Abstract
The emergence and spread of mobilized colistin resistance (mcr) genes have triggered extensive concerns worldwide. Here, we characterized the global distribution of mcr-9, a newly-identified variant of mcr, by assembling the data set of mcr-9-positive isolates from GenBank database and the literature available. Genetic features of all the mcr-9-harboring plasmids were determined by bioinformatic analysis. We showed that mcr-9 is globally distributed in 21 countries across six continents, with a wide dissemination among various species of Enterobacteriaceae strains from human, animal, food and environment. IncHI2-ST1 plasmids were found to be the predominant replicon type carrying mcr-9. Comparative genomics highlighted that IncHI2-type plasmids may also serve as a critical reservoir of mcr-9, from which different types of circulating plasmids acquired the mcr-9. Results revealed that the rcnR-rcnA-pcoE-pcoS-IS903-mcr-9-wbuC structure was consistent in most mcr-9 cassettes, suggesting a relatively unitary model involved in the mobilization of mcr-9. It is most likely that the spread of mcr-9 was mainly attributed to the conjugation and recombination events of mcr-9-carrying plasmids. In summary, our results provide a comprehensive picture of the distribution and genetic environment of mcr-9, and demonstrate the central roles played by IncHI2 plasmids in the worldwide dissemination of mcr-9.
Similar content being viewed by others
Introduction
Antibiotic resistance poses a great threat to global public health and carbapenem-resistant Enterobacteriaceae is triggering a health crisis worldwide1,2. Colistin, a cationic cyclic-peptide, is one of the last-resort antibiotics to defend against severe infections caused by carbapenem-resistant Enterobacteriaceae3. However, since the initial discovery of a plasmid-mediated mobilized colistin resistance gene (mcr-1) in China in late 20154, a number of diversified bacterial strains carrying mcr-1 have been detected across over 50 countries covering six continents5. The prevalent plasmid-borne MCR enzyme can catalyze chemical addition of phosphoethanolamine to lipid A moiety of bacterial lipopolysaccharides, the target of colistin, which consequently promotes colistin resistance6.
In recent years, a growing number of mcr-like genes (namely, from mcr-2 to mcr-10) have been identified7,8,9,10,11,12,13,14. These ongoing discoveries indicate a rapid evolution of MCR family under selective pressures, which raise global health concerns. mcr-9 is a newly emerging variant of the mobilized colistin resistance determinants, which was first identified in a clinical Salmonella enterica isolate in the USA in May, 201913. Since its initial identification, mcr-9 has been reported in several other countries, such as China15, Sweden16, and France17. Not only that, in silico analysis using the GenBank database indicated that mcr-9 had already been presented in a number of Enterobacteriaceae isolates recovered worldwide13,17. The high prevalence of mcr-9 suggests one more threat to public health. However, little information is available about the global epidemiology and dissemination patterns of mcr-9.
To explore these issues, here we studied the geographic and host distribution of mcr-9-carrying strains, and investigated the genomic features of various types of mcr-9-harboring plasmids from an extensive collection of publicly available sequence data sourced from the NCBI repository by bioinformatics analysis. Our findings may contribute to a better understanding of the prevalence and dissemination of the newly identified mcr-9 gene, and be helpful for developing better strategies to manage its spread.
Results and discussion
Geographic and isolation source of mcr-9
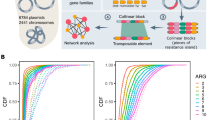
To obtain a comprehensive picture of the distribution of mcr-9, we blasted for mcr-9 in the NCBI GenBank database (as of 12th February 2020, n = 78) using a 99% identity cutoff and retrieved 71 complete plasmids and 6 chromosomes containing a full-length hit to mcr-9. A systematic review of the literature on mcr-9 published until 12 February 2020 resulted in the inclusion of 6 articles13,15,16,17,18,19. As a result, a total of 138 mcr-9-harboring strains were included for statistics. The statistical results showed that isolates carrying mcr-9 were identified in 21 countries across six continents (Fig. 1a), including Europe (n = 72; 52.2%), Asia (n = 27; 19.6%), North-America (n = 27; 19.6%), Oceania (n = 10; 7.2%), South-America (n = 1; 0.7%) and Africa (n = 1; 0.7%). The large number of mcr-9-positive isolates from Europe was notable, which can be largely ascribed to the inclusion of 30 isolates from horses in Sweden16 and 28 from human in Europe by a previous clinical surveillance program18. Anyway, the high incidence of mcr-9 in Europe is alarming, which demands more attention. In addition, the prevalence of mcr-9 around the world might be underestimated, due to the limited availability of its epidemiological investigations. Large-scale surveillance and molecular epidemiological studies are urgently required to better understand the global distribution and dissemination of mcr-9, thereby facilitating establishment of effective treatments to control its spread.
