Abstract
The DNA molecule is the most evolved and most complex molecule created by nature. The primary role of DNA in medicine is long-term storage of genetic information. Genetic modifying is one of the most critical challenges that scientists face. On the other hand, it is said that under the influence of acoustic, electromagnetic, and scalar waves, the genetic code of DNA can be read or rewritten. In this article, the most accurate and comprehensive dynamic model will be presented for DNA. Each of the two strands is modeled with an out of plane curved beam and then by doubling this two strands with springs, consider the hydrogen bond strength between this two strands. Beams are traditionally descriptions of mechanical engineering structural elements or building. However, any structure such as automotive automobile frames, aircraft components, machine frames, and other mechanical or structural systems contain beam structures that are designed to carry lateral loads are analyzed similarly. Also, in this model, the mass of the nucleobases in the DNA structure, the effects of the fluid surrounding the DNA (nucleoplasm) and the effects of temperature changes are also considered. Finally, by deriving governing equations from Hamilton’s principle method and solving these equations with the generalized differential quadrature method (GDQM), the frequency and mode shape of the DNA is obtained for the first time. In the end, validation of the obtained results from solving the governing equations of mathematical model compared to the obtained results from the COMSOL software is confirmed. By the help of these results, a conceptual idea for controlling cancer with using the DNA resonance frequency is presented. This idea will be presented to stop the cancerous cell’s protein synthesis and modifying DNA sequence and genetic manipulation of the cell. On the other hand, by the presented DNA model and by obtaining DNA frequency, experimental studies of the effects of waves on DNA such as phantom effect or DNA teleportation can also be studied scientifically and precisely.
Similar content being viewed by others
Introduction
Deoxyribonucleic acid, more commonly known as DNA, is an evolved molecule that contains the genetic code of organisms. Every living thing has DNA within their cells. It is important for inheritance, coding for proteins and the genetic instruction guide for life and its processes. DNA holds the instructions for an organism or each cell development processes, reproduction and ultimately death. Over the past decades, empirical discussions have been proposed to modify the genes in the DNA. These changes have been much discussed in the medical field by drawing on applications such as treatment, preventing the development of cancer, or erupting an organ (for example, a tooth). On the other hand, it should be noted that only 3% of DNA capacity is considered in medical fields. In the last two decades, a topic called “wave genome” has been raised by Russian scientists, which states that 97% of other DNA is not only inapplicable but also has a more significant role; because DNA can be affected by acoustic, electromagnetic, and scalar waves. Under the influence of these waves, the genetic code can be read or rewritten. Another claim of Russian scientists is that DNA is a biological network that binds all humans. About impressionability of DNA from the wave frequency, many experimental research studies have been carried out which have opened a new branch in science, called wave genome. Konstantin Meyl adapted the scalar waves described by Nicola Tesla to biology and proposed the relationship between the scalar waves and DNA1. Greg Braddon and colleagues in 3 experiments investigated the impressionability of DNA from human emotions2. Rein and Mccraty studied the impact of music on the DNA3,4,5. Another study was carried out on the effect of sound waves on the synthesis and genes of chrysanthemum6. Peter Garjajev and his research group proved that DNA can be reprogrammed by words and using the correct resonant frequencies of DNA7,8. Russian quantum biologist Poponin tried to prove that human DNA has a direct effect on the physical world using some experiments9. Also, he found out that our DNA can cause disturbing patterns in the vacuum, thus producing magnetized microscopic wormholes10. Nobel Prize-winning scientist Luc Montagnier known for his study on HIV and AIDS, claims to have demonstrated that DNA can be generated by teleportation through quantum imprint and also showed that DNA emits electromagnetic signals that teleport the DNA to other places, such as water molecules11,12.
