Abstract
Wastewater treatment plants (WWTPs) are necessary to protect ecosystems quality and human health. Their function relies on the degradation of organic matter and nutrients from a water influent, prior to the effluent release into the environment. In this work we studied the bacterial community dynamics of a municipal WWTP with a membrane bioreactor through 16S rRNA gene sequencing. The main phyla identified in the wastewater were Proteobacteria, Bacteroidetes, Chloroflexi, Planctomycetes and Actinobacteria. The WWTP is located in Spain and, like other studied WWTP in temperate climate zones, the temperature played a major role in community assembly. Seasonal community succession is observed along the two years sampling period, in addition to a continual annual drift in the microbial populations. The core community of the WWTP bioreactor was also studied, where a small fraction of sequence variants constituted a large fraction of the total abundance. This core microbiome stability along the sampling period and the likewise dissimilarity patterns along the temperature gradient makes this feature a good candidate for a new process control in WWTPs.
Similar content being viewed by others
Introduction
Contaminant removal from industrial and urban wastewater is a capital issue for the protection of the natural environment and human health. Most Wastewater Treatment Plants (WWTPs) rely on conventional biological treatment systems, such as activated sludge processes. In combination with different redox conditions (anaerobic, anoxic and aerobic), these biological processes assure the removal of organic matter and nutrients1. As the most common biological wastewater treatment application, activated sludges are complex microbial ecosystems composed of Bacteria, Archaea, Eukarya and viruses2,3. Recently, Membrane Bioreactor (MBR) technology in wastewater treatment has become a common practice globally. This process combines efficient biological degradation by activated sludge with a direct solid-liquid membrane separation, offering several advantages over other systems such as a superior effluent quality, high biodegradation capacity and low sludge production4,5. Developed activated sludge heavily depends on its microbial populations for the removal of nutrients and organic pollutants and hence, the depuration performance of WWTPs. Understanding the composition, structure and functioning of these microbial communities is essential for constructing improved wastewater treatment plants6.
Activated sludge bacterial communities have been widely studied either by culture-dependent methods or by molecular approaches revealing, on one side a high bacterial diversity, on the other the fact that a small fraction of taxa accounted for the majority of total abundance7,8. In previous works, Proteobacteria was the most abundant phylum in WWTPs (accounting for 30–80% of the total abundance), followed by Bacteroidetes, Firmicutes and Actinobacteria9,10,11,12,13. These communities include taxa involved in different metabolic pathways (nitrogen fixation, nitrification, denitrification, sulphur oxidation, etc.); physiological groups like anaerobic, aerobic, phototrophic, heterotrophic, etc.; and inter-species relationships systems (i.e. quorum sensing)14. Furthermore, microbial populations inhabiting WWTP bioreactors can differ among different stations and over time, and their biodiversity is thought to play an important role in enabling and facilitating particular ecosystem functions, such as nutrients removal. Predicting the behaviour of particular populations and communities under different situations and how these are linked to the performance of a particular ecosystem process, will improve the efficiency of critical processes in WWTP communities15.
WWTP performance should guarantee a certain effluent quality which will vary according to the posterior uses of water. Different techniques have been developed to measure activated sludge quality, such as Sludge Biotic Index (based on the presence and abundance of certain protists) or the Sludge Index (considering macroscopic and microscopic activated sludge features)16,17,18. A complementary tool to control microbial community and activated sludge stability based on the bacterial populations is necessary to assess WWTP performance and prevent activated sludge disruptions. High-throughput sequencing (HTS) of conserved regions in microbial genomes (i.e. bacterial 16S rRNA), are nowadays considered the most reliable and cost-effective method to study the microbial composition and ecological dynamics of WWTPs2,19.
The seasonality of natural ecosystems such as oceans, freshwater or soils has been widely studied with HTS, but little attention has been given to anthropic ecosystems. Indeed, in the activated sludge microbial community, seasonal dynamics may affect the performance and stability of organic material and nutrient removal (nitrogen and phosphorus) in WWTPs8,20. This aspect should be studied in more detail with the aim of using tag-sequencing as routine technology as a means of learning about microbial ecology in WWTPs. Furthermore, it is necessary to understand the dynamics of microbial communities and their seasonal variations for predicting undesirable changes in the functional diversity of activated sludges, for a better control over operational parameters of WWTPs.
