Abstract
Fragile X associated tremor/ataxia syndrome (FXTAS) is a late adult-onset neurodegenerative disorder that affects movement and cognition in male and female carriers of a premutation allele of 55–200 CGG repeats in the Fragile X mental retardation (FMR1) gene. It is currently unknown if and when an individual carrier of a premutation allele will develop FXTAS, as clinical assessment fails to identify carriers at risk before significant neurological symptoms are evident. The primary objective of this study was to investigate the alternative splicing landscape at the FMR1 locus in conjunction with brain measures in male individuals with a premutation allele enrolled in a very first longitudinal study, compared to age-matched healthy male controls, with the purpose of identifying biomarkers for early diagnosis, disease prediction and, a progression of FXTAS. Our findings indicate that increased expression of FMR1 mRNA isoforms, including Iso4/4b, Iso10/10b, as well as of the ASFMR1 mRNAs Iso131bp, are present in premutation carriers as compared to non-carrier healthy controls. More specifically, we observed a higher expression of Iso4/4b and Iso10/10b, which encode for truncated proteins, only in those premutation carriers who developed symptoms of FXTAS over time as compared to non-carrier healthy controls, suggesting a potential role in the development of the disorder. In addition, we found a significant association of these molecular changes with various measurements of brain morphology, including the middle cerebellar peduncle (MCP), superior cerebellar peduncle (SCP), pons, and midbrain, indicating their potential contribution to the pathogenesis of FXTAS. Interestingly, the high expression levels of Iso4/4b observed both at visit 1 and visit 2 and found to be associated with a decrease in mean MCP width only in those individuals who developed FXTAS over time, suggests their role as potential biomarkers for early diagnosis of FXTAS.
Similar content being viewed by others
Introduction
The fragile X mental retardation (FMR1) gene consists of 17 exons spanning approximately 38 kb of genomic DNA. A trinucleotide repeat expansion, greater than 200 CGG, with consequent methylation of the 5′UTR (untranslated region) of FMR1 gene, leads to Fragile X Syndrome (FXS), the most common form of intellectual disability and known monogenic cause of Autism Spectrum Disorder1. Expansions between 55 and 200 CGG repeats (known as premutation carriers) confer the risk of developing Fragile X-associated tremor/ataxia syndrome (FXTAS), a late-onset neurodegenerative disorder characterized by intention tremor, gait ataxia, autonomic dysfunction, and Parkinsonism2. In addition, females premutation carriers are at risk of developing Fragile X-associated primary ovarian insufficiency (FXPOI) that affects ovary function in women leading to early menopause and irregular elevation of follicle-stimulating hormone (FSH)3. The prevalence of the premutation allele among the general population is 1:110–200 females and 1:430 males with an estimated 40–75% of males and 8–16% of females developing FXTAS4,5.
At the molecular level, FXTAS is characterized by an increased level of FMR1 mRNA containing expanded CGG repeats6. The proposed molecular mechanisms of FXTAS pathogenesis include the sequestration of CGG binding proteins by the elevated levels of FMR1 mRNA leading to RNA toxicity, the production of toxic FMRPolyG protein due to RAN translation and the chronic activation of DNA damage response [reviewed in7].
Extensive alternative splicing of the FMR1 gene has been observed8,9,10,11,12,13,14,15,16. Alternative splicing (AS) is a regulated process occurring during gene expression that increases protein diversity and represents a powerful evolutionary resource. It regulates the protein localization, enzymatic properties, stability, interaction with ligands and membranes17 and is common in the nervous system playing a major role in neurogenesis, brain development18,19 and cell survival20. It is increasingly recognized that disruption of the splicing process, which is regulated by different splicing factors, can contribute to a number of neurological disorders19,21,22 including autism spectrum disorder (ASD)23, Parkinson’s disease24, dementia25, spinal muscular atrophy (SMA)26, Prader-Willi syndrome (PWS)27, schizophrenia28, myotonic dystrophy29, amyotrophic lateral sclerosis30,31 and Alzheimer’s disease32.
In the FMR1 gene, altered splicing has been observed in premutation carriers where increased levels of the FMR1 isoforms have been detected15,16. Of the many FMR1 mRNA isoforms that were demonstrated to exist in both humans and mice11,14,15,16, Iso10/10b showed the highest levels of expression in premutation samples which, together with the Iso4/4b (Fig. 1), result in truncated proteins that lack the function of the nuclear export signal (NES) and RGG box15. Of the 49 different FMR1 isoforms identified, 30 appeared to be expressed only in premutation carriers compared to controls16. Additionally, two novel isoforms IsoPB1.50 and IsoPB1.21 retain a portion of the intronic sequence between exons 9 and 10, causing a frameshift, which leads to a premature stop codon and consequently encodes for truncated proteins. A differential increase of specific FMR1 mRNA isoforms has been observed in premutation carriers, suggesting their potential functional relevance in the pathology of FXTAS due to RNA toxicity15,16.
Schematic overview of the FMR1 and ASFMR1 isoforms. Diagram representing the FMR1 locus (top), 4 FMR1 isoforms (Iso10, Iso10b, Iso4 and Iso4b) and the ASFMR1 Iso131bp. Exons are represented in different colors and the alternative splice sites are depicted for exon 12, 14,15 and 17. FMR1 isoforms Iso10, and Iso10b both miss exon 12 and 14 but differ for the splicing acceptor site in exon 17. Likewise, the FMR1 Iso4, and Iso4b both miss exon 14 but differ for the splicing acceptor site in exon 17.
