Abstract
In this study, we used shotgun metagenomic sequencing to characterise the microbial metabolic potential for lignocellulose transformation in the gut of two colonies of Argentine higher termite species with different feeding habits, Cortaritermes fulviceps and Nasutitermes aquilinus. Our goal was to assess the microbial community compositions and metabolic capacity, and to identify genes involved in lignocellulose degradation. Individuals from both termite species contained the same five dominant bacterial phyla (Spirochaetes, Firmicutes, Proteobacteria, Fibrobacteres and Bacteroidetes) although with different relative abundances. However, detected functional capacity varied, with C. fulviceps (a grass-wood-feeder) gut microbiome samples containing more genes related to amino acid metabolism, whereas N. aquilinus (a wood-feeder) gut microbiome samples were enriched in genes involved in carbohydrate metabolism and cellulose degradation. The C. fulviceps gut microbiome was enriched specifically in genes coding for debranching- and oligosaccharide-degrading enzymes. These findings suggest an association between the primary food source and the predicted categories of the enzymes present in the gut microbiomes of each species. To further investigate the termite microbiomes as sources of biotechnologically relevant glycosyl hydrolases, a putative GH10 endo-β-1,4-xylanase, Xyl10E, was cloned and expressed in Escherichia coli. Functional analysis of the recombinant metagenome-derived enzyme showed high specificity towards beechwood xylan (288.1 IU/mg), with the optimum activity at 50 °C and a pH-activity range from 5 to 10. These characteristics suggest that Xy110E may be a promising candidate for further development in lignocellulose deconstruction applications.
Similar content being viewed by others
Introduction
Over the past two decades, the global interest in the development of renewable energies has increased dramatically, particularly in the context of the climate change and the depletion of fossil fuels1,2. Bioethanol is considered a valuable renewable energy source capable of providing an alternative to petrol through blending with gasoline2,3. The main drawback with the production of bioethanol from lignocellulosic material is the cost of enzymatic hydrolysis because of low catalytic efficiencies of the enzymes currently in use1,2,4. A low-cost lignocellulose hydrolysis strategy is also of interest in other processes such as found in the textile, food, animal feed and paper industries1,5.
Lignocellulosic biomass mainly consists of polysaccharide polymers, cellulose and hemicellulose, and the phenolic polymer lignin. Termites are among the most efficient lignocellulose decomposers on earth, with hydrolysis efficiencies of up to 90%6,7,8,9. The ability of termites to degrade lignocellulose is more efficient than the digestion of less lignified forage grasses in ruminants6,10,11. This capacity to degrade lignocellulose with very high efficiency is due to a dual system that includes the mechanical and enzymatic machinery of the termite host, together with the action of intestinal symbionts6,9,12. This metabolic potential makes termites an ideal target to search for microbial lignocellulosic enzymes that might be used in the textile, food, animal feed, paper and biofuel industries9,13.
A wide range of microbial enzymes within the termite gut have been categorized in the different Carbohydrate-Active enZYmes (CAZy) classes (http://www.cazy.org/)14, including glycoside hydrolases (GHs), glycosyltransferases (GTs), polysaccharide lyases (PLs), carbohydrate esterases (CEs), carbohydrate-binding modules (CBMs) and auxiliary activity enzymes (AAs)15,16,17. Among these enzymes, some GHs families are particularly important for lignocellulosic biomass deconstruction, most especially in cellulose and hemicellulose degradation. For example, endo-β-1,4-glucanases, exo-β-1,4-glucanases or cellobiohydrolases, and β-glucosidases18,19 act in different sections of the cellulose polymer and its derived products, whereas endo-β-1,4-xylanases participate in the degradation of the xylan backbone, the main component of hemicellulose, to xylose. Other GHs families attack different substrates depending on the composition of the polysaccharide or its side chains, including α-L-arabinofuranosidases, endo-α-1,5-arabinanases, endo-β-1,4-mannanases, α-glucuronidases and α-L-fucosidases20,21. To date, there have been no reports of termite gut microbiome-derived enzymes that participate in lignin degradation, although this degradation process is known to occur20,22,23. The high alkalinity in termite gut segments may act as an alkaline pretreatment to facilitate subsequent lignin degradation24,25,26. It is noted that the anoxic conditions in some regions of the termite digestive tract may not support aerobic lignin degradation pathways10,27. It has been proposed that the ligninolytic capacity may be provided by termite host-derived enzymes, although this suggestion remains unresolved13,20.
Over the past 15 years, a substantial number of metagenomic, metatranscriptomic and metaproteomic studies of the gut microbiomes of termites and other wood-feeding insects have been reported, with the discovery of numerous enzymes involved in lignocellulose degradation20,21,28,29,30,31,32. A number of these genes have been cloned and their enzymatic activities characterized33,34,35,36,37. However, until July 2019 only two works referenced in PubMed has reported the cloning and expression of genes encoding GHs from termite gut microbiomes using shotgun metagenome sequencing15,36.
In this study, we selected two species of higher termites belonging to the Nasutermitinae subfamily with different life habits and diets (Cortaritermes fulviceps and Nasutitermes aquilinus). C. fulviceps builds mounds and is a polyphagous insect that feeds on leaves, roots and stems of various gramineous plants, as well as wood. By contrast, N. aquilinus inhabits live and dead trees and is a strict wood-feeder (monophagous) that consumes hardwoods or softwoods in dry, wet or decaying state38.
Our strategy involved an analysis of the gut bacterial diversity of the two termite colonies by using a sequence-driven metagenomic approach. This analysis allowed the identification of candidate genes coding for lignocellulose degrading enzymes. We also performed a comparative analysis of protein structures in a subset of GHs and selected and cloned a GH10 for further biochemical characterisation of enzyme functionality.
Materials and Methods
Termite sampling site
Worker caste specimens of the termite species N. aquilinus and C. fulviceps were collected from single colonies in the province of Corrientes, Argentina (S 27°28′30″: W 58°46′59.43″ and S 27°26′58.26″: W 58°44′17.64″, respectively). N. aquilinus and C. fulviceps were collected from live Enterobolium contortisiliquun trees and from inside a mound located in Elionurus muticus grassland, respectively.
The termites were collected with the authorization of the Direction of Natural Resources of Ministry of Tourism of the province of Corrientes (permission number 845/13). No endangered or protected species were used in this study. The specimens were stored at −20 °C until processing. A complete description of the sampling site is reported in Ben Guerrero et al.39.
DNA extraction and shotgun sequencing library preparation
The surface of the insects was disinfected with 70% ethanol before dissecting the whole guts under a binocular microscope. Ten guts per termite species were pooled and placed in tubes containing RNA-later (Ambion, Grand Island, USA).
For DNA shotgun sequencing, total genomic DNA was isolated using the DNeasy Blood and Tissue kit (Qiagen, USA) following the manufacturer’s instructions. Crude DNA extracts were further purified according to the QIAmp DNA micro protocol for tissues (Qiagen, USA). The DNA concentration was determined with a Qubit fluorometer (Qiagen) and the size range assessed using a bioanalyzer and gel electrophoresis. The extracted DNA from each sample was diluted to a final concentration of 200 ng for the preparation of a metagenomic library at the QB3 Vincent J. Coates Genomics Sequencing Laboratory (Berkeley, USA). Briefly, DNA was sheared using Covaris and the library preparation was performed using the Kapa Biosystems Library Prep system on the IntegenX Apollo 324 robot. Each sample was sequenced using an Illumina platform (HiSeq 2500 Rapid Run) to generate 150 bp paired-ends reads.
