Abstract
Laboratory strains, cell lines, and other genetic materials change hands frequently in the life sciences. Despite evidence that such materials are subject to mix-ups, contamination, and accumulation of secondary mutations, verification of strains and samples is not an established part of many experimental workflows. With the plummeting cost of next generation technologies, it is conceivable that whole genome sequencing (WGS) could be applied to routine strain and sample verification in the future. To demonstrate the need for strain validation by WGS, we sequenced haploid yeast segregants derived from a popular commercial mutant collection and identified several unexpected mutations. We determined that available bioinformatics tools may be ill-suited for verification and highlight the importance of finishing reference genomes for commonly used laboratory strains.
Similar content being viewed by others
Introduction
The frequent transfer of genetic materials between life science organizations introduces opportunities for quality control issues. Genetic mutations accumulate naturally over time, and human errors in labeling and sample preparation are unavoidable. Anecdotally, it is not uncommon for researchers to complain of samples exhibiting unexpected behaviors, only to later discover that the genetic material they’re working with is not as expected.
Laboratory strains, cell lines, and mutant collections exhibit considerable nucleotide variation and background mutations even among lines thought to be isogenic1,2,3,4. Despite a growing awareness, cell-line contamination and misidentification are persistent problems, particularly in mammalian cell research5,6,7,8,9. Comparably, much less attention has been paid to the potential for similar issues in non-mammalian samples. Yet even commonly used plasmids have been shown to vary dramatically from their published sequence10. The problem of plasmid verification has been addressed through the development of a web-based application for assembly of Sanger sequencing reads and alignment of the assembled plasmids with a reference11. Strain verification by a similar method will be orders of magnitude more challenging.
The methods currently used to verify samples/strains are biased towards a particular goal. For instance, diagnostic techniques such as PCR, targeted sequencing, or restriction enzyme-based methods are often used to identify whether or not a marker gene or known sequence variant is present, or for analysis of variable repeat regions such as in 16 s rRNA profiling12,13. These approaches are limited to particular regions of the genome or are insufficiently sensitive for capturing many types of sequence variations2,14,15.
In addition to wasted time and reagents, undetected genetic variation can lead to severe consequences including delays in publishing or patenting and misplaced conclusions that result in product recalls or retractions16. Given reports that the life sciences are facing a reproducibility crisis17, it is more important than ever for researchers to verify the samples and strains they work with.
As the cost and turnaround time of next generation sequencing continues to decrease, sample and strain verification by whole genome sequencing (WGS) is becoming a more feasible approach13. Many tools have been developed for assembling sequenced genomes and detecting variants by aligning sequencing reads to a reference genome18, but these tools have been largely developed and validated using human sequencing data. The same tools may not perform well when analyzing microbial sequencing data due to differing ploidy, genome size, and mutation rates19. The applicability of assembly and, especially, variant calling tools to microbial sample and strain verification has not been thoroughly explored.
In order to identify the practical obstacles that must be overcome to ultimately implement WGS as a regular part of genetics workflows, we used haploid yeast strains with an unexpected phenotype derived from a mutant collection as a test case.
Results
Yeast strain sequencing
As part of a series of yeast cell cycle experiments, we crossed two mutant lines from a knockout collection20 to produce cln3∆ mbp1∆ double mutants in S. cerevisiae. mbp1 and cln3 knockouts are each individually known to result in an increased critical cell size at the start of S phase21,22,23. When the cln3∆::kanMX and mpb1∆::natMX mutant lines were crossed, half of the double mutant progeny had a wild-type G1 cell size. When we examined the cln3 mutant strain, it too had a wild-type-like cell size. Three of the cln3∆ mbp1∆ double mutant segregants were sequenced using an Illumina MiSeq sequencer: one segregant, 1691, exhibited the unexpected wild-type-like phenotype, the others, 1693 and 1694 (which exhibited the mutant phenotype) were used for comparison.