Comprehensive analysis of the transcriptional landscape of the human FMR1 gene revealed the presence of long non-coding RNAs (FMR433, FMR5 and FMR6)34. Importantly, a unique antisense transcript at the FMR1 locus (ASFMR1), that similarly to the FMR1 gene, is upregulated in premutations and not expressed in full mutations, has been identified. Thus, the bidirectional expression of the FMR1 and the ASFMR1 genes has been suggested to potentially contribute to the clinical phenotype of FXTAS35. Interestingly, the ASFMR1 also exhibits a premutation specific alternative splicing, the Iso131bp (Fig. 1), which is mainly expressed in premutation carriers compared to controls, providing a molecular abnormality potentially associated with FXTAS15,36,37,38. However, no studies have been conducted to determine whether altered expression FMR1 and ASFMR1 isoforms are biomarkers of incipient FXTAS, particularly in relation to neurological and neuroanatomical changes.
In this study, we evaluated male premutation carriers enrolled in a longitudinal study at the UC Davis MIND Institute who have been followed for at least two longitudinal time points, and for whom neuroimaging, neuropsychological, and molecular measurements, as well as medical and neurological examinations, were collected. We have recently reported that the middle cerebellar peduncle (MCP) width decreased in a subgroup of these individuals who developed symptoms of FXTAS at subsequent visits (converters) compared to those who did not (non-converters) and compared to normal age-matched controls. Further, we reported reduced midbrain and pons cross-sectional areas in patients with FXTAS compared to both premutation carriers without FXTAS and controls39. These regions play an important role within the cortico-cerebellar pathway, which is necessary for the learning and coordination of various movements40. Measurements of these areas have been shown previously to successfully differentiate subcortical movement disorders, such as Parkinson’s disease41, which presents with tremor similar to that seen in FXTAS.
In the current study, we sought to determine whether the expression levels of alternative splicing isoforms at the FMR1 locus were significantly different both in premutation carriers who did and did not develop symptoms of FXTAS over time compared to non-carrier healthy controls. In addition, we also investigated whether the changes in molecular measures were associated with changes in brain measures.
Materials and methods
Study participants
As part of two continuing longitudinal studies, male participants over the age of 40 years were recruited from the Sacramento, CA area, and throughout the United States and Canada. The study and all experimental protocols were carried out in accordance with the Institutional Review Board (IRB) at the University of California, Davis with written informed consent obtained from all participants in accordance with the Declaration of Helsinki. Participants were fluent in English, with no history of any serious medical or neurological conditions, including a history of alcoholism or drug abuse. FXTAS stage scoring was based on the clinical descriptions as previously described42.
Three groups were included in this study: converters, non-converters, and healthy controls. They were matched by age and the length of the interval between visit 1 (V1) and visit 2 (V2); the converters and non-converters groups were also matched by CGG repeat number and they were selected on the basis of the brain measures availability. After two brain scans, on the basis of neurological assessment, FXTAS stage, and CGG repeat length, 15 participants were classified as “Converters” as they developed clear FXTAS symptomology between visits (FXTAS stage score was 0–1 at visit 1 and ≥ 2 at visit 2; 15 were defined as “Non-converters” because they continued to show no signs of FXTAS at V2 (FXTAS stage score was 0–1 at both V1 and V2) and 15 non-carrier healthy controls (FXTAS stage score was 0 at both V1and V2).
CGG repeat length
Genomic DNA (gDNA) was isolated from 5 mL of peripheral blood leukocytes using the Gentra Puregene Blood Kit (Qiagen, Valencia, CA, United States). CGG repeat allele size and methylation status were assessed using a combination of Southern Blot analysis and PCR amplification. For Southern blot analysis, 5–10 μg of isolated genomic DNA was digested with EcoRI and NruI, run on an agarose gel, transferred on a nylon membrane and hybridized with the FMR1-specific dig-labeled StB12.3 as detailed in43. PCR analysis was performed using FMR1 specific primers (AmplideX PCR/CE, Asuragen, Inc.); amplicons were visualized by capillary electrophoresis, as previously reported an analyzed using Gene Mapper software44.
mRNA expression levels
Total RNA was isolated from 2.5 ml of peripheral blood collected in PAXgene Blood RNA tubes using the PAXgene Blood RNA Kit (Qiagen, Valencia, CA, United States) and quantified using Agilent 2,100 Bioanalyzer system. RNA isolation was performed in a clean and RNA designated area. cDNA was synthesized as previously described45. FMR1 transcript levels measured by performing reverse transcription followed by real-time PCRs (qRT-PCR). qRT-PCR was performed using both Assays-On-Demand from Applied Biosystems (Applied Biosystems, Foster City, CA, United States) and custom-designed TaqMan primers and probe assays,45. Probe and primer assays designed to quantify FMR1 transcripts for the isoform IsoPB1.21 and IsoPB1.50, isoform Iso4/4b and isoform Iso10/10b were as previously reported16. Custom designed primers and probe were also designed to quantify the ASFMR1 gene and ASFMR1 131 bp splice isoform35.