Overview of the distribution of mcr-9. (a) Worldwide distribution of mcr-9-positive isolates. The map was generated using the online software dituhui (https://g.dituhui.com/). (b) Distribution of host species harboring mcr-9. (c) Distribution of the isolation source of mcr-9-positive strains.
According to the current data, various kinds of Enterobacteriaceae isolates were found to disseminate mcr-9, among which Enterobacter spp. were the most common ones (n = 51, 37.0%), followed by Klebsiella spp. (n = 41, 30.0%), Salmonella spp. (n = 14, 10.1%), Escherichia spp. (n = 13, 9.4%), Citrobacter spp. (n = 9, 6.5%), Leclercia spp. (n = 6, 4.3%), Cronobacter spp. (n = 2, 1.4%), Raoultella spp. (n = 1, 0.7%), and Phytobacter spp. (n = 1, 0.7%) (Fig. 1b). Of these mcr-9 isolates, 77 were of human origin (55.8%), 40 of animal origin (29.0%), 5 of environment origin (3.6%) and 4 strains (2.9%) were isolated from food. The source for 12 isolates (8.7%) could not be obtained for an incomplete strain information (Fig. 1c). These data point towards a widespread dissemination of mcr-9 among clinically important pathogens, which has serious implications on clinical therapies and public health. Active surveillance for clinical discovery of mcr-9-harboring pathogens is needed to provide a clue to defend against infections by superbugs with colistin resistance.
Overall landscape of the mcr-9-carrying plasmids
Seventy-one mcr-9-carrying plasmids with complete sequences were retrieved from the GenBank database (accessed on 12th February 2020, Table S1). Among them, IncHI2 was the dominant replicon type carrying mcr-9, accounting for 90.1% (64/71) of the plasmids. Of the 64 IncHI2 mcr-9-harboring plasmids, 60 plasmids had the IncHI2 replicon alone, 3 had IncHI2 plus IncR, and 1 had IncHI2 plus IncA/C2, with plasmid size varying from 222 kb to 477 kb. We found that mcr-9 plasmids with IncHI2 replicon alone had a worldwide distribution, while plasmids containing both IncHI2 and IncR replicons had only been recovered from China, suggesting that IncHI2-IncR hybrid plasmids may represent another important vehicle in mediating the dissemination of mcr-9 in China. Phylogenetic analysis showed that three IncHI2-IncR hybrid plasmids, pMCR-SCNJ07 (accession no. MK933279), pT5282-mphA (KY270852) and pN1863-HI2 (MF344583) were clustered with several IncHI2 plasmids (Fig. 2), implying that they may have diverged from a common ancestor and a subsequent genetic recombination event leading to the formation of hybrid plasmids. And, the IncHI2-IncA/C2 hybrid plasmid pGMI14-002_1 (accession no. CP028197), which was recovered from a Salmonella enterica isolate in Czech Republic in 2018, followed a similar pattern. From the phylogenetic tree, we also found that mcr-9-harboring plasmids recovered from human isolates were interspersed with plasmids from animals and the environment (Fig. 2), indicating that plasmid-borne mcr-9 might have circulated in the entire ecosystem.
