Abstract
TAC1 and LAZY1 are members of a gene family that regulates lateral shoot orientation in plants. TAC1 promotes outward orientations in response to light, while LAZY1 promotes upward shoot orientations in response to gravity via altered auxin transport. We performed genetic, molecular, and biochemical assays to investigate possible interactions between these genes. In Arabidopsis they were expressed in similar tissues and double mutants revealed the wide-angled lazy1 branch phenotype, indicating it is epistatic to the tac1 shoot phenotype. Surprisingly, the lack of TAC1 did not influence gravitropic shoot curvature responses. Combined, these results suggest TAC1 might negatively regulate LAZY1 to promote outward shoot orientations. However, additional results revealed that TAC1- and LAZY1 influence on shoot orientation is more complex than a simple direct negative regulatory pathway. Transcriptomes of Arabidopsis tac1 and lazy1 mutants compared to wild type under normal and gravistimulated conditions revealed few overlapping differentially expressed genes. Overexpression of each gene did not result in major branch angle differences. Shoot tip hormone levels were similar between tac1, lazy1, and Col, apart from exceptionally elevated levels of salicylic acid in lazy1. The data presented here provide a foundation for future study of TAC1 and LAZY1 regulation of shoot architecture.
Similar content being viewed by others
Introduction
Lateral organ orientation in both shoots and roots plays a key role in a plant’s interaction with the environment and its ability to access resources such as light and water. The IGT gene family members TILLER ANGLE CONTROL 1 (TAC1) and the related set of LAZY genes are important regulators of lateral organ orientation1,2. These genes share four conserved amino acid regions or domains, and LAZY genes share an additional C-terminal domain3,4. TAC1 generally occurs as a single copy gene, but many species possess multiple LAZY genes, with six identified in Arabidopsis2,3,4,5,6,7,8. While the LAZY1 gene of Arabidopsis thaliana (Arabidopsis) contributes almost exclusively to shoot architecture, the remaining Arabidopsis LAZY genes primarily control root architecture7,8. However, GUS reporter activity and additive shoot phenotypes in plants with combinatorial lazy1, lazy2, and lazy4 mutations suggest LAZY2 and LAZY4 also have a role in regulating shoot orientation7,8. LAZY2, 3, and 4 are also known as DEEPER ROOTING (DRO3, 2, & 1) and NEGATIVE GRAVITROPIC RESPONSE OF ROOTS (NGR1, 3, & 2), respectively, as they were separately identified as regulators of lateral root orientation6,9,10,11,12. Further, an alternate LAZY gene nomenclature denotes LAZY3 as LZY2 and LAZY4 as LZ3Y8.
Lateral branches, tillers, leaves, and flower buds of plants with loss-of-function tac1 mutations or reduced TAC1 expression exhibit upright orientations3,13,14,15,16,17,18,19. In contrast, lazy1 mutants have wider branch or tiller angles and lazy4/dro1 mutants display prostrate lateral root orientations4,5,6,7,8,9,20,21,22,23,24. Plants with multiple lazy/dro mutations exhibit even wider lateral shoot/root angles and, in some cases, downward shoot growth and/or upward root growth7,8,10,11. Together, those findings demonstrate that TAC1 promotes horizontal lateral organ orientations, while LAZY genes promote vertical orientations. Despite their sequence similarity and opposing roles in shoot architecture, functional relationships between TAC1 and the LAZY/DRO genes have not been identified to date.
Currently, very little is known about TAC1 function, but significant progress has been made uncovering LAZY/DRO functional mechanisms. LAZY/DRO family members regulate gravitropic responses in shoots and/or roots downstream of amyloplast sedimentation and upstream of the establishment of the asymmetric auxin hormone gradients generated by PIN auxin-efflux proteins4,5,7,8,12,20,21,25,26. LAZY/DRO genes are expressed in vasculature and gravity sensing tissues4,7,8,12,21. Single lazy1 and multiple lazy/dro mutants have impaired, or in some cases reversed, gravitropism phenotypes, yet exhibit normal amyloplast development and sedimentation in response to gravistimulation4,5,7,8,12,20,21,25,26. Oryza sativa (rice) and Zea mays (maize) lazy1 mutants have increased basipetal (root-ward) polar auxin transport and reduced lateral auxin transport, including gravity-induced transport in response to gravistimulation5,20,21,26. Additionally, lazy1 maize showed decreased expression of PIN1c, and agravitropic Arabidopsis lazy1;lazy2;lazy4 and lazy2;lazy3;lazy4 roots exhibited reversed auxin gradients and polar PIN3-GFP localization upon gravistimulation5,8,12
An N-terminal transmembrane domain (TMD) and two C-terminal Nuclear Localization Signals (NLS) have been identified in LAZY1 and other LAZY proteins4,27. Yet, the importance or specific role of each domain may vary between species. Heterologously expressed GFP-tagged full-length and truncated LAZY1 proteins, along with in vitro assays, indicated monocot and dicot LAZY1 proteins, as well as Arabidopsis AtLAZY2, 3 and 4, associate with the plasma membrane and, in some cases, microtubules and nuclei4,5,20,28,29. The LAZY1 TMD was needed for membrane localization of transiently expressed rice and maize LAZY1-GFP in onion20,27. But an AtLAZY1 truncation lacking the TMD still localized to membranes in tobacco28. In addition, Arabidopsis protein fractionation assays suggested AtLAZY1 is a peripheral membrane protein, not a transmembrane one28. Lastly, the AtLAZY1 N-terminal Region I, upstream of the TMD was found to be important for both plasma membrane localization and branch angle control, as illustrated by site directed mutagenesis29. AtLAZY1 Regions II and V are also essential for LAZY1-directed branch angle control29. Interestingly, in Arabidopsis, nuclear localization is not needed for LAZY1-mediated branch angle control4. The lazy1 shoot phenotype was rescued by overexpressing a LAZY1 sequence containing a mutation in the NLS, which was shown to prevent nuclear localization in tobacco4.
The last ~14 amino acids of the LAZY protein C-terminus (which is also called domain/region V, the Conserved C-terminus in LAZY1, or the CCL, as well as peptide VI) contains an Ethylene-responsive Amphiphilic Repression (EAR) transcriptional repressor motif and this region seems to be important for LAZY protein function in many species3,5,8,29. It was required to rescue Arabidopsis lazy1;lazy2;lazy4 mutant root gravitropism phenotypes8. Overexpression of this sequence in the triple mutant resulted in upward growing roots8. Further, site-directed mutagenesis of several amino acids within the EAR motif in region V reduced or eliminated rescue of the lazy1 and lazy1;lazy2;lazy4 phenotypes29,30. Lastly, the EAR motif mediated interactions between wheat LAZY4/DRO1 protein and auxin signaling repressor TOPLESS, suggesting it is functional in this species11.
LAZY1 transcription in rice is directly and positively regulated by the HEAT STRESS TRANSCRIPTION FACTOR 2D (HSF2D) protein upstream of auxin transport in response to gravistimulation31. Yeast-one-hybrid assays suggested LAZY4/DRO1 expression in wheat is directly regulated by AUXIN RESPONSE FACTOR 1 (ARF1) transcription factors11,31. In addition, yeast-two-hybrid and BiFC experiments determined that the maize LAZY1 (ZmLAZY1) protein bound to both a protein kinase (ZmPKC) which may be involved in PIN localization at the plasma membrane, and to a nuclear-localized Aux/IAA auxin signaling repressor protein (ZmIAA17)5. However, species-wide interpretations of results from maize should be cautioned because in addition to its role in gravitropism and shoot architecture, LAZY1 is also essential for tassel and ear development in maize5.
