Abstract
Background Deep learning has presented considerable potential and is gaining more importance in computer assisted diagnosis. As the gold standard for pathologically diagnosing cervical intraepithelial lesions and invasive cervical cancer, colposcopy-guided biopsy faces challenges in improving accuracy and efficiency worldwide, especially in developing countries. To ease the heavy burden of cervical cancer screening, it is urgent to establish a scientific, accurate and efficient method for assisting diagnosis and biopsy. Methods The data were collected to establish three deep-learning-based models. For every case, one saline image, one acetic image, one iodine image and the corresponding clinical information, including age, the results of human papillomavirus testing and cytology, type of transformation zone, and pathologic diagnosis, were collected. The dataset was proportionally divided into three subsets including the training set, the test set and the validation set, at a ratio of 8:1:1. The validation set was used to evaluate model performance. After model establishment, an independent dataset of high-definition images was collected to further evaluate the model performance. In addition, the comparison of diagnostic accuracy between colposcopists and models weas performed. Results The sensitivity, specificity and accuracy of the classification model to differentiate negative cases from positive cases were 85.38%, 82.62% and 84.10% respectively, with an AUC of 0.93. The recall and DICE of the segmentation model to segment suspicious lesions in acetic images were 84.73% and 61.64%, with an average accuracy of 95.59%. Furthermore, 84.67% of high-grade lesions were detected by the acetic detection model. Compared to colposcopists, the diagnostic system performed better in ordinary colposcopy images but slightly unsatisfactory in high-definition images. Implications The deep learning-based diagnostic system could help assist colposcopy diagnosis and biopsy for HSILs.
Similar content being viewed by others
Introduction
Ranking as the second most common cancer in the female reproductive system, cervical cancer still manifests high morbidity and mortality in developing countries including China, imposing a strong impact on the body health and quality of life of women1. Fortunately, studies have demonstrated that high-risk human papillomavirus (HPV) infection can be the definite etiology of cervical cancer2,3,4,5, making it possible to detect 80.7–98.7% of cervical intraepithelial neoplasia (CIN) early through screening combined with HPV testing and cytology6,7,8. As the gold standard for diagnosing cervical cancer and its precancerous lesions, colposcopy-guided biopsies play a key role in the early detection of cervical cancer.
In clinical practice, many factors affect the accuracy of cervical biopsies including the experience of the colposcopist, the location, size and depth of the lesions, and the menstrual status of the woman. Even by experienced colposcopists, the sensitivity of colposcopy varies from 81.4% to 95.7% to detect CIN, with a specificity of 34.2% to 69%9,10,11,12,13. Improving the accuracy of colposcopy is a key issue in the management of CIN.
Based on artificial intelligence and deep learning, computer assisted medical diagnosis can efficiently and scientifically deal with a large quantity of clinical data and achieve comparable performance on various medical tasks. Studies have suggested that medical artificial intelligence and computer assisted diagnosis can help detect lesions and improve diagnosis accuracy by using deep learning and medical image processing technology plus some possible physiological and pathological knowledge14,15,16. Studies in optical coherence tomography17, radiology18, computerized tomography scan19, colonoscopy20 and pathologic slides15 have indicated that computer algorithms can approach and even surpass the diagnostic accuracy of clinicians after training a large number of medical images in a convolutional neural network (CNN).
Early in 2009, Acosta et al. 21 used K-NN algorithm to automatically distinguish normal and abnormal cervical tissue in aceto-white pattern, and gained a sensitivity of 71% and the specificity of 59%. Years later, Asiedu et al.22 achieved the sensitivity, specificity, and accuracy of 81.3%, 78.6%, and 80.0% to distinguish CIN+ and benign tissues apart. Liming Hu et al.23 established a cohort and followed up for 7 years, using images shot by cervicography, to train and validate deep learning algorithm and gained higher accuracy compared to pap smear. Besides, Bing Bai et al.24 used K-means algorithm to automatically segment cervical region, indicating that cervical segmentation was feasible.
