Abstract
Alterations to the gut microbiota can cause an amplification of the inflammatory response to intestinal pathogens. We assessed the effect of Bacteroides thetaiotaomicron and Lactobacillus johnsonii on the elimination of Candida species and whether restoration of these two anaerobic bacteria could attenuate the development of colitis in mice. In this study, L. johnsonii and B. thetaiotaomicron interacted directly with Candida species and induced a degradation of the fungal cell wall, mediated via chitinase-like and mannosidase-like activities, which promoted the inhibition of Candida species growth. In the DSS-induced colitis model, oral administration of L. johnsonii and B. thetaiotaomicron to mice reduced the overgrowth of Escherichia coli, Enterococcus faecalis and Candida glabrata populations and resulted in a significant reduction in inflammatory parameters. L. johnsonii and B. thetaiotaomicron decreased pro-inflammatory mediators and enhanced the anti-inflammatory cytokine response with high TLR9 expression and chitinase-like protein-1 activation, which promoted the elimination of C. glabrata from the gut. Overall, these findings provide evidence that L. johnsonii and B. thetaiotaomicron decrease the development of colitis mediated by TLR9 and promote the elimination of C. glabrata from the gut via chitinase-like and mannosidase-like activities.
Similar content being viewed by others
Introduction
The gut microbiota plays a crucial role in protecting the host from colonization/infection by pathogens and in stimulating/modulating the immune system1,2. Alterations to the gut microbiota can occur via changes in the diversity of the bacterial community or by microbiota-host interactions and can be directly correlated with several diseases, in particular inflammatory bowel disease (IBD), which includes Crohn’s disease (CD) and ulcerative colitis (UC)3. Ulceration from CD can occur anywhere in the gastrointestinal tract while the hallmark of UC is ulceration beginning in the rectum and limited to the colon4. Although the pathogenesis of IBD is unknown, different studies have reported that changes in gut microbiota biodiversity, genetic susceptibility, environmental factors and the immune system are all involved5,6,7. Numerous studies have shown that the quality and composition of the microbiota are altered in IBD patients, with a reduction of Firmicutes and Bacteroidetes and an increase in Proteobacteria and Enterobacteria8. These alterations to the gut microbiota can cause an amplification of the inflammatory response to intestinal pathogens and trigger a range of mechanisms including increased epithelial permeability and decreased luminal IgA concentrations9. Additionally, ineffective bacterial clearance leads to excessive Toll-like receptor (TLR) stimulation, secretion of pro-inflammatory cytokines and activation of the immune response. In a model of dextran sulphate sodium (DSS)-induced colitis, overgrowth of Escherichia coli and Enterococcus faecalis populations, which are representative of the Enterobacteria and Proteobacteria phyla respectively, are associated with high levels of expression of TLR and pro-inflammatory cytokines10. Furthermore, increased E. coli or E. faecalis populations are also associated with clinically active CD11,12,13. In contrast, Bacteroides thetaiotaomicron and Lactobacillus johnsonii, which are representative of Bacteroidetes and Firmicutes respectively, are reduced during the devolvement of colitis and this reduction is more pronounced for L. johnsonii in colitic mice challenged with the opportunistic yeast Candida glabrata14,15.
Like Candida albicans, C. glabrata is an endosaprophytic yeast that can cause invasive fungal infection (IFI)16. Systemic C. glabrata infections are associated with higher mortality than C. albicans infections16,17. C. glabrata interacts with the host at the level of the fungal cell wall, which consists mainly of polysaccharides associated with proteins and lipids18. Fungal polysaccharides are released into the blood during infection and their detection enables the early diagnosis of IFI19. Clinical and experimental studies have shown that infection by Candida species can generate anti-glycan antibodies known as ASCA (anti-Saccharomyces cerevisiae mannan antibodies), ACCA (anti-chitobioside carbohydrate antibodies) and ALCA (anti-laminaribioside carbohydrate antibodies), which are described as serological markers of CD, suggesting a link between CD gut dysbiosis and endogenous opportunistic yeast species20,21. The DSS-induced colitis model promoted an abundance of C. glabrata which in turn increased the severity of inflammation leading to intense bleeding and diarrhoea, colonic epithelial damage and inflammatory cell infiltrates18.
Given that B. thetaiotaomicron and L. johnsonii populations are highly affected during the development of colitis and overgrowth of C. glabrata, we assessed whether restoration of these two anaerobic bacteria can attenuate the development of colitis and C. glabrata growth14,15. We then measured inflammatory parameters, changes in aerobic bacteria populations and modulation of receptor and cytokine expression. In addition to the DSS-colitis model, the direct effect of these two bacteria on C. glabrata growth/viability were analysed in vitro.
Our results show that L. johnsonii and B. thetaiotaomicron interact directly with Candida species and induce degradation of the fungal cell wall, mediated via chitinase-like and mannosidase-like activities, which promotes the inhibition of Candida species growth. In the DSS-induced colitis model, oral administration of L. johnsonii and B. thetaiotaomicron to mice restored the imbalance between aerobic and anaerobic populations and resulted in a significant reduction in inflammatory parameters, revealed by a decrease in pro-inflammatory mediators, and enhanced the anti-inflammatory cytokine response with high TLR9 expression and chitinase-like protein-1 activation, which promoted C. glabrata elimination from the gut.
Results
Impact of B. thetaiotaomicron and L. johnsonii on C. glabrata and C. albicans growth
Based on our previous observations that intestinal inflammation and Candida species promoted a significant decrease in anaerobic bacteria in a murine model of DSS-induced colitis, in particular B. thetaiotaomicron and L. johnsonii, we first assessed whether B. thetaiotaomicron + L. johnsonii could inhibit the growth of C. albicans and C. glabrata, which are the two most frequent causes of human yeast infections14,15. To determine the viability of C. albicans in real-time, a bioluminescent C. albicans strain was incubated with either B. thetaiotaomicron or L. johnsonii or both for different periods of time and at different concentrations. We observed a significant decrease in bioluminescence of C. albicans in the presence of L. johnsonii or both bacterial strains starting a few minutes after co-incubation of the bacteria with yeasts (at a ratio of 1:1 or 1:5). The immediate effect of these bacteria on C. albicans was not observed when the ratio was 5:1. Furthermore, C. albicans bioluminescence decreased significantly after 2 h co-incubation with the two bacterial strains at all ratios when compared to C. albicans unchallenged with bacteria (Fig. 1A–D). The effect of these bacteria on C. glabrata growth was assessed using culture methods based on agar plates. A significant decrease in C. glabrata colonies was observed in the presence of these bacteria after 2 h co-incubation (Fig. 1E). Confocal microscopy revealed that B. thetaiotaomicron and L. johnsonii interacted closely with the C. glabrata cell wall (Fig. 2).