Metagenomic assembly and annotation
Illumina adapters were removed from the reads and their quality checked using FastQC with the default settings. Paired-end reads were exported to the KBase40 (KBase: The U.S. Department of Energy Systems Biology Database) for co-assembly and binning. A random subsample of 20 million paired-end reads per sample was used for assembly using the IDBA-UD assembler (http://i.cs.hku.hk/~alse/h7kubrg/projects/idba_ud/)41. Gene abundance was assessed using Bowtie 2 version 2.3.342 to map sequence reads to the assembled contigs and to quantify the number of reads per contig. The coverage was calculated on every contig using gene length estimates, on a coding sequence basis. The coverage information and taxonomy classification were used to investigate the compositions of gut prokaryote communities of the two termite colonies.
Obtained scaffolds were RPKM (Reads per kilobase per Million) normalized to account for sequencing depth and scaffold length. The number of reads for each assembled contig in each sample was normalized to reads per kilobase per million reads mapped. RPKM-normalized coverage values were used as proxies for the abundance of each scaffold/contig in a sample. Open reading frames were predicted from the final set of scaffolds using Prodigal’s43 meta procedure (-p meta).
All obtained contigs were assigned a taxonomic classification label using the Kaiju web application (https://github.com/bioinformatics-centre/kaiju)44 and coding sequences annotated using Prokka (http://www.vicbioinformatics.com/software.prokka.shtml)45.
The obtained contigs with a minimum length of 1,000 base-pairs were binned using MaxBin246 and the generated genome populations (here referred to as bins) were analysed using CheckM47 to assess genome quality. Thirty-three bins were obtained and 17 of this bins likely originated from a single bacterial strain/population with completeness ranging from 5–97%. They were further taxonomically phylotyped according to their corresponding predicted proteins using CheckM reference markers. Only these 17 genome bins and their associated glycoside hydrolases (GHs) were used to analyse the differences in GH abundances between the studied termites. Coverage information for the scaffolds of each genome was extracted from the calculated coverage data RPKM normalized for each scaffold in the metagenome. Bins abundances in each gut sample were calculated as the average RPKM-coverage value over all the scaffolds in a bin. Statistical significance of the coverage distribution of the identified genes was assessed using the Kruskal-Wallis test and pairwise comparisons, which were carried out using the Wilcox test and the Benjamini-Hochber method for p-value adjustment using R software.
Additional functional and KEGG (Kyoto Encyclopedia of Genes and Genomes) metabolic pathway annotations were determined with the KAAS web tool (http://www.genome.jp/tools/kaas/)48, to identify the high level functions and utilities of the biological systems. Putative genes involved in plant biomass degradation were identified by comparing the predicted ORFs with the protein families classified in the CAZy database using the web server dbCAN (http://csbl.bmb.uga.edu/dbCAN/annotate.php)49.
Data handling and statistical analyses
The R statistical software version 3.4 was used to filter, process and consolidate data obtained from different servers and software and for all statistical analyses.
3D modelling of glycosyl hydrolases
A 3D modelling analysis was performed for 26 putative GHs. Phylogenetic analysis of each GH family was done for the selection of protein-coding gene sequences that would be used for structural modelling. CAZy reference sequences were included together with sequences from our dataset that were assigned to the same family according to dbCAN. One sequence per family was chosen based on its dissimilarity to the reference set (phylogenetic trees not shown).
Subsequently, 3D modelling rounds with no restrictions were made using the I-TASSER server (https://zhanglab.ccmb.med.umich.edu/I-TASSER/)50. A 3D model was built for each sequence obtained from the gut metagenomic data, and close structural neighbours were identified. The Visual Molecular Dynamic program was used to visualize the 3D models51.
Cloning, expression and enzymatic activities assays
Assessments of the quality of sequence assembly for the identification of lignocellulose-degrading enzymes were performed by selecting a predicted GH-coding gene KBCPBGKF 45352, here termed Xyl10E. The criteria for the selection was as follows: the gene a) was predicted as able to deconstruct hemicellulose, which is one of the main components of plant cell walls, b) encoded a protein from a highly enriched GH family in the metagenomes, c) had a complete coding sequence containing identifiable start and stop codons and a complete open reading frame, d) contained a signal peptide in its gene product, which suggests it may encode a secreted enzyme, e) lacked transmembrane regions.
For PCR amplification and cloning of the Xyl10E gene, DNA was extracted from N. aquilinus gut samples using the QIAamp DNA Stool kit (Qiagen) with modifications. Briefly, pooled gut samples from six individuals were heated at 95 °C in 1 mL of kit lysis buffer, and then ground with a FastPrep protocol (3 cycles of 20 sec. at 6000 rpm) using 300 mg of 150–212 µm glass beads (Sigma, USA). After elution of DNA, an additional purification step was performed with Agencourt AMPure XP magnetic beads (Beckman Coulter, USA). For this purpose, 1.5 volumes of bead solution were added per sample, followed by 5 min magnet incubation and two ethanol 80% washes. Finally, the samples were incubated for 5 min in 50 µL Qiagen elution buffer before eluting the supernatants. DNA concentration and purity were assessed with Qubit® fluorometer. The Xyl10E sequence was amplified, without the native signal peptide, using specific primers (designed from assembled contigs) containing BamHI and XhoI restriction enzyme sites: Xyl10E-F: 5′ GGATCCTTCTGCGCCTGACA 3′, Xyl10E-R: 5′ CTCGAGCTATTCCACCAATTTCC 3′, for N-terminal fusion to a 6xHis tag (restriction sites are shown underlined). The amplification product was first cloned in pGEM-T Easy vector using E. coli DH5-α competent cells. Then, the plasmid inserts from selected colonies were cloned into pET28b(+) vector (BamHI/XhoI) and transformed into competent E. coli Rossetta cells. Xyl10E protein expression was induced with 0.5 mM IPTG for 16 h at 37 °C. After cell lysis and sonication (six pulses of 10 s, 28% amplitude), recombinant protein was purified in the soluble fraction with Ni-NTA agarose resin (Qiagen), using 50 mM NaH2PO4, 300 mM NaCl, 250 mM imidazole, pH 8 as elution buffer. Typically, 1.5 mg/mL of recombinant Xyl10E protein was obtained from 90 mL induced E. coli culture.
Enzyme activity assays were performed using the enzyme (diluted in appropriate buffer) at a final concentration of 7.5 μg/mL. Endo-β-1,4-xylanase and endo-β-1,4-glucanase activities were determined in triplicate in microtube assays. For this purpose, 50 μL of purified protein were combined with 50 μL of beechwood xylan (Sigma) (1% w/v) or 50 μL of carboxymethyl cellulose (CMC) (Sigma) (2% w/v) in 0.1 M citrate buffer (pH 6) and incubated for 20 min at 50 °C in a Thermomixer (Eppendorf). Reducing sugars released from the polysaccharide hydrolysis were measured by dinitrosalicylic acid (DNS)52 with xylose or glucose standard curves. Specific activity was calculated per mg of total protein (IU/mg). For all enzymatic assays, one international unit (IU) was defined as the amount of enzyme that released 1 µmol of product per minute under the specified assay conditions.
Results
Assembly and analysis of metagenomic sequencing data
The analysis of the intestinal DNA extracted from C. fulviceps and N. aquilinus specimens generated 52 Gb of sequence reads. The reads were assembled into 86,012 and 72,572 contigs for C. fulviceps and N. aquilinus, respectively (Table 1). Eubacteria accounted for 92.7% (C. fulviceps) and 98.2% (N. aquilinus) of all classified contigs. Other less represented taxonomic groups included Eukaryote (protists (1.6%), fungi (1.7%) and viridiplantae (0.2%)), virus (0.2%) and Archaea (0.5%) on average for both termite microbiomes. DNA insect contamination was low (4%).
