Abstract
Genome editing has become one of the key technologies for plant breeding. However, in polyploid species such as chrysanthemum, knockout of all loci of multiple genes is needed to eliminate functional redundancies. We identified six cDNAs for the CmDMC1 genes involved in meiotic homologous recombination in chrysanthemum. Since all six cDNAs harbored a homologous core region, simultaneous knockout via TALEN-mediated genome editing should be possible. We isolated the CmDMC1 loci corresponding to the six cDNAs and constructed a TALEN-expression vector bearing a CmDMC1 target site containing the homologous core region. After transforming two chrysanthemum cultivars with the TALEN-expression vector, seven lines exhibited disruption of all six CmDMC1 loci at the target site as well as stable male and female sterility at 10–30 °C. This strategy to produce completely sterile plants could be widely applicable to prevent the risk of transgene flow from transgenic plants to their wild relatives.
Similar content being viewed by others
Introduction
Chrysanthemum (Chrysanthemum morifolium Ramat.) is one of the best-known cultivated ornamental flowers worldwide. Contemporary chrysanthemum cultivars are hexaploids with loss or gain of several chromosomes1 and display self-incompatibility2. Many agronomical traits have been introduced via conventional cross- and mutation breeding. However, these technologies are potentially limited by the gene resources available and the pathways modifiable by crossing and/or mutation.
Genetic transformation technologies are useful for introducing agronomical traits that cannot be achieved by conventional mutation breeding. Almost 50 ornamental flowers have been transformed to express modified traits3. In many countries, growing genetically modified (GM) plants is tightly regulated under the guidelines and directives of the Cartagena Protocol on Biosafety4,5. In particular, ornamentals require assessment for outcrossing to wild relatives prior to practical use.
Transgenic carnations (Dianthus caryophyllus L.) and roses (Rosa x hybrida) producing blue flowers are now marketed under the terms of the Cartagena Protocol. Carnation flowers generate little mature pollen, and no hybrids between carnations and their wild relatives have been reported so far6. Transgenic bluish roses are genetic chimeras whose transgenes are not transmitted to pollen7. In chrysanthemum, many useful traits have been introduced by transformation, including disease resistance8,9, resistance against insects and fungal disease10 and modified flower color11. However, transgenic chrysanthemums are not yet sanctioned for open field cultivation because of their cross-compatibility with wild relatives. F1 plants between non-GM commercial chrysanthemums and their wild relatives are known to be distributed widely in several habitats of the wild species12.
To inhibit transgene flow, Aida et al.13 reported periclinal L1 chimeric plants in chrysanthemum, as transgenes in the L1 layer are rarely transmitted to progeny. While this is very useful for modification of plant surface characteristics such as flower color, in order to use plants genetically modified to confer insect- or disease-resistance, induction of both male and female sterility by suppressing or disrupting genes involved in gametogenesis is required.
The protein DMC1 plays a critical role in meiotic homologous recombination. Knocking out or mutating of DMC1 causes serious defects in meiotic DNA recombination, asynapsis and sterility in yeast14, mouse15, Arabidopsis16 and rice17. Shinoyama et al.18 introduced an RNA interference (RNAi) construct of a DMC1 gene (designated CmDMC1a in this study) in chrysanthemum. Regenerated CmDMC1a-RNAi plants exhibited severe male sterility and a significant reduction in female fertility. However, due to their hexaploid nature and self-incompatible trait2, transgenic chrysanthemum plants showing both male and female sterility are needed to completely inhibit transgene flow. This study thus aimed to produce chrysanthemum plants with simultaneous knockout of all six endogenous CmDMC1 loci using transcription activator-like effector nucleases (TALEN)-mediated genome editing technology.
To simultaneously disrupt all DMC1 genes, we exploited the extraordinary level of conservation among DMC1 protein and DMC1 cDNA sequences19, identifying suitable target sequences common to all six chrysanthemum DMC1 genes. The resulting plants exhibited both male and female sterility, suggesting that knockout of all DMC1 alleles could be used to induce stable male and female sterility in chrysanthemum and other highly polyploid plants, and thus prevent transgene flow from GM plants to their wild relatives.
Results
Isolation of cDNAs and partial sequences of DMC1 genes from chrysanthemum
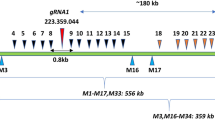
Based on the cDNA sequence of CmDMC1a (submitted to DDBJ under the accession number: LC575211)18, five additional CmDMC1 cDNAs (CmDMC1b–CmDMC1f, the accession numbers: LC575212, LC575213, LC575214, LC575215 and LC575216, respectively) were isolated from chrysanthemum cultivar ‘Shuho-no-chikara’ (Supplementary Fig. S1). Nucleotide sequence similarities (homology) of each cDNA for CmDMC1b to CmDMC1f compared to CmDMC1a were 99.2%, 99.5%, 99.1%, 99.0% and 99.3%, respectively. Amino acid sequence identities between CmDMC1b to CmDMC1f and CmDMC1a were 99.1%, 99.1%, 98.3%, 98.0% and 99.1%, respectively (Supplementary Fig. S2). These six cDNA clones were also isolated from the other nine cultivars (Supplementary Table S1). No novel cDNA clones were found in any of the nine cultivars, suggesting that there are six active DMC1 loci in chrysanthemum. Respective genomic DNAs corresponding to nucleotide positions 234–815 in the six cDNAs were isolated from ‘Shuho-no-chikara’. These sequences include the homologous core region used for the CmDMC1a-RNAi study18. Numbers of nucleotides varied among the six loci: 1782 bp (CmDMC1a), 1722 bp (CmDMC1b), 1727 bp (CmDMC1c), 1713 bp (CmDMC1d), 1709 bp (CmDMC1e), and 1714 bp (CmDMC1f) (Fig. 1), possibly due to differences in the number of nucleotides in intron(s) between exons among CmDMC1 genes. Since an exon sequence at nucleotide positions 1363 to 1452 bp in CmDMC1a (Fig. 1) is located in the conversed core region and identical to corresponding sequences in the other five CmDMC1 genes, the target sequence for the TAL effector repeat array was designed in this region for simultaneous disruption of all CmDMC1 alleles using the system TAL Effector Nucleotide Targeter 2.0 system developed by Cornell University (https://tale-nt.cac.cornell.edu/node/add/talen-old) (Fig. 1, Supplementary Table S2). This target sequence includes the multimer site (BRC) interface located upstream of the Walker B motif20.
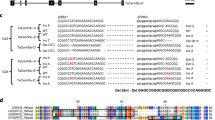
Partial genomic DNA sequences of the six CmDMC1 genes of Chrysanthemum morifolium cultivar ‘Shuho-no-chikara’. Genomic DNAs corresponding to nucleotide positions 234–815 in the six cDNAs (Supplementary Fig. S1) are aligned with the DNASIS version 3.7 (Hitachi Software Engineering). Uppercase letters in blue indicate exon, and lower-case letters indicate intron. Asterisks (*) indicate the same nucleotides as in CmDMC1a; red letters indicate different nucleotides compared to CmDMC1a; red bar (-) indicates nucleotide deletion. TAL-L and TAL-R indicate the TALEN pair targeting CmDMC1 genes. A primer pair, DMC1-RNAi F1 and DMC1-RNAi R1, was used to amplify partial DNA fragments of individual CmDMC1 genes that correspond to cDNA nucleotide positions 234–815 of ‘Shuho-no-chikara’ (Supplementary Fig. S1). A primer pair, DMC1-gF1 and DMC1-gR1, was designed for the detection of mutations around the recognition sequences of the TALENs.