In all previous studies, only cervical acetic acid images were collected for training and validation. In the present study, we collected a quantity of both acetic images and iodine images with clinical information, and utilized them to train three models to separately classify, segment cervical squamous intraepithelial lesions (SILs) and detect high-grade squamous intraepithelial lesions (HSILs) to assist colposcopy-guided biopsy. Furthermore, an independent dataset of cases with high-definition colposcopy images was collected as a whole to evaluate the accuracy of the models for the second time. The performance of the models in the two datasets was compared with that of clinical colposcopists. The aim of the study was to establish a novel colposcopy diagnostic system to efficiently and accurately recognize and detect HSIL in colposcopy images and to assist colposcopists in diagnosis and biopsy.
Results
The basic information of the modelling dataset
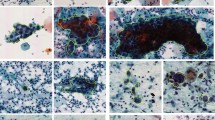
After enrolment, 22,330 cases were selected for model training and evaluation including 10,365 normal cases, 6,357 LSIL cases and 5,608 HSIL cases. Representative images of normal cases, LSIL cases and HSIL cases are presented in Fig. 1. The distributions of age, HPV infection status, cytology results and TZ type are presented in Fig. 2A.
(A) a. The cytology distribution of the modeling dataset used in the research. b. The HPV status distribution of the modeling dataset used in the research. c. The age distribution of the modeling dataset used in the research. d. The TZ type distribution of the modeling dataset used in the research. (B) The ROC curve of the validation set of the modeling dataset using the classification model.
The ResNet model can simply classify colposcopy images into two categories
In the classification model, one acetic image, one iodine image, age, HPV testing result, cytology result and TZ type were used as input indices. Pathology diagnoses were used as the output indices. Of those, 10,365 normal cases, 6,357 LSIL cases and 5,608 HSIL cases were proportionally divided into the training set, the test set and the validation set at a ratio of 8:1:1. The final result of the valid set is listed in Table 1.
The area under the curve (AUC) of the classification model reached 0.93 in the validation set (Fig. 2B), presenting a sensitivity of 85.38%, a specificity of 82.62% and an accuracy of 84.10%. In addition, the positive predictive value (PPV) and the negative predictive value (NPV) of the model were 85.02% and 83.03% respectively.
The U-Net model can precisely segment the lesions in the cervix
In total, 11,198 acetic images and 11,198 iodine images were separately input into the segmentation model. Since the U-Net model was trained at the pixel level after annotation, the segmentation model outputs a prediction area consisting of pixels that may possibly be the SIL at the end. Figure 3 presents the ground truth area (right) and the predicted area (left) in both acetic images and iodine images. The representative results showed high consistency between the two areas. Representative failed images were shown in Fig. S1. Most of the missed lesions were finally pathologically diagnosed as LSIL, and the reason of misdiagnosed lesions was not clear.
In acetic images, the recalls of normal pixels and SIL pixels were 96.14% and 84.73%, respectively, and the DICE of normal pixels and SIL pixels were 97.53% and 61.64% respectively. The average accuracy of the U-Net model in acetic images was 95.59%. The results in the iodine model were almost the same. The recalls of normal and SIL pixels were 96.03% and 87.78% respectively, and the DICE of normal and SIL pixels were 97.58% and 63.80% respectively, with a total accuracy of 95.70% in iodine images.
The MASK R-CNN model can detect HSIL lesion
In total, 22,396 images of 11,198 cases were utilized in the detection model. Nevertheless, the acetic images and the iodine images were separately trained. Finally, several rectangular prediction frames are presented with the confidence coefficient to be HSIL. The distribution of IoU and the mean IoU of prediction frames in acetic images and in iodine images were shown in Fig. 4. To control the biopsy number, only the top 3 confidence HSIL prediction frames were adopted as the final results. More specifically, circular labels of fixed diameter were utilized to mark the most suspicious area to assist biopsy (Fig. 5). Representative failed images were shown in Fig. S2.
The results of the validation set with 1,120 cases are listed in Table 2 (in acetic images) and Table 3 (in iodine images). The recalls to detect HSIL in acetic images and in iodine images were 84.67% and 84.75%, respectively. The PPV for HSIL was 19.98% in acetic images and 21.22% in iodine images. Putting LSIL and HSIL together, the detection model can recall 82.55% and 82.45% of SIL in acetic mages and iodine images respectively, with a PPV for SIL of 62.09% and 64.41%.
