Abstract
We present the first complete, closed genome sequences of Streptococcus pyogenes strains NCTC 8198T and CCUG 4207T, the type strain of the type species of the genus Streptococcus and an important human pathogen that causes a wide range of infectious diseases. S. pyogenes NCTC 8198T and CCUG 4207T are derived from deposit of the same strain at two different culture collections. NCTC 8198T was sequenced, using a PacBio platform; the genome sequence was assembled de novo, using HGAP. CCUG 4207T was sequenced and a de novo hybrid assembly was generated, using SPAdes, combining Illumina and Oxford Nanopore sequence reads. Both strategies yielded closed genome sequences of 1,914,862 bp, identical in length and sequence identity. Combining short-read Illumina and long-read Oxford Nanopore sequence data circumvented the expected error rate of the nanopore sequencing technology, producing a genome sequence indistinguishable to the one determined with PacBio. Sequence analyses revealed five prophage regions, a CRISPR-Cas system, numerous virulence factors and no relevant antibiotic resistance genes. These two complete genome sequences of the type strain of S. pyogenes will effectively serve as valuable taxonomic and genomic references for infectious disease diagnostics, as well as references for future studies and applications within the genus Streptococcus.
Similar content being viewed by others
Introduction
Streptococcus pyogenes, within the β-haemolytic, Lancefield group A Streptococcus (GAS)1, is an important clinically-relevant and strictly-human pathogen causing a wide range of diseases, including local and invasive infections (e.g., throat, skin infections, meningitis), severe toxin-mediated diseases (e.g., necrotizing fasciitis, scarlet fever, streptococcal toxic shock syndrome) and immune-mediated diseases (e.g., rheumatic fever, rheumatic heart disease, post-streptococcal glomerulonephritis)2. In 2005, it was estimated that more than 500,000 people were dying every year from severe diseases caused by GAS, as well as an estimated 600 million new cases of pharyngitis and 100 million new cases of pyoderma3. Thus, S. pyogenes is among the top-10 infectious causes of mortality in humans4. Moreover, S. pyogenes is the type species of the genus Streptococcus, the type genus of the family Streptococcaceae, and as a clinically-relevant bacterium, S. pyogenes has been continuously studied since it was first described5.
In recent decades, several next-generation and third-generation (i.e., long-read) sequencing technologies have emerged and are now widely used in many settings6. For instance, Illumina has led the field in high-throughput DNA sequencing, by providing highly accurate and relatively inexpensive sequence reads. However, their short lengths (few hundred base-pairs) have restricted efficacy to resolve problematic genomic regions (e.g., repeats, ribosomal operons, long sequence motifs), sometimes yielding fragmented and incomplete assemblies7. Meanwhile, PacBio provides long reads (several kilobase-pairs) with high consensus accuracy, generally yielding complete bacterial genome sequences. However, high capital costs of PacBio platforms have constrained accessibility to users, who normally access them via commercial/institutional sequencing services. Additionally, requirements of large quantities of high-quality DNA make PacBio sequencing relatively laborious, time-consuming and impractical for some applications. More recently, Oxford Nanopore Technologies launched the MinION portable sequencer, which provides ultra-long reads of as many as two million base-pairs8, requiring simple, rapid and cost-effective DNA library preparation protocols. Nanopore-sequencing has been demonstrated to resolve very-long repetitive regions that not even PacBio-sequencing could resolve9. However, inaugural high error rates (> 30%; currently ~ 7%)10,11,12 caused some degree of doubt within the scientific community, although more recent developments and studies have allayed much of the initial scepticism.
Resulting from these technological developments, in 2019-06-29, 1,883 genome sequences of S. pyogenes were publicly available in GenBank, of which 195 were complete. However, of those 195, only the complete genome sequences presented in this study represented the type and an important reference strain of the species.
Here, we present the first complete genome sequences of the type strain of S. pyogenes (NCTC 8198T = CCUG 4207T), determined by two different approaches: S. pyogenes NCTC 8198T completed using only PacBio reads; and S. pyogenes CCUG 4207T completed by combining Illumina and Oxford Nanopore reads. Both assemblies were identical in length and sequence nucleotide content, demonstrating the possibility of surpassing the inherent error rate of the Nanopore sequencing technology, by combining Illumina reads and, thus, obtaining an assembly as accurate as the one obtained with the PacBio approach.
Materials and methods
Cultivation conditions and DNA extractions
S. pyogenes NCTC 8198T was cultivated at the laboratories of the National Collection of Type Cultures (NCTC, London, UK) on Columbia blood agar (Columbia Agar Base plus 5% horse blood, Thermo Fisher Scientific, Waltham, MA, USA), at 37 °C. Genomic DNA was isolated, using the MasterPure Gram Positive DNA Purification Kit (Epicentre, Madison, WI, USA), for PacBio sequencing. S. pyogenes CCUG 4207T was cultivated at the laboratories of the Culture Collection University of Gothenburg (CCUG, Gothenburg, Sweden) on Chocolate agar medium (Brain Heart Infusion Agar with 10% heat-lysed defibrinated horse-blood, 15% horse-serum and fresh yeast extract, prepared by the Substrate Unit, Department of Clinical Microbiology, Sahlgrenska University Hospital), with 5% CO2, at 36 °C. Genomic DNA was extracted from fresh pure biomass, using a Wizard Genomic DNA Purification Kit (Promega, Madison, WI, USA), for Illumina sequencing, and a modified version13 of a previously described protocol14, for Oxford Nanopore sequencing (Fig. 1).
Sequencing workflows. Illustration showing the origin, the strain passage and indicating the whole-genome sequencing workflows performed, in parallel, by the NCTC and the CCUG to determine the complete genome sequence of the type strain of S. pyogenes. Map created using the on-line server MapChart (www.mapchart.net).
PacBio sequencing
Genomic DNA of S. pyogenes NCTC 8198T was sheared with a 26 G blunt Luer-Lok needle and used to prepare two 10 to 20-kb PacBio SMRT libraries, following the manufacturer’s recommendations. The libraries were sequenced using the P6-C4 chemistry on a Single Molecule, Real-Time (SMRT) cell, using a PacBio RSII platform (Pacific Biosciences of California, Inc., Menlo Park, CA, USA) (www.pacb.com), at the Wellcome Trust Sanger Institute (Hinxton, UK).
Illumina sequencing
Genomic DNA of S. pyogenes CCUG 4207T was used to prepare a standard Illumina library, with an insert size ranging from 130 to 680 bp, following an optimized protocol (GATC Biotech, Konstanz, Germany) and using standard Illumina adapter sequences. The library was sequenced at GATC Biotech (Konstanz, Germany), using an Illumina HiSeq 2500 instrument (Illumina, Inc., San Diego, CA, USA) (www.illumina.com) to generate paired-end reads of 126 bp.
Oxford Nanopore sequencing
High-molecular weight DNA of S. pyogenes CCUG 4207T was used to prepare a sequencing library, using a Rapid Sequencing Kit (SQK-RAD003) (Oxford Nanopore Technologies, Ltd., Oxford, UK), according to manufacturer’s instructions. The library was sequenced at the CCUG laboratories, on a MinION sequencer (Oxford Nanopore Technologies, Ltd., Oxford, UK) (www.nanoporetech.com), using a Flow Cell model FLO-MIN 106 version R9.4. The sequencing was performed with the software MinKNOWN, version 1.10.23 (Oxford Nanopore Technologies, Ltd., Oxford, UK), selecting the 48-h sequencing script (NC_48Hr_sequencing_FLO-MIN106_SQK-RAD003), with default parameters.
