Abstract
Lysophosphatidylserine (LysoPS) is a lysophospholipid, its generating enzyme, phosphatidylserine-specific phospholipase A1 (PS-PLA1), reportedly plays roles in stomach and colon cancers. Here, we examined the potential roles of LysoPS in hepatocellular carcinoma (HCC). The ninety-seven HCC patients who underwent surgical treatment were enrolled in this study and approved by the institutional review board. Among LysoPS-related enzymes and receptors, increased PS-PLA1 or LysoPS receptor 1 (LPS1) mRNA was observed in HCC tissues compared to non-HCC tissues. PS-PLA1 mRNA in HCC was associated with no clinical parameters, while LPS1 mRNA in HCC was correlated inversely with tumor differentiation. Furthermore, higher serum PS-PLA1 was observed in HCC patients compared to healthy control and correlated with PS-PLA1 mRNA in non-HCC tissues and with serum AST or ALT. Additionally, serum levels of PS-PLA1 were higher in HCC patients with HCV-related liver injury than in those with HBV or non-HBV-, non-HCV-related liver diseases. In conclusion, among LysoPS-related enzymes and receptors, PS-PLA1 and LPS1 mRNA were increased in HCC. Based on the correlation between the serum PS-PLA1 and the mRNA level of PS-PLA1 in non-HCC tissues, the liver may be the main source of serum PS-PLA1, and serum PS-PLA1 levels may be a useful marker for liver injury.
Similar content being viewed by others
Introduction
Over the last two decades, lysophospholipids (LPLs) have been studied widely as biologically active molecules that play multiple roles in carcinogenesis, neurogenesis, vascular development and immunity1,2,3,4. Among LPLs, the crucial roles of lysophosphatidic acid (LPA) and sphingosine 1-phosphate (S1P) have been extensively unveiled, exerting multiple cellular responses with G-protein coupled receptors (LPA1–6 and S1P1–5). We recently reported the involvement of LPA and S1P in the pathogenesis of hepatocellular carcinoma (HCC) and colon cancer5,6,7. On the other hand, the evidence has been relatively scarce showing the actions and the clinical roles of other LPLs, such as lysophosphatidylglycerol, lysophosphatidylserine (LysoPS), lysophosphatidylinositol, lysophosphatidylethanolamine or lysophosphatidylcholine.
Among these LPLs, we have been paying attention to LysoPS, the binding receptors of which were recently identified as LPS1 (GPR34), LPS2 (P2Y10) and LPS3 (GPR174) in humans and LPS2L (A630033H20 or LPS2-like receptor) in rodents8,9. LysoPS is known to affect several cellular responses, including mast cell degranulation10,11, lymphocyte proliferation12, fibroblast migration13,14 and neurite outgrowth15. Regarding its receptors, LPS1 is expressed in many tissues, mostly in mast cells16 but it is not involved in the mast cell degranulation induced by LysoPS17. The roles of LPS2 and LPS3 remain to be elucidated at present, but the expressions of which were reported in lymphoid organs for LPS2, LPS3 and in the spleen and thymus for LPS218,19. Among LysoPS receptors (LPSRs), the ectopic expression of LPS1 was reported in several solid tumors, such as stomach cancer20, with a potential correlation with clinico-pathological characteristics, tumor differentiation or prognosis. LysoPS stimulates the migratory ability of colon cancer cell lines through the LPS1 and PI3K/Akt pathways21.
Phosphatidylserine specific-phospholipase A1 (PS-PLA1), one of the enzymes that is responsible for LysoPS production and acts on PS in the cell bilayer, has the roles in the oncogenesis and metastasis of colorectal cancer in humans22. We have found evidence suggesting that the PS-PLA1-mediated pathway might be involved in the production of LysoPS in stomach cancer23. Furthermore, we have reported that the serum levels of PS-PLA1 are higher in melanoma subjects and associated with its clinical stages24. Because these lines of evidence may suggest a role for LysoPS in cancer, we sought to examine the potential involvement of LysoPS in the pathophysiology of HCC.
Results
mRNA levels of LysoPS-related enzymes and receptors in HCC
A total of 97 patients with HCC who underwent surgical treatment were analyzed, among whom, 61 were diagnosed with primary HCC and 36 with recurrent HCC. It is well known that HCC recurrence is frequently ectopic because the underlying chronic liver diseases continue to increase the risk for hepatocarcinogenesis in patients25. The clinical and laboratory characteristics of the patients are shown in Table 1.
