Abstract
Insomnia is one of the most prevalent and burdensome mental disorders worldwide, affecting between 10–20% of adults and up to 48% of the geriatric population. It is further associated with substance usage and dependence, as well other psychiatric disorders. In this study, we combined electronic health record (EHR) derived phenotypes and genotype information to conduct a genome wide analysis of insomnia in a 18,055 patient cohort. Diagnostic codes were used to identify 3,135 patients with insomnia. Our genome-wide association study (GWAS) identified one novel genomic risk locus on chromosome 8 (lead SNP rs17052966, p = 4.53 × 10−9, odds ratio = 1.28, se = 0.04). The heritability analysis indicated that common SNPs accounts for 7% (se = 0.02, p = 0.015) of phenotypic variation. We further conducted a large-scale meta-analysis of our results and summary statistics of two recent insomnia GWAS and 13 significant loci were identified. The genetic correlation analysis yielded a strong positive genetic correlation between insomnia and alcohol use (rG = 0.56, se = 0.14, p < 0.001), nicotine use (rG = 0.50, se = 0.12, p < 0.001) and opioid use (rG = 0.43, se = 0.18, p = 0.02) disorders, suggesting a significant common genetic risk factors between insomnia and substance use.
Similar content being viewed by others
Introduction
Insomnia is a highly prevalent sleep disorder characterized by the inability to fall asleep or maintain sleep1 and affects 10–20% of the adult population2,3. It is characterized by heterogeneous phenotypes and equifinality, which might reflect different underlying causal mechanisms4, including life style, stress and molecular mechanisms (for a review, see5). It is commonly comorbid with other physical and psychiatric disorders6,7.
Genetic contributions to insomnia have been demonstrated in both family and twin studies with the reported heritability being estimated at 25–45%8. Candidate gene studies have highlighted genetic variants in numerous systems including the circadian gene CLOCK9, the GABAergic system10, the adenosinergic system11, and the serotonergic system12.
A number of genome-wide association studies (GWAS) have been conducted examining the insomnia phenotype. In two recent studies, large-scale cohorts were developed using data from UK Biobank and the combination of UK Biobank and 23andMe yielding 57 and 202 significant loci, respectively13,14. Another study using survey data of soldiers in the Army Study To Assess Risk and Resilience in Servicemembers (STARRS) study identified one significant locus15. These studies also identified genetic correlations between insomnia and various clinical conditions, such as schizophrenia, type 2 diabetes, and depression13,15. Other studies have identified several insomnia related genes, such as CACNA1C16, RBFOX317, PAX818 and MEIS119.
In most previous studies, insomnia phenotypes were assessed through self-report, which could miss useful information and reflect only part of disorder status. Since insomnia can be a chronic process with different trajectories and multiple complications in clinical settings, it is important to conduct studies specifically targeting clinical patient populations20. Because of complex underlying mechanisms of insomnia and its various clinical manifestations, obtaining a clinically well-defined subject cohort is critical for genetic association analysis. Electronic health records (EHRs) from large medical institutes comprise a uniquely valuable data source to help identify genetic associations within very specific clinical conditions21.
In this study, we utilized a large-scale clinical database to explore the genetic underpinnings of insomnia and calculated the genetic correlation between insomnia and various clinical conditions. Further, we conducted a meta-analysis of our results combined with recent insomnia GWAS to discover novel genomic loci.
Methods
Clinical database
All the clinical data and genetic data in this study were obtained from the Partners Biobank22. The Partners Biobank is a large integrated database which contains clinical data from Partners HealthCare for approximately 90,000 consented patients, and genomic data for approximately 25,000 of them. The clinical data including patient family history, demographic information, diagnosis, medication records, lab test results and clinical notes. The clinical data is derived from the electronic health records, which have been collecting patient data since 1990. The informed consent was obtained from all study participants and/or their legal guardians. The study’s protocol was reviewed and approved by Partners Human Research Committee. All methods were performed in accordance with the relevant guidelines and regulations.
Electronic health record-derived phenotypes
We generated an ICD 9 and ICD10 code list for insomnia, three major substance use disorders and a series of relevant clinical conditions, including multiple psychiatric disorders and type 2 diabetes, then used these codes to identify our case cohort (Supplementary Table 1).
