Abstract
Macular Telangiectasia type 2 (MacTel) is an uncommon bilateral retinal disease, in which glial cell and photoreceptor degeneration leads to central vision loss. The causative disease mechanism is largely unknown, and no treatment is currently available. A previous study found variants in genes associated with glycine–serine metabolism (PSPH, PHGDH and CPS1) to be associated with MacTel, and showed low levels of glycine and serine in the serum of MacTel patients. Recently, a causative role of deoxysphingolipids in MacTel disease has been established. However, little is known about possible other metabolic dysregulation. Here we used a global metabolomics platform in a case–control study to comprehensively profile serum from 60 MacTel patients and 58 controls. Analysis of the data, using innovative computational approaches, revealed a detailed, disease-associated metabolic profile with broad changes in multiple metabolic pathways. This included alterations in the levels of several metabolites that are directly or indirectly linked to glycine–serine metabolism, further validating our previous genetic findings. We also found changes unrelated to PSPH, PHGDH and CPS1 activity. Most pronounced, levels of several lipid groups were altered, with increased phosphatidylethanolamines being the most affected lipid group. Assessing correlations between different metabolites across our samples revealed putative functional connections. Correlations between phosphatidylethanolamines and sphingomyelin, and glycine–serine and sphingomyelin, observed in controls, were reduced in MacTel patients, suggesting metabolic re-wiring of sphingomyelin metabolism in MacTel patients. Our findings provide novel insights into metabolic changes associated with MacTel and implicate altered lipid metabolism as a contributor to this retinal neurodegenerative disease.
Similar content being viewed by others
Introduction
Macular telangiectasia type 2 (MacTel) is an uncommon, bilateral neurodegenerative retinal disease affecting between 0.004 and 0.1% of the population1,2. It is characterized by alterations of the macular capillary network and neurosensory atrophy beginning temporal to the fovea, eventually affecting the so-called “MacTel area”; an oval area approximately 3 mm across the temporal-nasal axis and 2 mm across the superior-inferior axis centred on the fovea and of similar size in all patients3. Symptoms typically start in the 5th and 6th decade of life, most commonly with reading difficulties and distortions3,4,5,6. The pathogenic mechanism of this disease is still not fully understood, but post-mortem histopathological studies show abnormalities in the retinal pigment epithelium (RPE) throughout the retina7 and a complete loss of Müller cells specifically in the MacTel area; which also contains some regions of photoreceptor loss8. A recent phase 2 clinical trial with an ocular implant secreting ciliary neurotrophic factor showed promising results in delaying disease progression, but no evidence of recovery9. No other therapeutic treatments are currently available.
Several factors suggest that MacTel has a substantial genetic component. The disease occurs bilaterally and is heritable based on studies of monozygotic twins, siblings and families3,10,11,12,13. In a Genome-Wide Association Analysis (GWAS) for MacTel we previously identified five genetic loci associated with the disease14; of which four are associated with glycine and serine abundance in serum15,16,17. We also observed lower glycine and serine levels in the serum of MacTel patients relative to controls14. One of the several metabolic pathways in which serine is need, is the biosynthesis of sphinganine, which forms the backbone of all sphingolipids18. Low serine can lead to the formation of atypical sphingolipids (deoxysphingolipids). We recently found that low serine in the serum of MacTel patients correlates with increased levels of deoxysphingolipids19, which are known to have cytotoxic properties18. However, how deoxysphingolipids (and low glycine/serine) contribute to the MacTel disease mechanism remains unclear; in particular, which factors contribute to the local retinal specificity of the disease in otherwise healthy individuals. The degree to which other metabolites may contribute to MacTel requires further investigation.
To comprehensively characterise the metabolomic profile of MacTel patients, we analysed hundreds of different serum metabolites using a global metabolomics platform. The vast data captured by this technology requires advanced data processing and statistical analysis20,21. Here we employed advanced statistical methodologies20,22,23, to characterise the impact of reduced glycine and serine in a broad metabolic context, and identify novel metabolic pathways associated with MacTel.
Materials and methods
Study participants and serum collection
From our MacTel Natural History and Observation Study (NHOS) blood serum was available from 60 random patients. Control samples were collected from 58 unrelated individuals without MacTel (confirmed by ophthalmic examination by an ophthalmologist using fundoscopy and optical coherence tomography). Control patients were selectively included to achieve similar demographics as in the MacTel patient cohort regarding age, gender, diabetic status and ethnicity. However, we did not achieve perfect symmetry between the two cohorts within the timeframe of this study (Table S1), and therefore used multiple regression modelling in our downstream statistical analysis to correct for potential confounders effects (see “Statistical analysis” section).
The number of samples in our study was based on previous experience by Metabolon (Durham, USA) regarding effect size of serum metabolomic analysis on the analytical platform used. Samples (patients and controls) were collected at different clinical sites (Moorfields Eye Hospital London, UK; Royal Victorian Eye and Ear Hospital and Ophthalmology, Melbourne, Australia; Scripps Health Facility, Scripps Clinic Torrey Pines, La Jolla, USA, and St. James’ Hospital, Leeds, UK) according to a standardised protocol. All individuals fasted overnight, and blood was taken before noon. Around 5 ml of blood was collected in a clot activating vacutainer tube (Vacutainer Plastic SST II Advance Tube with Gold Hemogard Closure, Becton Dickinson), left at room temperature for 30 min and then centrifuged for 5 min at 1,200g. The supernatant was collected, frozen and stored at − 80 °C.
Metabolite measurements
Serum metabolites were measured by Metabolon (Durham, USA). Briefly, this involved initial protein precipitation with methanol under vigorous shaking for 2 min (Glen Mills GenoGrinder 2000) followed by centrifugation. The resulting extract was divided into five fractions: two for analysis by two separate reverse phase (RP)/UPLC–MS/MS methods with positive ion mode electrospray ionization (ESI); one for analysis by RP/UPLC-MS/MS with negative ion mode ESI; and one for analysis by HILIC/UPLC-MS/MS with negative ion mode ESI. Raw data was extracted, peak-identified and QC processed using Metabolon’s hardware and software. Compounds were identified by comparison to library entries of purified standards and peaks were quantified using the area-under-the-curve technique, providing relative abundances of 946 individual metabolites.
