Abstract
The Chinese cordyceps, a complex of the fungus Ophiocordyceps sinensis and its species-specific host insects, is also called “DongChongXiaCao” in Chinese. Habitat degradation in recent decades and excessive harvesting by humans has intensified its scarcity and increased the prices of natural populations. Some counterfeits are traded as natural Chinese cordyceps for profit, causing confusion in the marketplace. To promote the safe use of Chinese cordyceps and related products, a duplex PCR method for specifically identifying raw Chinese cordyceps and its primary products was successfully established. Chinese cordyceps could be precisely identified by detecting an internal transcribed spacer amplicon from O. sinensis and a cytochrome oxidase c subunit 1 amplicon from the host species, at a limit of detection as low as 32 pg. Eleven commercial samples were purchased and successfully tested to further verify that the developed duplex PCR method could be reliably used to identify Chinese cordyceps. It provides a new simple way to discern true commercial Chinese cordyceps from counterfeits in the marketplace. This is an important step toward achieving an authentication method for this Chinese medicine. The methodology and the developmental strategy can be used to authenticate other traditional Chinese medicinal materials.
Similar content being viewed by others
Introduction
The ascomycetes fungus Ophiocordyceps sinensis (anamorph: Hirsutella sinensis)1, which is synonymous with Cordyceps sinensis in the historical classification system2, colonises and mummifies the larvae of moth species in the family Hepialidae in the winter1,2. Then, during the next late spring or early summer, the multiplied fungi breaks the host’s exoskeleton at the head, forming a parasitic complex that comprises the remains of the caterpillar and a fungal sexual stroma bud3,4,5,6. This whole fruiting-body complex, with the characteristics “winter worm–summer grass”, is well known as Chinese cordyceps7 or “DongChongXiaCao” in Chinese3,7,8,9. It has a long history of being used for health and medicinal purposes in traditional Chinese medicine (TCM) as a tonic and renowned medicinal material7,8,9. Chinese cordyceps contains diverse bioactive ingredients10,11 and is widely used to enhance immunity or treat respiratory, bronchial and lung inflammation1,6,12.
Chinese cordyceps is distributed mainly in alpine regions at high altitudes(>3,000 m) on the Qinghai-Tibetan Plateau in southwestern China13. Owing to its strict host-specificity and confined geographic distribution14,15, the natural production of wild Chinese cordyceps is very limited. Habitat degradation in recent decades and excessive harvesting by humans have intensified its scarcity and increased the prices of natural populations6. Additionally, various counterfeit versions having a similar morphology, such as natural counterfeits Cordyceps hawkesii Gray (C. gunni), Cordyceps. liangschanensis, Cordyceps. militaris, Cordyceps. barnesii, Cordyceps. gracilis, and the root of the Starchys geobombycis plant, as well as artificial mimics made from dough, are traded as natural Chinese cordyceps for profit, thereby causing confusion in marketplace and affecting the safe use of Chinese cordyceps16,17,18,19,20. Furthermore, some fermented mycelial products from O. sinensis or C. militaris have been accidentally or intentionally labelled as “DongChongXiaCao”, which also causes confusion16,18.
The Chinese cordyceps is often used directly as a raw material, but it is also processed into a powder that is used in patented Chinese medicinal and health products, which makes it nearly impossible to identify based on morphological appearance16. To ensure the medicinal and edibility safety of Chinese cordyceps products in the marketplace, many detection methods for monitoring and verifying the Chinese cordyceps ingredients have been developed. The traditional methods, including macroscopic and microscopic observation19,20, which depend on the tester’s subjective judgment and professional experience, could effectively recognise the crude Chinese cordyceps, except as powdered products. Molecular biology techniques based on the internal transcribed spacer (ITS) sequence of Chinese cordyceps, such as DNA barcoding17,21,22, DNA barcoding coupled with high resolution melting23, loop-mediated isothermal amplification24 and nested PCR25, have been developed and used for species identification in recent years. However, none of these methods could clearly differentiate Chinese cordyceps from fermented mycelia in powdered form16,17,25. Because Chinese cordyceps is a parasitic complex, both the remains of the host larvae and the fungal sexual stroma are integral parts, as a commodity and raw medicinal material. Neither the fungus O. sinensis nor its host species are equivalent to the TCM Chinese cordyceps. Thus, a scheme that identifies both the O. sinensis fungus and its host species is theoretically reasonable and necessary, and it should more accurately identify Chinese cordyceps.