Brain measures
The following methods including MRI acquisition and MRPI analysis were originally described in our previous report39. High resolution structural magnetic resonance imaging (MRIs) acquisition was obtained on a 3 T Siemens Trio scanner using a 32-channel head coil and a T1-weighted 3D MPRAGE sequence with the following parameters: TR = 2170 ms, TE = 4.86 ms, flip angle = 7º, FoV = 256mm2, 192 slices, 1 mm slice thickness. The scans were first aligned along the anterior–posterior commissure line using acpcdetect (https://www.nitrc.org/projects/art)46 or manually using DTI Studio (www.mristudio.org)47. Then MRI bias field correction was performed using N4 (https://stnava.github.io/ANTs/)48.
A series of independent raters (two per measure) who were blinded to the participant age, group, and time point, quantitatively assessed all MR images for four measurements of brain morphology: MCP and superior cerebellar peduncle (SCP) widths as well as pons and midbrain cross-sectional areas based on methods previously described49,50 and detailed below.
The pons and midbrain areas were assessed on the mid-sagittal slice, where horizontal lines were drawn through the superior and inferior pontine notches. The midbrain was measured as the area above the superior pontine line – midbrain tegmentum, while the pons was the area between the two horizontal lines of the superior and inferior notches. The width of both left and right MCPs were measured on parasagittal slices. The linear distance of the MCP was delineated by the peripeduncular cerebrospinal fluid spaces of pontocerebellar cisterns, where the pons was still ‘intact’ and the cerebellum was fully formed (white matter connecting the cerebellar tonsil was present). Finally, the widths of both the left and right SCPs were measured on oblique coronal slices, at the midpoint of the SCP, when it first became separated from the inferior colliculi. The linear distance between the medial and lateral SCP borders was measured. For both MCP and SCP widths, a mean score was calculated by averaging the left and right measurements for each participant. The interrater reliability coefficients were excellent, greater than 0.98, for each of the four measurements. The mean score of the raters was used for further analysis.
Statistical analysis
Statistical Analyses were conducted blind to treatment groups using R, version 3.6.0. Age and CGG repeat numbers were compared between groups using ANOVA F-tests, followed by pairwise comparisons in the event of a significant F test. The association between mRNA expression and CGG repeat length in each group, adjusted for age, was analyzed using linear models with CGG repeat length, group, the interaction between CGG repeat length and group, and age as covariates. mRNA expression was compared between groups, adjusting for age, using linear models with group and age as covariates. The change between timepoint 1 and time point 2 in mRNA expression was compared between groups, adjusting for age and visit interval, using linear models with the group, age, and visit interval as covariates. The association between each brain measure and each molecular measure was analyzed at each time point within each group, adjusting for age, using linear models with molecular measure, group, the group–molecular measure interaction, and age as covariates. The association between changes in brain measures and changes in molecular measures, within each group, adjusting for age and visit interval, was analyzed using linear models including change in molecular measure, group, the group–molecular measure change interaction, age, and visit interval as covariates. mRNA and protein expression values were log-transformed prior to analysis in order to more closely satisfy model assumptions; brain measures were analyzed on their original scale. Due to the exploratory nature of this study, no adjustment for multiple testing was conducted51,52. The tables of all analyses conducted are reported in the supplemental materials.
Results
Demographics
Numbers of participants (N) for each group, age and CGG repeat number are as reported in (Table 1). Participants ages did not differ significantly between the three groups. CGG repeat numbers were significantly lower in healthy controls than in all other groups, as expected (p < 0.001 in both comparisons) and were not significantly different between converters and non-converters (p = 0.445).
FMR1 mRNA isoforms expression analysis
We measured the expression of FMR1 mRNA, FMR1 isoforms Iso10/10b, Iso4/4b, IsoPB1.21, ASFMR1, and ASFMR1 Iso131 at V1, V2 and between the visits (Supplementary Material Table S2a, S2b and S2a). As expected, the expression of FMR1 mRNA was significantly higher in both converters and non-converters as compared to non-carrier healthy controls both at V1 (p < 0.001) and V2 (p < 0.001) but no significant differences were observed between the two premutation groups (Fig. 2a.). FMR1 expression levels were associated with CGG repeat length, with greater expression levels being associated with longer CGG repeat length, in both converters and non-converters compared to non-carrier healthy controls (p < 0.001 for both comparisons; Fig. 2b.).
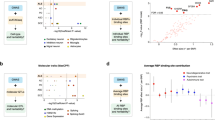
FMR1 mRNA expression among groups and by CGG repeat length. a Box plots showing increased expression levels of FMR1 mRNA in both premutation groups (converters and non-converters) compared to non-carrier healthy controls at V1 (p < 0.001) and V2 (p < 0.001) but no significant differences were observed between the two premutation groups. The heavy line in each box represents the median, the lower and upper box edges represent the 25th and 75th percentiles, respectively, and the lower and upper whiskers represent the smallest and largest observations, respectively. b Scatter plots showing FMR1 mRNA expression as function of CGG repeat number at V1 in the three groups.