About Mathematical Modeling of DNA, three eminent mathematical models have been proposed for describing DNA. The most famous model presented is the PBD (Peyrard-Bishop-Dauxois) model13, although this model has been updated by various researchers14,15,16,17,18, the models presented in these studies usually have at least three significant weaknesses such as being discrete, not being outside the plane, not being spiral, and not considering the position of nucleobases. Examples of the PBD model are displayed in Fig. 1. The other model is a rod model looking at DNA on a larger scale19,20,21, with significant weaknesses, including the lack of attention to the nucleobase positions and the hydrogen bond, as well as considering DNA with one strand (Fig. 2). There is another model called SIDD (stress-induced DNA duplex destabilization) that is entirely mathematical and applied in the field of molecular dynamics22,23. These models mentioned were mostly designed to investigate the vibration of DNA, and there are several models available in other fields such as DNA’s entropic elasticity24 and bending of a DNA25,26. A beam is a structural element that primarily resists loads applied laterally to the beam axis. Its mode of deflection is primarily by bending. The Timoshenko beam theory takes the shear deformation and rotational bending effects into consideration for describing the behavior of thick beams. On the other hand, all previous studies related to curved beam vibrations focus on out of plane vibration of curved beams (with inline coordinates) and do not study out of plane vibration of the curved beams. A.Y.T. Leung is the only reference derived from the governing equations for a helical beam with rectangular cross-sections with pre-twist27.
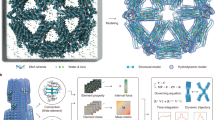
According to the above contents based on the necessity of DNA vibration analysis and the weaknesses of the previously proposed models, the necessity of carrying out this research becomes more evident. The dynamic model presented for DNA here has been named GMDM (Ghadiri Marvi DNA Model), it is provided by connecting two out of plane nano curved beams with spring and a damper. Each of the beams is a model for one of the two DNA strands (sugar-phosphate backbone). Also, spring and damper is a model for hydrogen bonds between nitrogen-containing nucleobases. The effects of the nucleobases (cytosine, guanine, adenine, and thymine) are also considered with their mass (Fig. 3). Also, the effects of DNA surrounding fluid (nucleoplasm) have been applied by using the Navier Stokes equations. Effects of temperature change on DNA are also applied to equations with external work. Finally, by using the relations of all the effects that mentioned above and using the Hamilton principle, the DNA equations will be extracted.
(a) Imagined shape for DNA43 © 2013 Nature Education Adapted from Pray, L. (2008) Discovery of DNA structure and function: Watson and Crick. Nature Education 1(1):100. All rights reserved. (b) The GMDM Mathematical model, which is presented in this paper for dynamics investigations of DNA. This image has been modeled with COMSOL Multiphysics 5.3a and edited with Adobe Photoshop CC 2018.
It should be noted that with the help of the theory of nonlocal, the effects of size were considered. By solving these equations, DNA natural frequency will be obtained for the first time. Numerical method can solve the equations derived from Hamilton’s principle method. The generalized differential quadrature method (GDQM) is one of the most numerical methods can be used for solving governing differential equations.
The idea brought up in this study is that if DNA is influenced by wave frequencies as much as DNA natural frequency then resonance occurs and DNA vibrates with large amplitude oscillations. With many shakes, DNA strands go up and down and at this moment a nucleobase in the nucleotide of one of the DNA strands establishes a hydrogen bond with the nucleobases lower or upper and the nucleobase aligns itself on the other DNA strands. This mechanism and idea which is presented schematically in Fig. 4, can cause disorganization in a sequence of DNA, and finally, with the help of a restriction enzyme (endonuclease), DNA in the cancerous cell loss of the ability to biosynthesis of proteins and the cancer is controlled just like the CRISPR/CAS9 technology.
(a) DNA’s shape before reaching resonant frequency. (b) DNA’s shape after reaching resonant frequency. (c) Establishes a hydrogen bond between the nucleobases upper with nucleobases lower. (d) Removal of additional and remaining nucleotides with the help of restriction enzyme. (e) New DNA’s shape and disorganization in a sequence of DNA. These images have been modeled with COMSOL Multiphysics 5.3a and edited with Adobe Photoshop CC 2018.