In this work, we have studied monthly the microbial community dynamics of the three reactors of a full-scale municipal WWTP located in North-East of Spain over a period of 2 years. We have defined the core microbiome of this WWTP, its inter-reactor variability and its evolution along time. The results showed a seasonal community variation observed in the full bacterial community that does not exist in the core microbiome. The core microbiome preserved a narrower range of stability while displaying the annual drift trend, making it a good candidate to be considered as a new WWTP process control indicator.
Results
Bacterial community composition
A total of 66 mixed liquor samples were analysed from a full scale MBR-WWTP. Samples were taken monthly along two years in the three compartments of the reactor, allowing the evaluation of annual community variability. A total of 3,454,577 good quality 16S rRNA sequences were obtained, resulting in 6,245 Amplicon Sequence Variants (ASVs). ASVs methods allow the distinction of sequences differing by as little as one nucleotide21.
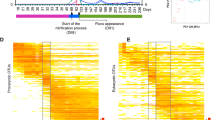
Taxonomic assignation against SILVA (release 132) allowed the identification of 39 phyla. We considered as main phyla those whose abundance was higher than 1% on at least one sample (Fig. 1). The average relative abundances, of sequence reads for the main phyla in all stages were as follows: Proteobacteria (considering Alpha-, Beta-, Delta- and Gammaproteobacteria) 30.18% ± 3.84%, Bacteroidetes 23.12% ± 5.12%, Chloroflexi 17.15% ± 3.55%, Planctomycetes 7.53% ± 2.10%, Actinobacteria 7.20% ± 3.25%, Acidobacteria 5.63% ± 1.52%, Patescibacteria 1.74% ± 1.17%, Firmicutes 1.66% ± 1.11%, Verrucomicrobia 1.25% ± 0.44%, Nitrospirae 0.51% ± 0.54%. All those phyla, except Nitrospirae, that is only present in spring and summer samples, tended to be persistent over the 2-year sampling period. The phylum Proteobacteria is more abundant in winter and spring samples, and is mainly represented by the classes Alpha (8.21% ± 2.49%), Beta- (9.71% ± 3.12%), Delta- (2.55% ± 1.06%) and Gammaproteobacteria (9.52% ± 2.21%). However, other phyla like Bacteroidetes, Chloroflexi or Planctomycetes were less abundant during these months (p < 0.05). This trend shows a seasonal distribution of key phyla in mixed liquors. Average relative abundance of Archaea was 0.25% ± 0.20%, being the phylum Euryarchaeota the most abundant (0.21% ± 0.13%), thus, those populations were not considered in subsequent analysis.
Mean relative abundance of dominant bacterial phyla with a frequency higher than 1% on at least one sample. “Other”, includes phyla of frequency <1%; and “unidentified”, taxonomically unassigned taxa. Sampling months are indicated, starting April 2017 and ending March 2019. There was no sampling in August. Phylum Proteobacteria is shown subdivided in classes Alpha-, Beta-, Delta- and Gammaproteobacteria.
Diversity, seasonal dynamics and functional predictions of the wastewater microbiome
To understand the annual variations found in the WWTP, the microbial community diversity (alpha diversity) was calculated, using the Simpson diversity index (D’). No differences were found in the communities inhabiting each compartment of the bioreactor (tested by analysis of similarity, ANOSIM, p = 0.975), so further statistical analysis was performed considering the sampling time. September, October and February were the months with lower diversity, while April and May were the months with higher diversity (Fig. 2a). Overall, autumn appeared to be the season with the lower alpha diversity values, popping a rebound during spring months.
Microbial diversity analysis. (a) Simpson diversity index as a measurement of alpha diversity. An ANOVA test and a LSD (Least Square Difference) test were conducted (a–c indicate significance groups). (b) Redundancy analysis (RDA). Dots and triangles correspond with each sample on the first or second sampling year, respectively. Black points stand for the centroid of the first and second sampling year. Colour is indicative of the temperature gradient, and lines show the physical-chemical factors constrained into the ordination. (c) pairs between predicted functional groups (Supplementary Table S2), and physical-chemical factors. Dark blue indicates stronger positive correlations and dark red stronger negative correlations. Asterisks denote the significance levels (***p < 0.001, **p < 0.01 and *p < 0.05). P values were adjusted for multiple testing using the Bonferroni correction.