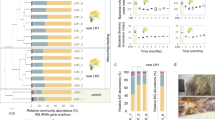
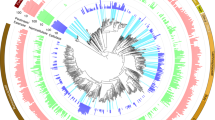
We then analysed the microbial community compositions and determined the abundances of bacterial phyla in the guts of both termite colonies. Spirochaetes, Firmicutes, Proteoabacteria, Fibrobacteres, Bacteroidetes and Actinobacteria were the most abundant phyla in both termite gut microbiomes (Fig. 1). Both microbiomes showed a low percentage of unclassified bacteria (~4.8%). The five dominant phyla were the same for both termites’ samples, with only minor differences in the relative abundances. In C. fulviceps, the most abundant phylum was Spirochaetes (47%), followed by Proteobacteria and Firmicutes (almost in equal proportions, ca. 17%), Bacteroidetes (5%) and Fibrobacteres (2%) and N. aquilinus with Spirochaetes (47%), Firmicutes (17%), Proteobacteria (10%), Fibrobacteres (8%) and Bacteroidetes (5%) as the predominant phyla (Fig. 1). Given that the analysis involved only single composite metagenomes from each termite species, no statistical significance can be assigned to these differences. The bacterial phyla, Actinobacteria, Acidobacteria, Cyanobacteria and Planctomycetes, all accounted for more than 2% of the total sequences reads.
Altogether, these nine phyla represented 92.5% and 91.1% of total reads in the gut microbiomes of the C. fulviceps and N. aquilinus specimens, respectively.
Comparative functional and metabolic analysis
To investigate the diversity of microbial enzymes in the gut samples, we annotated open reading frames (ORFs) in contigs using Prokka and assigned the predicted proteins to metabolic pathways with KEGG. We assessed the metabolic functions of gut microbiomes from the two termite colonies and classified 20 different categories with frequency of more than 2% (Fig. 2).
Annotations relating to amino acid metabolism (AAM) and carbohydrate metabolism (CM) accounted for ca. 30% of all predicted ORFs, regardless of the termite sample (Fig. 2). Interestingly, predicted AAM genes were the most abundant in the C. fulviceps gut microbiome, whereas ORFs involved in CM dominated in the N. aquilinus sample.
Cellulose and xylan degradation
To further explore the potential capacity of the gut microbiota to degrade lignocellulosic substrates, we screened the putative encoded protein sequences using the Carbohydrate-Active Enzymes (CAZymes) catalogue via the dbCAN annotation web server. We detected 585 and 1,967 putative CAZymes corresponding to 117 and 160 different families, in the C. fulviceps and N. aquilinus gut microbiome samples, respectively (Table 2). GHs, GTs, CBMs and CEs classes were the most abundant CAZyme categories, together represented 95.9% (C. fulviceps) and 97.4% (N. aquilinus) of all the CAZyme related annotations. Glycoside hydrolases were the most highly represented enzyme class with 40.3% (C. fulviceps) and 37.6% (N. aquilinus) of total ORFs. According to the dbCAN database analysis, the 975 GHs corresponded to 71 different CAZy families.
The grouping proposed by Allgaier et al.53 arranges all known GHs according to their main functional role (cellulases, hemicellulases, debranching enzymes, oligosaccharide degradation, and cell wall elongation). Following this approach, we identified 21 and 26 families that were potentially involved in lignocellulose degradation in the C. fulviceps and N. aquilinus samples, respectively (Table 3). The frequencies of these different functional groups were similar for the gut microbiome samples of the two termites. For both termite gut samples, the most abundant groups were cellulases and oligosaccharide-degrading enzymes. Three cellulase families (GH5, GH9 and GH45) were present in both termite specimens, whereas GH44 was only found in the N. aquilinus gut sample. In both termite colonies, the most abundant ORFs corresponded to GH5 cellulases. Other GHs included in the top eight lignocellulose-degrading enzymes were hemicellulases (GH10 and GH11), oligosaccharide-degrading enzymes (GH43, GH3 and GH1), cellulases (GH9) and cell wall elongation enzymes (GH74) (Fig. 3).
An association between a higher relative abundance of Spirochaetes and an increased level of cellulose degrading enzymes has been previously reported. To test this, we extracted genomes from the metagenomes of the analysed termites based on coverage and composition. A total of 33 genome populations were binned, with 17 of these genomes showing a completeness of 5–97% and identified as consisting only of bacterial elements. Glycoside hydrolases and their corresponding normalized coverage were extracted from the binned genomes and differences in their abundances between the two termite samples analysed (Fig. S1). The obtained results showed a significantly higher abundance of genome-associated GHs in N. aquilinus than in C. fulviceps. This finding was further evaluated by comparing the normalized average-coverage of the extracted genomes (Fig. S2).
3D modelling analysis of glycosyl hydrolases
Twenty six putative GH sequences obtained from the metagenomic analysis were modelled using I-TASSER (Table 4). Each modelled sequence was associated to a GH family based on both the templates and structural analogues identified, as deposited in the PDB database. All 26 sequences were identified as GHs and grouped in 23 different families, according to their structural properties. For most, the identification was in accordance with the CAZy classification reported in Table 3. Nevertheless, the predicted proteins AFHCADON 03545, KBCPBGKF 10146 and KBCPBGKF 23895, which were identified as GH 53, 106 and 38 (CAZy classification), respectively, were identified as members of GH1 and GH26 families according to protein modelling.
In general, the 3D models showed C-score values between −2.88 to 2.00 and TM values from 0.39 to 0.99. The C-score is estimated based on the significance of threading template alignments and the parameters obtained from the structure assembly simulations. The C-score values range from −5 to 2 where higher values indicate a higher confidence model. TM, which varies from 0 to 1, is calculated from the C-score, where the optimum TM value for a 3D model is 1. According to these parameters, the 3D models were generally estimated to be of high quality. However, some 3D models (i.e. models obtained from sequences: KCPBGKF 24894, KBCPBGKF 39042, KBCPBGKF 08463) showed low quality scores according to the C-score and TM values.
As previously noted, GH families 3, 5, 10, and 43 were the most abundant in both termite gut metagenomes. Members of GH family 10 have been extensively studied in terms of their capacity for hemicellulose deconstruction54,55,56,57,58,59,60,61,62,63,64. Accordingly, from the 26 modelled GHs, we selected the sequence KBCPBGKF 45352, a GH family 10 enzyme henceforth termed Xyl10E, for further structural analysis and biochemical characterization. The sequence presented 45% identity with a endo-β-1,4-xylanase from Treponema azotonutricium (GenBank: AEF82584.1)65.
The first attempt to model Xyl10E yielded five possible models (best model: C-score = −0.17; TM = 0.69 ± 0.12 and RMSD = 7.2 ± 4.2 Å). PDBs 2FGL, 6FHF, 4W8L, 2Q8X, 5OFJ and 2UWF (where the code identifies a crystallographic structure deposited in the Protein Data Bank) were the templates used by i-ITASSER to produce Xyl10E 3D-models. Proteins identified as both templates and structural analogues belonged to GH family 1066. To obtain a more accurate 3D-model, we performed a second round of modelling and obtained six models by using each identified structural analogue as a separate template. PDB 6FHF was found to be the best template to produce an optimum Xyl10E 3D-model. However, the quality values (C-score = 0; TM = 0.71 ± 0.11; RMSD = 6.8 ± 4.1 Å) obtained during this second modelling round suggested some important structural differences with those PDB identified as structural neighbours. Curiously, 6FHF is an unusual GH10 sequence, because it was designed by using rational protein design approaches, and generated by automated combinatorial backbone assembly and sequence design67. This observation supports the importance of using metagenomic approaches for the discovery of enzymes with unusual sequences.
The structural analysis of the Xyl10E model (Fig. 4) confirmed the characteristic folding topology of the GH10 family, an eight-fold TIM-barrel structure. Xyl10E showed the catalytic residues, E180 and E314, that are conserved in this family (Fig. 4A,B), as well as the seven conserved residues of the active site (Fig. 4C). Furthermore, this enzyme had one (V169) of three residues involved in the sensitivity to alkaline pH described in the active alkaline xylanase of Bacillus halodurans S7 PDB 2UWF68 (Fig. 4D) and an aromatic box, which is also characteristic of the GH10 family (Fig. 4E). We also assessed the electrostatic potential of Xyl10E (Fig. 4F) and found that the negative charge is concentrated in the centre of the sequence, which is consistent with the active alkaline xylanase of PDB 2UWF.