Production of CmDMC1 knockout plants using TALEN-mediated targeted mutagenesis
Infection of chrysanthemum leaf discs with Agrobacterium tumefaciens strain EHA10521 harboring a binary vector pBIK201DMC-TAL containing the TALEN pair targeting CmDMC1 loci (Fig. 2), and selection and regeneration of transgenic plants were performed as shown in Supplementary Table S3. From the two cultivars ‘Shuho-no-chikara’ and ‘Yamate-shiro’, 23 and 126 plantlets, respectively, were regenerated from calli resistant to 20 mg l−1 G418, giving regeneration rates of 3.2% and 17.5% from 719 and 720 initial leaf segments (Supplementary Table S4).
Structure of binary vector pBIK201DMC-TAL for disruption of six CmDMC1 genes in chrysanthemum. RB, right border; LB, left border; Pmas201, bidirectional promoter cassette from mannopine synthase 1′ and 2′ (mas1′-2′) genes; T35S, Cauliflower mosaic virus 35S terminator; Tnos, nopaline synthase terminator; nptII, neomycin phosphotransferase II gene for the selection of transgenic plants; TAL-L and TAL-R, the TALEN pair targeting CmDMC1 genes; Fok I, gene encoding a restriction enzyme. T2A, encoding a peptide for self-cleavage and ribosome skipping69. Red bar indicates the nptII-specific probe (about 800 bp) for Southern blotting in Supplementary Fig. S3.
The presence and number of T-DNA copies in individual regenerated plants (defined as the T1 generation) were confirmed by Southern blot analysis of XbaI-digested genomic DNA probed with a fragment of the neomycin phosphotransferase II (nptII). Theoretically, a unique fragment over 1.5 kbp (Fig. 2) could be detected when a single copy of T-DNA is integrated into the chrysanthemum genome. Single to multiple bands hybridizing to the nptII probe were observed in the regenerated plantlets. Among them, the results for four lines (SH#12, SHa#13, YS#16 and YSa#12) are shown in Supplementary Fig. S3. No hybridization signals were detected in non-transgenic chrysanthemum controls.
For the detection of mutations in individual CmDMC1 genes (CmDMC1a–f), a fragment of about 300 bp containing the TALEN recognition sequences was amplified by PCR using the primer pair DMC1-gF1 and DMC1-gR1 (see Fig. 1), and genomic DNA from a CmDMC1-TALEN line as a template. Each DNA amplicon was cloned and sequenced to identify individual CmDMC1 genes. Sequences of the clones carrying particular DNA amplicons were analyzed to identify and classify mutations in individual CmDMC1 genes. Typically, three mutation patterns were detected in each TALEN-targeted locus. The first was a mutation in a single allele of a specific CmDMC1 gene (monoallelic mutation). The rest were mutations in both alleles of a CmDMC1 gene (biallelic mutations) that consisted of two types: an identical mutation in both alleles, and different mutations in the two alleles of a single CmDMC1 gene.
DNA sequence analysis of six CmDMC1 genes in the 23 CmDMC1-TALEN lines of ‘Shuho-no-chikara’ revealed that five lines showed biallelic mutations in all six loci (genotype e.g. aabbccddeeff, where each lowercase letter indicates a mutated allele for each CmDMC1 locus). The remaining lines showed biallelic mutations in five loci, and a monoallelic mutation in one locus (genotype e.g. aaBbccddeeff, where the uppercase B indicates the wild-type allele and Bb denotes a monoallelic mutation in CmDMC1b) (Supplementary Table S4). Similarly, among 126 CmDMC1-TALEN lines of ‘Yamate-shiro’, two lines showed mutations in all six target loci (biallelic mutations) (Supplementary Table S4). For each cultivar, we selected seven lines that possessed biallelic mutations in all six CmDMC1 genes, or in five genes with one wild-type sequence to elucidate the effect of each gene. Among a total of 84 CmDMC1 genes (168 alleles) from 14 TALEN lines, 72 genes (144 alleles) were mutated. Deletion mutations were detected most frequently, and insertion mutations including a combination of insertion and deletion were detected only rarely (Table 1, Supplementary Table S5 and Supplementary Fig. S4). Biallelic mutation patterns with identical mutations in both alleles of a particular CmDMC1 gene (e.g. CmDMC1a, b, e and f in SH#12) were detected in 37 genes (74 alleles). Biallelic and different mutation patterns in the two alleles of a CmDMC1 gene (e.g. CmDMC1c and d in SH#12) were found in 35 genes (70 alleles) —a frequency similar to that of the identical mutation patterns (Supplementary Table S5). Frameshift mutations were caused by INDELs (insertion or deletion) of a number of nucleotides that is not divisible by three; in this case, stop codons appeared just downstream of the mutated site (Supplementary Table S6). On the other hand, INDELs involving multiples of 3 nucleotides caused insertion/deletion of a few amino acids (e.g. CmDMC1d in SHa#13 and SHf#20, and CmDMC1e in SHb#14 and YSc#27). Mutation patterns at each locus were examined in two different tissues, leaf and root, and found to be identical in these two tissues in every transgenic line analyzed (Table 1). These results suggested that the tissues (leaves and roots) in each regenerated transgenic plant were differentiated from a small callus possibly derived from a single or a few transgenic cells, and that each respective CmDMC1 locus is represented by a specific mutation pattern following TALEN-mediated genome editing. To investigate whether additional mutation(s) occurred outside the CmDMC1-TALEN recognition sites, approximately 1.8 kbp fragments containing the 300 bp regions in the CmDMC1 genes of the 14 TALEN lines were PCR-amplified using total DNAs from leaves or roots of respective lines as templates and the primer pair DMC1-RNAi F1 and DMC1-RNAi R1 (Supplementary Fig. S4), cloned and sequenced. Comparing the 1.8 kbp sequences for CmDMC1a in each TALEN-line with that of wild-type revealed that no additional mutations were generated outside the TALEN target regions (Supplementary Table S5 and Supplementary Fig. S4).
Transgenic chrysanthemums carrying the TALENs construct for CmDMC1 and non-transgenic chrysanthemums were grown at 20 °C and analyzed for CmDMC1 transcripts in anthers and ovaries by northern blotting. The expected size of the CmDMC1 mRNA was ca. 1.3 kbp. A strong hybridization signal was detected in the non-transgenic controls, while no signals were detected in anthers and ovaries of the transgenic lines carrying mutations in all 12 alleles (SH#12 and YS#16 in Fig. 3 and Supplementary Fig. S5). Lines SHc#15 and YSc#27 bore biallelic mutations in five CmDMC1 genes including CmDMC1a and CmDMC1b, and showed no signals in anthers, but weak signals in ovaries. Lines SHa#13 and YSa#12 did not possess mutation except in CmDMC1a, and exhibited moderate signals in both anthers and ovaries (Fig. 3, Supplementary Fig. S5).
Expression levels of CmDMC1 genes in CmDMC1-TALEN chrysanthemum plants. To detect endogenous CmDMC1 transcripts, total RNA was isolated from anthers and ovaries at early meiotic division stage before tetrad formation of CmDMC1-TALEN and non-transgenic control plants grown at 20 °C. Twenty μg of total RNA was applied to each lane. The RNA blots were probed with a 1032-bp fragment of the CmDMC1a cDNA as in Supplementary Fig. S1, and 1134-bp of the ACTIN cDNA of Ch. morifolium. SH and YS: RNA from non-transgenic controls ‘Shuho-no-chikara’ and ‘Yamate-shiro’, respectively. SH#12, SHa#13 and SHc#15 in panels A, C, E and G: TALEN lines from ‘Shuho-no-chikara’, and YS#16, YSa#12 and YSc#27 in panels B, F, D and H: those from ‘Yamate-shiro’. Panels A, B, C and D show RNA blots of anthers and panels E, F, G and H show those of ovaries. Their full-length blots are presented in Supplementary Fig. 5.
Growth characteristics of the transgenic lines SH#12 and YS#16 were compared to non-transgenic controls in a bio-safety containment greenhouse (hereinafter referred to as greenhouse). No obvious differences in growth and morphology were detected (Supplementary Table S7, Supplementary Fig. S6).