The multidimensional ordination (NMDS) of the sequence data based on Bray-Curtis dissimilarities was used to compare the ASV profiles of communities (Supplementary Fig. S1). This analysis revealed seasonal clustering of communities, where samples were sorted monthly, from April 2017 (first sample) to March 2019 (last sample), showing an annual cycle in both years (Supplementary Fig. S1). Despite that the annual cycle is repeated in both sampling years, permutational multivariate analysis of variance (PERMANOVA) test showed significant differences between them in terms of community structure (p = 0.001). PERMANOVA tests were also used to study the effect of physical-chemical parameters on the bacterial community, identifying the temperature of the bioreactor as the principal factor impacting microbial populations (R2 = 0.162, p = 0.001). The significant parameters (p < 0.01), along with temperature, explained a 54.95% of the variation observed between samples. Figure 2b shows a redundancy analysis (RDA) where some constrained features (influent biochemical oxygen demand-I.BOD) contributes to the first component in the same direction than temperature, as the most relevant variable, and some others (pH, effluent ammonia concentration-E.NH4-N, and hydraulic retention time-HRT) contributes in an opposite direction in shaping bacterial communities composition.
Through a functional prediction based on 16S rRNA sequences, it is possible to estimate functional traits of the bacterial population in the wastewater. We analysed correlations of 57 environmental functions from 5 pathways (involved in nitrogen and phosphorous metabolism, biodegradation and quorum sensing) and the wastewater physical-chemical factors (Fig. 2c). The temperature was significantly correlated with the presence of quorum sensing related genes (p < 0.001) and inversely correlated with denitrification (p < 0.01). Nitrification was significantly correlated with chemical oxygen demand (COD) in the effluent (p < 0.01) and its reduction (inversely, p < 0.001).
Wastewater core bacterial community
The core bacterial community was defined based on the high occurrence frequency of the ASVs. About 0.88% of total ASVs (55 ASVs) were present in 100% of the activated sludge samples and constituted a core that accounted for 36.5 ± 5.2% of the sequence reads. Most of the core community members belonged to Alpha-, Beta- and Gammaproteobacteria classes; and Bacteroidetes and Chloroflexi phyla (Fig. 3a). Spearman’s rank correlation coefficient revealed that changes of activated sludge community composition, at the family level, were significantly correlated with physical-chemical parameters. Out of the 30 families that form the core microbiome, 12 were significantly correlated (adjusted p < 0.05) to at least one physical-chemical parameter (Fig. 3b). Ammonia removal is correlated with Pirellulaceae and Fimbriimonadaceae families (p < 0.001), while temperature of the bioreactor is inversely correlated with Burkholderiaceae, Hyphomonadaceae, Ruminococcaceae, Chitinophagaceae and Rhodanobacteraceae families (p < 0.05).
Composition, functional correlations and multidimensional ordination based on core ASVs in the activated sludge samples. (a) Taxonomic composition of core ASVs at phylum level and class level. Proteobacteria group is divided in inferior taxonomic levels. (b) Spearman’s rank correlation coefficient between core ASVs, at family level assignation, and physical-chemical factors. Only core families and physical-chemical parameters with at least one significative correlation are shown. (c) Beta diversity of core community based on a non-metric multidimensional scaling analysis (NMDS, stress = 0.152).
The beta diversity of the core community showed a clear clustering and evolution based on the temperature of the sample (PERMANOVA test, R2 = 0.176, p = 0.001) (Fig. 3c). As occurred with the beta-diversity of the full community, PERMANOVA test showed a significant difference between the core microbiome composition the two years sampled (R2 = 0.053, p = 0.001).
Indeed, when calculating the distance of each sample to their sampling year centroid (Fig. 4) we observed that when the average temperatures rose, these distances also increased. This comparison indicated that the higher the temperature the higher the changes in the core bacterial community. Both years presented a similar pattern of temperature/community dispersion, making it possible to define a new system stability measure (Supplementary Fig. S2).
Mean distance to centroid of bacterial population dissimilarities of samples taken on each sampled month, accounting for both sampling years. (a) Mean distances are represented along the sampling months. (b) Correlation between mean distances and temperature (r = 0.26, p = 0.04). Each sampling year were considered independent, and both centroids were calculated. The colour gradient shows the average monthly temperature (°C) on both sampling years. For a desegregate visualization of the two year sampling monitoring see Supplementary Fig. S3.