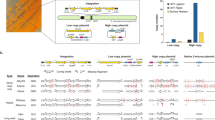
We conducted three different sequence analyses (Fig. 1): (A) Reads were assembled both de novo and against the S288C reference genome. (B) Variant finding tools were used to call single nucleotide polymorphisms (SNPs) that varied between strains 1691 and 1693. (C) Copy number variant tools were used to confirm the presence of strain-specific deletions and marker gene insertions (INDELs) and check for additional structural variations.
Data analysis pipeline. Software tools compared for each type of analysis are outlined in white. Icons are used to denote whether the analysis was conducted against the S288C reference genome (flag), the BY4741 draft genome (triangle), and or de novo (star). (A) Reads were assembled using SPAdes (S288C reference-based and de novo assembly) and SOAPdenovo (de novo only). (B) SNPs were analyzed using GATK and Samtools against the S288C reference strain and the BY4741 genetic background. (C) INDELs and other structural variants were analyzed against the S288C reference using cnv.kit, CNVnator, and Breakdancer. Software versions and parameters used are detailed in Supplementary Information.
Because variant calling via genome alignment and variant calling via mapped reads can result in different types of errors19, we tested both approaches. For each stage of the analysis, at least two different popular bioinformatic tools were tested. We selected tools partly based on Pabinger, et al.’s survey of 205 tools18. We tested additional tools to those described herein, but only results from the best performing tools were included in the manuscript. Known and unknown variants were confirmed visually (see Supplementary Figs. 1, 2 for examples) by aligning trimmed reads to the S288C reference genome using Integrated Genomics Viewer (IGV) software (http://www.broadinstitute.org/igv).
Assembling a genome for verification
An ideal approach to sample verification by WGS would be to sequence the sample, assemble the genome, and then compare the assembled genome to the exact reference genome (the genetic background plus any known variations). Unfortunately, there is not a finished reference genome available for the genetic background used in our analysis (BY4741), despite the fact that it is a commonly used laboratory strain. We thus conducted reference-based assemblies using the closely related S288C genome.
Analyzing a genome assembled de novo has a number of additional advantages. For instance, reads that do not align to the reference, such as those for a marker gene insertion or transgene, might be trimmed from the analysis. As such, we repeated the assemblies de novo.
The quality of each assembly was compared using the QUAST quality assessment tool for genome assemblies24 (Table 1 and Supplementary Table 2). In all metrics, the reference-based assembly exceeded the de novo assemblies, and SPAdes25 out-performed SOAPdenovo226. Although comparable to the previously published BY4741 draft genome27, the quality metrics of all assemblies varied markedly from the S288C reference in number of contigs and N50 values.
Cost is a major barrier to using WGS for sample verification and cost directly relates to coverage. In this study, strains were sequenced on a MiSeq for a total of ~2 million, 250 bp paired-end reads, corresponding to a predicted 80x coverage of the 12 Mb yeast genome. Coverage metrics produced with Picard tools show that ~95% of genome was covered with at least 30 reads in all three samples (http://broadinstitute.github.io/picard/).
To determine the minimal cost for which a comparable assembly might have been achieved, we repeated the SPAdes assemblies simulating varying levels of coverage by randomly subsampling the read library. For the majority of the quality assembly metrics compared for both reference-based and de novo assembly (Fig. 2 and Supplementary Tables 3 and 4 respectively), the assembly quality began plateauing at around 500,000 read pairs.
Assembly subsampling analysis. Comparison of the effect of sequencing depth on various metrics (from top to bottom: N50, total number of contigs, and genome fraction) for assemblies against the S288C reference (left) and de novo (right). In each case, the X-axis is number of read pairs. The red, green, and blue lines correspond to reads from 1691, 1693, and 1694 respectively.