In contrast to the mechanistic knowledge about LAZY1, much less is known about TAC1. Its function in promoting wide lateral shoot angles is dosage dependent and light regulated3,13,14,15,16,17,18,19,32,33,34. TAC1 expression has been detected in shoot tissues in peach, poplar, and Arabidopsis, with higher expression detected in apical regions3,16,34. Further, expression in Arabidopsis was induced by light and eliminated after prolonged darkness or application of photosynthetic inhibitors18. Additionally, the narrow branch angle phenotype in Arabidopsis tac1 plants phenocopies the branch angles of wild type plants exposed to continuous shade or the chlorophyll biosynthesis inhibitor Norflurazon18. Lastly, AtTAC1 overexpression could not fully rescue the Arabidopsis mutant phenotype and it only partially prevented upright reorientation of branches in wild type plants in response to darkness3,18. These findings suggest additional pathways promote outward shoot orientations in Arabidopsis and/or the existence of post-transcriptional regulation of TAC1.
Here we examined the functional relationship between TAC1 and LAZY1 in Arabidopsis at the molecular and genetic level as well as in connection with gravitropic responses. Our experimental results suggest that, while these genes have several commonalities, they also have distinct functions. In addition, while they do not appear to directly regulate each other’s expression, an indirect negative regulation of LAZY1 function via TAC1 at the protein level cannot be ruled out.
Results
GUS reporter constructs revealed TAC1 and LAZY1 have similar expression patterns
Tissue-specific TAC1 and LAZY1 gene expression patterns were assessed in Arabidopsis plants throughout development using a GUS reporter system. Arabidopsis plants containing the Arabidopsis LAZY1 promoter driving GUS (promAtLAZY1::GUS) previously revealed LAZY1 promoter activity in shoots, vasculature, endodermis, petioles, and shoot apical meristem tissue, as well as a faint signal in the root which was generally restricted to root vasculature4,7,8. We generated an Arabidopsis line containing the TAC1 promoter driving GUS (promAtTAC1::GUS) to compare spatiotemporal expression pattering between LAZY1 and TAC1. We found TAC1 promoter activity was highly similar to LAZY1 in the shoot (Fig. 1; Supplementary Figures S1 and S2). Two- and four-week-old seedlings had strong expression in the vasculature of cotyledons and young leaves (Fig. 1a–d). Strong staining was consistently visible in the primary and most prominent (thicker) secondary roots of promAtTAC1::GUS seedlings as well as in lateral root tips (Fig. 1b; Supplementary Figure S2). In contrast promAtLAZY1::GUS seedling roots had fainter staining overall, although strong signal was sometimes observed in primary roots of older seedlings and adult plants (Fig. 1a; Supplementary Figure S1). Little to no expression was observed in mature promAtTAC::GUS and promAtLAZY1::GUS rosette leaves, but signal was present in rosette petioles as well as younger leaves to varying degrees (Fig. C-F; Supplementary Figures S1 and S2).
GUS staining of Arabidopsis containing PromAtLAZY1::GUS and PromAtTAC1::GUS reporter constructs revealed similar tissue-specific gene expression patterns. Two-week old PromAtLAZY1::GUS (a) and PromAtTAC1::GUS (b) seedlings. Four-week old PromAtLAZY1::GUS (c) and PromAtTAC1::GUS (d) seedlings. Mature AtPromLAZY1::GUS (e) and PromAtTAC1::GUS (f) plants. Mature PromAtLAZY1::GUS (g) and PromAtTAC1::GUS (h) flowers.
In mature plants, staining for TAC1 and LAZY1 expression was observed in shoots, with the highest signal in the apical regions (Fig. 1e,f, Supplementary Figures S1 and S2). As the distance from the shoot apices decreased, the signal became increasingly faint and was often absent from the base of the shoots (Fig. 1e,f, Supplementary Figures S1 and S2). For both promAtTAC1::GUS and promAtLAZY1::GUS plants, GUS expression in cauline leaves varied but was similar in intensity and localization and often strongest in the vasculature and at the base of the leaves (Fig. 1e,f, Supplementary Figures S1 and S2). TAC1 and LAZY1 reporter lines also had strong staining in newly emerging axillary shoots (Supplementary Figures S1 and S2).
The most noticeable difference in the localization patterns for the two genes was in the flowers, where TAC1 was highly expressed in filaments but LAZY1 signal was absent. GUS expression for both promoters was apparent in sepal vasculature, style, carpel base, and pedicels (Fig. 1g,h, Supplementary Figures S1 and S2). However, the TAC1 signal was consistently stronger in these tissues (Fig. 1g,h; Supplementary Figures S1 and S2). On occasion, promAtTAC1::GUS signal was observed in the carpel valves and septum of developing siliques (Supplementary Figure S2).
Arabidopsis lazy1 branch phenotype was epistatic to the tac1 branch phenotype
The similarity between TAC1 and LAZY1 promoter::GUS expression patterns along with the opposing phenotypes of tac1 and lazy1 Arabidopsis suggest a possible intersection between their control of branch orientation. To determine the combined effect of their non-functional alleles on shoot architecture, tac1 and lazy1 single mutants were crossed and the shoot orientation phenotypes of at least two hundred homozygous double mutants from multiple lines and generations were characterized. The tac1;lazy1 double mutants clearly and consistently displayed a lazy1-like wide branch angle phenotype and could only be distinguished by genotyping (Fig. 2). Thus, the tac1;lazy1 mutant phenotype suggests lazy1 is epistatic to tac1 with regards to branch orientation control.
tac1 mutants did not exhibit an altered gravitropic shoot bending response
Previous studies found that lazy1 mutants had delayed gravitropic bending responses to shoot reorientation4,5,21,26. Here we tested if TAC1 also has a role in gravitropic shoot bending. We hypothesized that tac1 mutants might exhibit faster gravitropic bending responses than wild type and lazy1 plants due to the more upward-oriented branch phenotype in tac1 plants. We also wanted to test if the lazy1 gravitropic response would be epistatic to tac1, just as the branch orientation phenotype was. To investigate these possibilities, we performed four separate 90° reorientation experiments with up to 30 Arabidopsis wild type Columbia-0 (Col), tac1, lazy1, and tac1;lazy1 double mutant plants in pots (See Supplementary Figure S3 for our set-up). Plants were affixed to a custom-built rack and reoriented 90° under green light to eliminate phototropic interference and imaged in a single frame every minute for up to 12 hours to generate time-lapse videos, as exemplified in Video V1. To reduce the influence of plant size and shoot weight, all plants were a similar height and had only one primary shoot with short branches. Time-lapse videos were studied to assess if obvious reorientation differences existed between genotypes. We observed that the tac1 plants fully reoriented, while lazy1 and tac1;lazy1 exhibited similarly slow bending responses compared to Col (Fig. 3, Supplementary Figure S3, Video V1). Obvious differences between tac1 and Col plants were not detectable by eye. To quantitatively compare gravitropic bending rates, we initially measured the angle between plant shoot tips and the shoot trajectory. At 30- and 60-minutes post reorientation, lazy1 and tac1;lazy1 were very minimally less upright than Col (their average tip angles were closer to the 180-degree horizontal), and there was no significant difference between tac1 and Col (Table S1). This agreed with our visual observation that tac1 did not reorient faster and lazy1 and tac1;lazy1 responded equally slower than Col (Table S1). However, imaging a large field of view reduced the resolution of individual shoots and shoot tips, increasing variability and subjectivity when choosing anchor points for angle measurements. In addition, the simulated dark conditions resulted in extensive shoot tip circumnutation (clearly visible in Video V1). Consequently, tip angles from the images spanning the majority of the 12-hour time period did not always correlate with the degree of bending observed in the shoots. Therefore, we subsequently developed a novel approach to compare gravitropic responses over time in relation to shoot curvature, the most obvious response to phenotype. Using ImageJ software35 and images from twenty time points spanning 1.5 to 12 hours after reorientation, we extracted the area of circles that, when overlaid, best fit the curvature of each stem at the site of bending. This method enabled relative comparisons between genotypes for the later-stage bending response; as time from reorientation increases, shoots become more upright and the areas of the best-fitting circles decrease (Fig. 3). While this method works well to quantify the overall curvature of a stem, it is unable to assess possible differences during very early shoot lifting.