Metabolomics data processing
Missing values for some metabolites indicated levels below the detection limit of the Metabolon platform. For this reason, missing abundances were imputed with the minimum value for each metabolite following Metabolon’s standard imputation protocol. In subsequent analyses, we discovered how imputed minimum values for each metabolite were driving a substantial amount of variance captured by principal components (PCs). Because of this, we decided to discard 194 metabolites which had more than 20% of their total values missing from the analysis; using the previously proposed “80% rule”24,25. Metabolomics missingness in controls correlated with missingness in patients, which ensured compatibility between patients and control (Fig. S2) and ensured that missingness would not confound the disease signal. The average missingness per subject was 13.3% with SD 1.5%. Boxplot outlier detection was performed on the distribution of missing values among subjects: this analysis did not detect any outlier subjects for missingness, and no particular subject was excluded for excessive amount of missingness.
A total of 738 metabolites passed quality control steps and were further analysed. We used the R software package limma26 to further quantile normalise the data between samples and ensure correction for any batch effects not captured by the initial area-under-the-curve normalization. This software package was initially developed for gene expression analysis but has been cross-purposed for metabolite analyses27,28,29. To interrogate whether the heterogeneity in the remaining metabolites was driven by the diversity between subject demographics we visually inspected the first two principal components against all available covariates. Visual exploration of principal components plots ensured that no unwanted variation was observable after the scaling and normalisation step (Fig. S3). Lastly, we scaled each metabolite to have zero mean and unit standard deviation to ensure comparability for each specific metabolomic result.
Statistical analysis
Differences in demographics between patients and controls were tested using Chi-squared tests (for dichotomous demographics) and t tests with Welch correction for unequal variances (for continuous variables). We tested each metabolite against disease status using the limma package for the statistical software, R26. This package has been widely used in gene expression study and is based on an empirical Bayes approach; whose properties for the analysis of both microarray data and RNAseq data have been described elsewhere26,30. For this study we used a false discovery rate cut-off of 5%. To this end, we corrected each p value for false discovery rate by applying the Benjamini–Hochberg p value correction procedure on the nominal uncorrected p values using the R function p.adjust31. All metabolites with a corrected p value less than the false discovery rate cut off of 0.05 were considered as significant. Each model corrected for all available covariates which included sex at birth, age, diabetes status, ethnicity and BMI. A clustered heatmap of significant metabolites is presented in (Fig. S4). To test for any residual bias due to sample ethnic imbalances we have compared the results using all samples with the results using Caucasian samples only (Fig. S5) and found virtually equivalent results.
After defining the groups, we tested for enrichment by using a popular tool for gene enrichment pathway analysis—ROAST—available in the limma package. The properties and details of this tool have been discussed elsewhere32. We corrected for false discovery rate by correcting each p value using the Benjamini–Hochberg procedure. All groups with a corrected p value of less than 0.05 were considered as significant. To account for groups in which metabolites were changed in opposite directions, we represented the abundance of grouped metabolites as a single value (the first principal component) and performed differential expression analysis using the same method applied to individual metabolites.
To test for differential co-abundance, we firstly calculated the correlation between pairs of metabolites within patients and within controls. However, metabolomics correlation can be confounded by a set of external factors. To account for this, we firstly residualised all metabolomics profiles for each covariate jointly. This was achieved by regressing each metabolite on the set of covariates and MacTel status using a linear regression model and sequentially extracting the model residuals. This approach ensured that no correlation between metabolite was either created or masked by the covariates. After residualisation, we excluded from the analysis all those metabolites pairs for which the absolute value of their correlation was less than 0.5. We then tested for differential co-abundance between pairs of metabolites similarly to previous work33 by using the Fisher R to Z transformation as defined in the Psych package34. We corrected for false discovery rate by correcting each retained p value using Benjamini–Hochberg procedure. Given the very high number of correlation pairs tested and the reduced sample size, all pairs with a corrected p value less than 0.1 were considered as significantly differentially co-abundant. Following the definition of previous work on gene expression by Jiang et al.33 we explored the hypothesis that specific groups might present an abnormal amount of differential co-abundance. Accordingly, we discarded all metabolites belonging to the xenobiotic super classification from this analysis. To test the hypothesis of groups enriched with differential co-abundance, we used a binomial test. The number of successes was the number of significant differential co-abundant pairs which contained a metabolite in that group. The number of tests was the total number of co-abundant pairs tested which contained a metabolite belonging to that group. Lastly, the probability parameter was defined as the ratio between all significantly differentially co-abundant pairs in the dataset—161—and all tested pairs 19.127 (p_parameter = 0.008). We corrected for false discovery rate by correcting each p value using Benjamini–Hochberg procedure. All metabolites with a corrected p value less than 0.05 were considered as enriched.
Ethics approval and consent to participate
All experiments were conducted according to the principles expressed in the Declaration of Helsinki. All participants gave informed consent and the study was approved by the Research Ethics Office Bromley, UK (study number 05/Q0504/101).
Results
Metabolite levels
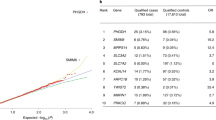
We conducted an untargeted global metabolomic analysis (Metabolon, Inc.) of serum from 60 extensively phenotyped MacTel patients and 58 healthy controls, measuring the relative levels of a total of 946 known metabolites, of which 738 survived quality control. The control cohort had similar demographics as the patient cohort (Table S1), and statistical tests (see “Materials and methods” section) revealed no significant differences between them, regarding average age (p = 0.06), gender (p = 0.14), body mass index (BMI, p = 0.34) and—since diabetes is a comorbidity of MacTel35,36—diabetic status (p = 0.22). However, there was a difference regarding ethnicity status between cases and controls (p = 0.009). Nevertheless, we used normalisation and multivariate regression strategies to correct for all available covariates (age, gender, diabetes status, ethnicity, BMI and collection site, as described in “Materials and methods” section). The fully normalised dataset (Table S2) was then analysed for differential abundance of individual metabolites in patients versus controls. This revealed 49 metabolites with significantly lower serum concentrations in MacTel patients compared to controls, and 72 with elevated serum concentrations (p < 0.05, corrected for false discovery rate, FDR) (Fig. 1).