The ICD codes of insomnia include the following definitions
307.4*: specific disorders of sleep of nonorganic origin; 327.0*: organic disorders of initiating and maintaining sleep; 780.51: insomnia with sleep apnea, unspecified; 780.52: insomnia, unspecified; G47.0*: insomnia; F51.0*: insomnia not due to a substance or known physiological condition.
We reviewed 15,750,104 diagnosis records, which were collected between 1991 and 2018, to identify patients meeting our insomnia phenotype definition. The control cohort consisted of patients not meeting the insomnia phenotype, and also excluded patients with any other kind of sleep disorders, including snoring, periodic limb movement, sleep related leg cramps, sleep related bruxism and hypersomnia.
For the three substance use disorders, the case cohort included patients with at least one corresponding ICD code of substance dependence, substance abuse or long-term substance use disorder. The control group consisted of 12,205 patients without any record of substance use disorder (nicotine, alcohol, opioid, cannabis, cocaine and amphetamine).
Genotyping, imputation and quality control
The genotyping was performed by Partners Biobank using the Illumina Multi-Ethnic Global (MEG) array (Illumina, Inc., San Diego, CA) including 1,779,763 SNPs. Prior to imputation, QC steps were conducted, including: a. sample-level filtration: any samples with a discrepancy between the reported and predicted sex were removed. b. SNP-level filtration: removal of sites with invalid alleles, duplicate, monomorphic, indel, allele mismatch, low call rate (less than 90%). The SNPs that were not in the reference panel were also removed. The imputation was performed using the Michigan Imputation Server with Minimac323. The HRC (Version r1.1 2016) reference panel consisting of 64,940 haplotypes of predominantly European ancestry was used24.
Post-imputation quality control was conducted to select high-quality SNPs and control for population stratification. In all analyses, only autosomal biallelic SNPs with minor allele frequencies (MAF) of at least 1%, an info score above 0.8 and call rates above 98% were retained, which led to 5,508,534 SNPs. The present analysis included only individuals of European ancestry, which were reported by patients, to minimize the risk for confounding due to ancestry differences. A principal components analysis (PCA) was applied to characterize population structure.
Statistical analysis
PLINK 1.90 was used to conduct the genome-wide association analysis, adjusted for age, sex and the top 10 principal components25.The Genome-based restricted maximum likelihood (GREML) method implemented in GCTA was used to estimate the percentage of variance explained by common SNPs and calculate the genetic correlations26,27. LD Score Regression (LDSC) was used to calculate the genetic correlations between our results and publicly available GWAS studies28. FUMA and MAGMA were used to conduct the gene-based test and pathway enrichment analysis29. METAL was used for the meta-analysis between our results and published insomnia GWAS30.
A standard genome-wide significance threshold of p < 5 × 10−8 was chosen for SNP identification and r2 = 0.6 was set as the cutoff to define LD block. All phenotyping analyses were conducted using R (version 3.3.3).
Results
We used diagnostic data from Partners Biobank to identify cases with insomnia and controls. The study cohort comprised of 21,310 patients of European ancestry with 11,420 females (53.6%) and 9,890 males (46.4%). The mean age was 59.7 (SD = 16.70). From a total of 15,750,104 patient visit records, we generated an ICD9/ICD10 list for the insomnia phenotype. The diagnosis definition for the cases included primary insomnia, insomnia due to medical conditions and insomnia due to psychiatric disorders. We removed patients with documented comorbid sleep disorder symptoms, including snoring, periodic limb movement, sleep related leg cramps, sleep-related bruxism and hypersomnia. Using this list, we obtained 3,135 case subjects. The control group consisted of 14,920 patients without any record of insomnia or other sleep disorder symptoms.