Thus, for a haploid yeast genome, a predicted coverage of 10x-30x is sufficient for a draft genome assembly and investing in NGS coverage beyond 10x may not yield notably better assemblies. To generate a more complete genome, it may thus be more prudent to invest in long read sequencing such as PacBio and Oxford Nanopore technologies in order to conduct hybrid assemblies28,29,30,31, as opposed to increasing short read depth.
A meaningful direct comparison of our assembled genomes with the reference would require a more complete assembly than we were able to accomplish using short reads alone. As such, we conducted the remainder of our analysis by aligning trimmed reads to the reference genome.
Variant calling from WGS data
The selection of tools for variant calling can drastically influences the results18,32. We called variants using33 and Samtools34 and attempted to confirm each variant visually by inspecting the reads in IGV (see Supplementary Fig. 1 for an example). To simplify the analysis, we focused on only those variants that were discordant between 1691 and 1693.
As has been previously observed32, Samtools identified substantially fewer SNPs than GATK (Supplementary Tables 5 vs 6, respectively). This is likely due to the fact that GATK HaplotypeCaller does local reassembly in regions with genomic variation35.
Table 2 highlights all of the SNPs that could be confirmed by aligning the reads to the S288C reference genome in IGV. All of these were identified by both GATK and Samtools. The nine variants called by Samtools that were not also called by GATK (unshaded rows in Supplementary Table 5) could be confirmed for both strain 1691 and 1693 by visual inspection of the aligned reads and thus were not truly discordant. The 58 variants identified by GATK but not Samtools (unshaded and lightly shaded rows in Supplementary Table 6) could all either be confirmed for both strains (not discordant) or for neither (possibly not true SNPs). Only two of the variants that were called by both GATK and Samtools could not be confirmed by inspecting the reads in IGV. The combination of GATK and Samtools, as has previously been proposed32, was thus a valuable approach for filtering out noise in this case.
Analysis with both tools was repeated using the draft genome for the genetic background strain of the parent, BY4741 (Supplementary Tables 7 and 8). This resulted in a substantially longer list of additional discordant SNPs, none of which could be validated by visual inspection of the reads in IGV.
The analysis using Samtools was also repeated using contigs generated by the SPAdes de novo assembly (Supplementary Table 9). This also resulted in a longer list of SNPs, but the quality of the calls could not be assessed, because the tool is designed to be used on dozens of reads, not a single contig. This common variant-finding tool is thus not well suited for use with draft genomes or assembled contigs, both of which could facilitate sample and strain verification.
Of the confirmed SNPs, half occurred in cell cycle related genes (Table 2). None of these are likely to account for the unexpected phenotype observed for 1691. The two variants that were specific to 1691 were located in genes DYN1 and SLA2, both of which are important for cytoskeletal functions36,37. dyn1 or sla2 loss-of-function slows the cell cycle and would not relieve the cell cycle delay in G1 caused by the cln3 mutation36,37. However, any of these SNPs could potentially impact the interpretation of cell-cycle experiments.
INDEL analysis
The segregants sequenced were expected to differ from the S288C reference strain at several auxotrophic marker deletions38, at their mating type genes (the reference strain is MATα mating type while 1691 and 1693 are MATa), and by marker gene insertions at CLN3 and MBP1 (shaded cells in Table 3). To confirm the presence or absence of these structural changes in 1691 and 1693 we performed copy-number variant analyses using three different tools: Breakdancer39, CNVkit40, and CNVnator41.
Table 3 lists all of the INDELs that could be confirmed visually by aligning the reads to the S288C reference genome in IGV (see Supplementary Fig. 2 for an example, duplications would be difficult to confirm in this manner). CNVkit found only one of the eight known INDELs in the two segregants but was one of the few tools to identify an event in the ATPase pump genes ENA1, ENA2, and ENA5. Without exploring the parameters to identify more known variants, Breakdancer and CNVnator performed comparably. It was only when we reduced the CNVnator search window down to just 20 bp that we were able to identify most of the known INDELs, even though most of the deletions are hundreds of bp in length (Table 3). These parameters also resulted in by far the longest list of variant calls (Supplementary Table 12 versus Supplementary Tables 10, 11, 13), most of which occurred in known repetitive regions (lightly shaded rows in Supplementary Table 12).