Nonlocal Elasticity Theory
In accordance with the nonlocal elasticity theory, the stress state at a reference point in an elastic continuum depends not only on the strain components at the same position but also a function of strains of all points in the neighbor regions. Therefore, the nonlocal stress tensor’s at point x is expressed as:
where T(x) is the classical, macroscopic stress tensor at point x related to strain by Hooke’s law with Eq. (1). C(x) is the fourth-order elasticity tensor, ε(x) is the strain tensor and K(|x′ − x|, τ) denotes nonlocal modulus. |x′ − x| represents the distance and τ is a material constant that depends on the internal and external characteristic length defined as \(\tau =\frac{{e}_{0}a}{L}\) where e0 is a constant appropriate to each material, a is an internal characteristics length (e.g., bonds length) and L is an external characteristics length (e.g., wavelength).
Solving the integral constitutive relation in Eq. (1) is difficult. Thus, equivalent relation in a differential form was proposed by Eringen28,29 as follows:
∇2 is the Laplacian operator
Beam theory and displacement of a double helical nanobeam
An out of plane curved beam is the beam having a twist and curvature around its central axis. The geometry situation of this model needs to choose the coordinate system that every moment changes its vector location (Fig. 5). Thus, for the analysis of the out of plane curved beam, Frenet triad must be used.
The base beam theory used to model an out of plane curved beam is Timoshenko beam theory. By following Timoshenko’s assumption, the displacements u are defined as consisting of two parts, part one is the displacements at the centerline along the local axes v and part 2 is the rotations of the cross-section θ. This two parts according to Timoshenko’s assumption, only defined at the x3 axis.
In Eq. (3) r is the vector of the main displacement functions.
In Fig. 6, a small cut of a cross section of a beam has been displayed.
To find the strain vector of out of plane curved beams must be differentiated the displacement concerning the arc length x3.
With using the Frenet triad and its special differential can be derived the displacement vector gradient of an out of plane curved beam as follows27:
Parameters \(\tau ,\kappa ,\mu \) used in the matrix Eq. (4) respectively represent tortuosity, curvature and pre-twist rate, in which, \(C=\,\cos (\mu s)\), \(S=\,\sin (\mu s)\) and \(\mu s\) is twist per length.
The non-vanishing strain matrix based on the Timoshenko’s assumption are derived as:
For simplicity in the use of nonlocal theory, strain matrix is separated into three matrices:
In which:
Now considering the nonlocal Eringen equation and replacing the strain term with the Eq. (6).
By defining axial force to form \(N=\int \sigma \,dA\) and bended moment as \(M=\int x\sigma \,dA\), we will have:
Where \({I}_{1}=\int {{x}_{2}}^{2}dA\) and \({I}_{2}=\int {{x}_{1}}^{2}dA\) and \(\{{M}_{1},{M}_{2}\}=\int \{{x}_{1},{x}_{2}\}\sigma dA\)
For convenience, Eqs. (8–10) are brought into a matrix.
The GMDM model consists of two out of plane curved beams that are used to model a DNA. These two beams are connected with springs and dampers. By considering the mass of the nucleobases (Fig. 3b) and then by writing the strain energy and kinetic energy equations of the components, and the use of Navier-Stokes equations to apply the effects of a nucleoplasm, the effects of the temperature increase as an external work, and putting all of these equations in the Hamilton equation, the governing equation for the DNA model will be derived.
To derive the governing equations in the first step, we need to find the strain energy. Strain energy for two beams and also damper and spring connecting two beams will be as follows.
where:
Finally, the strain energy of all the components is brought together and the strain energy of the whole system is obtained as
For simplicity, Eqs. (12) and (13) can be demonstrated in the form of a matrix as the following:
It should be noted that in order to prepare the conditions for using variational method form, the matrix \({[\begin{array}{c}{e}_{0}\\ {e}_{1}\\ {e}_{2}\end{array}]}_{9\ast 12}\) was separated and rearranged to this matrix \({[\begin{array}{cc}{e}_{01} & {e}_{02}\\ {e}_{11} & {e}_{12}\\ {e}_{21} & {e}_{22}\end{array}]}_{9\ast (6+6)}\), in which:
The kinetic energy of two beams and nucleobases with vibration frequency \(\omega \) is given by:
where \(\rho \) is the density of strands of DNA and:
It should be noted that in Fig. 3 and GMDM model, the mass of nucleobases have been considered. Adenine always provides hydrogen bond with Thymine and also Guanine always makes hydrogen bond with Cytosine. Therefore, by the averages of weight of Adenine and Thymine, also weight of Guanine and Cytosine, attach this average mass to the both beams. Depending on the genetic sequence in the DNA code, it is determined that which one of the nucleobases kinetic energy (\({{\bf{T}}}_{1AT},\,{{\bf{T}}}_{1CG},\,{{\bf{T}}}_{2AT},\,{{\bf{T}}}_{2CG}\)) should be used.