Discussion
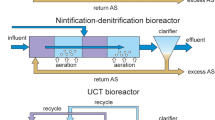
In this work, we studied the bacterial community dynamics of a municipal waste water treatment plant with a membrane bioreactor (MBR-WWTP). This bioreactor is functionally divided into three sections, attending to their oxygenation conditions (anoxic, oxic-anoxic, anoxic); besides it has a recirculation system connecting the third stage with the first. Through 16S rRNA gene sequencing we identified Proteobacteria, Bacteroidetes, Chloroflexi, Planctomycetes and Actinobacteria as the main phyla in the bacterial community studied, making 85.17% ± 2.43% of the total sequence reads. The most common phyla found in WWTPs around the globe are Proteobacteria and Bacteroidetes. The two Proteobacteria most common families in the WWTP studied were Burkholderiaceae (Betaproteobacteria), that has members known to degrade PCBs22; and Rhodobacteraceae (Alphaproteobacteria), involved in sulfur and carbon cycles23. Within Bacteroidetes phyla, the most abundant families were Chitinophagaceae, aerobic heterotrophs degraders of organic matter24; and Saprospiraceae which includes many bacteria associated with protein hydrolysis and epiphytic bacteria attached on some filamentous bacteria (Supplementary Fig. S3)25. In addition, the core bacterial community was determined based on the occurrence of sequence variants (ASVs) in the samples. Despite consisting in a relatively low number of ASVs (55 or 0.88% of the total), these accounted for a 36.5% of the total abundance, implying a hyper-dominance pattern similar to that found in a global WWTP core bacterial community study8.
Recent studies suggest that activated sludge microbial communities are shaped by deterministic (environmental and interspecies relationships) and neutral or stochastic (random events like colonization/extinction causing microbial dispersal) factors26,27. In natural aquatic ecosystems, a seasonal succession of the community is normally found, however, it is not as common to occur in artificial ecosystems28. Interestingly, we have found that in activated sludge systems the seasonal dynamics of microbial communities greatly affect the performance and stability of pollutant removal, and the main deterministic factor for this community dynamic in temperate climatic zones is the seasonal temperature fluctuation15,20. In the WWTP studied, samples were significantly different from each other, although, the samples of the same month of different years were more similar between them than with the other (Supplementary Fig. S1). This suggests a seasonal community succession but continual annual drift to the activated sludge at the studied facility, since both sampling years are significantly different (p = 0.001). The functional estimation performed in this work suggest a notable impact of temperature in certain bacterial-related functions such as a decrease in the presence of genes related with denitrification or a notable increase in those genes related with quorum sensing (Fig. 2c). This is of particular interest as quorum sensing regulates different metabolic mechanisms and coordinated behaviours at a community level; some of them of particular interest in WWTPs such as bacterial granulation and the maintenance of granular structures, or the undesired biofilm formation in MBR membranes29,30. This aspect should be studied in more detail for a better understanding of the impact of temperature in quorum sensing-mediated metabolisms in WWTPs.
As we have seen, the deterministic factors present in the studied WWTP greatly affects the bacterial community structure of its activated sludge (54.95% of the difference is explained by the physical-chemical parameters studied), with the temperature of the bioreactor as the main factor. Giving special attention to the families that compose the core microbiome, we observed that the ammonia removal is significantly correlated with the Fimbriimonadaceae and Pirellulaceae families. Pirellulaceae are ammonia-oxidizing bacteria31, while Fimbriimonadaceae belong to the Armatinomonadetes phylum, detected in ANAMMOX (ANaerobic AMMonium OXidation) consortia32. The two families aforementioned are positively correlated (p < 0.001, Spearman’s rank correlation coefficient = 0.495), implying that the Fimbriimonadaceae family either contains ammonia oxidizing taxons or has positive interactions with ammonia oxidizing bacteria, favouring the ammonia oxidizing processes. As we can see, bacteria present in the core microbiome are essential in the pollutant removal processes carried out in the activated sludge. Further research is necessary to study the biological interactions between both clades.