PacBio de novo assembly
PacBio sequence reads from both SMRT sequencing runs were used in the assembly. Read quality was assessed, using NanoPlot version 1.13.015. Sequence reads were auto-error-corrected and assembled de novo, using the Hierarchical Genome Assembly Process (HGAP) version 316. The assembled sequence was polished with the consensus calling algorithm, Quiver, version 1. The ends of the final assembly were trimmed (i.e., eliminating sequence redundancy) manually, to circularize the genome, and the genome sequence reorganized to start with the dnaA gene, which encodes the chromosomal replication initiator protein, DnaA. Assembly statistics were obtained, using QUAST, version 4.517.
Illumina-Nanopore hybrid de novo assembly
The quality of the Illumina paired-end reads was analysed with FastQC, version 0.11.3 (www.bioinformatics.babraham.ac.uk/projects/fastqc/). Subsequently, the reads were sub-sampled and trimmed, using Sickle, version 1.3318, with a Phred quality score threshold of Q30. Meanwhile, the FAST5 files containing the raw data generated by the Oxford Nanopore sequencing run were processed with the Oxford Nanopore basecalling pipeline, Albacore, version 2.0.2., and the quality was analysed with NanoPlot version 1.13.015. Only reads with a quality score greater than Q7 were used for the assembly (i.e., classified as Pass by Albacore). Afterwards, a hybrid de novo assembly, using both Illumina and Oxford Nanopore reads was performed with SPAdes, version 3.11.019,20. The assembly was performed with the flag --careful enabled, to map the Illumina reads back to the assembly with BWA, version 0.7.12-r103921 and to reduce the number of mismatches and short indels. The ends of the assembly were trimmed manually, and the sequence reorganized to start with dnaA, as was done for the PacBio assembly. Assembly statistics were obtained, using QUAST, version 4.517.
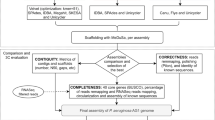
Genome sequence comparisons
Once assembled, closed and completed, the genome sequences of S. pyogenes NCTC 8198T and S. pyogenes CCUG 4207T were compared (Fig. 1). Firstly, both genome assemblies were aligned, using BLASTN, version 2.2.1022. Secondly, all the raw Illumina paired-end reads were mapped against the complete and closed genome sequences, using CLC Genomics Workbench, version 10.0 (Qiagen Aarhus A/S, Aarhus, Denmark), and a Basic Variant Detection 2.0 analysis was performed, using the same software, using a minimum frequency of 35% (default).
Genome annotations and characterization
The genome sequence of S. pyogenes NCTC 8198T was initially annotated with Prokka23, and submitted to the European Nucleotide Archive24. Afterwards, the genome sequence was re-annotated, with the NCBI Prokaryotic Genome Annotation Pipeline (PGAP), version 4.125, and deposited in the NCBI Reference Sequence (RefSeq) database26. The genome sequence of S. pyogenes CCUG 4207T was submitted to GenBank27. Subsequently, the sequence was annotated with PGAP, version 4.7, and deposited in RefSeq. The latest of these annotations (i.e., PGAP version 4.7) was used for further analyses and to construct a genome atlas with the on-line server GView (www.gview.ca)28.
The on-line tool, PHASTER (www.phaster.ca)29, was used to search for prophages inserted in the chromosome, while the tool CRISPRFinder30 was used to search clustered, regularly interspaced short palindromic repeat (CRISPR) arrays. The consensus sequences of the direct repeats were classified, using CRISPRmap v2.1.3-201431,32, and the crRNA-encoding strand determined, using CRISPRstrand32, implemented in CRISPRmap. Additionally, the tool CRISPRone33 was used to confirm the detected CRISPR arrays and to identify possible CRISPR-associated genes (cas). Spacer sequences of CRISPR arrays were analysed with BLASTN, version 2.2.1022, against the complete genome sequence of S. pyogenes NCTC 8198T (= CCUG 4207T). Searches for virulence factors across the genome were done against the protein sequences of the curated core dataset (3,200 protein sequences) of the Virulence Factors Database (VFDB; www.mgc.ac.cn/VFs/)34, using BLASTP, version 2.2.1022. Only hits with ≥ 50% of identity over ≥ 50% of the DNA sequence length were considered. For subtyping, the emm gene sequence was analysed, with BLASTN 2.2.27 +, against the emm gene database of the Streptococcus Laboratory (Centers for Disease Control and Prevention, CDC, USA; www2a.cdc.gov/ncidod/biotech/strepblast.asp). Potential antibiotic resistance genes were searched with the tool, Resistance Gene Identifier (RGI), of the Comprehensive Antibiotic Resistance Database (CARD; https://card.mcmaster.ca/)35. Only results classified as “perfect” or “strict” were considered.
Results
Whole-genome sequencing
The complete genome sequence of the type strain of S. pyogenes (NCTC 8198T = CCUG 4207T) was determined, using three different sequencing technologies. The yield of each sequencing run is summarized in Table 1.
PacBio
The two PacBio sequencing runs yielded a total of 420 and 694 Mb of raw data, distributed in 163,469 and 163,274 sequence reads, with average lengths of 2,570 and 4,250 bp, respectively. The sequence read length N50s were 4,555 and 9,809 bp, with longest reads of 35,267 and 41,160 bp and mean read Phred quality scores of 2.7 and 3.8, respectively.
Illumina
The Illumina genome sequencing of S. pyogenes CCUG 4207T yielded 1,292.9 Mb distributed in 10,260,942 paired-end reads of 126 bp with 94.1% of the reads exhibiting an average Phred quality score (i.e., for each sequence read, the arithmetic mean of every base quality) ≥ Q30. The total average Phred quality score was 35. Down-sampling and trimming with Sickle left 355.4 Mb distributed in 3,083,010 paired-end reads of an average length of 115 bp.
Oxford Nanopore
The Oxford Nanopore whole-genome sequencing of S. pyogenes CCUG 4207T was done, using the Rapid Sequencing Kit SQK-RAD003 with a MinION sequencer, on a Flow Cell (FLO-MIN 106, R9.4) with 465 active channels. The run yielded 1,121.6 Mb distributed in 135,020 reads with a mean length of 8,307 bp; the read length N50 was 14,598 bp and the longest read was 97,527 bp with a mean read Phred quality score of 11.5. In total, 130,764 reads (96.85%) were classified as Pass by Albacore and used for the hybrid assembly.
Genome sequence assemblies
The assembly of the PacBio reads with HGAP, followed by a polishing step performed with Quiver, yielded a complete and closed sequence. Trimming of the ends (i.e., overlapping redundant sequences) resulted in a final sequence of 1,914,862 bp, representing the genome of S. pyogenes NCTC 8198T. In parallel, the de novo hybrid assembly of the trimmed Illumina sequence reads plus the basecalled Oxford Nanopore reads also resulted in a complete and closed sequence. The trimming of the ends yielded a final sequence of 1,914,862 bp. Analysis performed with QUAST confirmed that both assemblies did not contain any gaps (i.e., no N’s).
The final complete and closed genome sequence of S. pyogenes CCUG 4207T was analysed, using BLASTN, against the complete genome sequence of S. pyogenes NCTC 8198T. The analysis yielded a match of 1,914,862 bp with 100% of identity. Afterwards, for further quality control, the entire set of raw Illumina paired-end reads (i.e., 1,292.9 Mb, coverage: 675 X) was mapped against the two complete genome sequences. A variant calling analysis was performed, and no variants were found in any of the cases. Thus, the two independent and parallel strategies of sequencing and assembly resulted in a genome sequence of identical length and identity of 1,914,862 bp and a GC content of 38.5% (Table 2).
