic acid, which is released from galactolipids of chloroplast membranes. Linoleic acid is then converted to OPDA through the enzymes Lipoxygenase (LOX), allene oxidase synthase (AOS), and allene oxide cyclase (AOC) in the chloroplast. OPDA is then transported to peroxisomes where it is reduced by OPDA reductase (OPR), and after three rounds of β-oxidation JA is produced. To note, among the mycorrhiza down-regulated genes in rice leaves, we identified genes involved in the different steps of JA biosynthesis, namely Phospholipases A (PLA1-IIδ, PLA1-Iβ1, PLA1-Iγ1, pPLA-IIIα, pPLA-IIϕ, pPLA-IIIβ, pPLA-IIIζ), Phospholipases C (PLC4, NPC1, NPC2, NPC3), Phospholipases D (PLDα2, PLDα3, PLDζ2, PLDφ), Lipoxygenases (OsLOX1, OsLOX2, OsLOX5, OsLOX6, OsLOX7, OsLOX8, OsHI-LOX, OsLOX), AOS (OsAOS1, OsAOS2), AOC (represented by a unique member in the rice genome), and OPR (OsOPR1, OsOPR7, OsOPR10) genes (Fig. 5A,B; Supplementary Table S4). Although the functional implication of this negative regulation of JA biosynthesis genes remains to be investigated, these findings suggest that JA biosynthesis is repressed in leaves of mycorrhizal rice plants which is consistent with previous observations indicating that JA biosynthesis is dispensable for AM symbiosis in rice39.
Differential expression of genes involved in biosynthesis of Jasmonic Acid and Ethylene in leaves of rice (cv Loto) plants. RNA-Seq was used to identify DEGs in leaves of F. mosseae-inoculated vs non-inoculated (+ F.m vs − F.m) plants at 23 dpi. (A) Heatmaps showing RNA-seq expression level and fold change as indicated in Fig. 4A legend. The full gene names and ID list are in listed in Supplementary Table S4. (B) Biochemical pathway for the synthesis of Jasmonic acid (JA). DEGs in leaves of AM-inoculated rice plants are indicated in purple (up-regulated) and green (down-regulated). Enzyme abbreviations are as follows: AOC Allene oxide cyclase, AOS Allene oxide synthase, COI Coronatine insensitive 1, HPL3 Hydroperoxide lyase 3, JAZ jasmonate ZIM-domain protein, LOX Lipoxygenase, NPC Non-specific phospholipase C, OPR 12-oxo-phylodienoic acid, PLA Phospholipase A, PLC Phospholipase C, PLD Phospholipase D, SCF SKp1, Cullin and F-box proteins, TPL TOPLESS protein.
Regarding JA signaling, it is known that perception of JA-Ile (the bioactive JA derivative) by CORONATINE INSENSITIVE 1 (COI1) and JASMONATE ZIM DOMAIN (JAZ) proteins triggers the degradation of JAZ proteins by the SCFCOI1 complex-mediated 26 proteasome for activation of downstream JA-responsive genes (Fig. 5). Whereas the Arabidopsis genome has a single COI1 gene (COI1), the rice genome contains three closely related COI genes (OsCOI1, OsCOI2, OsCOI3). The three COI genes play a role in JA signaling in rice by interacting with JAZ proteins in rice41. Our transcriptome analysis revealed up-regulation of OsCOI2, whereas 6 OsJAZ genes (OsJAZ1, OsJAZ3, OsJAZ4, OsJAZ6, OsJAZ7, OsJAZ11) were repressed in leaves of mycorrhizal plants (Fig. 5; Supplementary Table S4).
Among the set of down-regulated genes in leaves of mycorrhizal rice, we also identified the three key genes involved in ethylene biosynthesis, namely the S-adenosyl-L-methionine synthetase (OsSAMS1, OsSAMS2), which catalyzes the conversion of S-adenosylmethionine (AdoMet) from methionine, the 1-aminocyclopropane-1-carboxylic acid (ACC) synthases (OsACS5) which catalyzes the conversion of AdoMet to ACC, the rate-limiting step in ET biosynthesis, and ACC oxidases (OsACO1, OsACO3, OsACO5) which catalyzes oxidation of ACC to form ethylene (Fig. 5A; Supplementary Table S4). Although not proven, this observation points to the repression of ethylene biosynthesis in rice leaves during AM symbiosis.
Expression pattern of genes involved in Pi signaling and homeostasis
As previously mentioned, genes in the “Cellular Response to Phosphate starvation” GO term were highly represented among down-regulated DEGs in leaves of mycorrhizal rice plants (Fig. 3B, right panel). This category included genes that mediate Pi signaling, transport, and remobilization. In particular, the expression of an important number of phosphate transporters was found to be down-regulated. They included: members of the PHT phosphate transporters belonging to the PHT1 (OsPHT1;1, PHT1;4, OsPHT1;8), PHT2 (OsPHT2;1), PHT3 (OsPHT3;1, OsPHT3;5) and PHT4 (OsPHT4;1, OsPHT4;3, OsPHT4;5) families, and PHOSPHATE1 (PHO1) transporters (OsPHO1-2, OsPHO1-3) (Fig. 6A; Supplementary Table S5). In Arabidopsis, phosphorylation of PHT1 phosphate transporters by CK2α3/β3 kinase negatively affects trafficking to the plasma membrane from the endoplasmic reticulum42. We noticed that the CK2α3 subunit gene was up-regulated in leaves of mycorrhizal rice pointing to a possible retention of PHT1 transporters in the ER (Fig. 6A; Supplementary Table S5). Figure 6B represents the signaling events involved in Pi homeostasis altered in mycorrhizal rice plants.