The overall TIM-barrel structure of Xyl10E. (A) Catalytic residues are shown in red (E180 and E314). (B) Catalytic residues of GH10 family are shown in red, blue and green in Xyl10E, PDB 6FHF, PDB 2W5F, respectively. (C) Another seven residues strictly conserved in GH10 are shown in orange and black in Xyl10E and PDB 2W5F, respectively. (D) Residues involved in the alkaline sensitive xylanases are shown in orange and blue in Xyl10E and PDB 2UWF, respectively. (E) Aromatic residues forming the aromatic cage that surround the catalytic pocket. W108, W326 and W334 (in blue) and W126, W368 and W376 (in orange) belong to PDB 2UWF and Xyl10E, respectively. (F) Electrostatic potential of Xyl10E. Negatively and positively charged surfaces are coloured in red and blue, respectively.
Verification of sequence assembly and evaluation of enzymatic activities
We subsequently validated the predicted data by evaluating the activity of the recombinant Xyl10E. Xyl10E was successfully cloned and expressed in E. coli as an N-terminal His-tag fusion protein, and purified in a soluble form that allowed the subsequent functional characterization.
The predicted molecular weight (MW) and isoelectric point (IP) of recombinant Xyl10E were 49.2 kDa and 6.31, respectively. The purified protein showed an apparent monomeric molecular weight of 49 kDa, in accordance with the predicted size (Fig. 5A,B).
Expression, purification and enzymatic characterization of soluble Xyl10E. (A) SDS-PAGE, M: molecular weight marker, T: total protein content of cell lysates without induction, S: soluble fraction of cell lysates, FT: flow through, W1 and W3: washed fractions with 20 mM imidazole, E1 to E4: serial elution fractions with 250 mM imidazole. The arrow indicates the band corresponding to Xyl10E (49.2 kDa). (B) Western blot revealed with anti-His antibody and peroxidase activity. (C,D) Effect of pH and temperature on the recombinant xylanase.
According to the enzymatic activity assays, Xyl10E had a specific endo-β-1,4-xylanase activity of 288.1 IU/mg of enzyme, with an optimum activity at around 50 °C and pH 6. Interestingly, this enzyme retained more than 50% of its optimum activity over a wide temperature range (30 to 60 °C) and more than 80% over a wide pH range (5 to 10) (Fig. 5C,D). Endo-β-1,4-glucanase activity was negligible.
Discussion
In recent years, the search for new enzymes that degrade lignocellulose has become essential. These enzymes are important for biofuel production and other industries, such as paper, food and textile. In this context, the ability of termites to feed on wood and other types of plant biomass makes them an ideal system to obtain efficient cell wall degrading enzymes6,9,69,70. In higher termites, many of the main cellulolytic enzymes are produced by their bacterial endosymbionts. For this reason, a comprehensive exploration of gut microbiota is essential to understand the processes involved in lignocellulose digestion. Here, we characterized the bacterial community hosted in the guts of C. fulviceps and N. aquilinus colonies and identified the relevant putative proteins involved in lignocellulose degradation. Spirochaetes, Firmicutes, Proteobacteria, Fibrobacteres and Bacteroidetes were the dominant bacterial phyla in both termite specimens and Spirochaetes accounted for almost half of the sequences present in both termite samples. The same dominant bacterial phyla have been observed in other wood and grass feeders termites such as Nasutitermes corniger, N. ephratae, Microcerotermes sp., N. takasagoensi and Mironasutitermes shangchengensis20,21,71,72,73,74.
Spirochaetes seems to be important for the survival of higher termites75,76, as this group is known to be involved in all of the major functions in the termite hindgut (fibre hydrolysis, fermentation, homoacetogenesis and nitrogen fixation)20,21,74,77,78,79. The dominance of Spirochaetes in the hindgut environment may be linked to their high mobility in viscous media and to the high surface to volume ratio of their cells6,77. Studies of the hindguts microbiota of Nasutitermes and Amitermes spp.20,21 have attributed the abundance of glycoside hydrolases putatively involved in cellulose degradation to Spirochaetes. Our results demonstrate that N. aquilinus harbours Spirochaetes in higher abundance than C. fulviceps (Fig. S2A), and that these Treponema sp. genome-bins contain a larger array of putative GHs with functions related to cellulose and hemicellulose degradation (Fig. S2B). Interestingly, some non-homoacetogenic Treponema spp. isolated from lower termites degrade cellobiose80,81. This finding indicates that they also have an important role in fibre digestion.
In our study, most of the sequences assigned to the phylum Spirochaetes belonged to the genus Treponema (71.6% in N. aquilinus colony and 68% in C. fulviceps). This genus includes protist ectosymbionts and free-living bacteria in the lumen of the hindgut, both participating in the process of reductive acetogenesis to produce acetate, the main nutrient for the termite host6,82,83.
N. aquilinus feeds exclusively on wood, whereas C. fulviceps consumes more nitrogen rich organic matter, including either living or decaying plant tissues84,85. This difference in diet may explain the metabolic pathway profiles of each termite colony. Whereas the amino acid metabolism (AAM) was the dominant metabolic function in the C. fulviceps gut microbiome, carbohydrate metabolism (CM) was the predominant pathway in N. aquilinus gut microbiome, according to the analysis of ORFs assigned after contig annotation.
Other relevant metabolic categories were broadly similar in both termite microbiomes, supporting the hypothesis that the termites gut microbiomes retain a stable core set of metabolic functions.
The CAZy database classifies cellulases and other plant cell wall polysaccharides degrading enzymes into GHs families. Previous reports show that roughly 34% of GHs families contain enzymes that contribute to plant cell wall deconstruction86. In this study, we identified 975 GHs, corresponding to 71 different CAZy families.
We subsequently sorted the GHs involved in lignocellulose degradation according to the arrangement proposed by Allgaier et al.53. This analysis revealed 21 and 26 distinct GHs families present in the C. fulviceps and N. aquilinus gut microbiomes, respectively. These results are consistent with those reported by He et al.20, where a study of the gut microbiomes of Amitermes wheeleri (dung feeder) and N. corniger (wood feeder) identified around 25 GHs families.
The GHs family classifications are based on protein sequence and structure and therefore does not necessarily accurately predict enzyme activities. Most GHs families comprise proteins with different enzymatic activities and proteins with similar activity can be found in different GHs families87. Endoglucanases and other GHs involved in cellulose degradation can be found in several GHs families. For example, β-glucosidases are found in six GHs families and cellobiohydrolases are distributed across three GHs families. In consequence, the prediction of CAZyme enzymatic activity based on their sequences alone is difficult88.
We identified multiple cellulases, especially from families GH5 and GH9, in the gut sample from N. aquilinus. This termite species has a largely wood-based diet, and the high abundance of cellulases in its gut microbiome is consistent with its dietary preference. C. fulviceps was collected from inside a mound located in grassland and its gut microbiome was enrich in debranching- and oligosaccharide-degrading enzymes, in particular α-L-arabinofuranosidases (GH3, GH42 and GH43). In general, grass is composed of cellulose fibres surrounded by hemicellulose, mainly xylans, annotated with arabinose residues in the form of arabinoxylans and glucuronoarabinoxylans89,90. The high abundance of these GHs families in the C. fulviceps gut microbiome is consistent with its feeding habits, which include grass foraging.
The analysis of cellulolytic GHs distribution suggested that the set of enzymes was different in the termite microbiomes. This could be determined by host factors, diet or taxonomic composition. Thus, further research on microbial enzymes from Nasutitermitinae is necessary to better understand how these variables influences the cellulolytic enzyme diversity.