Visual representation of all 121 significantly differentially abundant metabolites, comparing MacTel patients against controls. Each row represents a metabolite. The x-axis represents the LogFC. Negative LogFC values indicate reduced metabolite levels in MacTel patients compared to controls, and positive LogFC values indicate increased levels in MacTel patients. The model results are presented as dots indicating the estimated logFC with 95% confidence interval bars. Metabolites are divided into coloured blocks by their metabolic group. Mtb metabolism.
Glycine and serine were the first and third most depleted metabolites in MacTel patients (log(2) fold changes (logFC) of − 1.31 and − 1.03, respectively). This agrees with our recent GWAS14, which also showed reduced glycine/serine concentrations in MacTel patients, using a subset of the samples presented here. The second and fourth most changed metabolites were gamma-glutamylglycine and alpha-ketoglutarate (logFC of − 1.08 and − 0.98, respectively), both of which are linked to glycine–serine metabolism (see “Discussion” section). However, we also found many changed metabolites belonging to other metabolism groups (Figs. 1, 2). For instance, we found reduced levels of arginine (logFC = − 0.76), ornithine (logFC = − 0.59) and guanidinoacetate (logFC = − 0.47), which—together with glycine—are needed for creatine biosynthesis (see “Discussion” section). Similarly, methionine (logFC = − 0.58) and betaine (logFC = − 0.53), which are linked to cysteine–methionine metabolism (see “Discussion” section), were reduced in MacTel patients. In contrast, the majority of measured lipids were increased. Furthermore, of the 121 significantly changed metabolites, 88 were lipids (73%), whilst the total dataset of 738 metabolites contained 45% lipids, suggesting a disproportional impact on lipid metabolism in MacTel.
Changes in abundance of 738 metabolites across the 50 metabolic groups. Each row represents a metabolic group. Significantly enriched metabolic groups are labelled with * for p < 0.05 (corrected for FDR). Each group row is composed of blocks representing metabolites contained in the group. The colour of each block represents the Log2 Fold-Change (logFC) of that metabolite comparing patients against controls. The colour blue represents depletion and magenta represents increased abundance. Mtb metabolism.
To fully assess whether the observed changes affected specific metabolic pathways, we divided all metabolites into 50 functional groups—largely reflecting pathways assigned by the Kyoto Encyclopaedia of Genes and Genomes (KEGG) database—and tested (described in “Materials and methods” section) whether any groups presented enrichment of differential abundance (Fig. 3, Table S3).
Graphical overview of the key metabolic pathways that were affected in MacTel patients. Metabolites in blue were reduced in patients (p < 0.05 in dark blue, 0.05 < p < 0.1 in light blue). Metabolites in red were increased in MacTel patients (p < 0.05 in dark red, 0.05 < p < 0.1 in light red). Grey indicates no change between patients and controls, and metabolites on a white background were not measured. Double borders around metabolites indicates multiple metabolites within a group. The gene names of enzymes mentioned in the text are in yellow ovals. Note the generally reduced metabolite levels in glycine–serine and adjacent metabolic pathways, and generally increased levels in glycerophospholipid metabolism.
The most differentially abundant metabolite group in MacTel patients was glycine–serine–threonine metabolism (p = 2.5E−5), with 7 of 11 measured metabolites depleted. The second most significantly different group was the phosphatidylethanolamines (p = 8.8E−5), with 13 out of 14 metabolites upregulated in MacTel patients. Other differentially abundant groups included long chain fatty acids (increased, p = 0.02) and diacylglycerols (increased, p = 0.012), as well as changes in alanine-asparagine (reduced, p = 0.012), methionine–cysteine (mixed, PC p = 0.018) and benzoate metabolism (reduced p = 0.002). We further detected several pronounced lipid group differences including increases in lysophosphatidylethanolamines (p = 0.0054), diacylglycerols (p = 0.012), monoacylglycerols (p = 0.463) and long chain fatty acids (p = 0.020) (Table S3). Although the majority of lipids were increased in MacTel patients, etherlipids with a choline headgroup were markedly reduced (p = 0.020). A second group of lipids showing a reverse trend of general lipid increases in MacTel patients were the sphingomyelins (p = 0.012), where 20 of the 21 measured species were reduced (7 of which were significant; Fig. 1).
Correlations between metabolites
Having established that the abundance of several metabolites was altered in MacTel patients we investigated how metabolites correlated with each other (co-abundance) across patients and controls. This technique can reveal molecular interactions that change in the context of disease37 and can inform potentially dysregulated biochemical mechanisms33,38,39. Differential correlation testing was limited to those metabolite pairs which were significantly correlated after correction for multiple comparisons in either patients or controls. Whereas most correlations between metabolites were similar across both groups (Fig. S1), three were significantly different in MacTel patients compared to the controls (Table S4). Orotate and orotidine were correlated in patients (r = 0.73) but not controls (r = 0.02; p = 0.001); and the correlation between xanthine and orotidine was negative in controls (r = − 0.43) but positive in patients (r = 0.43; p = 0.001). These metabolites all function within pyrimidine metabolism. However, closer inspection of these results revealed a potential xenobiotic confound, driven by four patients with elevated oxypurinol levels (not shown), likely due to Allopurinol drug exposure, which is known to disturb pyrimidine metabolism.