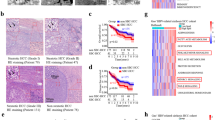
We first examined the mRNA levels of LysoPS-related enzymes and receptors in HCC tissues in comparison with adjacent non-HCC tissues. As depicted in Fig. 1a,b increased mRNA levels in HCC compared to non-HCC tissues were observed for PS-PLA1 (P < 0.05) and LPS1 (P < 0.01). Higher mRNA levels of PS-PLA1 and LPS1 in HCC tissues were observed in 42 of 97 (43.3%) and 61 of 97 patients (62.8%), respectively. The mean mRNA levels of PS-PLA1 and LPS1 were 1.4- and 1.7-fold higher, respectively, in HCC tissues than in non-HCC tissues (Fig. 1a,b). In contrast, no differences were observed between HCC tissues and non-HCC tissues regarding the mRNA levels of the other two LysoPS receptors, LPS2 (P = 0.57) and LPS3 (P = 0.16) (Fig. 1c).
mRNA levels of LysoPS-generating enzyme and its receptors. (a) PS-PLA1 mRNA levels were increased (*p < 0.05) in HCC tissues in comparison to non-HCC tissues in 42 of 97 patients, being 1.4-fold higher in HCC tissues than non-HCC tissues. (b) Higher mRNA levels of LPS1 in HCC tissues were observed in 61 of 97 patients (**p < 0.01), being 1.7-fold higher in HCC tissues than non-HCC tissues. (c) LPS2, LPS3 mRNA levels did not differ between HCC tissues and non-HCC tissues. (d) LPS1 mRNA levels in HCC were correlated with tumor differentiation.
To explore the features of HCC with enhanced mRNA levels of PS-PLA1 or LPS1, potential relationships with clinical parameters were analyzed, where there were no correlation between mRNA levels of PS-PLA1 in HCC and the clinical parameters. On the other hand, the increase in mRNA levels of LPS1 in HCC tissues, expressed as fold, was correlated with tumor differentiation; the differentiation levels of HCC were poorer with higher mRNA levels of LPS1 (Fig. 1d); however, prognosis was not different according to mRNA levels of LPS1.
Serum levels of PS-PLA1 in HCC patients
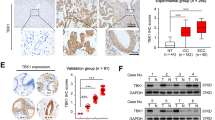
In these patients with HCC, serum levels of PS-PLA1 were measured as 23.3 ± 9.4 μg/L, which were significantly higher than those in 59 healthy subjects, 15.4 ± 10.7 μg/L, reported in our previous study24 (P < 0.01). Although increased mRNA levels in HCC tissues may be responsible for the increase in serum PS-PLA1 levels, at least in part, serum PS-PLA1 levels were not correlated with PS-PLA1 mRNA levels in HCC tissues but were correlated with those in non-HCC, background tissues (Fig. 2a). Furthermore, serum PS-PLA1 levels were not correlated with any parameters for HCC, including its size or HCC markers, such as AFP or PIVKA II (Table 2). Among the parameters in general, serum PS-PLA1 levels were strongly correlated with serum levels of AST and ALT (Fig. 2b, Table 2) but not with platelet counts and serum albumin levels (Table 2). Histological findings consistently revealed no association between the serum PS-PLA1 levels and the fibrosis stages of non-HCC, background tissues (Fig. 2c).
Serum PS-PLA1 levels and their correlation with various parameters. (a) Serum PS-PLA1 levels were correlated with the mRNA levels of PS-PLA1 in non-HCC tissues but not in HCC tissues. (b) The relationship between serum PS-PLA1 levels and clinical markers. A strong correlation was observed with serum levels of AST and ALT (b) but not with background liver fibrosis stages (c). (d) Serum PS-PLA1 levels in HCC patients with different etiologies. Serum PS-PLA1 levels in HCC patients with HCV infection were compared to those with HBV infection or non-HBV, non-HCV-related liver disease. (e) Serum PS-PLA1 levels in liver injuries without HCC. Serum PS-PLA1 levels were increased in non-HCC patients with liver cirrhosis caused by HCV (23.5 ± 5.9 μg/L) or non-alcoholic fatty liver disease (23.4 ± 9.1 μg/L) in comparison to healthy subjects (15.4 ± 10.7 μg/L).