At the patient level, the HSIL cases were regarded as “hit” when at least one HSIL lesion was accurately predicted. The detection model could “hit” 439 HSIL cases in all 503 HSIL cases in the validation set through acetic images and could “hit” 445 HSIL cases through iodine images. The sensitivity to predict HSIL cases was 87.27% and 88.47%, respectively.
The validation results in high-definition images
After selection, 5,384 cases were enrolled in the independent dataset from a total of 9,060 cases. All the images were shot by a high-definition electronic colposcope including 3,375 normal cases, 1,246 LSIL cases and 763 HSIL cases. The distributions of age, the results of HPV testing and cytology, and the TZ types are presented in Fig. 6A.
(A) a. The cytology distribution of the validation dataset used in the research. b. The HPV status distribution of the validation dataset used in the research. c. The age distribution of the validation dataset used in the research. d. The TZ type distribution of the validation dataset used in the research. (B) The ROC curve of the validation dataset using the classification model and the ROC curve of the colposcopists.
In the classification model, the sensitivity, specificity and accuracy in differentiating positive cases and negative cases in high-definition images were 73.37%, 58.16%, and 63.83%, respectively (Table 4). The PPV and NPV were 51.07% and 78.58%, respectively, with an AUC of 0.7127 (Fig. 6B). As a comparison, the sensitivity, specificity and accuracy of five colposcopy experts in women’s hospital, school of medicine, Zhejiang University to differentiate positive cases and negative cases were calculated and are presented in Table 5. The ROC curve of the experts was shown in Fig. 6B with an AUC of 0.715. Expert 1 and expert 2 were senior colposcopists with more than 10 years of experience, expert 3 and expert 4 were intermediate colposcopists who had almost 5 years of experience in colposcopy, and expert 5 was a senior colposcopist with 1 year of experience in colposcopy. From Table 5, we concluded that the performance of the classification model in ordinary images was much better than that of all five colposcopists, while the performance in high-definition images was comparable to that of intermediate and junior colposcopists. And the AUC of the experts were almost the same as that of the classification model in high-definition images, and were lower than that of the classification model in ordinary images.
In the segmentation model, the total accuracy, normal recall and SIL recall were 94.32%, 96.84%, 85.35% in the high-definition acetic images and 94.52%, 94.92%, 85.87% in the high-definition iodine images. The detection model, detected 84.76% and 82.61% HSIL regions in high-definition acetic images and iodine images, respectively, with a PPV for HSIL of 20.62% and 20.56%. The distribution of IoU and mean IoU of high-definition images were shown in Fig. 7. Moreover, the model could “hit” 691 cases through high-definition acetic images and 685 cases through high-definition iodine images among 763 HSIL cases. The prediction sensitivity at the patient level reached 90.56% in acetic images and 89.78% in iodine images. In Table 6, experts presented a higher HSIL biopsy accuracy of 22.22–30.57% than the detection model in ordinary images and high definition images. Compared to colposcopists, the biopsy number taken in each case using the detection model was slightly higher (2.79 vs. 2.39).
Discussion
Studies of deep learning in colposcopy images are quite limited, and most of them focused only on the classification of the acetic images collected from the existing database. Tao Xu et al.25 combined the pap test result, HPV test result, age, PH value and the cervicography observation result with the acetic image to output the result and gained a sensitivity of 87.83% and a specificity of 90%. However, this kind of algorithm could only be realized on the basis of an accurate cervicography observation result, representing less clinical value. A research study in Germany enrolled 198 women who had received colposcopy examination and biopsied and extracted 211 CIN1 annotations and 164 CIN2+ annotations. The deep learning model reached an accuracy of 80%, a sensitivity of 85% and a specificity of 75%26. A research study in Japan27 collected 485 colposcopy images and divided them into three categories of atypia, carcinoma in situ and invasive cervical cancer. The final accuracy of the deep learning model was 50%, which was higher than the accuracy of 33% in random classification. The research also demonstrated that the classification of CIN2+ with CIN1 was more significant in clinical practice. Another study collected 330 patients who received colposcopy guided biopsy to train a CNN model to identify HSIL images. The sensitivity, specificity and accuracy were 82.3%,79.7% and 80.0% , respectively, with an AUC of 0.82628.