The effects of the nucleoplasm on the vibration of DNA are also considered as an external work, which the Navier Stokes equations are also used to apply this effect. The effects of the nucleoplasm for one beam and for two directions \({x}_{1}\) and \({x}_{2}\) that are perpendicular to the cross-section are shown below:
In which ρn, P and μv are density of nucleoplasm, intracellular pressure and viscosity of nucleoplasm respectively.
The effects of fluid are considered as an external work.
In which:
Regardless of gravity and nonlinear term, the effects of fluid on DNA strands will be as follows:
where \({W}_{1{S}_{1}}\) represents the external work applied to number 1 strand in \({x}_{2}\) directions and \({W}_{2{S}_{1}}\) represents the external work applied to number 2 strand in \({x}_{1}\) directions. Also, the effects of temperature increase on DNA as linear in term of thickness of the beam will also be applied as an external work as below (since strands are larger than nucleobases, we neglected the effect of temperature on nucleobases).
In which:
By replacing the Eq. (24) in to the Eq. (23), we will have:
The Eq. (25) can be written as follows:
In which:
By substituting the strain energy (Eq. (16)) kinetic energy (Eq. (19)) external works (Eqs. (21, 26)) and also using variation method and put them in Hamilton relation, we will have:
By merging Eq. (11) in the Eq. (27), the governing equations of DNA with considering the effects of the fluid and temperature change will be as:
This model is the most comprehensive model to investigate the dynamic behavior of DNA considering mass of nucleobases, their hydrogen bond and effect of the surrounding fluid of DNA, is extracted to 12 PDE equations.
Also boundary condition equation of this system is as below:
The special boundary conditions that was used in are defined as clamped-clamped (at x = 0 & x = L), and simply supported (at x = 0 & x = L) as follows:
Clamped:
Simply supported:
Solution with using GDQM
Different numerical techniques can be used to solve the governing equations and related boundary conditions. In this paper the GDQ method that is introduced by Shu30 is used. This technique has been successfully employed to solve a variety of problems in vibration analysis and dynamical systems. The GDQM is a powerful method that can be used to solve partial differential equations extended and generalized with high accuracy and convergence and performance.
According to GDQM the \(r-th\) order derivative of function of \(f(r)\) with respect to x at \({x}_{i}\) is:
Where N is the number of total grid points and C(n) is the respective weighting coefficients matrix that can be obtained as follows
where M(x) is defined as below:
The weighting coefficients for \(r-th\) order are derived from:
The Chebyshev-Gauss-Lobatto technique can be used for the distribution of grid points in domain:
By writing the governing equations as a matrix form:
whereas M is the mass matrix and K is stiffness matrix
With coupling this six matrix equations and using computer software, natural frequency is found.
Geometry and mechanical properties
Dimensions and mechanical properties of DNA are depicted in Tables 1 and 2 according to various scientific references.
One of the challenges ahead of the present study was to obtain a DNA cross section. This work was carried out in a way that with an average weighted gain from the diameter of the atoms participating in the phosphate and sugar structure31,32, the thickness of the DNA strands was measured. In addition, by using the length of the atomic bonds in the sugar and phosphate structure33, and with the help of the angles between bonds34, the width of the DNA backbone was obtained. The results of these calculations are presented in the Table 3.
The sectional average radius of nucleobases is also obtained by an average weighted gain from the diameter of the atoms participating in the nucleobases structure. The sectional radius of nucleobases are shown in Table 4.
