Abstract
Loss-of-function homozygous or compound heterozygous mutations in IL36RN, which encodes interleukin-36 receptor antagonist (IL-36Ra), have been implicated in the pathogenesis of various skin disorders. Previous findings showed that IL-36γ promoted wound healing in mice; however, the pathogenic role of IL-36Ra in wound healing remains unclear. We elucidated the role of IL-36Ra, a regulator of IL-36 in tissue repair by investigating the recruitment of inflammatory cells and cytokine production in the absence of IL-36Ra. Full-thickness excisional wounds were made on the back of Il36rn−/− mice and healing was assessed by monitoring macroscopic wound sizes, numbers of infiltrated cells, and gene expression of inflammatory cytokines. Macroscopic wound healing, re-epithelialization, and granulation tissue formation were delayed by 3 days post-injury in Il36rn−/− mice. This delay was associated with increased infiltrations of neutrophils and macrophages, and increased expression of cytokines, such as IL-36γ, C-X-C motif chemokine ligand 1 (CXCL1), and transforming growth factor (TGF)-β. Importantly, administration of TAK-242, a toll-like receptor 4 (TLR4) inhibitor, caused normalization of wound healing in Il36rn−/− mice, abrogating the initial delay in tissue repair. These results showed that targeting TLR4- mediated infiltrations of immune cells and cytokine production could be beneficial in regulating wound healing in IL-36Ra-deficient skin disorders.
Similar content being viewed by others
Introduction
Interleukin-36 (IL-36) is a member of the IL-1 superfamily, with subfamily members identified as IL-36α, IL-36β, IL-36γ, interleukin-36 receptor antagonist (IL-36Ra) and IL-381,2,3. Unlike IL-36α, IL-36β, and IL-36γ that act as receptor agonists for pro-inflammatory functions4, IL-36Ra is an anti-inflammatory mediator5. The IL36RN gene encodes IL-36Ra, a protein responsible for the tight regulation of IL-36 signaling. IL-36 related cytokines are increasingly associated with various inflammatory diseases such as inflammatory bowel disease, rheumatoid and psoriatic arthritis, and inflammatory skin disorders. Among the IL-36 associated skin diseases, psoriasis is the most well-known disease.
Responsible for the pathogenesis of psoriasis-related pustular rash is caused by homozygous or compound heterozygous IL36RN gene mutations6,7,8. Loss-of-function mutations in IL36RN is defined as a recessive hereditary autoinflammatory disease. It was a type of autoinflammatory keratinization disease and called “deficiency of IL-36Ra” (DITRA)9,10. In the previous report, Il36rn−/− mice was generated and DITRA mouse model was established11. Aberrant interleukin-36 receptor (IL-36R) signaling causes transient skin inflammation including hyperkeratosis, acanthosis, and neutrophil-dominant mixed-cell infiltration11,12,13.
According to the Human Genetic Variation Database, two IL36RN founder mutations (c.28C > T (p.Arg10X) and c.115 + 6T > C (p.ArgfsX1)) are seen in about 2% of Japanese population14. The pathogenic roles of IL-36 in skin inflammatory diseases including atopic dermatitis, blistering disease, and allergic dermatitis have studied15,16,17. We found that deficiency of IL-36Ra increased the contact hypersensitivity response by eliciting excessive infiltration of neutrophils into the skin, a result of the activation of IL-36R-mediated sustained inflammatory signaling13. Thus, loss-of function mutations of IL36RN may potentially be an underlying cause of various skin inflammatory diseases including generalized pustular psoriasis. Although IL-36γ was shown to promote wound healing in mice18, little is known about the pathogenic role of IL-36Ra in the wound healing process. Therefore, we hypothesized that IL-36Ra deficiency may play an important role in wound healing process by regulating proper inflammatory cell recruitment.
Wound healing is one of the most complex biological processes19. The infiltration of inflammatory cells is very important in the inflammatory phase of the process. Wound healing is prolonged with excessive or insufficient inflammatory cell infiltration20,21,22. Excessive neutrophil infiltration and increased chemokines and growth factor mRNA expression that resulted in impaired wound healing have been observed in IL-1 receptor antagonist (IL-1ra) deficient mice20. Therefore, we hypothesized that IL-36 cytokine may play an important role in wound healing process by regulating proper inflammatory cell recruitment. To verify this hypothesis, we studied the wound healing process in Il36rn−/− mice.
Results
Deficiency of IL-36Ra caused delayed wound healing and excessive granulation tissue formation
To assess the effects of deficiency of IL-36Ra on wound healing and tissue repair, areas of open wounds were measured at 3 and 7 days after wounding Il36rn−/− and wildtype control mice. At both 3 and 7 days after injury, the open wound area was larger in Il36rn−/− mice than in wild type (Fig. 1A,B). We assessed re-epithelialization by measuring the epithelial gap, the distance between the migrating edges of keratinocytes, under the microscope. The epithelial gap was wider in Il36rn−/− mice compared with that in wild type mice at 3 days after wounding (Fig. 1A,C). Granulation tissue formation was highly promoted in Il36rn−/− mice compared with that in wild type at both 3 and 7 days after wounding (Fig. 1D). Thus, IL-36Ra deficiency led to excessive granulation tissue formation and delayed wound healing.