Analysis of male sterility in CmDMC1-TALEN chrysanthemum plants
Tubular flowers were collected from the CmDMC1-TALEN lines and non-transgenic controls 1 day before flowering (Supplementary Fig. S7A, B) and stained by Alexander staining solution22 for analysis of male sterility (Figs. 4 and 5, Supplementary Table S8, Supplementary Fig. S8). No pollen grains were observed in anthers of the non-transgenic controls grown at 30 °C, whereas viable pollen grains were observed in anthers of non-transgenic controls at 10–25 °C. Rates of viable pollen grains in anthers of the CmDMC1-TALEN lines were evaluated according to Alexander’s method22. Rates of viable pollen grains in the controls at 20 °C were over 80%, but the rates declined to about 60% at 25 °C, 50% at 15 °C and 30–40% at 10 °C (Supplementary Table S8). No viable pollen grains were observed in the two CmDMC1-TALEN lines SH#12 and YS#16 with mutation in all the CmDMC1 genes and without CmDMC1 transcript at all temperature ranges tested. Two CmDMC1-TALEN lines, SHc#15 and YSc#27 bearing biallelic mutation in five genes including CmDMC1a and CmDMC1b (Table 1 and Supplementary Table S5), displayed no pollen grains at all temperature ranges tested (Fig. 4, Supplementary Table S8, Supplementary Fig. S8). In contrast, CmDMC1-TALEN lines SHa#13 and YSa#12, carrying biallelic mutations in five of the six genes with the exception of CmDMC1a (Table 1), produced viable pollen grains when grown at 15–25 °C (Figs. 4 and 5, Supplementary Fig. S8). Viable pollen produced by the non-transgenic control was somewhat sporadic. However, aborted pollen grains were observed more frequently than viable pollen grains in CmDMC1-TALEN lines such as SHa#13 and YSa#12. Aborted pollen grains seemed to be at the tetrad stage and were stuck to each other, so they looked larger than viable pollen. The rates of viable pollen in TALEN-mutated lines were significantly lower than those in non-transgenic controls according to ANOVA test.
Evaluation of male sterility in CmDMC1-edited plants and non-transgenic controls by Alexander staining of pollen grains. (A) CmDMC1-edited plants of ‘Shuho-no-chikara’ and (B) CmDMC1-edited plants of ‘Yamate-shiro’. Red and blue bars indicate viable and aborted pollen grains, respectively. ** significant at 1% by ANOVA.
Assessment of pollen viability in CmDMC1-edited chrysanthemum plants. One day before flowering, anthers were stained according to Alexander22. Red- and green-colored pollen grains were judged to be viable and aborted, respectively. Scale bars indicate 0.2 mm (anthers) and 0.1 mm (magnified images of pollen at the center). SH and YS indicate non-transgenic controls ‘Shuho-no-chikara’ and ‘Yamate-shiro’. SH#12, SHa#13 and SHc#15 indicate CmDMC1-TALEN lines from ‘Shuho-no-chikara’, and YS#16, YSa#12 and YSc#27 from ‘Yamate-shiro’.
Analysis of female sterility in CmDMC1-TALEN chrysanthemum plants
Viable pollen grains from the wild relatives grown at 20 °C were collected and pollinated to stigmas of CmDMC1-TALEN lines and non-transgenic controls. In addition, these TALEN lines and non-transgenic controls were self-pollinated. Two types of achenes (F1 seeds) were observed 2 months after crossing and self-pollination (Supplementary Fig. S9). We evaluated the F1 seeds with germination ability as viable (A, C, G and I in Supplementary Fig. S9) and the ones without germination ability as aborted (B, D, E, F, H, J, K and L in Supplementary Fig. S9). Seed coats of the viable F1 seeds were hard and relatively bulgy, whereas the aborted F1 seeds were very small and fragile. The ratios of viable F1 seeds were 68.4–79.6% for non-transgenic controls that were grown, pollinated and matured at 20 °C, but these ratios declined to ca. 30% at 25 °C, 20% at 30 °C and 10% at 15 °C (Fig. 6, Supplementary Table S9). These results indicated that the ovules of non-transgenic controls are fertile. On the other hand, the two CmDMC1-TALEN lines SH#12 and YS#16 with biallelic mutations in all alleles gave no viable F1 seeds at any of the temperature ranges used for growth and pollination (Fig. 6, Supplementary Table S9), indicating that ovules of SH#12 and YS#16 are sterile. The ratios of viable F1 seeds were ca. 1%, and low levels of transcripts were detected by northern blot analysis of RNA from ovaries of CmDMC1-TALEN lines grown and pollinated at 20 °C with biallelic mutations in five CmDMC1 genes including CmDMC1a and CmDMC1b (data for SHc#15 and YSc#27 in Fig. 3 and Supplementary Fig. S5). The four CmDMC1-TALEN lines SHa#13, SHb#14, YSa#12 and YSb#13 with biallelic mutations in five CmDMC1 genes and without mutation in one of the two genes, CmDMC1a or CmDMC1b (Table 1), exhibited a ratio of viable F1 seeds of 1–2% when grown, pollinated and matured at 15 °C and about 6–7% at 20 °C (Supplementary Table S9), and intermediate levels of CmDMC1 transcripts were detected by northern blot analysis (Fig. 3, Supplementary Fig. S5). The ratios of viable F1 seeds in the CmDMC1-TALEN lines were significantly lower than those in non-transgenic controls, according to Tukey–Kramer’s HSD test (Fig. 6, Supplementary Table S9).
Evaluation of female sterility by crossing between transgenic lines and their wild relatives. Crossing between CmDMC1-TALEN plants of ‘Shuho-no-chikara’ and Ch. wakasaense (A) or Ch. japonense (B). Crossing between CmDMC1-edited plants of ‘Yamate-shiro’ and Ch. wakasaense (C) or Ch. japonense (D). Each value followed by the same letter is not significantly different at 5% level by Tukey–Kramer’s HSD.
Thus, our results indicate that the biallelic knockout by TALEN of six CmDMC1 loci confers to reduce both female and male fertility on chrysanthemums over a wide range of flowering temperatures, including the optimum growth temperature for chrysanthemums, i.e., between 17 and 22°C23.
Discussion
The introduction of targeted DNA double-strand breaks (DSBs) via sequence-specific nucleases (SSNs), such as meganucleases24, zinc finger nucleases (ZFNs)25, transcription activator-like effector nucleases (TALENs)26, and the bacterial clustered regularly interspaced short palindromic repeat (CRISPR)/CRISPR-associated protein 9 (Cas9) system27, results in deletions, insertions, and substitutions around the nuclease cleavage sites in the target genes. Recently, SSNs have become useful tools for genome engineering in plants, and these “genome editing” technologies have been established in many plants including Arabidopsis28, rice29,30, wheat31, soybean32, barley33, maize34, potato35,36, sugarcane37, etc.
Although the number of papers reporting successful genome editing in plants is increasing rapidly, there are still few reports mutating multi-alleles in highly polyploid plants using TALENs. For example, bread wheat (Triticum aestivum) is a hexaploid with the genomic constitution AABBDD (2n = 6x = 42) in which each constituent subgenome originated from a different ancestral species. Allopolyploidization leads to the generation of duplicated homoeologous genes (homoeologs) and, consequently, the hexaploid wheat genome contains triplicated homoeologs derived from the three ancestral diploid species. Wang et al.31 succeeded in mutating all three sets of homoeoalleles for MILDEW-RESISTANCE LOCUS (MLO) to produce wheat resistant to powdery mildew. Cultivated potato (Solanum tuberosum) is a highly heterozygous autotetraploid (AAAA, 2n = 4x = 48). Disruption of the gene for sterol side chain reductase (SSR2) in potatoes resulted in low SSR2 activity and a reduction in the levels of two toxic steroidal glycoalkaloids, α-solanine and α-chaconine35,36. Modern sugarcane varieties are complex interspecific hybrids (Saccharum spp.) and highly heterozygous allopolyploid ranging from decaploid to tridecaploid (2n = 10 × to 13x = 100–130). To reduce lignin content and improve biofuel yield, Jung and Altpeter37 induced mutations by TALENs in the gene for caffeic acid O-methyltransferase (COMT), involved in lignin biosynthesis.