The second challenge ahead of this study was to obtain a suitable nonlocal parameter for DNA. Parameter \({e}_{0}\) is directly related to the aspect ratio of geometry. Depending on the ratio of the length of the DNA to its thickness, and by extrapolating the results presented by Ghavanloo35, \({e}_{0}\) is obtained. Also Parameter a depends on the length of atomic bonds in the target objects, therefore, with an average of length of all atomic bonds in a nucleotide, a will also be obtained. These two parameters have been brought in the Table 5. It should be noted that the calculated value for e0a, applies in the general relation presented for the nonlocal parameter36,37. This relationship is in this form: \(0 < \frac{{e}_{0}a}{L} < 0.8\).
Nuclear DNA does not appear in free linear strands. DNA is wrapped around histones in order to fit within the nucleus and participate in the chromatin structure. According to the cause, the supports in this study are assumed clamped-clamped.
Convergence
The number of grid points affects the natural frequency of DNA. In Table 6 evaluating convergence have been proved with respect to number of elements for the first three frequencies of DNA.
It is observed that for 20 node later, the answers will converge.
Study of validation
Using simulations in COMSOL software, we obtained the natural frequency of DNA. Then the natural frequency obtained using mathematical modeling is compared with COMSOL simulations in Table 7.
In short articles, DNA natural frequency is mentioned approximately with simulations (Table 8).
It should be noted that the primary modes of frequency of DNA obtained in the present study, lie within the range considered in those studies.
Results due to changing parameters
The derived equations, are solved with the mathematical model to find the natural frequencies and mode shapes for DNA and a parametric study is performed on the frequency of the DNA.
The effect of the DNA length on frequency
As mentioned, the length of the DNA varies between the two histones and has 10 and 80 nucleobases. These results which are presented in Table 9 show that with increasing number of nucleobases, the natural frequency decreases. The reason for this change is that by increasing the length of the body, it has more mobility and less rigidity.
The effect of supporting conditions on frequency
It can be seen that clamped-clamped supporting, has the highest frequency (Table 10) because by increasing constraint in supporting (reducing degrees of freedom), rigidity and frequency increase.
The effect of DNA embedding fluid on frequency
As we know, chromatins and DNA are floating in a fluid called the nucleoplasm. In Table 11, the effects of the presence and absence of this fluid on DNA frequency, have been shown. It is observed that due to the low density of the nucleoplasm, the DNA frequency decreases slightly in the presence of this fluid. The reason for this topic can be found in the absorption of vibrating energy of DNA by the fluid.
The effects of temperature rise on frequency
Today, the huge effects of temperature increase on the stability of DNA strands, is proven. As DNA close to the melting temperature, displacement of the base pairs increases sharply and hydrogen bound becomes unbound13,38. DNA persistence length and internal base pair conformations, strongly depends on temperature39,40. Increasing temperature enhances flexibility of the double-stranded DNA chain41. As it is said in 354 K, the DNA denatures42. In Table 12 the effects of temperature rise on the frequency of DNA, was investigated. It is observed that the DNA frequency decreases with increasing temperature. The reason for this event is that the DNA rigidity is reduced by temperature rise.
It is also observed that at 44 °C increase in temperature (reaching 354 K), the DNA natural frequency, drops sharply.
Mode shape of DNA in COMSOL
As mentioned earlier the results were verified with the help of the DNA modeling in COMSOL. In this software, both strands of DNA are modeled with two spiral geometries and the nucleotides are also designed to be cylinders with different radius and height according to their dimensions. It should be noted that the distance between the nucleobases is taken as the length of the hydrogen bond. The nucleotides are arbitrarily arranged in sequence ATCGTAGCATCGTAGCATCG. Hydrogen bonds also have been modeled as springs. Since the DNA strands are inside the nucleobases and in order to apply the properties and forces of fluid in the COMSOL, we used a sphere larger than DNA in which DNA is enclosed by fluid (Fig. 7).
In Fig. 8, the mode shape of the DNA vibrations, obtained in COMSOL software are presented.
