Abstract
In this epidemiologic study, 16 S rRNA sequencing was used to investigate the changes of diversity and composition profile of gut microbiota resulting from long-term exposure to multiple metals, including arsenic (As), cadmium (Cd), cuprum (Cu), lead (Pb), and zinc (Zn). Due to long-term exposure to various metals, the relative abundances of Lachnospiraceae, Eubacterium eligens, Ruminococcaceae UGG-014, Erysipelotrichaceae UCG-003, Tyzzerella 3, Bacteroides, Slackia, italics, and Roseburia were found to become much higher, whereas the abundance of Prevotella 9 presented an opposite trend. Additionally, differences between males and female groups were found, such as the greater richness and evenness of bacteria for men subjected to long-term metal exposure in polluted areas. The changes of men’s microbiomes were more significant as a result of higher daily intake, mining and smelting activity, and living habits. This research presents a new theoretical basis for the correlation between long-term metal exposure and gut health for people living in contaminated areas.
Similar content being viewed by others
Introduction
Recently, the entire gut microecosystem has begun to be regarded as an essential organ in the human body1. The intestinal microbiome plays an important role, and the microorganisms contained in the microbiome are vast in number, consisting of populations of up to 100 trillion2. The intestinal microbiome has been proven to be profoundly responsible for maintaining human health, such as energy metabolism, environmental adaptation, immune adjustment, and even brain functions3,4,5. In addition, some related metabolic diseases (obesity, cirrhosis, hypertension, etc.) have been associated with the changed structure of the microbiome6,7,8.
The daily diet is the most critical factor in maintaining the symbiotic relationship between intestinal flora and the host throughout the host’s life9. An epidemiologic study has confirmed that gut microbiota can respond to various dietary styles. Rural children who consumed low-sugar, high-fiber diets were found to have higher microbial richness and biodiversity compared with European children10. Healey et al. have also found inter-individual variability in gut microbiota response to dietary interventions11. Some other influencing factors include age, sex, and exercise, as well as prebiotic and probiotic agents12,13,14.
16 S ribosomal RNA (rRNA) gene sequence analysis has been widely used to identify bacterial species and perform taxonomy, and massively parallel sequencing (MPS) is also commonly utilized15. There are nine “hypervariable regions” (V1–V9) in the sequence that can show considerable sequence diversity and are regarded as diagnostic assays for different bacteria16. Dethlefsen et al.17 determined the pervasive effects of antibiotics on gut microbiota by tag pyrosequencing of the 16 S rRNA V3 region. By pyrosequencing over 40,000 16 S rRNA gene V4 region amplicons per subject, links between health conditions and intestinal microbiota in elderly Irish subjects were reported18. Additionally, the V3–V4 regions of the 16 S rDNA gene were used in a comparison between patients with acne vulgaris and healthy controls19.
Dietary exposure is the major pathway of the accumulation of potential toxic metals in the body20. Some metals have shown different abilities to change the bacterial profile by altering its diversity and composition, and some host metabolisms have even been found to be altered21. Inorganic As exposure has been found to induce gut microbe perturbations, and even hepatocellular carcinoma22. The inhibited growth rate of the bacteria population, an altered overall gut microbial census, and an impaired gut barrier were found in mice fed with CdCl2 for 3 weeks23. Other toxic metals (Al, Pb, Co, etc.) have also been found to have negative effects on the gut microbiome24. In addition, the deficiency or excess of essential elements such as Zn and Fe have also exhibited the capability to shift gut microbiota composition and function25,26,27.
Hezhang County in Guizhou Province, China is most famous for its Pb-Zn mine28. The mineral mine in this area consists of mountain deposits, which account for over 70% of the plowed soils. Because production and marketing principally occur locally, contaminated crops, vegetables, and animal foodstuff cause mineral metal accumulations in the bodies of local residents within the region29,30. In a previous study by the authors, exposure levels and the harmful health effects of multiple metals in local subjects were reported31. However, the effect of multiple metals on the human microbiome is still unclear. Therefore, this paper reports an investigation of the synergistic effects of multiple metals on human gut microbiota, and fills in the gaps of the epidemiological studies in the crosstalk between heavy metals and intestinal flora.