Duplex PCR technology is a good option for this purpose, because it can not only detect both targets simultaneously but is also simple and reliable. Minor genetic variations in O. sinensis from different geographical regions have been reported26, and there are up to 60 host species of the fungus O. sinensis13,16,27. Here, degenerate primer pairs were identified and designed, and a duplex PCR technique for Chinese cordyceps detection was developed. Additionally, it was confirmed that this strategy was applicable and could provide a new practical way to identify the TCM Chinese cordyceps.
Results
The development and optimisation of the duplex PCR method
In total, 78 ITS nucleotide sequences of O. sinensis were aligned and showed high identity levels, and they differed from those of the counterfeit species (Fig. 1A). Areas that were conserved and stable among O. sinensis species but differed greatly in the sequences of other species were selected as targets for designing species-specific identification primers. Because a few base variation still existed in the conserved and stable regions within these species, degenerate oligonucleotide primers for the ITS region were designed for good compatibility. The same was true for the host species’ mitochondrial cytochrome oxidase c subunit 1 (COI) gene primers (Fig. 1B). Owing to the existence of single-base differences, degenerate primers were designed for the detection of the COI gene. Then, only 1 out of 17 primer pair combinations was selected for the detection of Chinese cordyceps. The CITS primer pair (CITS-F10′: 5′-GTTGCCTCGGCGGGAC-3′/CITS-R10′-2: 5′-CMTTTGCTTGCTTCTTGACTGAG-3′) and the COI primer pair (COI-F: 5′-GGAAATCCHGGATCTTTAATT-3′/COI-R: 5′-GATGCCCCMGARTGTGCAAT-3′), were further tested for their species-specificity and retained for subsequent experiments. The primer pairs, in same PCR reactions, at a 0.3-μM primer concentration and at an annealing temperature of 54 °C always generated clean band of the expected size for O. sinensis (113 bp) and its host Hepialiae (302 bp) in the positive control template. Then, the duplex PCR assay was successfully developed for Chinese cordyceps identification.
The sequence alignments used to design the primer pairs for Chinese cordyceps (DongChongXiaCao) detection. All the sequences were subjected to a BLAST search using DNAMAN software, and the black boxes indicate identical nucleotides. The nucleotide sequences of primers employed in this duplex PCR system are listed below each alignment. (A) Alignment of internal transcribed spacer sequences from Ophiocordyceps sinensis and its related species. In total, 78 ITS nucleotide sequences of O. sinensis and other related species taken from GenBank were aligned and compared. The positions of the CITS-F10′ and CITS -R10′-2 primers are indicated by bold black boxes. (B) Alignment of COI gene sequences from its host Hepialiae species. In total, eight nucleotide sequences of the COI gene were aligned and compared. The positions of the COI-F and COI -R primers are indicated by bold black boxes.
Specificity of the duplex PCR
As shown in Fig. 2, the predicted 185-bp amplicon was amplified from all the samples tested, except for the negative control. Thus, DNAs over a wide taxonomic range can be detected, and the validity of the extracted DNAs for PCR can be assessed using the T18S-F/T18S-R primer pair. The amplicon could be used effectively as an analytical control to evaluate the authenticity of the template DNA.