We additionally tested for enrichment in differential co-abundance at the metabolite group (pathway) level. Tests of enrichment for differential co-expression were performed using a binomial framework33. Of 45 metabolite groups, seven were enriched in differentially correlated metabolite groups in patients compared to controls (p < 0.05; Fig. 4, Table S5). Among these, methionine–cysteine metabolism and the sphingomyelin group overlapped with our metabolite abundance results. The sphingomyelin group exhibited the most homogeneous pattern of differential co-abundance, characterised by a unique enrichment in disrupted connections. Specifically, differential correlation between sphingomyelins and the most differentially abundant groups—phosphatidylethanolamines and glycine–serine–threonine pathway metabolites—were driving this result. Positive correlations between the sphingomyelins and serine–glycine pathway metabolites, observed in controls, were lost in MacTel patients; while negative correlations between sphingomyelins and the phosphatidylethanolamines, were reduced or lost in patients (Fig. 5). This striking result represents disruption of the normal metabolic links between serine, phosphatidylethanolamines and sphingomyelins in sphingolipid metabolism of MacTel patients.
Circos plot displaying 152 significantly differentially co-abundant metabolite pairs across 46 metabolic groups. Differential co-abundance between metabolic groups is represented by a line connecting the relevant groups. Thickness indicates the number of significant differential co-abundances. Correlations that were lost in patients are displayed in blue (positive correlation in controls and correlation lost in patients) and cyan (negative correlation in controls and correlation lost in patients). Newly formed connections in patients are presented in red (positive correlation in patients not observed in controls) and magenta (negative correlation in patients not observed in controls). The transparency of the lines represents correlation magnitude (more transparency = lower magnitude). Note that connections involving the sphingomyelin group are most strongly suppressed in patients. Mtb metabolism.
Hive plots comparing co-abundance between metabolites in the sphingomyelin (13), phosphatidylethanolamine (14) and glycine–serine metabolism (11) groups in controls and MacTel patients. Specific metabolites are represented by circles. Co-abundance correlation between metabolites is represented by a line. Line transparency represents the correlation magnitude. Red lines represent positive correlations while blue lines represent negative correlations. Note that the majority of connections evident in controls are lost or markedly reduced in MacTel patients. Mtb metabolism.
Discussion
Data analysis tools
Careful data collection and demographic balance between cases and control by demographic information and batch control is an important part of any metabolomics study. As a perfect study design is rarely achieved for rare diseases such MacTel, statistical approaches must be used to address any potential confounding arising from imperfections. As such, this study uses a wide variety of statistical tools and techniques to reach maximal discovery power while ensuring reproducibility as well as additional sensitivity analyses to assess the key results presented in the paper.
Isolating relevant signals in metabolomics data requires advanced statistical methods. For this study we used limma software26, widely used in gene expression analysis, to normalize for confounding factors, construct sample quality weights and perform multivariate modelling. Additionally, by using an empirical Bayesian framework that “borrows” information between metabolites, the software ensures maximal discovery power even for extremely variable metabolites26,30. To extend our finding beyond simple biomarkers, we performed enrichment analysis using ROAST32 which takes into account metabolite abundances rather than summary statistics30,32. In addition, we conducted semi-supervised principal components analysis to assess significance in groups containing metabolites with mixed correlations—which might have been missed by ROAST—as well as exploring metabolic network changes through differential co-abundance analysis. Although the computational tools we employed here have so far not been commonly used for metabolomics data analysis, in this study we demonstrate that their deployment in the field of metabolomics can be extremely useful and powerful.
Genetic variants in PSPH, PHGDH and CPS1 deeply impact metabolic profiles in MacTel patients
Our previous GWAS14 identified MacTel disease risk-associated single nucleotide polymorphisms (SNPs) within the three genes, PSPH (encoding phosphoserine phosphatase), PHGDH (encoding phosphoglycerate dehydrogenase) and CPS1 (encoding carbamoyl-phosphate synthase), which are all known to contribute to glycine–serine metabolism. PSPH and PHGDH catalyse consecutive steps in the serine biosynthesis pathway (Fig. 2), and mutations in PSPH are associated with reduced plasma serine levels40. Similarly, SNP rs477992 reduces PHGDH transcription41 and serine plasma levels16,17. This matches well with the clear reductions in glycine and serine in MacTel patient serum we are presenting here. Furthermore, PHGDH is also known to convert alpha-ketoglutarate to 2-hydroxyglutarate42,43. While the former was only marginally increased in patient serum (logFC = 0.40), the latter was strongly depleted (logFC = − 0.98), further supporting a likely PHGDH defect in MacTel patients. Serine depletion, caused by dysfunctional PSPH and PHGDH, increases conversion of glycine to serine (Fig. 2)44, explaining the reduced glycine levels, at least in part.
Another likely mechanism contributing to low glycine in MacTel patients is based on the activity of CPS1, which is connected to the urea cycle and creatine biosynthesis (Fig. 2). CPS1 converts ammonia and bicarbonate into carbamoyl phosphate, which then feeds the urea cycle. The C allele of SNP rs715 (located in the 3′ UTR of CPS1; estimated MAF = 0.24) has been found in a previous independent study to be strongly associated with increased glycine serum levels16. CPS1 is also implicated, via GWAS studies, in modulating creatine16, arginine and ornithine levels45. These effects are based on the role of CPS1 in creatine biosynthesis, which requires urea metabolites. In a key intermediary step, glycine and arginine are converted to guanidinoacetate and ornithine (Fig. 2), all four of which we found to be reduced in MacTel serum (logFC = − 1.31, logFC = − 0.76, logFC = − 0.46 and logFC = − 0.59, respectively). Reduced CPS1 activity slows the urea cycle and the linked guanidino-acetate production. This likely results in reduced glycine consumption and may explain the protective role of rs715(C) in MacTel14.
Mutations in CPS1 might also explain the depleted threonine levels (logFC = − 0.96) in our MacTel patients (Figs. 1, 2) since a previous GWAS has linked CPS1 with threonine plasma levels45. However, the biochemical connections between CPS1 and threonine are not clear. Whilst most mammals can directly convert glycine to threonine, the enzyme responsible for this reaction (threonine aldolase) has lost function in humans46, making threonine an essential amino acid. Alternatively, it is conceivable that microbiome influences may contribute to glycine–threonine conversion47.