Results
Individual information for the two study areas
From screening using a questionnaire, a total of 40 healthy residents took part in the stool collection activity (Table 1). Twenty participants (11 female and 9 male) from Maomaochang Village were assigned the group label T (Test), and the remaining participants (13 female and 7 male) from Yemachuan Village were given the group label CK (Control check). Compared to the population of CK, people in T were younger on average (Student’s Test, p < 0.01). The mean body mass index (BMI) value of the two groups was equal. Due to differences of intestinal flora between men and women, four groups divided by gender (labels: TM and CKM, TF and CKF) were used for comparative analysis20. The mean age of men in T was lower than that in CK, and there were no significant differences in body weight. Furthermore, the percentages of smokers and drinkers in both T and TF were larger than those in the corresponding groups from the control village population; nevertheless, the population of male smokers was larger in the control area.
Long-term exposure to various heavy metals changes the diversity of gut microbiota
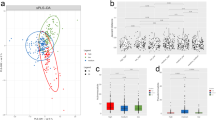
A total of 1,449,804 (1.45 million) 16 S rRNA reads were generated from the stool samples in T, with an average of 74,690 (±5,962 SD) reads in TM and 70,690 (±5,322 SD) reads in TF. A slightly lower number of reads (1,400,222) was detected in CK, with an average of 72,203 (±5,100 SD) reads in CKM and 68,830 (±6,594 SD) reads in CKF. Rarefaction analysis was conducted on the number of OTUs per sample, as presented in Figure S1. The plots tend towards a plateau but do not reach it, suggesting that new OTUs may occur as the number of samples increases. In contrast, the species accumulation curves all reach peaks, suggesting that new species may emerge as the sequencing depth increases, while causing little effect on the diversity of intestinal flora. The analysis of α-diversity metrics revealed the effects of multiple metals on the microbiota (Table 2). There were no significant differences in community richness (Chao1 and ACE indices) between T and CK. However, significantly higher values of the Shannon-Wiener Index, and much lower values of the Simpson Index, were found in group T, suggesting that the multiple metals accumulated in the participants’ bodies, thereby increasing the bacteria richness and evenness (Wilcox Test, p < 0.05). Similarly, differences between the groups (TM and CKM, TF and CKF) were also compared. The microbiota of the TM subjects had observably larger Shannon-Wiener index values compared with those of CKM (Wilcox Test, p < 0.01). Additionally, much lower values of the Simpson Index were also found for TM (Wilcox Test, p < 0.05). These findings all imply that long-term metal exposure increased the richness and evenness of bacteria for men who live in the polluted area. No significant changes were found between the female populations. As another means of identifying changes in the microbiota induced by metal exposure, PCoA was performed for all the samples in this study using the unweighted UniFrac distance method (Fig. 1); a higher clustering of the microbial population in CK than in T, which had greater dispersion, was observed. The same discrete situations were also found in the gender-based groups. These results suggest that the metal exposures changed the gut microbial communities of the local residents, with varying effects on different individuals. The PCA analysis also shows the qualitatively similar results in these groups (Fig. S2).