Phylogenetic analysis of IncHI2 mcr-9-harboring plasmids. Core-genome alignments and phylogenetic reconstruction of IncHI2 mcr-9-carrying plasmids were performed using Parsnp. The heatmap denotes the presence of ESBL or carbapenemase genes as determined by ABRicate. The annotation denotes (from left to right) locations, host species, isolation sources, and sequence types of plasmids. NA, not available. ND, not determined.
IncHI2-type plasmids have a broad host range, which represent one of the most commonly encountered plasmid groups in the Enterobacteriaceae family20. Among them, IncHI2-ST1 plasmids are known to disseminate clinically important antimicrobial resistance genes and are most frequently reported to play a critical role in the evolution of complex resistance phenotypes within disease-causing strains of Enterobacteriaceae21. Our presentation here highlighted that IncHI2-ST1 plasmids are a major vehicle carrying mcr-9 around the world using the pMLST tool. In addition to IncHI2 plasmids, there was one IncFII (FIIY3: FIA-: FIB-) plasmid p1_045523 (accession no. CP032893) carrying mcr-9. And, the replicon type for 6 mcr-9-bearing plasmids could not be determined, with plasmid size ranging from 56 kb to 133 kb. These findings revealed that multiple types of plasmids are involved in the spread of mcr-9.
IncHI2 plasmids are known carriers of determinants for resistance not only to antibiotics such as β-lactams, quinolones and aminoglycosides, but also to heavy metals such as copper, silver ions, and mercuric ions22. Analysis of the resistance genes revealed that IncHI2 mcr-9-carrying plasmids contained between 0 and 5 additional ESBL or carbapenemase genes (Table S1). Among these resistance genes, blaSHV-12 was the most common one (32/71, 45.1%) that co-existed with mcr-9, followed by blaTEM-1 (29/71, 40.8%). In particular, 23 out of the 64 (35.9%) IncHI2 mcr-9-carrying plasmids harbored one of the carbapenemase genes, namely blaIMP- [1, 4, 8, 26, 70, 79], blaKPC-4, blaNDM-1, blaVIM- [1,4], and blaOXA-436. Co-transfer of the colistin resistance and carbapenemase genes by a single plasmid could raise the risk of dissemination of these resistance genes, which is of great concern for clinical therapies.
Genetic characterization of plasmids carrying mcr-9
Comparative analysis was performed to obtain a comprehensive view of the genetic features of 60 IncHI2-type mcr-9-bearing plasmids (hybrid plasmids were excluded). Results showed that these plasmids were diverse in terms of genetic structure, except that tra1 and tra2 regions as well as the ter region were conserved among all the plasmids (Fig. 3a). mcr-9 was located in the sil-cop region of IncHI2-type plasmids. In this accessory resistance region, the structure and genetic content immediately upstream of mcr-9 were consistent among IncHI2 mcr-9-bearing plasmids. While, genes downstream of mcr-9 were genetically diverse, with silver resistance determinants and qseB/qseC two-component regulators being absent in most plasmids (Fig. 3a,b). The wbuC gene, which was proposed to have transferred together with mcr-9 as a whole fragment from Buttiauxella spp.17, was lacking downstream of mcr-9 in some plasmids (Fig. 3a,b).
Genetic characterization of IncHI2-type mcr-9-harboring plasmids (hybrid plasmids are excluded). (a) Sequence alignment of 60 IncHI2 mcr-9 plasmids. pRH-R27(accession no. LN555650) was used as a reference to compare with other plasmids. Gaps indicate regions that were missing in the respective plasmid compared to the reference plasmid. The outer circle with dark blue arrows denotes annotation of reference plasmid. mcr-9 gene is highlighted by red arrows. Backbone (tra1 and tra2 region) and four accessory resistance regions (ter region, the sil–cop region, MDR region 1 and MDR region 2) are indicated by red curves. The mcr-9-harboring region is indicated by the dotted box. Information about the IncHI2 plasmids tested is provided in Table S1. (b) Linear comparison of the mcr-9-harboring region in pRH-R27, p17277A_477 (CP043927) and pMRVIM0813 (KP975077). The corresponding region on non-mcr-9-carrying plasmid R478 (BX664015, top) is shown for comparison. Grey shading denotes regions of shared homology among different plasmids ranging from 80% to100%. Colored arrows represent open reading frames, with brown, green, and red arrows representing heavy metal resistance genes, mobile elements, and the mcr-9 gene, respectively. Other plasmid backbone genes and hypothetical genes are colored gray.