Specificity test of duplex PCR using primers specific for Ophiocordyceps sinensis and its host Hepialiae species. Lane M: DNA Marker II. Lane 1: positive control (No.121201); Lane 2: negative control (water); Lanes 3–19: raw Chinese cordyceps samples (1-QH-GL, 2-XZ-NQ, 3-XZ-LS, 4-XZ-LS, 5-XZ-SN, 6-XZ-LZ, 7-XZ-LZ, 8-XZ-CD, 9-XZ-LS, 10-QH-GL, 11-QH-YS, 12-QH-YS, 13-QH-HB, 14-QH-HN, 15-QH-TR, 16-SC-KD and 17-AB); Lanes 20 and 21: mycelial samples (23-JZ1 and 24-JZ2); Lane 22: moth (25-HY); Lane 23: XinJiangChongCao (22-XJCC); Lane 24: Stachys geobombycis (26-DC); Lane 25: Euphorbia kansui (27-GS); Lane 26: Cordyceps militaris (28-YCC); Lane 27: YuNanChongCao (18-YN); Lane 28: Cordyceps gunni (19-YXB); Lane 29: Cordyceps liangschanensis (21-LS); Lane 30: Cordyceps sbarnesii (20-XB); Lane 31: Mixed powder of several conventional crops (29-MC). The simultaneous appearance of the two target bands, one of the expected size for O. sinensis (113 bp) and one for the host Hepialiae species (302 bp), was considered a positive result and marked “+”. Others marked “−” were negative.
Duplex PCR assays were further performed, and the expected fragments for the ITS (113 bp) and COI gene (302 bp) were successfully amplified from 17 Chinese cordyceps samples using the two primer pairs (CITS-F10′/CITS-R10′-2 and COI-F/COI-R), and two clear electrophoretic strips displayed the GelRed stain (Fig. 2). A single band of 113 bp appeared in the mycelial samples, and a band of 302 bp appeared in the moth and C. gracilis samples. No PCR products were detected from the other species tested, including the five readily confused varieties [C. hawkesii (C. gunni), C. liangschanensis, C. barnesii and C. militaris], the two counterfeit plants (S. geobombycis and Euphorbia kansui), one sample of a powdered crop mixture and the negative water control. The results showed that when the two target bands appeared from the same reaction, the test must be considered positive, and when none or only one target was detected, the test must be considered negative. In our specificity tests, two target amplicons were detected only in those samples identified preliminarily as Chinese cordyceps by the experts.
Cordyceps gracilis, named “XinJiangChongCao” in Chinese, is shaped like Chinese cordyceps. It is a complex containing the parasitic ergot fungus C. gracilis and its host lepidopteron larvae. The larvae of the moth Hepialus altaicola28, which is the host component of “XinJiangChongCao”, belongs to a species of Hepialidae family, which is the same as that of the host of Chinese cordyceps. As shown in Fig. 2, only the 302-bp host-specific amplicon appeared in the C. gracilis sample. This result adequately verified and confirmed the assay’s specificity.
A sequence analysis showed that the two amplified fragments were more than 99% identical to the corresponding COI gene of Hepialidae species. (GenBank Accession No: KM197522, KF696585, JQ325935, etc.) and the ITS region of O. sinensis (GenBank Accession No: KY321845~321856, MF598754, MF403011, KX082968, etc.), respectively. It was strong evidence for the specificity of the duplex PCR assay. Thus, the duplex PCR method developed in this study specifically identified the Chinese cordyceps.
Sensitivity
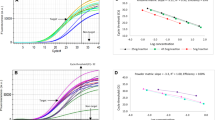
In the sensitivity test, 50.23 ng/μL total genomic DNA was initially extracted from the reference material, powdered Chinese cordyceps, and nine serial DNA dilutions were generated by fivefold serially diluting the initial extract. Then, 2 µL of each DNA dilution was added as the template in a PCR reaction, corresponding to approximately 20,000, 4,000, 800, 160, 32, 6.4, 1.3, 0.25 and 0.05 pg of the specific amplicon of the Chinese cordyceps per PCR reaction. As shown in Fig. 3, both target amplicons were simultaneously detected down to the 32 pg level in all the parallel reactions. The amplicons could not be detected in all the reactions at a lower template amount. Thus, the limit of detection (LOD), with a 95% confidence interval, for the duplex PCR assay was 32 pg in accordance with the criteria reported by Žel et al.29. Thus, the Chinese cordyceps was detected with an acceptable level of precision and accuracy. These results confirmed that the duplex PCR assay established in this study was a good approach for detecting and authenticating Chinese cordyceps.