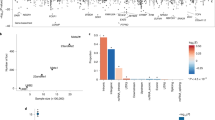
Using high-quality imputed SNPs, a genome wide association analysis was conducted for the insomnia phenotype. Setting the p-value threshold at 5 × 10−8, one novel genomic risk locus was identified on chromosome 8p21.2 (Fig. 1a, Supplementary Fig. 1a, Genomic Inflation Factor λ 1.007). The leading SNP was rs17052966 (p = 4.53 × 10−9) (Table 1), located inside the gene region of the long non-coding RNA (lncRNA), CTD-2168K21.1. Using FUMA (Functional Mapping and Annotation) and MAGMA (Multi-marker Analysis of GenoMic Annotation) pipeline, 8 protein-coding genes were identified in the 10 kb distance window, including LOXL2, ENTPD4, ADAMDEC1, ADAM7, NEFM, EBF2, BNIP3L and ADRA1A. Previous research has linked these genes to sleep related disorders, psychiatric disorders and neurodegenerative disorders (Table 2)31,32,33,34,35,36,37,38. 27 other SNPs reached suggestive threshold (5 × 10−6) were also identified (Table 1). Among them, multiple SNPs on Chromosome 4 were close to gene SORCS2, which functions as a receptor for the precursor form of neurotrophin39.
We also attempted to replicate previous GWAS study reported sleep disorder associated variants13,40. Among reported significant SNPs, 5 SNPs (rs8180817, 7q31.1; rs7044885, 9q31.32; rs113851554, 2p14; rs12187443, 5q21.1; and rs701394, 5q14.1) showed significances between 3.50 × 10−4 and 9.70 × 10−3 in our samples (Table 1). In addition, 8 SNPs that showed suggestive significances in our study had marginal p values in previous studies13,14 (Supplementary Table 2).
GCTA was used to estimate the proportion of phenotypic variance explained by common SNPs. The common SNPs could explain 7% (se = 0.02, p = 0.015) of the phenotypic variability. This is consistent with several previous GWAS studies on insomnia15,41. Using GCTA, we also calculated the genetic correlation between insomnia and three substance use disorder phenotypes, namely alcohol (3,594 cases, 12,205 controls), nicotine (4,896 cases, 12,205 controls) and opioid (1,039 cases, 12,205 controls) use disorders, which were also extracted from the same study cohort using ICD codes (Supplementary Table 1). The strongest correlation was found between insomnia and alcohol use disorder (rG = 0.56, se=0.14, p < 0.001), followed by nicotine use disorder (rG = 0.50, se=0.12, p < 0.001) and opioid use disorder (rG = 0.43, se = 0.18, p = 0.02) (Table 3). Furthermore, we evaluated the genetic correlations between insomnia and a series of clinical conditions extracted from Partners Biobank using codified data. Among them, a moderate correlation was observed between insomnia and anxiety or type 2 diabetes (rG = 0.76, se = 0.38, p = 0.17; rG = 0.31, se = 0.14, p = 0.25) (Table 3). Limited by the sample size, we did not observe the significant correlations.
To gain more statistical power and further validate our results, we obtained the summary statistics from two recent insomnia GWAS, using data from UK Biobank or STARRS dataset13,15. We calculated the pair-wise genetic correlations between results from the Partners Biobank and these two studies and observed a moderate correlation between our results and Jansen et al. 2019 study (rG = 0.68, se = 0.36, p = 0.18), while no significant correlation was found between Partners Biobank and Stein’s study (rG = 0.57, se = 1.28, p = 0.86). Lastly, a moderate correlation was observed between Jansen’ study and Stein’s study (rG = 0.35, se = 0.16, p = 0.07). We also checked the top two SNPs identified in Partners Biobank (rs17052966 and rs117915572) in both Stein’s and Jansen’s studies, but did not observe significant signals (STRASS: rs17052966: p = 0.95, beta = 0.002; rs117915572: p = 0.34, beta = 0.062; UKBB: rs17052966: p = 0.73, beta = 0.003; rs117915572: p = 0.83, beta = −0.002).