We further explored the genetic structures of the IncHI2-IncR mcr-9-carrying plasmids, pMCR-SCNJ07, pT5282-mphA and pN1863-HI2, by comparing them with the IncHI2 reference plasmid pRH-R27 and IncR reference plasmid pHN84KPC (accession no. KY296104). Results showed that 1) three IncHI2-IncR hybrid plasmids had similar genetic structures, 2) the hybrid plasmids showed high similarity to backbone elements of IncHI2 on a large scale (>77% query coverage, 99.99% nucleotide identity), 3) the hybrid plasmids showed low sequence homology (<9% coverage) with the IncR reference plasmid, and 4) mcr-9 was located on the IncHI2 plasmid region (Fig. 4a). These findings suggested that multiple recombination events involving several plasmids probably contributed to the integration of foreign regions into mcr-9-bearing IncHI2 backbones. In a similar way, IncHI2-IncA/C2 hybrid plasmid pGMI14-002_1 was formed by fusion of a mcr-9-bearing IncHI2 plasmid and an IncA/C2 plasmid, like pR55 (accession no. JQ010984) (Fig. 4c). Sequence comparison of the mcr-9-harboring regions showed that the flanked upstream regions of mcr-9 in IncHI2-IncR hybrid plasmids were highly similar (>93% coverage,>99.98% identity) to that of the IncHI2 reference plasmid pRH-R27, except that pcoS was truncated by an insertion element in pMCR-SCNJ07 and pT5282-mphA (Fig. 4b). Whereas, in comparison to pRH-R27, IncHI2-IncR plasmids lacked the sil resistance module downstream of mcr-9, with only silR survived, implying a relic for frequent recombination events (Fig. 4b). In the IncHI2-IncA/C2 plasmid, the sil resistance module, the qseC-qseB regulatory genes, and the insertion elements between them were all missing from the downstream of mcr-9, while the neighboring genes upstream of mcr-9 (from orf to IS903) were identical to those in pRH-R27 (Fig. 4d).
Genetic characterization of mcr-9-harboring hybrid plasmids. (a) Comparison of the IncHI2-IncR hybrid plasmids, pMCR-SCNJ07, pT5282-mphA and pN1863-HI2, with the IncHI2 plasmid pRH-R27 and IncR plasmid pHN84KPC. pN1863-HI2 was used as a reference to compare with other plasmids. (b) Linear comparison of the mcr-9-harboring region in pRH-R27, pMCR-SCNJ07, pT5282-mphA and pN1863-HI2. (c) Comparison of the IncHI2-IncA/C2 hybrid plasmid, pGMI14-002_1 with the IncHI2 plasmid pRH-R27 and IncA/C2 plasmid pR55. pGMI14-002_1 was used as a reference to compare with other plasmids. (d) Linear comparison of the mcr-9-harboring region in pRH-R27 and pGMI14-002_1. Gaps in the circular maps refer to plasmid regions that were missing in the respective plasmid compared to the reference plasmid. The outer circle with dark blue arrows denotes annotation of reference plasmid. mcr-9 gene is highlighted by red arrows. The foreign region that was proposed to be an insertion was indicated by red curves. The mcr-9-harboring region is indicated by the dotted box. Grey shading in the linear maps denotes regions of shared homology among different plasmids ranging from 80% to 100%. Colored arrows represent open reading frames, with brown, green, and red arrows representing heavy metal resistance genes, mobile elements, and the mcr-9 gene, respectively. Other plasmid genes are colored gray.