In addition, several further metabolites linked to CPS1 via GWAS45,48 were reduced in our MacTel samples (asparagine, logFC = − 0.85; glutamine, logFC = − 0.74 and betaine, logFC = − 0.53). Asparagine and glutamine have been linked to CPS1 by a GWAS45, matching the correlations found in our study (glycine–asparagine, r = 0.65 and glycine–glutamine, r = 0.51, in controls). As glutamine is needed to create intracellular asparagine, which in turn is needed for serine uptake49, depletion of these metabolites—as observed in our study—is likely to additionally contribute to the low serine availability observed in MacTel. However, the correlation between these metabolites was not changed in MacTel patients compared to controls (glycine–asparagine, r = 0.63 and glycine–glutamine, r = 0.56). Of interest, there was a trend towards reduced correlation between glycine and betaine (r = 0.56 in controls, r = 0.16 in MacTel patients), but this change did not attain statistical significance (p = 0.22).
In addition, we observed a trend towards changed correlations between serine and pyruvate, which rose from r = − 0.16 in controls to r = 0.39 in patients. Although the change did not reach statistical significance (p = 0.13), it is interesting that these two metabolites are linked via serine dehydratase (SDS), which converts serine to pyruvate (Fig. 2). An increased correlation between serine and pyruvate possibly indicates a more pronounced usage of this pathway in MacTel patients, which would reduce serine levels. Of note, SDS can also degrade threonine to α-ketobutyrate and ammonia (Fig. 2), which might be reflected by the increased α-ketobutyrate levels in MacTel serum due to a potential increase of SDS activity in our patients.
Changes in metabolites related to cysteine/methionine metabolism imply oxidative stress and phospholipid species bias in MacTel
GWAS has also linked CPS1 to the metabolites betaine, choline and homocysteine48,50, which are all relevant for cysteine/methionine metabolism (Fig. 2). In MacTel patients, betaine and choline were both reduced (logFC = − 0.53, logFC = − 0.46, respectively), which might relate to the fact that methionine was also lower in MacTel patients (logFC = − 0.58). The strong correlations we observed between methionine and asparagine (r = 0.73 in controls, r = 0.69 in MacTel patients) and between methionine and threonine (r = 0.61 in controls, r = 0.74 in MacTel patients) support the notion of a potential involvement of CPS1. However, the mechanism by which CPS1 could influence methionine levels is not known.
Methionine is an essential amino acid and can be recycled from homocysteine via two different pathways. One depends on betaine (as mentioned above, low in MacTel patients), while the other requires 5,10-methylenetetrahydrofolate (CH2-THF in Fig. 2), which was not directly measured in our study, but is likely to be reduced given its close metabolic links to serine and glycine51. Additionally, low choline and betaine observed in MacTel patients increases the reliance of the methionine cycle on one-carbon metabolism, adding a further demand on glycine. Furthermore, histidine—also used to add one carbon to tetrahydrofolate—was also reduced in MacTel patients (logFC = − 0.55). Together, these findings suggest lower methionine cycle capacity in MacTel patients compared to controls.
In this context it is interesting that MacTel patients exhibited increased concentrations of alpha-ketobutyrate (logFC = 0.59) and 2-hydroxybutyrate (logFC = 0.63), which may relate either to increased threonine degradation or to increased glutathione production (Fig. 2). Possible reasons for the latter are increased oxidative stress or detoxification in the liver52. Increased glutathione synthesis consumes serine and glycine. Furthermore, it also limits the supply of cysteine, diverting homocysteine away from the transmethylation pathway towards glutathione synthesis, leading to a stressed transmethylation pathway.
A key product of the methionine cycle is S-adenosylmethionine (SAM), required for transmethylation reactions. The conversion of phosphatidylethanolamine to phosphatidylcholine requires three SAM molecules and is, therefore, particularly affected by methionine cycle limitations. This strongly agrees with the aforementioned increase of phosphatidylethanolamines found in MacTel patients. Although phosphatidylcholine levels were nominally elevated (logFC = 0.38, p = 0.076), the phosphatidylethanolamine increase was much more pronounced (logFC = 0.81, p = 8.8e−05), apparent in nearly all measured phosphatidylethanolamine species (Fig. 2, Table S3). Interestingly, ether lipids with an ethanolamine head group were less severely reduced, mirroring the shift in the ethanolamine/choline ratio mentioned here.
The lower abundance of methionine observed in MacTel patients may be connected to changes in both glutathione and phosphatidylcholine synthesis. As a relative lack of methionine may impair glutathione synthesis, this might expose MacTel patients to a higher oxidative stress load. Additionally, the methionine derivative SAM provides the methyl group substrates required to produce choline from ethanolamine. In fact, we found increased phosphatidylethanolamine in MacTel patients, suggesting an imbalance in the phosphatidylethanolamine/phosphatidylcholine ratio. Further, substantial reduction in choline relative to ethanolamine etherphosphoilipids in patients is also consistent with methionine depletion.
Lipid dysregulation is a novel disease risk factor for MacTel
Several lipid groups were significantly changed in MacTel patients; with increased phosphatidylethanolamines, lysophosphatidylethanolamines and diacylglycerols, and decreased sphingomyelins (Table S3). These changes cannot be linked (directly or indirectly) to the activity of PSPH, PHGDH and CPS1, based on current understanding of human metabolic pathways. It is, therefore, plausible that the observed lipid dysregulation in MacTel patients represents a novel MacTel risk factor, in addition to the previously identified risk factors related to glycine–serine metabolism. The abnormal lipid levels in MacTel patients may be caused by as yet unidentified genetic risks, as well as by environmental or dietary influences.