The meta-analysis was then conducted by combining our results from Partners Biobank and these two studies. Since the sample size of UK Biobank is significantly larger than our cohort and STARRS cohort, which can lead to a UK Biobank dominated meta-analysis result, we divided the meta-analysis into two steps: combining our results with the STARRS data alone (meta-1, N = 35,706) or combining all three studies (meta-2, N = 422,239) (Fig. 1b,c, Supplementary Fig. 1b,c, Supplementary Tables 3, 4). Two significant genomic loci were identified from meta-1 on chromosome 7 and 9, (Supplementary Tables 5 and 7). The leading SNPs, rs147549871 (p = 9.10 × 10−9) and rs7855172 (p = 1.32 × 10−8), from the identified loci were the top SNPs of the original study using the STARRS dataset. Also, the top SNPs rs17052966 and rs117915572 from Partners Biobank GWAS showed suggestive significances in the meta-1 analysis (p = 2.87 × 10−5 and p = 9.61 × 10−6). In meta-2 analysis, we identified 13 significant genomic loci with 31 independent significant SNPs, in which 11 loci were novel (Supplementary Tables 4 and 6). The top SNP, rs113851554 (p = 1.37 × 10−21), is on chromosome 2 and close to the MEIS1 gene. MEIS1 is a homeobox gene and plays an important role in neural crest development42. Multiple studies showed its relationship with sleep disorder, as well as restless legs syndrome (RLS)13,14,19,43.
In meta-analysis 2, using position mapping, we also identified 118 related genes within 10 kb region of significant SNPs. MAGMA tissue expression results suggested that genes from central nervous system tissues were highly enriched for expression (Supplementary Fig. 2). GWAS catalog analysis showed a series of previously reported sleep disorder genes, such as MEIS1, CUL9 and FOXP2 (Supplementary Table 8)40,44,45.
Discussion
Insomnia is one of the most prevalent mental disorders world-wide, affecting 10–20% of population. The strong genetic impact on insomnia has been repeatedly reported from different data sources. In many of these studies, self-reported insomnia symptoms were used to identify cases from the general population, which could limit our understanding of the complexity of this disease.
The current study used electronic health records and genomic information from a large patient cohort to conduct a GWAS on clinically defined insomnia phenotype. We discovered one novel genomic risk locus on chromosome 8. The leading SNP is in the region transcribes a long non-coding RNA, which has not been reported for insomnia. Differential expressions of several lncRNAs were shown to be associated with sleep deprivation46. In addition, among the eight genes mapped by our most highly significant SNP, 7 genes have been shown to be related with neuronal functions and psychiatric disorders, suggesting the possible significance of the genome region surrounding the discovered risk genomic locus.
We also conducted a large-scale meta-analysis by combining our results and 2 recent insomnia GWAS using data from UK Biobank and STARRS. The top identified SNP rs113851554 (p = 1.37 × 10−21) was among the top SNPs from Jansen et al. (p = 1.56 × 10−51) and Lane et al. (p = 9.76 × 10−30) 2019 studies13,14. Since the UKBB sample size is significantly larger than cohorts from Partners Biobank and STARRS, the result of meta-analysis was mainly driven by UKBB samples and the top SNPs from Partners GWAS did not show significances. However, we observed a moderated genetic correlation between study of Partners Biobank and Jansen’s report. Also, multiple significant SNPs we identified showed moderate significances in other GWAS, suggesting common components across them.
Substance use disorders, such as alcohol, nicotine and opioid, can also affect sleep patterns through various neurotransmitters and were shown to be significantly genetically associated with insomnia47. We found a strong positive genetic correlation between insomnia and these major substance use disorders among the same study population, providing more evidence for the relationship between psychiatric disorders and insomnia. Sleep patterns and multiple other clinical conditions were also showed to be closely connected. Studies have shown that sleep disorders affect more than 50% of adults with anxiety disorders48. Consistently, a moderate genetic correlation between insomnia and anxiety condition was observed in the current study. However, we did not observe significant correlations with depression, type 2 diabetes (we observed a moderate correlation) and schizophrenia which were previously reported13,15. Considering the previous correlation studies were mainly using summary statistics from UK Biobank, the different results we obtained could be caused by different definitions of these traits or the smaller sample size in our study.
Because of the broad definitions of insomnia, the phenotypes targeted by genome-wide association analysis have varied significantly across studies, ranging from primary insomnia to measurements of sleep length, sleep quality and early morning awakening. This could be one of the reasons for the fewer identified significant SNPs for insomnia and lack of consistent findings across studies. In this regard, electronic health records containing rich information about patient status and diagnostic information, can serve as an important data source of disease phenotypes.