Conclusion
The presented study on DNA is the most accurate dynamic model that was named GMDM. This model is provided by connecting two out of plane nano curved beams with spring and a damper. Each of the beams is a model for one of the two DNA strands, also spring and damper is a model for hydrogen bonds between nitrogen-containing nucleobases. DNA vibrations and DNA natural frequency is investigated for the first time with considering thermal, mass of nucleobases and fluid effect on GMDM. These effects were applied using the equations like Navier Stokes in to the Hamilton principle. With solving these equations by GDQM, DNA natural frequency will be obtained for the first time. The novelty of this model are as follows:
-
Being out of plane, spiral with twist and curvature
-
Being continuous model
-
Considering the effects of the mass and viscoelastic of the nucleoplasm
-
Considering the effects of temperature rise
-
Considering the position of the nucleobases.
As the energy absorption by the nucleoplasm is very insignificant, it can be concluded that during the resonance, the DNA amplitude oscillations range is very large and with severe shakes and using a restriction enzyme, sequence of DNA might be disorganized, and DNA in cancerous cells loses its ability for proteinization and consequently in this way cancer may be controlled.
Change history
31 August 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41598-021-97512-z
04 July 2022
This article has been retracted. Please see the Retraction Notice for more detail: https://doi.org/10.1038/s41598-022-15829-9
References
Konstantin meyl, DNA and Cell Resonanse, 2nd Ed. Villingen-Schwenningen: INDEL GmbH (2011).
Braden, G. L’effetto Isaia: decodificare la scienza perduta della preghiera e della profezia. Macro Edizioni (2001).
Rein, G. Effect of conscious intention on human DNA. Proc. Internat. Forum New Sci. Denver, CO, pp. 1–12 (1996).
Rein, G. The In Vitro Effect of Bioenergy on The Conformational States of Human Dma in Aqueous Solutions. Acupunct. Electrother. Res., 20(3), 173–180 (Jan. 1995).
Rein, G., Ph, D. & Mccraty, R. Local and Non-Local Effects of Coherent Heart Frequencies on Conformational Changes of Dna. Proc. Jt. USPA/IAPR Psychotronics Conf., no. 1, 1–6 (1993).
Xiujuan, W., Bochu, W., Yi, J., Chuanren, D. & Sakanishi, A. Effect of sound wave on the synthesis of nucleic acid and protein in chrysanthemum. Colloids Surfaces B Biointerfaces 29(2), 99–102 (2003).
Gariaev, P. P., Marcer, P. J., Leonova-gariaeva, K. A. & Kaempf, U. DNA as Basis for Quantum Biocomputer. DNA Decipher 1(1), 25–46 (2011).
Garjajev, P. et al. The DNA-wave biocomputer. fourth Int. Conf. Comput. Anticip. Syst. 10, 290–310 (2000).
Poponin, V. The DNA phantom effect: direct measurement of a new field in the vacuum substructure. In Ann. Conf. on Treatment and Res. Experienced Anomalous Trauma, 1995.
Pitk, M. Wormhole Magnetic Fields, In Quantum Hardware of Living Matter, 2006. [Online]. Available, http://tgdtheory.com/public_html/bioware/bioware.html#wormc.
Montagnier, L. et al. Transduction of DNA information through water and electromagnetic waves. Electromagn. Biol. Med. 8378(2), 10 (2014).
Montagnier, L. et al. DNA waves and water. J. Phys. Conf. Ser. 306(1), 012007 (2011).
Peyrard, M. & Bishop, A. R. Statistical mechanics of a nonlinear model for DNA denaturation. Phys. Rev. Lett. 62(23), 2755–2758 (1989).
Zdravković, S. Helicoidal Peyrard–Bishop Model of Dna Dynamics. J. Nonlinear Math. Phys. 18(sup2), 463–484 (2011).
Silva, R. A. S., Filho, E. D. & Ruggiero, J. R. A model coupling vibrational and rotational motion for the DNA molecule. J. Biol. Phys. 34(5), 511–519 (2008).
Zdravković, S., Tuszyński, J. A. & Satarić, M. V. Peyrard-Bishop-Dauxois model of DNA dynamics and impact of viscosity. J. Comput. Theor. Nanosci. 2(2), 263–271 (2005).
Farzadian, O. & Niry, M. D. Delocalization of mechanical waves in the ladder chain of DNA with correlated disorder. Phys. A Stat. Mech. its Appl. 450, 95–103 (2016).