All the above studies enrolled a limited number of colposcopy cases and focused only on the classification task. Our classification model gained a sensitivity of 85.38% with an acceptable specificity of 82.62%, which performed better than the above studies. In addition, with an AUC of 0.9261, the classification model we established qualified for primary triage in colposcopy.
Research on the accuracy of clinical experts in colposcopy varies greatly. Prajakta Adsul et al. calculated the colposcopy diagnosis and the biopsy pathologic results in 1,482 subjects and found that the agreement rate of the two results was only 65.5%, and the colposcopists would always underestimate the lesions29. Margaret E. Baum et al. compared the diagnostic accuracy of different colposcopy operators and found that the nurse practitioners obtained the highest accuracy of 92%, and the accuracies of R2, R3 and R4 residents were 77%, 75% and 73%, respectively, with an average accuracy of 77%30. A meta-analysis of 86 studies demonstrated that the average sensitivity of colposcopists was 96% and the average specificity was 48%, with an AUC of 0.8 to differentiate normal and abnormal cases; the average sensitivity, specificity and AUC were 85%, 69% and 0.82 to distinguish HSIL cases from LSIL cases and normal cases, respectively31. In conclusion, the diagnostic accuracy of colposcopy relied greatly on the experience of operators, and the accuracy of most colposcopists remained below 80%. The classification model used in this article achieved an accuracy of 84.10% in ordinary images, which was higher than the accuracy of colposcopists in the literature and the accuracy of the five experts in Women’s Hospital.
For the high-definition images, the diagnostic accuracy was 63.83%, lower than that in ordinary images. Tracing back, we found that the ordinary images and the high-definition images had different distributions of standard deviation and variance in the image features including brightness, contrast, RGB colour, saturation and other factors we may not focus on now. The higher saturation and brighter colour in high definition images might lead to the highlight of acetic white and iodine nonstaining areas in normal cases, which might be mistaken with LSIL cases, accounting for the unsatisfactory performance of the classification model. Besides, the detection model could “hit” approximately 88% of HSIL patients in ordinary images and approximately 90% of HSIL patients in high-definition images. The better performance may also be attributed to the highlighting of lesions with high saturation and brightness.
Our study combined the multimodal classification model, segmentation model, and detection model to build a comprehensive system to cope with colposcopy images and to assist diagnosis and biopsy for HSIL for the first time. The ordinary images enrolled in the study were shot by three main colposcope brands, including the electronic colposcope and the photoelectric colposcope. All three types of images presented perfect receptivity to our models. Besides, in the high definition images shot by another two electronic colposcopes, the models we established could also reach the diagnostic accuracy equal to the junior experts and presented better ability to detect HSIL.
Nevertheless, more investigations need to be carried out in the future. A large number of images from colposcopes of various brands, especially high-definition images, were required to improve the existing models. A prospective, large-scale, multicentre clinical trial needs to be carried out to evaluate the clinical value.
Methods
Data resource
All the colposcopy images of the modeling dataset were collected in women’s hospital, school of medicine, Zhejiang University from August 2013 to March 2019. Those who met the following conditions were excluded: without complete clinical and pathological information (age, result of HPV testing and cytology); without biopsies; pathologically diagnosed as invasive cervical cancer or glandular intraepithelial lesions; with poor-quality colposcopy images (blur, over-reflection, incomplete cervix exposure, severe bleeding, lesions covered by vaginal discharge). For each qualified case, her colposcopy images including saline image, acetic image and iodine image at the magnification of 7.5 were collected, as well as the corresponding clinical data including patient’s age, results of HPV testing and cytology, type of transformation zone (TZ),and pathologic diagnosis. All the colposcopy images were from ordinary electronic colposcope and photoelectric integrated colposcope (hereafter called ordinary images).The flowchart of case collection was shown in Fig. 8A. The research was approved by the Medical Ethics Committee of Women’s Hospital, School of Medicine, Zhejiang University, and written informed consent was obtained from all subjects. All the methods were performed in accordance with the relevant guidelines and regulations.