Detection limit of the duplex PCRs for both Ophiocordyceps sinensis and its host Hepialiae species. The template was genomic DNA extracted from reference material (No. 121201) and subjected to a fivefold serial dilution, resulting in approximately 20,000, 4,000, 800, 160, 32, 6.4, 1.3, 0.25 and 0.05 pg per PCR reaction. The DL2000 DNA Marker was used as the size scale for the target DNA. Sterile water was used as the negative blank control. The simultaneous appearance of the two target bands was considered a positive result and marked “+”. Others marked “−” were negative.
Applicability
Some commercial samples were tested. As shown in Table 1, in total nine raw samples (Samples 30-T, 32-T–38-T and 40-T) produced positive amplicons, and the target bands for the ITS from O. sinensis and the COI gene from the host Hepialidae species were synchronously amplified. No bands were detected in sample 31-T and three health-care products (Samples 42-T-44-T). Additionally, a single amplicon of O. sinensis was observed in sample 39-T and two other health-care products (Samples 41-T and 45-T). This confirmed that spurious products exist on the market and that the presence of Chinese cordyceps in the samples could be diagnosed accurately. The duplex PCR method developed in this study could distinguish products made from powdered mycelia of O. sinensis fungi from raw natural Chinese cordyceps. Thus, it is a quick and efficient way to identify and authenticate Chinese cordyceps.
Conclusions and Discussions
Degenerate primer pairs targeting the ITS region of O. sinensis and the COI gene of its species-specific host were designed, and a duplex PCR assay for Chinese cordyceps was successfully established that can easily be performed in any molecular biology laboratory. It could specifically detect raw Chinese cordyceps and its primary components in products in which the genomes are undamaged. The results of the specificity analysis were in accordance with the preliminary identifications provided by experts. The LOD for Chinese cordyceps DNA could be as low as 32 pg. The accuracy and precision of the established duplex PCR method were confirmed and validated in this study. The applicability test further confirmed that the developed duplex PCR method can reliably identify and authenticate Chinese cordyceps. In addition, the duplex PCR method developed in this study is very simple to operate and cost-effective, which will be beneficial for its popularisation and utilisation. We believe that this method will be useful in the quality control and the identification of Chinese cordyceps and related products. The methodology and developmental strategy could be applied to other TCM materials as well, which will make authenticating TCMs easier. However, the method is not perfect. We were not able to differentiate mixed powders containing O. sinensis fungus mycelia and its insect host caterpillar from powders of the natural Chinese cordyceps using this method, although it was difficult to artificially raise the specific host insects required. In the future, chromatography methods that can detect the specific components of the Chinese cordyceps could be used together with the duplex PCR method. This may increase the reliability and accuracy of the identification of true Chinese cordyceps in any form.