Characterization of the complete genome sequence
Annotation
The latest annotation (i.e., S. pyogenes CCUG 4207T annotated with PGAP version 4.7 and available in RefSeq) revealed a total of 2,009 genes, of which 1,920 were coding sequences (CDSs). The annotation detected 89 RNA genes, which included 67 tRNA genes, four non-coding RNA genes and 18 ribosomal genes distributed in six complete ribosomal operons. Additionally, 60 pseudogenes were annotated. A total of 306 genes (14.8%) were annotated as “hypothetical proteins”. A genome atlas of this annotation version is depicted in Fig. 2.
Genome atlas of the type strain of Streptococcus pyogenes. The atlas was built with the genome sequence annotated with PGAP 4.7 and available in RefSeq (NZ_CP028841.1), using the on-line server GView. Labelling, from outside to inside: backbone; CDSs and prophage regions (coloured in blue); GC content deviations (GC-rich towards outside, GC-poor towards inside); GC skew (excess of guanine over cytosine towards outside, and vice versa) and CDSs coloured by COG categories (if assigned).
Prophages
Five putative prophages were detected using the software PHASTER, four marked by the software as ‘intact’ (completeness score > 90) and one as ‘questionable’ (completeness score = 70–90), with a GC content ranging from 37.4 to 39.1% (Table 3). The largest prophage region was 56,926 bp and the shortest 41,886 bp. Overall, the five regions add up to 234,671 bp, which represents 12.3% of the genome size (Fig. 2). In total, according to the PGAP annotation, the five prophage regions encompass 321 CDSs, which represents a 16.72% of the total 1,920 CDSs annotated by PGAP in the genome sequence.
CRISPR-Cas systems
The analysis of the genome with CRISPRFinder revealed the presence of a CRISPR array (positions: 1,317,644–1,317,938 bp), composed of five direct repeats of 32 bp and four spacers of sizes ranging from 33 to 35 bp. The consensus sequence of the direct repeats was classified, with CRISPRmap, into the family 5 and structure motif 3 (Fig. 3). The analysis with CRISPRone revealed seven CRISPR-associated (cas) genes, located adjacent to the CRISPR array (cas3, cas5, cas8c, cas7, cas4, cas1 and cas2; locus tags: DB248_RS07080–DB248_RS07050). This architecture corresponds to the Class 1, subtype I-C of the updated evolutionary classification of CRISPR-Cas systems36. However, cas3 was frame-shifted due to a single-nucleotide deletion. The frameshift was confirmed by manually inspecting the mapped Illumina reads.
CRISPR-Cas system of the type strain of S. pyogenes. (A) The genomic architecture of the S. pyogenes CCUG 4207T (= NCTC 8198T) CRISPR-Cas subtype I-C system, formed by seven cas genes followed by six direct repeats (DR) and five spacers. The stepped line crossing the cas3 gene indicates frameshift. (B) The hairpin structure motif of the consensus sequence of the direct repeats of the CRISPR array of S. pyogenes CCUG 4207T (= NCTC 8198T).
The BLASTN analyses of the spacer sequences against the whole genome sequence revealed that the sequence of the second spacer of the CRISPR array (positions 1,317,807–1,317,839 bp) shows 100% identity against a sequence of the prophage region SF130.1 (positions: 551,171–551,203; identities = 33/33), located in a gene encoding a phage predominant capsid protein (DB248_RS03050). Additionally, the sequence of the third spacer shows high similarity to a sequence of the prophage region SF130.5 (positions: 1,388,239–1,388,206 bp; identities: 32/34), which is part of a gene encoding a hypothetical protein (DB248_RS07380). Finally, the sequence of the fourth spacer presents a high degree of homology to a sequence of the prophage region SF130.2 (positions: 850,316–850,348; identities = 32/33), which is also part of a gene encoding a hypothetical protein (DB248_RS04545).
Additionally, four putative cas genes (cas9, cas1, cas2, csn2; locus tags: DB248_RS04305–DB248_RS04320) were located between positions 812,387 and 818,352, adjacent to a gene encoding a phage integrase of the prophage region 2 (DB248_RS04325). The architecture of this locus corresponds to the Class 2, subtype II-A of the CRISPR-Cas systems classification, although no CRISPR arrays were found in the surrounding region.
Virulence factors
The protein sequences of S. pyogenes CCUG 4207T (= NCTC 8198T) were analysed with BLASTP against the core dataset of the VFDB. Numerous genes encoding putative virulence factors were detected (Supplementary Table S1), several of them within the prophage regions. One of the genes was emm, encoding the surface protein M (DB248_RS09295), one of the major virulence factors of S. pyogenes, which provides protection against the immune system and has been used for strain serological typing37. The BLATN analysis of the sequence of the emm gene against the emm database of the Streptococcus Laboratory (CDC, USA) confirmed that it is type 1.0. Genes related with the synthesis of the hyaluronic acid capsule were also found (hasA, DB248_RS09950; hasB, DB248_RS09955; hasC, DB248_RS09965). This polysaccharide capsule is a key virulence factor involved in adhesion, tissue invasion38 as well as in molecular mimicry for immune evasion39. The sic gene encoding the hypervariable streptococcal inhibitor of complement-mediated lysis (SIC) was also found (DB248_RS09285)40. Two genes encoding DNAses were also found: mf/spd (DB248_RS09395), encoding the DNaseB41,42 and mf3/spd3 (DB248_RS06505), within the prophage region SF130.4, encoding the DNaseC43. A gene encoding the hyaluronate lyase HylA was detected (DB248_RS04240). HylA has been suggested to facilitate spread of large molecules and to play a nutritional role for S. pyogenes, by disrupting host-tissue as well as its own capsule, allowing growth on hyaluronic acid as carbon source44. In addition, four prophage-associated hyaluronidase-encoding genes were found (DB248_RS03100, DB248_RS04540, DB248_RS06545, DB248_RS07275), located in the prophage regions SF130.1, SF130.2, SF130.4 and SF130.5, respectively, which may act as additional spreading factors45. These enzymes seem to be useful for phages to penetrate the capsule of hyaluronic acid46. Additionally, an ideS/mac gene, encoding an immunoglobulin G-degrading enzyme, was found (DB248_RS03795). This exoenzyme shields the cells from being opsonized by IgG antibodies, by cleaving their heavy chain47. A gene encoding a C5a peptidase was also found (scpA, DB248_RS09275). This peptidase degrades the chemotactic complement factor C5a48, thus preventing the C5a-based recruitment of neutrophils and other inflammatory cells to the site of infection49. Moreover, a gene encoding the streptokinase A (ska, DB248_RS09135) was found, which catalyses the conversion of plasminogen to plasmin, a serine protease that facilitates tissue invasion by degrading proteins of the extracellular matrix50. Additionally, several genes encoding putative streptococcal exotoxins were detected: speA (DB248_RS05660), encoding a pyrogenic exotoxin type A and located in the prophage region SF130.3; speB (DB248_RS09375), encoding a pyrogenic exotoxin type B; speG (DB248_RS01135), encoding a pyrogenic exotoxin type G; speJ (DB248_RS01990) encoding a pyrogenic exotoxin type J precursor; and smeZ (DB248_RS09220), encoding the streptococcal mitogenic exotoxin Z. Genes for pili biosynthesis were also detected (cpa, DB248_RS00770; lepA, DB248_RS00775; fctA, DB248_RS00780; srtC1, DB248_RS00785; fctB, DB248_RS00790), which have been shown to play roles in biofilm formation and attachment to pharyngeal cells51. Moreover, several genes encoding putative adhesion-related proteins were found (e.g., fibronectin binding proteins), such as fbaA (DB248_RS09270), encoding an F2-like fibronectin-binding protein, or fbp54 (DB248_RS04160). FBP54 has been shown to play a role in the adhesion of GAS to host cells52,53. An lbp gene was also detected (DB248_RS09265), codifying the Lbp laminin-binding protein, involved in adhesion to epithelial cells54 and suggested to play a role in zinc homeostasis55. A grab gene, encoding a G-related α2-macroglobulin-binding protein (GRAB), has also been detected (DB248_RS06200). GRAB is a surface protein that inhibits unwanted proteolysis through a high affinity for α2-macroglobulin, a proteinase inhibitor of human plasma56.