Next, we examined serum PS-PLA1 levels in HCC patients according to the background liver diseases. Figure 2d demonstrates that serum PS-PLA1 levels were higher in HCC patients with hepatitis C-related chronic liver disease than those with hepatitis B-related chronic liver disease or those with non-HBV, non-HCV-related chronic liver disease. The current results suggest that the background liver of HCC may be involved in the regulation of serum PS-PLA1 levels as the site of production. Thus, we further wondered whether serum PS-PLA1 levels might be increased in liver injury without HCC. Indeed, serum PS-PLA1 levels were higher in patients with non-alcoholic fatty liver disease (NAFLD), 23.4 ± 9.1 μg/L (P < 0.01, n = 19), compared to those in healthy subjects (Fig. 2e), and tended to be higher in patients with liver cirrhosis caused by hepatitis C virus, 23.5 ± 5.9 μg/L, though not significant due to the small sample size (P = 0.053, n = 4). Although the exact mechanism(s) should be further elucidated, the high levels of serum PS-PLA1 in non-alcoholic fatty liver disease are noted.
Discussion
In the current study, the increased mRNA levels of a generating enzyme, PS-PLA1, and a receptor (LPS1) of LysoPS were observed in HCC tissues compared to non-HCC tissues. We then examined the potential relationships between the mRNA levels of PS-PLA1 or LPS1 in HCC and its clinical parameters. We found only a correlation between higher LPS1 mRNA levels in HCC and poorer degree of differentiation, but no other correlations between PS-PLA1 mRNA levels and the clinical parameters were observed. However, the prognosis of HCC was not different according to LPS1 mRNA levels. This is because the curative operation was performed in the patients enrolled, obscuring the effects of tumor differentiation and hence LPS1 on prognosis.
In rodents, PS-PLA1 is reportedly released from activated platelets26, but regarding humans, its release is to be elucidated. The current study suggests that the liver may play a role in the regulation of serum PS-PLA1 levels, based on the evidence that serum PS-PLA1 levels were correlated with mRNA levels of PS-PLA1 in non-HCC, background liver tissue but not with those in HCC tissue. We would assume that because the size of HCC treated with surgical resection in the current study was relatively small, enhanced PS-PLA1 production in HCC might not affect serum PS-PLA1 levels. It should be clarified whether serum PS-PLA1 levels might be useful as a marker of HCC with larger sized tumors. Nonetheless, in line with the current finding, serum PS-PLA1 levels varied according to the etiology of the background liver tissue, and serum PS-PLA1 levels were increased in patients with liver injury in the absence of HCC. Because serum PS-PLA1 levels were correlated with serum levels of AST and ALT, it is speculated that hepatocyte injury may cause active production of PS-PLA1 in the liver and hence may lead to an increase in serum PS-PLA1 levels.
As described in the introduction, the evidence showing the clinical significance of PS-PLA1 has been very scarce. As a limitation of our current study, we could just find the regulatory mechanism of serum PS-PLA1 levels, at least in part, without clarifying the clinical significance of PS-PLA1. In the current study, we observed elevated serum PS-PLA1 in patients with NAFLD, which suggests the involvement of the PS-PLA1 in the metabolic syndrome. Similar to PS-PLA1, upregulation of LPS1 in HCC has been unveiled in the current study, however, its significance remains obscure. Although we could find the supportive data for a link between HCC and PS-PLA1 or LPS1, the data are phenomenological at present and should be elucidated further.
Patients and Methods
Patients
Patients with HCC who were treated at the Hepatobiliary Pancreatic Surgery Division, Department of Surgery, the University of Tokyo Hospital, between January 2013 and June 2015 provided consent to be enrolled in this study. All of the enrolled patients underwent liver resection, among whom, 61 patients developed primary HCC and 36 had recurrent HCC, from whom sufficient HCC tissues and adjacent non-tumorous tissues for the analyses were obtained. None of the patients received pre-op treatment.
This study was conducted in accordance with the ethical guidelines of the 1975 Declaration of Helsinki and was approved by the Institutional Research Ethics Committee of the University of Tokyo. Written informed consent was obtained from the patients for the use of clinical samples.