This study has several limitations. First, insomnia is a common clinical symptom associated with multiple psychiatric disorders, which makes it very challenging to accurately define clinical insomnia. For the same reason, the genetic architecture identified by genome wide association studies can only reflect certain aspects of the complex insomnia phenotype. In this study, we used a simple ICD-code-based phenotype definition, and did not attempt to stratify the sample into multiple insomnia sub-phenotypes for GWAS due to the limitation of our sample size and the accuracy of the phenotyping method. We are planning to conduct following-up studies to further address these questions with larger sample size and other sources of phenotype information in the EHR, such as problem lists and clinical notes. Second, the study cohort is derived from a patient population, which could reflect more severe stage of insomnia. This could be one of the reasons we did not replicate several known insomnia related SNP from previous studies. Third, the cohort we extracted from Partners Biobank has a relatively small sample size compared with UK Biobank, which caused a significant imbalanced signal when conducting the meta-analysis.
In summary, we used clinical diagnosis information to identify insomnia cases among hospitalized patients. Our study cohort consists of clinically defined insomnia and provides a novel reference for insomnia genetic studies. Due to the heterogeneous clinical stages and complexity of the EHR data mining methods, we only utilized diagnostic codes in the development of our cohort in the current study. Based on this exploration, our developed pipeline will facilitate future research for more comprehensive genetic studies based on clinical records.
Data availability
The datasets generated and/or analyzed during the current study are not publicly available due to IRB regulations. The summary statistics are available from the corresponding author on request.
References
Association, A. P. Diagnostic and statistical manual of mental disorders (DSM-5®). (American Psychiatric Pub, 2013).
Roth, T. et al. Prevalence and perceived health associated with insomnia based on DSM-IV-TR; international statistical classification of diseases and related health problems, tenth revision; and research diagnostic criteria/international classification of sleep disorders, criteria: results from the America insomnia survey. Biological psychiatry 69, 592–600 (2011).
Ohayon, M. M. Epidemiology of insomnia: what we know and what we still need to learn. Sleep medicine reviews 6, 97–111 (2002).
Bonnet, M., Burton & Arand. Bonnet, M. H., Burton, G. & Arand, D. L. Physiological and Medical Findings in Insomnia: Implications for Diagnosis and Care. Sleep Medicine Rev, 2014, 18, 111-122. Vol. 18 (2014).
Morin, C. M. et al. Insomnia disorder. Nature Reviews Disease Primers 1, 15026 (2015).
ten Have, M. et al. Insomnia among current and remitted common mental disorders and the association with role functioning: results from a general population study. Sleep medicine 25, 34–41 (2016).
Roth, T. et al. Sleep Problems, Comorbid Mental Disorders, and Role Functioning in the National Comorbidity Survey Replication. Vol. 60 (2007).
Wing, Y. et al. Familial aggregation and heritability of insomnia in a community-based study. Sleep medicine 13, 985–990 (2012).
Serretti, A. et al. Genetic dissection of psychopathological symptoms: insomnia in mood disorders and CLOCK gene polymorphism. American Journal of Medical Genetics Part B: Neuropsychiatric Genetics 121, 35–38 (2003).
Buhr, A. et al. Functional characterization of the new human GABA A receptor mutation β3 (R192H). Human genetics 111, 154–160 (2002).
Retey, J. et al. A genetic variation in the adenosine A2A receptor gene (ADORA2A) contributes to individual sensitivity to caffeine effects on sleep. Clinical Pharmacology & Therapeutics 81, 692–698 (2007).
Deuschle, M. et al. Association between a serotonin transporter length polymorphism and primary insomnia. Sleep 33, 343 (2010).
Jansen, P. R. et al. Genome-wide analysis of insomnia in 1,331,010 individuals identifies new risk loci and functional pathways. Nat Genet 51, 394–403, https://doi.org/10.1038/s41588-018-0333-3 (2019).
Lane, J. M. et al. Biological and clinical insights from genetics of insomnia symptoms. Nat Genet 51, 387–393, https://doi.org/10.1038/s41588-019-0361-7 (2019).
Stein, M. B. et al. Genome-wide analysis of insomnia disorder. Mol Psychiatry 23, 2238–2250, https://doi.org/10.1038/s41380-018-0033-5 (2018).
Byrne, E. M. et al. A genome-wide association study of sleep habits and insomnia. Am J Med Genet B Neuropsychiatr Genet 162B, 439–451, https://doi.org/10.1002/ajmg.b.32168 (2013).