Women with different ages may manifest different degrees of reliability in HPV infection status and colposcopy impression. They are also applied to different screening strategies32. Thus, the ages of the patients were divided into three groups for better management (Table 7).
HPV testing was performed by food and drug administration (FDA) approved techniques, including Hybrid Capture 2 (HC2) hr-HPV DNA Test33 (Qiagen, Gaithersburg, MD), Cobas 4,800 Test34 (Roche Molecular system Inc, Pleasanton, CA), Cervista HPV HR Test35 (Hologic Inc., Madison, WI), and Aptima HPV Assay36 (Hologic Gen-Probe Inc., San Diego, CA).
The results of HPV testing were divided into negative and positive (Table 8).
The cytology diagnoses were divided into six categories, according to the Bethesda 2014 classification 37 (Table 9).
TZ types were divided into three categories according to the International Federation for Cervical Pathology and Colposcopy (IFCPC) colposcopy terminology38 (Table 10).
The pathological diagnoses were divided into normal, low-grade squamous intraepithelial lesion (LSIL, including the condylomatous variant), high-grade squamous intraepithelial lesion (HSIL) based on the 2014 World Health Organization (WHO) Classification of Tumors of the Female Genital Tract39.
The independent validation dataset of colposcopy images and the corresponding age, HPV testing results, cytology results, type of transformation zone and pathologic diagnosis were collected in order to better evaluate the established models. All cases were collected in women’s hospital, school of medicine, Zhejiang University from March 1st, 2019 to September 12nd 2019. And all colposcopy images were from high definition electronic colposcope (hereafter called high definition images). The exclusion criterions were the same as above.
The retrospective study was approved by the Medical Ethical Committee of Women’s Hospital, School of Medicine, Zhejiang University. (No. 20180059).
Data pre-process
One acetic image and one iodine image of each qualified case were kept and resized into 512*512 pixels. All the resized images of the modeling dataset were divided into 100 categories by K-means algorithm and randomly relocated into three sets of the training set, the valid set and the test set with the ratio of 8:1:1. Normal images, LSIL images and HSIL images were relocated separately so that they would be equally distributed into the three sets. Only the valid set was calculated to evaluate the performance of models. Pathologically diagnosed lesions were annotated by labelme40 software in every acetic image and every iodine image (Fig. 8B).
Text information including age, results of HPV testing and cytology, and TZ type were coded by the methods represented in Tables 1 to 4. As an example, a 45 year-old patient with HR-HPV positive and ASCUS cytology result, type 3 TZ, her texting code is 01001010000001.
Transfer learning model
In order to get high efficiency, a pretrained deep learning model was obtained by training a ResNet41 model from a database called ImageNet, which contains more than 1 million images of over 1,000 categories. On that basis, colposcopy images were input to fine-tune multi-modal ResNet classification model, U-Net42 segmentation model and Mask R-CNN43 detection model, which use the pre-trained ResNet model as backbone.
Multi-modal ResNet classification model to simply classify the images into two groups
Two ResNet-50 models were used for acetic image and iodine image, respectively. Cervix regions were firstly extracted due to other undesired information on the acetic and iodine images such as text, equipment and non-cervix tissues. Since clinical diagnosis were often made after a long comparison of the acetic and the iodine images, fusing the acetic image features and iodine image features during the training process can better capture cervical lesions and to offer a more scientific diagnosis. In the end, the coded non-image information of age, HPV testing result, cytology result and TZ type were input into the model and integrated with the fused image features. All the images will be classified into two groups: the negative group which means no squamous intraepithelial lesion (SIL, including LSIL and HSIL) in the cervix and the positive group which means one or more SIL were found in the cervix (Fig. 8C).
For classification model, the input image was scaled to 512 on the shorter edge. We used BCE loss with positive weith of 10. Batch size was set to 16. SGD optimizer was used with learning rate 1e−4, weight decay 1e−4 and momentum 0.9. Learning rate was multiplied with 0.9 when training loss was no longer reduced during 10 epochs.