Nutrient and organic matter removal processes carried out in the activated sludge are based on the activity of the studied microorganisms, thus maintaining optimal conditions for their growth and development is necessary for a better process control and management18. The dynamics and organization of microbial communities, along with biotic components of the activated sludge, determine its quality and the efficiency of the depuration process16. Based on this premise, the operation of the reactor is currently controlled through various indicator organisms and various indices to assess the quality of the active sludge. These types of indices offer invaluable information about the effluent characteristics and the adequacy of the operational conditions33. Previous studies suggest that highly diverse ecosystems are more stable, due to the presence of species able to adapt to a changing ecosystem. The maintenance of stable activated sludge ecosystem requires understanding the functional stability of the bacterial communities34. These species representing the core microbiome are essential for wastewater treatment processes35. In addition, MBR systems, from this work perspective, have additional problems such as the biofouling onto filtration membranes, causing failures in the depuration performance. Among other reasons, biofouling is caused by quorum sensing-controlled biofilm formation on filtration membranes, so understanding keystone species in biofilm formation also represents an interesting topic for future research36. Thus, estimating the stability of the mentioned population would indicate a correct system function, so we propose monitoring this stability along time as a new control parameter on WWTP functioning. To quantify this stability, we calculated the difference between each sample and the centroid of their sampling year. Thus, we observed that in the warmer months populations tend to be more dissimilar, returning to baseline differences in the winter months. It is the effect we observed of the temperature on the population. Further analyses are needed to implement this monitoring measure, such as in the case of a problem detection in the plant checking how it would affect the dissimilarities of the populations. Monitoring with longer time series would help to verify the continued annual drift.
Material and Methods
Site description, samples collection and basic water characterization
Samples were collected over a period of two years from a full-scale WWTP with a membrane bioreactor system (MBR) located in Barcelona (Spain) and described in previous studies37. The 66 samples were taken monthly and consisted of collected mixed liquor from the three functional stages in which the bioreactor was divided (Supplementary Fig. S4). Physical-chemical parameters were measured in the influent (I.-) and the effluent (E.-) of the WWTP, according to standard methods: biochemical oxygen demand, BOD (UNE-EN-1899); chemical oxygen demand, COD (ISO-6060); ammonia, NH4-N (ISO-7150); total nitrogen, TN (ISO-11905); total phosphorous, TP (ISO-6878) and suspended solids, SS (UNE-EN-872). The removal rate (R.-) was calculated accordingly. Mixed liquor suspended solids (MLSS) were measured according to UNE-EN-872 and hydraulic retention time, HRT was also calculated. Detailed information concerning plant physical-chemical parameters is summarized in Supplementary Table S1.
DNA extraction and 16S rRNA sequencing
Mixed liquor samples were analysed following a 16S metabarcoding strategy. Samples were stored at −80 °C until DNA extraction was performed using DNA Power Soil extraction kits. The V4 region of the 16S rRNA gene was amplified by PCR using the primers 515F (GTGYCAGCMGCCGCGGTAA) and 806R (GGACTACNVGGGTWTCTAAT). Libraries were prepared following the two-step PCR Illumina® protocol and these were subsequently sequenced on Illumina® MiSeq instrument (Illumina®, San Diego, CA, USA) using 2 × 300 paired-end reads38,39 and then it was analysed by amplifying and sequencing the 16S rRNA V4 gene using custom primers40.
Sequence processing
DADA2 algorithm41 implemented in R pipeline42 was used to perform sequence analysis, such as denoise, filter, align pairs and filter out chimeras. This algorithm implements an error correction model that allows the differentiation of even a single nucleotide21, giving as a final output an amplicon sequence variant (ASV) table. A total of 3,454,577 good quality reads were obtained. The taxonomic assignment was performed using the naïve Bayesian classifier implemented in DADA2 using as reference database Silva (release 132), with a bootstrap cut-off of 80%43.
Microbial diversity and statistical analysis
Microbial diversity and statistical analysis were performed using phyloseq, version 1.26.144 and vegan, version 2.5.545 R packages. Simpson index of diversity46 was calculated, per sample, as an estimation of community alpha diversity. Beta-diversity (differences between samples) was calculated using a Bray-Curtis dissimilarity matrix on Hellinger transformed data47,48 and permutational multivariate analysis of variance (PERMANOVA). Redundancy analysis (RDA) was conducted from the dissimilarity matrices to compress dimensionality into two dimensional plots, constraining physical-chemical information in the plot.