Metal-induced change in microbial composition
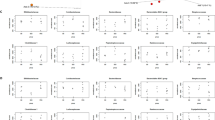
After species annotation analysis, 490 OTUs were divided into the following seven phyla: Bacteroidetes, Firmicutes, Proteobacteria, Cyanobacteria, Actinobacteria, Fusobacteria, and Verrucomicrobia (Fig. 2). Bacteroidetes and Firmicutes were the most dominant taxa of feces both in T and CK, and their sum was over 90%. This result was consistent with previous reports on the main dominant phyla of Bacteroidetes and Firmicutes in Chinese people. However, based on the results of the Kruskal-Wallis test, there were no statistical differences in all phyla between the two groups. The same result also occurred for the comparisons between TM and CKM, and between TF and CKF. At the family level, the two groups (T and CK) had twenty equally dominant bacteria (Fig. 3C). The relative abundances of Lachnospiraceae, Erysipelotrichaceae, Acidaminococcaceae, and Porphyromonadaceae families in T were significantly higher than those in CK (Fig. 3A). Only the abundance of Prevotellaceae showed markedly lower amounts in T (p < 0.05). These five families also showed changes in the comparisons between the gender-based groups, although these changes were not statistically significant. In addition, five genera (Lachnospiraceae, Eubacterium eligens, Ruminococcaceae UGG-014, Erysipelotrichaceae UCG-003, and Tyzzerella 3) presented significantly higher values in T (p < 0.05) (Fig. 3B). At the family level, the two groups (TM and CKM, TF and CKF) had twenty equally dominant bacteria (Fig. 4Aa,Ba). However, there were different changes at the genus level revealed by the comparison between groups classified by gender. Bacteroides, Roseburia, Pseudobutyrivibrio, and Parabacteroides were enriched in TM, although these changes were not statistically significant (Fig. 4Ab); by contrast, only the relative abundance of Alloprevotella was larger in the females in group CK (Fig. 4Bb). Compared with CK, uncultured species in the Ruminococcus UCG-014, Phascolarctobacterium, and Tyzzerella 3 genera exhibited higher values for the T population (Fig. 4Ac,Bc), while only uncultured Alloprevotella sp. was found to have a larger abundance in CKF (Fig. 4Bc). A cladogram representative of the structure of the gut microbiota and their predominant bacteria is presented in Fig. 5. The specific bacteria taxa were concentrated in the Clostridia and Bacteroida classes. Alterations of Lachnospiraceae, Slackia, and Prevotellaceae were consistent with the abovementioned results. However, new findings were also discovered. Greater abundances of Bacterioides, Sarcina, and Roseburia were found in T, which also had a lower abundance of Prevotella 9 (Fig. 5A). Similar changes at the class level also occurred, as revealed by the comparison between TM and CKM (Fig. 5B). Intriguingly, Proteus mirabilis, the only species in genus Proteus, was decreased in TM. In a comparison of the two female groups, only one change happened in the Lachnospiraceae family (Fig. 5C). Taken together, the results indicate that presence of multiple metals significantly alters the compositional profile of the gut microbiome.
Linear discriminant analysis effect size (LEfSe) cladogram that shows the most differentially abundant taxons between these groups. Taxonomic cladogram obtained from LEfSe analysis of pyrosequencing sequences. Blue indicates taxa enriched in CK, and orange indicates taxa enriched in exposures. Only taxa meeting a linear discriminant analysis (LDA) significant threshold > 4 are shown. (A) Taxons significantly different between groups CK and T. (B) Taxons significantly different between men from groups CK and T. (C) Taxons significantly different between women from groups CK and T.
Discussion
A healthy microbiota in the intestine can protect against bacterial infection and contribute to intestinal functions including digestion and absorption. Moreover, it is also an effective barrier to protect against the threat of toxic metals32. The present study provides novel evidence that long-term exposure to multiple metals including As, Cd, Cu, Pb, and Zn is associated with alterations in the gut microbiome of people living in metal-polluted areas, especially for men.
Metals such as As, Cd, and Cu, as well as excessive amounts of Zn, have been proven to reduce community diversity in previous studies23,33,34. Although no effect on α-diversity has been found, Pb was shown to change complexity (β-diversity) in an animal study35. However, the results of single exposure in the present study were different from those of previous studies. Multiple metals significantly increased α-diversity in the gut microbiome (Shannon-Wiener and Simpson indices), indicating that the abundance and evenness of bacteria in people who live in the polluted area were higher. Perturbation in the community was also found via the analysis of β-diversity (PCoA and PCA).
Various effects of metals at the phylum level have been found in previous studies; for example, one epidemiological and animal study suggested that early-life lead exposure can significantly affect Bacteroides and Firmicutes populations, which may be responsible for the expression of the genes involved in lipid metabolites, isooflavone metabolites, and bile acid metabolites36. It has also been found by individual studies on As, Cr, and Cd that toxic metal exposure can produce alterations of Proteobacteria and Verrcuomicrobia21. In the results of the present study, the proportions of the three phyla Bacteroides, Firmicutes, and Proteobacteria showed similar changes in people who had suffered from long-term exposure to the metals, albeit without any statistical significance.