Differential expression of genes involved in Pi signaling and homeostasis in leaves of rice (cv Loto) plants. RNA-Seq was used to identify DEGs in leaves of F. mosseae-inoculated vs non-inoculated (+ F.m vs − F.m) plants at 23 dpi. (A) Heatmap showing RNA-seq expression level and fold change as indicated in Fig. 4A legend. The full gene names and ID list are in listed in Supplementary Table S5. (B) Regulatory network of Pi starvation responses. The PHR and related transcription factors are negatively regulated by the SPX proteins, which can sense Inositol polyphosphates (InsPs). Under P limitation, PHR relieves negative post-transcriptional control of Pi transporters responsible of Pi uptake through the induction of MIR399 (targeting PHO2). In rice, miR827 targets SPX-MFS1 and SPX-MSF2 transcripts. The SPX-MFS1 and SPX-MSF2 proteins, as well as the Vacuolar Efflux Proteins (VPEs) modulate Pi homeostasis by regulating Pi influx and efflux from vacuoles. Under sufficient Pi-conditions, the CK2α3/β3 kinase phosphorylates PHT1 resulting in retention of PHT1 transporters in the endoplasmic reticulum. InsPs are sensed by the SPX domain of SPX-EXS proteins (e.g., PHO1-2, PHO1-3) and SPX-MFS proteins. Most of these studies have been done in Arabidopsis43.
PHR1 (PHOSPHATE STARVATION RESPONSE 1), a member of the MYB superfamily of transcription factors, has been shown to control the transcriptional activation and repression responses to phosphate starvation in plants43,44,45,46 In particular, PHR1 and related transcription factors are known to be key players in Pi signaling in Arabidopsis and rice43,45,46. To note, the expression of several PHR transcription factors were regulated in leaves of mycorrhizal plants (Fig. 6; Supplementary Table S5). Of them, OsPHR4 was strongly down-regulated, indicating that this particular PHR family member might have a specific function in regulating Pi homeostasis in leaves of mycorrhizal rice. Other transcription factors that are involved in the phosphate starvation responses were also regulated in leaves of mycorrhizal rice, such as OsMYB4P and OsWRKY74 (down-regulated), or PTF1 (up-regulated) (Fig. 6; Supplementary Table S5)47,48,49. Furthermore, the expression of several SPX domain-containing protein genes was repressed in leaves of mycorrhizal plants (OsSPX1, OsSPX2, OsSPX3, OsSPX5, OsSPX6). SPX domain-containing proteins are known to function as a sensor for Pi level in controlling Pi homeostasis50,51.
Pi import and export from the vacuole, the primary intracellular compartment for Pi storage in plant cells, serve as a critical mechanism for the maintenance of appropriate cytosolic Pi levels. In rice, three members of SPX proteins possess an MFS (Major Facilitator Superfamily) domain, and are involved in Pi transport across the tonoplast, SPX-MSF1, SPX-MSF2, and SPX-MSF3. An opposite behavior in the expression of these genes could be observed in leaves of mycorrhizal rice plants (down-regulation of OsSPX-MSF2; up-regulation of OsSPX-MSF1 and OsSPX-MSF3). In this respect, Pi deficiency was reported to induce (OsSPX-MSF2) or repress (OsSPX-MSF1, OsSPX-MSF3) the expression of these genes in rice51. Expression of the vacuolar Pi efflux transporters OsVPE1 and OsVPE2 was also down-regulated in leaves of mycorrhizal rice (Fig. 6A,B; Supplementary Table S5).
Genes that have proven to be implicated in the remobilization of Pi from organic P resources, such as purple acid phosphatases (PAPs), acid phosphatase (ACP1), and ribonucleases (RNS3, PR10a) were repressed in leaves of mycorrhizal rice plants, indicating that these plants have an adequate P status (Fig. 6A; Supplementary Table S5). In this respect, it is well known that membrane phospholipids (a major source for internal Pi supply) are degraded under Pi limiting conditions, which are then replaced by non-phosphorus lipids (galactolipids and sulfolipids). Galactolipids are essential for a functional photosynthetic apparatus in the chloroplast, and include monogalactosyldiacylglycerol (MGDG) and digalactosyldiacylglycerol (DGDG) lipids, which are synthesized by MGDG synthase (or MGD) and DGDG synthase (or DGD). Plants have evolved various acclimation responses to cope with phosphate depletion, including Pi remobilization from endogenous phosphorus-containing resources like phospholipids52. Thereby membrane phospholipids are dephosphorylated and can be used as an internal phosphate source, while galactolipids are incorporated into the membrane to maintain membrane functionality. Remarkably, the expression of genes involved in the biosynthesis of these major galactolipids, MGDG (OsMGD1, OsMGD2, OsMGD3) and DGDG (OsDGD1-1, OsDGD1-2, OsDGD2-1, OsDGD2-2) was regulated in leaves of mycorrhizal rice, their expression being mainly down-regulated in leaves of mycorrhizal rice (Fig. 6; Supplementary Table S5). These observations suggest that membrane lipid remodeling toward galactolipids, most probably, does not occur in leaves of mycorrhizal plants because of the greater capacity of mycorrhizal roots to take up Pi. On the other hand, the synthesis of sulfolipids is catalyzed by uridine diphosphate-sulfoquinovose synthase (encoded by OsSQD1) and sulfoquinovosyldiacylglycerol synthase (OsSQD2). The expression of OsSQD1, OsSQD2.1 and OsSQD2.2 was repressed in leaves of mycorrhizal rice.