Páez, C. J., Rey-González, R. & Schulz, P. A. Macroscopic localization lengths of vibrational normal modes in a heuristic DNA model. Phys. Rev. B - Condens. Matter Mater. Phys. 81(2), 1–5 (2010).
Hirsh, T. D. Lillian, Lionberger, Taranova, Andricioaei, and Perkins, A Model for Highly Strained DNA Compressed Inside a Protein Cavity A Model for Highly Strained DNA Compressed Inside a Protein Cavity. J. Comput. Nonlinear Dyn 8, 031001–8 (2013).
Lillian, T. D., Taranova, M., Wereszczynski, J., Andricioaei, I. & Perkins, N. C. A multiscale dynamic model of DNA supercoil relaxation by topoisomerase IB. Biophys. J. 100(8), 2016–2023 (2011).
Goyal, S. A DYNAMIC ROD MODEL TO SIMULATE MECHANICS OF CABLES AND DNA, PhD Dissertation in Mechanical Engineering. The University of Michigan, Ann Arbor, Michigan, 2006.
Benham, C. J. Destabilization in Long Genomic DNA Sequences. J. Comput. Biol. 11(4), 519–543 (2004).
Sershen, C. L. A Dynamic Model of DNA Structure and Function, University of California, Davis, 2009.
Haijun, Z., Yang, Z. & Zhong-Can, O. Y. Bending and base-stacking interactions in double-stranded DNA. Phys. Rev. Lett. 82(22), 4560–4563 (1999).
Xu, X., Thio, J. R. & Cao, J. Correlated Local Bending of a DNA Double Helix and Its Effect on DNA Flexibility in the Sub-Persistence-Length Regime. J. Phys. Chem. Lett. 5(16), 2868–2873 (2014).
Yan, J. & Marko, J. F. Localized single-stranded bubble mechanism for cyclization of short double helix DNA. Phys. Rev. Lett. 93(10), 3–6 (2004).
Leung, A. Y. T. Vibration of thin pre-twisted helical beams. Int. J. Solids Struct. 47(9), 1177–1195 (2010).
Eringen, A. C. & Edelen, D. G. B. On nonlocal. Int. J. Engng. Sci. 10, 233–248 (1972).
Eringen, A. C. Nonlocal Continuum Field Theories. Springer Science & Business Media, 2001.
Shu, C. Differential Quadrature and Its Application in Engineering. Springer Science & Business Media, 2000.
Clementi, E. Atomic Screening Constants from SCF Functions. II. Atoms with 37 to 86 Electrons. J. Chem. Phys. 47(4), 1300 (1967).
Slater, J. C. Atomic radii in crystals. J. Chem. Phys. 41(10), 3199–3204 (1964).
Silberberg, M. S. & Amateis, P. Chemistry: The Molecular Nature of Matter and Change with Advanced Topics 8th Ed. McGraw-Hill, 2018.
Lu, X.-J. Torsion angles of Nucleic Acid Structures. NIH, 2010. Available, http://x3dna.org/highlights/torsion-angles-of-nucleic-acid-structures.
Ghavanloo, E. & Fazelzadeh, S. A. Evaluation of nonlocal parameter for single-walled carbon nanotubes with arbitrary chirality. Meccanica 51(1), 41–54 (2016).
Lu, P., Lee, H. P., Lu, C. & Zhang, P. Q. Dynamic properties of flexural beams using a nonlocal elasticity model. J. Appl. Phys. 99, 7 (2006).
Wang, C. M., Zhang, Y. Y. & He, X. Q. Vibration of nonlocal Timoshenko beams. Nanotechnology 18, 10 (2007).
li Zhang, Y., Zheng, W. M., Liu, J. X. & Chen, Y. Z. Theory of DNA melting based on the Peyrard-Bishop model. Phys. Rev. E 56(6), 7100–7115 (1997).
Geggier, S., Kotlyar, A. & Vologodskii, A. Temperature dependence of DNA persistence length. Nucleic Acids Res. 39(4), 1419–1426 (2011).