Materials and Methods
Samples and materials
In total, 16 dried Chinese cordyceps samples (1-QH-GL, 2-XZ-NQ, 3-XZ-LS, 4-XZ-LS, 5-XZ-SN, 6-XZ-LZ, 7-XZ-LZ, 8-XZ-CD, 10-QH-GL, 11-QH-YS, 12-QH-YS, 13-QH-HB, 14-QH-HN, 15-QH-TR, 16-SC-KD and 17-AB) and one fresh Chinese cordyceps sample (9-XZ-LS) were collected from different sites in Qinghai, Tibet and Sichuan Provinces, where The Chinese cordyceps are mainly distributed. Four other varieties, C. hawkesii Gray (C. gunni) (19-YXB), C. liangschanensis Zang (21-LS), C. barnesii Thwaites (20-XB) and YunNanChongCao (18-YN) were collected in a local market in Qinghai. XinJiangChongCao [C. gracilis (Grev.) duretMont., 22-XJCC] was purchased from a local market in Urumqi, Xingjiang. The C. militaris (28-YCC) and two dried roots of S. geobombycis (26-DC) and E. kansui (27-GS) were purchased from a local market. A moth specimen (25-HY) of the family Hepialiae was provided by Harvard University. All the samples above were identified and confirmed by traditional morphological and DNA bar-coding method. A powdered reference material of Chinese cordyceps (No.121201) was purchased from National Institutes for Food and Drug Control, China. The mixed powder sample (29-MC) of conventional crop seeds including Pisum sativum, Glycine max, Zea mays, Triticum aestivum, Oryza sativa and Vicia faba were preserved and provided by Institute of Quality Standard and Testing Technology Research, Sichuan Academy of Agricultural Sciences. Two fungal strains (23-JZ1 and 24-JZ2) were isolated from stromal tissue of natural Chinese cordyceps specimens (9-XZ-LS) and were cultivated under previously described conditions30. The mycelia were then collected and conserved at −80 °C for extraction of genomic DNA. In total, 11 items (30-T~40-T), which were dried raw materials, labelled as Chinese cordyceps, were purchased as test sample from Jingdong Mall (an online shopping platform) (https://www.jd.com/) and Hehuachi Professional Market for Chinese Herbal Medicine, a local market in Chengdu. Two health-care products (41-T and 42-T) labelled with “DongChongXiaCao” were purchased from local TCM pharmacies in Chengdu, and three healthcare products (43-T–45-T) were purchased from Jingdong Mall. Detailed information regarding all the samples are shown in Table 2.
DNA extraction and purification
All the samples were treated and homogenised in liquid nitrogen. Genomic DNAs were isolated and purified from all the samples, except for two mycelia specimens, using a Plant Genomic DNA Kit (DP305-02, TIANGEN Biotech Co., Ltd., Beijing, China) in accordance with the manufacturer’s instructions. However, before that, the homogenised samples (50–100 mg each) were pre-processed in Proteinase K buffer [20-mM Tris-HCl (pH 8.0), 5-mM ethylenediaminetetraacetic acid (EDTA), 400-mM NaCl, 0.3% sodium dodecyl sulfate and 200 μg/mL Proteinase K (P6556, Sigma-Aldrich, Saint Louis, MO, USA)]. The resultant mixtures were incubated at 55 °C for 30 min, followed by 95 °C for 5 min. An equal volume of phenol/chloroform (1:1) was added to the suspensions and mixed fully for 5 min, and then, the mixtures were centrifuged at 12,000 × g for 5 min. The supernatants were transferred to fresh tubes and treated once with chloroform.
The total genomic DNAs of the two mycelia samples were prepared using the benzyl chloride method31.
The concentrations and qualities of the DNA samples were measured and evaluated using a NanoDrop ND-1000 UV/Vis Spectrophotometer (Thermo Scientific, Waltham, MA, USA) by examining the OD260/OD280 and OD260/OD230.