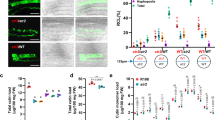
On the other hand, the implication of the microRNAs miR399 and miR827 in regulating Pi starvation responses in plants is well documented50,53. Our transcriptome analysis revealed down-regulation of miR827, and up-regulation of its target gene SPX-MSF1 encoding a vacuolar Pi transporter (Fig. 6; Supplementary Table S5), which was further confirmed by qRT-PCR and statistical analysis of miR827 precursor transcripts and SPX-MSF1 (Fig. 7A). miR827 transcripts were barely detectable in mycorrhizal rice. Thus, a miR827-mediated cleavage of SPX-MSF1 transcripts would regulate the intracellular distribution of Pi between the cytoplasm and vacuoles50,51.
Accumulation of miR399 and miR827 precursor transcripts in leaves of mycorrhizal plants. RNA was obtained from leaves of non-inoculated (− F.m) and AM inoculated (+ F.m) rice (cv. Loto) plants at 23 dpi. The expression level was determined by qRT-PCR. Primers used sequences are listed in Supplementary Table S6. Data are mean ± SE (n = 3; each sample consisted of a pool of 3 individual leaves; *p < 0.05, ANOVA test). (A) Accumulation of precursor transcripts for miR827 and its target genes OsSPX-MFS1. (B) Accumulation of precursor transcripts for miR399 family members and its target genes OsPHO2.
On the other hand, the rice miRNA399 family consists of 11 members (miRNA399a to miR399k; miRBASE v22.1) (Supplementary Fig. S3), all of then targeting the PHOSPHATE2 (OsPHO2, also named LTN, Leaf Tip Necrosis) gene, which encodes a ubiquitin-conjugating E2 enzyme that mediates the degradation of Pi transporters54,55. As for most rice miRNAs, no RAP-DB gene ID is currently assigned to any of the miR399 family members, and, accordingly, this miRNA was not identified as DEG in our RNA-seq analysis. Nevertheless, the miR399 target OsPHO2 was significantly up-regulated in leaves of mycorrhizal rice (Fig. 6; Supplementary Table S5). Accordingly, the expression of each miR399 member was analyzed by qRT-PCR. Precursor transcripts for seven out of the eleven miR399 family members (MIR399a, MIR399b, MIR399c, MIR399d, MIR399e, MIR399i, MIR399j) accumulated in leaves of non-mycorrhizal control plants, these plants being grown under low Pi supply (Fig. 7A). Of them, MIR399a, MIR399b, MIR399c, MIR399d, MIR399j were barely detectable in leaves of mycorrhizal plants. Conversely, MIR399e and MIR399i precursor transcripts accumulated at higher or similar level, respectively, in mycorrhizal plants compared with non-mycorrhizal plants. These results demonstrated a statistically significant differential regulation (from repression to activation) of miR399 family members in leaves of mycorrhizal rice plants. In this study, precursor transcripts for MIR399f, MIR399g, and MIR399h were undetectable. As expected and in accordance with RNA-seq data, anticorrelation between MIR399 and OsPHO2 occurred in leaves of mycorrhizal rice (Fig. 7B).
Overall, comparative analysis of leaf transcriptomes of mycorrhizal and non-mycorrhizal rice plants revealed systemic regulation of distinct members of genes involved in Pi signaling and homeostasis. Repression of phosphate starvation-responsive genes and those involved in the biosynthesis of non-phosphorus membrane lipids is consistent with an improved Pi nutrition, as revealed by the observation that mycorrhizal Loto plants have a higher Pi content in leaves than non-mycorrhizal plants (Fig. 1D).
Discussion
Although rice is the staple food of more than half of the world's population, very few studies focused on the impact of root colonization by AM fungi on rice. Here, we show that inoculation with F. mosseae stimulates growth and increases Pi content in leaves of rice plants (cv Loto), supporting that Loto plants are effectively colonized by this AM fungus. We also show that root colonization by F. mosseae is associated to massive systemic transcriptional alterations in the rice leaf transcriptome. The transcriptional responses in leaves of mycorrhizal rice plants appear to differ substantially from those previously reported in roots of rice plants colonized by R. irregularis19,56. Here, alterations in the expression of genes related to mineral transport were described in lateral roots, whereas secondary cell wall and phytohormone metabolism genes are regulated in adult crown roots19. As AM symbiosis improves Pi nutrition with the potential to optimize rice production, a deeper understanding of transcriptional regulatory mechanisms underlying AM symbiosis in rice is a requisite to ultimately use this symbiotic association in rice cultivation while avoiding the adverse impact of chemical fertilizers.
Diverse processes were regulated in leaves of mycorrhizal rice plants, such as those related to phospholipid and hormone signaling, as well as regulatory mechanisms controlling Pi homeostasis (summarized in Fig. 8). Reprogramming of host gene expression in leaves is potentially important to maintain an appropriate Pi status in the plant and a stable symbiotic relationship.
Summary of systemically regulated genes in leaves of mycorrhizal rice plants. Genes involved in phosphatidylinositol biosynthesis and phospholipid-based signaling events are indicated. Heatmaps showing log2 fold change (AMF-inoculated vs non inoculated) rice (cv. Loto) plants are included. Up-regulated (purple) and down-regulated (green) DEGs. The expression of genes involved in AM symbiosis has been mostly studied in roots of mycorrhizal plants57. In this study, several AM symbiosis-related genes were also found to be up-regulated in leaves of mycorrhizal rice plants, such as two cation channels OsCASTOR (Os03g0843600) and OsPOLLUX (Os01g0870100), and two nuclear porins OsNUP85 (Os01g074620) and OsNUP133 (Os03g022550).
