Abstract
Early diagnosis of dementia including Alzheimer’s disease (AD) is an urgent medical and welfare issue. However, to date, no simple biometrics have been available. We reported that blood DNA methylation levels of the COASY gene, which encodes coenzyme A synthase, were increased in individuals with AD and amnestic mild cognitive impairment (aMCI). The present study sought to replicate these findings with larger numbers of samples. Another objective was to clarify whether COASY methylation is associated with neurodegeneration through a comparison of AD, AD with cardiovascular disease (CVD), and vascular dementia (VaD). We measured blood COASY methylation levels in normal controls (NCs) (n = 200), and individuals with aMCI (n = 22), AD (n = 151), and VaD (n = 21). Compared with NCs, they were significantly higher in individuals with aMCI and AD. Further, they were significantly higher in AD patients without cardiovascular diseases compared to AD patients with them. These findings suggest that COASY methylation levels may be related to neurodegeneration in AD.
Similar content being viewed by others
Introduction
In recent years, the increased incidence of dementia has become a social problem for the aging of society and aging is a major risk factor for dementia. Alzheimer’s disease (AD)1 is the most common type of dementia, followed by vascular dementia (VaD)2. AD is a neurodegenerative disease characterized by progressive neuronal loss with deposition of abnormal proteins. The neurodegenerative disease goes through the early stage called mild cognitive impairment (MCI) and ultimately develops into dementia. On the other hand, in the case of VaD, cognitive impairment is caused by cerebrovascular disease (CVD). Differential diagnosis in the early stage of dementia is crucial because the therapeutic strategy depends on the type of dementia.
Neuropathological changes in AD begin with the deposition of amyloid β-protein (Aβ), followed by the deposition of phosphorylated tau protein1,3. These changes precede cognitive impairment. For the early diagnosis of AD, amyloid positron emission tomography (PET) and tau PET are useful but these modalities are costly and difficult to use in all individuals in the clinical setting. Although decreases in Aβ42 and increases in tau protein in cerebrospinal fluid are useful biomarkers3, measurements are invasive and insufficient for differential diagnosis at an early stage of AD. Therefore, viable and less invasive methods are required in the context of feasibility. Interacting with various environmental factors, DNA methylation, a known epigenetic mechanism, affects gene expression and phenotypes without any changes in gene sequences4. Since DNA methylation changes with aging5,6 blood DNA methylation levels could be a potential diagnostic biomarker for dementia7.
The COASY gene encodes coenzyme A (CoA) synthase and it is involved in the biosynthesis of CoA from pantothenic acid. CoA synthase is mainly present in the mitochondrial matrix and mutations in this gene alter enzymatic activity. It has been reported that there is an association between mutations in COASY and neurodegeneration with brain iron accumulation (NBIA)8 and that a single-nucleotide polymorphism of COASY exon 4 is a risk factor for early onset of AD in females with Down syndrome9. Moreover, via hyper-acetylation of kinases associated with mitosis, COASY knock-down causes prolonged mitosis and failure of cytokinesis10. Thus, the aforementioned findings suggest that COASY may play an important role in neurodegeneration. In a previous pilot study using methylation-sensitive high-resolution melting (MS-HRM) analysis11, we reported that DNA methylation in the COASY gene promoter region is increased in individuals with amnestic MCI (aMCI) (n = 28) and AD12,13 (n = 30) in comparison with normal controls (NCs) (n = 30), suggesting that DNA methylation in the COASY gene promoter region could be a diagnostic marker for aMCI and AD.
However, it still remains unclear what roles COASY plays in neurodegeneration, and VaD, a disease differing from AD. Moreover, changes in COASY DNA methylation in the brain remain unclear because it would be necessary to measure them through postmortem examination of the brains of individuals with AD and NCs. Therefore, the aim of the present study was to expand the scope of our previous study and investigate changes of COASY DNA methylation in individuals with AD, VaD, and NCs. Through these experiments, we further aimed to demonstrate that changes in COASY methylation levels are associated with the upstream cascade of the pathophysiology of AD and could be a useful blood biomarker for the diagnosis of AD.