Only one of the unexpected INDELs, which CNVnator identified as a 1660 bp deletion in WHI5, differed between 1691 and 1693. Deletion of WHI5 has been previously shown to partially suppress the large cell phenotype seen in cln3 mutants42,43,44. It is, therefore, very likely that the unexpected wild-type-like phenotype observed for 1691 is due to suppression by a mutation in WHI5. A visual inspection of the aligned reads at WHI5 (Fig. 3) suggested that there is a transposon insertion interrupting the WHI5 reading frame in 1691. The fact that half of the progeny in the cross exhibited the same cell size phenotype as 1691 suggests that the transposon insertion was already present in the cln3∆::kanMX mutant parent obtained from the knock-out collection.
Reads aligned at the WHI5 locus. Each elongated block arrow is a different read. Reads that are colored (not grey) indicate that the mate pair matches a different location in the genome. For each of the colored reads highlighted in the red dashed circle, the mate pair matches a transposon (TY elements).
Discussion
Using WGS, we identified eight unexpected SNPs and one unexpected INDEL that differed between segregants derived from a commercial mutant collection. Because this mutant collection is popular in studies of the budding yeast cell cycle, it is pertinent that four of the SNPs identified occurred in cell cycle related genes. This test case demonstrates the value of verifying strains and cell lines from mutant collections by WGS. While this approach was successful in identifying the mutation that was likely the cause for an unexpected phenotype, there may be more changes to the genome that were missed. Clearly, unexpected mutations in common laboratory cell lines cannot be ignored, but the technology needed to get a clear vision of the magnitude of the problem is underdeveloped.
The variant-finding tools used in this analysis were not ideally suited to verification workflows. Most data analysis pipelines, including those described here, rely on ad-hoc or heuristic decision points that require an advanced understanding of the software tools used for analysis19. Analyzing the results required manually validating the calls by visualizing the reads, as well as looking up the function of each individual gene – processes that are tedious, time consuming, and potentially error-prone. Additionally, the SNPs/INDELs called differed dramatically depending on the tools and parameters used. None of the tools and parameters tested successfully identified all of the known INDELs (Table 3). It was only when we adjusted the parameters to find the known INDELS, that we identified a large transposon insertion in an important gene. In conclusion, commonly used software tools could not reliably return expected outcomes, were individually too narrow in focus, and collectively too sensitive to parameters to be integrated into a consistent pipeline for verification by WGS.
Before WGS can be used for routine sample and strain validation, genome finishing also needs to be streamlined and made more affordable, such that reference genomes are available for all commonly used laboratory strains. The shortcomings of using short read sequencing in genome assembly have been well reported45. In the described use case, the use of short reads significantly hampered our ability to resolve repetitive regions of the genome. This is evident in the fact that most contigs were flanked by transposons. Read alignment across repetitive regions was also ambiguous (for example, Supplementary Fig. 2), complicating the variant analysis; many of variants called were located within transposable elements and telomeres, and near genes encoding tRNAs and ribosomal RNAs (lightly shaded rows in Supplementary Tables 10–13). PacBio sequencing would likely have provided an improved resolution, but it remains prohibitively expensive for verification purposes. And while Oxford Nanopore sequencers are affordable, the reagents and flow cells are costly, and the associated software and algorithms are even less accessible to life science researchers without bioinformatics expertise.
In light of the findings presented in this paper, we would like to suggest a call to action for the development of tools and approaches specifically focused on verification by WGS, in order to ultimately implement WGS as a regular part of genetics workflows, such that all genetic materials are verified by WGS prior to experimentation to improve experimental reproducibility. For instance, it is important that variant finding tools be developed, trained, and validated with microbial sequencing data specifically19.