The expression levels of Iso10/ Iso10b, CGG dependent (Fig. 3d) were significantly higher in converters at V1 (p = 0.022) and V2 (p = 0.046) respectively, as compared to non-carrier healthy controls. Importantly, these markers were not significantly different in non-converters as compared to non-carrier healthy controls at either V1 (p = 0.401) or V2 (p = 0.592) (Fig. 3a). For Iso4/4b we observed significantly higher expression in converters as compared to non-carrier healthy controls at V1 (p = 0.032) but no difference was found at V2 (p = 0.247) and with the CGG repeat number (Fig. 3e).The expression of these isoforms was not significantly different in non-converters as compared to non-carrier healthy controls both at V1 (p = 0.542) and V2 (p = 0.684) (Fig. 3b). The expression levels of additional transcripts encoding for truncated proteins, IsoPB1.21 and IsoPB1.50 were not significantly different among groups both at V1 or V2.
Isoforms mRNA expression levels among three groups and as function of the CGG repeat number. a Box plots showing increased levels of FMR1 Iso10/10b mRNA at both V1 and V2 (p = 0.022 and p = 0.046 respectively) only in the converter group but not in the non-converter group (p = 0.401 at V1 and p = 0.592 at V2) compared to non-carrier healthy controls. b Box plots showing increased levels of FMR1 Iso4/4b mRNA at V1 (p = 0.032) but not at V2 (p = 0.247) in the converter group compared to non-carrier healthy controls. No differences were observed in the non-converter group at both visits (p = 0.542 at V1 and p = 0.684 at V2). c Box plots showing increased levels of ASFMR1 Iso131bp mRNA at both V1 (p < 0.001) and V2 (p < 0.001) in both the premutation groups compared to non-carrier healthy controls but not statistically significant differences between the two premutation groups. The heavy line in each box represents the median, the lower and upper box edges represent the 25th and 75th percentiles, respectively, and the lower and upper whiskers represent the smallest and largest observations, respectively. Scatter plots showing d FMR1 e of FMR1 Iso4/4b and f of ASFMR1 Iso131bp mRNA expression levels as function of the CGG repeat number at V1 in the three groups.
Significantly increased expression levels of ASFMR1 Iso131 bp, CGG dependent (p = 0.012 for the non-converter group and p = 0.05 for the converter group; Fig. 3f) were observed in the premutation groups, both converters, and non-converters, as compared to non-carrier healthy controls both at V1 (p < 0.001) and V2 (p < 0.001) with no difference in levels between the two premutation groups (Fig. 3c).
However, when comparing the changes in the gene expression levels between V1 and V2 we found a significant greater change in the expression of ASFMR1 Iso131 bp in converters (p = 0.006; Fig. 4) as compared to non-carrier healthy controls while no significant difference was observed between non-converters and non-carrier healthy controls (p = 0.102; Fig. 4).Finally, when comparing the expression levels at V1 and V2 among the groups, no significant differences were detected for any of the FMR1 splicing isoforms or the ASFMR1 gene.
Changes in ASFMR1 Iso131bp expression levels between visits. Boxplots showing higher changes between V1 and V2 in the expression level of the ASFMR1 Iso131bp in the converter group as compared to non-carrier healthy controls while no significant change in non-converter versus control was observed. The heavy line in each box represents the median, the lower and upper box edges represent the 25th and 75th percentiles, respectively, and the lower and upper whiskers represent the smallest and largest observations, respectively.
Correlation between brain measures and molecular measures
We compared brain measures, including MCP and SCP width, midbrain and pons cross-sectional area with the molecular measures at V1, V2 and between the visits (Supplementary Material Table S3a, S3B and S3c). Changes in the measures of brain morphology were associated with the expression of some FMR1 isoforms. Specifically, we found that the higher level of expression of the Iso4/4b mRNAs was associated with smaller MCP width in converters both at V1 (p = 0.028, beta = -0.421; Fig. 5a). and V2 (p = 0.048, beta = -0.531; Fig. 5b) but not in the non-converters at both V1( p = 0.611, beta = -0.064; Fig. 5a) or V2 (p = 0.333, beta = – 0.392; Fig. 5b) and not in the healthy controls (p = 0.530, V1; beta = 0.085; p = 0.365, beta = – 0.376). Further, the expression of the ASFMR1 isoform (131 bp) also increased significantly with a decrease in changes in the pons (p = 0.047, beta = – 0.012) only in the converter group between V1 and V2.
Finally, we found a positive correlation, between the change in brain measures, including the mean SCP (p = 0.017, beta = 0.726) and midbrain area (p = 0.031, beta = 0.316) with increased expression of the FMR1 mRNA, between the two visits only in the converter group. We also found that the expression of Iso10/10b increased significantly with an increase with changes in the midbrain area (p = 0.045, beta = 0.076).
Discussion
Alternative Splicing (AS) is a common process in the central nervous system and crucial for the differentiation and physiology of cells, particularly neurons. It is estimated that AS occurs in about 95% of human genes53, contributing greatly to the regulation of mRNA levels and to proteomic diversity. Different studies have reported that AS events take place at the FMR1 locus and have an impact on the expression and function of FMRP10,11,12,14,54,55,56,57 We recently reported on the characterization of the FMR1 isoforms and showed differential expression and distribution as a function of the CGG repeat number in premutation carriers. Differently from the 24 predicted FMR1 mRNA variants, we also reported on the existence, of at least 49 different ones in several human tissues, 30 of which detected only in premutation carriers15,16. Thus, an altered alternative splicing phenomenon is present in premutation carriers.