We also investigated the most abundant annotated ORFs belonging to GHs families in the two metagenome sequence datasets. In both termite gut metagenomes, the most abundant cellulolytic ORFs belonged to family GH5. This is a large multigene family that includes endoglucanases (cellulases) and endo-mannanases, as well as exo-glucanases, exo-mannanases, β-glucosidases and β-mannosidases91. Our results are comparable with previous reports in Amitermes wheeleri and N. corniger termites, where GH5 was identified among the most abundant GHs families20. Furthermore, a high proportion of hemicellulases identified in the gut microbiomes of both termites belonged to the GH10 family, followed by members of GH11.
The 3D modelling of the 26 selected protein sequences yielded high quality structural models in most cases, with only three proteins yielding low quality 3D models. The sequences of proteins that were identified and 3D modelled revealed similarity with proteins with a wide range of enzymatic activities, including glucosidases, xylanases, cellulases, rhamnogalactosidases, mannanases, xylosidases, laminarinases and arabinofuranosidases. This result demonstrates that the GH sequences identified in these termites represent a valuable resource for the identification of new genes and gene products for possible use in lignocellulose deconstruction.
A structural analysis of the sequence KBCPBGKF 45352 (Xyl10E) revealed that this protein belongs to GH family 10, showing the correct folding topology, the key catalytic residues and the conserved active site residues typical of proteins in the GH10 family. This analysis, in conjunction with the activity data, confirmed conclusively that Xyl10E is an endo-β-1,4-xylanase of the GH10 family capable of functioning at an alkaline pH.
The analysis of the enzymatic activity of xylanase Xyl10E showed that the enzyme had a specific activity of 288 IU/mg. This value is 3-and 22-fold higher than the experimental values for endo-xylanases from Paenibacillus sp., rGH10XynA (~100 IU/mg) and HC1 (~13 IU/mg), respectively55,63 and 5- and 15- fold higher than that of Cohnella laevirobosi HY-21 (~58 IU/mg)58 and Massilia sp. XynRBM26 (~20 IU/mg)64, respectively. Conversely, the specific activity of Xyl10E was of the same order as several xylanases recovered from functional metagenomics analyses, Xyn10N18 derived from bovine rumen (~242 IU/mg)56 and SCXyl extracted from sugarcane soil bacteria (~200 IU/mg)54 and Xyl-ORF19, from the gut microbiome of termite Globitermes brachycerastes exhibited a specific activity of (~114 IU/mg)57.
Most of the reported GH10 family xylanases have been recovered from isolated bacteria or from functional screening of metagenomic libraries56,57,58,59,60,61,62,63,64,92,93,94,95. This study shows the value of metagenome DNA assemblages as a source of novel enzymes.
The optimal temperature of Xyl10E was 50 °C and this enzyme retained more than 90% of its optimum activity at 60 °C. These temperatures are in the same range as those found in other metagenome-derived xylanases of the GH10 family54. Kim et al.60,61,62 characterised several endo-xylanases from GH10 family, all of which were cloned from insect endosymbiont bacteria, and showed that their optimal temperatures range from 50 to 70 °C.
The endo-β-1,4-xylanase Xyl10E showed an optimum activity at pH 6 and retained around 80% of its activity between pH 5 and pH 10. This is consistent with the broad pH-activity ranges reported in xylanases of bacterial origin59,61,64,95,96.
Many GH10 xylanases showed both endo-β-1,4-xylanase and endo-β-1,4-glucanase activities93,94,97, although some enzymes of this family exhibit only xylanase activity: e.g. Xyn10N18 from a bovine rumen metagenomic library56 and XynAMG1 from chicken cecum92. The endoglucanase activity of Xyl10E against CMC was negligible.
This study has demonstrated that the gut microbiomes of these neotropical higher termite species encode a high diversity of enzymes that are potentially involved in plant cell wall degradation. Further study of these genes and their products might reasonably be expected to produce novel sequences, novel enzyme activities and even novel specificities.
Data availability
All sequence data were deposited in the NCBI Sequence Read Archive under the BioProject accession number PRJNA475948.
References
Batista-García, R. A. et al. From lignocellulosic metagenomes to lignocellulosic genes: trends, challenges and prospects. Biofuels, Bioprod. Bioref. 10, 864–882, https://doi.org/10.1002/bbb.1709 (2016).
Robak, K. & Balcerek, M. Review of Second Generation Bioethanol Production from Residual Biomass. Food Technol. Biotechnol. 56, 174–187, https://doi.org/10.17113/ftb.56.02.18.5428 (2018).
Hill, J., Nelson, E., Tilman, D., Polasky, S. & Tiffany, D. Environmental, economic, and energetic costs and benefits of biodiesel and ethanol biofuels. Proc. Natl Acad. Sci. USA 103, 11206–11210, https://doi.org/10.1073/pnas.0604600103 (2006).
Gao, D. et al. Hemicellulases and auxiliary enzymes for improved conversion of lignocellulosic biomass to monosaccharides. Biotechnology for Biofuels 4, https://doi.org/10.1186/1754-6834-4-5 (2011).
Kuhad, R. C., Gupta, R. & Singh, A. Microbial Cellulases and Their Industrial Applications. Enzyme Res. 2011, 1–10, https://doi.org/10.4061/2011/280696 (2011).
Brune, A. Symbiotic digestion of lignocellulose in termite guts. Nat. Rev. Microbiology 12, 168–180, https://doi.org/10.1038/nrmicro3182 (2014).
Shi, P. et al. Comparative Genomic Analysis of the Endosymbionts of Herbivorous Insects Reveals Eco-Environmental Adaptations: Biotechnology Applications. PLoS Genetics 9, e1003131, https://doi.org/10.1371/journal.pgen.1003131 (2013).
Sun, J. Z., Ding, S. Y. & Peterson, J. D. In Biological Conversion of Biomass for Fuels and Chemicals: Exploration from Natural Utilization Systems (eds. Sun, J. Z., Ding, S. Y. & Peterson, J. D.) 195–222 (Royal Society of Chemistry, 2013).
Talia, P. & Arneodo, J. In Termites and Sustainable Management Vol. 1 Sustainability in Plant and Crop Protection (eds. Aslam. Khan & Wasim. Ahmad) Ch. 5, 101–117 (Springer series, 2018).
Breznak, J. A. & Brune, A. Role of microorganisms in the digestion of lignocellulose by termites. Annu. Rev. Entomology 39, 453–487, https://doi.org/10.1146/annurev.en.39.010194.002321 (1994).
Watanabe, H. & Tokuda, G. Cellulolytic Systems in Insects. Annu. Rev. Entomology 55, 609–632, https://doi.org/10.1146/annurev-ento-112408-085319 (2010).
König, H., Li, L. & Fröhlich, J. The cellulolytic system of the termite gut. Applied Microbiology and Biotechnology, https://doi.org/10.1007/s00253-013-5119-z (2013).
Scharf, M. E. Termites as targets and models for biotechnology. Annu. Rev. Entomology 60, 77–102, https://doi.org/10.1146/annurev-ento-010814-020902 (2015).
Lombard, V., Golaconda Ramulu, H., Drula, E., Coutinho, P. M. & Henrissat, B. The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res. 42, 490–495, https://doi.org/10.1093/nar/gkt1178 (2014).
Tokuda, G. et al. Fiber-associated spirochetes are major agents of hemicellulose degradation in the hindgut of wood-feeding higher termites. Proc Natl Acad Sci USA 115, E11996–E12004, https://doi.org/10.1073/pnas.1810550115 (2018).
Zhang, D. et al. Carbohydrate-active enzymes revealed in Coptotermes formosanus (Isoptera: Rhinotermitidae) transcriptome. Insect Mol. Biol. 21, 235–245, https://doi.org/10.1111/j.1365-2583.2011.01130 (2012).