Gravitropic responses of wildtype (Col), tac1, lazy1, and tac1;lazy1 Arabidopsis plants after a 90-degree reorientation. (A) Images of the same representative Col, tac1, lazy1, and tac1;lazy1 plants at 30-minute, 6-hour, and 12-hour time points. (B) The rate of bending in response to reorientation represented by the decreasing area of circles that best fit the curve of each inflorescence meristem bend between 1.5 and 12 hours after reorientation. Each data point represents the average area from four independent experiments, each with five to nine plants per genotypes. Student’s t-test results found significant differences between the Col and lazy1 and Col and tac1;lazy1 values at every time point (p < 0.05). No significant differences were found between Col and tac1 or lazy1 and tac1;lazy1 at any time point. Data points were fit with a power trendline and error bars indicate SEM.
Using this ‘best-fit circle’ relative assessment method, we identified that lazy1 and tac1;lazy1 bending (curvature) responses were slower than Col and tac1. The circles fitting their shoots at all time points were consistently larger than those for Col and tac1 (p < 0.05, determined by t-test) (Fig. 3). No differences were found between Col and tac1 or lazy1 and tac1;lazy1 at any time point. The lazy1 and tac1;lazy1 genotypes exhibited a comparable delayed gravitropic curvature response, suggesting the loss of TAC1 does not alter this lazy1 gravitropism phenotype (Fig. 3). In addition, we found no differences in overall rate of reorientation between tac1 and Col using this method (Fig. 3).
Ectopic LAZY1 expression causes an EAR-domain dependent curled leaf phenotype
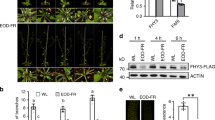
We previously reported that overexpression of TAC1 in plum trees resulted in wider branch angles17. However, we also reported TAC1 overexpression only partially complemented the tac1 Arabidopsis mutant phenotype, and in 24-hour light or dark conditions TAC1 overexpression in Col had little effect on branch angle3,18. In contrast, 35 S::AtLAZY4/DRO1 in Arabidopsis exhibited more vertically oriented lateral shoots as a result of narrower branch angles, as well as upward curled leaves6. Further, the overexpression of a LAZY4/DRO1 sequence lacking the EAR motif (35 S::AtLAZY4/DRO1ΔEAR) did not result in these phenotypes6. Here, we investigated the effect of overexpressing the full-length AtLAZY1 CDS (35 S::LAZY1) as well as a truncated sequence lacking the EAR motif (35 S::LAZY1ΔEAR) in the Col background (Fig. 4, Supplementary Figure S4). For comparison, 35 S::TAC1 in Col were grown alongside the LAZY1 and LAZY1ΔEAR overexpression plants (Fig. 4B,F; Supplementary Figure S2). We found the 35 S::TAC1 plants exhibited a very slight, but significant increase in branch angle (Supplementary Figure S4). In addition, in contrast to the published 35 S::AtLAZY4/DRO1 phenotype, overexpression of full length LAZY1 surprisingly did not produce obvious branch angle differences compared to Col (Fig. 4A–D). However, when measured, the 35 S::LAZY1 plants also had a slight increase in branch angle compared to Col (Supplementary Figure S4). Compared to Col, 35 S::LAZY1 plants also exhibited upward curling of rosette leaves, with some rolling up completely as they aged (Fig. 4I,J). Upward curled cauline leaves were also observed sporadically in the 35 S::LAZY1 plants but not in Col. These leaf phenotypes were comparable to those in 35 S::AtLAZY4/DRO1 plants6. Curled leaves were not observed in 35 S::AtLAZY1ΔEAR or 35 S::TAC1 (Fig. 4). In addition, young rosette leaves from 35 S::LAZY1 plants were more rounded than the Col, 35 S::TAC1, and 35 S::AtLAZY1ΔEAR plants (Fig. 4E–H). The lack of a 35 S::LAZY1 branch angle phenotype, in contrast to the narrower branch angles exhibited in published 35 S::LAZY4/DRO1 lines, suggests possible spatial regulatory divergence regarding lateral shoot orientations, as LAZY4/DRO1 primarily regulates root angle. However, the shared leaf curl phenotype, and absence of this phenotype in both LAZY1ΔEAR and LAZY4/DRO1ΔEAR overexpression plants, suggests some functional conservation, as well as highlights the importance of the EAR motif for LAZY family members.
Overexpression of LAZY1 resulted in leaf curl but not altered branch angles. Arabidopsis Columbia (A,E), 35 S::TAC1 (B,F), 35 S::LAZY1 (C,G), and 35 S:LAZY1∆EAR (D,H) plants and young rosettes prior to bolting. (I) Mature 35 S::LAZY1 rosette with bolt removed (J) Enlarged image of curled rosette leaf from (I). (K) 35 S::LAZY1 node with a rare upward curled cauline leaf. (L) Columbia node with cauline leaf.
lazy1 plants exhibit substantially elevated SA levels
Hormone concentrations and gradients play a key role in directing shoot architectures, including lateral shoot orientations. Prior studies established that LAZY1 acts upstream of auxin transport5,7,20,21,26,31. Pillar peach trees (tac1 mutants) were found to have elevated auxin concentrations in their shoot tips relative to standard cultivars36. Further, altered Abscisic Acid (ABA) localization in branches was associated with a weeping mulberry tree architecture, and tac1 is epistatic to a weeping peach architecture32,37. To investigate if TAC1- or LAZY-mediated shoot orientations are associated with hormone concentrations, LC/MS/MS was performed on extracts from Arabidopsis tac1, lazy1, and Col shoot tips, stems (just below the tip), and branch tips from plants (Fig. 5; Supplementary Figure S5). Salicylic Acid (SA), Jasmonic Acid (JA), ABA, and auxin (IAA) concentrations were quantified. These tissues were chosen because they are regions where TAC1 and LAZY1 gene expression is highest.
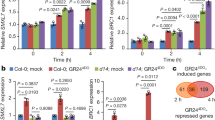
LC/MS/MS hormone analysis from shoot tissues. Hormones were extracted from (A) the shoot tip, (B) the stem immediately below the shoot tip segment, and (C) shoot apex of lateral branches. Bars represent standard deviation of biological replicates, each of which contained pooled tissue from multiple plants. *Indicates hormone concentration was significantly different than the same tissues from other genotypes according to ANOVA, p < 0.01 with Tukey’s ad hoc test.