Chrysanthemums are hexaploids (2n = 6x = 54) with a loss or gain of several chromosomes1, originated by crossing and duplicating between Ch. zawadskii var. latilobum (Maxim.) Kitamura (2n = 2x = 18) and Ch. indicum var. procumbense (Lour.) Kitamura (2n = 4x = 36)38. Very recently, Kishi-Kaboshi et al.39 succeeded in knocking out multiple transgenes previously integrated in transgenic chrysanthemums using the CRISPR/Cas9 system. They used transgenic chrysanthemum lines harboring more than four or five copies of a gene for yellow-green fluorescent protein from Chiridius poppei (CpYGFP) as materials for gene disruption, and edited all the CpYGFP-transgene alleles simultaneously. In this study, we isolated six CmDMC1 loci and intended to mutagenize the BRC multimer interface just upstream of the Walker B motif20, since Walker A and/or B motifs are functionally essential ATP-binding sites40. Disruption of these motifs by frameshift mutations could destroy DMC1 function, resulting in abnormal meiosis and, consequently, a male and female sterility phenotype. Interestingly, changing even one amino acid at these motifs caused ablation of ATP binding and inactivation of human DMC141,42. Similarly, Dresser et al.43 and Masson and West44 reported that a Saccharomyces cerevisiae DMC1 mutant carrying a single amino acid substitution in the ATP-binding site confers a null mutation.
Previous studies have indicated that the sizes of the INDELs mediated by TALENs are mostly less than 300 bp including the TALEN target sites28,29,30,31,32,33,34,35,36,37. We designed a pair of primers generating amplicons of about 300 bp including the TALEN target sites for CmDMC1 genes to identify deletions in the sites in addition to various insertions and substitutions. Using TALEN-mediated targeted mutagenesis of six CmDMC1 loci, three patterns of mutation were induced. The frequency of monoallelic mutations was lower than that of biallelic mutations. Moreover, biallelic mutations carrying the same mutation in both alleles of a CmDMC1 gene were detected with approximately the same frequency as biallelic mutations carrying different mutations in two alleles of the gene. In calli of rice cultivars, most of the TALEN-induced deletion mutations at Waxy locus were less than 100 bp, and deletions larger than 300 bp occurred rarely30. The largest deletion observed in SSR2-TALEN lines of tetraploid potato was 283 bp36. Accordingly, sequences of larger regions (~ 1.8 kbp) surrounding the TALEN target sites for CmDMC1 loci were analyzed in the TALEN-induced mutant lines of chrysanthemum. The sequencing results detected no novel mutations outside the target site of CmDMC1a locus in each TALEN lines, indicating only small INDELs and substitutions could occur in our system (Table 1, Supplementary Table S5 and Supplementary Fig. S4).
In our TALEN-mediated editing of CmDMC1, both in-frame mutations conferring insertion or deletion of amino acid(s) and frameshift mutations were detected. Looking for putative CmDMC1 products, we noted that the frameshift mutants could generate premature termination codons (PTCs) just downstream of the mutated sites (Supplementary Table S6). Northern blot analysis of CmDMC1 transcripts detected markedly decreased or no hybridization signals in four out of six CmDMC1-TALEN lines: SH#12, SHc#15, YS#16 and YSc#27 (Fig. 3 and Supplementary Fig. S5) bearing frameshift mutations in at least the CmDMC1a and CmDMC1b loci. All the frameshift mutations found in those lines potentially generate aberrant mRNAs carrying PTCs that would be recognized and rapidly degraded by nonsense-mediated mRNA decay (NMD). NMD, which is conserved among eukaryotes, is a mechanism of quantity and quality control of mRNAs that prevents accumulation of detrimental truncated proteins45,46. Regulation of NMD activity plays a crucial role in plant growth and responses to the environment47,48.
In contrast, the allele harboring a deletion of three nucleotides potentially generates deletion of a single amino acid in the respective DMC1 proteins encoded by a single allele of CmDMC1d (lines SHa#13 and SHf#20) or CmDMC1e (SHb#14 and YSc#27). In these four lines, each CmDMC1d or CmDMC1e mutant protein carries a deletion of Lys at position 215 (K215) located in the BRC multimer interface domain, which was proposed to be involved in monomer–monomer interaction20. This Lys residue—in equivalent positions in DMC1 proteins—is shared among mammals and higher plants49 that bear both BRCA and DMC1 genes. Deletion of such a positively charged amino acid at the interface domain might affect DMC1–DMC1 and/or DMC1–BRCA2 interactions50,51,52,53.
The male and female sterility of the 14 CmDMC1-TALEN lines bearing biallelic mutations in five or all six CmDMC1 genes differed significantly from non-transgenic controls. In fact, rates of male and female sterility seem to depend on the mutated CmDMC1 locus. Biallelic mutations of all CmDMC1 genes or both CmDMC1a and CmDMC1b genes caused complete male sterility at 10–30 °C. However, when one of the two genes was not mutated, mature pollen grains were formed at 15–25 °C (Supplementary Table S8, Supplementary Fig. S8).
When CmDMC1a or CmDMC1b was not mutated in CmDMC1-TALEN lines as the female parents, viable F1 seeds with the ability to germinate were formed at 15 and 20 °C. When any one of the CmDMC1c, CmDMC1d, CmDMC1e or CmDMC1f genes was not mutated in the female parents, ratios of viable F1 seeds were much lower only at 20 °C (Fig. 6, Supplementary Table S9). These results showed that female fertility is influenced strongly by CmDMC1a and/or CmDMC1b, and moderately by CmDMC1c, CmDMC1d, CmDMC1e and/or CmDMC1f, and that loss-of-function mutations in all the CmDMC1 loci are required for the loss of crossing ability in female reproductive organs that is independent of ambient temperature. As a future study, microscopic observation of egg cells from the CmDMC1-TALEN lines would be necessary to verify the effects of CmDMC1 mutagenesis by genome editing.
The genes for TALENs and the nptII selection marker were driven by a bidirectional promoter from the mannopine synthase-2′ and -1′ (mas2′-1′) genes of an A. tumefaciens strain as pathogen of the Compositae family. Mannopine synthase is composed of two enzyme conjugates encoded by the mas1′ gene and a reductase encoded by the mas2′ gene54. These two genes are located on the T-DNAs of certain octopine-type Ti and Ri plasmids55,56, and the mas2′ and mas1′ promoters are oriented in a head-to-head manner on a 483-bp fragment of pTiAch555. The mas2′-1′ promoter conferred high level of expression to the modified cry1Ab gene from Bacillus thuringiensis, the nptII gene and the CmDMC1a-RNAi construct in leaves, stems, roots and pollen, and weaker expression levels in ovaries18. The difference in the degree of sterility between male (pollen) and female (ovaries) in CmDMC1a-RNAi chrysanthemums might be due to the differential activity of the promoter controlling RNAi expression in the organs. The CmDMC1-TALEN lines were all mutated at the target sequence, and 5 of 23 regenerated plants in ‘Shuho-no-chikara’ and 2 of 126 regenerated plants in ’Yamate-shiro’ were mutated in all six loci (Supplementary Table S4). According to DNA sequence analysis of the six CmDMC1 loci, these seven lines showed one or two mutation patterns without wild-type sequences at each CmDMC1 locus (ex. SH#12 and YS#16 in Table 1, Supplementary Tables S5 and S6). The frequency of biallelic mutations in the transgenic plants was much higher than that reported in other polyploidy plants mutagenized by TALENs31,35,36,37. This seemed to be due to the use of a mas2′-1′ promoter, whose expression is relatively high at the site of Agrobacterium infection in leaves and in calluses of chrysanthemum. Thus, a stable male and female sterility phenotype could be introduced into chrysanthemum by the proper selection of a TALENs target sequence that is highly conserved among CmDMC1 multi-genes, and by the use of a mas2′-1′ promoter that can express TALENs strongly in chrysanthemum.