There were nine genera that responded obviously to metal exposure, and many of them have proven associations with the functions of the host. Numbers of Lachnospiraceae and Erysipelotrichaceae, which are linked to intestinal inflammation, were all increased, which is consistent with a previous study on the effect of heavy metals on commensal communities21. Another bacterium, Eubacterium eligens, which presents a negative correlation with intestinal inflammation, was also enriched in feces from the metal-exposed population. In some previous studies, beneficial bacteria, such as Roseburia (associated with effects of colonic motility, immunity maintenance, and anti-inflammatory properties) and Ruminococcaceae (mainly associated with mucosa), have presented lower amounts in feces from a population living in a metal-contaminated area37. However, in the results of the present study, these populations were all markedly enriched for subjects from the metal-contaminated area. Many studies have reported either reduced abundance of Prevotella in relapsing remitting multiple sclerosis (RRMS) patients, or an increased abundance of Prevotella after treatment with disease-modifying therapies38,39,40,41,42. These data also suggest that Prevotella might have an important anti-inflammatory role in RRMS patients. Additionally, Ashutosh et al.43 reported that Prevotella can suppresses disease through the modulation of systemic immune responses. Interestingly, the results of the present study show that an opportunistic pathogen (Prevotella 9) was decreased in people burdened with higher metal loads. These people also have a risk of inflammation and other diseases, such as RRMS. This evidence suggests that the homeostasis of gut flora in the residents in the mining and smelting area was broken by the complex toxicity of these metals.
Some differences were also found in the group comparisons based on gender. The same changes of three bacteria (Bacteroides, Roseburia, and Prevotella 9) were found in the male groups. Proteus, a gram-negative pathogenic bacterium, was significantly depleted in men who live in the polluted area. All these findings indicate that these metals synergistically operate on the male microbiome. For women, among all the bacteria, only Lachnospiraceae showed significant changes. This exceptional gender-based disparity is noteworthy. This is the first time that greater shifts in male microbiome have been demonstrated, and there is unconformity with previous views based on the physical characteristics of women44. The lower metal burdens of women living in the contaminated site were proven in the authors’ previous study31; this could be the most important reason for the variation of the bacteria in the intestinal tract. The Different living habits (smoking and drinking) and work categories (farming and smelting) between men and females may contribute to this difference. In summation, the intestinal flora in females may retain a large proportion of resistance against long-term metal exposure, while the confusion of the males is already apparent.
The results reveal for the first time that long-term multiple As, Cd, Cu, Pb, and Zn metal exposure can perturb the normal gut community in local inhabitants of contaminated areas, especially in male subjects. Given the importance of gut microbiota to human health, further research is needed to confirm the contribution of the microbiome to metal-related diseases.
Materials and Methods
Study area
Two areas from Hezhang County were selected due to their similar geographical factors, life habits, and daily diet. The study area was a Pb-Zn mining and smelting location, called Maomaochang Village (104°10′E to 105°35′E, 26°46′N to 27°28′E), and the surrounding area, Yemachuan Village, was selected as the control area. The distance between Maomaochang Village and Yemachuan is 120 kilometers, and the drinking water of residents come from different sources. Table 3 indicates that the mean values of five metals (As, Cd, Cu, Pb, and Zn) in the plowed soil of the study area were higher than those of the control area, while the Cr and Ni values were similar between the two areas.
Sample collection
In 2016, 85 volunteers took part in a personal information questionnaire. Bowel disease patients, people who had recently used antibiotics or other medicines, and other sick persons were eliminated from among these subjects. Using stool boxes, fecal samples were collected from the remaining subjects in the group. The samples were stored in a 4 °C icebox and then sent back to the lab, after which they were stored at −80 °C in a freezer45.