Wound closure and granulation tissue formation in wild type mice (WT), Il36rn−/− mice (KO) at 3 and 7 days after wounding. (A) Representative photographs of open wounds and histology of wound tissues (epithelial gap and granulation tissue) in WT and KO (magnification original × 40 and × 400). (B) The area of open wound was measured at 3 and 7 days after wounding by tracing of the wound openings onto a transparency. The epithelial gap (C), which represents the distance between the leading edge of migrating keratinocytes, and the area of granulation tissue (D) were both microscopically measured in the tissue sections. We identified areas of newly formed capillaries with a collection of fibroblasts and inflammatory cells as granulation tissues. Each histogram shows the mean ± SEM values obtained from 15 mice (51 wounds) in the WT group, and 15 mice (47 wounds) in KO group (*p < 0.05 versus WT mice).
Infiltration of inflammatory cells was intensive in wounded Il36rn−/− mice
Next, we evaluated infiltration of inflammatory cells into the wound of Il36rn−/− mice. The distribution of neutrophils in the lesions of mice was detected by histological haematoxylin and eosin staining and immunofluorescence. Numbers of neutrophils that migrated out of the blood vessels were larger in Il36rn−/− mice compared with those in wild type mice at 3 days (p < 0.05), but not 7 days (p = 0.3) after wounding (Fig. 2A,B,D).
Inflammatory cell recruitment in wounded skin of wild type mice (WT), and Il36rn−/− mice (KO) at 3 and 7 days after injury. (A–C) Representative histological skin sections from WT and KO at day 3 and day 7 after wounding. (A) Representative histological of haematoxylin and eosin staining in WT and KO (original magnification, × 100; scale bar 50 μm). (B) Immunofluorescence staining of MPO (red) in skin lesions. Nuclei were counterstained with DAPI (blue) (original magnification, × 100; scale bar 50 μm). (C) Representative histological of F4/80 staining in WT and KO (× 200; scale bar 20 μm). (D) Numbers of neutrophils per section were determined by counting H&E-stained sections under the microscope. (E) Numbers of F4/80 staining macrophages per section were also counted under the microscope. Each histogram shows the mean ± SEM values obtained from 10 wounds in each group (*p < 0.05 versus WT mice).
Macrophage numbers were larger in Il36rn−/− mice compared with those in wild type mice at 3 days (p < 0.05) but not 7 days (p = 0.48) after injury (Fig. 2C,E). There was no significant difference in numbers of CD3-positive T cells migrated out of the blood vessels in Il36rn−/− mice and wild type mice (data not shown). Collectively, IL-36Ra deficiency resulted in increased infiltration of neutrophils and macrophages into the wound tissues at the early phase of skin injury.
Cytokine mRNA expression modulated in Il36rn−/− mice
Expression levels of IL-36α, IL-36β, IL-36γ, IL-6, IL-10, IL-4, TNF-α, CXCL1, and TGF-β mRNA in the wounded skin tissue were examined by real-time RT-PCR. At 3 days after wounding, Il36rn−/− mice showed increased mRNA expression levels of IL-36γ (p < 0.05), TGF-β (p < 0.001), and CXCL1 (p < 0.05) compared with wild type mice (Fig. 3), whereas at 7 days after wounding, there was no significant difference in the mRNA expression between wild type and Il36rn−/− mice (data not shown). Thus, IL-36Ra deficiency was considered to increase the expression of IL-36γ, TGF-β, and CXCL1 in skin tissue at 3 days after wounding.
Increase in cytokine expression by IL-36Ra deficiency in the wounded skin. Expression levels of IL-6, IL-10, TNF-α, TGF-β, bFGF, CXCL1, IL-36α, IL-36β, and IL-36γ mRNA wild type mice (WT) and Il36rn−/− mice (KO) (n = 7–10; *p < 0.05; ***p < 0.001 versus wild-type mice). GAPDH mRNA was used as an internal control.
Hyaluronic acid (HA) accumulated in extracellular matrix of wounded skin
IL-36Ra deficiency resulted in delayed wound healing, enhanced infiltration of neutrophils and macrophages, and an altered mRNA expression of cytokines after 3 days of wounding. These results suggest that IL-36Ra deficiency play a crucial role in early stage of the wound-healing process. Therefore, we hypothesized that IL-36Ra modulate wound-healing through the activation of innate immunity including toll-like receptor (TLR) signal pathways. A recent study demonstrated an essential role of TLR4 in early wound repair23. In non-infectious inflammation, certain molecules, called endogenous danger signals, activate TLRs24,25,26. For example, HA binds TLR2 and TLR4 to initiates inflammatory responses and promotes recovery from acute injury27. HA is one of the major structural components of the extracellular matrix and undergoes rapid degradation at sites of tissue injury and inflammation28.
To examine the contribution of HA to delayed wound healing in Il36rn−/− mice, we assessed HA compositions in normal and wounded skin by immunohistochemistry. Although HA was slightly stained in normal skin (Fig. 4A), we found a significant increase in its levels in wounded skins at 3 and 7 days after injury. In addition, HA staining was mainly observed extracellularly (Fig. 4B,C). There was no difference in HA accumulation between wild type and Il36rn−/− mice (data not shown).
HA accumulation in wild type mice (WT) during wound healing. HA accumulation in normal skin (A) and in wounded skin at 3 days (B) and 7 days (C) after injury was assessed by immunohistochemistry using biotinylated HA binding protein (original magnification, × 100; scale bar 100 μm). No difference was observed in HA accumulation between WT and KO mice.