In all three tissue types, SA levels were substantially higher in lazy1 compared to both Col and tac1, with statistically significant increases in the primary shoot and branch tips (Fig. 5). Arabidopsis tac1 plants had less free IAA than Col in both shoot tips and stems, with the latter being statistically significant (Fig. 5A,B). In addition, lazy1 shoot tips had significantly greater IAA concentrations (Fig. 5A,B). No statistically significant differences in JA or ABA concentrations were detected between genotypes for the tissues we tested, although ABA levels in tac1 were slightly lower in stems and branch tips (Fig. 5).
Transcriptomes suggest tac1 and lazy1 mutants have altered gene regulation in response to gravistimulation
To compare influences of TAC1 and LAZY1 on transcriptional regulation, RNAseq transcriptome profiles were generated for inflorescence shoot tips from Col, tac1, and lazy1 under normal conditions (t0) and after 45 minutes of gravistimulation by 90° reorientation (t45) (Supplementary Data File D1). Shoot tips were again used because promoter::GUS and qPCR studies revealed TAC1 and LAZY1 are highly expressed in shoot tips and often absent from the middle and base of primary shoots and branches (Fig. 1; Supplementary Figures S1 and S2)3.
After comparing expression profiles of each mutant to Col under the same conditions (normal or gravistimulated), a combined total of 829 differentially expressed genes (DEGs) were identified (Supplementary Data File D2). Fewer than 200 of these genes had expression changes greater than 2-fold, and very few DEGs had changes greater than 5-fold (Fig. 6A; Supplementary Data File D2). Most of the largest differences of expression were in response to gravistimulation. Interestingly, although no gravitropic bending differences between tac1 and Col were detected, in response to gravistimulation Col shoot tips had over two times the number of DEGS than tac1 (Fig. 6A, Supplementary Data File D2).
Differentially expressed genes (DEGs) between upright and gravistimulated Columbia (Col), tac1, and lazy1 arabidopsis. (A) Numbers of DEGs from different comparisons of RNAseq data from upright (t0) and gravistimulated (t45) plants. (B,C,D) Venn diagrams indicating numbers of overlapping DEGs between t0, t45, and t45 vs. t0 comparisons. (E,F) Numbers of DEGs in significantly enriched MapMan gene categories for upright (E) and gravistimulated (F) comparisons. All categories have statistically significant representation of DEGs with a p < 0.05. ** indicates very high significance (p < 0.0001) and *** indicates extremely high significance (p < 10e-10).
Due to their opposing phenotypes and sequence similarity, we hypothesized that TAC and LAZY1 may act through common pathways. Accordingly, we anticipated that tac1 and lazy1 transcriptomes might have overlapping of DEGs, and potentially some with opposite directional changes. However, there were only three DEGs in common between the genotypes with upright orientations (Fig. 6B–D, Supplementary Data Files D2 and D3). They were AtLFNR2, CSP41S, and HTB4 and they all had minimal fold changes, the changes were not in opposite directions, and the three genes have no known functional relationships to each other (Supplementary Data Files D2 and D3). Similar results were observed for the DEG comparisons from the gravistimulated plants (Fig. 6C,D; Supplementary Data Files D2 and D3).
A very small number of DEGs stood out as exhibiting exceptionally large differences in expression between genotypes. These included the uncharacterized gene At3g01345, upregulated nearly 400-fold in upright (t0) tac1 and METHIONINE RESPONDING DOWN 1 (MRD1), repressed over 300-fold in upright (t0) lazy1 (Supplementary Data File D2). Secondary cell wall formation genes TRACHEARY ELEMENT DIFFERENTION 6 (TED6) and TED7 were also greatly downregulated in the gravistimulated (t45) lazy1 tissues (−82 and −43-fold, respectively) but not in t45 Col (Supplementary Data File D2). Another notable result was LAZY1 was not differentially expressed in tac1 nor was TAC1 differentially expressed in lazy1 under either normal or gravistimulated conditions (Supplementary Data File D2). Further, TAC1 and LAZY1 were not differentially expressed in Col shoot tips in response to gravistimulation (t45 Col vs. t0 Col) (Supplementary Data File D2). This is consistent with a prior report of LAZY1 expression increasing only after several hours of 90° reorientation in rice seedlings31. Thus, our collection time may have been too early after 90° reorientation to detect differential expression of TAC1 and/or LAZY1. In addition, the transcriptome analyses for gravistimulated plants come with the caveat that they are from whole shoot tips; any asymmetric (upper vs. lower) expression differences in the shoot tips would not be detectable.
Despite the higher concentration of SA in lazy shoot tips and stems, DEGs associated with SA were not identified in lazy1 plants grown under normal conditions (Supplementary Data File D2). However, there was a 3-fold decrease in BSMT1 expression in gravistimulated lazy1. This methyltransferase is induced by biotic and abiotic stress to produce methyl-salicylate (MeSA), a mobile signal for systemic acquired resistance38. BSMT1 downregulation might be a response to a negative feedback mechanism triggered by high SA levels. Yet, this same gene was upregulated 8-fold in in gravistimulated Col (compared to upright Col) (Supplementary Data File D2). SA-degradation gene MO1 was also upregulated ~8-fold in t45 Col, as well as WAK1, a cell wall associated kinase known to respond to SA. Another DEG with a SA connection (ADH2) was downregulated in the t45 lazy1 transcriptome, but to a lesser degree. ADH2 expression is reduced in response to high SA and, interestingly, adh2 mutants exhibit dwarfing and dark-grown seedlings do not elongate their hypocotyls (phenotypes reported by ABRC; arabidopsis.org)39. In the t45 Col samples, a few additional SA-responsive genes were also slightly differentially expressed (Supplementary Data File D2).
A MapMan categorical analysis of the DEGs from each comparison revealed both expected and unexpected enrichments. This included an enrichment of only stress-related genes for upright tac1 plants compared to Col (Fig. 6E, Supplementary Data File D4). A similar enrichment was reported for tac1 (pillar) peaches17. Supporting the known connection between LAZY1 and auxin transport, and in accordance with the lazy1 mutant phenotype, t0 lazy1 transcriptomes (when compared to t0 Col) were enriched for DEGs associated with hormone metabolism, the cell wall, and photosynthesis, among other categories (Fig. 6E, Supplementary Data File D4). This included downregulated auxin efflux transporters PIN3 and PIN4, auxin responsive SAUR16, multiple expansins (EXP3, 5, and 8), FASCICILIN-LIKE ARABINOGALACTANS (FLA2 AND 11), wall degradation enzymes (GH98B and a pectin lyase), a cellulose-synthase-like gene (CSLC4) and 18 photosynthesis-related genes (Supplementary Data Files D2 and D4).
MapMan categorizations of DEGs from gravistimulated mutant to Col comparisons and t45 vs. t0 plants of the same genotype identified enrichment in additional gene categories, (Supplementary Figures S4, S6 and Supplementary Data File D4). These categories included various types of metabolism as well as the general ‘Protein’ and ‘RNA’ categories (Fig. 6F, Supplementary Figures S4, S6 and Supplementary Data File D4).