Sphingomyelins link dysregulated lipids and serine metabolism
Differential co-abundance analysis identified sphingomyelins as an important metabolic group for MacTel. Not only were sphingomyelins depleted in MacTel, but metabolic connections—indicated by correlations of metabolites—between this group with the glycine–serine metabolism group, and the phosphatidylethanolamine group, were lost (Fig. 5). Although serine was depleted in MacTel patients, phosphatidylethanolamines and their fatty acid constituents were enriched. This striking change in correlation is particularly interesting in context of our recent finding that deoxysphingolipids are an important component in MacTel19. How exactly deoxysphingolipids contribute to retinal pathology in MacTel is not fully understood yet, but an intriguing possibility is that part of their toxic properties might be mediated by dysregulation of sphingomyelins, which are well known to differentially regulate apoptosis and autophagy53. In this context it is also interesting that ceramides were partially enriched in MacTel patients (2 out of 6 measured). Ceramides can act as pro-apoptotic signalling molecules, and it has been shown that blocking ceramide biosynthesis prevents photoreceptor cell death in a mouse model of retinitis pigmentosa54.
MacTel metabolic dysregulation might affect type 2 diabetes risk
Previous studies have observed a high prevalence of type 2 diabetes among MacTel patients35,36. However, the mechanism explaining this association is not understood. We therefore roughly matched for diabetic status in our MacTel and control cohorts (Table S1), and used a computational approach (described in “Materials and methods” section) to correct for any remaining imbalances. Despite this, our study identified an intriguing overlap between MacTel metabolites and previously described metabolic phenotypes associated with type 2 diabetes55,56,57. For instance, MacTel patients displayed increased fatty acids, diacylglycerols and phosphatidylethanolamines, as well as reduced etherlipids, glycine and glutamine, a similar pattern to that observed in type 2 diabetes metabolomics studies, hinting at potentially shared mechanistic links between MacTel and diabetes. Additionally, the recent finding of increased deoxy-sphingolipids in MacTel patients also aligns with previous findings of deoxy-sphingolipids abundance among Type 2 Diabetes patients58,59,60, further strengthening our observation of possible metabolic overlap between the two diseases.
Using serum metabolomics to study a complex eye disease
MacTel is a genetically complex and moderately clinically heterogeneous disease involving focal degeneration of retinal glia and photoreceptors, and vascular damage of the central retina (macula). MacTel heterogeneity may arise from a complex mechanism impacting multiple biological pathways. Our previous study14 provided initial evidence that the disease was associated with global alterations of metabolism, determined in part by complex genetic contributions. To explore this hypothesis, we comprehensively compared the metabolome of MacTel patients to that of healthy controls. Metabolomics is an emerging field and has been recognised as a powerful tool in ophthalmological research58,59. In this study, we used serum samples, although MacTel is a retinal disease not characterised by any known systemic pathology. This may appear counterintuitive, as retinal metabolism is considered to be isolated from the peripheral blood circulation due to the blood-retina barrier. However, MacTel is associated with systemic hyperglycaemia35,36, and metabolomics profiles of blood samples have been used to study other retinal diseases, identifying changes as potential metabolic biomarkers or interrogating broad dysregulations60,61,62,63. Furthermore, serum samples are more readily available than retina samples in people with the disease, and metabolomics platforms for analysis are readily and commercially available. We recently also showed in mice that systemic serine and glycine changes (induced by removing serine and glycine from the diet) were clearly reflected in the retina19. Finally, we note that although our study could not profit from a validation set, our findings are strengthened by the clear alignment with our previous publications on MacTel genetics14 and some metabolic features, such as reduced glycine and serine19.
It is important to keep in mind that, although our study has identified systemic risk factors for the disease, the metabolic changes in MacTel retina are so far unknown. It is also unknown by what route metabolic changes might affect the retina. It is possible that the underlying genotypes that cause the systemic changes—for example due to altered liver metabolism – affect the retina indirectly via the circulation. On the other hand, it is also possible that the genes that contribute to the systemic changes may have independent functions in the retina where they directly contribute to the disease.
Conclusion
In summary, in this study we have revealed putative functional relationships between multiple metabolite groups and MacTel disease using global metabolomic analyses. Several of the changes we observed in this study confirm our previously identified genetic MacTel findings where we implicated PSPH, PHGDH and CPS1; however, many other changes are novel and represent novel risk factors for this disease. Our study provides not only a foundation for future genetic and experimental analyses of MacTel pathobiology, but also serves as a template for the use of computational approaches to global metabolomics data to investigate diseases with complex aetiologies.
Data availability
Raw data not included in this published article is included as supplementary information.
References
Aung, K. Z., Wickremasinghe, S. S., Makeyeva, G., Robman, L. & Guymer, R. H. The prevalence estimates of macular telangiectasia type 2: the Melbourne collaborative cohort study. Retina 30, 473–478 (2010).
Klein, R. et al. The prevalence of macular telangiectasia type 2 in the Beaver Dam eye study. Am. J. Ophthalmol. 150, 2705–2710 (2010).
Charbel, P. et al. Macular telangiectasia type 2. Prog. Retin. Eye Res. 34, 49–77 (2013).
Finger, R. P. et al. Reading performance is reduced by parafoveal scotomas in patients with macular telangiectasia type 2. Investig. Ophthalmol. Vis. Sci. 50, 1366–1370 (2009).
Clemons, T. E. et al. The national eye institute visual function questionnaire in the macular telangiectasia (MacTel) project. Investig. Ophthalmol. Vis. Sci. 49, 4340–4346 (2008).
Heeren, T. F. C., Holz, F. G. & Issa, P. C. First symptoms and their age of onset in macular telangiectasia type 2. Retina 34, 916–919 (2014).
Powner, M. B. et al. Fundus-wide subretinal and pigment epithelial abnormalities in macular telangiectasia type 2. Retina 38, S105–S113 (2018).
Powner, M. B. et al. Loss of Müller’s cells and photoreceptors in macular telangiectasia type 2. Ophthalmology 120, 2344–2352 (2013).
Chew, E. Y. et al. Effect of ciliary neurotrophic factor on retinal neurodegeneration in patients with macular telangiectasia type 2: a randomized clinical trial. Ophthalmology 126, 540–549 (2019).
Parmalee, N. L. et al. Analysis of candidate genes for macular telangiectasia type 2. Mol. Vis. 16, 2718–2726 (2010).
Parmalee, N. L. et al. Identification of a potential susceptibility locus for macular telangiectasia type 2. PLoS ONE 7, 1–10 (2012).