Antifungal effect of B. thetaiotaomicron and L. johnsonii on C. albicans and C. glabrata. (A–C) A bioluminescent C. albicans strain was treated with either B. thetaiotaomicron and/or L. johnsonii at a yeast-bacteria ratio of 1:1, 5:1 and 1:5, respectively, and monitored at 30 and 120 min. CTL corresponds to PBS and substrate only. Ca corresponds to C. albicans alone without bacteria. CaBt, CaLj and CaBtLj correspond to co-incubation of C. albicans with B. thetaiotaomicron and/or L. johnsonii. The results were obtained from three independent experiments. (D) Visualization of the bioluminescence of C. albicans in real-time. Bioluminescence of C. albicans incubated with B. thetaiotaomicron and/or L. johnsonii at a ratio of 1:1 and monitored at 30, 60 and 120 min. Line CTL: PBS without yeasts. Line Ca: C. albicans alone. Line 1 corresponds to C. albicans treated with L. johnsonii at a ratio of 1:1. Line 2 represents C. albicans treated with both B. thetaiotaomicron and L. johnsonii at a bacteria-yeast ratio of 1:1. (E) Viability of C. glabrata in the presence of B. thetaiotaomicron and/or L. johnsonii. C. glabrata was challenged with B. thetaiotaomicron and/or L. johnsonii at a ratio of 1:1. The results were obtained from three independent experiments.
Confocal microscopy of the interaction between C. glabrata, B. thetaiotaomicron and L. johnsonii. Controls (a) C. glabrata alone labelled with Con A, (b) B. thetaiotaomicron alone labelled with calcein, (c) L. johnsonii alone labelled with calcein. (d) Co-incubation of C. glabrata with (e) B. thetaiotaomicron: (f) merged images. (g) Co-incubation of C. glabrata with (h) L. johnsonii: (i) merged images. (j) Co-incubation of C. glabrata with (k) both B. thetaiotaomicron and L. johnsonii: (l) merged images. The scale bars correspond to 5 μm.
Antifungal activity of bacterial enzymes against C. glabrata
After monitoring in real-time how B. thetaiotaomicron and L. johnsonii were able to interact directly with Candida and inhibit fungal growth, we then assessed whether B. thetaiotaomicron and L. johnsonii were able to induce cell wall degradation of C. glabrata, α-mannan and chitin (Fig. 3). In contrast to L. johnsonii which did not show any mannosidase-like activity against C. glabrata, B. thetaiotaomicron was able to induce degradation of C. glabrata α-mannan in the cell wall after 2 h co-incubation (Fig. 3A–D). To confirm that mannans are degraded by B. thetaiotaomicron, C. glabrata mannan was extracted and co-incubated with B. thetaiotaomicron. Mannans derived from C. glabrata alone form a smear in SDS-acrylamide gels as do all other heavily glycosylated yeast glycoproteins22,23 (Supplementary data). This mannan smear was no longer visible after incubation of B. thetaiotaomicron with mannans. These data correlate closely with those showing degradation of cell wall mannan after co-incubation of C. glabrata cells with B. thetaiotaomicron and support the idea that B. thetaiotaomicron is able to induce degradation of C. glabrata cell wall mannan and exert a mannosidase-like activity that promotes C. glabrata elimination. In terms of chitin, L. johnsonii exhibited chitinase-like activity, which was correlated with the elimination of C. glabrata. In contrast, B. thetaiotaomicron did not show any chitinase-like activity against C. glabrata (Fig. 3F).
Antifungal activities of B. thetaiotaomicron and L. johnsonii against C. glabrata. (A and B) Determination of C. glabrata cell wall α-mannan levels after co-incubation of yeasts + bacteria at a ratio of 1:1. (C and D) Western blot analysis of C. glabrata cell wall α-mannan labelled with either GNL or ConA, respectively. Line 1: B. thetaiotaomicron extract. Line 2: C. glabrata extract. Line 3: extract of both C. glabrata and B. thetaiotaomicron incubated for 2 h. Line 4: extract of both C. glabrata and B. thetaiotaomicron incubated for 48 h. Different exposures of the blots were performed (Supplementary data). (F) Determination of chitinase activity of L. johnsonii and B. thetaiotaomicron in the presence or absence of C. glabrata.
Impact of B. thetaiotaomicron and L. johnsonii on intestinal inflammation, C. glabrata elimination and modulation of bacteria populations
In order to assess whether these two bacteria exert antifungal and/or inflammatory properties, the effect of B. thetaiotaomicron and L. johnsonii on intestinal inflammation and C. glabrata elimination was assessed in the DSS-induced colitis model. Mice were administered a single oral dose of C. glabrata and were exposed to DSS treatment for 2 weeks in order to induce acute colitis. These two bacteria were then administered orally to mice for 5 days after C. glabrata challenge.
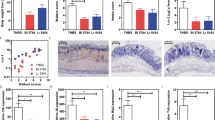
In terms of inflammatory parameters, control mice (CTL), BtLj (B. thetaiotaomicron + L. johnsonii) and Cg (C. glabrata) groups, did not show any inflammatory signs. In the presence of DSS, the D (DSS) and DCg (DSS + C. glabrata) groups had a 15% loss in body weight starting from day 12 when compared to that of control mice (Fig. 4A). This decrease in body weight was inversely correlated with the increase in clinical score in the D and DCg groups (Fig. 4A,B). Although inflammatory parameters increased during the development of colitis and C. glabrata overgrowth, treatment with the two bacteria significantly decreased the clinical signs of inflammation, body weight and clinical score for inflammation in D or DCg mice.