Primer design
ITS sequences from O. sinensis (GenBank Accession No: AJ243775, AJ243776, AJ243778, AJ245559, AJ309354, AJ309357, AJ309361, AJ413178–AJ413184, AJ507399–AJ507405, FJ654148, FJ654149 and FJ654206–FJ654259) and its related species C. militaris (GenBank Accession No: EU825996), C. hawkesii (GenBank Accession No:AJ536571), Hirsutella sp. (GenBank Accession No: EF469126) and C. liangshanensis (GenBank Accession No: AJ238503) were aligned using DNAMAN Version 6.0 (Lynnon Corporation, San Ramon, CA, USA) to select the species-specific ITS amplicons. Four kinds of Lepidoptera larvae, Thitarodes, Ahamus, Hepialus and Parahepialus, are the main host species of the fungus O. sinensis27. The sequences of COI genes from the host insects of O. sinensis, Hepialus xiaojinensis (GenBank Accession No: KP772242), Hepialus menyuanicus (GenBank Accession No: HM595858), Hepialus armoricanus (GenBank Accession No: HM595856), Ahamus yushuensis (GenBank Accession No: HM595854, HM595849, HM595840 and HM595850) and Thitarodes renzhiensis (GenBank Accession No: HM744694) were selected and aligned for species-specific amplicons of the COI gene. Owing to the minor variations that exist among the above sequences, degenerate primer pairs for the detection and identification of the commercial Chinese cordyceps were designed using Primer Premier Software (Version 5.0) (PREMIER Biosoft International, Palo Alto, CA, USA) and Primer Express Software Version 3.0 (Applied Biosystems, Foster City, CA, USA). They were synthesised by Sangon Biotech (Shanghai) Co. Ltd. (Shanghai, China). In addition, 18S ribosomal RNA gene sequences of H. armoricanus, and its closely related species and host plants (GenBank Accession No: JN036435, AF286273, KT343381, AJ830762, AJ830755, AF286298, KR068931, AJ830832, AJ830770, AJ830767, AJ830761, L37582 and U42535) were aligned. A specific universal primer pair (T18S-F: 5′-CGGAGAGGGAGCCTGAGAA-3′/T18S-R: 5′-GCACCAGACTTGCCCTCC-3′), producing a 185-bp amplicon, was designed and synthesised based on the highly conserved regions to validate the extracted DNAs for PCR.
PCR conditions
Each PCR reaction was performed in a 25-μL reaction volume with 1× HotStar Taq® Plus Master Mix (QIAGEN GmbH, Hilden, Germany), various concentrations of forward and reverse primers and 50 ng genomic DNA as the template. The tested primer concentrations were 0.16, 0.24, 0.32, 0.40 and 0.8 μM. Two primer pairs (CITS-F10′/CITS- R′-2 and COI-F/COI-R) were simultaneously added into the PCR reaction system to develop a duplex PCR assay for Chinese cordyceps identification. Primer combinations at different concentrations were tested to determine the optimal proportions that influenced the amplification efficiency. A gradient PCR was performed with annealing temperatures ranging from 50 °C to 60 °C in an ABI Veriti 96 thermal cycler (Applied Biosystems). Six temperatures in a gradient, 50 °C, 52 °C, 54 °C, 56 °C, 58 °C and 60 °C, were tested to determine the optimal annealing temperature. The program was as follows: one step of 5 min at 95 °C, 35 cycles of 30 s at 95 °C, 30 s at 50 °C–60 °C and 30 s at 72 °C; followed by one step of 7 min at 72 °C. The PCR products were size separated using electrophoresis on 2.5% agarose gel in 1 × Tris-acetate-EDTA buffer and stained with GelRed Nucleic Acid Stain (Cat. 41003, Biotium, Inc. Fremont, CA, USA) for visualisation. The amplicons were further confirmed by cloning and sequencing (Sangon Biotech (Shanghai) Co. Ltd.).
Specificity test
To determine the specificity of the primer pairs and the obtained method, the duplex PCR was tested three times using both the ITS and COI gene primer mixtures and template DNAs from Chinese cordyceps samples (17), several easily confused varieties (5), counterfeit plants (2), mycelial samples (2), a moth from the family Hepialiae (1) and a powdered mixture of conventional crops seeds (1), as mentioned in Section 2.1.
Additionally, DNA from the Chinese cordyceps reference material was used as the positive control, and sterile water was used as the negative control.
Sensitivity analysis
To further evaluate the sensitivity of the established method, genomic DNA from raw Chinese cordyceps was prepared and fivefold serially diluted using nuclease-free Tris-EDTA buffer to 0.05 pg/μL, and the gradient dilution technique previously published by Zhang et al.32. The duplex PCR assays were performed as described in Section 2.4, and the serial dilutions were used as the templates. All the reactions were repeated three times, with 15 parallel runs for each template DNA, to determine the LOD of the duplex PCR assay based on the positive result criteria29.