IL-36γ mRNA expression and protein levels in HaCaT cells
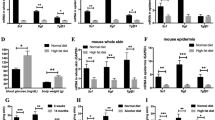
IL-36α, IL-36β, and IL36γ can be produced by various cells such as keratinocytes, epithelial cells, monocytes, and lymphocytes29. However, keratinocytes are the major source of IL-36γ30,31,32. Therefore, we stimulated HaCaT cells with LPS or low molecular weight HA (LMW HA), and examined IL-36α, β, and γ mRNA expression and protein levels by real-time RT-PCR and ELISA analysis, respectively. Both LPS and LMW HA stimulations significantly increased IL-36γ mRNA expression and protein levels in HaCaT cells, compared with media alone (Fig. 5A,B), while there was no significant change in IL-36α and IL-36β mRNA expression levels (data not shown). The increase in IL-36γ mRNA expression and protein levels in HaCaT cells stimulated with LPS or LMW HA was inhibited by TAK-242, an antagonist of TLR4.
IL-36γ mRNA expression and protein levels in HaCaT cells stimulated with LMW HA and Poly(I:C). Confluent HaCaT cells were stimulated with either media alone (Media), LPS, LWM HA (HA), LPS plus TAK-242 (LPS + TAK) or LMW HA plus TAK-242 (HA + TAK). IL-36γ mRNA expression and protein levels were quantified by real-time RT-PCR and ELISA. Each histogram shows the mean ± SEM values (n = 6; *p < 0.05; **p < 0.01).
Given that delayed wound healing has been reported in TLR3−/− mice33, we also examined IL-36γ mRNA expression and protein levels in HaCaT cells with TLR3 stimulation. TLR3 ligand, Poly(I:C) stimulation increased mRNA expression and protein levels of IL-36γ, and TLR3/dsRNA complex inhibitor suppressed IL-36γ production (Figs. 5C,D). Therefore, HaCaT cells produced IL-36γ by both LMW HA and Poly(I:C) stimulation.
TGF-β mRNA expression and proteins levels by macrophages
We observed an increase in the expression of TGF-β in the wounded skin of Il36rn−/− mice at 3 days after injury (Fig. 3). TGF-β is abundant in damaged skin and plays an important role in wound repair27,34. Further, HA increases TGF-β expression by stimulating macrophages21. We investigated the mechanism of HA stimulation of TGF-β production by HA, which may result in increased infiltration of macrophages in the wound of Il36rn−/− mice. Peritoneal macrophages from wild type mice were stimulated with LPS or LMW HA, and TGF-β mRNA expression and protein levels were examined by real-time RT-PCR and ELISA (Fig. 6).
TGF-β mRNA expression and protein levels by macrophage stimulated with LMW HA. Peritoneal macrophages were purified from wild type mice (WT) and stimulated with either media alone (Media), LPS, LMW HA (HA), LPS plus TAK-242 (LPS + TAK) and LMW HA plus TAK-242 (HA + TAK). TGF-β mRNA expression and protein levels were quantified by real-time RT-PCR and ELISA. Each histogram shows the mean ± SEM values (n = 6; **p < 0.01).
We also examined TGF-β mRNA expression and protein levels in peritoneal macrophages by Poly (I:C) stimulation and found no significant differences (data not shown). Thus, we speculated that low molecular HA increased TGF-β production by peritoneal macrophages via TLR4 signalling.
Inhibition of HA prohibited wound healing delay in Il36rn−/− mice
We examined the effect of TAK-242 on delayed wound healing observed in Il36rn−/− mice. Intraperitoneal injection of TAK-242 at the early phase of skin injury prohibited the delay in wound healing observed in the control Il36rn−/− mice without TAK-242 treatment at 3 and 7 days after injury (Fig. 7).
Wound closure in wild type mice (WT), Il36rn−/− mice (KO), and KO treated with TAK-242 (KO + TAK) at 3 and 7 days after wounding. (A) Representative photographs of open wounds in WT, KO, and TAK. (B) Areas of open wound were measured at 3 and 7 days after wounding. (C) Numbers of neutrophils at 3 days after wounding per section were determined by counting H&E-stained sections under the microscope. Each histogram shows the mean ± SEM values obtained from 3 mice (12 wounds) in the WT group, and 3 mice (12 wounds) in KO group, and 3 mice (12 wounds) in TAK group (*p < 0.05; ***p < 0.001).
Discussion
The current study is the first to demonstrate a critical role of IL-36Ra in the wound healing process. Macroscopic wound healing, re-epithelialization and granulation tissue formation were inhibited due to IL-36Ra deficiency by 3 days after wounding (Fig. 1). Delayed wound healing in Il36rn−/− mice was associated with increased infiltration of neutrophils and macrophages (Fig. 2). In addition, Il36rn−/− mice also showed increased expression of cytokines, such as IL-36γ, CXCL1, and TGF-β in the skin, especially at 3 days after injury (Fig. 3). In addition, keratinocytes and macrophages were stimulated with HA to release IL-36γ and TGF-β cytokines (Figs. 5, 6). Also, keratinocytes was stimulated with Poly(I:C) to release IL-36γ cytokines (Fig. 5). Taken together, the results of the present study indicate that IL-36Ra deficiency-induced excessive infiltrations of neutrophils and macrophages, and various cytokine production through TLR4 activation, resulted in delayed wound healing.