DNA extraction and 16 S rRNA sequencing
DNA was extracted using the CTAB method with QIAamp genomic DNA and RNA kits (Qiagen, Crawley, West Sussex, UK), and its concentration and purity was monitored on 1% agarose gels. Upon concentration determination, the DNA was diluted to 1 ng/μl and immediately stored at −20 °C. The V3 and V4 regions of 16 S rRNA genes were amplified; the forward primer was CCTAYGGGRBGCASCAG, and the reverse primer was GGACTACNNGGGTATCTAAT. All PCR reactions were carried out with Phusion@ High-Fidelity PCR Master Mix (New England Biolabs). The same volume of 1× loading buffer (containing SYBR green) was mixed with the PCR products, and electrophoresis on 2% agarose gel was carried out for detection. Samples with principal bright bands between 400 and 450 bp were selected for further experiments. Equimolar ratios of the total samples were purified using a Qiagen Gel Extraction Kit (Qiagen, Germany). Sequencing libraries were generated using a TruSeq® DNA PCR-Free Sample Preparation Kit (Illumina, USA) following the manufacturer’s recommendations, and index codes were added; the quality of the library was assessed on Qubit@2.0 Fluorometer (Thermo Scientific) and Agilent Bioanalyzer 2100 systems. Finally, the library was sequenced on an IlluminaHiSeq2500 platform, and 250 bp paired-end reads were generated.
Bioinformatic analysis
Clean data was attained by jointing raw tags, removing the barcode and primer, and filtering. All effective tags were clustered into operational taxonomical units (OTUs) at the 97% identification level by Uparse v7.0.1001. The Mothur method and SILVA SSU rRNA database were used for species annotation to obtain information on the types of classification levels (Woo et al., 2008). Alpha diversity (α-diversity) was obtained to describe the sample complexity; R software (Version 2.15.3) was used to draw the curves of rarefaction and calculate the indices of richness and diversity (ACE, Chao 1, Shannon-Wiener, and Simpson). Excel and SPSS 25.0 were used to evaluate significant differences in these indices. Beta diversity (β-diversity) was used to evaluate the differences between samples in species complexity via unweighted UniFrac distances, which were calculated by QIIME software (Version 1.7.0). Principal coordinate analysis (PCoA) and principal components analysis (PCA) were used to elucidate the distances among the samples. The linear discriminant analysis effect size (LEfSe) and Kruskal-Wallis tests were utilized to analyze the significantly different bacteria.
Ethical statement
We declare that all experimental protocols were approved by the Human Ethics Committee of China Agricultural University. We also confirm that all methods were performed in accordance with the relevant guidelines and regulations.
Informed consent
We declare that all the participants were adults. All the subjects knew that drinking water, diet information, urine, and hair samples were collected by our team. We also informed them of the scientific research purposes of all the experimental samples (Appendix S4).
References
de Almada, C. N., De Almada, C. N., Martinez, R. C. R. & de Souza Sant’Ana Characterization of the intestinal microbiota and its interaction with probiotics and health impacts. Appl. Microbiol. Biotechnol. 99, 4175–4199 (2015).
Prakash, S., Rodes, L., Coussa-Charley, M. & Tomaro-Duchesneau Gut microbiota: next frontier in understanding human health and development of biotherapeutics. Biologics: targets & therapy 5, 71 (2011).
Aziz, Q., Doré, J., Emmanuel, A., Guarner, F. & Quigley, E. M. M. Gut microbiota and gastrointestinal health: current concepts and future directions. Neurogastroenterol. Motil. 25, 4–15 (2013).
Adamovsky, O. et al. The gut microbiome and aquatic toxicology: An emerging concept for environmental health. Environmental toxicology and chemistry 37, 2758–2775 (2018).
Harris, V. C. et al. Effect of antibiotic-mediated microbiome modulation on rotavirus vaccine immunogenicity: a human, randomized-control proof-of-concept trial. Cell Host Microbe 24, 197–207. e194 (2018).
Xu, W.-T., Nie, Y.-Z., Yang, Z. & Lu, N. The crosstalk between gut microbiota and obesity and related metabolic disorders. Future Microbiol. 11, 825–836 (2016).
Yang, T. et al. Gut dysbiosis is linked to hypertension. Hypertension 65, 1331–1340 (2015).
Usami, M., Miyoshi, M. & Yamashita. Gut microbiota and host metabolism in liver cirrhosis. World J. Gastroenterol. 21, 11597 (2015).