In the genetic transformation of chrysanthemum, transgenic plants with chimeric nature were reported. Early studies, that used direct plant regeneration from leaf or stem segments, often showed chimeric nature in the transgenic plants57,58. Transgenic plants generated from somatic embryos, possibly with the single transgenic-cell origin, displayed non-chimerism59. However, number of cultivars suitable for somatic embryogenesis is limited60, and the regeneration system through somatic embryogenesis would not be widely applicable for chrysanthemum transformation. We circumvented the chimerism using a callus induction and regeneration system61 (Supplementary Table S3). By applying relatively strict antibiotic selection under 20 mg l−1 G418 for 2.5 to 3 months during callus formation. Non-transformed calli (escapes) were effectively eliminated and only transformed calli survived on the medium. The formation of chimeras may also occur during the process of producing genome-edited plants62,63. We obtained CmDMC1-TALEN chrysanthemum plants according to the above-mentioned system, and analyzed partial DNA sequences of the six CmDMC1 genes for over 13 individual clones bearing genomic PCR products surrounding TALEN recognition sites (about 300 bp and 1.8 kbp) amplified from total DNAs of leaves or roots of the TALEN lines. The results indicated that no more than two mutation patterns were detected for each CmDMC1 locus (Supplementary Table S5) and that no other mutations were detected outside the target sites for each CmDMC1a locus (Supplementary Fig. S4). Though these results support the non-chimeric nature of our CmDMC1-TALEN lines, more detailed studies would be needed for further confirmation such as the elucidation of the relationship between mutation patterns in the six CmDMC1 genes (genotypes) and sterility phenotypes in the clonal plants.
Since chrysanthemum exhibits self-incompatibility, it is hard to obtain null segregants by self-fertilizaion. Recently, DNA-free genome editing methods have been developed in potato64 and bread wheat65. It is expected that DNA-free methods will be more advantageous for environmental safety assessments than DNA-based methods introducing transgene(s) into the genome.
In this study, we determined six CmDMC1 cDNA sequences and six corresponding partial genomic DNA fragments from 10 chrysanthemum cultivars. However, because some chrysanthemum cultivars have unstable and variable chromosome numbers that form a hexaploid complex with aneuploidy (2n = 6x = 54 ± 7–10)1, a search for other CmDMC1 loci from other cultivars or Compositae family will be needed, as well as an analysis of allelic configurations, including these six loci, to assess the efficiency of mutagenesis by genome editing in the prevention of transgene flow.
The strategy reported here should be useful in preventing transgene flow via pollen. A stable sterility phenotype will be a key technology for the practical use of chrysanthemums transgenic for characteristics such as pest/disease resistance and flower color modifications under the terms of the Cartagena Protocol on Biosafety.
Methods
Plant materials and culture conditions
Ten chrysanthemum (Ch. morifolium Ramat.) cultivars (double flower type: ‘Shuho-no-chikara’, ‘Seiun’, ‘Shinba No. 2’ and ‘Summer Yellow’; single flower type: ‘Yamate-shiro’, ‘Kosuzu’, ‘Utage’, ‘Monroe’, ‘Kofuku-no-tori’ and ‘Kin-fusha’) were used. Shoot tips of plants grown in a greenhouse were surface-sterilized in 70% ethanol and then in a 1% sodium hypochlorite solution for 15 min. They were rinsed three times with sterile distilled water. Shoot tip explants were cultured in vitro (meristem culture) on Murashige and Skoog (MS) basal medium66 containing 3% sucrose and 0.3% gellan gum (FUJUFILM Wako Pure Chemical Corporation, Osaka, Japan). The medium was adjusted to pH 5.8 before autoclaving at 121 °C for 15 min. The explants were cultured at 25 °C under a 16-h photoperiod using cool white fluorescent lamps or at 25 °C in darkness. The lamps provided a photosynthetic photon flux density (PPFD between 400 and 700 nm) of 60 μmol m−2 s−1.
Isolation of CmDMC1 loci from chrysanthemum cultivars
The 10 chrysanthemum cultivars described above were used. In a previous study18, we cloned a partial cDNA encoding a DMC1 protein from chrysanthemum cultivar ‘Shuho-no-chikara’. Based on this sequence, we isolated cDNAs encoding DMC1 homologs from ‘Shuho-no-chikara’ and the other nine cultivars using a SMART RACE cDNA Amplification Kit (Clontech Laboratories, Mountain View, CA, USA) and cDNA libraries of individual cultivars (Supplementary Fig. S1). Partial DNA fragments of individual CmDMC1 genes that correspond to cDNA nucleotide positions 234–815 of ‘Shuho-no-chikara’ (Supplementary Fig. S1) were amplified from genomic DNAs (Fig. 1) using the primer pair DMC1-RNAi F1 (5′-aatctgtgaagctgct-3′) and DMC1-RNAi R1 (5′-tttgtcatatacac-3′) (Fig. 1). Multiple sequence alignment of the conserved core regions of CmDMC1 genes was performed using DNASIS software version 3.7 (Hitachi Software Engineering, Tokyo, Japan). The target sequence for TAL effector repeat array (TALEN recognition sequence 5′ upstream: TAL-L; TALEN recognition sequence 3′ downstream: TAL-R) was designed using tools from the TAL Effector Nucleotide Targeter 2.067 for multigene disruption (Fig. 1). Sequences of 48 clones per cDNA and genomic DNA (over 7 clones per cDNA and genomic DNA sequence) were analyzed for isolation of CmDMC1 loci.
Plasmid construction
The TAL effector repeat array was synthesized using a Golden Gate TALEN and TAL Effector Kit 2.0 (addgene, Cambridge, MA, USA). The TALEN pair targeting the CmDMC1 homologs was cloned into the binary vector pBIK201G10 by substituting the β-d-glucuronidase A (gusA) gene cassette. The TALEN-pair segment and the nptII gene encoding neomycin phosphotransferase II as selectable marker were placed under the control of bi-directional promoters from the mannopine synthase 1′ and 2′ (mas1′-2′) genes. The resulting vector, designated ‘pBIK201DMC-TAL’ (Fig. 2), was introduced into A. tumefaciens strain EHA10521, kindly provided by Dr. L. S. Melchers.
Plant transformation
Two popular cultivars, ‘Shuho-no-shikara’ and ‘Yamate-shiro,’ which demonstrated high transformation rates61, were used. Preparation of Agrobacterium culture, cocultivation, selection of G418-resistant calluses, and plant regeneration were performed as described in Supplementary Table S3 and Shinoyama et al.61.
Southern blot hybridization
Total DNA was extracted from 100 mg of fresh young leaves of CmDMC1-TALEN lines and non-transgenic control plants according to Shinoyama et al.61. A 25-μg aliquot of DNA digested with XbaI was subjected to electrophoresis and blotted onto a Hybond N+ nylon membrane (GE Healthcare, Tokyo, Japan). Southern blot hybridization68 was carried out using a digoxigenin (DIG)-labeled nptII fragment (about 800 bp) as a probe (Fig. 2) and a DIG DNA Labeling and Detection Kit (Roche Diagnostics, Mannheim, Germany) according to the supplier’s instructions.