Materials and methods
Subjects
Individuals in the present study were independent of those in our previous study12,13. Individuals with aMCI, AD, and VaD were enrolled from the memory clinics of the following: the Jikei University Hospital, Tokyo (n = 51); the Jikei University Kashiwa Hospital, Kashiwa City, Chiba Prefecture (n = 80)14; and Aira-no-mori Hospital, Aira-gun, Kagoshima Prefecture (n = 63). Diagnosis of aMCI was determined by the criteria defined by Peterson15 and included both aMCI-single domain- and MCI-multiple domain-type individuals. AD was diagnosed based on the US National Institute of Neurological and Communicative Disorders and Stroke and the Alzheimer's Disease and Related Disorders Association (NINCDS-ADRDA) criteria16. The AD group was further stratified for presence of ischemic changes (hereafter, this category referred to as AD with CVD subgroup)17. All individuals with AD underwent 1.5 T magnetic resonance imaging (MRI) including T1 weighted images (T1WI), T2 weighted images (T2WI), and fluid attenuated inversion recovery (FLAIR). AD with CVD was defined as having, in addition to AD, cerebrovascular lesions such as lacunar infarctions, white matter lesions (WMLs), and microbleeding on MRI by expert neuropsychiatrists. VaD was diagnosed according to the criteria of the National Institute of Neurological Disorders and Stroke and Association Internationale pour la Recherche et l’Enseignement en Neurosciences (NINDS-AIREN)2,18. The exclusion criteria for patients included (1) being 91 years and older, (2) having been diagnosed with severe mental illness, or (3) having history of head trauma or substance use disorders.
NCs were recruited from inhabitants of Arakawa Ward in the eastern Tokyo metropolitan area as a part of the Arakawa 65 + Study and Japan Prospective Studies Collaboration for Aging and Dementia (JPSC-AD; https://www.eph.med.kyushu-u.ac.jp/jpsc/en/), which was a prospective cohort study that aimed to explore healthy longevity in elderly individuals between 65 and 84 years of age19. From the individuals (n = 1,099) recruited for the research, 200 were selected at random and enrolled in the present study. NCs had normal cognitive function and did not meet the clinical criteria for any types of dementia or MCI.
The Mini-Mental State Examination (MMSE) was administered to all of the patient groups by expert clinical psychologists20.
APOE genotyping
Genomic DNA was extracted from peripheral blood cells using a standard method14. APOE genotypes (rs429358 and rs7412) were determined by allelic discrimination on an Applied Biosystems 7,300 real-time PCR System (Thermo Fisher Scientific). The amplifications were performed in duplicate in a total volume of 25 µL containing 12.5 µL of 2x TaqMan Genotyping Master Mix (Thermo Fisher Scientific), 0.625 µL of 40 × Primer and TaqMan Probe dye mix (assay ID C—3084793_20 or C—904973_10) (Thermo Fisher Scientific), 0.2 µL of the genomic DNA, and 11.675 µL of PCR-grade water. The thermal profile was 95 °C for 10 min, followed by 40 cycles of 95 °C for 15 s and 60 °C for 1 min. Data analysis used Sequence Detection Software version 1.4 (Thermo Fisher Scientific).
Un-methylated DNA and methylated DNA
The unmethylated human genomic DNA used was EpiScope Unmethylated HCT116 DKO gDNA (Takara Bio), DNA derived from double knock-out HCT116 cells (DNMT1 and DNMT3B knocked out). The methylated human genomic DNA was EpiScope Methylated HCT116 gDNA (Takara Bio), made by high-degree methylation of un-methylated DNA using CpG methylase.
2 µg of each genomic DNA sample was bisulfite-converted using an EpiTect Plus DNA Bisulfite Kit (Qiagen) and purified. Various mixtures of bisulfited unmethylated DNA and bisulfited methylated DNA were made to produce a calibration curve for methylation rates of 100%, 75%, 50%, 25%, 5%, and 0%.
Methylation-sensitive high-resolution melting (MS-HRM) analysis
500 ng–1 µg of each genomic DNA sample was bisulfite-converted using an EpiTect Plus DNA Bisulfite Kit (Qiagen). Samples were bisulfited at the same concentration as for the bisulfited control DNA used for the calibration curve, and then purified.