Discussion
IGT family genes TAC1 and LAZY1 have opposite but essential roles in directing lateral shoot orientations in vascular plants. TAC1, which is upregulated in response to light, promotes outward shoot orientations (via establishing wide branch or tiller angles). LAZY1, acting in the gravitropism pathway upstream of lateral auxin transport, promotes upward shoot orientations (via establishing narrow branch or tiller angles). We previously proposed that TAC1 and LAZY1 may act in the same pathway but one (or possibly both) can regulate the other’s function2. This was logically based on their opposing mutant phenotypes, the presence of LAZY genes in primitive plants and the apparent evolutionary loss of the EAR motif in TAC1, and gene dosage-dependent phenotypes in tac1 Arabidopsis and peach heterozygotes3. However, genetic interactions between TAC1 and LAZY1 as well as a direct connection between TAC1 and gravity response have not been established. Here, we found that LAZY1 and TAC1 were both expressed in similar shoot tissues at the same stages of development, as assessed using GUS reporters. Second, we found that lazy1 was epistatic to tac1 as double tac1;lazy1 mutants displayed a lazy1 branch phenotype. In addition, loss of TAC1 had no detectable impact on gravitropic bending or curvature responses, and the double tac1;lazy1 mutant showed very similar bending dynamics to lazy1 alone. Lastly, auxin levels were also inverse in the two mutants with lazy1 shoot tips exhibiting slightly higher IAA levels and tac1 shoot tips having lower IAA levels. While these results are consistent with TAC1 potentially serving as a negative regulator of LAZY1, the collective data were not entirely consistent with a simple negative regulatory model.
If TAC1 was a direct negative regulator of LAZY1, we might identify opposite directional changes of common DEGs in the respective mutants, which we did not observe. In addition, LAZY1 expression was not differentially expressed in tac1 plants nor was TAC1 expression altered in lazy1 mutants. Further, although TAC1 and LAZY1 overexpression in some species significantly enhanced branch/tiller angles, this was not the case in Arabidopsis13,16,17. 35 S::TAC1 lines were only minimally wider than Col while 35 S::LAZY1 also exhibited slightly wider branch angles, a strong contrast to previously reported narrow branch angle phenotype of 35 S::LAZY4/DRO1 Arabidopsis6. Only the curled leaf phenotype of LAZY4/DRO1 overexpression was recapitulated when using LAZY1. This result may originate from minor functional differences between species for both genes and among LAZY-family members. While the branch angle directional change in 35 S::LAZY1 contradicts known function, a recent publication demonstrated inherent branch angle inconsistencies when expressing a LAZY1 transgene in Arabidopsis29. Together these findings suggest TAC1 and LAZY proteins and their interactions may be subject to complex regulatory mechanisms.
Although the precise nature of the relationship between these two genes remains unclear, the data presented here provide several important insights. First, relatively few genes were differentially expressed in tac1 and lazy1. Second, the majority of the DEGS were downregulated. Thus, if the LAZY1 protein can act as a repressor via its EAR motif, it likely targets few genes, and may repress one or multiple repressors. Also, potential transcriptional repression by LAZY1 in Arabidopsis is likely secondary to its gravitropism role, as nuclear localization was not required for maintaining branch orientation in this species4. Third, our RNAseq data are consistent with a role of LAZY1 in influencing gravitropic responses. Genes repressed in lazy1 plants included cell wall-related genes such as expansins and FLAs (whose expression was downregulated in lazy1), thereby promoting asymmetric cell elongation in shoots for positioning purposes. The lateral auxin transport genes PIN3, PIN4, and auxin influx gene AUX1 were also downregulated in lazy1.
Interestingly, one gene showed extreme differential expression. MTO 1 RESPONDING DOWN1 (MRD1) was repressed by over 300-fold in upright lazy1 shoot tips. MRD1, is minimally characterized as of now, with the exception that it is downregulated in mto1-1 mutants that over-accumulate methionine. However, this gene in the root system architecture mutant agb1-2 was found to be co-expressed with At3g01345, the most highly induced gene in shoot tips from upright tac1 plants40. Connections between MRD1, At3g01345, TAC1, and LAZY1 warrant further investigation, as the functions of both are not understood.
One of our most surprising findings, and one we cannot at this time explain, was the tremendous increase in SA concentrations (~5 fold) in shoot tips, stems, and branch tips of lazy1 mutants. SA is primarily associated with biotic stress and no prior connections between this hormone and any lazy mutants have been reported. However, several studies have shown that SA can regulate shoot elongation. SA accumulation in response to cold temperatures inhibits cell elongation and reduces plant growth, and SA signaling (but not concentration) is important for petiole elongation in response to shade41,42. It’s possible that, in addition to auxin transport impairments, the lazy1 branch angle phenotype is connected to reduced cell elongation in abaxial branch tissues as a result of high SA concentrations. The Arabidopsis LAZY1 promoter region also has several SA responsive W-Box and WLE1 elements — (T)TGAC(C/T) and TGACA respectively — suggesting SA may have a direct role in regulating LAZY1 expression43,44. However, our RNAseq data uncovered DEGs more tangentially, rather than directly, related to SA biosynthesis and signaling. Given these results and the presence of SA-responsive elements, connections between LAZY1 and SA warrant further exploration.
In summary, the data described here highlight molecular and genetic aspects of TAC1- and LAZY1 function and their relationship to each other. Recent findings linking LAZY proteins to PIN3 localization via physical interactions with BRX-domain proteins suggest a possible mechanistic area where TAC1 and LAZY1 function could intersect30,45. One possibility is that TAC1 acts to integrate light perception into an ancient LAZY1-mediated gravity response pathway, thereby optimizing shoot positions for light capture. However, extensive experimentation is needed to build a comprehensive understanding of their independent and/or interdependent roles in regulating lateral shoot orientations in plants. The data provided here fills in some key knowledge gaps and provides new directions for future studies on TAC1 and LAZY1 branch angle control.
Materials and methods
Cloning
The promAtTAC1::GUS construct was generated by cloning a 2-kb fragment of the AtTAC1 promoter sequence, including the 5’ untranslated region (5′UTR), upstream of the GUS CDS in a pBI101 vector, using SalI and SmaI restriction sites. The 35 S::LAZY1 plasmid was made by amplifying the AtLAZY1 CDS from Arabidopsis cDNA using the forward primer 5′-CGA GTC GAC ATG AAG TTT TGG GGC TGG-3′ and reverse primer 5′-AAT GGA TCC TTA CAG TTC CAA CAC GAA ATA GTC TTC-3′ and cloning it into pCR8/GW/TOPO. After sequence confirmation, the CDS was inserted downstream of the 35 S promoter in 35S-pBIN-AFRS (a modified pBINPLUS/ARS vector) using the SalI and BamHI restrictions sites46. Similarly, the 35 S::LAZY1∆EAR construct was made by amplifying the AtLAZY1 CDS, using a reverse primer (5′-AAT GGA TCC TTA ATA GTC TTC GTC TGT CTT GAT CCA G-3′) that excluded the sequence for the last 15 amino acids of the full-length protein.