U-Net segmentation model to segment the lesions apart from the normal areas
Same as the classification model, the U-Net model was also fine-tuned on the basis of the transfer-learning ResNet model. The colposcopy images were resized to 512*512 pixels, and each pixel was labeled as “1” for “lesion” or “0” for “normal” according to the annotations made by the colposcopy experts. In the end, all the lesions will be highlighted, representing the possible biopsy sites (Fig. 8D).
Taking every pixel as the object, recall and dice were calculated to evaluate the two models using the following formula: Recall = true positive pixels/ predicted positive pixels. DICE = 2*true predicted pixels/(predicted positive pixels + true positive pixels).
For segmentation model, the input image was scaled to 512 on the shorter edge. Focal loss was used. Batch size was set to 8. SGD optimizer was used with learning rate 1e-2, weight decay 1e-4 and momentum 0.9. Learning rate was multiplied with 0.9 when training loss was no longer reduced during 10 epochs.
Mask-R-CNN detection model to offer the final HSIL biopsy sites
Based on the transfer-learning ResNet model, the Mask R-CNN model detected lesion regions on colposcopy images according to the ground truth of delineating bounding boxes on existing segmentation annotation. Compared to the corresponding ground truth bounding boxes, the predicted ones offered by the detection model were considered positive when their Intersection over Union (IoU) value is more than 0.5. The loU is defined as the area of the intersection divided by the area of the union of a predicted bounding box (Bp) and a ground truth box (Bgt): loU = area (Bp ∩ Bgt)/area (Bp ∪ Bgt). In order to decrease the biopsy number, the model chose only the top 3 possible HSIL predicted bounding boxes in the premise of acceptable accuracy (Fig. 8E).
For detection model, the input image was scaled to 600 on the shorter edge. Batch size was set to 4. SGD optimizer was used with learning rate 2e−3, weight decay 1e−4 and momentum 0.9. We trained the model for 80 k iteration and learning rate was divided by 10 on 50 k, 70 k iteration.
For all the three models, we applied random color, random contrast, random saturation, and random hue transformation.
References
Bruni LAG, Serrano B, Mena M, Gómez D, Muñoz J, Bosch FX, de Sanjosé S.ICO/IARC Information Centre on HPV and Cancer (HPV Information Centre). Human papillomavirus and related diseases in china. Summary report. 17 June 2019.
Goodman A. Hpv testing as a screen for cervical cancer. BMJ, 2015,350(h2372).
Jin, J. Hpv infection and cancer. JAMA 319(10), 1058 (2018).
Castellsague, X. Natural history and epidemiology of hpv infection and cervical cancer. Gynecol. Oncol. 110(3 Suppl 2), S4-7 (2008).
Walboomers, J. M. M. et al. Human papillomavirus is a necessary cause of invasive cervical cancer worldwide. J. Pathol. 189, 12–19 (1999).
Snijders, P. J. et al. High-risk hpv testing on self-sampled versus clinician-collected specimens: a review on the clinical accuracy and impact on population attendance in cervical cancer screening. Int. J. Cancer 132(10), 2223–2236 (2013).
Wang, J. Analysis of the application values of different combination schemes of liquid-based cytology and high-risk human papilloma virus test in the screening of high-grade cervical lesions. Braz. J. Med. Biol. Res. 52(1), e7517 (2018).
Arbyn, M. et al. Accuracy of human papillomavirus testing on self-collected versus clinician-collected samples: a meta-analysis. Lancet Oncol. 15(2), 172–183 (2014).
Brown, B. H. & Tidy, J. A. The diagnostic accuracy of colposcopy - a review of research methodology and impact on the outcomes of quality assurance. Eur. J. Obstetrics Gynecol. Reprod. Biol. 240, 182–186 (2019).
Underwood, M. et al. Accuracy of colposcopy-directed punch biopsies: a systematic review and meta-analysis. BJOG: Int. J. Obstetrics Gynaecol. 119(11), 1293–1301 (2012).