The primers for the human COASY promoter region were the same as those used in our previous research13. Regarding the procedure, we designed primers with Methyl Primer Express Software v1.0 (Thermo Fisher Scientific) in a region including the sequences of the probe (Target ID cg01756799) used with the Illumina Infinitum HD Methylation Assay within Homo sapiens COASY, transcript variant 1, mRNA and Genomic Sequence (GenBank accession number NM_025233, Gene ID: 80,347). We made fine adjustments to the sequences manually. Amplifications were performed in a total volume of 20 µL containing 10 µL of 2x MeltDoctor HRM Master Mix (Applied Biosystems), 0.12 µL of 50 µM forward primer, 0.12 µL of 50 µM reverse primer, 0.6 µL of the bisulfited DNA, and 9.16 µL of PCR-grade water. The thermal profile was 95 °C for 10 min, 50 cycles of 95 °C for 15 s, and 60 °C for 60 s. The primers were as follows: human COASY forward primer, 5′-GATTATGGGATAGGAGAAGTGTT-3′, human COASY reverse primer, 5′-CCTAATCCAAAATCCCTCTTAC-3′. The amplification size was 264 bp and included 14 CpG sites (Fig. 1). MS-HRM was performed with the Applied Biosystems QuantStudio 12 K Flex Real-Time PCR System (Thermo Fisher Scientific). Using the bisulfited control DNA values, the calibration curve was obtained by regression using a cubic function and DNA methylation levels of samples were quantified. The aligned melt curves and calibration curve are shown in Fig. 2A,B.
Statistical analysis
The Shapiro–Wilk test was used to assess the normality of data. Multiple-group comparisons of non-parametric data were conducted using the Kruskal–Wallis test and the Dunn–Bonferroni correction was used for subsequent testing as needed. Sex and APOE genotype were compared using the chi-squared test. Spearman's rank correlation coefficients were used to investigate correlations between individual demographic characteristics and DNA methylation levels. Multiple linear regression analysis was then conducted with COASY DNA methylation levels in the AD group as the dependent variable, and age, sex, and MMSE score, for which differences (P-values less than 0.1) were noted using Spearman’s rank correlation coefficients, as forced entry variables. The Mann–Whitney U test was used for two-group comparisons. P < 0.05 was considered statistically significant.
Statistical analysis was conducted using SPSS Statistics 21.0 for Windows (IBM) and Prism 8 for macOS (GraphPad Software).
Ethics statement
The study was approved by the Ethics Committees of the Jikei University School of Medicine and Keio University School of Medicine and written informed consent was obtained from all individuals. For participants whose capacity to consent was compromised, caregivers who were the spouse or a relative within the second degree consented on their behalf as a substitute decision-maker only when they provided assent for the participation. Our present study was performed in accordance with the principles of the Declaration of Helsinki and the Ethical Guidelines for Medical and Health Research Involving Human Subjects in Japan.
Results
Participant characteristics
Participant characteristics are presented in Table 1. The AD group consisted of 115 individuals diagnosed as AD without CVD and 36 individuals diagnosed as AD with CVD.
Parameters that were not normally distributed were age in the NC, aMCI, AD, and VaD groups, duration of disease in the aMCI, AD, and VaD groups, age at onset in the AD and VaD groups, education in the NC, AD, and VaD groups and MMSE score in the NC, and AD groups. Age, female-male ratio, education history, MMSE score, and APOE ε4 carrier frequency were significantly different among groups (Table 1). No significant differences were found in APOE ε2 carrier frequency and APOE ε3 carrier frequency among groups (Table 1). A post hoc analysis showed age was significantly higher in the AD and VaD groups compared to the NCs (P < 0.0001 and P < 0.0001, respectively). The female-male ratio was significantly different as compared with NCs only in the AD group (P = 0.018). Education history was significantly lower in the AD and VaD groups as compared with NCs (P < 0.0001 and P < 0.0001, respectively). MMSE score in the AD and VaD groups were significantly lower as compared with NCs (P < 0.0001 and P < 0.0001, respectively). Compared with NCs, the APOE ε4 carrier frequency was significantly higher in the aMCI and AD groups (P = 0.014 and P < 0.0001, respectively).
Blood COASY DNA methylation levels
COASY promoter DNA methylation levels in the AD group did not follow a normal distribution. COASY DNA methylation levels were significantly different among groups (Fig. 3A, P < 0.0001). A post hoc analysis showed methylation levels in the COASY promoter region were higher in the aMCI and AD groups as compared with NCs (Fig. 3A, aMCI, P < 0.0001, AD, P < 0.0001, VaD, P = 0.59).