Sequencing of genomic DNA from transgenic lines
Total DNAs were prepared from leaves and roots of CmDMC1-TALEN lines as described in Shinoyama et al.61. At first, fragments of about 300 bp, including the TALEN recognition sequences (TAL-L and TAL-R; Fig. 1) for CmDMC1 genes, were amplified by PCR using a total DNA preparation as a template, ExTaq DNA polymerase (Takara-Bio, Shiga, Japan) and primer pair DMC1-gF1 (5′-ggcatggatcctggagctgtac-3′) and DMC1-gR1 (5′-ctgcaagttctcctcttccagtg-3′) (Fig. 1). Fragments of about 1.8 kbp including the 300 bp sequence were amplified by PCR using total DNA preparations as templates and primer pair DMC1-RNAi F1 (5′-aatctgtgaagctgct-3′) and DMC1-RNAi R1 (5′-tttgtcatatacac-3′) (Fig. 1). The amplicon DNA was ligated to a T-vector (pMD20, Takara-Bio) and transformed into Escherichia coli JM109. Sequences of the individual inserts were read by M13 primers M4 or RV using an ABI PRISM 3100 Genetic Analyzer with POP-4 polymer and an 80-cm capillary array (Thermo Fisher Scientific, Waltham, MA, USA). Sequences of over 13 clones per locus were analyzed for each CmDMC1-TALEN line.
Northern blot hybridization
Total RNA was extracted from 150 mg fresh weight of anthers and ovaries of CmDMC1-TALEN lines and a non-transgenic control grown at 20 °C. Each sample was homogenized in liquid nitrogen using a ceramic mortar and pestle. Total RNA was prepared from the homogenate using an RNeasy Plant Mini Kit (Qiagen, Hilden, Germany). A 20-μg aliquot of RNA was fractionated by formaldehyde gel electrophoresis through 0.8% agarose containing ethidium bromide in MOPS buffer (20 mM MOPS, 5 mM sodium acetate, 1 mM EDTA, pH 7.0). Equal loading was confirmed by examining the gel with ultraviolet light. The gel was then blotted onto a nylon membrane (Nylon Membranes, positively charged; Roche Diagnostics). Prehybridization, hybridization and detection conditions were as described above for Southern blot hybridization68 using a CmDMC1a cDNA fragment (1032 bp, Supplementary Fig. S1) and an ACTIN gene fragment (1134 bp, accession number: LC576411) from Ch. morifolium as probes.
Analysis of growth characteristics of CmDMC1-TALEN and non-transgenic chrysanthemum plants
CmDMC1-TALEN lines bearing biallelic mutations in all (six) or five CmDMC1 genes and a non-transgenic control were acclimatized in a greenhouse at 20 °C and propagated by using stem cuttings in vermiculite. After rooting, these plants (height ~ 5 cm) were exposed to 10 °C for 40 days for vernalization under a 16-h photoperiod and cool white fluorescent lamps at a PPFD (400–700 nm) of 200 μmol m–2 s–1 in a temperature gradient incubator (TG-100-A, Nippon Medical & Chemical Instruments, Osaka, Japan). They were then cultivated in the greenhouse at 20 °C under natural daylength. After flowering, factors related to growth characteristics such as stem length and number of leaves were measured. Ten plants per line were used for the observation and measurements.
Evaluation of male sterility in CmDMC1-TALEN and non-transgenic chrysanthemum plants
Male sterility was judged by pollen viability using a stain technology developed by Alexander22. CmDMC1-TALEN lines bearing biallelic mutations in all (six) or five CmDMC1 genes and a non-transgenic control were cultivated in the greenhouse at 20 °C under natural daylength. At the early meiotic division stage before tetrad formation (when diameters of flower buds were about 6 mm), the plants were transferred to the temperature gradient incubator and flowered at temperatures ranging from 10 to 30 °C with an 8-h photoperiod. Tubular florets were separated from the head flower just before flowering (Supplementary Fig. S7A) and immersed in staining solution. The tubular florets were incubated in Alexander solution22 at 50 °C overnight. Anthers were cut from the tubular florets using a scalpel and observed with a digital microscope (RH-2000, HiROX, Tokyo, Japan). Numbers of viable pollen grains (stained red with Alexander solution) and aborted pollen grains (stained green) were then counted. The percentage of pollen viability was calculated as the rate of viable grains to total pollen grains. A total of 100 head flowers each and about 10 receptive tubular florets from each head flower were used for staining of the CmDMC1-TALEN and non-transgenic control plants of ‘Shuho-no-chikara’ and ‘Yamate-shiro’. All pollen grains from each anther (a tubular floret has five anthers) were counted.
Evaluation of female sterility in CmDMC1-TALEN and non-transgenic chrysanthemums plants
CmDMC1-TALEN and non-transgenic plants were crossed with two wild relatives, Ch. wakasaense and Ch. japonense, kindly supplied by Dr. K. Taniguchi, Hiroshima University with National BioResource Project (NBRP) Chrysanthemum (https://shigen.nig.ac.jp/chrysanthemum/). The cultivar ‘Yamate-shiro’ is a quantitative short-day plant and flowers in August, whereas ‘Shuho-no-chikara’ and its wild relatives are qualitative short-day plants and flower October to November under natural daylength in Japan. To adjust the timing of flowering, CmDMC1-TALEN and non-transgenic plants and the wild relatives were exposed in June to appropriate low-temperature treatment at 10 °C for 40 days under a 16-h photoperiod and cool white fluorescent lamps (200 μmol m−2 s−1) in the temperature gradient incubator. CmDMC1-TALEN and non-transgenic plants of ’Shuho-no-chikara’ and ‘Yamate-shiro’ were then cultivated in the greenhouse at 20 °C under natural daylength. When flower buds of the CmDMC1-TALEN and non-transgenic control plants grew to a diameter of 6 mm at the early meiotic division stage, they were transferred to the temperature gradient incubator at temperatures ranging between 10 and 30 °C with an 8-h photoperiod. The pollen parents (wild relatives) were grown in the greenhouse at 20 °C, and pollen grains were collected from the dehiscent anthers of tubular florets using a small brush and placed into a Petri dish (60 mm in diameter). For the seed parents (CmDMC1-TALEN lines and non-transgenic controls), immature tubular florets were removed from head flowers with protruding stigmas; 100 flowers each and about 10 receptive stigmas each (the top of a stigma is Y-shaped as shown in Supplementary Fig. S7C) from a head flower were used for the CmDMC1-TALEN and non-transgenic control plants of ‘Shuho-no-chikara’ and Yamate-shiro’. The collected pollen grains were placed immediately on stigmas using a small brush, and each flower was covered with a paper bag. The flowers were then grown in an incubator (temperature range: 10–30 °C). After 2 months, all achenes (F1 hybrid seeds) were collected from the seed parents, sown on vermiculite and then incubated in the greenhouse at 20 °C. After 2 weeks, numbers of germinated and non-germinated seeds were counted, and assessed as viable and aborted, respectively.
Statistical comparison of CmDMC1-TALEN and non-transgenic chrysanthemums plants
For analysis of male sterility, the average percentages of viable and non-viable pollen in each head flower were arcsine-transformed prior to analysis by ANOVA. For analysis of female sterility, the percentages of F1 seed production in each head flower were arcsine-transformed prior to analysis by Tukey–Kramer’s HSD.
Data availability
All data generated or analyzed during this study are available from the corresponding author on reasonable request.
References
Shibata, M. & Kawata, J. Chromosomal variation of recent chrysanthemum cultivars for cut flower. In Development of New Technology for Identification and Classification of Tree Crops and Ornamentals (eds Kitaura, K. et al.) 41–45 (Fruit Tree Research Station, Ministry of Agriculture, Forestry and Fisheries, Government of Japan, Tokyo, 1986).
Drewlow, L. W., Ascher, P. D. & Widmer, R. E. Genetic studies of self incompatibility in the garden chrysanthemum Chrysanthemum morifolium Ramat. Theor. Appl. Genet. 43, 1–5 (1973).
Chandler, S. F. & Sanchez, C. Genetic modification; the development of transgenic ornamental plant varieties. Plant Biotechnol. J. 10, 891–903 (2012).
Kikuchi, A., Watanabe, K., Tanaka, Y. & Kamada, H. Recent progress on environmental biosafety assessment of genetically modified trees and floricultural plants in Japan. Plant Biotechnol. 25, 9–15 (2008).
Strauss, S. H. Why are regulatory requirements a major impediment to genetic engineering of horticultural crops? In Transgenic Horticultural Crops; Challenges and Opportunities (eds Mou, B. & Scorza, R.) 249–262 (CRC Press, Boca Raton, FL, 2011).