Predictive functional analysis based on 16S rRNA was performed using an adaptation of the Tax4Fun routine49 software package. To obtain the proportion of each community containing each specific function, we filtered a total of 57 KEGGs functional orthologues (Supplementary Table S2) within 5 pathways related to nitrogen and phosphorous metabolisms, biodegradation and quorum sensing50,51,52.
We also investigated the core microbial community, defining core community members as the ASVs occurring in the whole dataset, regardless of their abundance. The Bray-Curtis dissimilarity matrix of the core community table was computed and the distance of the beta diversity of the samples to their sampling year centroid was calculated using betadisper function (vegan package).
Data availability
Raw files are available in the National Center for Biotechnology (NCBI) repository under the project code PRJNA588045.
References
Alvarino, T., Suarez, S., Lema, J. & Omil, F. Understanding the sorption and biotransformation of organic micropollutants in innovative biological wastewater treatment technologies. Sci. Total Environ. 615, 297–306 (2017).
Ju, F. & Zhang, T. Bacterial assembly and temporal dynamics in activated sludge of a full-scale municipal wastewater treatment plant. ISME J. 9, 683–695 (2015).
Xia, Y., Wen, X., Zhang, B. & Yang, Y. Diversity and assembly patterns of activated sludge microbial communities: a review. Biotechnol. Adv. 36, 1038–1047 (2018).
Wan, C. Y. et al. Biodiversity and population dynamics of microorganisms in a full-scale membrane bioreactor for municipal wastewater treatment. Water Res. 45, 1129–1138 (2011).
Zhang, S., Zhou, Z., Li, Y. & Meng, F. Deciphering the core fouling-causing microbiota in a membrane bioreactor: low abundance but important roles. Chemosphere. 195, 108–118 (2018).
Widder, S. et al. Challenges in Microbial Ecology: Building Predictive Understanding of Community Function and Dynamics. ISME J. 10, 2557–2568 (2016).
Calusinska, M. et al. A year of monitoring 20 mesophilic full-scale bioreactors reveals the existence of stable but different core microbiomes in bio-waste and wastewater anaerobic digestion systems. Biotechnol. Biofuels. 11, 196 (2018).
Wu, L. et al. Global diversity and biogeography of bacterial communities in wastewater treatment plants. Nat. Microbiol. 4, 1183 (2019).
Abzazou, T. et al. Characterization of nutrient-removing microbial communities in two full-scale WWTP systems using a new qPCR approach. Sci. Total Environ. 618, 858–865 (2018).
Huo, Y., Bai, Y. & Qu, J. Unravelling riverine microbial communities under wastewater treatment plant effluent discharge in large urban areas. Appl. Microbiol. Biotechnol. 101, 6755–6764 (2017).
Meerbergen, K. et al. Assessing the composition of microbial communities in textile wastewater treatment plants in comparison with municipal wastewater treatment plants. Microbiologyopen. 6, e00413 (2016).
Xia, S. et al. Bacterial community structure in geographically distributed biological wastewater treatment reactors. Environ. Sci. Technol. 44, 1043–1045 (2010).
Zhang, T., Shao, M. F. & Ye, L. 454-Pyrosequencing reveals bacterial diversity of activated sludge from 14 sewage treatment plants. ISME J. 6, 1137–1147 (2012).
Numberger, D. et al. Characterization of bacterial communities in wastewater with enhanced taxonomic resolution by full-length 16S rRNA sequencing. Sci. Rep. 9, 9673 (2019).
Johnson, D. R. et al. Association of biodiversity with the rates of micropollutant biotransformations among full-scale wastewater treatment plant communities. Appl. Environ. Microb. 81, 666–675 (2015).
Arregui, L. et al. Analysis of the usefulness of biological parameters for the control of activated sludge wastewater treatment plants in an interlaboratory study context. J. Environ. Monit. 14, 1444–1452 (2012).
Madoni, P. A sludge biotic index (SBI) for the evaluation of the biological performance of activated sludge based on the microfauna analysis. Water Res. 28, 67–75 (1994).
Pedrazzani, R., Menoni, L., Nembrini, S., Manili, L. & Bertanza, G. Suitability of Sludge Biotic Index (SBI), Sludge Index (SI) and filamentous bacteria analysis for assessing activated sludge process performance: The case of piggery slaughterhouse wastewater. J. Ind. Microbiol. Biotechnol. 43, 953–964 (2016).