Applicability test
To test the applicability of the duplex PCR method for analysing practical samples, 16 samples were purchased from a local market or Jingdong Mall, as described in Section 2.1. DNAs were isolated from 11 raw Chinese cordyceps samples and 5 health-care products containing Chinese cordyceps, according to the labels. Then, the duplex PCR assays were performed in triplicate using the obtained DNAs as templates. The genomic DNA isolated from the Chinese cordyceps reference material was used as the positive control. Sterile water was used as the negative control.
References
Lo, H. C. et al. A Systematic Review of the Mysterious Chinese cordyceps Ophiocordyceps sinensis in DongChongXiaCao (冬蟲夏草Dōng Chóng Xià Cǎo) and Related Bioactive Ingredients. Journal of Traditional and Complementary Medicine 3, 16–32 (2013).
Sung, G. H. et al. Phylogenetic classification of Cordyceps and the clavicipitaceous fungi. Studies in Mycology 57, 5–59 (2007).
Hu, X. et al. Genome survey uncovers the secrets of sex and lifestyle in Chinese cordyceps. Chinese Science Bulletin 58, 2846–2854 (2013).
Zeng, W. et al. The growth of Cordyceps sinensis (Berk.) Sacc. In the infection and parasitic phases. Mycosystema 25, 646–650 (2006).
Lei, W. et al. The latent infection process of Ophiocordyceps sinensis in host Thitarodes larvae. Journal of Environmental Entomology 37, 387–392 (2015).
Xu, M., Xu, M. & Li, R. Q. Progress and prospects of several crucial aspects in the biological and ecological research on the The Chinese cordyceps, Ophiocordyceps sinensis. Acta Ecologica Sinica 39(5), 1853–1862 (2019).
Shrestha, B. et al. What is the The Chinese cordyceps Ophiocordyceps sinensis (Ophiocordycipitaceae)? Mycology 1, 228–236 (2010).
Zhang, S. et al. Ophiocordyceps sinensis and Cordyceps militaris: research advances, issues and perspectives. Mycosystema 32, 577–597 (2013).
Han, R. C. et al. Research on Chinese cordyceps during the past 70 years in China. Chinese Journal of Applied Entomology 56(5), 849–883 (2019).
Li, H. Y. et al. Quantitative analysis of five nucleosides in Cordyceps by HPLC and quality control of Cordyceps militaris. Chinese Traditional and Herbal. Drugs 22, 5410–5417 (2018).
Hu, X. D. et al. Progress in Pharmacological Research of Cordyceps Polysaccharides from Ophiacordyceps sinensis. 6, 224–229 (2016).
Nakamura, K., Shinozuka, K. & Yoshikawa, N. Anticancer and antimetastatic effects of cordycepin, an active component of Cordyceps sinensis. Journal of Pharmacological Sciences 127, 53–56 (2015).
Li, Y. et al. A survey of the geographic distribution of Ophiocordyceps sinensis. The Journal of Microbiology 49, 913–919 (2011).
Wichadakul, D. et al. Insights from the genome of Ophiocordyceps polyrhachis-furcata to pathogenicity and host specificity in insect fungi. BMC Genomics 16, 881 (2015).
Zhang., G. R. et al. Factors influencing the occurrence of Ophiocordyceps sinensis. Acta Ecologica Sinica 31, 4117–4125 (2011).
Su, Y. Y. et al. Rapid identification of Cordyceps and its products using DNA barcoding method. China Journal of Chinese Materia Medica. 2, https://doi.org/10.19540/j.cnki.cjcmm.20190131.002 (2019).
Xiang, L. et al. DNA barcoding the commercial The Chinese cordyceps. FEMS Microbiol Letters 347, 156–162 (2013).
Hsu, T. H. et al. A comparison of the chemical composition and bioactive ingredients of the Chinese medicinal mushroom DongChongXiaCao, its counterfeit and mimic, and fermented mycelium of Cordyceps sinensis. Food Chemistry 78, 463–469 (2002).