Antibiotic resistances
The analysis of the genome sequences with RGI (CARD) revealed one gene related with antibiotic resistance and classified as “strict”. The gene encodes a putative ABC transporter ATP-binding protein (locus tag: DB248_01185), located downstream of a gene encoding another ABC transporter ATP-binding protein (DB248_01180). Gene products show 67 and 66% sequence identity to ABC transporters PatB and PatA of S. pneumoniae TIGR4, whose overexpression has been linked to fluoroquinolone resistance57.
Discussion
Here we present the first complete genome sequence of S. pyogenes NCTC 8198T = CCUG 4207T, the type strain of the type species of the genus Streptococcus, the type genus of the family Streptococcaceae. The sequence has been determined twice, using two fully independent but parallel strategies: PacBio-solo and Illumina plus Oxford Nanopore sequencing. Both strategies have yielded 100% identical complete genome sequences, thus demonstrating that hybrid approaches can completely mitigate the error rate of long read sequences.
The type strain of S. pyogenes (NCTC 8198T = CCUG 4207T) was isolated as strain S.F. 130 in Manchester, UK, from a throat swab of a scarlet fever case. The strain was provided by William W. C. Topley (University of Manchester) to Frederick Griffith (Pathological Laboratory of the Ministry of Health), who used it for the preparation of Type 1 agglutination sera, in a study of scarlatinal streptococci58. In 1950, the strain was deposited at the NCTC by Robert E. O. Williams (Central Public Health Laboratory, Colindale, London, UK) and, in 1974, the NCTC strain was deposited at the CCUG (Fig. 1). After decades being available to the scientific community, the strain has served as a taxonomic reference point and has been used in numerous studies. Today, the strain is also preserved and publicly available in other culture collections, e.g., ATCC 12344T (USA) = BCRC 14758T (Taiwan) = CECT 985T (Spain) = CIP 56.41T (France) = DSM 20565T (Germany) = JCM 5674T (Japan) = LMG 14700T (Belgium) = S.F. 130T. To date, nearly 2,000 genome sequences of S. pyogenes are available in GenBank/ENA/DDBJ, of which nearly 200 are complete genome sequences. In fact, the first complete genome sequence of a strain of S. pyogenes was determined almost 20 years ago59. However, despite the clinical relevance of the species and the taxonomic importance of the type strain, this is the first complete genome sequence of the type strain of S. pyogenes that has been determined.
Following sequencing, two different sequence assembly strategies were used. While S. pyogenes NCTC 8198T was assembled de novo, using only PacBio reads, S. pyogenes CCUG 4207T was assembled de novo, using both short Illumina and long Oxford Nanopore reads. Surprisingly, both approaches yielded fully identical complete genome sequences of 1,914,862 bp. Recently, numerous studies have reported high quality genome assemblies, obtained by combining high-quality Illumina reads and long Oxford Nanopore reads60,61,62. However, to our knowledge, this is the first study reporting two identical complete genome sequences determined with different methodologies.
The fact that two wholly independent approaches have yielded identical sequences demonstrates the high quality of these genome sequences. On the one hand, the NCTC strain was sequenced, using a PacBio RSII platform, which, as expected, yielded long and relatively inaccurate raw sequence reads (indicated in Table 1, by the low Phred quality scores). However, due to their random distribution, sequencing errors can be corrected during assembly, by high coverage63. On the other hand, the CCUG strain was sequenced with the highly accurate Illumina HiSeq 2500 and the Oxford Nanopore MinION device, which, as expected, yielded long raw sequence reads with low accuracy (indicated in Table 1), Interestingly, Nanopore reads exhibit a higher average Phred quality score than PacBio reads. However, because of the less random distribution of the errors (e.g., misinterpretation of homopolymers)64, the inclusion of the high-quality Illumina reads, during or after the assembly, is crucial to obtain an accurate genome sequence.
These results demonstrate how a hybrid strategy, combining Illumina and Oxford Nanopore sequencing, can provide results as accurate as high coverage PacBio-solo sequencing. This should help to reduce the scepticism generated by the initially high error rate of Oxford Nanopore10,11,12. In this study, the identical results obtained by both strategies are an indicator of the high quality and accuracy of this genome sequence, which makes it a definitive genomic reference of the species as well as a good model candidate for being used in future evaluations of bacterial genome assemblers.
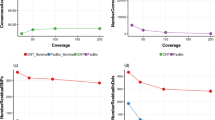
Furthermore, it is noteworthy that the hybrid assembly was performed with SPAdes (i.e., a relatively user-friendly, well-established and widely-used de novo genome assembler), as the assembly itself only required a single command line and did not involve complex and tedious methodologies. Nevertheless, further strategies have been developed in recent years to perform de novo hybrid assemblies combining short and long reads (e.g., Unicycler and MaSuRCA), aiming to cover the shortcomings of each technology with the advantages of the other65,66. Alternative strategies involve only-Nanopore de novo assemblies67, which can be afterwards ‘polished’ by other pieces of software (e.g., Pilon and/or Racon) in order to improve their accuracy68,69. In addition, all these strategies and methodologies can be complementary to each other, as one protocol might be more or less useful under particular conditions, while the other one could be the opposite. For instance, SPAdes relies on the short reads to create a first draft assembly and afterwards perform a scaffolding step, while Canu performs an only-long-read de novo assembly67 which can be optionally followed by a polishing step with short reads, using software like Pilon68.
In any case, despite the lack of differences between the two determined genome sequences, genomic variations could have been expected, as there have been several passages between strain NCTC 8198T and strain CCUG 4207T, and cultivation conditions and DNA preparation methods were different between both culture collections. These circumstances evidently increase the probability of having natural genotypic and phenotypic changes70,71,72. As a practical example, the ATCC recommends users to do no more than five passages from ATCC Genuine Cultures. For this reason, i.e., to reduce risk of natural alterations, the same starting material and DNA preparation should had been used. Nonetheless, this genome sequence will be a definitive reference of the type strain of S. pyogenes.
The vast amount of genomic data that the current “next-generation” and “third-generation” sequencing platforms generate, can be used for bacterial systematics and taxonomy, which traditionally has been based on observations of phenotypic features, DNA G + C content, DNA-DNA hybridization similarities and sequence determinations and analyses of marker genes, such as 16S rRNA73. Recently, numerous studies have shown the effectivity, high resolution and discriminative power of whole-genome sequence-based comparative studies74,75. In fact, several methods and tools have been developed for analysing whole-genome sequence similarities, i.e., Average Nucleotide Identity (ANI)76, which can be calculated, using JSpecies77,78 and in silico DNA-DNA hybridization, which can be calculated, using the Genome-to-Genome Distance Calculator (GGDC)79. Other interesting tools include the Type Strain Genome Server80 and TrueBac ID81, both high-throughput on-line servers for genome sequence-based taxonomy, dependent upon curated databases of bacterial species type strain genome sequences. Despite the availability of such tools, public databases contain numerous misclassified genome sequences74,75,82, most likely due to disregard for taxonomic controls, but also because genome sequences of the type strains of many species have not yet been determined83 or are not reliable, even because genome sequences may have been erroneously labelled as “type strains”84. Global efforts and initiatives are underway to curate public databases85, as well as to sequence the genomes of type strains83,86. Since January 2018, the International Journal of Systematic and Evolutionary Microbiology (IJSEM), the official publication of the International Committee on Systematics of Prokaryotes (ICSP) and the journal of record for publication of novel microbial taxa, has required authors describing new taxa to provide the genome sequence data; the genome of an organism encodes the basis of its biology and, therefore, is the fundamental basis of information for understanding the organism. Furthermore, genome sequences of the type strains of bacterial and archaeal species are crucial as reference points for identifying and classifying genetic and metagenomic data87,88.