The main shortcoming in the described workflow was its inability to resolve repetitive regions such as transposons. Tools for finding transposons specifically have been developed, but are not routinely employed46. Pipelines incorporating tools with different strengths should be used to overcome the false positive and false negatives associated with a particular approach. Variability and repetitive sequences (such as at telomeres, transposons, and ribosomal RNA genes), on the one hand, complicate analysis by WGS, but, on the other hand, emphasize the importance of frequently verifying strains, because the genome is a living dynamic structure, not a rigid set of permanent instructions.
Methods
Generating and phenotyping yeast mutants
The mutant collection from which the parental strains used in this study were obtained was generated using background strains BY4741 and BY4742, which differ only in their mating type and the auxotrophic markers MET15 and LYS238. Both were derived from Saccharomyces cerevisiae strain FY2 which is a direct descendent of S288C. Both BY4741 and BY4742 are known to differ from S288C by the deletion of four auxotrophic markers. According to a recently prepared draft genome, BY4741 additionally differs from S288C by fewer than 5 SNPs per 100,000 bp27.
One of the haploid parents obtained from the collection has the CLN3 ORF replaced with a marker for G418 resistance (MATa cln3∆::kanMX MBP1). The other has the MBP1 ORF replaced with a marker for nourseothricin resistance by marker switching of the commercial deletion strain (MATα CLN3 mbp1∆::natMX)47. These strains were crossed to yield the haploid progeny we analyzed by whole-genome sequencing. The list of strains used in this study is reported in Table 4.
Sequencing
Each of the strains was sequenced on a Miseq for a total of ~2 million, 250 bp paired-end reads, corresponding to a predicted 80x coverage of the 12 Mb yeast genome. Coverage metrics produced with Picard tools show that ~95% of genome was covered with at least 30 reads in all three samples (http://broadinstitute.github.io/picard/). In addition, the percentage of aligned reads versus total reads was in the range of 97–98% for all three samples.
Analysis
The following software tools were used in the described analysis: FastQC (https://www.bioinformatics.babraham.ac.uk/projects/fastqc/), Trimgalore! (https://www.bioinformatics.babraham.ac.uk/projects/trim_galore/), cutadapt (https://cutadapt.readthedocs.io/en/stable/), bwa (http://bio-bwa.sourceforge.net), Picard (https://broadinstitute.github.io/picard/), ENSEMBL (http://ensembl.org), CNVnator (https://github.com/abyzovlab/CNVnator), Breakdancer (http://breakdancer.sourceforge.net/), cnv.kit (https://cnvkit.readthedocs.io/en/stable/), Samtools (http://www.htslib.org/doc/samtools.html), BCFtools (https://samtools.github.io/bcftools/bcftools.html), Genome Analysis Toolkit (GATK) (https://software.broadinstitute.org/gatk/), snpEFF (http://snpeff.sourceforge.net/), SOAPdenovo2 (http://soap.genomics.org.cn/soapdenovo.html), SPAdes (http://cab.spbu.ru/software/spades/), and QUAST (http://bioinf.spbau.ru/quast), BLAST (http://doi.org/10.1186/1471-2105-10-421). Default parameters were used unless otherwise noted.
FastQC was used to calculate and visualize sequence quality metrics before and after trimming with Trimgalore!. Samples were aligned to Saccharomyces cerevisiae genome assembly R64-1-1 (Ensembl release 92), corresponding to strain S288C (baker’s yeast), using BWA (v. 0.7.15) with default parameters. Alignment quality of the resulting bam files was assessed using Picard (v.2.9).
CNVnator (v0.3.3) was used for structural variant calling with a bin size of 20 and 100. In addition, copy number variation was assessed with CNVkit (v0.9.3) and Breakdancer (v. 1.3.6).