Results here presented revealed systemic alterations in the expression of genes involved in phosphatidylinositol metabolism and signaling, including major phospholipids (phosphoinositides), inositol polyphosphates, and non-phosphorus lipids (galactolipids and sulfolipids). Whereas most genes involved in phosphoinositide and inositol polyphosphate biosynthesis were up-regulated in leaves of mycorrhizal rice, genes involved in the biosynthesis of non-phosphorus lipids (galactolipids and sulfolipids) were essentially down-regulated. Studies in Arabidopsis have shown that plants adapt to low P status by replacing phospholipids by galactolipids and sulfolipids. Accordingly, non-phospholipid biosynthesis is induced while phospholipid biosynthesis is repressed in Pi-starved plants58,59. Our transcriptome analysis showed activation of phospholipid biosynthesis genes (Fig. 4) and repression of non-phosphorus lipid biosynthesis genes (Fig. 6) in leaves of mycorrhizal rice plants (summarized in Fig. 8). These findings suggest that inoculation with F. mosseae provides an adequate Pi level to the rice plant and that Pi acquired in AM colonized roots is used efficiently. Regulation of phospholipid biosynthesis genes reinforces the notion that phospholipids might play a central role in the AM symbiosis.
Transcript profiling also revealed up-regulation of kinase genes involved in phosphatidylinositol biosynthetic processes. Plant phosphoinositides and inositol polyphosphates are formed via phosphorylation of the phospholipid PtdIns by the activity of specific kinases. Dynamic changes in phosphoinositide levels occur in plant tissues in which enzymes mediating biosynthesis and degradation of phosphoinositides are continuously synthesized and degraded27. With this in mind, it is not surprising to find that kinase and phosphatase genes regulating a particular reaction opposite one another are both up-regulated. In comparison to enzymes of phosphatidylinositol biosynthesis, however, there is only limited information about phosphatases and/or phospholipases involved in phosphatidylinositol degradation. Whereas biochemical assays can only reveal global kinase activity, transcript profiling allowed us to discriminate among members of the same family that are specifically regulated in leaves of mycorrhizal rice (e.g., IPTKs or inositol kinases).
As phospholipids are one of the main structural components of membranes, alterations in the expression of phospholipid biosynthesis genes might influence membrane structure and/or function. But phospholipids and inositol polyphosphates are also important signaling molecules that function as cofactors of partner proteins. It is tempting to hypothesize that alterations in phospholipid biosynthesis genes are accompanied by changes in the level of these metabolites, which would then have an impact on diverse cellular processes. However, the functional implications of alterations in phospholipid biosynthesis genes in AM symbiosis in rice have yet to be clarified. It will now be of interest to investigate whether mycorrhiza-induced alterations in phospholipids have a consequence in long-distance signaling. In this respect, the presence of phospholipids and phospholipid-binding proteins, in the phloem of several plant species has long been reported60,61,62. A phospholipid-based signaling mechanism might then operate during AM symbiosis.
Moreover, alterations in inositol polyphosphate biosynthesis genes might have an effect on hormonal signaling processes, particularly with regard to JA signaling. In favor of this hypothesis, InsP5 and InsP4 were found to interact with the COI/JAZ complex involved in JA perception in Arabidopsis36. In other studies, InsP7 and InsP8 were reported to promote the binding between COI1 and JAZ1 for the regulation of JA-dependent responses30. The VIP gene functioning in phosphorylation of PP-IP5 (named as VIH2 in Arabidopsis) regulates the synthesis of InsP8, the cofactor of the COI-JAZ receptor complex30. Results here presented, together with those obtained by other authors, broadens our view on the existence of cross-talk between phospholipid metabolism and JA signaling processes during AM symbiosis. Clearly, many efforts are still needed to understand the connections between these signaling pathways in AM symbiosis.
Plants associate with AM fungi when grown under low Pi soil conditions and, accordingly, AM symbiosis is generally studied in plants grown in low Pi. In line with this, down-regulation of Pi starvation-responsive genes occurs in leaves of mycorrhizal rice compared with non-mycorrhizal plants (the later ones responding to the situation of low Pi supply). Among the down-regulated genes in leaves of mycorrhizal rice, we identified genes belonging to different families of Pi-responsive genes. They included genes involved in: (i) Pi transport (e.g. several PHT, vacuolar transporters; PHO1); (ii) signaling processes (SPX domain, CK2α3, PHO2); (iii) transcriptional regulation (PHR, PTF1 transcription factors); (iv) post-transcriptional regulation (miR399, miR827); (v) Pi recycling (phosphatases, RNases); and (vi) non-phosphorus membrane lipid biosynthesis (galactolipids, sulpholipids). Pi starvation-responsive genes whose expression is regulated in leaves of mycorrhizal rice are summarized in Fig, 8. A differential regulation of genes belonging to each one of these families points to a specialization of these mycorrhiza-regulated family members in controlling Pi homeostasis in rice leaves. As an example, a functional specialization of miR399 species in regulating Pi homeostasis in leaves of mycorrhizal plants can be anticipated. Also, down-regulation of genes involved in Pi starvation responses or Pi remobilization from macromolecules (PAPs, RNases, galactolipid and sulfolipid biosynthesis genes) further supports that rice plants have a normal level of Pi.