Tanaka, Y. et al. Flower color modification by engineering of the flavonoid biosynthetic pathway: practical perspectives. Biosci. Biotechnol. Biochem. 74, 1760–1769 (2010).
Nakamura, N. et al. Molecular based evidence for a lack of gene-flow between Rosa x hybrida and wild Rosa species in Japan. Plant Biotechnol. 28, 245–250 (2011).
Takatsu, Y., Nishizawa, Y., Hibi, T. & Akutsu, K. Transgenic chrysanthemum (Dendranthema grandiflorum (Ramat.) Kitamura) expressing a rice chitinase gene shows enhanced resistance to gray mold (Botrytis cinerea). Sci. Hort. 82, 113–123 (1999).
Toguri, T., Ogawa, T., Kakitani, M., Tukahara, M. & Yoshioka, M. Agrobacterium-mediated transformation of chrysanthemum (Dendranthema grandiflora) plants with a disease resistance gene (pac1). Plant Biotechnol. 20, 121–127 (2003).
Shinoyama, H., Mitsuhara, I., Ichikawa, H., Kato, K. & Mochizuki, A. Transgenic Chrysanthemums (Chrysanthemum morifolium Ramat.) carrying both insect and disease resistance. Acta Hort. 1087, 485–497 (2015).
Noda, N. et al. Generation of blue chrysanthemums by anthocyanin B-ring hydroxylation and glucosylation and its coloration mechanism. Sci. Adv. 3, e1602785 (2017).
Taniguchi, K., Nakata, M. & Kusaba, M. Chrysanthemum –Genome invasion and role of genetic resource. Biophilla 5, 55–60 (2009) (in Japanese).
Aida, R., Sasaki, K. & Ohtsubo, N. Production of chrysanthemum periclinal chimeras through shoot regeneration from leaf explants. Plant Biotechnol. 33, 45–49 (2016).
Bishop, D. K., Park, D., Xu, L. & Kleckner, N. DMC1: a meiosis-specific yeast homolog of E. coli recA required for recombination, synaptonemal complex formation, and cell cycle progression. Cell 69, 439–456 (1992).
Pittman, D. L. et al. Meiotic prophase arrest with failure of chromosome synapsis in mice deficient for Dmc1, a germline-specific RecA homolog. Mol. Cell 1, 697–705 (1998).
Doutriaux, M. P., Couteau, F., Bergounioux, C. & White, C. Isolation and characterisation of the RAD51 and DMC1 homologs from Arabidopsis thaliana. Mol. Gen. Genet. 257, 283–291 (1998).
Wang, H. et al. OsDMC1 is not required for homologous pairing in rice meiosis. Plant Physiol. 171, 230–241 (2016).
Shinoyama, H. et al. Induction of male sterility to GM chrysanthemum plants to prevent transgene flow. Acta Hort. 937, 337–346 (2012).
Kobayashi, T., Hotta, Y. & Tabata, S. Isolation and characterization of a yeast gene that is homologous with a meiosis-specific cDNA from a plant. Mol. Gen. Genet. 237, 225–232 (1993).
Etedali, F., Kohnehrouz, B. B., Valizadeh, M., Gholizadeh, A. & Malboobi, M. A. Genome wide cloning of maize meiotic recombinase Dmc1 and its functional structure through molecular phylogeny. Genet. Mol. Res. 10, 1636–1649 (2011).
Hood, E. E., Gelvin, S. B., Melchers, L. S. & Hoekema, A. New Agrobacterium helper plasmids for gene transfer to plants. Transgenic Res. 2, 208–218 (1993).
Alexander, M. P. Differential staining of aborted and nonaborted pollen. Stain Technol. 44, 117–122 (1969).
van der Ploeg, A. & Heuvelink, E. The influence of temperature on growth and development of chrysanthemum cultivars. J. Hortic. Sci. Biotechnol. 81, 174–182 (2006).
Chevalier, B. S. et al. Design, activity, and structure of a highly specific artificial endonuclease. Mol. Cell 10, 895–905 (2002).
Kim, Y. G., Cha, J. & Chandrasegaran, S. Hybrid restriction enzymes: zinc finger fusions to Fok I cleavage domain. Proc. Natl. Acad. Sci. USA 93, 1156–1160 (1996).
Christian, M. et al. Targeting DNA double-strand breaks with TAL effector nucleases. Genetics 186, 757–761 (2010).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Cermak, T. et al. Efficient design and assembly of custom TALEN and other TAL effector-based constructs for DNA targeting. Nucl. Acids Res. 39, e82 (2011).
Li, T., Liu, B., Spalding, M. H., Weeks, D. P. & Yang, B. High-efficiency TALEN-based gene editing produces disease-resistant rice. Nat. Biotechnol. 30, 390–392 (2012).
Nishizawa-Yokoi, A. et al. A Defect in DNA ligase4 enhances the frequency of TALEN-mediated targeted mutagenesis in Rice. Plant Physiol. 170, 653–666 (2016).
Wang, Y. et al. Simultaneous editing of three homoeoalleles in hexaploid bread wheat confers heritable resistance to powdery mildew. Nat. Biotechnol. 32, 947–951 (2014).
Haun, W. et al. Improved soybean oil quality by targeted mutagenesis of the fatty acid desaturase 2 gene family. Plant Biotechnol. J. 12, 934–940 (2014).
Gurushidze, M. et al. True-breeding targeted gene knock-out in barley using designer TALE-nuclease in haploid cells. PLoS ONE 9, e92046 (2014).
Char, S. N. et al. Heritable site-specific mutagenesis using TALENs in maize. Plant Biotechnol. J. 13, 1002–1010 (2015).
Sawai, S. et al. Sterol side chain reductase 2 is a key enzyme in the biosynthesis of cholesterol, the common precursor of toxic steroidal glycoalkaloids in potato. Plant Cell 26, 3763–3774 (2014).
Yasumoto, S. et al. Efficient genome engineering using Platinum TALEN in potato. Plant Biotechnol. 36, 167–173 (2019).
Jung, J. H. & Altpeter, F. TALEN mediated targeted mutagenesis of the caffeic acid O-methyltransferase in highly polyploid sugarcane improves cell wall composition for production of bioethanol. Plant Mol. Biol. 92, 131–142 (2016).
Kitamura, S. Chrysanthemum. In The Encyclopedia of Horticulture (ed Ishii, Y.) 576–585 (Seibundo-Shinkosha Publishing co. ltd., Tokyo, 1950) (in Japanese).
Kishi-Kaboshi, M., Aida, R. & Sasaki, K. Generation of gene-edited Chrysanthemum morifolium using multicopy transgenes as targets and markers. Plant Cell Physiol. 58, 216–226 (2017).
Walker, J. E., Saraste, M., Runswick, M. J. & Gay, N. J. Distantly related sequences in the alpha- and beta-subunits of ATP synthase, myosin, kinases and other ATP-requiring enzymes and a common nucleotide binding fold. EMBO J. 1, 945–951 (1982).
Sharma, D. et al. Role of the conserved lysine within the Walker A motif of human DMC. DNA Repair 12, 53–62 (2013).
Chang, H. Y. et al. Functional relationship of ATP hydrolysis, presynaptic filament stability, and homologous DNA paring activity of the human meiotic recombinase DMC1. J. Biol. Chem. 290, 19863–19873 (2015).
Dresser, M. E. et al. DMC1 functions in a Saccharomyces cerevisiae meiotic pathway that is largely independent of the RAD51 pathway. Genetics 147, 533–544 (1997).
Masson, J. Y. & West, S. C. The Rad51 and Dmc1 recombinases: a non-identical twin relationship. Trends Biochem. Sci. 26, 131–136 (2001).
Karousis, E. D., Nasif, S. & Mühlemann, O. Nonsense-mediated mRNA decay: novel mechanistic insights and biological impact. WIREs RNA 7, 661–682 (2016).