Liu, H. et al. Morphological and microscopic identification studies of Cordyceps and its counterfeits. Acta Pharmaceutica Sinica B 1, 189–195 (2011).
Au, D. et al. Application of microscopy in authentication of valuable Chinese medicine i-Cordyceps sinensis, its counterfeits, and related products. Microscopy Research and Technique 75, 54–64 (2012).
Zhang, W. J., Wei, F. & Ma, S. C. Application of ITS1 barcode sequence for the identification of Cordyceps sinensis from its counterfeit species. Chinese Journal of Pharmaceutical Analysis 35, 1716–1721 (2015).
Chen, S. et al. A renaissance in herbal medicine identification: from morphology to DNA. Biotechnology Advances 32, 1237–1244 (2014).
Sun, W. et al. The potential power of Bar-HRM technology in herbal medicine identification. Frontiers in Plant Science 7, 367 (2016).
Zhao, M. et al. Rapid authentication of the precious herb saffron by loop-mediated isothermal amplification (LAMP) based on internal transcribed spacer 2 (ITS2) sequence. Scientific Reports 6, 25370 (2016).
Jin, G. S. et al. Development of conventional and nested PCR assays for the detection of Ophiocordyceps sinensis[J]. J. Basic Microbiol 53(4), 340 (2013).
Chen, Y. Q. et al. Genetic variation of Cordyceps sinensis, a fruit-body-producing entomopathogenic species from different geographical regions in China. FEMS Microbiology Letters 230, 153–158 (2004).
Wang, X. L. & Yao, Y. J. Host insect species of Ophiocordyceps sinensis: a review. ZooKeys 127, 43–59 (2011).
Wang, Y. et al. Non-destructive methods for sex determination of the live pupae of Hepinlus altaicola Wang (Lepidoptera: Hepialidae). Journal of Environmental Entomology 40, 30–35 (2018).
Žel, J. et al. How to Reliably Test for GMOs. Springer Briefs in Food, Health, and Nutrition, 1–95 (2012).
Zhang, Y. J., Liu, X. Z. & Wang, M. Cloning, expression, and characterization of two novel cuticle-degrading serine proteases from the entomopathogenic fungus Cordyceps sinensis. Research in Microbiology 159, 462–469 (2008).
Zhu, H., Qu, F. & Zhu, L. H. Isolation of genomic DNAs from fungi using benzyl chloride. Acta Mycologica Sinica 13, 34–40 (1994).
Zhang, F. L. et al. A new construct specific real-time PCR method for screening GMO ingredients with gat-tpinII cassette in foods and seeds. Food Control 86, 266–274 (2018).
Acknowledgements
This work was financially supported by the Special Funds of Ability Promotion Project from Sichuan Provincial Finance (2016GYSH-032). We thank Dr. Youqing Bai from Tibet, Mr. Ling Ma from Qinghai Province, Zhi-Qiong Lan from Chengdu University of Traditional Chinese Medicine and Dr. Zhengyang Wang from Sichuan who helped us during sample collection. We thank Prof. Liangke Song and Jia-Yu Zhou from Southwest Jiaotong University for their expert help in identifying materials. Additionally, we thank all the reviewers for useful suggestions during the preparation of this manuscript. We thank Lesley Benyon, PhD, from Liwen Bianji, Edanz Group China (www.liwenbianji.cn/ac), for editing the English text of a draft of this manuscript. Thanks to our family and friends for all the support.
Author information
Authors and Affiliations
Contributions
Zhang F.-L., Lei S.-R., Guo L.-A. and Song J. conceived the study. Zhang F.-L. and Yang X.-F. carried out the experiments and prepared the manuscript. Wang D. and Liu W.-J. provided suggestions about the manuscript. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, FL., Yang, XF., Wang, D. et al. A simple and effective method to discern the true commercial Chinese cordyceps from counterfeits. Sci Rep 10, 2974 (2020). https://doi.org/10.1038/s41598-020-59900-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-59900-9
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.