CNVkit was run with automatic binning and with p-value threshold of 0.000005 for accepting segments and their breakpoints. Copy numbers were called with log2 ratio thresholds of −1.0000000, 0.5849625, 1.3219281, 1.8073549, and 2.1699250 for copy numbers 0 to 4, with thresholds being the upper limits of log2 coverage ratio for each copy number. Thresholds were calculated by adding 0.5 to the integer copy number value for rounding, dividing by ploidy (1), and log2 transforming the result.
Breakdancer was run with default parameters and configuration files generated using script bam2cfg.pl, included in its distribution.
Variant calling was run both with the GATK pipeline and a pipeline consisting of Samtools (v1.8) and Bcftools (v.1.8).
The GATK pipeline was built on GATK version v4.beta.5, except for function CombineGVCFs, which was run with GATK version v3.8, as there was no working version of this function in GATK 4 at the time of setting up the analyses. GATK was run with default parameters and using GATK HaplotypeCaller for calling variants with —sample_ploidy set to 1.
Variant calling with Samtools and Bcftools was run with —ploidy set to 1 and using multi-allelic calling mode (Bcftools flag -m).
Variants called with GATK and Bcftools were annotated using snpEFF (v.4.3 T), a software for variant annotation and predicting effects. Samples aligned to R64-1-1 were annotated using a database for strain S288C provided by snpEFF authors. Samples aligned to BY4741 were annotated using snpEFF database custom built on annotation files downloaded from Saccharomyces Genome Database48.
De novo assemblies were annotated by BLASTing assembled contigs against the reference genome of strain S288C. In addition, variant calling was performed using our de novo assembled genome as a reference. Called variants were annotated by BLASTing their flanking sequences against the reference genome of S288C to find corresponding gene annotations. Function blastn of BLAST toolbox (v2.7.1+) was run with following parameters: -outfmt 6 -max_target_seqs. 1 -max_hsps 1 -num_threads 10 -strand plus.
De novo assemblies were performed with SOAPdenovo2 (v. 2.04) and SPAdes (v. 3.9.0), with SPAdes used for downstream analyses and assembly by subsampling. Reference-based de novo assembly, with S288C chromosome sequences used as trusted contigs for i.a. gap closure and repeat resolution, with and without subsampling was also performed with SPAdes (v. 3.9.0).
Qualities of all assemblies were assessed using QUAST, with the reference genome of strain S288C used for benchmarking.
Data availability
All data needed to repeat the analysis described in this manuscript as well as descriptions of the software tools and parameters used is available in the GitHub repository peccoud/strain-verification.
References
Ajjawi, I., Lu, Y., Savage, L. J., Bell, S. M. & Last, R. L. Large-scale reverse genetics in Arabidopsis: case studies from the Chloroplast 2010 Project. Plant physiology 152, 529–540 (2010).
Kleensang, A. et al. Genetic variability in a frozen batch of MCF-7 cells invisible in routine authentication affecting cell function. Scientific reports 6, 1–11 (2016).
Sarin, S. et al. Analysis of multiple ethyl methanesulfonate-mutagenized Caenorhabditis elegans strains by whole-genome sequencing. Genetics 185, 417–430 (2010).
Watkins-Chow, D. E. & Pavan, W. J. Genomic copy number and expression variation within the C57BL/6J inbred mouse strain. Genome research 18, 60–66 (2008).
Lorsch, J. R., Collins, F. S. & Lippincott-Schwartz, J. Fixing problems with cell lines. Science 346, 1452–1453 (2014).
Masters, J. R. End the scandal of false cell lines. Nature 492, 186–186 (2012).
Freedman, L. P. et al. Reproducibility: changing the policies and culture of cell line authentication. Nature methods 12, 493–497 (2015).
Capes‐Davis, A. et al. Check your cultures! A list of cross‐contaminated or misidentified cell lines. International journal of cancer 127, 1–8 (2010).
Lacroix, M. Persistent use of “false” cell lines. International journal of cancer 122, 1–4 (2008).
Peccoud, J. et al. Targeted Development of Registries of Biological Parts. Plos one 3, e2671 (2008).