Five prophage regions have been detected in the genome sequence of S. pyogenes CCUG 4207T (= NCTC 8198T), encompassing 17% of the CDSs of the genome. These results agree with initial reports of genome sequences of S. pyogenes, which already confirmed a high prevalence of prophages inserted in chromosomes of the species59,89,90. Overall, numerous studies have shown the crucial role of bacteriophages in the ecology, pathogenicity and the evolution of S. pyogenes strains. In fact, prophages have been linked with the recent resurgence of M1-GAS-associated invasive diseases91.
CRISPR-Cas systems are adaptive immune systems that are widely spread in bacteria36,92. These systems have been previously found in several strains of S. pyogenes, and an inverse correlation has been observed between them and the number of prophages inserted in the genome93. Contrary to this observation, we found a complete CRISPR-Cas system together with five putative prophages in the genome sequence of S. pyogenes CCUG 4207T (= NCTC 8198T). Additionally, high similarity was found between the sequences of the spacers of the CRISPR arrays and three of the prophage regions, one of them being 100% identical. However, we detected a frameshift, caused by a single-nucleotide deletion in the cas3 gene, which encodes an endonuclease/helicase that is essential for CRISPR-Cas interference94. Therefore, the CRISPR-Cas system most likely is non-functional due to this truncation, thus leaving a freeway for infection by bacteriophages, which could explain the presence of five prophage regions inserted within the chromosome. In addition, the short number of spacers also suggests that the CRISPR-Cas system might also not be active in acquisition of new spacers.
As a major human pathogen, S. pyogenes has a great repertoire of virulence factors, some of which are intrinsic and shared among almost all strains, while others might be present only in certain strains34 or serotypes (e.g., EndoS2, exclusive of serotype M49)95. In this study, the analysis of the protein sequences of S. pyogenes CCUG 4207T (= NCTC 8198T) against the VFDB has provided insight into the virulence potential of this strain, revealing the presence of numerous prominent virulence factors. In particular, this strain was isolated from a throat swab from a scarlet fever patient58. For many years, scarlet fever has been associated with S. pyogenes strains producing pyrogenic toxins96. In accordance to this, several streptococcal pyrogenic exotoxins have been found, one of them encoded in a prophage region. Additionally, several of the other virulence factors have also been found encoded in prophage regions, highlighting the role of prophages in the pathogenicity of S. pyogenes strains. In any case, further studies will be needed to uncover the full pathogenic potential of this strain, with emphasis in revealing the role of the still high number of CDSs annotated as ‘hypothetical proteins’, which represent 15% of the 2,009 genes annotated.
The only antibiotic resistance-gene detected in the genome sequence was a gene encoding an ABC transporter ATP-binding protein; overexpression has been associated with fluoroquinolone resistance for homologues of this gene in Streptococcus pneumoniae57. This lack of significant antibiotic resistance genes was expected, as the type strain of S. pyogenes was isolated before the antibiotic era (i.e., before 1928). In addition, the current antibiotic resistance problem is not that great yet among S. pyogenes as it is among other bacterial species and taxa, with penicillin remaining the drug of choice, despite numerous decades of use97.
Conclusions
Here we present the first complete genome sequences of the type strain of S. pyogenes (NCTC 8198T = CCUG 4207T = S.F. 130T), the type species of the genus Streptococcus, the type genus of the family Streptococcaceae and a major human pathogen. These genome sequences represent the reference genomic material to be used in taxonomic studies involving this family and its members. Additionally, we have shown how the combination of high-quality, short, Illumina sequence reads with long Oxford Nanopore sequence reads is able to generate a complete genome sequence, identical to the one obtained with only PacBio sequencing.
Data availability
The complete genome sequence of S. pyogenes NCTC 8198T is publicly available in DNA DataBank of Japan (DDBJ)/European Nucleotide Archive (ENA)/GenBank under the accession number LN831034. The version described in this article is LN831034.1. The PacBio sequence reads of S. pyogenes NCTC 8198T are available in the Sequence Read Archive (SRA)98 under the accession numbers ERR550482 and ERR550487. The complete genome sequence of S. pyogenes CCUG 4207T is publicly available in DDBJ/ENA/GenBank under the accession number CP028841. The version described in this article is CP028841.1. The Illumina and Oxford Nanopore raw sequence reads of S. pyogenes CCUG 4207T are available in the SRA under the accession numbers SRR8631872 and SRR10092043, respectively. The Illumina and Oxford Nanopore sequence reads sets used in the assembly are available in the SRA under the accession numbers SRR10092042 and SRR8608127.
References
Lancefield, R. C. A serological differentiation of human and other groups of hemolytic streptococci. J. Exp. Med. 57, 571–595 (1933).
Ralph, A. P. & Carapetis, J. R. Group a streptococcal diseases and their global burden. Curr. Top. Microbiol. Immunol. 368, 1–27. https://doi.org/10.1007/82_2012_280 (2013).
Carapetis, J. R., Steer, A. C., Mulholland, E. K. & Weber, M. The global burden of group A streptococcal diseases. Lancet Infect. Dis. 5, 685–694. https://doi.org/10.1016/S1473-3099(05)70267-X (2005).
Barnett, T. C., Bowen, A. C. & Carapetis, J. R. The fall and rise of Group A Streptococcus diseases. Epidemiol. Infect. https://doi.org/10.1017/S0950268818002285 (2019).
Rosenbach, F. J. Mikro-organismen bei den Wund-Infections-Krankheiten des Menschen (J.F. Bergmann, München, 1884).
Loman, N. J. & Pallen, M. J. Twenty years of bacterial genome sequencing. Nat. Rev. Microbiol. 13, 787–794. https://doi.org/10.1038/nrmicro3565 (2015).
Goodwin, S., McPherson, J. D. & McCombie, W. R. Coming of age: Ten years of next-generation sequencing technologies. Nat. Rev. Genet. 17, 333–351. https://doi.org/10.1038/nrg.2016.49 (2016).
Payne, A., Holmes, N., Rakyan, V. & Loose, M. Whale watching with BulkVis: A graphical viewer for Oxford Nanopore bulk FAST5 files. bioRxiv https://doi.org/10.1101/312256 (2018).
Schmid, M. et al. Pushing the limits of de novo genome assembly for complex prokaryotic genomes harboring very long, near identical repeats. Nucleic Acids Res. https://doi.org/10.1093/nar/gky726 (2018).
Greninger, A. L. et al. Rapid metagenomic identification of viral pathogens in clinical samples by real-time nanopore sequencing analysis. Genome Med. 7, 99. https://doi.org/10.1186/s13073-015-0220-9 (2015).
Laver, T. et al. Assessing the performance of the Oxford Nanopore technologies MinION. Biomol. Detect. Quant. 3, 1–8. https://doi.org/10.1016/j.bdq.2015.02.001 (2015).
Goodwin, S. et al. Oxford Nanopore sequencing, hybrid error correction, and de novo assembly of a eukaryotic genome. Genome Res. 25, 1750–1756. https://doi.org/10.1101/gr.191395.115 (2015).
Salvà-Serra, F. et al. A protocol for extraction and purification of high-quality and quantity bacterial DNA applicable for genome sequencing: A modified version of the Marmur procedure. Protoc. Exchange https://doi.org/10.1038/protex.2018.084 (2018).