As an additional level of complexity in the regulation of Pi homeostasis in plants, there is growing evidence on a regulatory role of inositol polyphosphates in Pi signaling33. Indeed, the SPX domain was recently defined as a Pi sensor in Arabidopsis plants that regulates Pi transport activity of SPX domain-containing proteins through binding inositol polyphosphate signaling molecules32,33. Additionally, the Arabidopsis PHR1 and rice PHR2 transcription factors, which are recognized as central components of the phosphate starvation response, have been shown to interact with SPX proteins63,64. The interaction between SPX proteins and the AtPHR1 transcription factor results in PHR1 inactivation, being this interaction promoted by inositol polyphosphates34. Results here presented demonstrated connections between inositol polyphosphate metabolism and Pi homeostasis in AM symbiosis. Furthermore, mechanisms involved in Pi homeostasis and hormonal signaling must be tightly regulated in leaves of mycorrhizal rice, which would then allow the host plant to maximize the benefits of Pi acquisition during association with the AM fungus. Additional studies are, however, needed to determine the exact mechanisms by which phospholipids might exert a regulatory function in maintaining Pi homeostasis and how this would affect hormonal processes in mycorrhizal rice plants.
In conclusion, our work provided new insights on the systemic alterations in gene expression during the symbiotic association of rice plants with an AM fungus. Phosphorus deficiency is widespread in all major rice ecosystems and is the major growth-limiting factor in acid upland soils where soil P-fixation capacity is often high (https://www.knowledgebank.irri.org). To circumvent this problem, P fertilizers are applied extensively in rice farming to improve plant performance, but their use has negative effects on the environment. Although there are still numerous issues that still need to be addressed, pieces of evidence here presented might accelerate research on the AM symbiosis in rice, which will open new avenues for improvement of sustainability in rice production. A more sustainable way to improve mineral nutrition in rice would be through the use of AM fungi, which would also avoid the overuse of chemical fertilizers.
Materials and methods
Plant and fungal material
The rice cultivar Loto (Oryza sativa ssp japonica) was used in this study. Loto rice is a heritage variety of short-grain rice widely cultivated in northern Italy (the first rice producer in Europe), also grown in southern Switzerland (the northern-most rice-growing region in Europe) and northern United States (Vermont State), making Loto a suitable rice variety for rice farming in northern latitudes.
The AMF fungus Funneliformis mosseae (formerly Glomus mosseae; FR140) was purchased from MycAgro (Bretenière, France; https://www.mycagrolab.com/) in a granular form composed of mineral inert solid particles (clay, zeolite) and F. mosseae propagules at a concentration of minimum 10 propagules/gram.
Inoculation with F. mosseae and growth of AMF-inoculated plants
Rice Loto seeds were dehusked, surface-sterilized twice with 5% sodium hypochlorite for 15 min, and washed extensively with sterile water. Seeds were germinated on Petri dishes with sterile water for 7 days and then transplanted to 150 ml-cones (20.5 cm; 2 plants/cone) containing a mix of 63.3% quartz sand (0.3–0.8 mm), 31.6% soil (turface and vermiculite 2:1), and 5% of granular inoculum of Funneliformis mosseae. No inoculum was added to the substrate for the non-inoculated, control plants. Transplanted seedlings were then grown under aerobic conditions at the greenhouse under a 14 h/10 h day/night cycle, 28 ºC/25 ºC, and 60% humidity for the required time. Rice seedlings were bottom-watered during the whole experiment. Fertirrigation was supplied at nine days after F. mosseae inoculation, using a modified Hoagland half-strength solution (15 ml, every two days) containing 2.5 mM KNO3, 2.5 mM Ca(NO3)2·4H2O, 1 mM MgSO4·7H2O, 0.5 mM NH4NO3, 25 μM KH2PO4, 23.15 μM H3BO3, 4.55 μM MnCl2·4H2O, 0.38 μM ZnSO4·7H2O, 0.1 μM CuSO4·5H2O, 0.14 μM Na2MoO4·2H2O, 26 μM Fe-EDDHA, pH 5.5)65.
Analysis of root colonization
Root fragments were collected at eight weeks post-inoculation with F. mosseae and stained with 0.1% cotton blue in acid lactic as previously described66. Fifty root fragments (1 cm) per biological replicate (n = 3) were examined with a light microscope, and the intensity of root cortex colonization was determined67. AMF structures were examined using an Axiophot Zeiss microscope equipped with a Digital color camera (DP70 Olympus) and 40 × magnification.
Transcriptome analysis by RNA-Seq
Total RNA was extracted from rice leaves of 30-day old plants that had been root-inoculated with F. mosseae (23 days post-inoculation), or not, using Maxwell(R) RSC Plant RNA Kit (Promega). Three biological replicates for each condition were examined, each biological replicate consisting of leaves from five individual plants (the youngest totally expanded leaf was selected from each individual plant). RNA concentration and purity were checked using a spectrophotometer (NanoDrop, ND-1000). RNA quality and integrity were evaluated using an Agilent 2100 Bioanalyzer (Agilent Technologies, Inc.), and only samples with an. RNA integrity number (RIN) ≥ 8 were used. RNA was analyzed by sequencing six libraries. An average of 32 736 498 clean reads/library were reported (Supplementary Table S6). Raw 125 bp paired-end reads processing and analysis were carried out as previously described65. RNA‐Seq raw data are available at the NCBI GEO database (https://www.ncbi.nlm.nih.gov/geo) under the accession number GSE148574.
Reads were mapped against the reference genome, Oryza sativa Japonica (IRGSP-1.0). To identify genes with significant differences in expression, an FDR cutoff < 0.01 and log2FC -0.5 ≤ or ≥ 0.5 was applied. Gene Ontology (GO) enrichment analysis (GOEA) was performed with AgriGO based on the hypergeometric test, and a minimum FDR of 0.05 was considered. Multiple testing corrections controlling false positives were also performed with the Benjamini–Hochberg method both for differential expression analysis and GOEA. (https://bioinfo.cau.edu.cn/agriGOv2/)68. Enriched GO terms were clustered and plotted with the online analysis tool ReviGO (https://revigo.irb.hr/)69.