Kurosaki, T. & Maquat, L. E. Nonsense-mediated mRNA decay in humans at a glance. J. Cell Sci. 129, 461–467 (2016).
Ohtani, M. & Watcher, A. NMD-based gene regulation—a strategy for fitness enhancement in plants?. Plant Cell Physiol. 60, 1953–1960 (2019).
Shaul, O. Unique aspects of plant nonsense-mediated mRNA decay. Trends Plant Sci. 20, 767–779 (2015).
Kant, C. R., Rao, B. J. & Sainis, J. K. DNA binding and pairing activity of OsDmc1, a recombinase from rice. Plant Mol. Biol. 57, 1–11 (2005).
Siaud, N. et al. BRCA2 is involved in meiosis in Arabidopsis thaliana as suggested by its interaction with Dmc1. EMBO J. 23, 1392–1401 (2004).
Day, E., Siaud, N., Dubois, E. & Doutriaux, M. P. Interaction between Arabidopsis BRCA2 and its partners RAD51, DMC1 and DSS1. Plant Physiol. 140, 1059–1069 (2006).
Thorslund, T., Esashi, F. & West, S. C. Interaction between human BRCA2 protein and the meiosis-specific recombinase DMC1. EMBO J. 26, 2915–2922 (2007).
Martinez, J. S. et al. BRCA2 regulated DMC1-mediated recombination though the BRC repeats. Proc. Natl. Acad. Sci. USA 113, 3515–3520 (2016).
Ellis, J. G., Ryder, M. H. & Tate, M. E. Agrobacterium tumefaciens TR-DNA encodes a pathway for agropine biosynthesis. Mol. Gen. Genet. 195, 466–473 (1984).
Velten, J., Velten, L., Hain, R. & Schell, J. Isolation of a dual plant promoter fragment from the Ti plasmid of Agrobacterium tumefaciens. EMBO J. 3, 2723–2730 (1984).
Bouchez, D. & Tourneur, J. Organization of the agropine synthesis region of the T DNA of the Ri plasmid from Agrobacterium rhizogenes. Plasmid 25, 27–39 (1991).
Ledger, S. E., Deroles, C. & Given, N. K. Regeneration and Agrobacterium-mediated transformation of chrysanthemum. Plant cell Rep. 10, 195–199 (1991).
Renou, J. P., Brochard, P. & Jaloizot, R. recovery of transgenic chrysanthemum (Dendranthema grandiflorua Tzvelev) after hygromycin resistance selection. Plant Sci. 89, 185–197 (1993).
Pavingerová, D., Dostál, J., Bísková, R. & Benetka, V. Somatic embryogenesis and Agrobacterium-mediated transformation of chrysanthemum. Plant Sci. 97, 95–101 (1994).
Shinoyama, H., Nomura, Y., Tsuchiya, T. & Kazuma, T. A simple and efficient method for somatic embryogenesis and plant regeneration from leaves of chrysanthemum [Dendranthema x grandiflorum (Ramat.) Kitamura]. Plant Biotechnol. 21, 25–33 (2004).
Shinoyama, H., Kazuma, T., Komano, M., Nomura, Y. & Tsuchiya, T. An efficient transformation system in chrysanthemum [Dendranthema × grandiflorum (Ramat.) Kitamura] for stable and non-chimeric expression of foreign genes. Plant Biotechnol. 19, 335–343 (2002).
Mao, Y., Botella, J. R. & Zhu, J.-K. Heritability of targeted gene modifications induced by plant-optimized CRISPR systems. Cell. Mol. Life Sci. 74, 1075–1093 (2017).
Feng, C. et al. High-efficiency genome editing using a dmc1 promoter-controlled CRISPR/Cas9 system in maize. Plant Biotechnol. J. 16, 1848–1857 (2018).
Andersson, M. et al. Genome editing in potato via CRISPR-Cas9 ribonucleoprotein delivery. Physiol. Plant. 164, 378–384 (2018).
Liang, Z. et al. Genome editing of bread wheat using biolistic delivery of CRISPR/Cas9 in vitro transcripts or ribonucleoproteins. Nat. Protoc. 13, 413–430 (2018).
Murashige, T. & Skoog, F. A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol. Plant. 15, 473–497 (1962).
Doyle, E. L. et al. TAL Effector-Nucleotide Targeter (TALE-NT) 2.0: tools for TAL effector design and target prediction. Nucl. Acids Res. 40, W117–W122 (2012).
Southern, E. M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J. Mol. Biol. 98, 503–517 (1975).
Halpin, C., Cooke, S. E., Barakate, A., El Amrani, A. & Rayan, M. D. Self-processing 2A-polyproteins – a system for co-ordinate expression of multiple proteins in transgenic plants. Plant J. 17, 453–450 (1999).
Acknowledgements
We give special thanks to Drs. H. Kamada, H. Ezura (University of Tsukuba), F. Kikuchi (Tokyo University of Agriculture), K. Taniguchi, M. Kusaba (Hiroshima University), M. Shibata, H. Kurumizaka (The University of Tokyo), S. Ohki (Kyoto Gakuen University), M. Mii (Chiba University), K. Wakasa, K. Kondo (Tokyo University of Agriculture), M. Nakata (Toyama Botanical Garden), Y. Yonezawa (Naruto University of Education), S. Fukai (Kagawa University), K. Murai (Fukui Prefectural University), F. Altpeter (University of Florida), A. Shimakov, M. Kutsev and S. Smirnov (Altai State University) and Mr. M. Komano (Fukui prefectural government) for their valuable information, comments and suggestions and to Dr. H. Rothnie for her valuable English editing and comments. We also thank Ms. R. Murai, C. Amaya, F. Nogawa and K. Matsuura (Fukui Agricultural Experiment Station) for their technical support throughout the study. And we would like to express our deep and sincere gratitude to all members in Fukui Agricultural Experiment Station. This work was supported by the Promotion of Regional Science and Technology Research Project of the Ministry of Education, Culture, Sports, Science and Technology (MEXT) of Japan from 2013 to 2016 to Fukui Agricultural Experiment Station.
Author information
Authors and Affiliations
Contributions
H.S. and A.N.-Y. designed the research, and together with H.I., M.S. and S.T. performed experiments and analyzed the data; A.N.-Y., H.I. and S.T. conceived and supervised the project; and H.S., H.I. and M.S. conceived the experiment and wrote the main manuscript text. A.N.-Y and H.I. prepared Fig. 1, Supplementary Figs. S1 and S2; H.S. and M.S. prepared Fig. 2, Supplementary Figs. S3, S5, S6 and S8, Supplementary Tables S1–S4 and S7–S9; H.I. and H.S. prepared Figs. 3–6, Supplementary Figs. S4, S7 and S9, Table 1 and Supplementary Tables S5 and S6. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors have no competing interests as defined by Scientific Reports, or other interests that might be perceived to influence the interpretation of the article.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Shinoyama, H., Ichikawa, H., Nishizawa-Yokoi, A. et al. Simultaneous TALEN-mediated knockout of chrysanthemum DMC1 genes confers male and female sterility. Sci Rep 10, 16165 (2020). https://doi.org/10.1038/s41598-020-72356-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-72356-1
This article is cited by
-
Establishment of an efficient genetic transformation system for Tanacetum cinerariifolium
Plant Cell, Tissue and Organ Culture (PCTOC) (2024)
-
Agrobacterium tumefaciens-Mediated Plant Transformation: A Review
Molecular Biotechnology (2024)
-
Combination of long-read and short-read sequencing provides comprehensive transcriptome and new insight for Chrysanthemum morifolium ray-floret colorization
Scientific Reports (2022)
-
Random mutagenesis in vegetatively propagated crops: opportunities, challenges and genome editing prospects
Molecular Biology Reports (2022)
-
A chromosome-level genome sequence of Chrysanthemum seticuspe, a model species for hexaploid cultivated chrysanthemum
Communications Biology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.