Sideri M, Garutti P, Costa S, et al. Accuracy of colposcopically directed biopsy: Results from an online quality assurance programme for colposcopy in a population-based cervical screening setting in Italy. BioMed Res. Int., 2015, 614035.
Massad, L. S. et al. The accuracy of colposcopic grading for detection of high-grade cervical intraepithelial neoplasia. J. Lower Genital Tract Disease 13(3), 137–144 (2009).
Huh, W. K., Papagiannakis, E. & Gold, M. A. Observed colposcopy practice in us community-based clinics: The retrospective control arm of the improve-colpo study. J. Lower Genital Tract Disease 23(2), 110–115 (2019).
Liston, C. J. L. C. New machine-learning technologies for computer-aided diagnosis. Nat. Med. 24(9), 1306–1307 (2018).
Coudray, N. et al. Classification and mutation prediction from non–small cell lung cancer histopathology images using deep learning. Nat. Med. 24(10), 1559–1567 (2018).
Ting, D. S. W. et al. Ai for medical imaging goes deep. Nat Med 24(5), 539–540 (2018).
Kermany, D. S. et al. Identifying medical diagnoses and treatable diseases by image-based deep learning. Cell 172(5), 1122–1131 (2018).
Hosny, A. et al. Artificial intelligence in radiology. Nat. Rev. Cancer 18(8), 500–510 (2018).
Chilamkurthy, S. et al. Deep learning algorithms for detection of critical findings in head ct scans: a retrospective study. Lancet 392(10162), 2388–2396 (2018).
Ahmad, O. F. et al. Artificial intelligence and computer-aided diagnosis in colonoscopy: current evidence and future directions. Lancet Gastroenterol. Hepatol. 4, 71–80 (2019).
Acosta-Mesa, H. G., Cruz-Ramirez, N. & Hernandez-Jimenez, R. Aceto-white temporal pattern classification using k-nn to identify precancerous cervical lesion in colposcopic images. Comput. Biol. Med. 39(9), 778–784 (2009).
Asiedu, M. N. et al. Development of algorithms for automated detection of cervical pre-cancers with a low-cost, point-of-care, pocket colposcope. IEEE Trans. Bio-Med. Eng. 66(8), 2306–2318 (2019).
Hu, L. et al. An observational study of deep learning and automated evaluation of cervical images for cancer screening. J. Natl. Cancer Inst. 111(9), 923–932 (2019).
Bai, B. et al. Automatic segmentation of cervical region in colposcopic images using k-means. Australas. Phys. Eng. Sci. Med. 41(4), 1077–1085 (2018).
T. Xu, H. Zhang, X. Huang, S. Zhang, D.N. Metaxas. Multimodal deep learning for cervical dysplasia diagnosis. MICCAI, 2016.
Mehlhorn, G. et al. Computer-assisted diagnosis in colposcopy: results of a preliminary experiment?. Acta Cytol. 56(5), 554–559 (2012).
Sato, M. et al. Application of deep learning to the classification of images from colposcopy. Oncol. Lett. 15(3), 3518–3523 (2018).
Miyagi, Y., Takehara, K. & Miyake, T. Application of deep learning to the classification of uterine cervical squamous epithelial lesion from colposcopy images. Mol. Clin. Oncol. 11(6), 583–589 (2019).
Adsul, P. et al. Implementing community-based cervical cancer screening programs using visual inspection with acetic acid in India: a systematic review. Cancer Epidemiol. 49, 161–174 (2017).
Baum, M. E. et al. Colposcopic accuracy of obstetrics and gynecology residents. Gynecol. Oncol. 103(3), 966–970 (2006).
Mitchell, M. F., Schottenfeld, D., Tortolero-Luna, G., Cantor, S. B. & Richards-Kortum, R. Colposcopy for the diagnosis of squamous intraepithelial lesions: a meta-analysis. Obstet. Gynecol. 91, 626–631 (1998).
Sawaya, G. F. & Smith-McCune, K. Cervical cancer screening. Obstet. Gynecol. 127(3), 459–467 (2016).