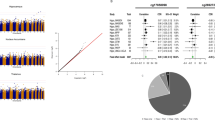
Blood COASY DNA methylation levels in dementia patients. This shows blood COASY DNA methylation levels in the dementia patients overall (A), females only (B), and males only (C). Red circles indicate females and blue circles males. Horizontal lines indicate medians and error bars interquartile range. *P < 0.05, ***P < 0.001, ****P < 0.0001. Dunn–Bonferroni correction.
We investigated the correlations of age, disease duration, age at onset, education years, and MMSE score with COASY DNA methylation levels. A significant positive correlation was observed between MMSE score and COASY DNA methylation levels in the AD group (Table 2, P < 0.0001). No significant correlations were found for the other parameters in any group (Table 2). Therefore, among the background characteristics with significant differences among the groups, age and education history had no influence on COASY DNA methylation levels. Further, COASY DNA methylation levels were compared by stratifying individuals based on sex and presence of APOE ε4. This revealed that COASY DNA methylation levels in the NC and AD groups were significantly higher in women [Table 2, NCs: female, 53.1 ± 0.9 (53.6), male, 49.7 ± 0.9 (48.9), P = 0.007, AD: female, 71.3 ± 1.2 (74.0), male, 60.4 ± 2.6 (60.6), P < 0.0001, mean ± S.E.M. (median)]. No significant differences were found in COASY DNA methylation levels between men and women for the other patient groups or due to presence of APOE ε4 in any group (Table 2). Further, multiple linear regression analysis found that among age, sex, and MMSE score (factors that could possibly affect COASY DNA methylation levels), both MMSE and sex were significantly associated with COASY DNA methylation levels in AD (Table 3, P < 0.0001, P < 0.0001, respectively).
Since we observed a significant difference in sex ratio in the disease groups and a significant difference in COASY methylation levels due to sex, we analyzed these data separately for men and women. COASY DNA methylation levels were significantly different among groups in both females and males (Fig. 3B,C, P < 0.0001 and P < 0.0001, respectively). A post hoc analysis revealed that methylation levels in the COASY promoter region were higher in the aMCI and AD groups as compared with NCs in both females and males (Fig. 3B,C, Female: aMCI, P < 0.0001, AD, P < 0.0001, and VaD, P = 0.54; Male: aMCI, P < 0.001, AD, P = 0.0005, and VaD, P = 1.00,).
Receiver operating characteristics (ROC) analysis
We carried out ROC analysis to determine whether COASY DNA methylation levels could be a useful blood biomarker in the diagnosis of MCI and AD. The results were: area under the curve (AUC): 0.84, 95% Confidence interval (CI): 0.79–0.88 (Fig. 4A). Stratifying the analysis by sex, the result for women was AUC: 0.87, 95% CI: 0.82–0.91 (Fig. 4B) and for men AUC: 0.75, 95% CI: 0.65–0.85 (Fig. 4C).
Comparison in COASY DNA methylation levels in AD without CVD, AD with CVD, and VaD
We also compared COASY promoter DNA methylation levels among the AD without CVD subgroup (n = 115), AD with CVD subgroup (n = 36), and VaD group (n = 21). COASY DNA methylation levels were significantly different among these groups (Fig. 5, P < 0.0001). COASY promoter region DNA methylation levels were significantly higher in the AD without CVD subgroup than in the AD with CVD subgroup and VaD group (Fig. 5, P < 0.001 and P < 0.0001, respectively). Also, there was a trend-toward tendency for COASY promoter region DNA methylation levels to be higher in the AD with CVD subgroup than in the VaD group (Fig. 5, P = 0.058).
COASY promoter region DNA methylation levels in AD without CVD, AD with CVD, and VaD. This shows blood COASY DNA methylation levels in AD without CVD, AD with CVD, and VaD. Horizontal lines indicate medians and error bars interquartile range. ***P < 0.001. ****P < 0.0001. Dunn–Bonferroni correction.
Discussion
The present study revealed that blood COASY DNA methylation levels were higher in AD and aMCI than those in NCs (Fig. 3A). When comparing the blood COASY DNA methylation levels among individuals with AD without CVD, AD with CVD, and VaD, the median value of the methylation levels was highest in individuals with AD without CVD (Fig. 5).