IL-36 expression induced during tissue damage can modulate immune cells including neutrophils35,36,37,38,52. In in vitro experiments, IL-36 binds to IL-36R in keratinocytes and macrophages and releases cytokines and chemokines, such as CXCL1, CXCL8, and G-CSF, promoting neutrophil migration39,40. Therefore, increased IL-36γ and CXCL1 expression in the injured skin may be reflective of excessive IL-36 and IL-36R signaling on neutrophils, macrophages, and keratinocytes. IL-36 is a member of the IL-1 cytokine family41 and IL-1ra deficiency increases inflammatory cell infiltration and delayed wound healing20. Since IL-36Ra binds to IL-36R, inhibiting the binding of IL36, and thus exhibiting an anti-inflammatory effect29, it is plausible that IL-36Ra deficiency may cause excessive inflammatory cell infiltration, excessive IL-36 and IL-36R signaling, increased cytokine and chemokine release, resulted in delayed wound healing as we observed in our study.
In Il36rn−/− mice, delayed wound healing was observed as early as day3; thus, we hypothesized that delayed wound healing was associated with innate immunity, such as TLR4 signaling. This model was in a sterile condition and wound healing occurred without microbial stimuli such as LPS. Therefore, we considered being a candidate that endogenous ligands for TLR4 including HA, HMGB1, and fibrinogen. It has been previously reported that trauma can release small molecular weight HA fragments from the extracellular matrix, thus it may present HA as an endogenous signal of inflammation24. Therefore, we focused on HA among the candidates for endogenous ligands in this study. Indeed, we confirmed that HA accumulated in injured skin tissue. HA fragments can signal through TLR4 and/or TLR2, and releasing the proinflammatory cytokines from endothelial cells, macrophages and dendritic cells19,42,43. On the other hand, CD44, known as the HA receptor, is expressed on many cells at injured skin44,45. Therefore, HA-CD44 signaling may also influence the process of skin wound healing, and to evaluate HA function in Il36rn−/− mice, HA-mediated signaling needs to be examined. In the current study, we speculated that the main factor that caused delayed wound healing in Il36rn−/− mice would be the different cytokine and chemokine production through TLR4 signaling by IL-36Ra deficiency, and that HA is one of possible candidates for endogenous TLR4 ligand.
We observed increased IL-36γ and TGF-β expression in the wounded skin of Il36rn−/− mice. We considered HA, an endogenous ligand for TLR4, as a trigger to induce IL-36γ and TGF-β production in the wounded skin. The concentration of HA has been shown to increase very rapidly in experimental skin wounds and peaks after 3 days of injury19, as we confirmed in our study. TLR4 plays an important role in the early stages of wound healing19,46. TLR4 is expressed in various cells including keratinocytes and macrophages47. TLR4 is important for activation of macrophages via HA stimulation21,27. When HA stimulates TLR4 expression in macrophages, it is followed by release of cytokines and chemokines such as TGF-β, CXCL1, CCL20, CCL5, causing neutrophils and macrophages to migrate to the wound site, promoting wound healing27,48,49,50,51,52. During these processes, IL-36 is released through the HA-TLR4 pathway53. We confirmed that upon stimulation with HA, there was an increase in expression of IL-36γ in HaCaT cells and TGF-β in macrophages. TAK-242, a TLR4 antagonist, effectively decreased HA-induced IL-36γ expression in HaCaT cells. Furthermore, intraperitoneal administration of TAK-242 normalized delayed wound healing in Il36rn−/− mice. These previous and current results suggest that TLR4 stimulation by HA increases IL-36γ and TGF-β expression from keratinocytes and macrophages and plays an important role in wound healing.
Delayed wound healing has been reported in TLR3−/− mice33. Studies also showed that skin injury increases IL-36γ expression in epidermal keratinocytes, and that TLR3 is required for the induction of IL-36γ18,54. Therefore, it is possible that in addition to TLR4, TLR3 signaling may also be involved in delayed wound healing in Il36rn−/− mice. In accordance with this possibility, administration of TLR3/dsRNA complex inhibitor showed significant improvement of macroscopic wound closure in Il36rn−/− mice (unsubmitted data). In addition, we examined cytokine production by HaCaT cells and macrophages with TLR3 ligand, Poly (I:C). The results showed that in HaCaT cells, Poly(I:C) stimulation increased mRNA expression and protein levels of IL-36γ, and TLR3/dsRNA complex inhibitor suppressed IL-36γ production. By contrast, peritoneal macrophages did not show a significant increase in TGF-β mRNA expression or protein levels by poly(I:C) stimulation (data not shown). Taken together, TLR3 signaling may also be involved in wound healing by a mechanism different from TLR4 signaling. Various additional analysis would be need to elucidate the detailed mechanism by which TLR3 signaling may be involved in Il36rn−/− mice.