Determination of inflammatory parameters in the DSS-induced colitis model. (A) Body weight. Control groups correspond to CTL (water), Cg (C. glabrata) and BtLj (B. thetaiotaomicron + L. johnsonii). Experimental groups correspond to D (DSS), DBtLj (DSS + B. thetaiotaomicron + L. johnsonii) and DBtLjCg (DSS + B. thetaiotaomicron + L. johnsonii + C. glabrata). Data are the mean ± SD of eight mice per group from two independent experiments. * p < 0.05, D and DCg groups vs. DBtLj and DBtLjCg. (B) Clinical score. * p < 0.0001, D and DCg groups vs. DBtLj and DBtLjCg. (C) Histological score. p < 0.01, D and DCg groups vs. DBtLj and DBtLjCg. (D) Histological analysis of colon sections from DSS-induced colitis. Panels (a), (c) and (e) correspond to colon sections from mice receiving water (control), C. glabrata alone and BtLj, respectively. Panel (g) corresponds to colon sections from mice receiving DSS. Panel (i) corresponds to colon sections from mice receiving DSS + BtLj. Panel (K) represents colon sections from mice receiving C. glabrata + DSS. Panel (m) represents colon sections from mice receiving C. glabrata + BtLj + DSS. Colon sections from either D or DCg show tissue destruction (θ), important inflammatory cell infiltrates (δ) and oedema in the mucosa and submucosa of colon wall structures (φ). Scale bars represent 50 µm (a, c, e, g, i, k and m) and 10 µm (b, d, f, h, j, l and n).
Microscopic observation of colon sections from control groups (CTL, BtLj and Cg groups) showed an intact epithelium with no inflammatory cell infiltrates in the colon mucosa (Fig. 4C,D). In contrast, D or DCg mice showed histopathological changes consistent with an inflammatory process including epithelial damage, scattered inflammatory cell infiltrates in the mucosa and submucosa with the presence of oedema. However, the histopathological features were less severe in DBtLj and DCgBtLj mice than in D or DCg mice and some colon sections were entirely normal with no evidence of inflammatory changes suggesting that BtLj treatment protected mice from DSS-mediated colitis (Fig. 4C,D).
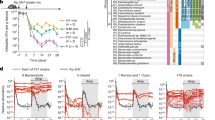
The number of C. glabrata colony-forming units (CFU) was counted in the stools and different segments of the digestive tract, in particular the colon and stomach. A significant decrease in C. glabrata CFU was observed in stool samples between day 1 and day 4 for all groups (Fig. 5A). This decrease continued until day 6 when significant elimination of C. glabrata was observed in the Cg group. Following DSS treatment, an increase in C. glabrata CFU was observed in the DCg group on days 12 and 14 and this fungal overgrowth was correlated with the development of colitis (Fig. 5A). In contrast, treatment with BtLj significantly decreased fungal overgrowth in mice. In addition, C. glabrata CFU were significantly reduced in the stomach and colon of DCgBtLj mice when compared to DCg mice (Fig. 5B,C). In terms of the impact of anaerobic bacteria on colitis, E. coli and E. faecalis populations increased significantly starting from day 8 to day 14 in D and DCg groups when compared to controls and this overgrowth occurred regardless of C. glabrata colonization. In contrast, a significant decrease in these two bacterial populations was observed in mice treated with BtLj (Fig. 5D,E).
Effect of B. thetaiotaomicron and L. johnsonii on C. glabrata elimination from the gut and faecal aerobic bacteria. (A–C) Number of C. glabrata colonies recovered from the stools, stomach and colon. Data are the mean ± SD of eight mice per group from two independent experiments. (D and E) Determination of the number of viable E. coli and E. faecalis colonies recovered from stools. For all experiments, stool bacteria were collected from each tagged mouse on day 0 before DSS treatment and C. glabrata challenge. Data are the mean ± SD of eight mice per group from two independent experiments.
Effect of B. thetaiotaomicron and L. johnsonii on modulation of the innate immune receptor and cytokine expression and on induction of IgA in the colon
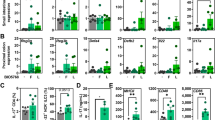
TLR and MBL-C play a crucial role in sensing Candida species and have a pivotal role in the pathogenesis of IBD. Following DSS treatment, significant increases in expression of TLR4, TLR5, TLR8, TLR9 and MBL-C were observed in D and DCg mice when compared to controls while BtLj treatment reduced TLR and MBL-C expression; the exception was the TLR9 transcript which was increased significantly in mice challenged with BtLj (Fig. 6). Expression of chitinase-like 3 protein, which is involved in the degradation and elimination of fungi, was highly increased in DCgBtLj mice. This high expression of chitinase-like 3 protein was correlated with the increased expression of TLR9 and elimination of C. glabrata from the gut. In terms of the signalling pathways mediated by TLR in particular, the MyD88-dependent pathway, which is common to all TLRs, and NF-κB expression increased significantly and an increase in expression of Myd88 and NF-kB signalling factors was observed following DSS treatment, while treatment with BtLj reduced this expression (Fig. 6). These data correlate with the expression of the pro-inflammatory cytokine IL-1β, which was decreased significantly in mice treated with BtLj, while the expression of the anti-inflammatory cytokine IL-10 was highly increased in these mice. In terms of IgA induction in the colon following BtLj treatment, a basal IgA concentration of approximately 600 ng/mL was observed in the control groups (CTL, Cg and BtLj). IgA level increased during the development of colitis, in particular in D or DCg mice where IgA levels were around 700 and 800 ng/mL, respectively (Fig. 7). However, the IgA level was even higher in groups treated with BtLj (800 and 1,000 ng/mL for DBtLj and DCgBtLj, respectively) (Fig. 7).