Wilson, M. L. et al. Sequence verification of synthetic DNA by assembly of sequencing reads. Nucleic Acids Research 41, e25 (2013).
Janda, J. M. & Abbott, S. L. 16S rRNA gene sequencing for bacterial identification in the diagnostic laboratory: pluses, perils, and pitfalls. Journal of clinical microbiology 45, 2761–2764 (2007).
Almeida, J. L., Cole, K. D. & Plant, A. L. Standards for cell line authentication and beyond. PLos biology 14, e1002476 (2016).
Zischewski, J., Fischer, R. & Bortesi, L. Detection of on-target and off-target mutations generated by CRISPR/Cas9 and other sequence-specific nucleases. Biotechnology advances 35, 95–104 (2017).
Hendel, A., Fine, E. J., Bao, G. & Porteus, M. H. Quantifying on-and off-target genome editing. Trends in biotechnology 33, 132–140 (2015).
Peccoud, J., Gallegos, J. E., Murch, R., Buchholz, W. G. & Raman, S. Cyberbiosecurity: From Naive Trust to Risk Awareness. Trends Biotechnol 36, 4–7 (2018).
Baker, M. 1,500 scientists lift the lid on reproducibility. Nature 533, 452–454 (2016).
Pabinger, S. et al. A survey of tools for variant analysis of next-generation genome sequencing data. Briefings in bioinformatics 15, 256–278 (2014).
Olson, N. D. et al. Best practices for evaluating single nucleotide variant calling methods for microbial genomics. Frontiers in genetics 6, 235 (2015).
Winzeler, E. A. et al. Functional characterization of the S. cerevisiae genome by gene deletion and parallel analysis. science 285, 901–906 (1999).
Bähler, J. Cell-cycle control of gene expression in budding and fission yeast. Annual review of genetics 39, 69–94 (2005).
Shi, L. & Tu, B. P. Acetyl-CoA induces transcription of the key G1 cyclin CLN3 to promote entry into the cell division cycle in Saccharomyces cerevisiae. Proceedings of the National Academy of Sciences 110, 7318–7323 (2013).
Koch, C., Moll, T., Neuberg, M., Ahorn, H. & Nasmyth, K. A role for the transcription factors Mbp1 and Swi4 in progression from G1 to S phase. Science 261, 1551–1557 (1993).
Gurevich, A., Saveliev, V., Vyahhi, N. & Tesler, G. QUAST: quality assessment tool for genome assemblies. Bioinformatics 29, 1072–1075 (2013).
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. Journal of computational biology 19, 455–477 (2012).
Luo, R. et al. SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. Gigascience 1, 2047-217X-1-18 (2012).
Song, G. et al. AGAPE (Automated Genome Analysis PipelinE) for pan-genome analysis of Saccharomyces cerevisiae. Plos one 10, e0120671 (2015).
Antipov, D., Korobeynikov, A., McLean, J. S. & Pevzner, P. A. hybridSPAdes: an algorithm for hybrid assembly of short and long reads. Bioinformatics 32, 1009–1015 (2016).
Au, K. F., Underwood, J. G., Lee, L. & Wong, W. H. Improving PacBio long read accuracy by short read alignment. Plos one 7, e46679 (2012).
Goodwin, S. et al. Oxford Nanopore sequencing, hybrid error correction, and de novo assembly of a eukaryotic genome. Genome research 25, 1750–1756 (2015).
Utturkar, S. M. et al. Evaluation and validation of de novo and hybrid assembly techniques to derive high-quality genome sequences. Bioinformatics 30, 2709–2716 (2014).
Altmann, A. et al. A beginners guide to SNP calling from high-throughput DNA-sequencing data. Human genetics 131, 1541–1554 (2012).
DePristo, M. A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nature genetics 43, 491–498 (2011).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Pirooznia, M. et al. Validation and assessment of variant calling pipelines for next-generation sequencing. Human genomics 8, 14 (2014).