Quantitative real-time PCR of LPSRs and PS-PLA1
Quantitative real-time PCR was performed as previously described6. Human LPS1, LPS2, LPS3, PS-PLA1 and eukaryotic ribosomal 18 S primers and probes (TaqMan Gene Expression Assays) were obtained from Applied Biosystems (Hs00271105_s1, Hs00274326_s1, Hs00261404_s1, Hs01056915_m1 and Hs99999901_s1). The reaction conditions were as follows: incubation for 10 min at 95 °C, followed by 40 cycles at 95 °C for 15 sec and 60 °C for 1 min. The target gene mRNA expression levels were quantified relative to the 18S ribosomal RNA using the 2−ΔΔCt method (Applied Biosystems, User Bulletin No. 2).
Measurement of serum PS-PLA levels
Serum samples were collected after patient admission and before surgical procedure. Serum levels of PS-PLA1 were measured using a two-site immunoenzymatic assay in the TOSOH AIA system (TOSOH, Tokyo, Japan)27.
Statistical analysis
All tests for significance were two-tailed, and P < 0.05 was considered statistically significant. The cumulative incidence of intra and extrahepatic recurrence was calculated using the Kaplan-Meier method, and differences among groups were assessed using the log-rank test. A paired t-test was used to analyze differences in mRNAs levels of LPSRs in tumors and corresponding nontumorous tissues. Data processing and analysis were performed using R statistic software version 3.3.1 (http://www.r-project.org).
The results of the in vitro experiments are expressed as the means and standard deviations (SDs). The results were considered significant when P-values were < 0.05.
References
Aikawa, S., Hashimoto, T., Kano, K. & Aoki, J. Lysophosphatidic acid as a lipid mediator with multiple biological actions. J. Biochem. 157, 81–89, https://doi.org/10.1093/jb/mvu077 (2015).
Takuwa, Y., Okamoto, Y., Yoshioka, K. & Takuwa, N. Sphingosine-1-phosphate signaling in physiology and diseases. Biofactors 38, 329–337, https://doi.org/10.1002/biof.1030 (2012).
Frasch, S. C. & Bratton, D. L. Emerging roles for lysophosphatidylserine in resolution of inflammation. Prog. Lipid Res. 51, 199–207, https://doi.org/10.1016/j.plipres.2012.03.001 (2012).
Grzelczyk, A. & Gendaszewska-Darmach, E. Novel bioactive glycerol-based lysophospholipids: new data–new insight into their function. Biochim. 95, 667–679, https://doi.org/10.1016/j.biochi.2012.10.009 (2013).
Enooku, K. et al. Higher LPA2 and LPA6 mRNA Levels in Hepatocellular Carcinoma Are Associated with Poorer Differentiation, Microvascular Invasion and Earlier Recurrence with Higher Serum Autotaxin Levels. Plos One 11, e0161825, https://doi.org/10.1371/journal.pone.0161825 (2016).
Uranbileg, B. et al. Increased mRNA Levels of Sphingosine Kinases and S1P Lyase and Reduced Levels of S1P Were Observed in Hepatocellular Carcinoma in Association with Poorer Differentiation and Earlier Recurrence. Plos One 11, e0149462, https://doi.org/10.1371/journal.pone.0149462 (2016).
Uranbileg, B. et al. Evidence Suggests Sphingosine 1-Phosphate Might Be Actively Generated, Degraded, and Transported to Extracellular Spaces With Increased S1P2 and S1P3 Expression in Colon Cancer. Clin. Colorectal Cancer 17, e171–e182, https://doi.org/10.1016/j.clcc.2017.11.004 (2018).
Inoue, A. et al. TGFalpha shedding assay: an accurate and versatile method for detecting GPCR activation. Nat. Methods 9, 1021–1029, https://doi.org/10.1038/nmeth.2172 (2012).
Makide, K. et al. Novel lysophosphoplipid receptors: their structure and function. J. Lipid Res. 55, 1986–1995, https://doi.org/10.1194/jlr.R046920 (2014).
Martin, T. W. & Lagunoff, D. Interactions of Lysophospholipids and Mast-Cells. Nat. 279, 250–252, https://doi.org/10.1038/279250a0 (1979).