Expression analysis by qRT-PCR
Total RNA was extracted as described above. Quantitative RT-PCR (qRT-PCR) analyses were performed using 1 μg of total RNA and poly-dT using the High Capacity cDNA Reverse Transcription kit (Life technology, Applied Biosystems). PCR amplification was carried out using LightCycler 480 thermocycler (Roche Diagnostics, Mannheim, Germany) from 2 μl cDNA (5 ng/μl) with SYBR Green I dye and gene-specific primers (Supplementary Table S7). The rice Ubiquitin1 (Ubi1) or Ubiquitin-conjugating Enzyme 1 (UBC1) were used as housekeeping gene for normalization with the same results. For statistical analysis, the means and standard errors were calculated using Microsoft Excel. Significant differences were assessed using ANOVA test (p-value ≤ 0·05).
Chlorophyll and Pi content
Rice plants were F. mosseae-inoculated or not, and grown under greenhouse conditions as described above. At 23 days after AMF inoculation, the youngest totally expanded leaves were harvested. Chlorophyll content of rice leaves was determined using a chlorophyll meter (SPAD 502 Plus Chlorophyll Meter, Spectrum Technologies) as previously described65. Pi content of rice leaves was determined using a colorimetric as previously described70.
Data availability
The RNA‐Seq data that support the findings of this study are available at the NCBI GEO repository (https://www.ncbi.nlm.nih.gov/geo) under the Accession Number GSE148574.
References
Parniske, M. Arbuscular mycorrhiza: the mother of plant root endosymbioses. Nat. Rev. Microbiol. 6, 763–775 (2008).
Bonfante, P. & Genre, A. Mechanisms underlying beneficial plant–fungus interactions in mycorrhizal symbiosis. Nat. Commun. 1, 48 (2010).
Smith, S. E. & Read, D. J. Mycorrhizal Symbiosis (Academic Press, Cambridge, 2008).
Choi, J., Summers, W. & Paszkowski, U. Mechanisms underlying establishment of arbuscular mycorrhizal symbioses. Annu. Rev. Phytopathol. 56, 135–160 (2018).
MacLean, A. M., Bravo, A. & Harrison, M. J. Plant signaling and metabolic pathways enabling arbuscular mycorrhizal symbiosis. Plant Cell 29, 2319–2335 (2017).
Müller, L. M. & Harrison, M. J. Phytohormones, miRNAs, and peptide signals integrate plant phosphorus status with arbuscular mycorrhizal symbiosis. Curr. Opin. Plant Biol. 50, 132–139 (2019).
Luginbuehl, L. H. et al. Fatty acids in arbuscular mycorrhizal fungi are synthesized by the host plant. Science 356, 1175–1178 (2017).
Begum, N. et al. Role of arbuscular mycorrhizal fungi in plant growth regulation: implications in abiotic stress tolerance. Front. Plant Sci. 10, 1068 (2019).
Campos-Soriano, L., García-Martínez, J. & Segundo, B. S. The arbuscular mycorrhizal symbiosis promotes the systemic induction of regulatory defence-related genes in rice leaves and confers resistance to pathogen infection. Mol. Plant Pathol. 13, 579–592 (2012).
Wang, Y.-Y., Yin, Q.-S., Qu, Y., Li, G.-Z. & Hao, L. Arbuscular mycorrhiza-mediated resistance in tomato against Cladosporium fulvum -induced mould disease. J. Phytopathol. 166, 67–74 (2018).
Pozo, M. J. & Azcón-Aguilar, C. Unraveling mycorrhiza-induced resistance. Curr. Opin. Plant Biol. 10, 393–398 (2007).
Gallou, A., Lucero Mosquera, H. P., Cranenbrouck, S., Suárez, J. P. & Declerck, S. Mycorrhiza induced resistance in potato plantlets challenged by Phytophthora infestans. Physiol. Mol. Plant Pathol. 76, 20–26 (2011).
Rivero, J., Álvarez, D., Flors, V., Azcón-Aguilar, C. & Pozo, M. J. Root metabolic plasticity underlies functional diversity in mycorrhiza-enhanced stress tolerance in tomato. New Phytol. 220, 1322–1336 (2018).
Fiorilli, V. et al. Omics approaches revealed how arbuscular mycorrhizal symbiosis enhances yield and resistance to leaf pathogen in wheat. Sci. Rep. 8, 1–18 (2018).
Wipf, D., Krajinski, F., Tuinen, D., Recorbet, G. & Courty, P. Trading on the arbuscular mycorrhiza market: from arbuscules to common mycorrhizal networks. New Phytol. 223, 1127–1142 (2019).
Ruiz-Sánchez, M., Aroca, R., Muñoz, Y., Polón, R. & Ruiz-Lozano, J. M. The arbuscular mycorrhizal symbiosis enhances the photosynthetic efficiency and the antioxidative response of rice plants subjected to drought stress. J. Plant Physiol. 167, 862–869 (2010).
Vallino, M., Greppi, D., Novero, M., Bonfante, P. & Lupotto, E. Rice root colonisation by mycorrhizal and endophytic fungi in aerobic soil. Ann. Appl. Biol. 154, 195–204 (2009).
Bernaola, L. et al. Natural colonization of rice by arbuscular mycorrhizal fungi in different production areas. Rice Sci. 25, 169–174 (2018).
Gutjahr, C. et al. Transcriptome diversity among rice root types during asymbiosis and interaction with arbuscular mycorrhizal fungi. Proc. Natl. Acad. Sci. USA 112, 6754–6759 (2015).