We proposed a possible mechanism for the role of IL-36Ra in the wound healing process driven by the HA generated during early days of skin injury. We used a relevant IL-36Ra deficient model to demonstrate that HA stimulated epithelial cells and macrophages via TLR4, causing the release of IL-36 related cytokines and growth factors that promote wound healing by activating other immune cells. IL-36Ra deficiency revealed that IL-36 is a specific activator of IL-36R that may overly induce inflammatory cells in the wounded skin, resulting in delayed wound healing. In addition, since IL-36 is associated with various inflammatory skin diseases such as psoriasis and AD, this result would be a clue to elucidate pathophysiology of various inflammatory skin diseases. The limitations of current study are as follows. First, it remains unknown whether IL-36α or IL-36β is involved in wound healing. Second, the association between increased neutrophil infiltration to the wound and neutrophil extracellular traps (NETs) formation remains unknown in this study. Third, the mechanism of wound healing in a mouse is different from that of humans, primarily due to process of contraction. In murine skin, the extensive subcutaneous striated muscle layer, called the panniculus carnosus, that is absent in the human skin allows the skin to move independently of the deeper muscles, resulting in rapid contraction of skin following wounding55. Understanding the contributions of IL-36Ra to the wound repair processes could provide new clues to innovate novel manners for regulation of wound healing.
Materials and methods
Animals
Il36rn−/− mice were generated as previously reported11. All mice were healthy, fertile, and did not display any evidence of infection or disease. All mice were backcrossed between 5 to 10 generations onto the C57BL/6NCr1 (Charles River Laboratories, Inc., Wilmington, Massachusetts, USA) background. For experiments, 8–14 weeks old Il36rn−/− mice and age-matched wild-type littermates or C57BL/6NCr1 controls were used. All mice were housed in a specific pathogen-free barrier facility and screened regularly for pathogens.
Wounding and macroscopic examination
Mice were anaesthetized with medetomidine hydrochloride, midazolam and butorphanol tartrate, and their backs were shaved and wiped with 70% alcohol. Four full-thickness excisional wounds were made per mouse, using a disposable sterile 6-mm biopsy punch (Maruho Co. Ltd, Osaka, Japan), as previously described34. At 3 and 7 days after wounding, mice were anaesthetized, and areas of open wounds were measured by tracing the wound openings onto a transparency. No signs suggestive of local infection were detected in the wounded skins. All wounds were macroscopically assessed for wound closure, although wounds that were extremely distorted or whose sizes were difficult to evaluate were excluded. For macroscopic analysis of wound closure, 51 wounds from 15 mice were used in wild type group, and 47 wounds from 15 mice were used in Il36rn−/− mice group.
Histological assessment
Histological assessment of wounds was performed as previously described34. Briefly, wounds were harvested and fixed in 3.5% paraformaldehyde and embedded in paraffin. 6-μm paraffin sections were stained with H&E. The epithelial gap, and area of open wound were measured under a light microscope. The number of infiltrated neutrophils and macrophages were counted in the H&E and F4/80 stained sections. All sections were examined independently by two investigators in a blinded fashion.
Immunohistochemical staining
Immunohistochemical staining of F4/80 and HA was performed as previously described34. Briefly, paraffin-embedded sections were stained with rabbit monoclonal antibody (mAb) specific for F4/80 (Cell signal Technology Japan K.K. Tokyo, Japan). HA staining using biotinylated HA binding protein (Hokudo Co. Ltd, Sapporo, Japan) was also performed according to manufacturer’s instructions.
Immunofluorescence
The neutrophils in the wounded skin tissue were detected by MPO immunofluorescent staining. Immunofluorescent staining was performed as follows: the skin tissues were incubated with primary antibodies against MPO (1:100, R&D Systems Minneapolis, USA) overnight at 4 °C, followed by incubation with fluorescein isothiocyanate-conjugated secondary antibody for 2 h. The nuclei were counterstained with 4′,6-diamidino-2-phenylindole (DAPI) for 5 min.
Measurement of mRNA expression by real-time RT-PCR
Total RNAs were extracted from wounded skin tissues using Qiagen RNeasy mini kit (QIAGEN, Valencia, CA) and reverse transcribed to cDNA using PrimeScript RT reagent Kit with gDNA Eraser (Perfect Real Time) (TAKARA BIO, Otsu, Japan). Expression levels of IL-6, IL-10, IL-36α, IL-36β, IL-36γ, CXCL1, bFGF, TGF-β, were measured by real-time RT-PCR using the Light Cycler System (F. Hoffmann-La Roche, Ltd, Basel, Switzerland), according to the manufacturer’s instructions. Primers were obtained by pre-validated PrimeTime qPCR Assays (Integrated DNA Technologies, Coralville, IA). GAPDH was used as an internal control. Relative mRNA expression levels were calculated using the 2−∆∆Ct method, as described elsewhere.
HaCaT cell culture and stimulation
We used immortalized HaCaT cells56,57. Cells were cultured in medium containing DMEM, 100 U/mL penicillin–streptomycin, and 10% FBS. For IL-36γ induction experiments, cells were grown to confluency without changing the medium and rendered quiescent by incubation in serum-free DMEM for 1 h. Medium was replaced with fresh DMEM containing LPS (10 μg/ml), LMW HA (500 μg/ml), LPS plus TAK-242 (50 μg/ml), LMW HA plus TAK-242, Poly (I:C) (10 μg/ml), Poly (I:C) plus TLR3/dsRNA complex inhibitor (10 μg/ml) for 4 h. Cells were then harvested for real-time RT-PCR analysis. Furthermore, protein levels of IL-36γ was measured in cells cultured/treated for 72 h by using ELISA kit (Thermo Fisher SCIENTIFIC, Massachusetts, USA) according to manufacturer’s protocol.