Marmur, J. A procedure for the isolation of deoxyribonucleic acid from micro-organisms. J. Mol. Biol. 3, 208. https://doi.org/10.1016/S0022-2836(61)80047-8 (1961).
De Coster, W., D’Hert, S., Schultz, D. T., Cruts, M. & Van Broeckhoven, C. NanoPack: Visualizing and processing long read sequencing data. Bioinformatics 34, 2666–2669. https://doi.org/10.1093/bioinformatics/bty149 (2018).
Chin, C. S. et al. Nonhybrid, finished microbial genome assemblies from long-read SMRT sequencing data. Nat. Methods 10, 563–569. https://doi.org/10.1038/nmeth.2474 (2013).
Gurevich, A., Saveliev, V., Vyahhi, N. & Tesler, G. QUAST: Quality assessment tool for genome assemblies. Bioinformatics 29, 1072–1075. https://doi.org/10.1093/bioinformatics/btt086 (2013).
Sickle: A sliding-window, adaptive, quality-based trimming tool for FastQ files v. 1.33 (2011).
Bankevich, A. et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477. https://doi.org/10.1089/cmb.2012.0021 (2012).
Antipov, D., Korobeynikov, A., McLean, J. S. & Pevzner, P. A. hybridSPAdes: An algorithm for hybrid assembly of short and long reads. Bioinformatics 32, 1009–1015. https://doi.org/10.1093/bioinformatics/btv688 (2016).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760. https://doi.org/10.1093/bioinformatics/btp324 (2009).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410. https://doi.org/10.1016/S0022-2836(05)80360-2 (1990).
Seemann, T. Prokka: Rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069. https://doi.org/10.1093/bioinformatics/btu153 (2014).
Harrison, P. W. et al. The European nucleotide archive in 2018. Nucleic Acids Res. 47, D84–D88. https://doi.org/10.1093/nar/gky1078 (2019).
Tatusova, T. et al. NCBI prokaryotic genome annotation pipeline. Nucleic Acids Res. 44, 6614–6624. https://doi.org/10.1093/nar/gkw569 (2016).
O’Leary, N. A. et al. Reference sequence (RefSeq) database at NCBI: Current status, taxonomic expansion, and functional annotation. Nucleic Acids Res. 44, D733-745. https://doi.org/10.1093/nar/gkv1189 (2016).
Sayers, E. W. et al. GenBank. Nucleic Acids Res. 47, D94–D99. https://doi.org/10.1093/nar/gky989 (2019).
Petkau, A., Stuart-Edwards, M., Stothard, P. & Van Domselaar, G. Interactive microbial genome visualization with GView. Bioinformatics 26, 3125–3126. https://doi.org/10.1093/bioinformatics/btq588 (2010).
Arndt, D. et al. PHASTER: A better, faster version of the PHAST phage search tool. Nucleic Acids Res. 44, W16-21. https://doi.org/10.1093/nar/gkw387 (2016).
Grissa, I., Vergnaud, G. & Pourcel, C. CRISPRFinder: A web tool to identify clustered regularly interspaced short palindromic repeats. Nucleic Acids Res. 35, W52-57. https://doi.org/10.1093/nar/gkm360 (2007).
Lange, S. J., Alkhnbashi, O. S., Rose, D., Will, S. & Backofen, R. CRISPRmap: An automated classification of repeat conservation in prokaryotic adaptive immune systems. Nucleic Acids Res. 41, 8034–8044. https://doi.org/10.1093/nar/gkt606 (2013).
Alkhnbashi, O. S. et al. CRISPRstrand: Predicting repeat orientations to determine the crRNA-encoding strand at CRISPR loci. Bioinformatics 30, 489–496. https://doi.org/10.1093/bioinformatics/btu459 (2014).
Zhang, Q. & Ye, Y. Not all predicted CRISPR-Cas systems are equal: Isolated Cas genes and classes of CRISPR like elements. BMC Bioinform. 18, 92. https://doi.org/10.1186/s12859-017-1512-4 (2017).
Liu, B., Zheng, D., Jin, Q., Chen, L. & Yang, J. VFDB 2019: A comparative pathogenomic platform with an interactive web interface. Nucleic Acids Res. 47, D687–D692. https://doi.org/10.1093/nar/gky1080 (2019).
Jia, B. et al. CARD 2017: Expansion and model-centric curation of the comprehensive antibiotic resistance database. Nucleic Acids Res. 45, D566–D573. https://doi.org/10.1093/nar/gkw1004 (2017).
Makarova, K. S. et al. An updated evolutionary classification of CRISPR-Cas systems. Nat. Rev. Microbiol. 13, 722–736. https://doi.org/10.1038/nrmicro3569 (2015).
Metzgar, D. & Zampolli, A. The M protein of group A Streptococcus is a key virulence factor and a clinically relevant strain identification marker. Virulence 2, 402–412. https://doi.org/10.4161/viru.2.5.16342 (2011).
Cywes, C. & Wessels, M. R. Group A Streptococcus tissue invasion by CD44-mediated cell signalling. Nature 414, 648–652. https://doi.org/10.1038/414648a (2001).
Wessels, M. R., Moses, A. E., Goldberg, J. B. & DiCesare, T. J. Hyaluronic acid capsule is a virulence factor for mucoid group A streptococci. Proc. Natl. Acad. Sci. U.S.A. 88, 8317–8321. https://doi.org/10.1073/pnas.88.19.8317 (1991).
Fernie-King, B. A. et al. Streptococcal inhibitor of complement (SIC) inhibits the membrane attack complex by preventing uptake of C567 onto cell membranes. Immunology 103, 390–398. https://doi.org/10.1046/j.1365-2567.2001.01249.x (2001).
Sriskandan, S., Unnikrishnan, M., Krausz, T. & Cohen, J. Mitogenic factor (MF) is the major DNase of serotype M89 Streptococcus pyogenes. Microbiology 146(Pt 11), 2785–2792. https://doi.org/10.1099/00221287-146-11-2785 (2000).
Iwasaki, M., Igarashi, H. & Yutsudo, T. Mitogenic factor secreted by Streptococcus pyogenes is a heat-stable nuclease requiring His122 for activity. Microbiology 143, 2449–2455. https://doi.org/10.1099/00221287-143-7-2449 (1997).
Sumby, P. et al. Extracellular deoxyribonuclease made by group A Streptococcus assists pathogenesis by enhancing evasion of the innate immune response. Proc. Natl. Acad. Sci. U.S.A. 102, 1679–1684. https://doi.org/10.1073/pnas.0406641102 (2005).
Starr, C. R. & Engleberg, N. C. Role of hyaluronidase in subcutaneous spread and growth of group A Streptococcus. Infect. Immunol. 74, 40–48. https://doi.org/10.1128/IAI.74.1.40-48.2006 (2006).
Hynes, W. Virulence factors of the group A streptococci and genes that regulate their expression. Front. Biosci. 9, 3399–3433. https://doi.org/10.2741/1491 (2004).
Smith, N. L. et al. Structure of a group A streptococcal phage-encoded virulence factor reveals a catalytically active triple-stranded β-helix. Proc. Natl. Acad. Sci. U.S.A. 102, 17652–17657. https://doi.org/10.1073/pnas.0504782102 (2005).
von Pawel-Rammingen, U., Johansson, B. P. & Björck, L. IdeS, a novel streptococcal cysteine proteinase with unique specificity for immunoglobulin G. EMBO J. 21, 1607–1615. https://doi.org/10.1093/emboj/21.7.1607 (2002).
Cleary, P. P., Prahbu, U., Dale, J. B., Wexler, D. E. & Handley, J. Streptococcal C5a peptidase is a highly specific endopeptidase. Infect Immun 60, 5219–5223 (1992).