Meyer, S. et al. Temperature dependence of the dna double helix at the nanoscale: Structure, elasticity, and fluctuations. Biophys. J. 105(8), 1904–1914 (2013).
Driessen, R. P. C. et al. Effect of temperature on the intrinsic flexibility of DNA and its interaction with architectural proteins. Biochemistry 53(41), 6430–6438 (2014).
Macedo, D. X., Guedes, I. & Albuquerque, E. L. Thermal properties of a DNA denaturation with solvent interaction. Phys. A Stat. Mech. its Appl. 404, 234–241 (2014).
Pray, L. A. Discovery of DNA Structure and Function: Watson and Crick. Nat. Educ. 1(1), 100 (2008).
Cocco, S., Marko, J. F. & Monasson, R. Theoretical models for single-molecule DNA and RNA experiments: From elasticity to unzipping. Comptes Rendus Phys. 3(5), 569–584 (2002).
Gupta, S. K., McEwan, A. & Lukačević, I. Elasticity of DNA nanowires. Phys. Lett. Sect. A Gen. At. Solid State Phys. 380(1–2), 207–210 (2016).
Sober, H. A. CRC Handbook of Biochemistry: Selected data for molecular biology. Boca Raton, FL: CRC Press, 1973.
Yakushevich, L. V. Nonlinear Physics of DNA, vol. 1. Weinheim, FRG: Wiley-VCH Verlag GmbH & Co. KGaA, 2004.
Sulaiman, A., Zen, F. P., Alatas, H. & Handoko, L. T. Dynamics of DNA breathing in the Peyrard-Bishop model with damping and external force. Phys. D Nonlinear Phenom. 241(19), 1640–1647 (2012).
Higgs, H. N. & Peterson, K. J. Phylogenetic analysis of the formin homology 2 domain. Mol. Biol. Cell 16(1), 1–13 (2005).
Liang, L., Wang, X., Xing, D., Chen, T. & Chen, W. R. Noninvasive determination of cell nucleoplasmic viscosity by fluorescence correlation spectroscopy. J. Biomed. Opt. 14(2), 024013 (2009).
Savin, A. V., Mazo, M. A., Kikot, I. P., Manevitch, L. I. & Onufriev, A. V. Heat conductivity of the DNA double helix. Phys. Rev. B 83(24), 245406 (2011).
Rouzina, I. & Bloomfield, V. A. Heat capacity effects on the melting of DNA. 2. Analysis of nearest- neighbor base pair effects. Biophys. J. 77(6), 3252–3255 (1999).
Watson, J. D. & Crick, F. H. D. A Structure for Deoxyribose Nucleic Acid. Nature 171(4356), 737–738 (1953).
Sinden, R. R., Pearson, C. E., Potaman, V. N. & Ussery, D. W. DNA: STRUCTURE AND FUNCTION, vol. 5. JAI Press, 1998.
Neidle, S. Principles of Nucleic Acid Structure. Elsevier, 2008.
Ikeda, M. et al. Frequency-dependent electrical characteristics of DNA using molecular dynamics simulation. Inst. Phys. Publ. 14, 123–127 (2003).
Adair, R. K. Vibrational Resonances in Biological Systems at Microwave Frequencies. Biophys. J. 82(3), 1147–1152 (2002).
Author information
Authors and Affiliations
Contributions
Mobin Marvi and Majid Ghadiri wrote the main manuscript text and prepared figures. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this Article was revised: The original version of this Article contained a typographical error in Table 9, where the value of the First mode in “Number of Nucleobases”, “30” was incorrect.
This article has been retracted. Please see the retraction notice for more detail:https://doi.org/10.1038/s41598-022-15829-9
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Marvi, M., Ghadiri, M. RETRACTED ARTICLE: A Mathematical Model for Vibration Behavior Analysis of DNA and Using a Resonant Frequency of DNA for Genome Engineering. Sci Rep 10, 3439 (2020). https://doi.org/10.1038/s41598-020-60105-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-60105-3
This article is cited by
-
A study of undulatory and rotational wave motion in the beam for the locomotion of underwater robots
Journal of Marine Science and Technology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.