Peritoneal macrophage culture and stimulation
Mouse peritoneal macrophages were harvested by washing the peritoneal cavity with ice cold PBS. The cells were allowed to adhere overnight in DMEM medium supplemented with 10% FBS and 1% penicillin–streptomycin 1% glutamine before use. Macrophages were cultured in serum-free DMEM medium in 24-well flat-bottom plates and stimulated for 6 h with LPS (10 ng/mL), LMW HA (100 μg/ml), LPS plus TAK-242 (10 μg/ml), LMW HA plus TAK-242, Poly(I:C) (10 μg/ml), Poly (I:C) plus TLR3/dsRNA complex inhibitor (10 μg/ml). Cells were then harvested for and real-time RT-PCR analysis. Furthermore, protein expression levels of TGF-β, was measured in cells cultured/treated for 72 h ELISA kit (Thermo Fisher SCIENTIFIC), according to manufacturer’s protocol. Culture cells from unstimulated or stimulated macrophages were harvested. RNA was extracted from peritoneal macrophage with RNeasy Mini Kit (QIAGEN) and analysed for the mRNA level of TGF-β, by real-time RT-PCR.
TLR4 inhibition with TAK-242
Il36rn−/− mice were treated with an intraperitoneal injection of 5 mg/kg TAK-242 or DW daily from Days 0 (the day at wounding) to Day 3 (3 days after wounding). TAK-242 dissolved in DMSO (10 mg/mL) was diluted in DW. Mice were anaesthetized 3 and 7 days after wounding and areas of open wounds were measured by tracing the wound openings onto a transparency, and tissue samples of the wound opening collected for IHC analysis. We analysed 12 wounds from three mice in each group (wild type, Il36rn−/−, or Il36rn−/− mice treated with TAK-242).
Statistical analysis
Data analysis were performed using GRAPHPAD PRISM software (version 7; GraphPad Software, La Jolla, CA). All data were presented as means ± SEM. Mann–Whitney U test was used to determine the statistical significance of two different samples, and one-way analysis of variance (ANOVA) was used for comparison among multiple experimental groups. Values of p < 0.05 were defined as significant.
Ethics statement
The mice were ethically treated according to the Regulations for the Management of Laboratory Animals at Fujita Health University. All studies and procedures for the ethical use of these animals have been approved by the Animal Care and Use Committee at Fujita Health University (Permit No.: AP16079).
Abbreviations
- IL:
-
Interleukin
- IL-36Ra:
-
IL-36 receptor antagonist
- CXCL1:
-
C-X-C motif chemokine ligand 1
- TGF-β:
-
Transforming growth factor-β
- TLR:
-
Toll-like receptor
- DITRA:
-
Deficiency of interleukin-36 receptor antagonist
- IL-36R:
-
IL-36 receptor
- IL-1ra:
-
IL-1 receptor antagonist
- TNF-α:
-
Tumour necrosis factor-α
- mRNA:
-
Messenger ribonucleic acid
- real-time RT-PCR:
-
Real-time reverse transcription-polymerase chain reaction
- HA:
-
Hyaluronic acid
- LPS:
-
Lipopolysaccharide
- CCL:
-
CC chemokine ligand
- H&E:
-
Haematoxylin and eosin
- PBS:
-
Phosphate buffered saline
- mAb:
-
Monoclonal antibody
- cDNA:
-
Complementary deoxyribonucleic acid
- bFGF:
-
Basic fibroblast growth factor
- PCR:
-
Polymerase chain reaction
- GAPDH:
-
Glyceraldehyde-3-phosphate
- DMEM:
-
Dulbecco's modified eagle's medium
- FBS:
-
Foetal bovine serum
- PBS:
-
Phosphate buffered saline
- DW:
-
Distilled water
- SEM:
-
Standard error of the mean
References
Sims, J. E. et al. A new nomenclature for IL-1-family genes. Trends Immunol. 22, 536–537 (2001).
Dinarello, C. et al. IL-1 family nomenclature. Nat. Immunol. 11, 973 (2010).
van de Veerdonk, F. L. & Netea, M. G. New insights in the immunobiology of IL-1 family members. Front. Immunol. 4, 167 (2013).
Towne, J. E. et al. Interleukin (IL)-1F6, IL-1F8, and IL-1F9 signal through IL-1Rrp2 and IL-1RAcP to activate the pathway leading to NF-kappa B and MAPKs. J. Biol. Chem. 279, 13677–13688 (2004).
Debets, R. et al. Two novel IL-1 family members, IL-1delta and IL-1 epsilon, function as an antagonist and agonist of NF-kappa B activation through the orphan IL-1 receptor-related protein 2. J. Immunol. 167, 1440–1446 (2001).
Marrakchi, S. et al. Interleukin-36-receptor antagonist deficiency and generalized pustular psoriasis. N. Engl. J. Med. 365, 620–628 (2011).
Onoufriadis, A. et al. Mutations in IL36RN/IL1F5 are associated with the severe episodic inflammatory skin disease known as generalized pustular psoriasis. Am. J. Hum. Genet. 89, 432–437 (2011).