The increased levels of blood COASY DNA methylation in AD and aMCI compared to NCs confirmed the reproducibility of our previous findings13. Further, in the present study, since the analyses were performed with an increased number of samples, the effect of the sex difference between the NC and AD groups could be assessed (Table 2). In the present study, blood COASY DNA methylation was significantly higher in women than in men in both the NC and AD groups (Table 2). Epidemiological research has reported a higher prevalence of AD in women21. This may be because the number of women in the elderly population is higher and the effects of hormonal changes are greater in women compared with men; however, the actual reasons remain unclear22. In the present study, we considered it possible that the high COASY DNA methylation levels in women were associated with the higher prevalence of AD in women. Further, in analysis by sex, compared to NCs, blood COASY DNA methylation levels were increased in AD and aMCI in both men and women (Fig. 3B,C). This suggested that COASY DNA methylation levels were associated with AD pathology itself, irrespective of sex.
The results of the ROC analysis suggested that, as a blood biomarker, COASY DNA methylation levels had adequate potential for use in the diagnosis of MCI and AD (Fig. 4A–C). Application as a clinical diagnostic marker in the early phase of AD onset or at the MCI stage could be foreseen, and in this case, sensitivity and specificity would be expected to increase since COASY DNA methylation levels were positively correlated with MMSE score (Table 2).
For VaD, in which cognitive dysfunction is due to cerebrovascular disorders, we observed no significant change in blood COASY DNA methylation levels as compared to NCs. When we compared blood COASY DNA methylation levels in the AD without CVD and AD with CVD subgroups, they were significantly higher in AD without CVD than AD with CVD and, compared with VaD, there was a tendency for levels to be higher in AD with CVD, although the difference was not significant (Fig. 5). In AD with CVD, although individuals met the clinical diagnosis criteria for AD, vascular elements may have been greatly involved. This suggests that COASY DNA methylation levels are higher in AD without CVD in which vascular impairment is minimal.
The above observations suggest that rises in blood COASY DNA methylation levels reflect an AD pathology. However, since there was a positive correlation between blood COASY DNA methylation levels and MMSE scores (Table 2) and in multiple linear regression analysis with adjustment for age and sex, the effect on MMSE was still observed (Table 3), we considered that, irrespective of the severity of cognitive dysfunction, this reflected neuropathological changes occurring before the appearance of overt symptoms. In addition, in AD, Aβ-plaque accumulation is thought to reach a plateau before the onset of cognitive symptoms23. Thus, when AD progresses after symptom appearance in patients and there is a drop in MMSE score, perhaps COASY plays an adaptive role.
The COASY gene encodes CoA synthase and CoA has very important roles in the human body and the mechanism of acetylcholine esterase inhibitors used to treat AD relies on an increase in acetylcholine in the synaptic cleft, the acetylcholine being synthesized with acetyl-CoA and choline as substrates. While we did not observe a clear change in mRNA levels due to changes in COASY DNA methylation levels, our findings suggested that the long-term effect of changes in COASY DNA methylation levels would be to cause neurotoxicity. As they were evident before amyloid plaque formation, altered methylation levels could be a useful early blood biomarker.
There are some limitations in the present study. First, the functional roles of CoA synthase and COASY DNA methylation in AD onset are unclear. Further studies will be needed to determine if they have a functional role in the cause of neurodegeneration or increases in COASY DNA methylation levels are observed in AD when they are used as a surrogate marker. Second, we did not determine the locations where the DNA methylation actually occurred. However, MS-HRM is capable of measuring epigenetic changes24. In addition, it is a low-cost method widely used for this purpose. Third, we were unable to investigate an association between Aβ deposition in the brain and COASY DNA methylation levels in humans. In the future, it will be necessary to demonstrate this through PET imaging and longitudinal research.
Conclusion
In the present study, we showed that blood COASY DNA methylation levels were increased in individuals with AD and aMCI. Our results suggested that increased COASY DNA methylation levels could be a useful biomarker that reflects neurotoxicity in the early stages of AD.