Cydzik-Kwiatkowska, A. & Zielińska, M. Bacterial communities in full-scale wastewater treatment systems. World J. Microbiol. Biotechnol. 32, 66 (2016).
Ju, F., Guo, F., Ye, L., Xia, Y. & Zhang, T. Metagenomic analysis on seasonal microbial variations of activated sludge from a full-scale wastewater treatment plant over 4 years. Environ. Microbiol. Rep. 6, 80–89 (2013).
Callahan, B. J., McMurdie, P. J. & Holmes, S. P. Exact sequence variants should replace operational taxonomic units in marker-gene data analysis. ISME J. 11, 2639–2643 (2017).
Parnell, J. J., Denef, V. J., Park, J., Tsoi, T. & Tiedje, J. M. Environmentally relevant parameters affecting PCB degradation: carbon source- and growth phase-mitigated effects of the expression of the biphenyl pathway and associated genes in Burkholderia xenovorans LB400. Biodegradation 21, 147–156 (2010).
Pujalte, M. J., Lucena, T., Ruvira, M. A., Arahal, D. R. & Macián, M. C. The family Rhodobacteraceae, In The Prokaryotes: Alphaproteobacteria and Betaproteobacteria, 4th Edn, eds Rosenberg, E., DeLong, E. F., Lory, S., Stackebrandt, E. & Thompson, F. L. (Berlin: Springer), 439–512 (2014).
Oh, S., Choi, D. & Cha, C. J. Ecological processes underpinning microbial community structure during exposure to subinhibitory level of triclosan. Sci. Rep. 9, 4958 (2019).
Xia, Y., Kong, Y. H., Thomsen, T. R. & Nielsen, P. H. Identification and ecophysiological characterization of epiphytic protein-hydrolyzing Saprospiraceae (Candidatus epiflobacter spp.) in activated sludge. Appl. Environ. Microbiol. 74, 2229–2238 (2008).
Meerburg, F. A. et al. High-rate activated sludge communities have a distinctly different structure compared to low-rate sludge communities, and are less sensitive towards environmental and operational variables. Water Res. 100, 137–145 (2016).
Valentin-Vargas, A., Toro-Labrador, G. & Massol-Deya, A. A. Bacterial community dynamics in full-scale activated sludge bioreactors: operational and ecological factors driving community assembly and performance. Plos One. 7, e42524 (2012).
Griffin, J. S. & Wells, G. F. Regional synchrony in full-scale activated sludge bioreactors due to deterministic microbial community assembly. ISME J. 11, 500–511 (2016).
Ren, T.-T., Yu, H.-Q. & Li, X.-Y. The quorum-sensing effect of aerobic granules on bacterial adhesion, biofilm formation, and sludge granulation. Appl. Microbiol. Biotechnol. 88, 789–797 (2010).
Lade, H., Paul, D. & Kweon, J. H. Isolation and molecular characterization of biofouling bacteria and profiling of quorum sensing signal molecules from membrane bioreactor activated sludge. Int. J. Mol. Sci. 15, 2255–2273 (2014).
Kellogg, C. A., Ross, S. W. & Brooke, S. D. Bacterial community diversity of the deep-sea octocoral Paramuricea placomus. PeerJ. 4, e2529 (2016).
Zhang, Z. & Liu, S. Insight into the overconsumption of ammonium by anammox consortia under anaerobic conditions. J. Appl. Microbiol. 117, 1830–1838 (2014).
Leal, A. L. et al. Implementation of the sludge biotic index in a petrochemical WWTP in Brazil: improving operational control with traditional methods. J. Ind. Microbiol. Biotechnol. 40, 1415–1422 (2013).
Saikaly, P. E. & Oerther, D. B. Diversity of dominant bacterial taxa in activated sludge promotes functional resistance following toxic shock loading. Microb. Ecol. 61, 557–567 (2011).
Zhang, B., Xu, X. Y. & Zhu, L. Structure and function of the microbial consortia of activated sludge in typical municipal wastewater treatment plants in winter. Sci. Rep. 7, 11 (2017).
Miura, Y., Watanabe, Y. & Okabe, S. Membrane biofouling in pilot-scale membrane bioreactors (MBRs) treating municipal wastewater: impact of biofilm formation. Environ.Sci. Technol. 41, 632–638 (2007).
Liébana, R. et al. Membrane bioreactor wastewater treatment plants reveal diverse yeast and protist communities of potential significance in biofouling. Biofouling. 31, 71–82 (2015).