Guo, R. F. & Ward, P. A. Role of C5a in inflammatory responses. Annu. Rev. Immunol. 23, 821–852. https://doi.org/10.1146/annurev.immunol.23.021704.115835 (2005).
Ringdahl, U. et al. Molecular co-operation between protein PAM and streptokinase for plasmin acquisition by Streptococcus pyogenes. J. Biol. Chem. 273, 6424–6430. https://doi.org/10.1074/jbc.273.11.6424 (1998).
Manetti, A. G. et al. Streptococcus pyogenes pili promote pharyngeal cell adhesion and biofilm formation. Mol. Microbiol. 64, 968–983. https://doi.org/10.1111/j.1365-2958.2007.05704.x (2007).
Courtney, H. S., Li, Y., Dale, J. B. & Hasty, D. L. Cloning, sequencing, and expression of a fibronectin/fibrinogen-binding protein from group A streptococci. Infect. Immunol. 62, 3937–3946 (1994).
Courtney, H. S., Dale, J. B. & Hasty, D. I. Differential effects of the streptococcal fibronectin-binding protein, FBP54, on adhesion of group A streptococci to human buccal cells and HEp-2 tissue culture cells. Infect. Immunol. 64, 2415–2419 (1996).
Terao, Y., Kawabata, S., Kunitomo, E., Nakagawa, I. & Hamada, S. Novel laminin-binding protein of Streptococcus pyogenes, Lbp, is involved in adhesion to epithelial cells. Infect. Immunol. 70, 993–997 (2002).
Linke, C., Caradoc-Davies, T. T., Young, P. G., Proft, T. & Baker, E. N. The laminin-binding protein Lbp from Streptococcus pyogenes is a zinc receptor. J. Bacteriol. 191, 5814–5823. https://doi.org/10.1128/JB.00485-09 (2009).
Rasmussen, M., Müller, H. P. & Björck, L. Protein GRAB of Streptococcus pyogenes regulates proteolysis at the bacterial surface by binding α2-macroglobulin. J. Biol. Chem. 274, 15336–15344. https://doi.org/10.1074/jbc.274.22.15336 (1999).
Garvey, M. I., Baylay, A. J., Wong, R. L. & Piddock, L. J. Overexpression of patA and patB, which encode ABC transporters, is associated with fluoroquinolone resistance in clinical isolates of Streptococcus pneumoniae. Antimicrob. Agents Chemother. 55, 190–196. https://doi.org/10.1128/AAC.00672-10 (2011).
Griffith, F. Types of haemolytic streptococci in relation to scarlet fever. J. Hyg. (Lond.) 25, 385–397 (1926).
Ferretti, J. J. et al. Complete genome sequence of an M1 strain of Streptococcus pyogenes. Proc. Natl. Acad. Sci. U.S.A. 98, 4658–4663. https://doi.org/10.1073/pnas.071559398 (2001).
Wick, R. R., Judd, L. M., Gorrie, C. L. & Holt, K. E. Completing bacterial genome assemblies with multiplex MinION sequencing. Microb. Genom. 3, e000132. https://doi.org/10.1099/mgen.0.000132 (2017).
De Maio, N. et al. Comparison of long-read sequencing technologies in the hybrid assembly of complex bacterial genomes. Microb. Genom. https://doi.org/10.1099/mgen.0.000294 (2019).
Goldstein, S., Beka, L., Graf, J. & Klassen, J. L. Evaluation of strategies for the assembly of diverse bacterial genomes using MinION long-read sequencing. BMC Genom. 20, 23. https://doi.org/10.1186/s12864-018-5381-7 (2019).
Koren, S. et al. Reducing assembly complexity of microbial genomes with single-molecule sequencing. Genome Biol. 14, R101. https://doi.org/10.1186/gb-2013-14-9-r101 (2013).
Lu, H., Giordano, F. & Ning, Z. Oxford Nanopore MinION sequencing and genome assembly. Genomics. Proteomics Bioinform. 14, 265–279. https://doi.org/10.1016/j.gpb.2016.05.004 (2016).
Wick, R. R., Judd, L. M., Gorrie, C. L. & Holt, K. E. Unicycler: Resolving bacterial genome assemblies from short and long sequencing reads. PLoS Comput. Biol. 13, e1005595. https://doi.org/10.1371/journal.pcbi.1005595 (2017).
Zimin, A. V. et al. Hybrid assembly of the large and highly repetitive genome of Aegilops tauschii, a progenitor of bread wheat, with the MaSuRCA mega-reads algorithm. Genome Res. 27, 787–792. https://doi.org/10.1101/gr.213405.116 (2017).
Koren, S. et al. Canu: Scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 27, 722–736. https://doi.org/10.1101/gr.215087.116 (2017).
Walker, B. J. et al. Pilon: An integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS ONE 9, e112963. https://doi.org/10.1371/journal.pone.0112963 (2014).
Vaser, R., Sović, I., Nagarajan, N. & Šikić, M. Fast and accurate de novo genome assembly from long uncorrected reads. Genome Res. 27, 737–746. https://doi.org/10.1101/gr.214270.116 (2017).
Somerville, G. A. et al. In vitro serial passage of Staphylococcus aureus: Changes in physiology, virulence factor production, and agr nucleotide sequence. J. Bacteriol. 184, 1430–1437. https://doi.org/10.1128/jb.184.5.1430-1437.2002 (2002).
Eberhard, T. H., Sledjeski, D. D. & Boyle, M. D. Mouse skin passage of a Streptococcus pyogenes Tn917 mutant of sagA/pel restores virulence, beta-hemolysis and sagA/pel expression without altering the position or sequence of the transposon. BMC Microbiol. 1, 33 (2001).
Rezcallah, M. S., Boyle, M. D. & Sledjeski, D. D. Mouse skin passage of Streptococcus pyogenes results in increased streptokinase expression and activity. Microbiology 150, 365–371. https://doi.org/10.1099/mic.0.26826-0 (2004).
Konstantinidis, K. T. & Tiedje, J. M. Towards a genome-based taxonomy for prokaryotes. J. Bacteriol. 187, 6258–6264. https://doi.org/10.1128/JB.187.18.6258-6264.2005 (2005).
Gomila, M., Peña, A., Mulet, M., Lalucat, J. & García-Valdés, E. Phylogenomics and systematics in Pseudomonas. Front. Microbiol. 6, 214. https://doi.org/10.3389/fmicb.2015.00214 (2015).
Jensen, A., Scholz, C. F. & Kilian, M. Re-evaluation of the taxonomy of the Mitis group of the genus Streptococcus based on whole genome phylogenetic analyses, and proposed reclassification of Streptococcus dentisani as Streptococcus oralis subsp. dentisani comb. nov., Streptococcus tigurinus as Streptococcus oralis subsp. tigurinus comb. nov., and Streptococcus oligofermentans as a later synonym of Streptococcus cristatus. Int. J. Syst. Evol. Microbiol. 66, 4803–4820. https://doi.org/10.1099/ijsem.0.001433 (2016).
Goris, J. et al. DNA-DNA hybridization values and their relationship to whole-genome sequence similarities. Int. J. Syst. Evol. Microbiol. 57, 81–91. https://doi.org/10.1099/ijs.0.64483-0 (2007).
Richter, M. & Rosselló-Móra, R. Shifting the genomic gold standard for the prokaryotic species definition. Proc. Natl. Acad. Sci. U.S.A. 106, 19126–19131. https://doi.org/10.1073/pnas.0906412106 (2009).
Richter, M., Rosselló-Móra, R., Oliver Glöckner, F. & Peplies, J. JSpeciesWS: A web server for prokaryotic species circumscription based on pairwise genome comparison. Bioinformatics 32, 929–931. https://doi.org/10.1093/bioinformatics/btv681 (2016).