Discussion
The intestinal microbiota act as a natural barrier against the proliferation of opportunistic pathogens including the opportunistic yeast Candida1,2. Bacteroidetes and Firmicutes are two dominant phyla, which mostly contain obligate anaerobic bacteria and are responsible for providing energy sources to colonic enterocytes and protection against pathogenic microorganisms24. Although the aetiology of IBD remains unclear, intestinal inflammation is often correlated with changes in the biodiversity of the gut microbiota in IBD patients3. These changes in the gut microbiota are referred to as dysbiosis25. Additionally, oxygen and reactive oxygen species play a crucial role in the pathogenesis of IBD by creating an imbalance between obligate and facultative anaerobes. Intestinal inflammation promotes an increase in release of haemoglobin carrying oxygen and reactive oxygen species into the intestinal lumen which creates a microenvironment for facultative anaerobes and a reduction in obligate anaerobes that leads to increased inflammation. The metabolic biofilm model shows that oxygen can play a crucial role in IBD dysbiosis by increasing the abundance of E. coli at the expense of the obligate anaerobe B. thetaiotaomicron26. In terms of pathogenic yeasts, it has been shown that intestinal inflammation promotes the overgrowth of Candida species, which in turn aggravates intestinal inflammation in mice21,27. Clinically, CD patients have high levels of ASCA and an abundance of Candida species in the gut when compared to healthy individuals. Additionally, in the DSS-induced colitis model, which mimics what happens in IBD patients, it has been reported that intestinal inflammation and C. glabrata overgrowth promote a modification of the gut microbiota biodiversity14. Among these aerobic bacteria, E. coli and E. faecalis populations are highly increased while two obligate anaerobic bacteria, L. johnsonii and B. thetaiotaomicron are reduced during the development of colitis, indicating that an alteration of the gut microbiota accelerates the disease process15. Furthermore, a decrease in these two anaerobic bacteria is correlated with the overgrowth of C. glabrata in the mouse gut. This observation led us to assess whether B. thetaiotaomicron and L. johnsonii can inhibit the growth of Candida species in vitro.
In the present study, we observed that co-incubation of Candida with L. johnsonii and B. thetaiotaomicron decreased the viability and growth of Candida species. These data are consistent with different studies showing that growth, adhesion and filamentation of C. albicans are inhibited by Lactobacillus species28. In addition, we showed that B. thetaiotaomicron induced the degradation of C. glabrata cell wall mannans. These data support those of a study showing that B. thetaiotaomicron utilizes complex machinery to digest and metabolize C. albicans cell wall mannans, enabling B. thetaiotaomicron to adapt to different competitive environments and to nutrients present in the intestinal tract29. Furthermore, we also observed that L. johnsonii produces enzymes with chitinase-like activity that promote the degradation of chitin and elimination of both C. albicans and C. glabrata. This in vitro approach reveals that these two anaerobic bacteria can inhibit the growth of Candida species through their antifungal enzyme activities.
In the DSS-induced colitis model, L. johnsonii and B. thetaiotaomicron were administered orally to mice for 5 days in order to restore the anaerobic bacteria and to promote fungal elimination. We observed a significant decrease in inflammatory parameters in terms of body weight, clinical score and histological score for inflammation in mice treated with anaerobic bacteria. In addition, a decrease in E. coli and E. faecalis and C. glabrata populations was observed in the mice indicating that these two anaerobic bacteria not only have a beneficial effect on C. glabrata elimination from the gut, but also on modulating intestinal inflammation and restoring the microbial balance.
IgA plays a crucial role in the elimination of pathogens by inhibiting their adherence and access to epithelial surfaces30. Germ-free mice have low levels of intestinal IgA whereas restoring their commensal microbiota promotes IgA production31,32. In our study, restoration of these two anaerobic bacteria was found to promote IgA secretion, which was correlated with a decrease in E. coli, E. faecalis and C. glabrata populations in mice.
Changes in gut microbiota biodiversity are known to affect the regulation of TLR and other intestinal receptors, in particular mannose-binding lectin (MBL)10,33. MBL is a collectin produced by hepatocytes and intestinal epithelial cells33. Sensing of the fungal cell wall by the MBL complex contributes to the elimination of Candida species from the gut33,34. Following DSS treatment and C. glabrata challenge, high expression of TLR4, 5, 8 and 9 and MBL-C was observed in the colon, indicating that the development of colitis and alterations in the gut microbiota drive TLR and MBL-C activation by different opportunistic bacteria and yeasts including E. coli, E. faecalis and C. glabrata. In contrast, treatment of mice with BtLj reduced the expression of TLR and MBL-C, except for TLR9 which was increased substantially. TLR9 is expressed intracellularly in immune cells and on the apical and basolateral membranes of intestinal epithelial cells35. Remarkably, this high TLR9 expression was correlated with low NF-κB transcripts in mice treated with BtLj. TLR9-deficient mice were highly susceptible to experimental colitis and had a lower NF-κB activation threshold when compared to wild-type and TLR2-deficient mice36.
IL-1β is a potent pro-inflammatory cytokine that is often upregulated in IBD patients and is involved in maintaining chronic gut inflammation, while the anti-inflammatory cytokine IL-10 has a crucial role in regulating inflammasome-mediated IL-1β production37,38,39. Patients with IL-10R deficiency develop severe IBD in the first months of life40. In the present study, the development of colitis and C. glabrata overgrowth were associated with high expression of IL-1β and down-regulation of IL-10. These findings were corroborated by high clinical and histological scores and an abundance of aerobic bacterial populations. In contrast, mice treated with BtLj displayed attenuated colitis, accompanied by a significant reduction in IL-1β and an increase in IL-10 transcripts. This IL-10 up-regulation was associated with increased chitinase-like 3 protein, which is involved in the degradation of fungal chitin. These data provide evidence that L. johnsonii and B. thetaiotaomicron, through their chitinase-like and mannosidase-like activities, can generate small mannose and chitin particles that are recognized by intestinal MBL-C and TLR and promote the activation of chitinase-like protein-1. This in turn enables chitin digestion and the generation of small sized chitin particles that drive IL-10 production via TLR-9 sensing, promoting the attenuation of colitis and C. glabrata elimination. These data are consistent with a previous study that demonstrated that the administration of fungal chitin activated chitinase-like protein-1, promoting C. glabrata elimination in a DSS-induced colitis model14.