In this study, we aimed to identify molecular biomarkers, specifically the expression levels of some alternatively spliced isoforms at the FMR1 locus, for risk prediction, early diagnosis, and progression of developing FXTAS. Increased and CGG dependent expression levels of FMR1 mRNA were observed in premutation groups (including both converters and non-converters) as compared to non-carrier healthy controls confirming many previous reports on this well-established altered molecular phenotype in premutation carriers. In addition to FMR1 mRNA, we also observed elevated expression of various alternative splicing isoforms as a function of CGG repeat suggesting their potential contribution to the RNA toxicity in premutation carriers.
Taking advantage of the longitudinal study design, we investigated the expression profile for these specific isoforms among our three groups and observed the higher expression of isoform Iso10/10b in converters as compared to non-carrier healthy controls at both visits. Importantly, the non-converter group did not show the differential expression for this isoform. This suggests that these isoforms, that encode for truncated proteins might be relevant in the pathogenesis of FXTAS and, pending replication and further confirmation, may eventually play a role in early testing and screening of premutation allele carriers at greater risk of developing the disorder.
Previous investigations on the FMR1 transcriptional landscape reported the highest expression of isoform Iso10/10b in premutation carriers with and without FXTAS15,16. However, this study shows a higher expression of these isoforms in individuals, in the converter group, who develop FXTAS over time. Similarly, we observed higher expression of the isoforms Iso4/4b, which encode for truncated proteins and are therefore missing the nuclear export signal and C-terminal RGG box, in the converter group, but not in the non-converter group, as compared to healthy controls, also suggesting the potential role of truncated proteins in the pathogenesis of FXTAS, which needs to be further investigated.
The midbrain, pons, and superior cerebellar peduncles (SCP) atrophy is the characteristic pathologic finding in patients with a number of neurodegenerative disorders including progressive supranuclear palsy (PSP)58,59, Parkinson's disease (PD), Multiple System Atrophy (MSA)49,60 and Alzheimer's disease61. These studies have proposed a number of midbrain metrics as potential biomarkers for differentiation of the patients with different neurodegenerative conditions. Shelton et al.39 suggested that the decreased MCP width observed in the premutation carriers who developed FXTAS over time (converters) could be a biomarker for early identification of incipient FXTAS. Of relevance, this study found a significant association of increased expression of Iso4/4b with decreased MCP width and of the expression of ASFMR1 isoform (131 bp) with decreased changes in pons, only in the converter group, which supports their potential role as biomarkers and support evidence of their potential contribution to the pathogenesis of the disorder. Interestingly, in the current study we found a positive association of FMR1 mRNA with mean SCP and midbrain and of isoform Iso10/10b with the midbrain, suggesting that some of the molecular measures may be linked with the changes in the brain structures. However, further studies are warranted to confirm these findings.
Generally, the mRNAs containing premature termination codons (PTCs) are degraded by the nonsense-mediated mRNA decay (NMD) system. However, this is not always the case as sometimes the mRNAs escape from the nonsense-mediated mRNA decay and results in truncated proteins62,63. Truncated proteins can form aggregates and can act in a dominant-negative manner. The accumulation of these abnormal truncated proteins leads to the gradual loss of function and structure of neurons including the death of neurons. These aggregations have been found to be associated with various neurodegenerative diseases such as Alzheimer’s, Huntington’s and Parkinson’s diseases, ASD, and amyotrophic lateral sclerosis. Additionally, a truncated form of DISC1 aggregates has been associated with major depression and schizophrenia64. Thus, it is possible that the translation of these isoforms might contribute to the pathogenesis of the FMR1 associated disorders. However, the exact mechanism and functional role are still unknown.
The previous report suggested that the antisense FMR1 (ASFMR1) gene and the premutation specific ASFMR1 Iso131 bp may contribute to the pathogenesis of FXTAS35. Although the expression of this splice isoform in premutation carriers was reported to be higher compared to controls, no difference in the expression levels was observed between non-FXTAS and FXTAS premutation carrier groups in previous studies36,37. The study presented here confirms the previous findings of a higher expression of this splice isoform in premutation carriers; however and importantly, the observed significant change of expression levels between V1 and V2 only in the converter group but not in the non-converter or control groups, suggest its potential role in the progression of FXTAS. These findings suggest the presence of an altered bidirectional transcription alternative splicing in premutation carriers and their potential role in the development and progression of FXTAS.
Finally, further studies are required to investigate the function of this specific isoform. The longitudinal follows up of these individuals may help our understanding of the potential role of such isoforms in FXTAS.
Conclusions
In conclusion, this is a first study that provides evidence of an elevated expression of some of the FMR1 and ASFMR1 mRNA splicing isoforms is present in premutation carriers who develop FXTAS compared to those who remain symptom free and to controls, providing support for these measures as potential biomarkers for early identification and monitoring of disease progression. In addition, the association of these molecular measures with brain measures provide us better insight regarding disease pathogenesis. However, due to limitation of the small sample size in the present study, further studies with larger sample size are required to replicate our initial findings and elucidate and confirm the role of these potential biomarkers.
Data Availbility
All statistical data generated during the study is available in the Additional File.xlsx titled “Supplementary Data”.