Tian, L. et al. Comparative study of the mycorrhizal root transcriptomes of wild and cultivated rice in response to the pathogen Magnaporthe oryzae. Rice 12, 35 (2019).
Siciliano, V. et al. Transcriptome analysis of arbuscular mycorrhizal roots during development of the prepenetration apparatus. Plant Physiol. 144, 1455–1466 (2007).
Watts-Williams, S. J. et al. Diverse Sorghum bicolor accessions show marked variation in growth and transcriptional responses to arbuscular mycorrhizal fungi. Plant. Cell Environ. 42, 1758–1774 (2019).
Pimprikar, P. & Gutjahr, C. Transcriptional regulation of arbuscular mycorrhiza development. Plant Cell Physiol. 59, 673–690 (2018).
Das, D. & Gutjahr, C. Role of phytohormones in arbuscular mycorrhiza development. in The Model Legume Medicago truncatula 485–500 (Wiley, Hoboken, 2020). https://doi.org/10.1002/9781119409144.ch61
Pozo, M. J., López-Ráez, J. A., Azcón-Aguilar, C. & García-Garrido, J. M. Phytohormones as integrators of environmental signals in the regulation of mycorrhizal symbioses. New Phytol. 205, 1431–1436 (2015).
Davidson, H. et al. Spatial effects and GWA mapping of root colonization assessed in the interaction between the rice diversity panel 1 and an arbuscular mycorrhizal fungus. Front. Plant Sci. 10, 633 (2019).
Heilmann, I. Phosphoinositide signaling in plant development. Development 143, 2044–2055 (2016).
Raboy, V. Approaches and challenges to engineering seed phytate and total phosphorus. Plant Sci. 177, 281–296 (2009).
Laha, D. et al. Arabidopsis ITPK1 and ITPK2 have an evolutionarily conserved phytic acid kinase activity. ACS Chem. Biol. 14, 2127–2133 (2019).
Laha, D. et al. VIH2 regulates the synthesis of inositol pyrophosphate InsP8 and jasmonate-dependent defenses in arabidopsis. Plant Cell 27, 1082–1097 (2015).
Zhu, J. et al. Two bifunctional inositol pyrophosphate kinases/phosphatases control plant phosphate homeostasis. Elife 8, 1 (2019).
Kuo, H. F. et al. Arabidopsis inositol phosphate kinases IPK1 and ITPK1 constitute a metabolic pathway in maintaining phosphate homeostasis. Plant J. 95, 613–630 (2018).
Dong, J. et al. Inositol pyrophosphate InsP8 acts as an intracellular phosphate signal in Arabidopsis. Mol. Plant 12, 1463–1473 (2019).
Jung, J. Y., Ried, M. K., Hothorn, M. & Poirier, Y. Control of plant phosphate homeostasis by inositol pyrophosphates and the SPX domain. Curr. Opin. Biotechnol. 49, 156–162 (2018).
Mosblech, A. et al. Phosphoinositide and inositolpolyphosphate signalling in defense responses of Arabidopsis thaliana challenged by mechanical wounding. Mol. Plant 1, 249–261 (2008).
Sheard, L. B. et al. Jasmonate perception by inositol-phosphate-potentiated COI1-JAZ co-receptor. Nature 468, 400–407 (2010).
Mosblech, A., Thurow, C., Gatz, C., Feussner, I. & Heilmann, I. Jasmonic acid perception by COI1 involves inositol polyphosphates in Arabidopsis thaliana. Plant J. 65, 949–957 (2011).
Bedini, A., Mercy, L., Schneider, C., Franken, P. & Lucic-Mercy, E. Unraveling the initial plant hormone signaling, metabolic mechanisms and plant defense triggering the endomycorrhizal symbiosis behavior. Front. Plant Sci. 871, 1800 (2018).
Gutjahr, C., Siegler, H., Haga, K., Iino, M. & Paszkowski, U. Full establishment of arbuscular mycorrhizal symbiosis in rice occurs independently of enzymatic jasmonate biosynthesis. PLoS ONE 10, e0123422 (2015).
Nguyen, H. T., To, H. T. M., Lebrun, M., Bellafiore, S. & Champion, A. Jasmonates—the master regulator of rice development, adaptation and defense. Plants 8, 339 (2019).
Lee, H. Y. et al. Oryza sativa COI homologues restore jasmonate signal transduction in Arabidopsis coi1-1 mutants. PLoS ONE 8, e52802 (2013).
Chen, J. et al. The rice CK2 kinase regulates trafficking of phosphate transporters in response to phosphate levels. Plant Cell 27, 711–723 (2015).
Puga, M. I. et al. Novel signals in the regulation of Pi starvation responses in plants: facts and promises. Curr. Opin. Plant Biol. 39, 40–49 (2017).
Zhou, J. et al. OsPHR2 is involved in phosphate-starvation signaling and excessive phosphate accumulation in shoots of plants. Plant Physiol. 146, 1673–1686 (2008).
Rubio, V. et al. A conserved MYB transcription factor involved in phosphate starvation signaling both in vascular plants and in unicellular algae. Genes Dev. 15, 2122–2133 (2001).
Guo, M. et al. Integrative comparison of the role of the PHOSPHATE RESPONSE1 subfamily in phosphate signaling and homeostasis in rice. Plant Physiol. 168, 1762–1776 (2015).
Dai, X., Wang, Y. & Zhang, W.-H. OsWRKY74, a WRKY transcription factor, modulates tolerance to phosphate starvation in rice. J. Exp. Bot. 67, 947–960 (2016).