Kashtanova, D. A. et al. Association between the gut microbiota and diet: Fetal life, early childhood, and further life. Nutrition 32, 620–627 (2016).
De Filippo, C. et al. Impact of diet in shaping gut microbiota revealed by a comparative study in children from Europe and rural Africa. Proc. Natl. Acad. Sci. USA 107, 14691–14696 (2010).
Healey, G. R., Murphy, R., Brough, L. & Butts, C. A. Interindividual variability in gut microbiota and host response to dietary interventions. Nutr. Rev. 75, 1059–1080 (2017).
Kumar, M., Babaei, P., Ji, B. & Nielsen Human gut microbiota and healthy aging: recent developments and future prospective. Nutrition and Healthy aging 4, 3–16 (2016).
Clarke, S. F. et al. Exercise and associated dietary extremes impact on gut microbial diversity. Gut 63, 1913–1920 (2014).
Takagi, T. et al. Differences in gut microbiota associated with age, sex, and stool consistency in healthy Japanese subjects. J. Gastroenterol. 54, 53–63 (2019).
Watanabe, K., Kodama, Y. & Harayama. Design and evaluation of PCR primers to amplify bacterial 16S ribosomal DNA fragments used for community fingerprinting. J. Microbiol. Methods 44, 253–262 (2001).
Chakravorty, S., Helb, D., Burday, M., Connell, N. & Alland. A detailed analysis of 16S ribosomal RNA gene segments for the diagnosis of pathogenic bacteria. J. Microbiol. Methods 69, 330–339 (2007).
Dethlefsen, L., Huse, S. & Sogin, M. L. The pervasive effects of an antibiotic on the human gut microbiota, as revealed by deep 16S rRNA sequencing. PLoS biology 6, e280 (2008).
Claesson, M. J. et al. Composition, variability, and temporal stability of the intestinal microbiota of the elderly. Proc. Natl. Acad. Sci. USA 108, 4586–4591 (2011).
Deng, Y. et al. Patients with Acne Vulgaris Have a Distinct Gut Microbiota in Comparison with Healthy Controls. Acta Derm.-Venereol. 98, 783–790 (2018).
Lu, Y. et al. Impacts of soil and water pollution on food safety and health risks in China. Environment international 77, 5–15 (2015).
Richardson, J. B. et al. Exposure to toxic metals triggers unique responses from the rat gut microbiota. Sci. Rep. 8 (2018).
Choiniere, J. & Wang, L. Exposure to inorganic arsenic can lead to gut microbe perturbations and hepatocellular carcinoma. Acta Pharm. Sin. B 6, 426–429 (2016).
Yehao, L., Yuhui, L., Kaiyong, L. & Jie, S. Exposing to cadmium stress cause profound toxic effect on microbiota of the mice intestinal tract. PLoS One 9, e85323 (2014).
Jin, Y., Wu, S., Zeng, Z. & Fu, Z. Effects of environmental pollutants on gut microbiota. Environ. Pollut. 222, 1–9 (2017).
Qin, G. et al. Effects of dietary iron modulation on gut microbial composition and function in monogastrics: a review. Environ. Pollut. 24, 1–1 (2017).
Spenser, R. et al. Chronic Zinc Deficiency Alters Chick Gut Microbiota Composition and Function. Nutrients 7, 9768–9784 (2015).
Zackular, J. P. et al. Dietary zinc alters the microbiota and decreases resistance to Clostridium difficile infection. Nat. Med. 22, 1330–1334 (2016).
Meryem, B. et al. Characterization, distribution, and risk assessment of heavy metals in agricultural soil and products around mining and smelting areas of Hezhang, China. Environ. Monit. Assess. 187, 1–21 (2015).
Mei, L. I., Sun, J. & Liya, Q. A Survey of Cadmium Pollution in Hezhang County and Its Effects on Human Health. Journal of Guiyang Medical College 32, 464–467 (2007).