BLASTn of the IncFII mcr-9-carrying plasmid p1_045523 against the NCBI’s nr database identified pOZ172 (accession no. CP016763) as the highest-scoring hit, with only 50% coverage and 99.96% identity, suggesting that p1_045523 was novel in structure. Analysis of the genetic structure showed that p1_045523 contained backbone elements of the IncFII plasmid pOZ172 that encoded functions for plasmid replication and horizontal transfer, and also harbored two additional resistance gene regions (13.2–29.7 kb and 37.6–64.0 kb) displaying high similarity to the IncHI2 reference plasmid pRH-R27, with 97.80% and 99.55% identity, respectively (Fig. 5a). It was noted that mcr-9 was exactly carried on one of the resistance regions. Comparison of the mcr-9-harboring regions between p1_045523 and pRH-R27 revealed that they shared regions flanking mcr-9 (from dcm to qseB) with 100% coverage and 98.88% identity (Fig. 5b). The genetic environment of mcr-9 in the six untyped mcr-9-bearing plasmids was investigated by comparing them with the IncHI2 plasmid pRH-R27. Results showed that an mcr-9-harboring region (from dcm to wbuC) was consistent among them, except in the plasmid pLEC-b38d, in which only the mcr-9-wbuC module retained (Fig. 5c,d). Our findings suggested that these untyped plasmids (excluding pLEC-b38d) were most likely to have acquired the mcr-9 gene from IncHI2-type plasmids by multiple recombination events, and pLEC-b38d might have evolved earlier with the possibility of different patterns for the acquisition and dissemination of mcr-9. In addition, mcr-9 was also found to have a chromosomal location in 6 Enterobacteriaceae strains (Fig. S1), with identical core structure of mcr-9 cassettes to the IncHI2 plasmid pRH-R27, implying a relic of recombination/integration between the plasmid and chromosome.
Genetic characterization of IncFII-type mcr-9-carrying plasmid p1_045523 and six untyped mcr-9-carrying plasmids. (a) Comparison of the p1_045523 with the IncHI2 plasmid pRH-R27 and IncFII plasmid pOZ172. p1_045523 was used as a reference to compare with other plasmids. (b) Linear comparison of the mcr-9-harboring region in pRH-R27 and p1_045523. (c) Comparative genome maps of pCAV1099-114 (accession no. CP011596), pCAV1335-115 (CP011617), pCAV1015-114 (CP017930), pSHV12-1301491 (CP031568), pCFSAN002050 (CP006057), pLEC-b38d (CP026168) and the IncHI2 plasmid pRH-R27. pCAV1099-114 was used as a reference to compare with other plasmids. (d) Linear comparison of the mcr-9-harboring region in six untyped mcr-9-carrying plasmids and pRH-R27. The outer circle with dark blue arrows denotes annotation of reference plasmid. Gaps in the circular maps refer to plasmid regions that were missing in the respective plasmid compared to the reference plasmid. mcr-9 gene is highlighted by red arrows. The mcr-9-harboring region is indicated by the dotted box. Colored arrows represent open reading frames, with brown, green, and red arrows representing heavy metal resistance genes, mobile elements, and the mcr-9 gene, respectively. The remaining genes are shown in gray. Grey shading in the linear maps denotes regions of shared homology among different plasmids ranging from 80% to 100%.
Analysis on the plasmid reservoirs of mcr-9 gene highlighted a leading role of IncHI2-type plasmid in the dissemination of mcr-9. Searches for IncHI2 mcr-harboring plasmids deposited in GenBank suggested that IncHI2 plasmids also served as an important vehicle in carrying mcr-1 and mcr-3. Unlike the scenario seen with the mcr-1 or mcr-3, which was inserted into different genetic loci in IncHI2 plasmids, mcr-9 was consistently located in the sil-cop region (Figs. S2, 3a). These results indicated that mcr-9 was inactive during dynamic gene transposition events, suggesting that transposon elements of mcr-9 could be lost soon after the mobilization of mcr-9 onto the IncHI2 backbone.