Qiao, Y.-l et al. A new hpv-DNA test for cervical-cancer screening in developing regions: A cross-sectional study of clinical accuracy in rural china. Lancet Oncol. 9(10), 929–936 (2008).
Isidean, S. D., Coutlee, F. & Franco, E. L. Cobas 4800 hpv test, a real-time polymerase chain reaction assay for the detection of human papillomavirus in cervical specimens. Expert Rev. Mol. Diagn. 1, 5–16 (2014).
Boers, A. et al. Clinical validation of the cervista hpv hr test according to the international guidelines for human papillomavirus test requirements for cervical cancer screening. J. Clin. Microbiol. 52(12), 4391–4393 (2014).
Iftner, T. et al. Longitudinal clinical performance of the rna-based aptima human papillomavirus (ahpv) assay in comparison to the DNA-based hybrid capture 2 hpv test in two consecutive screening rounds with a 6-year interval in germany. J. Clin. Microbio. 57(1), 1–12 (2019).
Nayar, R. & Wilbur, D. C. The pap test and Bethesda 2014. Cancer Cytopathol. 123(5), 271–281 (2015).
Tatti, S., Bornstein, J. & Prendiville, W. Colposcopy: a global perspective: Introduction of the new ifcpc colposcopy terminology. Obstet. Gynecol. Clin. North Am. 40(2), 235–250 (2013).
Reich, O. et al. Precancerous lesions of the cervix, vulva and vagina according to the 2014 who classification of tumors of the female genital tract. Geburtshilfe Frauenheilkd. 75(10), 1018–1020 (2015).
Wada K. Labelme: Image polygonal annotation with python. https://github.com/wkentaro/labelme, 2016.
Fulton, L. V. et al. Classification of alzheimer’s disease with and without imagery using gradient boosted machines and resnet-50. Brain Sci. 9(9), 212 (2019).
Tang, P. et al. Efficient skin lesion segmentation using separable-unet with stochastic weight averaging. Comput. Methods Prog. Biomed. 178, 289–301 (2019).
He K, Gkioxari G, Dollar P, et al. Mask r-cnn. IEEE transactions on pattern analysis and machine intelligence, 2018.
Acknowledgements
This study was supported by grants from the National Key Research and Development Program of China (No. 2016YFC1302903); the Key Research and Development Program of Zhejiang province (No. 2020C03025) and the Fundamental Research Funds for the Central Universities (No. 2017XZZX001-01). The retrospective study was approved by the Hospital Ethical Committee (No. 20180059).
Author information
Authors and Affiliations
Contributions
W.L., X.X., J.W., X.W. and C.Y. cooperated to design the research. X.X., W.L., X.W. and X.C. offered constructive advice throughout the research. C.Y., Y.Y., B.C., Y.C., Y.L. and Y.L. collected the data and labeled images. C.Y., Y.Y. and X.L. analyzed and interpreted the data. X.L. and J.W. implemented the network architecture and offered technical support. All authors reviewed the manuscript. All the authors listed have approved the manuscript publication.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the articleΓÇÖs Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the articleΓÇÖs Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yuan, C., Yao, Y., Cheng, B. et al. The application of deep learning based diagnostic system to cervical squamous intraepithelial lesions recognition in colposcopy images. Sci Rep 10, 11639 (2020). https://doi.org/10.1038/s41598-020-68252-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-68252-3
This article is cited by
-
Usability of a smartphone-compatible, confocal micro-endoscope for cervical cancer screening in resource-limited settings
BMC Women's Health (2024)
-
Development and validation of artificial intelligence-based analysis software to support screening system of cervical intraepithelial neoplasia
Scientific Reports (2024)
-
Augmenting Cervical Cancer Analysis with Deep Learning Classification and Topography Selection Using Artificial Bee Colony Optimization
SN Computer Science (2024)
-
Exploring data mining and machine learning in gynecologic oncology
Artificial Intelligence Review (2024)
-
Proceedings of the North American Society of Head and Neck Pathology Companion Meeting, New Orleans, LA, March 12, 2023: Oral Cavity Dysplasia: Why Does Histologic Grading Continue to be Contentious?
Head and Neck Pathology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.