In conclusion, L. johnsonii and B. thetaiotaomicron can interact directly with Candida species and induce degradation of the fungal cell wall, mediated via chitinase-like and mannosidase-like activities, which promotes inhibition of Candida species growth. In the DSS-induced colitis model, oral administration of L. johnsonii and B. thetaiotaomicron restored the imbalance between aerobic and anaerobic populations following DSS treatment and C. glabrata challenge and resulted in a significant reduction in inflammatory parameters revealed by a reduction in clinical and histological scores for inflammation. L. johnsonii and B. thetaiotaomicron also decreased pro-inflammatory mediators and enhanced anti-inflammatory cytokine responses. Modulation of cytokine expression is associated with high TLR9 expression and chitinase-like protein-1 activation, which promotes the elimination of C. glabrata from the gut. Overall, these findings provide evidence that L. johnsonii and B. thetaiotaomicron not only attenuated the development of colitis mediated by TLR9 but were also involved in the elimination of C. glabrata from the gut via chitinase-like and mannosidase-like activities.
Methods
Fungal and bacterial strains
C. glabrata BG2 wild-type (ATCC; Cg), C. albicans SC 5,314 (Ca) and bioluminescent C. albicans CEC749 Gluc strains were used in the present study18,41. Fungal culture was carried out in YPD medium (yeast extract 1%, peptone 1%, dextrose 1%) on a rotary shaker for 18 h at 37 °C. The culture obtained was then centrifuged at 2,500 rpm for 5 min and washed twice in PBS (phosphate-buffered saline). For bacterial culture, B. thetaiotaomicron and L. johnsonii were isolated from mouse stools on Bacteroides Bile Esculin agar and De Man, Rogosa and Sharpe (MRS) culture media, respectively, at 37 °C for 48 h under anaerobic conditions14,15. Identification of B. thetaiotaomicron and L. johnsonii was performed by MALDI-TOF mass spectrometry (Maldi-TOF, Microflex-Bruker Daltonics)14.
Determination of the antifungal effect of B. thetaiotaomicron and L. johnsonii on Candida species
The viability of C. albicans cells was tracked in real-time using a bioluminescent yeast strain challenged with B. thetaiotaomicron and L. johnsonii. A suspension of 106 bioluminescent C. albicans in 150 µL RPMI media was added to each well of a black 96-well plate (Greiner bio-one, Chimney well). B. thetaiotaomicron and/or L. johnsonii were then added to each well at a ratio of 1:1, 1:5 or 5:1 yeast-bacteria. Coelenterazine was then added to each well at a concentration of 2 μM. Bioluminescence kinetics were measured at 30 and 120 min and analysed with a FLUOstar Omega Fluorometer (BMG Labtech). The positive control consisted of C. albicans strain alone. The kinetics were also measured with a Xenogen device. Viability of C. glabrata in the presence of B. thetaiotaomicron and/or L. johnsonii was determined by serial dilution. A suspension of C. glabrata and B. thetaiotaomicron and/or L. johnsonii at 1:1, 1: 5 or 5:1 yeast-bacteria ratios was incubated for 30 or 120 min and an aliquot of 100 µL of each dilution was plated on Sabouraud dextrose agar and incubated for 24 h. For observation by confocal microscopy, C. glabrata cells were labelled with concanavalin A (ConA)-rhodamine and bacterial cells with calcein (Invitrogen, France). Specific slides (well 6.7 mm; Thermo Scientific, France) were used in this experiment and were examined by confocal microscopy (Zeiss LSM710, Zeiss Airyscan SR mode × 6,311.4).
Antifungal activities of bacterial enzymes
To determine the chitinase activity of L. johnsonii, a suspension of 106 yeasts in the presence of L. johnsonii (at a ratio of 1:1) was incubated overnight at 37 °C in 500 µL RPMI. After co-incubation, chitinase activity was measured as described previously42. Briefly, 0.5 mL of sample, 1 mL of acetate buffer (0.2 M at pH 5.0) and 5 mg of chitin-azur (Sigma, France) were mixed and incubated at 50 °C for 3 h. The samples were then centrifuged. The supernatant was read at 550 nm using a spectrophotometer. The positive control consisted of pure chitinase (chitinase from Streptomyces griseus at a concentration of 200 U/g; Sigma, France). For mannosidase activities of B. thetaiotaomicron, a suspension of 107 yeasts in the presence of B. thetaiotaomicron (at a ratio of 1:1) was incubated overnight at 37 °C in 500 µL RPMI. The samples were then centrifuged and washed several times with PBS. Proteins and glycoconjugates were recovered by alkaline extraction under reducing conditions. The protein content of each extract was estimated using a bicinchoninic acid protein assay (Pierce) and was adjusted to the same protein concentration prior to analysis by SDS-PAGE on a 10% polyacrylamide gel18. The glycoproteins were then probed with either biotinylated lectin ConA (Sigma-Aldrich, France) or GNL (biotinylated galanthus nivalis; Vector Laboratories, California USA), both diluted 1:1,000. Horseradish peroxidase (HRP)-labelled streptavidin (1:2000 dilution) (Southern Biotech) was used to detect mannans. The glycoproteins of interest were revealed and detected using a reading device (Carestream, Image station, 4,000 MM Pro) and different exposures of the blots were performed (Supplementary data). For the determination of C. glabrata mannan levels by ELISA, a C. glabrata suspension of 105 cells was co-incubated with 105 B. thetaiotaomicron in 200 µL RPMI for 2 h or 48 h in 96-well plates (Greiner bio-one, Chimney well). After centrifugation, the supernatant was removed and the plate was saturated with 1% PBS-bovine serum albumin (BSA; Sigma-Aldrich, France) for 1 h. The supernatant was then removed and either ConA (Sigma-Aldrich, France) or GNL was added to each well at a concentration of 10 μg/mL for 1 h. After different washes with PBS, 100 μL of streptavidin-HRP was added to each well for 1 h. Absorbance was measured at 450 nm. Bacteria alone were used as controls.