References
McKhann, G. M. et al. The diagnosis of dementia due to Alzheimer’s disease: recommendations from the National Institute on Aging-Alzheimer’s Association workgroups on diagnostic guidelines for Alzheimer’s disease. Alzheimer’s Dementia J. Alzheimer’s Assoc. 7, 263–269. https://doi.org/10.1016/j.jalz.2011.03.005 (2011).
O’Brien, J. T. & Thomas, A. Vascular dementia. Lancet 386, 1698–1706. https://doi.org/10.1016/S0140-6736(15)00463-8 (2015).
Selkoe, D. J. & Hardy, J. The amyloid hypothesis of Alzheimer’s disease at 25 years. EMBO Mol. Med. 8, 595–608. https://doi.org/10.15252/emmm.201606210 (2016).
Feil, R. & Fraga, M. F. Epigenetics and the environment: emerging patterns and implications. Nat. Rev. Genet. 13, 97–109. https://doi.org/10.1038/nrg3142 (2012).
Horvath, S. DNA methylation age of human tissues and cell types. Genome Biol. 14, R115. https://doi.org/10.1186/gb-2013-14-10-r115 (2013).
Hannum, G. et al. Genome-wide methylation profiles reveal quantitative views of human aging rates. Mol. Cell 49, 359–367. https://doi.org/10.1016/j.molcel.2012.10.016 (2013).
Fransquet, P. D. et al. Blood DNA methylation as a potential biomarker of dementia: a systematic review. Alzheimer’s Dementia J. Alzheimer’s Assoc. 14, 81–103. https://doi.org/10.1016/j.jalz.2017.10.002 (2018).
Dusi, S. et al. Exome sequence reveals mutations in CoA synthase as a cause of neurodegeneration with brain iron accumulation. Am. J. Hum. Genet. 94, 11–22. https://doi.org/10.1016/j.ajhg.2013.11.008 (2014).
Lee, J. H. et al. Polymorphisms in HSD17B1: early onset and increased risk of Alzheimer’s disease in women with down syndrome. Curr. Gerontol. Geriatr. Res. 2012, 361218. https://doi.org/10.1155/2012/361218 (2012).
Lin, C. C. et al. CoA synthase regulates mitotic fidelity via CBP-mediated acetylation. Nat. Commun. 9, 1039. https://doi.org/10.1038/s41467-018-03422-6 (2018).
Wojdacz, T. K. & Dobrovic, A. Methylation-sensitive high resolution melting (MS-HRM): a new approach for sensitive and high-throughput assessment of methylation. Nucl. Acids Res. 35, e41. https://doi.org/10.1093/nar/gkm013 (2007).
Kobayashi, N. et al. Development of biomarkers based on DNA methylation in the NCAPH2/LMF2 promoter region for diagnosis of alzheimer’s disease and amnesic mild cognitive impairment. PLoS ONE 11, e0146449. https://doi.org/10.1371/journal.pone.0146449 (2016).
Kobayashi, N. et al. Usefulness of DNA methylation levels in COASY and SPINT1 gene promoter regions as biomarkers in diagnosis of alzheimer’s disease and amnestic mild cognitive impairment. PLoS ONE 11, e0168816. https://doi.org/10.1371/journal.pone.0168816 (2016).
Nagata, T., Shinagawa, S., Nukariya, K., Yamada, H. & Nakayama, K. Association between BDNF polymorphism (Val66Met) and executive function in patients with amnestic mild cognitive impairment or mild Alzheimer disease. Dement. Geriatr. Cogn. Disord. 33, 266–272. https://doi.org/10.1159/000339358 (2012).
Petersen, R. C. et al. Current concepts in mild cognitive impairment. Arch. Neurol. 58, 1985–1992 (2001).
McKhann, G. et al. Clinical diagnosis of Alzheimer’s disease: report of the NINCDS-ADRDA Work Group under the auspices of Department of Health and Human Services Task Force on Alzheimer’s Disease. Neurology 34, 939–944 (1984).
Korczyn, A. D. Mixed dementia–the most common cause of dementia. Ann. N. Y. Acad. Sci. 977, 129–134. https://doi.org/10.1111/j.1749-6632.2002.tb04807.x (2002).
Roman, G. C. et al. Vascular dementia: diagnostic criteria for research studies. Report of the NINDS-AIREN International Workshop. Neurology 43, 250–260. https://doi.org/10.1212/wnl.43.2.250 (1993).