Amin, N. et al. Genetic variants in RBFOX3 are associated with sleep latency. Eur J Hum Genet 24, 1488–1495, https://doi.org/10.1038/ejhg.2016.31 (2016).
Gottlieb, D. J. et al. Novel loci associated with usual sleep duration: the CHARGE Consortium Genome-Wide Association Study. Mol Psychiatry 20, 1232–1239, https://doi.org/10.1038/mp.2014.133 (2015).
Hammerschlag, A. R. et al. Genome-wide association analysis of insomnia complaints identifies risk genes and genetic overlap with psychiatric and metabolic traits. Nat Genet 49, 1584–1592, https://doi.org/10.1038/ng.3888 (2017).
Seow, L. S. E. et al. Evaluating DSM-5 Insomnia Disorder and the Treatment of Sleep Problems in a Psychiatric Population. J Clin Sleep Med 14, 237–244, https://doi.org/10.5664/jcsm.6942 (2018).
Song, W., Huang, H., Zhang, C. Z., Bates, D. W. & Wright, A. Using whole genome scores to compare three clinical phenotyping methods in complex diseases. Sci Rep 8, 11360, https://doi.org/10.1038/s41598-018-29634-w (2018).
Gainer, V. S. et al. The Biobank Portal for Partners Personalized Medicine: A Query Tool for Working with Consented Biobank Samples, Genotypes, and Phenotypes Using i2b2. J Pers Med 6, https://doi.org/10.3390/jpm6010011 (2016).
Fuchsberger, C., Abecasis, G. R. & Hinds, D. A. minimac2: faster genotype imputation. Bioinformatics 31, 782–784, https://doi.org/10.1093/bioinformatics/btu704 (2015).
McCarthy, S. et al. A reference panel of 64,976 haplotypes for genotype imputation. Nat Genet 48, 1279–1283, https://doi.org/10.1038/ng.3643 (2016).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81, 559–575, https://doi.org/10.1086/519795 (2007).
Lee, S. H., Yang, J., Goddard, M. E., Visscher, P. M. & Wray, N. R. Estimation of pleiotropy between complex diseases using single-nucleotide polymorphism-derived genomic relationships and restricted maximum likelihood. Bioinformatics 28, 2540–2542, https://doi.org/10.1093/bioinformatics/bts474 (2012).
Yang, J. et al. Common SNPs explain a large proportion of the heritability for human height. Nat Genet 42, 565–569, https://doi.org/10.1038/ng.608 (2010).
Bulik-Sullivan, B. K. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat Genet 47, 291–295, https://doi.org/10.1038/ng.3211 (2015).
Watanabe, K., Taskesen, E., van Bochoven, A. & Posthuma, D. Functional mapping and annotation of genetic associations with FUMA. Nat Commun 8, 1826, https://doi.org/10.1038/s41467-017-01261-5 (2017).
Sanna, S. et al. Common variants in the GDF5-UQCC region are associated with variation in human height. Nat Genet 40, 198–203, https://doi.org/10.1038/ng.74 (2008).
Chuang, S. M., Wang, Y., Wang, Q., Liu, K. M. & Shen, Q. Ebf2 marks early cortical neurogenesis and regulates the generation of cajal-retzius neurons in the developing cerebral cortex. Dev Neurosci 33, 479–493, https://doi.org/10.1159/000330582 (2011).
Gao, F. et al. The mitochondrial protein BNIP3L is the substrate of PARK2 and mediates mitophagy in PINK1/PARK2 pathway. Hum Mol Genet 24, 2528–2538, https://doi.org/10.1093/hmg/ddv017 (2015).
Liu, Q. et al. Neurofilamentopathy in neurodegenerative diseases. Open Neurol J 5, 58–62, https://doi.org/10.2174/1874205X01105010058 (2011).
Maletic, V., Eramo, A., Gwin, K., Offord, S. J. & Duffy, R. A. The Role of Norepinephrine and Its alpha-Adrenergic Receptors in the Pathophysiology and Treatment of Major Depressive Disorder and Schizophrenia: A Systematic Review. Front Psychiatry 8, 42, https://doi.org/10.3389/fpsyt.2017.00042 (2017).