References
Hagerman, R. J. et al. Fragile X syndrome. Nat. Rev. Dis. Primers. 3, 17065 (2017).
Hall, D. A. & Berry-Kravis, E. Fragile X syndrome and fragile X-associated tremor ataxia syndrome. Handb. Clin. Neurol. 147, 377–391 (2018).
Fink, D. A. et al. Fragile X associated primary ovarian insufficiency (FXPOI): case report and literature review. Front Genet. 9, 529 (2018).
Jacquemont, S. Penetrance of the fragile X–associated tremor/ataxia syndrome in a premutation carrier population. JAMA. https://doi.org/10.1001/jama.291.4.460 (2004).
Tassone, F. et al. FMR1 CGG allele size and prevalence ascertained through newborn screening in the United States. Genom. Med. https://doi.org/10.1186/gm401 (2012).
Tassone, F. et al. Elevated levels of FMR1 mRNA in carrier males: a new mechanism of involvement in the fragile-X syndrome. Am. J. Hum. Genet. https://doi.org/10.1086/302720 (2000).
Hagerman, R. J. & Hagerman, P. Fragile X-associated tremor/ataxia syndrome — features, mechanisms and management. Nat Rev. Neurol. https://doi.org/10.1038/nrneurol.2016.82 (2016).
Ashley, C., Wilkinson, K., Reines, D. & Warren, S. FMR1 protein: conserved RNP family domains and selective RNA binding. Science. https://doi.org/10.1126/science.7692601 (1993).
Verheij, C. et al. Characterization and localization of the FMR-1 gene product associated with fragile X syndrome. Nature. https://doi.org/10.1038/363722a0 (1993).
Verkerk, A. J. M. H. et al. Alternative splicing in the fragile X gene FMR1. Hum. Mol. Genet. https://doi.org/10.1093/hmg/2.4.399 (1993).
Huang, T., Li, L.-Y., Shen, Y., Qin, X.-B., Pang, Z.-L., Wu, G.-Y. Alternative splicing of the FMR1 gene in human fetal brain neurons. Am. J. Med. Genet. https://doi.org/10.1002/(SICI)1096-8628(19960809)64:2<252::AID-AJMG3>3.0.CO;2-W (1996).
Sittler, A., Devys, D., Weber, C.-L. & Mandel, J. Alternative splicing of Exon 14 determines nuclear or cytoplasmic localisation of FMR1 protein isoforms. Hum. Mol. Genet. https://doi.org/10.1093/hmg/5.1.95 (1996).
Xie, W., Dolzhanskaya, N., LaFauci, G., Dobkin, C. & Denman, R. B. Tissue and developmental regulation of fragile X mental retardation 1 exon 12 and 15 isoforms. Neurobiol. Dis. 35, 52–62 (2009).
Brackett, D. M. et al. FMR1 transcript isoforms: association with polyribosomes; regional and developmental expression in mouse brain. PLoS ONE 8, e58296 (2013).
Pretto, D. I. et al. Differential increases of specificFMR1mRNA isoforms in premutation carriers. J. Med. Genet. https://doi.org/10.1136/jmedgenet-2014-102593 (2015).
Tseng, E., Tang, H.-T., AlOlaby, R.R., Hickey, L., Tassone, F. Altered expression of the FMR1 splicing variants landscape in premutation carriers. Biochim. Biophysica Acta (BBA) Gene Regul. Mech. https://doi.org/10.1016/j.bbagrm.2017.08.007. (2017).
Kelemen, O. et al. Function of alternative splicing. Gene 514, 1–30 (2013).
Su, C.-H., Dhananjaya, D. & Tarn, W.-Y. Alternative splicing in neurogenesis and brain development. Front. Mol. Biosci. https://doi.org/10.3389/fmolb.2018.00012 (2018).
Raj, B. & Blencowe, B. J. Alternative splicing in the mammalian nervous system: recent insights into mechanisms and functional roles. Neuron 87, 14–27 (2015).
Paronetto, M. P., Passacantilli, I. & Sette, C. Alternative splicing and cell survival: from tissue homeostasis to disease. Cell Death Differ. 23, 1919–1929 (2016).
Feng, D. & Xie, J. Aberrant splicing in neurological diseases. Wiley Interdiscipl. Rev. RNA. https://doi.org/10.1002/wrna.1184 (2013).
Cieply, B. & Carstens, R. P. Functional roles of alternative splicing factors in human disease. Wiley Interdiscipl. Rev. RNA. https://doi.org/10.1002/wrna.1276 (2015).
Quesnel-Vallières, M. et al. Misregulation of an activity-dependent splicing network as a common mechanism underlying autism spectrum disorders. Mol. Cell. 64, 1023–1034 (2016).
Cognata, V. L., La Cognata, V., D’Agata, V., Cavalcanti, F. & Cavallaro, S. Splicing: is there an alternative contribution to Parkinson’s disease?. Neurogenetics. https://doi.org/10.1007/s10048-015-0449-x (2015).
Hutton, M. et al. Association of missense and 5′-splice-site mutations in tau with the inherited dementia FTDP-17. Nature. https://doi.org/10.1038/31508 (1998).
Kashima, T., Rao, N. & Manley, J. L. An intronic element contributes to splicing repression in spinal muscular atrophy. Proc. Natl. Acad. Sci. https://doi.org/10.1073/pnas.0700343104 (2007).