Animals
The animals used were 3–4-month-old wild-type female C57BL/6 mice, certified disease-free (Janvier Laboratories, France). The mice were housed in the pet store of the Faculty of Medicine, Lille. The temperature of the room was maintained at 21 °C and the mice had free access to water and food with exposure to light 12 h/day. All animal experiments were approved by the subcommittee for Research Animal Care, Regional Hospital Centre, Lille, France (00550.05) and in accordance with institutional (86/609/CEE) and European guidelines for the care and use of laboratory animals. The animals were tagged and separated into seven groups. Control groups were: CTL group, receiving only water; Cg group, receiving only C. glabrata; and BtLj group, receiving only B. thetaiotaomicron + L. johnsonii. The experimental groups were: D (group treated with DSS only), which represented the control group for intestinal inflammation; DCg (DSS + C. glabrata), which served as a control group for fungal colonization and intestinal inflammation; DBtLj (DSS + B. thetaiotaomicron + L. johnsonii), group to assess the effect of anaerobic bacteria on intestinal inflammation; and DCgBtLj (DSS + C. glabrata + B. thetaiotaomicron + L. johnsonii), group of interest.
Inoculum preparation and induction of colitis
Mice were given 200 µL PBS containing 108 live C. glabrata cells on day 1 by oral gavage. From day 1 to day 14, mice were also treated with DSS (36–50 kDa; stock solution 2%, diluted solution 1.5%, MP Biomedicals, LLC, Germany) in drinking water in order to induce colitis. For restoration of the two anaerobic bacterial populations, mice were given 106 live B. thetaiotaomicron and/or 106 live L. johnsonii in 200 µL PBS orally and daily for 5 days, starting on day 1. The presence of C. glabrata in the gut was assessed by counting the number of colonies in faeces (approximately 0.1 g/sample) collected from each tagged animal on day 14. Faecal samples were suspended in 1 mL PBS and then cultured on Candi-Select medium (Bio-Rad Laboratories, France). The results are shown as colony-forming units (CFU)/mg faeces21. For the isolation of bacteria, mouse faecal samples were collected and serial dilutions were performed daily. The samples were cultured on non-selective bacterial media (AC agar) focusing on the most representative cultivable anaerobic and aerobic bacteria that can undergo changes during intestinal inflammation. Isolation of aerobic and anaerobic bacteria was carried out on MacConkey (MCK) and Bile Esculin azide culture media, respectively. Identification of bacteria isolated on selective media was performed by MALDI-TOF mass spectrometry (Maldi-TOF; Microflex-Bruker Daltonics)14.
Clinical and histological scores for inflammation
Body weight of each tagged mouse was measured daily and the presence of blood in the rectum and stool consistency were also determined14. Clinical scores, as described previously, were analysed independently by two investigators blinded to the protocol27,41. Two scores (stool consistency and bleeding) were added, resulting in a total clinical score ranging from 0 (healthy) to 12 (maximum colitis activity). For the histological score, the entire colon from the caecum to the anus was collected. The colon samples were fixed overnight in 4% paraformaldehyde-acid, embedded in paraffin, cut in a microtome (Leica RM2245, Leica Biosystems, Germany) and affixed onto slides for histological staining. The colon sections (4 µm thick) were stained with periodic acid–Schiff stain (Merck, France) and then viewed with a digital slide scanner (Axio-Scan.Z1, Zeiss, Jena, Germany). Histological scoring was performed by two independent investigators blinded to the protocol. The score was based on the infiltration of inflammatory cells into the submucosa and epithelial damage of the colon and ranged from 0 (no changes) to 6 (extensive cell infiltration and tissue damage)14,43.
Secretory IgA in the mouse colons
Mouse colons were flushed with 2 mL PBS and the intestinal content was vortexed and assayed for IgA using an ELISA kit (Invitrogen, France)30. Briefly, the first antibody (purified monoclonal anti-mouse IgA) was fixed to the bottom of the wells of the 96-well plate (overnight at 4 °C). After different washings, mouse colon contents were deposited in each well for 2 h and the wells were then washed. The second antibody (HRP-conjugated monoclonal anti-mouse IgA) was added to each well for 1 h. After washing, substrate solution was added and incubated for 15 min followed by the addition of stop solution. Optical density was measured with a spectrophotometer at 450 nm (Biorad, France). The results were analysed by normalizing to standard concentrations of IgA.
Real-time mRNA quantification of innate immune receptors
Extraction of total RNA from the colon was carried out using a commercial kit (Nucleospin RNA/Protein; Macherey–Nagel, France). RNA was quantified by spectrophotometry (Nanodrop; Nyxor Biotech, France). mRNA reverse transcription was performed in a final volume of 20 µL from 1 µg total RNA (high capacity cDNA RT kit; Applied Biosystems). Fast SYBR green (Applied Biosystems) was used in PCR to amplify the cDNA in a one-step system (Applied Biosystems). SYBR green dye intensity was assessed using one-step software. All results were normalized to the reference gene POLR2A15,33.
Statistical analysis
All data are presented as the mean ± standard deviation (SD) of individual experimental groups. Pairs of groups were compared using the Mann–Whitney U test. Differences were considered significant when the P value was as follows: p < 0.05; p < 0.01; p < 0.001. Statistical analyses were carried out using Prism 4.0 from GraphPad and XLSTAT.
References
Pultz, N. J., Stiefel, U., Subramanyan, S., Helfand, M. S. & Donskey, C. J. Mechanisms by which anaerobic microbiota inhibit the establishment in mice of intestinal colonization by vancomycin-resistant Enterococcus. J. Infect. Dis. 191, 949–956. https://doi.org/10.1086/428090 (2005).
Kamada, N., Chen, G. Y., Inohara, N. & Nunez, G. Control of pathogens and pathobionts by the gut microbiota. Nat. Immunol. 14, 685–690. https://doi.org/10.1038/ni.2608 (2013).