Yang, W. T. et al. Overexpression of OsMYB4P, an R2R3-type MYB transcriptional activator, increases phosphate acquisition in rice. Plant Physiol. Biochem. 80, 259–267 (2014).
Yi, K. et al. OsPTF1, a novel transcription factor involved in tolerance to phosphate starvation in rice. Plant Physiol. 138, 2087–2096 (2005).
Lin, S.-I. et al. Complex regulation of two target genes encoding SPX-MFS proteins by rice miR827 in response to phosphate starvation. Plant Cell Physiol. 51, 2119–2131 (2010).
Wang, C. et al. Functional characterization of the rice SPX-MFS family reveals a key role of OsSPX-MFS1 in controlling phosphate homeostasis in leaves. New Phytol. 196, 139–148 (2012).
Pfaff, J., Denton, A. K., Usadel, B. & Pfaff, C. Phosphate starvation causes different stress responses in the lipid metabolism of tomato leaves and roots. Biochim. Biophys. Acta 186, 158763 (2020).
Chiou, T.-J. et al. Regulation of phosphate homeostasis by microRNA in Arabidopsis. Plant Cell 18, 412–421 (2006).
Fujii, H., Chiou, T.-J., Lin, S.-I., Aung, K. & Zhu, J.-K. A miRNA involved in phosphate-starvation response in Arabidopsis. Curr. Biol. 15, 2038–2043 (2005).
Hu, B. et al. LEAF TIP NECROSIS1 plays a pivotal role in the regulation of multiple phosphate starvation responses in rice. Plant Physiol. 156, 1101–1115 (2011).
Güimil, S. et al. Comparative transcriptomics of rice reveals an ancient pattern of response to microbial colonization. Proc. Natl. Acad. Sci. U. S. A. 102, 8066–8070 (2005).
Gutjahr, C. et al. Arbuscular mycorrhiza-specific signaling in rice transcends the common symbiosis signaling pathway. Plant Cell 20, 2989–3005 (2008).
Lan, P., Li, W. & Schmidt, W. Complementary proteome and transcriptome profiling in phosphate-deficient arabidopsis roots reveals multiple levels of gene regulation. Mol. Cell. Proteomics 11, 1156–1166 (2012).
Woo, J. et al. The response and recovery of the Arabidopsis thaliana transcriptome to phosphate starvation. BMC Plant Biol. 12, 62 (2012).
Hoffmann-Benning, S. Transport and function of lipids in the plant phloem. AOCS Lipid Library, Plant Biochemistry. (2015). https://lipidlibrary.aocs.org/chemistry/physics/plant-lipid/transport-and-function-of-lipids-in-the-plant-phloem.
Guelette, B., Benning, U. & Hoffmann-Benning, S. Identification of lipids and lipid-binding proteins in phloem exudates from Arabidopsis thaliana. J. Exp. Bot. 63, 3603–3616 (2012).
Barbaglia, A. M., Tamot, B., Greve, V. & Hoffmann-Benning, S. Phloem proteomics reveals new lipid-binding proteins with a putative role in lipid-mediated signaling. Front. Plant Sci. 7, 563 (2016).
Puga, M. I. et al. SPX1 is a phosphate-dependent inhibitor of phosphate starvation response 1 in Arabidopsis. Proc. Natl. Acad. Sci. USA 111, 14947–14952 (2014).
Wang, Z. et al. Rice SPX1 and SPX2 inhibit phosphate starvation responses through interacting with PHR2 in a phosphate-dependent manner. Proc. Natl. Acad. Sci. USA 111, 14953–14958 (2014).
Sánchez-Sanuy, F. et al. Osa-miR7695 enhances transcriptional priming in defense responses against the rice blast fungus. BMC Plant Biol. 19, 563 (2019).
Berruti, A. et al. Application of laser microdissection to identify the mycorrhizal fungi that establish arbuscules inside root cells. Front. Plant Sci. 4, 135 (2013).
Trouvelot, A. Mesure du taux de mycorhization VA d’un systeme radiculaire. Recherche de methodes d’estimation ayant une significantion fonctionnelle (1986).
Du, Z., Zhou, X., Ling, Y., Zhang, Z. H. & Su, Z. AgriGO: a GO analysis toolkit for the agricultural community. Nucleic Acids Res. 38, W64-70 (2010).
Supek, F., Bosnjak, M., Skunca, N. & Smuc, T. REVIGO Summarizes and visualizes long lists of gene ontology terms. PLoS ONE 6, 1–9 (2011).
Ames, B. N. Assay of inorganic phosphate, total phosphate and phosphatases. Methods Enzymol. 8, 115–118 (1966).
Acknowledgements
This research was supported by the FACCE-JPI project GreenRice (Sustainable and environmental friendly rice cultivation systems in Europe, ref. 618105), funded by the Instituto Nacional de Investigación y Tecnología Agraria y Alimentaria (INIA), and Project RTI2018-101275-B-I00 funded by the “Ministerio de Ciencia, Innovación y Universidades (MCIU)-Agencia Estatal de Investigación (AEI)” and “Fondo Europeo de Desarrollo Regional (FEDER). We also acknowledge support from the CERCA Programme (“Generalitat de Catalunya”), and the “Severo Ochoa Programme for Centres of Excellence in R&D” 2016-2019 (SEV-2015-0533).
Author information
Authors and Affiliations
Contributions
S.C. did the experimental work and analyzed the data; S.C. and B.S.S. conceived the experiments and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Campo, S., San Segundo, B. Systemic induction of phosphatidylinositol-based signaling in leaves of arbuscular mycorrhizal rice plants. Sci Rep 10, 15896 (2020). https://doi.org/10.1038/s41598-020-72985-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-72985-6
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.