Ronquillo, C. C., Wegner, K., Calvo, C. M. & Bernstein, P. S. Genetic penetrance of macular telangiectasia type 2. JAMA Ophthalmol. 136, 1158 (2018).
Szental, J. A. et al. Analysis of glutathione S-transferase Pi isoform (GSTP1) single-nucleotide polymorphisms and macular telangiectasia type 2. Int. Ophthalmol. 30, 645–650 (2010).
Scerri, T. S. et al. Genome-wide analyses identify common variants associated with macular telangiectasia type 2. Nat. Genet. 49, 559–567 (2017).
Xie, W. et al. Genetic variants associated with glycine metabolism and their role in insulin sensitivity and type 2 diabetes. Diabetes 62, 2141–2150 (2013).
Shin, S. Y. et al. An atlas of genetic influences on human blood metabolites. Nat. Genet. 46, 543–550 (2014).
Suhre, K. et al. Human metabolic individuality in biomedical and pharmaceutical research. Nature 477, 54–62 (2011).
Lone, M. A., Santos, T., Alecu, I., Silva, L. C. & Hornemann, T. 1-Deoxysphingolipids. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 1864, 512–521 (2019).
Gantner, M. L. et al. Serine and lipid metabolism in macular disease and peripheral neuropathy. N. Engl. J. Med. 381, 1422–1433 (2019).
Considine, E. C., Thomas, G., Boulesteix, A. L., Khashan, A. S. & Kenny, L. C. Critical review of reporting of the data analysis step in metabolomics. Metabolomics 14, 7 (2017).
Johnson, C. H., Ivanisevic, J. & Siuzdak, G. Metabolomics: beyond biomarkers and towards mechanisms. Nat. Rev. Mol. Cell Biol. 17, 451–459 (2016).
Ren, S., Hinzman, A. A., Kang, E. L., Szczesniak, R. D. & Lu, L. J. Computational and statistical analysis of metabolomics data. Metabolomics 11, 1492–1513 (2015).
Alonso, A., Marsal, S. & Julià, A. Analytical methods in untargeted metabolomics: state of the art in 2015. Front. Bioeng. Biotechnol. 3, 1–20 (2015).
Bijlsma, S. et al. Large-scale human metabolomics studies: a strategy for data (pre-) processing and validation. Anal. Chem. 78, 567–574 (2006).
Wei, R. et al. Missing value imputation approach for mass spectrometry-based metabolomics data. Sci. Rep. 8, 1–10 (2018).
Smyth, G. K. Limma: linear models for microarray data. In Statistics for Biology and Health (eds Gentleman, R. et al.) 397–420 (Springer, Berlin, 2005). https://doi.org/10.1007/0-387-29362-0_23.
Frediani, J. K. et al. Plasma metabolomics in human pulmonary tuberculosis disease: a pilot study. PLoS ONE 9, e108854 (2014).
Fischer, R. et al. Discovery of candidate serum proteomic and metabolomic biomarkers in ankylosing spondylitis. Mol. Cell. Proteomics 11, 1–11 (2012).
Putluri, N. et al. Metabolomic profiling reveals potential markers and bioprocesses altered in bladder cancer progression. Cancer Res. 71, 7376–7386 (2011).
Ritchie, M. E. et al. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. 57, 289–300 (2007).
Wu, D. et al. ROAST: Rotation gene set tests for complex microarray experiments. Bioinformatics 26, 2176–2182 (2010).
Jiang, Z., Dong, X., Li, Z., He, F. & Zhang, Z. Differential coexpression analysis reveals extensive rewiring of arabidopsis gene coexpression in response to Pseudomonas syringae infection. Publ. Gr. Nat. https://doi.org/10.1038/srep35064 (2016).
Revelle, W. An Overview of the Psych Package. (Accessed 10 July 2020); https://personality-project.org/r/overview.pdf (2017).
Clemons, T. E. et al. Medical characteristics of patients with macular telangiectasia type 2 (MacTel Type 2) MacTel Project Report No. 3. Ophthalmic Epidemiol. 20, 109–113 (2013).
Chew, E. Y., Newsome, D. A. & Fine, S. L. Parafoveal telangiectasis and diabetic retinopathy. Arch. Ophthalmol. 104, 71–75 (1986).
Hsu, C., Juan, H. & Huang, H. Functional analysis and characterization of differential coexpression networks. Publ. Gr. Nat. https://doi.org/10.1038/srep13295 (2015).
Choi, J. K., Yu, U., Yoo, O. J. & Kim, S. Differential coexpression analysis using microarray data and its application to human cancer. Bioinformatics 21, 4348–4355 (2005).
Gov, E. & Arga, K. Y. Differential co-expression analysis reveals a novel prognostic gene module in ovarian cancer. Sci. Rep. 7, 1–10 (2017).
Veiga-da-Cunha, M. et al. Mutations responsible for 3-phosphoserine phosphatase deficiency. Eur. J. Hum. Genet. 12, 163–166 (2004).
Battle, A., Brown, C. D., Engelhardt, B. E. & Montgomery, S. B. Genetic effects on gene expression across human tissues. Nature 550, 204–213 (2017).
Zhao, G. & Winkler, M. E. A novel alpha-ketoglutarate reductase activity of the serA-encoded 3-phosphoglycerate dehydrogenase of Escherichia coli K-12 and its possible implications for human 2-hydroxyglutaric aciduria. J. Bacteriol. 178, 232–239 (1996).
Fan, J. et al. Human phosphoglycerate dehydrogenase produces the oncometabolite D-2-hydroxyglutarate. ACS Chem. Biol. 10, 510–516 (2015).
Pacold, M. E. et al. A PHGDH inhibitor reveals coordination of serine synthesis and one-carbon unit fate. Nat. Chem. Biol. 12, 452–458 (2016).
Imaizumi, A. et al. Genetic basis for plasma amino acid concentrations based on absolute quantification: a genome-wide association study in the Japanese population. Eur. J. Hum. Genet. 27, 621–630 (2019).