Feld, L., Nielsen, T. K., Hansen, L. H., Aamand, J. & Albers, C. N. Establishment of bacterial herbicide degraders in a rapid sand filter for bioremediation of phenoxypropionate-polluted groundwater. Appl. Environ. Microbiol. 82, 878–887 (2016).
Albers, C. N., Ellegaard-Jensen, L., Hansen, L. H. & Sørensen, S. R. Bioaugmentation of rapid sand filters by microbiome priming with a nitrifying consortium will optimize production of drinking water from groundwater. Water Res. 129, 1–10 (2018).
Becares, A. A. & Fernandez, A. F. Microbiome based identification, monitoring and enhancement of fermentation processes and products. 15/779,531, US Patent App. 15/779,531 (2018).
Callahan, B. J. et al. DADA2: high-resolution sample inference from Illumina amplicon data. Nat. Methods. 13, 581–583 (2016).
Callahan, B. J., Sankaran, K., Fukuyama, J. A., McMurdie, P. J. & Holmes, S. P. Bioconductor Workflow for Microbiome Data Analysis: from raw reads to community analyses. F1000Research. 5, 1492 (2016).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucl. Acids Res. 41, D590–D596 (2013).
McMurdie, P. J. & Holmes, S. phyloseq: An R package for reproducible interactive analysis and graphics of microbiome census data. PLoS One. 8, e61217 (2013).
Oksanen, J. et al. vegan: Community Ecology Package. R package version 2.5–5 https://CRAN.R-project.org/package=vegan (2019).
Simpson, E. H. Measurement of diversity. Nature 163, 668 (1949).
Bray, R. J. & Curtis, J. T. An ordination of the upland forest communities of Southern Wisconsin. Ecol. Monogr. 27, 325–349 (1957).
McMurdie, P. J. & Holmes, S. Waste not, want not: why rarefying microbiome data is inadmissible. PLoS Comput. Biol. 10, 1003531 (2014).
Wemheuer, F. et al. Tax4Fun2: a R-based tool for the rapid prediction of habitat-specific functional profiles and functional redundancy based on 16S rRNA gene marker gene sequences. bioRxiv 490037 (2018).
Johnston, J., Lapara, T. & Behrens, S. Composition and dynamics of the activated sludge microbiome during seasonal nitrification failure. Sci. Rep. 9, 4565 (2019).
Fang, H. et al. Exploring bacterial communities and biodegradation genes in activated sludge from pesticide wastewater treatment plants via metagenomic analysis. Environ. Pollut. 243, 1206–1216 (2018).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Acknowledgements
This research has been funded by the Spanish Ministry of Economy, Industry and Competitiveness, in the framework of the project CTM2016-76491-P. Miguel de Celis holds a PhD grant associated to this project (FPI grant: BES-2017-080024). The authors also want to thank to Biome Makers’s laboratory team for their efforts in NGS analysis of wastewater samples.
Author information
Authors and Affiliations
Contributions
L.A., D.M., S.S. and A.S. conceived the work. M.C., I.B. and A.S. designed the experimental and analytical approaches M.C., I.B. and R.O.-A. performed the data analysis. M.C., I.B., L.A., D.M., S.S. and A.S. discussed the results. M.C., I.B. and A.S. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
de Celis, M., Belda, I., Ortiz-Álvarez, R. et al. Tuning up microbiome analysis to monitor WWTPs’ biological reactors functioning. Sci Rep 10, 4079 (2020). https://doi.org/10.1038/s41598-020-61092-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-61092-1
This article is cited by
-
Niche differentiation drives microbial community assembly and succession in full-scale activated sludge bioreactors
npj Biofilms and Microbiomes (2022)
-
Coexistence of antibiotic resistance genes, fecal bacteria, and potential pathogens in anthropogenically impacted water
Environmental Science and Pollution Research (2022)
-
Multi-omics analysis on seasonal variations of the biofilm microbial community in a full-scale pre-denitrification biofilter
Environmental Science and Pollution Research (2022)
-
Bacterial diversity and predicted enzymatic function in a multipurpose surface water system – from wastewater effluent discharges to drinking water production
Environmental Microbiome (2021)
-
Community and single cell analyses reveal complex predatory interactions between bacteria in high diversity systems
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.