Marynowska, M. et al. Optimization of a metatranscriptomic approach to study the lignocellulolytic potential of the higher termite gut microbiome. BMC Genomics 18, https://doi.org/10.1186/s12864-017-4076-9 (2017).
Lynd, L. R., Weimer, P. J., Van Zyl, W. H. & Pretorius, I. S. Microbial cellulose utilization: fundamentals and biotechnology. Microbiol. Mol. Biol. Rev. 66, 506–577, https://doi.org/10.1128/mmbr.66.3.506-577 (2002).
Ezeilo, U. R., Zakaria, I. I., Huyop, F. & Wahab, R. A. Enzymatic breakdown of lignocellulosic biomass: the role of glycosyl hydrolases and lytic polysaccharide monooxygenases. Biotechnol. Biotechnological Equip. 31, 647–662, https://doi.org/10.1080/13102818.2017.1330124 (2017).
He, S. et al. Comparative Metagenomic and Metatranscriptomic Analysis of Hindgut Paunch Microbiota in Wood- and Dung-Feeding Higher Termites. PLoS One 8, e61126, https://doi.org/10.1371/journal.pone.0061126 (2013).
Warnecke, F. et al. Metagenomic and functional analysis of hindgut microbiota of a wood-feeding higher termite. Nat. 450, 560–565, https://doi.org/10.1038/nature06269 (2007).
Li, H. et al. Lignocellulose pretreatment in a fungus-cultivating termite. Environ. Sci. 114, 4709–4714, https://doi.org/10.1073/pnas.1618360114 (2017).
Zhou, J. et al. Diversity, Roles, and Biotechnological Applications of Symbiotic Microorganisms in the Gut of Termite. Curr. Microbiology 76, 755–761, https://doi.org/10.1007/s00284-018-1502-4 (2019).
Köhler, T., Dietrich, C., Scheffrahn, R. H. & Brune, A. High-Resolution Analysis of Gut Environment and Bacterial Microbiota Reveals Functional Compartmentation of the Gut in Wood-Feeding Higher Termites (Nasutitermes spp.). Appl. Environ. Microbiology 78, 4691–4701 (2012).
Brune, A., Emerson, D. & Breznak, J. A. The Termite Gut Microflora as an Oxygen Sink: Microelectrode Determination of Oxygen and pH Gradients in Guts of Lower and Higher Termites. Appl. Environ. Microbiology 61, 2681–2687, https://doi.org/10.1128/AEM.61.7.2681-2687 (1995).
Brune, A. & Kühl, M. pH profiles of the extremely alkaline hindguts of soil-feeding termites (Isoptera: Termitidae) determined with microelectrodes. J. Insect Physiol. 42, 1121 (1996).
Brune, A. & Ohkuma, M. In Biology of Termites: a Modern Synthesis (eds Bignell, D., Roisin, Y. & Lo, N.) 439–444 (Springer, 2010).
Do, T. H. et al. Mining biomass-degrading genes through Illumina-based de novo sequencing and metagenomic analysis of free-living bacteria in the gut of the lower termite Coptotermes gestroi harvested in Vietnam. J. Biosci. Bioeng. 6, 665–671, https://doi.org/10.1016/j.jbiosc.2014.05.010 (2014).
Liu, N. et al. Metagenomic insights into metabolic capacities of the gut microbiota in a fungus-cultivating termite (Odontotermes yunnanensis). PLoSONE, e69184, https://doi.org/10.1371/journal.pone.0069184 (2013).
Rajarapu, S. P., Shreve, J. T., Bhide, K. P., Thimmapuram, J. & Scharf, M. E. Metatranscriptomic profiles of Eastern subterranean termites, Reticulitermes flavipes (Kollar) fed on second generation feedstocks. BMC Genomics 16, 332, https://doi.org/10.1186/s12864-015-1502-8 (2015).
Sethi, A., Slack, J. M., Kovaleva, E. S., Buchman, G. W. & Scharf, M. E. Lignin-associated metagene expression in a lignocellulose-digesting termite. Insect Biochem. Mol. Biol. 43, 91–101, https://doi.org/10.1016/j.ibmb.2012.10.001 (2013).
Ceja-Navarro, J. A. et al. Gut anatomical properties and microbial functional assembly promote lignocellulose deconstruction and colony subsistence of a wood-feeding beetle. Nat. microbiology 4, 864–875, https://doi.org/10.1038/s41564-019-0384-y (2019).
Bastien, G. et al. Mining for hemicellulases in the fungus-growing termite Pseudacanthotermes militaris using functional metagenomics. Biotechnol. Biofuels 6, 78, https://doi.org/10.1186/1754-6834-6-78 (2013).
Cairo, J. P. L. F. et al. Functional characterization and target discovery of glycoside hydrolases from the digestome of the lower termite Coptotermes gestroi. Biotechnol. Biofuels 4, 1–10 (2011).
Nimchua, T., Thongaram, T., Uengwetwanit, T., Pongpattanakitshote, S. & Eurwilaichitr, L. Metagenomic analysis of novel lignocellulose-degrading enzymes from higher termite guts inhabiting microbes. J. Microbiol. Biotechnol. 22, 462–469 (2012).
Woon, J. S. et al. Cloning, Production and Characterization of a Glycoside Hydrolase Family 7 Enzyme from the Gut Microbiota of the Termite Coptotermes curvignathus. Mol. Biotechnol. 59, 271–283, https://doi.org/10.1007/s12033-017-0015-x (2017).
Zhang, M. et al. Phylogenetic and Functional Analysis of Gut Microbiota of a Fungus-Growing Higher Termite: Bacteroidetes from Higher Termites Are a Rich Source of β-Glucosidase Genes. Microb. Ecol. 68, 416–425, https://doi.org/10.1007/s00248-014-0388-3 (2014).
Laffont, E. R., Torales, G. J., Coronel, J. M., Arbino, M. O. & Godoy, M. C. Termite (Insecta, Isoptera) fauna from National Parks of the north-east region of Argentina. Sci. Agricola 61, 665–670, https://doi.org/10.1590/S0103-90162004000600016 (2004).
Ben Guerrero, E. et al. Prospection and Evaluation of Cellulolytic and Hemicellulolytic Enzymes Using Untreated and Pretreated Biomass in two Argentinean native Termites. PLoS One 10, e0136573, https://doi.org/10.1371/journal.pone.0136573 (2015).
Arkin, A. P. et al. KBase: The United States Department of Energy Systems Biology Knowledgebase. Nat. Biotechnol. 36, 566–569, https://doi.org/10.1038/nbt.4163 (2018).
Peng, Y., Leung, H. C., Yiu, S. M. & Chin, F. Y. IDBA-UD: a de novo assembler for single-cell and metagenomic sequencing data with highly uneven depth. Bioinforma. 28, 1420–1428, https://doi.org/10.1093/bioinformatics/bts174 (2012).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359, https://doi.org/10.1038/nmeth.1923 (2012).
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinf. 11, 119, https://doi.org/10.1186/1471-2105-11-119 (2010).
Menzel, P., Ng, K. L. & Krogh, A. Fast and sensitive taxonomic classification for metagenomics with Kaiju. Nat. Commun. 13, 11257, https://doi.org/10.1038/ncomms11257 (2016).
Seemann., T. Rapid prokaryotic genome annotation. Bioinforma. 15, 2068–2069, https://doi.org/10.1093/bioinformatics/btu153 (2014).
Wu, Y.-W., Simmons, B. A. & Singer, S. W. MaxBin 2.0: an automated binning algorithm to recover genomes from multiple metagenomic datasets. Bioinforma. 32, 605–607, https://doi.org/10.1093/bioinformatics/btv638 (2016).
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 25, 1043–1055, https://doi.org/10.1101/gr.186072.114 (2015).