Plant material, Genotyping, and Branch Angle Measurements
The Arabidopsis tac1, lazy1, promLAZY1::GUS, and 35 S::TAC1 plants were previously described3,4,18. The tac1;lazy1 homozygous double mutant lines were generated by crossing the single mutants. Genotyping for tac1 and lazy1 was performed using standard PCR methods. To detect the lazy1 mutant allele, the primer AtLAZY-Fwd (5′-CAA GGT TTT TAC AAC ACA GAG CAA) was with AtLAZY-T-DNA Rev (5′-ATA TTG ACC ATC ATA CTC ATT GC) primer. To detect the wild type LAZY1, AtLAZY-Fwd was used with At-LAZY-genotype-Rev (5′ GCT GAC ACC CAT CAT AAT GCT T). The mutant tac1 allele was detected using AtTAC1-F-genotype (5′TCA ATT GTT CGT GTG CGT TT) and AtTAC1-R-genotype (5′AAA AGC TCC GCA AGT GTT GT). Wild type TAC1 was detected with AtTAC1-F-genotype and AtTAC1mut-Rev (5′TTA AAA ACG TCC GCA ATG TG). The 35 S::LAZY1, 35 S::LAZY1∆EAR, and the promAtTAC1::GUS plants were generated by transforming their respective constructs, described above, into Arabidopsis (cv. Columbia) via agrobacteria-mediated floral dip transformation and kanamycin selection. Multiple transgenic lines were phenotyped and/or stained and average lines were used for characterizations. All plants were grown in either 2- or 4-inch pots with one plant per pot and 16-hour light (~100 µmol m−2 s−1) in growth chambers set to ~22 °C. Branch angle measurements for Col, 35 S::TAC1 and 35 S::LAZY1 plants were determined by measuring from the tip of each branch (just below the floral cluster) to the branch node and then up the stem to the node above each branch.
GUS staining
Tissues for GUS staining were harvested in cold 90% acetone prior to a 20-minute room temperature incubation. Afterwards, tissues were rinsed with a staining buffer solution containing 50 mM NaHPO4 buffer (pH 7.2), 0.2% triton, 0.5 mM potassium ferrocyanide, and 0.5 mM potassium ferricyanide. Next, samples were submerged in staining buffer containing a 2 mM final concentration of X-gluc, vacuumed on ice for ~20 minutes, and incubated in dark for up to 16 hours at 37 °C. The next day, tissues underwent 30-minute incubations in the following solutions: 20%, 30%, and 50% ethanol, FAA, and then 70% ethanol. Several warm 95% ethanol washes followed until tissues cleared and samples were stored in 95% ethanol until imaged using a digital camera or a Nikon SMZ-800N stereomicroscope with a Nikon DS-Fi3 camera.
Gravitropism experiment
For the gravitropism experiments, on four different dates 4-5-week-old plants in 2″ square pots with a single inflorescence shoot approximately ~10–12 cm tall were placed in a custom-built rack at the same time of day, adapted to dark conditions under green light in a sealed room for one hour, and then rotated clockwise 90° to stimulate shoot bending. Prior to dark treatment, a metal pin was placed on the right side of each shoot, between the tip and base, to minimize initial shoot dropping. Between six and nine plants of each genotype were included for each experiment and the placement of each genotype was randomized. A total of 28 Columbia, 27 tac1, 30 lazy1, and 23 tac1;lazy1 plants were used for quantification. Images were taken every minute for between 12 and 36 hours after rotation with a Canon EOS REBEL T3 digital camera using a time-lapse intervalometer shutter release control. A timer, counting from zero, was started immediately after reorientation and placed in the field of view of the camera to ensure accurate correlations between images and time after reorientation. Time-lapse videos were generated using Microsoft Movie maker or Microsoft Video Editor. To quantify bending, images from twenty time points between 1.5 hours and ending at 12 hours (at 30- or 60-minute intervals) were selected. For plants on these images, ImageJ was used to draw circles that best fit the curvature at the bending point in the stem and then determine the area of that circle. A total of 2,160 measurements were taken. For each time point, a Student’s t-test was used to test for statistically significant differences between the average area for each genotype to each other.
RNA isolation for transgenic plants and qPCR analyses
Total RNA was extracted from Arabidopsis leaves using the Zymo Research Direct-zol RNA MiniPrep kit (Irvine, CA). qPCR was conducted using 50 ng total RNA per sample using the iTaq Universal SYBR Green One-Step Kit (BioRad, Hercules, CA) in a 15 µl total reaction volume in a 384 well plate on a BioRad CFX384 Touch with the following settings: 50 °C for 10 min for the RT step followed by a 95 °C 1 minute denaturation and then 35 cycles of 95 °C for 10 seconds and 60 °C for 20 seconds. A melt curve was subsequently performed to confirm the presence of only one amplicon. LAZY1 primers were Fwd: TCC GGG AGA ATA GCA AAG AGC CAT and Rev: 5′ TGT ATT GCT TGG GTC CTG CGA A; TAC1 primers were Fwd 5′AGC TGG TCA TGT CAA AGT CCA and Rev: 5′ TCA CAG TTC CGA GTT GGC TTG T). For 35 S::LAZY1, 35 S::LAZY1∆EAR, and 35 S::TAC1 lines between three and six biological reps were tested, each with three technical reps. Samples were analyzed for relative expression based on a standard curve using Columbia RNA.
Hormone analysis
Phytohormone extraction and LC/MS/MS methods were adapted from an established protocol47. Approximately 1 cm samples were collected from the tip of the primary inflorescence shoot, 1 cm below the tip, and the tip of the first sub-apical branch of Arabidopsis Columbia, lazy1, and tac1 plants (Supplementary Table S2). At the time of collection, the plants were approximately 12 cm tall and had only one shoot and no more than three branches. Three biological replicates were generated, each containing a pool of samples from 12-15 plants. These samples were flash-frozen in liquid nitrogen and the fresh weight of each pool was recorded. Phytohormones were extracted by grinding samples with a drill-driven blue pestle in buffer composed of 80% MeOH, 20% H2O, 0.1% formic acid, 0.1 g/L butylated hydroxytoluene, and internal standards of 100 nM [D6]-ABA (Toronto Research Chemicals) and 100 nM [D7]IAA (CDN Isotopes), using 1 mL of extraction buffer per 100 mg of sample. Samples were then rotated at 4 °C for 18 hours, centrifuged to pellet debris, the supernatant was filtered through a 0.2 m centrifugal filter unit (Millipore), and transferred to an autosampler vial. Samples (10 µL injection) were separated on an Ascentis Express C18 column (5 cm ×2.1 mm, 2.7 µm pore size (Supelco) using a linear gradient of 0.1% Formic Acid (solvent A) to 100% methanol (solvent B) with a flow rate of 0.4 mL/min, column temperature of 50 °C, and a 5 minute program. The separated samples directly interfaced with a Quattro Premier XE (Micromass Technologies) tandem quadrupole mass spectrometer via electrospray ionization. Capillary voltage, cone voltage and extractor voltage were 3 kV, 25 V, and 5 V, respectively. Source temperature was 120 °C and desolvation temperature was 350 °C. Cone gas flow was 50 L/hr, desolvation gas flow was 600 L/hr, and collision gas flow was 0.15 mL/min. Optimized detection parameters for each analyte is listed in Supplementary Table S2. Detection was performed by multiple reaction monitoring (MRM) in positive ion mode for IAA and IAA conjugates and negative ion mode for ABA, JA, and SA. Data acquisition and calculation was performed in Masslynx 4.1 software. A 4-fold dilution series from 1 µM to 1 nM was used to generate a standard curve, which then allowed for quantification of each phytohormone by peak area rations related to their corresponding internal standard ([D6]-ABA or [D7]-IAA).