To further characterize the genetic contexts of mcr-9, representative mcr-9-haboring regions from different types of plasmids and chromosomes were subjected to comparison analysis. mcr-9 was found in a variety of genetic contexts, which were differed by the truncation status of neighboring genes (from rcnR to qseB) encompassing mcr-9. Results also revealed that the core structure of all known mcr-9 cassettes was rcnR-rcnA-pcoE-pcoS-IS903-mcr-9-wbuC (Fig. 6), and it was identical in most (86.0%, 61/71) of the study plasmids, indicating that these core elements encompassing mcr-9 are most likely to be required for the evolution of mcr-9 cassettes. We failed to identify potential transposon elements (intact or truncated) mediating the transfer of mcr-9, posing the high possibility that conjugation and recombination events of mcr-9-carrying plasmids are mainly responsible for the spread of mcr-9. The lacking of the downstream two-component regulatory genes qseC and qseB, which were proposed to be involved in the induction of colistin resistance mediated by mcr-9, was commonly observed among the mcr-9-haboring plasmids (Figs. 3–6). More research should be performed to confirm the essential role of qseC-qseB module or the involvement of other genes in mcr-9 induction.
A summary of representative genetic contexts of mcr-9 from different types of plasmid and chromosomes available to date. The genetic contexts of mcr-9 were differed by the truncation status of neighboring genes (from rcnR to qseB) encompassing mcr-9. The ‘rcnR-rcnA-pcoE-pcoS-IS903-mcr-9-wbuC’ structure was revealed as the core structure of mcr-9 cassettes by comparison analysis. Colored arrows represent open reading frames, with brown, green, and red arrows representing heavy metal resistance genes, mobile elements, and the mcr-9 gene, respectively. The remaining genes are shown in gray. Grey shading denotes genetic regions that exhibit sequence homology among different segments.
Conclusion
In conclusion, we assembled the data set to date of mcr-9-positive sequenced plasmids and chromosomes through an exhaustive search of publicly available databases. This study allowed us to obtain a clear picture of the global distribution of mcr-9-harboring isolates, which covered 21 countries across six continents and were distributed in at least 9 Enterobacteriaceae species of diverse origins. The co-transfer of mcr-9 and carbapenemase genes in clinically important pathogens posed a high risk to the public health and clinical treatment. IncHI2-ST1 plasmid was found to be the predominant replicon type carrying mcr-9, implying that IncHI2-ST1 plasmids may be a major vehicle in mediating the dissemination mcr-9. Besides, our results extended significantly our understanding of genetic features of different types of mcr-9-bearing plasmids, and highlighted that IncHI2-type plasmids serve as a critical reservoir of mcr-9. In addition, the highly similarity of mcr-9 cassette in different plasmids posed the possibility that the dissemination of mcr-9 across bacterial populations was mainly mediated by conjugation and recombination events of mcr-9-carrying plasmids. A limitation of this study lies in that the phenotypes of the mcr-9-harboring strains are not available, which makes it difficult to correlate phenotypic with genotypic features. Whole-genome sequencing based surveillance and epidemiological studies are needed to achieve better insights into the genetic features of mcr-9-positive strains, which might eventually allow us to develop effective strategies to manage their spread.
Materials and Methods
Retrieval of genomic data and literature search
To obtain the mcr-9-harboring genome sequences, a nucleotide BLAST with standard options was performed in the NCBI GenBank database with the nucleotide base sequence of mcr-9 as a query (GenBank accession no. MK791138). Models and uncultured environmental samples were excluded. Complete plasmids and chromosomes containing at least one contig with a full-length hit to mcr-9 with 100% query coverage and at least 99% identity were selected and exported. Information about geographic distribution and isolation source of the mcr-9-positive strains were investigated. Additionally, relevant papers that published on mcr-9 were identified in PubMed and Google Scholar using the query ‘mcr-9 AND colistin’, and non-repetitive information of the geographic and isolation source of mcr-9 strains were included in the statistics.
Phylogenetic analysis
The complete sequences of all available mcr-9-carrying plasmids as of 12th February 2020 were retrieved from the GenBank. The plasmid replicon type and MLST were determined using the Plasmidfinder (https://cge.cbs.dtu.dk/services/PlasmidFinder/) and pMLST (https://cge.cbs.dtu.dk/services/pMLST/). ResFinder (http://genomicepidemiology.org/) was employed for the identification of antimicrobial resistance genes. All the IncHI2 mcr-9-carrying plasmids were used for phylogenetic analysis using Parsnp in the Harvest package23, and the phylogenetic tree was visualized by iTOL (https://itol.embl.de/).