Eshel, D. et al. Cytoplasmic dynein is required for normal nuclear segregation in yeast. Proceedings of the National Academy of Sciences 90, 11172–11176 (1993).
Holtzman, D. A., Yang, S. & Drubin, D. G. Synthetic-lethal interactions identify two novel genes, SLA1 and SLA2, that control membrane cytoskeleton assembly in Saccharomyces cerevisiae. The Journal of cell biology 122, 635–644 (1993).
Baker Brachmann, C. et al. Designer deletion strains derived from Saccharomyces cerevisiae S288C: a useful set of strains and plasmids for PCR‐mediated gene disruption and other applications. Yeast 14, 115–132 (1998).
Chen, K. et al. BreakDancer: an algorithm for high-resolution mapping of genomic structural variation. Nature methods 6, 677–681 (2009).
Talevich, E., Shain, A. H., Botton, T. & Bastian, B. C. CNVkit: genome-wide copy number detection and visualization from targeted DNA sequencing. PLoS computational biology 12, e1004873 (2016).
Abyzov, A., Urban, A. E., Snyder, M. & Gerstein, M. CNVnator: an approach to discover, genotype, and characterize typical and atypical CNVs from family and population genome sequencing. Genome research 21, 974–984 (2011).
De Bruin, R. A., McDonald, W. H., Kalashnikova, T. I., Yates, J. III. & Wittenberg, C. Cln3 activates G1-specific transcription via phosphorylation of the SBF bound repressor Whi5. Cell 117, 887–898 (2004).
Palumbo, P. et al. Whi5 phosphorylation embedded in the G 1/S network dynamically controls critical cell size and cell fate. Nature communications 7, 1–14 (2016).
Schmoller, K. M., Turner, J., Kõivomägi, M. & Skotheim, J. M. Dilution of the cell cycle inhibitor Whi5 controls budding-yeast cell size. Nature 526, 268–272 (2015).
Hardwick, S. A., Deveson, I. W. & Mercer, T. R. Reference standards for next-generation sequencing. Nature Reviews Genetics 18, 473 (2017).
Goerner-Potvin, P. & Bourque, G. Computational tools to unmask transposable elements. Nature Reviews Genetics 19, 688–704 (2018).
Tong, A. H. Y. & Boone, C. In Yeast Protocol 171–191 (Springer, 2006).
Cherry, J. M. et al. SGD: Saccharomyces Genome Database. Nucleic Acids Research 26, 73–79 (1998).
Acknowledgements
The research reported in this publication was supported by NIH Award R01GM078989, NSF Award #1832320 “EAGER: Modeling DNA Manufacturing Processes Using Extensible Attribute Grammars”, NSF Award #1759900 “Collaborative Research: ABI Innovation: Automated Prioritization and Design of Experiments to Validate and Improve Mathematical Models of Molecular Regulatory Systems”, and Colorado State University’s Office of the Vice President for Research Catalyst for Innovative Partnerships Program.
Author information
Authors and Affiliations
Contributions
J.G. wrote the manuscript text, prepared all figures and tables, and managed the project. S.H. conducted the bioinformatics analysis, prepared the GitHub documentation, and generated the graphs used in Figure 2. NA carried out the yeast genetic experiments and prepared the samples for W.G.S. J.P. devised, oversaw the project, and edited the manusccript.
Corresponding author
Ethics declarations
Competing interests
The authors declare the following competing interests: J.P. holds an equity stake in GenoFAB, Inc. a company that may benefit or may be perceived to benefit from the publication of this article.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gallegos, J.E., Hayrynen, S., Adames, N.R. et al. Challenges and opportunities for strain verification by whole-genome sequencing. Sci Rep 10, 5873 (2020). https://doi.org/10.1038/s41598-020-62364-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-62364-6
This article is cited by
-
Genetic interactions derived from high-throughput phenotyping of 6589 yeast cell cycle mutants
npj Systems Biology and Applications (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.