Cassidy, S. B., Schwartz, S., Miller, J. L. & Driscoll, D. J. Prader-Willi syndrome. Genet. Med. https://doi.org/10.1038/gim.0b013e31822bead0 (2012).
Stefansson, H. et al. Large recurrent microdeletions associated with schizophrenia. Nature 455, 232–236 (2008).
Jiang, H., Mankodi, A., Swanson, M. S., Moxley, R. T. & Thornton, C. A. Myotonic dystrophy type 1 is associated with nuclear foci of mutant RNA, sequestration of muscleblind proteins and deregulated alternative splicing in neurons. Hum. Mol. Genet. https://doi.org/10.1093/hmg/ddh327 (2004).
Sreedharan, J. et al. TDP-43 Mutations in familial and sporadic amyotrophic lateral sclerosis. Science. https://doi.org/10.1126/science.1154584 (2008).
Yan, J. et al. Frameshift and novel mutations in FUS in familial amyotrophic lateral sclerosis and ALS/dementia. Neurology. https://doi.org/10.1212/wnl.0b013e3181f07e0c (2010).
Koch, L. Altered splicing in Alzheimer transcriptomes. Nat. Rev. Genet. https://doi.org/10.1038/s41576-018-0064-4 (2018).
Khalil, A. M., Faghihi, M. A., Modarresi, F., Brothers, S. P. & Wahlestedt, C. A novel RNA transcript with antiapoptotic function is silenced in fragile X syndrome. PLoS ONE 3, e1486 (2008).
Pastori, C. et al. Comprehensive analysis of the transcriptional landscape of the human FMR1 gene reveals two new long noncoding RNAs differentially expressed in Fragile X syndrome and Fragile X-associated tremor/ataxia syndrome. Hum. Genet. 133, 59–67 (2014).
Ladd, P. D. et al. An antisense transcript spanning the CGG repeat region of FMR1 is upregulated in premutation carriers but silenced in full mutation individuals. Hum. Mol. Genet. https://doi.org/10.1093/hmg/ddm293 (2007).
Olaby, R. R. A. et al. Assessment of molecular measures in non-FXTAS male premutation carriers. Front. Genet. https://doi.org/10.3389/fgene.2018.00302 (2018).
Vittal, P. et al. ASFMR1splice variant. Neurol. Genet. https://doi.org/10.1212/nxg.0000000000000246 (2018).
Loesch, D. Z. et al. Evidence for the toxicity of bidirectional transcripts and mitochondrial dysfunction in blood associated with small CGG expansions in the FMR1 gene in patients with parkinsonism. Genet. Med. 13, 392–399 (2011).
Shelton, A. L. et al. Middle cerebellar peduncle width—a novel MRI biomarker for FXTAS?. Front. Neurosci. https://doi.org/10.3389/fnins.2018.00379 (2018).
Ramnani, N. The primate cortico-cerebellar system: anatomy and function. Nat. Rev. Neurosci. 7, 511–522 (2006).
Emamzadeh, F. N. & Surguchov, A. Parkinson’s disease: biomarkers, treatment, and risk factors. Front. Neurosci. https://doi.org/10.3389/fnins.2018.00612 (2018).
Bacalman, S. et al. Psychiatric phenotype of the fragile X-associated tremor/ataxia syndrome (FXTAS) in males: newly described fronto-subcortical dementia. J. Clin. Psychiat. 67, 87–94 (2006).
Tassone, F., Pan, R., Amiri, K., Taylor, A. K. & Hagerman, P. J. A rapid polymerase chain reaction-based screening method for identification of all expanded alleles of the fragile X (FMR1) gene in newborn and high-risk populations. J. Mol. Diagnost. https://doi.org/10.2353/jmoldx.2008.070073 (2008).
Filipovic-Sadic, S. et al. A novel FMR1 PCR method for the routine detection of low abundance expanded alleles and full mutations in fragile X syndrome. Clin. Chem. 56, 399–408 (2010).
Tassone, F., Hagerman, R. J., Chamberlain, W. D., Hagerman, P. J. Transcription of the FMR1 gene in individuals with fragile X syndrome. Am. J. Med. Genet. https://doi.org/10.1002/1096-8628(200023)97:3<195::AID-AJMG1037>3.0.CO;2-R (2000).
Ardekani, B. A. & Bachman, A. H. Model-based automatic detection of the anterior and posterior commissures on MRI scans. NeuroImage. https://doi.org/10.1016/j.neuroimage.2009.02.030 (2009).
Mori, S., Crain, B. J., Chacko, V. P., Van Zijl, P. C. M. Three-dimensional tracking of axonal projections in the brain by magnetic resonance imaging. Ann. Neurol. p. 265–9. https://doi.org/10.1002/1531-8249(199902)45:2<265::AID-ANA21>3.0.CO;2-3 (1999).
Tustison, N. J., Avants, B. B., Cook, P. A., Zheng, Y., Egan, A., Yushkevich, P. A., et al. N4ITK: improved N3 bias correction. IEEE Trans. Med. Imaging. p. 1310–20. Available from: https://doi.org/10.1109/tmi.2010.2046908. (2010).