Malinovsky, A. V. Reason for indispensability of threonine in humans and other mammals in comparative aspect. Biochemistry 82, 1055–1060 (2017).
Metges, C. C. Contribution of microbial amino acids to amino acid homeostasis of the host. J. Nutr. 130, 1857S-1864S (2000).
Hartiala, J. A. et al. Genome-wide association study and targeted metabolomics identifies sex-specific association of CPS1 with coronary artery disease. Nat. Commun. 7, 2705–2710 (2016).
Krall, A. S., Xu, S., Graeber, T. G., Braas, D. & Christofk, H. R. Asparagine promotes cancer cell proliferation through use as an amino acid exchange factor. Nat. Commun. 7, 2705–2710 (2016).
Lange, L. A. et al. Genome-wide association study of homocysteine levels in Filipinos provides evidence for CPS1 in women and a stronger MTHFR effect in young adults. Hum. Mol. Genet. 19, 2050–2058 (2010).
Kikuchi, G., Motokawa, Y., Yoshida, T. & Hiraga, K. Glycine cleavage system: reaction mechanism, physiological significance, and hyperglycinemia. Proc. Jpn. Acad. Ser. B. Phys. Biol. Sci. 84, 246–263 (2008).
Gall, W. E. et al. Alpha-hydroxybutyrate is an early biomarker of insulin resistance and glucose intolerance in a nondiabetic population. PLoS ONE 5, e10883 (2010).
Young, M. M., Kester, M. & Wang, H. G. Sphingolipids: regulators of crosstalk between apoptosis and autophagy. J. Lipid Res. 54, 5–19 (2013).
Strettoi, E. et al. Inhibition of ceramide biosynthesis preserves photoreceptor structure and function in a mouse model of retinitis pigmentosa. Proc. Natl. Acad. Sci. U. S. A. 107, 18706–18711 (2010).
Meikle, P. J. & Summers, S. A. Sphingolipids and phospholipids in insulin resistance and related metabolic disorders. Nat. Rev. Endocrinol. 13, 79–91 (2017).
Guasch-Ferré, M. et al. Metabolomics in prediabetes and diabetes: a systematic review and meta-analysis. Diabetes Care 39, 833–846 (2016).
Meikle, P. J. et al. Plasma lipid profiling shows similar associations with prediabetes and type 2 diabetes. PLoS ONE 8, 2705–2710 (2013).
Tan, S. Z., Begley, P., Mullard, G., Hollywood, K. A. & Bishop, P. N. Introduction to metabolomics and its applications in ophthalmology. Eye 30, 773–783 (2016).
Laíns, I. et al. Metabolomics in the study of retinal health and disease. Prog. Retin. Eye Res. 69, 57–79 (2019).
Laíns, I. et al. Human plasma metabolomics in age-related macular degeneration (AMD) using nuclear magnetic resonance spectroscopy. PLoS ONE 12, 1–18 (2017).
Laíns, I. et al. Human plasma metabolomics study across all stages of age-related macular degeneration identifies potential lipid biomarkers. Ophthalmology 125, 245–254 (2018).
Burgess, L. G. et al. Metabolome-wide association study of primary open angle glaucoma. Investig. Ophthalmol. Vis. Sci. 56, 5020–5028 (2015).
Li, X., Luo, X., Lu, X., Duan, J. & Xu, G. Metabolomics study of diabetic retinopathy using gas chromatography-mass spectrometry: a comparison of stages and subtypes diagnosed by Western and Chinese medicine. Mol. Biosyst. 7, 2228–2237 (2011).
Acknowledgements
We thank the volunteers who participated in this study. We are grateful to Saskia Freytag, Anna Quaglieri, Terry Speed, Gordon Smyth and Mari Gantner for helpful discussion. Support for this study came from the Lowy Medical Research Institute, National Institute of Health Research (NIHR) Biomedical Research Centre for Ophthalmology, Moorfields Eye Hospital NHS Foundation Trust, London, United Kingdom (partial funding, CE and MF), and National Institute for Health Research (NIHR). The views expressed are those of the authors and not necessarily those of the NHS, the NIHR or the Department of Health. This work was also made possible through the Victorian State Government Operational Infrastructure Support and Australian Government National Health and Medical Research Council (NHMRC) independent research Institute Infrastructure Support Scheme (IRIISS). RB was supported by the Melbourne International Research Scholarship. BREA was supported by an NHMRC early career Fellowship (1157776). RB and TFCH were supported the Lowy Medical Research fellowship. MB was supported by an NHMRC Senior Research Fellowship (1102971) and Program Grant (1054618). RG was supported by an NHMRC Senior Research Fellowship (1103013). KNK was supported by an NIHR-Rare Disease Fellowship. The funding organizations had no role in the design or conduct of this research.
Author information
Authors and Affiliations
Contributions
M.F.(3), C.A.E., K.N.K., R.B., M.B. conceived the study, S.M.W., T.F.C.H., C.A.E., K.N.K., R.G., J.T., M.F.(8) collected and prepared samples, R.B., S.M.W., B.R.E.A., M.B., M.F.(3) analysed data, and all authors contributed to the writing and final approval of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bonelli, R., Woods, S.M., Ansell, B.R.E. et al. Systemic lipid dysregulation is a risk factor for macular neurodegenerative disease. Sci Rep 10, 12165 (2020). https://doi.org/10.1038/s41598-020-69164-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-69164-y
This article is cited by
-
Unraveling the mysteries of macular telangiectasia 2: the intersection of philanthropy, multimodal imaging and molecular genetics. The 2022 founders lecture of the pan American vitreoretinal society
International Journal of Retina and Vitreous (2023)
-
New insight of metabolomics in ocular diseases in the context of 3P medicine
EPMA Journal (2023)
-
Spatial distribution of metabolites in the retina and its relevance to studies of metabolic retinal disorders
Metabolomics (2023)
-
Insulin-regulated serine and lipid metabolism drive peripheral neuropathy
Nature (2023)
-
Ciliary neurotrophic factor-mediated neuroprotection involves enhanced glycolysis and anabolism in degenerating mouse retinas
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.