Sugiura, K. et al. The majority of generalized pustular psoriasis without psoriasis vulgaris is caused by deficiency of interleukin-36 receptor antagonist. J. Invest. Dermatol. 133, 2514–2521 (2013).
Akiyama, M. et al. Autoinflammatory keratinization diseases. J. Allergy Clin. Immunol. 140, 1545–1547 (2017).
Akiyama, M. et al. Autoinflammatory keratinization diseases: An emerging concept encompassing various inflammatory keratinization disorders of the skin. J. Dermatol. Sci. 90, 105–111 (2018).
Shibata, A. et al. Toll-like receptor 4 antagonist TAK-242 inhibits autoinflammatory symptoms in DITRA. J. Autoimmun. 80, 28–38 (2017).
Blumberg, H. et al. Opposing activities of two novel members of the IL-1 ligand family regulate skin inflammation. J. Exp. Med. 204, 2603–2614 (2007).
Fukushima, H. et al. TAK-242 ameliorates contact dermatitis exacerbated by IL-36 receptor antagonist deficiency. Sci. Rep. 10, 734 (2020).
Sugiura, K. The genetic background of generalized pustular psoriasis: IL36RN mutations and CARD14 gain-of-function variants. J. Dermatol. Sci. 74, 187–192 (2014).
Moniaga, C. S., Watanabe, S., Honda, T., Nielsen, S. & Hara-Chikuma, M. Aquaporin-9-expressing neutrophils are required for the establishment of contact hypersensitivity. Sci. Rep. 5, 15319 (2015).
Christensen, A. D., Skov, S. & Haase, C. The role of neutrophils and G-CSF in DNFB-induced contact hypersensitivity in mice. Immun. Inflamm. Dis. 2, 21–34 (2014).
Goebeler, M. et al. Differential and sequential expression of multiple chemokines during elicitation of allergic contact hypersensitivity. Am. J. Pathol. 158, 431–440 (2001).
Jiang, Z. et al. IL-36γ Induced by the TLR3-SLUG-VDR Axis Promotes Wound Healing via REG3A. J. Invest. Dermatol. 137, 2620–2629 (2017).
Geoffrey, C. Gurtner. Wound repair and regeneration. Nature 453, 314–321 (2018).
Ishida, Y. et al. Absence of IL-1 receptor antagonist impaired wound healing along with aberrant NF-kappa B activation and a reciprocal suppression of TGF-beta signal pathway. J. Immunol. 176, 5598–5606 (2006).
Suga, H. et al. TLR4, rather than TLR2, regulates wound healing through TGF-β and CCL5 expression. J. Dermatol. Sci. 73, 117–124 (2014).
Oshio, T. et al. Nuclear expression of IL-33 in epidermal keratinocytes promotes wound healing in mice. J. Dermatol. Sci. 85, 106–114 (2017).
Chen, L. et al. Toll-like receptor 4 has an essential role in early skin wound healing. J. Invest. Dermatol. 133, 258–267 (2013).
Taylor, K. R. et al. Hyaluronan fragments stimulate endothelial recognition of injury through TLR4. J. Biol. Chem. 279, 17079–17084 (2004).
Foell, D., Wittkowski, H., Vogl, T. & Roth, J. S100 proteins expressed in phagocytes: a novel group of damage-associated molecular pattern molecules. J. Leukoc. Biol. 81, 28–37 (2007).
Sims, G. P. et al. HMGB1 and RAGE in inflammation and cancer. Annu. Rev. Immunol. 28, 367–388 (2010).
Jiang, D. et al. Regulation of lung injury and repair by Toll-like receptors and hyaluronan. Nat. Med. 11, 1173–1179 (2005).
Muto, J., Sayama, K., Gallo, R. L. & Kimata, K. Emerging evidence for the essential role of hyaluronan in cutaneous biology. J. Dermatol. Sci. 94, 190–195 (2019).
Ding, L. et al. IL-36 cytokines in autoimmunity and inflammatory disease. Oncotarget 9, 2895–2901 (2017).
Ritchlin, C. et al. Patterns of cytokine production in psoriatic synovium. J. Rheumatol. 25, 1544–1552 (1998).
Chustz, R. T. et al. Regulation and function of the IL-1 family cytokine IL-1F9 in human bronchial epithelial cells. Am. J. Respir. Cell Mol. Biol. 45, 145–153 (2011).
Berglöf, E. et al. IL-1Rrp2 expression and IL-1F9 (IL-1H1) actions in brain cells. J. Neuroimmunol. 139, 36–43 (2003).
Lin, Q. et al. Impaired wound healing with defective expression of chemokines and recruitment of myeloid cell in TLR3-deficient mice. J. Immunol. 186, 3710–3717 (2011).
Iwata, Y. et al. CD19, a response regulator of B lymphocytes, regulates wound healing through hyaluronan-induced TLR4 signaling. Am. J. Pathol. 175, 649–660 (2009).
Clancy, D. M. et al. Neutrophil extracellular traps can serve as platforms for processing and activation of IL-1 family cytokines. FEBS J. 284, 1712–1725 (2017).
Afonina, I. S. et al. Granzyme B-dependent proteolysis acts as a switch to enhance the proinflammatory activity of IL-1α. Mol. Cell. 44, 265–278 (2011).