Quattrone, A. et al. MR imaging index for differentiation of progressive supranuclear palsy from parkinson disease and the parkinson variant of multiple system atrophy. Radiology. https://doi.org/10.1148/radiol.2453061703 (2008).
Nicoletti, G. et al. MR imaging of middle cerebellar peduncle width: differentiation of multiple system atrophy from parkinson disease. Radiology. https://doi.org/10.1148/radiol.2393050459 (2006).
Althouse, A. D. Adjust for multiple comparisons? It’s not that simple. Ann. Thorac. Surg. https://doi.org/10.1016/j.athoracsur.2015.11.024 (2016).
Hurlbert, S. H., Levine, R. A. & Utts, J. Coup de grâce for a tough old bull: “statistically significant” expires. Am. Stat. https://doi.org/10.1080/00031305.2018.1543616 (2019).
Pan, Q., Shai, O., Lee, L. J., Frey, B. J. & Blencowe, B. J. Deep surveying of alternative splicing complexity in the human transcriptome by high-throughput sequencing. Nat Genet. https://doi.org/10.1038/ng.259 (2008).
Ashley, C. T. et al. Human and murine FMR-1: alternative splicing and translational initiation downstream of the CGG–repeat. Nat. Genet. https://doi.org/10.1038/ng0793-244 (1993).
Ding, J., Huang, T., Li, L. & Fan, Y. Shen Y [Alternative splicing of the FMR1 gene in human fetal tissues]. Zhongguo Yi Xue Ke Xue Yuan Xue Bao 19, 241–246 (1997).
Banerjee, P. et al. Short- and long-term memory are modulated by multiple isoforms of the fragile X mental retardation protein. J. Neurosci. https://doi.org/10.1523/jneurosci.6369-09.2010 (2010).
Fu, X. et al. Alternatively spliced products lacking exon 12 dominate the expression of fragile X mental retardation 1 gene in human tissues. Mol. Med. Rep. https://doi.org/10.3892/mmr.2015.3574 (2015).
Saeed, U. et al. Imaging biomarkers in Parkinson’s disease and Parkinsonian syndromes: current and emerging concepts. Transl. Neurodegener. https://doi.org/10.1186/s40035-017-0076-6 (2017).
Mangesius, S. et al. MR planimetry in neurodegenerative parkinsonism yields high diagnostic accuracy for PSP. Parkinsonism Relat. Disord. 46, 47–55 (2018).
Whitwell, J. L. et al. Radiological biomarkers for diagnosis in PSP: Where are we and where do we need to be?. Mov. Disord. 32, 955–971 (2017).
Lee, J. H., Ryan, J., Andreescu, C., Aizenstein, H. & Lim, H. K. Brainstem morphological changes in Alzheimer’s disease. NeuroReport. https://doi.org/10.1097/wnr.0000000000000362 (2015).
Fernandes, R., Nogueira, G., da Costa, P. J., Pinto, F. & Romão, L. Nonsense-mediated mRNA decay in development, stress and cancer. mRNA Metabol. Hum. Disease. https://doi.org/10.1007/978-3-030-19966-1_3 (2019).
Jaffrey, S. R. & Wilkinson, M. F. Nonsense-mediated RNA decay in the brain: emerging modulator of neural development and disease. Nat. Rev. Neurosci.. https://doi.org/10.1038/s41583-018-0079-z (2018).
Korth, C. Aggregated proteins in schizophrenia and other chronic mental diseases. Prion. https://doi.org/10.4161/pri.18989 (2012).
Acknolwedgements
We thank the participants of the community‐based studies who donated their time and sample for this study. This paper is dedicated to the memory of Matteo. This study was funded by the National Institute of Health RO1NS110100-11/RO1MH078041, and grants of UL1 TR001860 and U54 HD079125.
Author information
Authors and Affiliations
Contributions
F.T., D.H. and S.M.R. designed the study; M.Z., H.-T.T. and E.F. performed the practical work and collected the data; B.D.-J. performed statistical analyses; M.Z. and F.T. drafted the manuscript. All listed authors have read and critically revised drafts for intellectual contents and provided full approval for publication.
Corresponding author
Ethics declarations
Competing interests
Dr. Flora Tassone received funding from Zynerba and Azrieli Foundation for carrying out studies in Fragile X Syndrome. Dr. David Hessl received the funding from Zynerba and Ovid Therapeutics regarding fragile X syndrome clinical trials. The other authors authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zafarullah, M., Tang, HT., Durbin-Johnson, B. et al. FMR1 locus isoforms: potential biomarker candidates in fragile X-associated tremor/ataxia syndrome (FXTAS). Sci Rep 10, 11099 (2020). https://doi.org/10.1038/s41598-020-67946-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-67946-y
This article is cited by
-
Language use predicts symptoms of fragile X-associated tremor/ataxia syndrome in men and women with the FMR1 premutation
Scientific Reports (2024)
-
Structure and Alternative Splicing of the Antisense FMR1 (ASFMR1) Gene
Molecular Neurobiology (2023)
-
Fmr1 exon 14 skipping in late embryonic development of the rat forebrain
BMC Neuroscience (2022)
-
Neuropsychological changes in FMR1 premutation carriers and onset of fragile X-associated tremor/ataxia syndrome
Journal of Neurodevelopmental Disorders (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.