Hoarau, G. et al. Bacteriome and mycobiome interactions underscore microbial dysbiosis in familial Crohn’s disease. mBio https://doi.org/10.1128/mBio.01250-16 (2016).
Panaccione, R. Mechanisms of inflammatory bowel disease. Gastroenterol. Hepatol. (N.Y.) 9, 529–532 (2013).
Darfeuille-Michaud, A. et al. Presence of adherent Escherichia coli strains in ileal mucosa of patients with Crohn’s disease. Gastroenterology 115, 1405–1413 (1998).
Hugot, J. P. et al. Association of NOD2 leucine-rich repeat variants with susceptibility to Crohn’s disease. Nature 411, 599–603. https://doi.org/10.1038/35079107 (2001).
Hart, A. L. et al. Characteristics of intestinal dendritic cells in inflammatory bowel diseases. Gastroenterology 129, 50–65 (2005).
Liguori, G. et al. Fungal dysbiosis in mucosa-associated microbiota of Crohn’s disease patients. J. Crohns Colitis 10, 296–305. https://doi.org/10.1093/ecco-jcc/jjv209 (2016).
Duerr, R. H. et al. A genome-wide association study identifies IL23R as an inflammatory bowel disease gene. Science 314, 1461–1463. https://doi.org/10.1126/science.1135245 (2006).
Lavelle, E. C., Murphy, C., O’Neill, L. A. & Creagh, E. M. The role of TLRs, NLRs, and RLRs in mucosal innate immunity and homeostasis. Mucosal Immunol. 3, 17–28. https://doi.org/10.1038/mi.2009.124 (2010).
Zhou, Y. et al. Increased Enterococcus faecalis infection is associated with clinically active Crohn disease. Medicine (Baltimore) 95, e5019. https://doi.org/10.1097/MD.0000000000005019 (2016).
Kim, S. C., Tonkonogy, S. L., Karrasch, T., Jobin, C. & Sartor, R. B. Dual-association of gnotobiotic IL-10-/- mice with 2 nonpathogenic commensal bacteria induces aggressive pancolitis. Inflamm. Bowel Dis. 13, 1457–1466. https://doi.org/10.1002/ibd.20246 (2007).
Darfeuille-Michaud, A. Adherent-invasive Escherichia coli: a putative new E. coli pathotype associated with Crohn’s disease. Int. J. Med. Microbiol. 292, 185–193. https://doi.org/10.1078/1438-4221-00201 (2002).
Charlet, R. et al. Remodeling of the Candida glabrata cell wall in the gastrointestinal tract affects the gut microbiota and the immune response. Sci. Rep. 8, 3316. https://doi.org/10.1038/s41598-018-21422-w (2018).
Charlet, R., Bortolus, C., Barbet, M., Sendid, B. & Jawhara, S. A decrease in anaerobic bacteria promotes Candida glabrata overgrowth while beta-glucan treatment restores the gut microbiota and attenuates colitis. Gut Pathog. 10, 50. https://doi.org/10.1186/s13099-018-0277-2 (2018).
Ryan, P. et al. Candidaemia in an Irish intensive care unit setting between 2004 and 2018 reflects increased incidence of Candida glabrata. J. Hosp. Infect. https://doi.org/10.1016/j.jhin.2019.01.017 (2019).
Lee, I. et al. Clinical and economic outcomes of decreased fluconazole susceptibility in patients with Candida glabrata bloodstream infections. Am. J. Infect. Control 38, 740–745. https://doi.org/10.1016/j.ajic.2010.02.016 (2010).
Jawhara, S. et al. Murine model of dextran sulfate sodium-induced colitis reveals Candida glabrata virulence and contribution of beta-mannosyltransferases. J. Biol. Chem. 287, 11313–11324. https://doi.org/10.1074/jbc.M111.329300 (2012).
Poulain, D. et al. Yeasts: neglected pathogens. Dig. Dis. 27(Suppl 1), 104–110. https://doi.org/10.1159/000268129 (2009).
Sendid, B. et al. Antibodies against glucan, chitin, and Saccharomyces cerevisiae mannan as new biomarkers of Candida albicans infection that complement tests based on C. albicans mannan. Clin. Vacc. Immunol. 15, 1868–1877. https://doi.org/10.1128/CVI.00200-08 (2008).
Jawhara, S. et al. Colonization of mice by Candida albicans is promoted by chemically induced colitis and augments inflammatory responses through galectin-3. J. Infect. Dis. 197, 972–980. https://doi.org/10.1086/528990 (2008).
Conde, R., Cueva, R., Pablo, G., Polaina, J. & Larriba, G. A search for hyperglycosylation signals in yeast glycoproteins. J. Biol. Chem. 279, 43789–43798. https://doi.org/10.1074/jbc.M406678200 (2004).
Fradin, C. et al. Beta-1,2 oligomannose adhesin epitopes are widely distributed over the different families of Candida albicans cell wall mannoproteins and are associated through both N- and O-glycosylation processes. Infect. Immun. 76, 4509–4517. https://doi.org/10.1128/IAI.00368-08 (2008).
Ramakrishna, B. S. Role of the gut microbiota in human nutrition and metabolism. J. Gastroenterol. Hepatol. 28(Suppl 4), 9–17. https://doi.org/10.1111/jgh.12294 (2013).
Carding, S., Verbeke, K., Vipond, D. T., Corfe, B. M. & Owen, L. J. Dysbiosis of the gut microbiota in disease. Microb. Ecol. Health Dis. 26, 26191. https://doi.org/10.3402/mehd.v26.26191 (2015).
Henson, M. A. & Phalak, P. Microbiota dysbiosis in inflammatory bowel diseases: in silico investigation of the oxygen hypothesis. BMC Syst. Biol. 11, 145. https://doi.org/10.1186/s12918-017-0522-1 (2017).
Jawhara, S. & Poulain, D. Saccharomyces boulardii decreases inflammation and intestinal colonization by Candida albicans in a mouse model of chemically-induced colitis. Med. Mycol. 45, 691–700. https://doi.org/10.1080/13693780701523013 (2007).