Comparative analysis of mcr-9-harboring genome sequences
The annotations of mcr-9-carrying plasmids were performed using Prokka24 combined with BLASTp/BLASTn searches against the NCBI database. Alignments among highly homologous complete plasmid sequences were performed using BLAST and visualized with the BRIG tool25. Sequence alignments among mcr-9-carrying plasmids or chromosomes were performed using BLAST and visualized with Easyfig v 2.2.326.
References
Laxminarayan, R. et al. UN High-Level Meeting on antimicrobials—what do we need? Lancet 388, 218–220, https://doi.org/10.1016/s0140-6736(16)31079-0 (2016).
Perez, F. & Bonomo, R. A. Carbapenem-resistant Enterobacteriaceae: global action required. The Lancet Infectious Diseases, https://doi.org/10.1016/s1473-3099(19)30210-5 (2019).
Poirel, L., Jayol, A. & Nordmann, P. Polymyxins: Antibacterial Activity, Susceptibility Testing, and Resistance Mechanisms Encoded by Plasmids or Chromosomes. Clin. Microbiol. Rev. 30, 557–596, https://doi.org/10.1128/CMR.00064-16 (2017).
Liu, Y. Y. et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: a microbiological and molecular biological study. Lancet Infect. Dis. 16, 161–168, https://doi.org/10.1016/S1473-3099(15)00424-7 (2016).
Zhang, H. et al. A Genomic, Evolutionary, and Mechanistic Study of MCR‐5 Action Suggests Functional Unification across the MCR Family of Colistin Resistance. Advanced Science, 1900034, https://doi.org/10.1002/advs.201900034 (2019).
Sun, J., Zhang, H., Liu, Y. H. & Feng, Y. Towards Understanding MCR-like Colistin Resistance. Trends Microbiol. 26, 794–808, https://doi.org/10.1016/j.tim.2018.02.006 (2018).
Xavier, B. B. et al. Identification of a novel plasmid-mediated colistin-resistance gene, mcr-2, in Escherichia coli, Belgium, June 2016. Euro Surveill 21, https://doi.org/10.2807/1560-7917.ES.2016.21.27.30280 (2016).
Liu, L., Feng, Y., Zhang, X., McNally, A. & Zong, Z. New Variant of mcr-3 in an Extensively Drug-Resistant Escherichia coli Clinical Isolate Carrying mcr-1 and bla NDM-5. Antimicrob Agents Chemother 61, https://doi.org/10.1128/AAC.01757-17 (2017).
Carattoli, A. et al. Novel plasmid-mediated colistin resistance mcr-4 gene in Salmonella and Escherichia coli, Italy 2013, Spain and Belgium, 2015 to 2016. Euro Surveill 22, https://doi.org/10.2807/1560-7917.ES.2017.22.31.30589 (2017).
Borowiak, M. et al. Identification of a novel transposon-associated phosphoethanolamine transferase gene, mcr-5, conferring colistin resistance in d-tartrate fermenting Salmonella enterica subsp. enterica serovar Paratyphi B. J. Antimicrob. Chemother. 72, 3317–3324, https://doi.org/10.1093/jac/dkx327 (2017).
Yang, Y. Q., Li, Y. X., Lei, C. W., Zhang, A. Y. & Wang, H. N. Novel plasmid-mediated colistin resistance gene mcr-7.1 in Klebsiella pneumoniae. J Antimicrob Chemother, https://doi.org/10.1093/jac/dky111 (2018).
Wang, X. et al. Emergence of a novel mobile colistin resistance gene, mcr-8, in NDM-producing Klebsiella pneumoniae. Emerg. Microbes Infect. 7, 122, https://doi.org/10.1038/s41426-018-0124-z (2018).