Meier-Kolthoff, J. P., Auch, A. F., Klenk, H.-P. & Göker, M. Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinform. 14, 60. https://doi.org/10.1186/1471-2105-14-60 (2013).
Meier-Kolthoff, J. P. & Göker, M. TYGS is an automated high-throughput platform for state-of-the-art genome-based taxonomy. Nat. Commun. 10, 2182. https://doi.org/10.1038/s41467-019-10210-3 (2019).
Ha, S. M. et al. Application of the whole genome-based bacterial identification system, TrueBac ID, using clinical isolates that were not identified with three matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS) systems. Ann. Lab. Med. 39, 530–536. https://doi.org/10.3343/alm.2019.39.6.530 (2019).
Beaz-Hidalgo, R., Hossain, M. J., Liles, M. R. & Figueras, M. J. Strategies to avoid wrongly labelled genomes using as example the detected wrong taxonomic affiliation for Aeromonas genomes in the GenBank database. PLoS ONE 10, e0115813. https://doi.org/10.1371/journal.pone.0115813 (2015).
Wu, L. et al. The global catalogue of microorganisms 10K type strain sequencing project: Closing the genomic gaps for the validly published prokaryotic and fungi species. GigaScience https://doi.org/10.1093/gigascience/giy026 (2018).
Salvà-Serra, F. et al. Beware of false “type strain” genome sequences. Microbiol. Resour. Announc. https://doi.org/10.1128/MRA.00369-19 (2019).
Federhen, S. et al. Meeting report: GenBank microbial genomic taxonomy workshop (12–13 May, 2015). Stand. Genom. Sci. 11, 15. https://doi.org/10.1186/s40793-016-0134-1 (2016).
Mukherjee, S. et al. 1,003 reference genomes of bacterial and archaeal isolates expand coverage of the tree of life. Nat. Biotechnol. 35, 676–683. https://doi.org/10.1038/nbt.3886 (2017).
Klenk, H. P. & Göker, M. En route to a genome-based classification of Archaea and Bacteria?. Syst. Appl. Microbiol. 33, 175–182. https://doi.org/10.1016/j.syapm.2010.03.003 (2010).
Whitman, W. B. Genome sequences as the type material for taxonomic descriptions of prokaryotes. Syst. Appl. Microbiol. 38, 217–222. https://doi.org/10.1016/j.syapm.2015.02.003 (2015).
Beres, S. B. et al. Genome sequence of a serotype M3 strain of group A Streptococcus: phage-encoded toxins, the high-virulence phenotype, and clone emergence. Proc. Natl. Acad. Sci. U.S.A. 99, 10078–10083. https://doi.org/10.1073/pnas.152298499 (2002).
Smoot, J. C. et al. Genome sequence and comparative microarray analysis of serotype M18 group A Streptococcus strains associated with acute rheumatic fever outbreaks. Proc. Natl. Acad. Sci. U.S.A. 99, 4668–4673. https://doi.org/10.1073/pnas.062526099 (2002).
Nasser, W. et al. Evolutionary pathway to increased virulence and epidemic group A Streptococcus disease derived from 3,615 genome sequences. Proc. Natl. Acad. Sci. U.S.A. 111, E1768-1776. https://doi.org/10.1073/pnas.1403138111 (2014).
Grissa, I., Vergnaud, G. & Pourcel, C. The CRISPRdb database and tools to display CRISPRs and to generate dictionaries of spacers and repeats. BMC Bioinform. 8, 172. https://doi.org/10.1186/1471-2105-8-172 (2007).
Nozawa, T. et al. CRISPR inhibition of prophage acquisition in Streptococcus pyogenes. PLoS ONE 6, e19543. https://doi.org/10.1371/journal.pone.0019543 (2011).
Brouns, S. J. et al. Small CRISPR RNAs guide antiviral defense in prokaryotes. Science 321, 960–964. https://doi.org/10.1126/science.1159689 (2008).
Sjögren, J. et al. EndoS2 is a unique and conserved enzyme of serotype M49 group A Streptococcus that hydrolyses N-linked glycans on IgG and α1-acid glycoprotein. Biochem. J. 455, 107–118. https://doi.org/10.1042/BJ20130126 (2013).
Dick, G. F. & Dick, G. H. The etiology of scarlet fever. JAMA 82, 301–302. https://doi.org/10.1001/jama.1924.02650300047013 (1924).
Spellerberg, B. & Brandt, C. in Streptococcus pyogenes: basic biology to clinical manifestations (eds Ferretti, J. J., Stevens, D. L. & Fischetti, V. A.) (2016).
Leinonen, R., Sugawara, H., Shumway, M. & International Nucleotide Sequence Database Consortium. The sequence read archive. Nucleic Acids Res. 39, 19–21. https://doi.org/10.1093/nar/gkq1019 (2011).
Acknowledgements
Open access funding provided by University of Gothenburg. This work was supported by the European Commission 7th Framework Programme: TAILORED-Treatment (Project No. 602860), by the Swedish Västra Götaland regional funding (Projects No. ALFGBG-437221 and ALFGBG-720761), the Swedish Västra Götaland FoU Grant No. VGFOUREG-665141, Laboratoriemedicin FoU (Project No. 51060-6268) and by the Wellcome Trust (Project No. 101503/Z/13/Z). DJL, RK and ERBM acknowledge the support of the Joint Programme Initiative—Anti-Microbial Resistance (JPIAMR) (Vetenskapsrådet Project No. 2016-06504). The Culture Collection University of Gothenburg (CCUG) was supported by the Department of Clinical Microbiology, Sahlgrenska University Hospital, Gothenburg, Sweden. FS-S was supported by a stipend for Basic and Advanced Research from the CCUG, through the Institute for Biomedicine, Sahlgrenska Academy, University of Gothenburg. FS-S, DJ-L, HEJ, LG-S, RK, NK and ERBM acknowledge the support from the Centre for Antibiotic Resistance Research (CARe) at the University of Gothenburg. Antonio Busquets was supported by a postdoctoral contract from the University of the Balearic Islands. The authors acknowledge Gemma Langridge, Julian Parkhill and Nick Grayson, at the Wellcome Trust Sanger Institute (Hinxton, United Kingdom), for the PacBio assembly and thank them and Timur Tunovic for valuable comments and suggestions.
Author information
Authors and Affiliations
Contributions
Conceptualization—F.S.-S., D.J.L., L.G.S., R.K., J.E.R., S.A. and E.R.B.M.; Methodology—F.S.-S., D.J.L., H.E.J., A.B., M.G., M.A.F. and S.A.; Validation—F.S.-S., D.J.L. and A.B.F.; Original draft preparation—F.S.-S.; Review & Editing—F.S.-S., D.J.L., H.E.J., L.G.S., R.K., A.B., A.B.F., J.E.R., M.A.F., S.A. and E.R.B.M.; Supervision—H.E.J., A.B.F., J.E.R. and E.R.B.M.; Project administration—R.K., J.E.R., S.A. and E.R.B.M.; Funding acquisition—R.K., J.E.R., S.A. and E.R.B.M.
Corresponding author
Ethics declarations
Competing interests
Author RK is affiliated with a company, Nanoxis Consulting AB. The company did not have influence on the conception, elaboration and decision to submit the present study for publication. The other authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Salvà-Serra, F., Jaén-Luchoro, D., Jakobsson, H.E. et al. Complete genome sequences of Streptococcus pyogenes type strain reveal 100%-match between PacBio-solo and Illumina-Oxford Nanopore hybrid assemblies. Sci Rep 10, 11656 (2020). https://doi.org/10.1038/s41598-020-68249-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-68249-y
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.