Transcriptome tissue collection
Arabidopsis plants containing only a single inflorescence shoot, with a height of ~15-18 cm, and no more than 3 lateral shoots (each no larger than ~1 cm) were used for expression profiling. Prior to collection, plants were left for at least 12 hours in a dark chamber with wooden stakes to support primary shoots to ensure they would be vertical at the time of collection. Primary shoot tips from upright (t0) plants and plants after 45 minutes of gravistimulation in the dark by a 90° reorientation (t45 plants) were collected on the same day. The t0 collections were at ~10 AM and t45 collections were at ~10:45 AM (having been reoriented at ~10 AM). From all plants, the upper most ~1 cm of primary shoot apical tissue (minus any leaves, flowers, or opening buds) was harvested and immediately frozen in liquid nitrogen. When t45 plants were reoriented to 90 degrees, their shoots were supported by microfuge tube racks to keep them in the same position, relative to the rosette, as when upright.
RNA extraction for transcriptomes, RNA sequencing and transcriptome analysis
Total RNA was extracted from apical tissues using the Qiagen Plant Mini RNeasy kit (Germantown, MD) including a DNAse treatment with Invitrogen Turbo DNA-free (Carlsbad, CA). Approximately 2 μg of total RNA for each sample was sent to the Cornell Weill Medical Genomics Resources Core Facility (New York, NY, USA) where RNA TruSeq. 50 bp unpaired barcoded libraries were prepared for each and sequenced with nine libraries per lane on an Illumina HiSeq. 2000. Raw sequencing data, now available in the NCBI GEO database under accession GSE147254, was uploaded to CLC Genomics Workbench Version 9.5.3 (Qiagen, Redwood City, CA) and trimmed based on a QC limit of 0.05, an ambiguity max of 2 nucleotides, and a minimum length of 40 nucleotides. The number of reads for each tissue sample before and after trimming can be found in Supplementary Table S3. Using the CLC RNAseq analysis tool trimmed reads were aligned to the arabidopsis genome (TAIR10) with the following parameters: 2 maximum mismatches, a minimum length fraction of 0.9, a minimum similarity fraction of 0.8, a nonspecific match limit of 10, a minimum exon coverage fraction of 0.2, and 10 minimum reads. Expression values were reported as RPKM. Next, the RNAseq Experiment function was used to perform differential expression analysis. Five separate experimental comparisons were performed. A multiple comparison of all three types of t0 plants (Col, tac1, and lazy1) with three biological replicates for each. A multiple comparison of the t45 samples for all three plants, which included 4 biological replicates of Col and lazy1 and 5 biological replicates for tac1. Individual analyses, comparing t0 to t45 samples of the same genome were also performed. All five experimental analyses were normalized by quantiles and then a proportion-based statistical analysis with Baggerley’s test as performed. Bonferroni corrected p-values and FDR-corrected p-values were also determined. Data sets for all comparisons containing DEGs with an FDR p-value correction ≤ 0.05 were then annotated using TAIR10 gene info and descriptions. For each of the five comparisons, the normalized relative fold-change values for genes with an FDR p-value ≤ 0.05 were used to identify significant MapMan Categories using the BAR Classification SuperViewer (https:bar.utoronto.ca)48,49.
Video V1. Time-lapse video of the 90° reorientation gravitropism experiment performed on May 5th, 2014. Video covers ~39 hours immediately following reorientation and is sped up 50×, with each image duration lasting 0.02 seconds. Shoot circumnutation is visible for some plants within the first 10 seconds (~2.5 hours into the experiment) and increases throughout the duration of the video. A key indicating the genotype of each plant can be found in Supplementary Figure S3.
References
Hollender, C. A. & Dardick, C. Molecular basis of angiosperm tree architecture. New Phytol. 206, 541–556 (2015).
Hill, J. L. & Hollender, C. A. Branching out: new insights into the genetic regulation of shoot architecture in trees. Curr. Opin. Plant Biol. 47, 73–80 (2019).
Dardick, C. et al. PpeTAC1 promotes the horizontal growth of branches in peach trees and is a member of a functionally conserved gene family found in diverse plants species. Plant J. 75, 618–630 (2013).
Yoshihara, T., Spalding, E. P. & Iino, M. AtLAZY1 is a signaling component required for gravitropism of the Arabidopsis thaliana inflorescence. Plant J. 74, 267–279 (2013).
Dong, Z. et al. Maize LAZY1 mediates shoot gravitropism and inflorescence development through regulating auxin transport, auxin signaling, and light response. Plant Physiol. 163, 1306–1322 (2013).
Guseman, J. M., Webb, K., Srinivasan, C. & Dardick, C. DRO1 influences root system architecture in Arabidopsis and Prunus species. Plant J. 89, 1093–1105 (2017).
Yoshihara, T. & Spalding, E. P. LAZY genes mediate the effects of gravity on auxin gradients and plant architecture. Plant Physiol. 175, 00942.2017 (2017).
Taniguchi, M. et al. The Arabidopsis LAZY1 family plays a key role in gravity signaling within statocytes and in branch angle control of roots and shoots. Plant Cell 29, tpc.00575.2016 (2017).
Uga, Y. et al. Control of root system architecture by DEEPER ROOTING 1 increases rice yield under drought conditions. Nat. Genet. 45, 1097–102 (2013).
Ge, L. & Chen, R. Negative gravitropism in plant roots. Nat. Plants 2, 17–20 (2016).
Ashraf, A. et al. Evolution of Deeper Rooting 1-like homoeologs in wheat entails the C-terminus mutations as well as gain and loss of auxin response elements. PLOS ONE 14, e0214145 (2019).
Ge, L. & Chen, R. Negative gravitropic response of roots directs auxin flow to control root gravitropism. Plant Cell Environ. 42, 2372–2383 (2019).
Yu, B. et al. TAC1, a major quantitative trait locus controlling tiller angle in rice. Plant J. 52, 891–8 (2007).
Ku, L. et al. Cloning and characterization of a putative TAC1 ortholog associated with leaf angle in maize (Zea mays L.). PLoS ONE 6, e20621 (2011).
Zhao, H. et al. Natural variation and genetic analysis of the tiller angle gene MsTAC1 in Miscanthus sinensis. Planta 240, 161–175 (2014).
Xu, D. et al. PzTAC and PzLAZY from a narrow-crown poplar contribute to regulation of branch angles. Plant Physiol. Biochem. 118, 571–578 (2017).
Hollender, C. A. et al. Alteration of TAC1 expression in Prunus species leads to pleiotropic shoot phenotypes. Hortic. Res. 5, 26 (2018).
Waite, J. M. & Dardick, C. TILLER ANGLE CONTROL 1 modulates plant architecture in response to photosynthetic signals. J. Exp. Bot. 69, 4935–4944 (2018).
González-Arcos, M. et al. A loss-of-function allele of a TAC1-like gene (SlTAC1) located on tomato chromosome 10 is a candidate for the Erectoid leaf (Erl) mutation. Euphytica 215, 95 (2019).
Li, P. et al. LAZY1 controls rice shoot gravitropism through regulating polar auxin transport. Cell Res. 17, 402–410 (2007).
Yoshihara, T. & Iino, M. Identification of the gravitropism-related rice gene LAZY1 and elucidation of LAZY1-dependent and -independent gravity signaling pathways. Plant Cell Physiol. 48, 678–688 (2007).
Uga, Y., Okuno, K. & Yano, M. Dro1, a major QTL involved in deep rooting of rice under upland field conditions. J. Exp. Bot. 62, 2485–2494 (2011).