Kida, H. et al. Approach-oriented coping strategy level may be related to volume of the whole hippocampus in the elderly. Psychiatry Clin. Neurosci. https://doi.org/10.1111/pcn.12981 (2020).
Folstein, M. F., Folstein, S. E. & McHugh, P. R. “Mini-mental state”. A practical method for grading the cognitive state of patients for the clinician. J. Psychiatry Res. 12, 189–198 (1975).
Ronquillo, J. G., Baer, M. R. & Lester, W. T. Sex-specific patterns and differences in dementia and Alzheimer’s disease using informatics approaches. J. Women Aging 28, 403–411. https://doi.org/10.1080/08952841.2015.1018038 (2016).
Medeiros, A. M. & Silva, R. H. Sex differences in Alzheimer’s disease: where do we stand?. J. Alzheimers. Dis. 67, 35–60. https://doi.org/10.3233/JAD-180213 (2019).
Jack, C. R. Jr. et al. Hypothetical model of dynamic biomarkers of the Alzheimer’s pathological cascade. Lancet Neurol. 9, 119–128. https://doi.org/10.1016/S1474-4422(09)70299-6 (2010).
Hussmann, D. & Hansen, L. L. Methylation-sensitive high resolution melting (MS-HRM). Methods Mol. Biol. 1708, 551–571. https://doi.org/10.1007/978-1-4939-7481-8_28 (2018).
Acknowledgements
The present work was supported by Japan Society for the Promotion of Science (JSPS) KAKENHI (grant number 17K10318, 17K10317), the Japan Agency for Medical Research and Development (AMED) (Grant No. 18dk0207025h0003), Mitsui Life Social Welfare Foundation and Private University Research Branding Project from the Ministry of Education, Culture, Sports, Science and Technology (MEXT). The funders were not involved in the design of the study or conducting it; in the collection, management, analysis, and interpretation of the data; or in the preparation, review and approval of the manuscript. Also, we would like to thank Mr. Alexander Cox for editorial assistance with the manuscript.
Author information
Authors and Affiliations
Contributions
N.K.: Methodology, Investigation, Formal analysis, Data curation, Visualization, Writing—Original Draft, Project administration, Conceptualization, Funding acquisition. S.S.: Resources, Investigation, Data curation, Writing—review and editing, Funding acquisition. H.N.: Resources, Investigation, Data curation, Writing—review and editing. H.K.: Resources, Investigation, Data curation, Writing—review and editing. T.N.: Resources, Investigation, Data curation, Writing—review and editing. K.T.: Resources, Investigation, Data curation, Writing—review and editing. K.S.: Investigation, Validation. N.O.: Investigation, Validation. R.S.: Resources, Investigation, Data curation. Y.N.: Resources, Writing—review and editing. S.N.: Resources, Writing—review and editing. M.M.: Supervision, Funding acquisition. M.S.: Supervision. K.K.: Supervision, Funding acquisition.
Corresponding author
Ethics declarations
Competing interests
K.K. has stock in Virus Ikagaku Kenkyusho Inc. However, Virus Ikagaku Kenkyusho Inc. was not involved in the design of the study or conducting it; in the collection, management, analysis, and interpretation of the data; or in the preparation, review and approval of the manuscript. The other authors report no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kobayashi, N., Shinagawa, S., Niimura, H. et al. Increased blood COASY DNA methylation levels a potential biomarker for early pathology of Alzheimer’s disease. Sci Rep 10, 12217 (2020). https://doi.org/10.1038/s41598-020-69248-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-69248-9
This article is cited by
-
Distinct CSF biomarker-associated DNA methylation in Alzheimer’s disease and cognitively normal subjects
Alzheimer's Research & Therapy (2023)
-
Identification of candidate DNA methylation biomarkers related to Alzheimer’s disease risk by integrating genome and blood methylome data
Translational Psychiatry (2023)
-
Distinct sex-specific DNA methylation differences in Alzheimer’s disease
Alzheimer's Research & Therapy (2022)
-
Cross-tissue analysis of blood and brain epigenome-wide association studies in Alzheimer’s disease
Nature Communications (2022)
-
Investigation of Mitochondrial Related Variants in a Cerebral Small Vessel Disease Cohort
Molecular Neurobiology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.