Peng, D. H., Pan, W. U., Cao, Z. X., Yang, S. Z. & Xie, H. H. J. Jo. A.-E. S. Spatial and Temporal Variation Characteristics of Heavy Metal Pollution of Water-sediments in Stream of Zinc Smelting Area,Hezhang of Guizhou,China. Environment Science 30, 979–985 (2011).
Shao, M. et al. The health hazards of potentially toxic metals in the daily diets of adults and children from a mining and smelting region (Hezhang County) in southwestern China. Environmental monitoring and assessment 190, 432 (2018).
Franziska, F. & Baumler, A. J. The impact of intestinal inflammation on the nutritional environment of the gut microbiota. Immunol. Lett. 162, 48–53 (2014).
Chi, L. et al. The Effects of an Environmentally Relevant Level of Arsenic on the Gut Microbiome and its Functional Metagenome. Toxicological Sciences 160, 193–204 (2017).
Ba, Q. et al. Sex-Dependent Effects of Cadmium Exposure in Early Life on Gut Microbiota and Fat Accumulation in Mice. Environ. Health Perspect. 125, 437–446 (2017).
Xia, J. et al. Chronic exposure to low concentrations of lead induces metabolic disorder and dysbiosis of the gut microbiota in mice. Sci. Total Environ. 631-632, 439 (2018).
Kun, L. et al. Arsenic exposure perturbs the gut microbiome and its metabolic profile in mice: an integrated metagenomics and metabolomics analysis. Environ. Health Perspect. 122, 284 (2014).
Tamanai-Shacoori, Z. et al. Roseburia spp.: a marker of health? Future Microbiol. 12, 157–170 (2017).
Chen, J. et al. Multiple sclerosis patients have a distinct gut microbiota compared to healthy controls. Sci. Rep. 6, 28484 (2016).
Miyake, S. et al. Dysbiosis in the gut microbiota of patients with multiple sclerosis, with a striking depletion of species belonging to clostridia XIVa and IV clusters. PloS one 10, e0137429 (2015).
Jangi, S. et al. Alterations of the human gut microbiome in multiple sclerosis. Nature communications 7, 12015 (2016).
Castillo Alvarez, F. et al. In Multiple Sclerosis Journal. 190–190 (Sage Publications Ltd 1 Olivers Yard, 55 City Road, London Ec1y 1Sp, England).
Shahi, S. K., Freedman, S. N. & Mangalam, A. K. Gut microbiome in multiple sclerosis: the players involved and the roles they play. Gut microbes 8, 607–615 (2017).
Mangalam, A. et al. Human gut-derived commensal bacteria suppress CNS inflammatory and demyelinating disease. Cell reports 20, 1269–1277 (2017).
Marie, I. et al. Systemic sclerosis and exposure to heavy metals: A case control study of 100 patients and 300 controls. Autoimmun. Rev. 16, 223–230 (2017).
Jonna, J. et al. Effects of bowel cleansing on the intestinal microbiota. Gut. 64, gutjnl-2014–307240 (2015).
Acknowledgements
The authors are extremely grateful to Dr. Ji for his constructive comments. We would also like to acknowledge the help extended to us by the staff of the Institute of Geochemistry, Chinese Academy of Sciences in sample collection.
Author information
Authors and Affiliations
Contributions
Z.Y. conceived and designed the experiment. S.M. performed high-throughput sequencing and analysis, and wrote the manuscript. Both authors discussed the results and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Shao, M., Zhu, Y. Long-term metal exposure changes gut microbiota of residents surrounding a mining and smelting area. Sci Rep 10, 4453 (2020). https://doi.org/10.1038/s41598-020-61143-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-61143-7
This article is cited by
-
Influence of grape consumption on the human microbiome
Scientific Reports (2023)
-
Individual and combined effects of fluoride and arsenic on gut bacteria: a recent update
The Nucleus (2023)
-
Microbiome differences between wild and aquarium whitespotted eagle rays (Aetobatus narinari)
Animal Microbiome (2022)
-
Impact of Heavy Metal Toxicity on the Gut Microbiota and Its Relationship with Metabolites and Future Probiotics Strategy: a Review
Biological Trace Element Research (2022)
-
Urolithin A attenuates arsenic-induced gut barrier dysfunction
Archives of Toxicology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.