Lefrancais, E. et al. IL-33 is processed into mature bioactive forms by neutrophil elastase and cathepsin G. Proc. Natl. Acad. Sci. USA 109, 1673–1678 (2012).
Afonina, I. S., Müller, C., Martin, S. J. & Beyaert, R. Proteolytic processing of interleukin-1 family cytokines: variations on a common theme. Immunity 42, 991–1004 (2015).
Foster, A. M. et al. IL-36 promotes myeloid cell infiltration, activation, and inflammatory activity in skin. J. Immunol. 192, 6053–6061 (2014).
Gabay, C. & Towne, J. E. Regulation and function of interleukin-36 cytokines in homeostasis and pathological conditions. J. Leukoc. Biol. 97, 645–652 (2015).
Arend, W. P., Malyak, M., Guthridge, C. J. & Gabay, C. Interleukin-1 receptor antagonist: role in biology. Annu. Rev. Immunol. 16, 27–55 (1998).
Johnson, G. B., Brunn, G. J. & Platt, J. L. Cutting edge: an endogenous pathway to systemic inflammatory response syndrome (SIRS)-like reactions through Toll-like receptor 4. J. Immunol. 172, 20–24 (2004).
Taylor, K. R. et al. Recognition of hyaluronan released in sterile injury involves a unique receptor complex dependent on toll-like receptor 4, CD44, and MD-2. J. Biol. Chem. 282, 18265–18275 (2007).
Aruffo, A. et al. CD44 Is the principal cell surface receptor for hyaluronate. Cell 61, 1303–1313 (1990).
Jiang, D. et al. Hyaluronan in tissue injury and repair. Annu. Rev. Cell Dev. Biol. 23, 435–461 (2007).
Neuman, M. G., Nanau, R. M., Oruña-Sanchez, L. & Coto, G. Hyaluronic acid and wound healing. J. Pharm. Pharm. Sci. 18, 53–60 (2015).
Chen, L. & DiPietro, L. A. Toll-like receptor function in acute wounds. Adv. Wound Care 16, 344–355 (2017).
Benomar, Y. et al. Central resistin overexposure induces insulin resistance through Toll-like receptor 4. Diabetes 62, 102–114 (2013).
Julian, M. W. et al. Nicotine treatment improves Toll-like receptor 2 and Toll-like receptor 9 responsiveness in active pulmonary sarcoidosis. Chest 143, 461–470 (2013).
Postlethwaite, A. E., Keski-Oja, J., Moses, H. L. & Kang, A. H. Stimulation of the chemotactic migration of human fibroblasts by transforming growth factor beta. J. Exp. Med. 165, 251–256 (1987).
Miyazono, K., Hellman, U., Wernstedt, C. & Heldin, C. H. Latent high molecular weight complex of transforming growth factor beta 1. Purification from human platelets and structural characterization. J. Biol. Chem. 263, 6407–6415 (1988).
Pircher, R., Jullien, P. & Lawrence, D. A. Beta-transforming growth factor is stored in human blood platelets as a latent high molecular weight complex. Biochem. Biophys. Res. Commun. 136, 30–37 (1986).
Babaioo, A. R. et al. Evaluation of the effect of IL-36γ expression on chronic periodontitis by enhancing the MAPK and TLR4 signaling pathways: a basic research. J. Dent. Res. Dent. Clin. Dent. Prosp. 12, 159–165 (2018).
Lai, Y. et al. Commensal bacteria regulate toll-like receptor 3-dependent infammation after skin injury. Nat. Med. 15, 1377–1382 (2009).
Wong, V. W. et al. Surgical approaches to create murine models of human wound healing. J. Biomed. Biotechnol. https://doi.org/10.1155/2011/969618 (2011).
Sasaki, H., Akamatsu, H. & Horio, T. Protective role of copper, zinc superoxide dismutase against UVB-induced injury of the human keratinocyte cell line HaCaT. J. Invest. Dermatol. 114, 502–507 (2000).
Boukamp, P. et al. Normal keratinization in a spontaneously immortalized aneuploid human keratinocyte cell line. J. Cell Biol. 106, 761–771 (1988).
Acknowledgements
This research was supported by AMED under Grant Number 19ek0109295h0003 and JSPS KAKENHI under Grant Numbers 15H04886, and 18K08281 to K.S. Additional support received by grants from the Lydia O’Leary Memorial Pias Dermatological Foundation and the Maruho Takagi Dermatology Foundation to K.S.
Author information
Authors and Affiliations
Contributions
K.S., Y.I., and K.S. wrote the manuscript. K.S., Y.I., H.F., and S.W. collected data and performed experiments. K.S., Y.I., M.A., and K.S. designed the study. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Saito, K., Iwata, Y., Fukushima, H. et al. IL-36 receptor antagonist deficiency resulted in delayed wound healing due to excessive recruitment of immune cells. Sci Rep 10, 14772 (2020). https://doi.org/10.1038/s41598-020-71256-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-71256-8
This article is cited by
-
Autoinflammatory Keratinization Diseases—The Concept, Pathophysiology, and Clinical Implications
Clinical Reviews in Allergy & Immunology (2023)
-
Role of Interleukin 36 in Generalised Pustular Psoriasis and Beyond
Dermatology and Therapy (2022)
-
IL-36 cytokines in inflammatory and malignant diseases: not the new kid on the block anymore
Cellular and Molecular Life Sciences (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.