Howard, T. P. et al. Identification of the maize gravitropism gene lazy plant1 by a transposon-tagging genome resequencing strategy. PLoS ONE 9, e87053 (2014).
Arai-Sanoh, Y. et al. Deep rooting conferred by DEEPER ROOTING 1 enhances rice yield in paddy fields. Sci. Rep. 4, 5563 (2015).
Abe, K., Takahashi, H. & Suge, H. Localization of Cells Containing Sedimented Amyloplasts in the Shoots of Normal and Lazy Rice Seedlings. Biological Sciences in Space 8, 221–225 (1994).
Godbolé, R. et al. The lazy mutation in rice affects a step between statoliths and gravity-induced lateral auxin transport. Plant Biol. 1, 379–381 (1999).
Dong, Z. et al. Maize LAZY1 Mediates Shoot Gravitropism and Inflorescence Development through Regulating Auxin Transport, Auxin Signaling, and Light Response. PLANT Physiol. 163, 1306–1322 (2013).
Sasaki, S. & Yamamoto, K. T. Arabidopsis LAZY1 is a peripheral membrane protein of which the carboxy-terminal fragment potentially interacts with microtubules. Plant Biotechnol. 32, 103–108 (2015).
Yoshihara, T. & Spalding, E. P. Switching the direction of stem gravitropism by altering two amino acids in AtLAZY1. Plant Physiol. pp.01144.2019 (2019) doi:10.1104/pp.19.01144.
Furutani, M. et al. Polar recruitment of RLD by LAZY1-like protein during gravity signaling in root branch angle control. Nat. Commun. 11, 76 (2020).
Zhang, N. et al. A core regulatory pathway controlling rice tiller angle mediated by the LAZY1-dependent asymmetric distribution of auxin. Plant Cell tpc.00063.2018, https://doi.org/10.1105/tpc.18.00063 (2018).
Werner, D. J. & Chaparro, J. X. Genetic interactions of pillar and weeping peach genotypes. HortScience 40, 18–20 (2005).
Dong, H. et al. A novel tiller angle gene, TAC3, together with TAC1 and D2 largely determine the natural bariation of tiller angle in rice cultivars. PLoS Genet. 12, 1–21 (2016).
Wang, L. et al. The isolation of the IGT family genes in Malus × domestica and their expressions in four idiotype apple cultivars. Tree Genet. Genomes 14, 46 (2018).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Tworkoski, T., Miller, S. & Scorza, R. Relationship of Pruning and Growth Morphology with Hormone Ratios in Shoots of Pillar and Standard Peach Trees. J. Plant Growth Regul. 25, 145–155 (2006).
Reches, S., Leshem, Y. & Wurzburger, J. On hormones and weeping: asymmetric hormone distribution and the pendulous growth habit of the weeping mulberry, morus alba var. Pendula. New Phytol. 73, 841–846 (1974).
Bishop, G., Sakakibara, H., Seo, M. & Yamaguchi, S. Biosynthesis of Hormones. in Biochemisctry & Molecular Biology of Plants 769–833 (Wiley Blackwell, 2015).
Díaz, M., Achkor, H., Titarenko, E. & Martínez, M. C. The gene encoding glutathione-dependent formaldehyde dehydrogenase/GSNO reductase is responsive to wounding, jasmonic acid and salicylic acid. FEBS Lett. 543, 136–139 (2003).
Mudgil, Y. et al. Photosynthate Regulation of the Root System Architecture Mediated by the Heterotrimeric G Protein Complex in Arabidopsis. Front. Plant Sci. 7 (2016).
Scott, I. M., Clarke, S. M., Wood, J. E. & Mur, L. A. J. Salicylate Accumulation Inhibits Growth at Chilling Temperature in Arabidopsis. Plant Physiol. 135, 1040–1049 (2004).
Nozue, K. et al. Network Analysis Reveals a Role for Salicylic Acid Pathway Components in Shade Avoidance. Plant Physiol. 178, 1720–1732 (2018).
Li, H.-Y., Wei, W. & Li, Y. Roles of Salicylic Acid-responsive Cis-acting Elements and W-boxes in Salicylic Acid Induction of VCH3 Promoter in Transgenic Tobaccos. Acta Biochim. Biophys. Sin. 38, 46–52 (2006).
Hwang, S.-H., Lee, I. A., Yie, S. W. & Hwang, D.-J. Identification of an OsPR10a promoter region responsive to salicylic acid. Planta 227, 1141–1150 (2008).
Li, Z. et al. OsBRXL4 Regulates Shoot Gravitropism and Rice Tiller Angle through Affecting LAZY1 Nuclear Localization. Mol. Plant 12, 1143–1156 (2019).
Belknap, W. R., Rockhold, D. R. & McCue, K. F. pBINPLUS/ARS: an improved plant transformation vector based on pBINPLUS. BioTechniques 44, 753–756 (2008).
Zeng, W. et al. A Genetic Screen Reveals Arabidopsis Stomatal and/or Apoplastic Defenses against Pseudomonas syringae pv. tomato DC3000. PLOS Pathog. 7, e1002291 (2011).
Thimm, O. et al. mapman: a user-driven tool to display genomics data sets onto diagrams of metabolic pathways and other biological processes. Plant J. 37, 914–939 (2004).
Provart, N. & Zhu, T. A browser-based Functional Classification SuperViewer for Arabidopsis genomics. Currents in Computational Molecular Biology 2003, 271–272 (2003).
Acknowledgements
The authors thank Allison Brown, Erik Miller, Emma Acly, and Michaela Dunn at USDA-ARS Appalachian Fruit Research Station as well as Faith Hatt and Andrea Kohler at Michigan State university for help with miscellaneous plant care, genotyping, and/or data collection. We also would thank Drs. Takeshi Yoshihara and Edgar Spalding for kindly sharing promAtLAZY::GUS seed. Lastly, we are grateful to our anonymous reviewers, whose constructive feedback significantly improved this manuscript. This work was supported by Agriculture and Food Research Initiative Competitive grant 10891264 from the United States Department of Agriculture National Institute of Food and Agriculture, the National Science Foundation grant number 1339211, the United States Department of Agriculture National Institute of Food and Agriculture HATCH project 1013242, the Michigan State University Department of Horticulture, and Michigan State University AgBioResearch.
Author information
Authors and Affiliations
Contributions
C.A.H., C.D. and J.L.H. wrote the manuscript. C.A.H., J.L.H., J.W. and performed experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hollender, C.A., Hill, J.L., Waite, J. et al. Opposing influences of TAC1 and LAZY1 on Lateral Shoot Orientation in Arabidopsis. Sci Rep 10, 6051 (2020). https://doi.org/10.1038/s41598-020-62962-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-62962-4
This article is cited by
-
IGT/LAZY genes are differentially influenced by light and required for light-induced change to organ angle
BMC Biology (2024)
-
Different contributions of PROG1 and TAC1 to the angular kinematics of the main culm and tillers of wild rice (Oryza rufipogon)
Planta (2024)
-
Genome-wide identification of peanut IGT family genes and their potential roles in the development of plant architecture
Scientific Reports (2023)
-
Photoperiod and gravistimulation-associated Tiller Angle Control 1 modulates dynamic changes in rice plant architecture
Theoretical and Applied Genetics (2023)
-
Analysis of controlling genes for tiller growth of Psathyrostachys juncea based on transcriptome sequencing technology
BMC Plant Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.