Matsubara, V. H., Wang, Y., Bandara, H. M., Mayer, M. P. & Samaranayake, L. P. Probiotic lactobacilli inhibit early stages of Candida albicans biofilm development by reducing their growth, cell adhesion, and filamentation. Appl. Microbiol. Biotechnol. 100, 6415–6426. https://doi.org/10.1007/s00253-016-7527-3 (2016).
Cuskin, F. et al. Human gut Bacteroidetes can utilize yeast mannan through a selfish mechanism. Nature 517, 165–169. https://doi.org/10.1038/nature13995 (2015).
Cao, A. T., Yao, S., Gong, B., Elson, C. O. & Cong, Y. Th17 cells upregulate polymeric Ig receptor and intestinal IgA and contribute to intestinal homeostasis. J. Immunol. 189, 4666–4673. https://doi.org/10.4049/jimmunol.1200955 (2012).
Talham, G. L., Jiang, H. Q., Bos, N. A. & Cebra, J. J. Segmented filamentous bacteria are potent stimuli of a physiologically normal state of the murine gut mucosal immune system. Infect. Immun. 67, 1992–2000 (1999).
Klaasen, H. L. et al. Intestinal, segmented, filamentous bacteria in a wide range of vertebrate species. Lab. Anim. 27, 141–150. https://doi.org/10.1258/002367793780810441 (1993).
Choteau, L. et al. Role of mannose-binding lectin in intestinal homeostasis and fungal elimination. Mucosal. Immunol. 9, 767–776. https://doi.org/10.1038/mi.2015.100 (2016).
Jensenius, H. et al. Mannan-binding lectin: structure, oligomerization, and flexibility studied by atomic force microscopy. J. Mol. Biol. 391, 246–259. https://doi.org/10.1016/j.jmb.2009.05.083 (2009).
Lee, J., Mo, J. H., Shen, C., Rucker, A. N. & Raz, E. Toll-like receptor signaling in intestinal epithelial cells contributes to colonic homoeostasis. Curr. Opin. Gastroenterol. 23, 27–31. https://doi.org/10.1097/MOG.0b013e3280118272 (2007).
Lee, J. et al. Maintenance of colonic homeostasis by distinctive apical TLR9 signalling in intestinal epithelial cells. Nat. Cell Biol. 8, 1327–1336. https://doi.org/10.1038/ncb1500 (2006).
Muro, M. & Mrowiec, A. Interleukin (IL)-1 gene cluster in inflammatory bowel disease: is IL-1RA implicated in the disease onset and outcome?. Dig. Dis. Sci. 60, 1126–1128. https://doi.org/10.1007/s10620-015-3571-6 (2015).
van Deventer, S. J., Elson, C. O. & Fedorak, R. N. Multiple doses of intravenous interleukin 10 in steroid-refractory Crohn’s disease. Crohn’s Disease Study Group. Gastroenterology 113, 383–389 (1997).
Shouval, D. S. et al. Interleukin 1beta mediates intestinal inflammation in mice and patients with interleukin 10 receptor deficiency. Gastroenterology 151, 1100–1104. https://doi.org/10.1053/j.gastro.2016.08.055 (2016).
Kotlarz, D. et al. Loss of interleukin-10 signaling and infantile inflammatory bowel disease: implications for diagnosis and therapy. Gastroenterology 143, 347–355. https://doi.org/10.1053/j.gastro.2012.04.045 (2012).
Bortolus, C. et al. A small aromatic compound has antifungal properties and potential anti-inflammatory effects against intestinal inflammation. Int. J. Mol. Sci. https://doi.org/10.3390/ijms20020321 (2019).
Gomez Ramirez, M., Rojas Avelizapa, L. I., Rojas Avelizapa, N. G. & Cruz Camarillo, R. Colloidal chitin stained with Remazol Brilliant Blue R, a useful substrate to select chitinolytic microorganisms and to evaluate chitinases. J. Microbiol. Methods 56, 213–219 (2004).
Jawhara, S. et al. Modulation of intestinal inflammation by yeasts and cell wall extracts: strain dependence and unexpected anti-inflammatory role of glucan fractions. PLoS ONE 7, e40648. https://doi.org/10.1371/journal.pone.0040648 (2012).
Acknowledgements
The authors thank Ms. Nadine François, Karine Lecointe and Meryem Tardivel for their excellent technical assistance. This work was funded by the FP7 Health 260338 ‘‘ALLFUN’’ Project ‘‘Fungi in the setting of inflammation, allergy and auto-immune diseases: translating basic science into clinical practices’’. This work was partially funded by the Agence Nationale de la Recherche (ANR) in the setting of Project "InnateFun", promotional reference ANR-16-IFEC-0003-05, in the "Infect-ERA" program.
Author information
Authors and Affiliations
Contributions
R.C., C.B. and S.J. performed the experiments. R.C., C.B., B.S. and S.J. analysed the data. R.C., C.B., B.S. and S.J. interpreted the results of the experiments. S.J. designed the experiments and drafted the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Charlet, R., Bortolus, C., Sendid, B. et al. Bacteroides thetaiotaomicron and Lactobacillus johnsonii modulate intestinal inflammation and eliminate fungi via enzymatic hydrolysis of the fungal cell wall. Sci Rep 10, 11510 (2020). https://doi.org/10.1038/s41598-020-68214-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-68214-9
This article is cited by
-
Biosorption and Bioprotective Potential of Levilactobacillus brevis in Mice Challenged by Lead-Induced Oxidative Stress
Biological Trace Element Research (2024)
-
The Antidepressant- and Anxiolytic-Like Effects of the Phosphodiesterase Type-5 Inhibitor Tadalafil are Associated with the Modulation of the Gut-Brain Axis During CNS Autoimmunity
Journal of Neuroimmune Pharmacology (2024)
-
Dectin-1 signaling on colonic γδ T cells promotes psychosocial stress responses
Nature Immunology (2023)
-
Oral microbial changes and oral disease management before and after the treatment of hematological malignancies: a narrative review
Clinical Oral Investigations (2023)
-
Gut microbiota-bile acid crosstalk contributes to the rebound weight gain after calorie restriction in mice
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.