Moriya, Y., Itoh, M., Okuda, S., Yoshizawa, A. C. & Kanehisa, M. KAAS: an automatic genome annotation and pathway reconstruction server. Nucleic Acids Res. 35, W182–185, https://doi.org/10.1093/nar/gkm321 (2007).
Yin, Y. et al. DbCAN: A web resource for automated carbohydrate-active enzyme annotation. Nucleic Acids Res. 40, 445–451, https://doi.org/10.1093/nar/gks479 (2012).
Roy, A., Kucukural, A. & Zhang, Y. I-TASSER: a unified platform for automated protein structure and function prediction. Nat. Protoc. 5, 725–738, https://doi.org/10.1038/nprot.2010.5 (2010).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 33–38, 27–38, https://doi.org/10.1016/0263-7855(96)00018-5 (1996).
Miller, G. L. Use of Dinitrosalicylic Acid Reagent for Determination of Reducing Sugar. Anal. Chem. 31, 426–428 (1959).
Allgaier, M. et al. Targeted discovery of glycoside hydrolases from a switchgrass-adapted compost community. PLoS One 5, e8812, https://doi.org/10.1371/journal.pone.0008812 (2010).
Alvarez, T. M. et al. Development and biotechnological application of a novel endoxylanase family GH10 identified from sugarcane soil metagenome. PLoS One 8, e70014, https://doi.org/10.1371/journal.pone.0070014 (2013).
Ghio, S. et al. Paenibacillus sp. A59 GH10 and GH11 extracellular endoxylanases: application in biomass bioconversion. BioEnergy. Res. 11, 174, https://doi.org/10.1007/s12155-017-9887-7 (2018).
Gong, X., Gruniniger, R. J., Forster, R. J., Teather, R. M. & McAllister, T. A. Biochemical analysis of a highly specific, pH stable xylanase gene identified from a bovine rumen-derived metagenomic library. Appl. Microbiology Biotechnol. 97, 2423–2431, https://doi.org/10.1007/s00253-012-4088-y (2013).
Han, Q. et al. Biochemical characterization and crystal structure of a GH10 xylanase from termite gut bacteria reveal a novel structural feature and significance of its bacterial Ig-like domain. Biotechnol. Bioeng. 110, 3093–3103, https://doi.org/10.1002/bit.24982 (2013).
Kim, D. Y. et al. Novel intracellular GH10 xylanase from Cohnella laeviribosi HY-21: Biocatalytic properties and alterations of substrate specificities by site-directed mutagenesis of Trp residues. Bioresour. Technol. 101, 8814–8821, https://doi.org/10.1016/j.biortech.2010.06.023 (2010).
Kim, D. Y. et al. Novel GH10 xylanase with a fibronectin type 3 domain, from Cellulosimicrobium sp. strain HY-13, a bacterium in the gut of Eisenia fetida. Appl. Env. Microbiol. 75, 7275–7279 (2009).
Kim, D. Y. et al. Genetic and functional characterization of a novel GH10 endo-ß- 1,4-xylanase with a ricin-type ß-trefoil domain-like domain from Luteimicrobium xylanilyticum HY-24. Int. J. Biol. Macromol. 106, 620–628, https://doi.org/10.1016/j.ijbiomac.2017.08.063 (2018).
Kim, D. Y. et al. Novel alkali-tolerant GH10 endo-ß-1,4-xylanase with broad substrate specificity from Microbacterium trichothecenolyticum HY-17, a gut bacterium of the mole cricket Gryllotalpa orientalis. J. Microbiol. Biotechnol. 24, 943–953, https://doi.org/10.4014/jmb.1405.05032 (2014).
Kim, D. Y. et al. Biocatalytic Properties and Substrate-binding Ability of a Modular GH10 ß-1,4-Xylanase from an Insect-symbiotic Bacterium, Streptomyces mexicanus HY-14. J. Microbiol. 52, 863–870, https://doi.org/10.1007/s12275-014-4390-8 (2014).
Lee, S. H. & Lee, H. E. Cloning and characterization of a multidomain GH10 xylanase from Paenibacillus sp. DG-22. J. Microbiology Biotechnol. 24, 1525–1535, https://doi.org/10.4014/jmb.1407.07077 (2014).
Xu, B. et al. Molecular and biochemical characterization of a novel xylanase from Massilia sp. RBM26 isolated from the feces of Rhinopithecus bieti. J. Microbiology Biotechnol. 26, 9–19, https://doi.org/10.4014/jmb.1504.04021 (2016).
Rosenthal, A. Z., Matson, E. G., Eldar, A. & Leadbetter, J. R. RNA-seq reveals cooperative metabolic interactions between two termite-gut spirochete species in co-culture. ISME J. 5, 1133–1142, https://doi.org/10.1038/ismej.2011.3 (2011).
Manikandan, K. et al. Crystal structures of native and xylosaccharide-bound alkali thermostable xylanase from an alkalophilic Bacillus sp. NG-27: structural insights into alkalophilicity and implications for adaptation to polyextreme conditions. Protein Sci. 15, 1951–1960, https://doi.org/10.1110/ps.062220206 (2006).
Lapidoth, G. et al. Highly active enzymes by automated combinatorial backbone assembly and sequence design. Nat. Commun. 9, 1–9, https://doi.org/10.1038/s41467-018-05205-5 (2018).
Mamo, G., Thunnissen, M., Hatti-Kaul, R. & Mattiasson, B. An alkaline active xylanase: Insights into mechanisms of high pH catalytic adaptation. Biochim. 91, 1187–1196, https://doi.org/10.1016/j.biochi.2009.06.17 (2009).
Ni, J. & Tokuda, G. Lignocellulose-degrading enzymes from termites and their symbiotic microbiota. Biotechnol. Adv. 31, 838–850, https://doi.org/10.1016/j.biotechadv.2013.04.005 (2013).
Scharf, M. E. & Tartar, A. Termite digestomes as sources for novel lignocellulases. Biofuels Bioprod. Bioref. 2, 540–552, https://doi.org/10.1002/bbb.107 (2008).
Dietrich, C., Köhler, T., Brune, A., Kohler, T. & Brune, A. The cockroach origin of the termite gut microbiota: patterns in bacterial community structure reflect major evolutionary events. Appl. Environ. Microbiology 80, 2261–2269, https://doi.org/10.1128/AEM.04206-13 (2014).
Hongoh, Y. et al. Phylogenetic diversity, localization, and cell morphologies of members of the candidate phylum TG3 and a subphylum in the phylum Fibrobacteres, recently discovered bacterial groups dominant in termite guts. Appl. And. Environ. Microbiology 72, 6780–6788, https://doi.org/10.1128/AEM.00891-06 (2006).
Hongoh, Y. et al. Intra- and interspecific comparisons of bacterial diversity and community structure support coevolution of gut microbiota and termite host. Appl. Env. Microbiol. 71, 6590–6599, https://doi.org/10.1128/AEM.71.11.6590-6599 (2005).
Su, L. et al. Comparative Gut Microbiomes of Four Species Representing the Higher and the Lower Termites. J. Insect Sci. 16, 1–9, https://doi.org/10.1093/jisesa/iew081 (2016).
Eutick, M. L., Veivers, P., O’Brien, R. W. & Slator, M. Dependence of the higher termite Nasutitermes exitiosus and the lower termite, Coptotermes lacteus on their gut flora. J. Insect Physiol. 24, 363–368 (1978).
Iida, T., Ohkuma, M., Ohtoko, K. & Kudo, T. Symbiotic spirochetes in the termite hindgut: phylogenetic identification of ectosymbiotic spirochetes of oxymonad protists. FEMS Microbiology Ecol. 34, 17–26, https://doi.org/10.1111/j.1574-6941.2000.tb00750.x (2000).
Breznak, N. A. Phylogenetic Diversity and Physiology of Termite Gut Spirochetes. Integr. Comp. Biol. 42, 313–318, https://doi.org/10.1093/icb/42.2.313 (2002).