Mesarwi, O. A. et al. Lysyl Oxidase as a Serum Biomarker of Liver Fibrosis in Patients with Severe Obesity and Obstructive Sleep Apnea. Sleep 38, 1583–1591, https://doi.org/10.5665/sleep.5052 (2015).
Saito, A. et al. An association study on polymorphisms in the PEA15, ENTPD4, and GAS2L1 genes and schizophrenia. Psychiatry Res 185, 9–15, https://doi.org/10.1016/j.psychres.2009.09.018 (2011).
Wei, X. et al. Analysis of the disintegrin-metalloproteinases family reveals ADAM29 and ADAM7 are often mutated in melanoma. Hum Mutat 32, E2148–2175, https://doi.org/10.1002/humu.21477 (2011).
Yang, L. et al. Polygenic transmission and complex neuro developmental network for attention deficit hyperactivity disorder: genome-wide association study of both common and rare variants. Am J Med Genet B Neuropsychiatr Genet 162B, 419–430, https://doi.org/10.1002/ajmg.b.32169 (2013).
Glerup, S. et al. SorCS2 is required for BDNF-dependent plasticity in the hippocampus. Mol Psychiatry 21, 1740–1751, https://doi.org/10.1038/mp.2016.108 (2016).
Lane, J. M. et al. Genome-wide association analyses of sleep disturbance traits identify new loci and highlight shared genetics with neuropsychiatric and metabolic traits. Nat Genet 49, 274–281, https://doi.org/10.1038/ng.3749 (2017).
Lind, M. J. & Gehrman, P. R. Genetic Pathways to Insomnia. Brain Sci 6, https://doi.org/10.3390/brainsci6040064 (2016).
Sarayloo, F., Dion, P. A. & Rouleau, G. A. MEIS1 and Restless Legs Syndrome: A Comprehensive Review. Front Neurol 10, 935, https://doi.org/10.3389/fneur.2019.00935 (2019).
Schormair, B. et al. Identification of novel risk loci for restless legs syndrome in genome-wide association studies in individuals of European ancestry: a meta-analysis. Lancet Neurol 16, 898–907, https://doi.org/10.1016/S1474-4422(17)30327-7 (2017).
Dashti, H. S. et al. Genome-wide association study identifies genetic loci for self-reported habitual sleep duration supported by accelerometer-derived estimates. Nat Commun 10, 1100, https://doi.org/10.1038/s41467-019-08917-4 (2019).
Doherty, A. et al. GWAS identifies 14 loci for device-measured physical activity and sleep duration. Nat Commun 9, 5257, https://doi.org/10.1038/s41467-018-07743-4 (2018).
Gaine, M. E., Chatterjee, S. & Abel, T. Sleep Deprivation and the Epigenome. Front Neural Circuits 12, 14, https://doi.org/10.3389/fncir.2018.00014 (2018).
Valentino, R. J. & Volkow, N. D. Drugs, sleep, and the addicted brain. Neuropsychopharmacology, https://doi.org/10.1038/s41386-019-0465-x (2019).
Oh, C. M., Kim, H. Y., Na, H. K., Cho, K. H. & Chu, M. K. The Effect of Anxiety and Depression on Sleep Quality of Individuals With High Risk for Insomnia: A Population-Based Study. Front Neurol 10, 849, https://doi.org/10.3389/fneur.2019.00849 (2019).
Acknowledgements
The authors would like to acknowledge contributions of Partners HealthCare Biobank for providing genomic data and health information data, Partners HealthCare Biobank Team for providing all the technique support. This work was supported by United States National Library of Medicine grant T15LM007092 and the National Heart, Lung, And Blood Institute of the National Institutes of Health under Award Number R01HL122225.
Author information
Authors and Affiliations
Contributions
W.S., A.W., J.T. and J.K. wrote the main manuscript text. W.S., A.W., C-Y.C. and H.H. designed the analysis pipeline. W.S. and A.W. designed the data integration procedure. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Song, W., Torous, J., Kossowsky, J. et al. Genome-wide association analysis of insomnia using data from Partners Biobank. Sci Rep 10, 6928 (2020). https://doi.org/10.1038/s41598-020-63792-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-63792-0
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.