Laura, M. Carroll, a. & Ahmed Gaballa, a. C. G., b Genevieve Sullivan,Lory O. Henderson,a,c Martin Wiedmanna. Identification of Novel Mobilized Colistin Resistance Gene mcr-9 in a Multidrug-Resistant, Colistin-Susceptible Salmonella enterica Serotype Typhimurium Isolate. MBio 10, e00853–00819, https://doi.org/10.1128/mBio (2019).
Wang, C. et al. Identification of novel mobile colistin resistance gene mcr-10. Emerg. Microbes Infect. 9, 508–516, https://doi.org/10.1080/22221751.2020.1732231 (2020).
Yuan, Y. et al. Coproduction Of MCR-9 And NDM-1 By Colistin-Resistant Enterobacter hormaechei Isolated From Bloodstream Infection. Infect. Drug. Resist. 12, 2979–2985, https://doi.org/10.2147/IDR.S217168 (2019).
Borjesson, S. et al. A link between the newly described colistin resistance gene mcr-9 and clinical Enterobacteriaceae isolates carrying bla SHV-12 from horses in Sweden. J Glob Antimicrob Resist, https://doi.org/10.1016/j.jgar.2019.08.007 (2019).
Kieffer, N. et al. mcr-9, an inducible gene encoding an acquired phosphoethanolamine transferase in Escherichia coli, and its origin. Antimicrob Agents Chemother, https://doi.org/10.1128/AAC.00965-19 (2019).
Wang, Y. et al. Detection of mobile colistin resistance gene mcr-9 in carbapenem-resistant Klebsiella pneumoniae strains of human origin in Europe. J Infect, https://doi.org/10.1016/j.jinf.2019.12.016 (2019).
Chavda, K. D. et al. First Report of bla VIM-4-and mcr-9-Coharboring Enterobacter Species Isolated from a Pediatric Patient. msphere 4, e00629–00619 (2019).
Fang, L.-X. et al. High genetic plasticity in multidrug-resistant sequence type 3-IncHI2 plasmids revealed by sequence comparison and phylogenetic analysis. Antimicrobial Agents Chemotherapy 62, e02068–02017 (2018).
Roy Chowdhury, P. et al. Identification of a novel lineage of plasmids within phylogenetically diverse subclades of IncHI2-ST1 plasmids. Plasmid 102, 56–61, https://doi.org/10.1016/j.plasmid.2019.03.002 (2019).
Liang, Q. et al. Sequencing and comparative genomics analysis of the IncHI2 plasmids pT5282-mphA and p112298-catA and the IncHI5 plasmid pYNKP001-dfrA. Int. J. Antimicrob. Agents 49, 709–718, https://doi.org/10.1016/j.ijantimicag.2017.01.021 (2017).
Treangen, T. J., Ondov, B. D., Koren, S. & Phillippy, A. M. The Harvest suite for rapid core-genome alignment and visualization of thousands of intraspecific microbial genomes. Genome Biol. 15, 524 (2014).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Alikhan, N.-F., Petty, N. K., Zakour, N. L. B. & Beatson, S. A. BLAST Ring Image Generator (BRIG): simple prokaryote genome comparisons. BMC genomics 12, 402 (2011).
Sullivan, M. J., Petty, N. K. & Beatson, S. A. Easyfig: a genome comparison visualizer. Bioinformatics 27, 1009–1010 (2011).
Acknowledgements
This work was supported by National Natural Science Foundation of China (31900125), the Science and Technology Strategic Cooperation Programs of Luzhou Municipal People’s Government and Southwest Medical University (2019LZXNYDJ47), Natural Science Foundation of Southwest Medical University (2018-ZRZD-011).
Author information
Authors and Affiliations
Contributions
L.Z. designed the study. X.D. and J.Z. collected the data. Y.G. and Z.Z. analyzed and interpreted the data. Y.L. wrote the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, Y., Dai, X., Zeng, J. et al. Characterization of the global distribution and diversified plasmid reservoirs of the colistin resistance gene mcr-9. Sci Rep 10, 8113 (2020). https://doi.org/10.1038/s41598-020-65106-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-65106-w
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.