Lilburn, T. G. et al. Nitrogen fixation by symbiotic and free-living Spirochetes. Sci. 292, 2495–2498, https://doi.org/10.1126/science.1060281 (2001).
Yamada, A., Inoue, T., Noda, Y., Hongoh, H. & Ohkuma, M. Evolutionary trend of phylogenetic diversity of nitrogen fixation genes in the gut community of wood-feeding termites. Mol. Ecol. 16, 3768–3777, https://doi.org/10.1111/j.1365-294X.2007.03326.x (2007).
Graber, J. R., Leadbetter, J. R. & Breznak, J. A. Description of Treponema azotonutricium sp. nov. and Treponema primitia sp. nov., the First Spirochetes Isolated from Termite Guts. Applied. Environ. Microbiology 70, 1315–1320, https://doi.org/10.1128/AEM.70.3.1315-1320 (2004).
Droöge, S., Reinhard, R., Radek, R. & König, H. Treponema isoptericolens sp. nov. a Nov. spirochaete hindgut termite Incisitermes tabogae Int. J. Syst. Evolut. Microbiology 58, 1079–1083, https://doi.org/10.1099/ijs.0.64699-0 (2008).
Benjamino, J. & Graf, J. Characterization of the Core and Caste-Specific Microbiota in the Termite, Reticulitermes flavipes. Front. Microbiology 7, 171, https://doi.org/10.3389/fmicb.2016.00171 (2016).
Graber, J. R. & Breznak, J. A. Physiology and nutrition of Treponema primitia, an H2/CO2-acetogenic spirochete from termite hindguts. Appl. Environ. Microbiology 70, 1307–1314, https://doi.org/10.1128/AEM.70.3.1307-1314.2004 (2004).
Chandrasekharaiah, M. et al. Molecular cloning, expression and characterization of a novel feruloyl esterase enzyme from the symbionts of termite (Coptotermes formosanus) gut. BMB Rep. 44, 52–57, https://doi.org/10.5483/BMBRep.2011.44.1.52 (2011).
Rashamuse, K. et al. Metagenomic mining of feruloyl esterases from termite enteric flora. Appl. Microbiol. Biotechnol. 98, 727–737, https://doi.org/10.1007/s00253-013-4909-7 (2014).
Gilbert, H. J. The biochemistry and structural biology of plant cell wall deconstruction. Plant. Physiol. 153, 444–455, https://doi.org/10.1104/pp.110.156646 (2010).
Nguyen, S. T. C., Freund, H. L., Kasanjian, J. & Berlemont, R. Function, distribution, and annotation of characterized cellulases, xylanases, and chitinases from CAZy. Appl. Microbiology Biotechnol. 102, 1629–1637, https://doi.org/10.1007/s00253-018-8778-y (2018).
Busk, P. K., Mette, L., Pilgaard, B. & Lange, L. Several Genes Encoding Enzymes with the Same Activity Are Necessary for Aerobic Fungal Degradation of Cellulose in Nature. PLoS One 9, e114138, https://doi.org/10.1371/journal.pone.0114138 (2014).
Scheller, H. V. & Ulvskov, P. Hemicelluloses. Annu. Rev. Plant. Biol. 61, 263–289, https://doi.org/10.1146/annurev-arplant-042809-112315 (2010).
Vogel, J. Unique aspects of the grass cell wall. Curr. Opin. Plant. Biol. 11, 301–307, https://doi.org/10.1016/j.pbi.2008.03.002 (2008).
Aspeborg, H., Coutinho, P. M., Wang, Y., Brumer, H. & Henrissat, B. Evolution, substrate specificity and subfamily classification of glycoside hydrolase family 5 (GH5). BMC Evol. Biol. 12, 1 (2012).
AL-Darkazali, H., Meevootisom, V., Isarangkul, D. & Wiyakrutta, S. Gene Expression and Molecular Characterization of a Xylanase from Chicken Cecum Metagenome. International Journal of Microbiology, 1–12, https://doi.org/10.1155/2017/4018398 (2017).
Chu, Y. et al. Insights into the roles of non-catalytic residues in the active site of a GH10 xylanase with activity on cellulose. J. Biol. Chem. 292, 19315–19327, https://doi.org/10.1074/jbc.M117.807768 (2017).
Jeong, Y. S. et al. Characterization of Xyn10J, a novel family 10 xylanase froma compost metagenomic library. Appl. Biochem. Biotechnol. 166, 1328–1339, https://doi.org/10.1007/s12010-011-9520-8 (2012).
Kim, D. Y. et al. Catalytic properties of a GH10 endo-β-1, 4- xylanase from Streptomyces thermocarboxydus HY-15 isolated from the gut of Eisenia fetida. J. Mol. Catal. B: Enzym. 62, 32–39, https://doi.org/10.1016/j.molcatb.2009.08.015 (2010).
Zhou, J. et al. Molecular and biochemical characterization of a novel xylanase from the symbiotic Sphingobacterium sp. TN19. Appl. Microbiol. Biotechnol. 85, 323–333, https://doi.org/10.1007/s00253-009-2081-x (2009).
Xue, X. et al. The N-Terminal GH10 Domain of a Multimodular Protein from Caldicellulosiruptor bescii Is a Versatile Xylanase/β-Glucanase That Can Degrade Crystalline Cellulose. Appl. Environ. Microbiology 81, 3823–3833, https://doi.org/10.1128/AEM.00432-15 (2015).
Acknowledgements
PT, JA and EC acknowledge CONICET as career research members. MRV, CE and OO acknowledge the CONICET Fellowships. The authors are grateful to Dr. Julia Sabio y García for linguistic improvement in the manuscript. This work was supported by grants from the Instituto Nacional de Tecnología Agropecuaria (INTA) (PNAIyAV-1130034), Agencia Nacional de Promoción Científica y Tecnológica (ANPCyT) Proyectos de Investigación Científica y Tecnológica (PICT) 2013 No.1454 (Argentina) and 2018 No. 4149. Part of this work was performed at the Lawrence Berkeley National Laboratory under the Department of Energy contract no. DE-AC02- 05CH11231. RABG appreciates the support of Conacyt, Project CB285816. LM y JQ acknowledge the Conacyt Fellowships.
Author information
Authors and Affiliations
Contributions
M.R.V. conceived the experiments, analysed the results and contributed to manuscript writing. M.A.S. analysed the results and contributed to manuscript writing. R.A.B.G. conceived and analysed the protein modelling results and contributed to manuscript writing. J.A.C.N. conceived the experiments and contributed to manuscript writing. S.V., M.O., O.O. and S.G. conceived the experiments. L.M.A. and O.J.Q.G. analysed the protein modelling results. C.E. collected the termites. E.C. analysed the results. D.C. contributed to manuscript writing. J.A. analysed the results and contributed to manuscript writing. P.M.T. conceived and conducted the experiments, analysed the results and was responsible for writing the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Romero Victorica, M., Soria, M.A., Batista-García, R.A. et al. Neotropical termite microbiomes as sources of novel plant cell wall degrading enzymes. Sci Rep 10, 3864 (2020). https://doi.org/10.1038/s41598-020-60850-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-60850-5
This article is cited by
-
Metagenomic analysis of gut microbiome illuminates the mechanisms and evolution of lignocellulose degradation in mangrove herbivorous crabs
BMC Microbiology (2024)
-
Analysis of gut microbiota of ladybug beetle (Harmonia axyridis) after feeding on different artificial diets
BMC Microbiology (2024)
-
Heterologous expression and structure prediction of a xylanase identified from a compost metagenomic library
Applied Microbiology and Biotechnology (2024)
-
Bacterial diversity from soil-feeding termite gut and their potential application
Annals of Microbiology (2023)
-
Production, purification, characterization and application of two novel endoglucanases from buffalo rumen metagenome
Journal of Animal Science and Biotechnology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.