Smith, G. A., Hesketh, T. R., Plumb, R. W. & Metcalfe, J. C. Exogenous Lipid Requirement for Histamine-Release from Rat Peritoneal Mast-Cells Stimulated by Concanavalin-A. Febs Lett. 105, 58–62, https://doi.org/10.1016/0014-5793(79)80887-X (1979).
Bellini, F. & Bruni, A. Role of a serum phospholipase A1 in the phosphatidylserine-induced T cell inhibition. FEBS Lett. 316, 1–4, https://doi.org/10.1016/0014-5793(93)81724-e (1993).
Lee, S. Y. et al. Lysophosphatidylserine stimulates chemotactic migration in U87 human glioma cells. Biochem. Bioph Res. Co. 374, 147–151, https://doi.org/10.1016/j.bbrc.2008.06.117 (2008).
Park, K. S. et al. Lysophosphatidylserine stimulates L2071 mouse fibroblast chemotactic migration via a process involving pertussis toxin-sensitive trimeric G- proteins. Mol. pharmacology 69, 1066–1073, https://doi.org/10.1124/mol.105.018960 (2006).
Lourenssen, S. & Blennerhassett, M. G. Lysophosphatidylserine potentiates nerve growth factor-induced differentiation of PC12 cells. Neurosci. Lett. 248, 77–80, https://doi.org/10.1016/s0304-3940(98)00275-4 (1998).
Sugo, T. et al. Identification of a lysophosphatidylserine receptor on mast cells. Biochem. Biophys. Res. Commun. 341, 1078–1087, https://doi.org/10.1016/j.bbrc.2006.01.069 (2006).
Liebscher, I. et al. Altered immune response in mice deficient for the G protein-coupled receptor GPR34. J. Biol. Chem. 286, 2101–2110, https://doi.org/10.1074/jbc.M110.196659 (2011).
Rao, S., Garrett-Sinha, L. A., Yoon, J. & Simon, M. C. The Ets factors PU.1 and Spi-B regulate the transcription in vivo of P2Y10, a lymphoid restricted heptahelical receptor. J. Biol. Chem. 274, 34245–34252, https://doi.org/10.1074/jbc.274.48.34245 (1999).
Qin, Y. et al. Quantitative expression profiling of G-protein-coupled receptors (GPCRs) in metastatic melanoma: the constitutively active orphan GPCR GPR18 as novel drug target. Pigment. Cell Melanoma Res. 24, 207–218, https://doi.org/10.1111/j.1755-148X.2010.00781.x (2011).
Yu, W. D. et al. Upregulation of GPR34 expression affects the progression and prognosis of human gastric adenocarcinoma by PI3K/PDK1/AKT pathway. Histol. Histopathol. 28, 1629–1638 (2013).
Iida, Y. et al. Lysophosphatidylserine stimulates chemotactic migration of colorectal cancer cells through GPR34 and PI3K/Akt pathway. Anticancer. Res. 34, 5465–5472 (2014).
Iida, Y. et al. Phosphatidylserine-specific phospholipase A1 (PS-PLA1) expression in colorectal cancer correlates with tumor invasion and hematogenous metastasis. Anticancer. Res. 35, 1459–1464 (2015).
Emoto, S. et al. Analysis of glycero-lysophospholipids in gastric cancerous ascites. J. Lipid Res. 58, 763–771, https://doi.org/10.1194/jlr.P072090 (2017).
Kurano, M. et al. Association between serum autotaxin or phosphatidylserine-specific phospholipase A1 levels and melanoma. J. Dermatol. 45, 571–579, https://doi.org/10.1111/1346-8138.14278 (2018).
El-Serag, H. B. Hepatocellular carcinoma. N. Engl. J. Med. 365, 1118–1127, https://doi.org/10.1056/NEJMra1001683 (2011).
Sato, T. et al. Serine phospholipid-specific phospholipase A that is secreted from activated platelets. A new member of the lipase family. J. Biol. Chem. 272, 2192–2198, https://doi.org/10.1074/jbc.272.4.2192 (1997).
Nakamura, K. et al. A novel enzyme immunoassay for the determination of phosphatidylserine-specific phospholipase A(1) in human serum samples. Clin. Chim. Acta 411, 1090–1094, https://doi.org/10.1016/j.cca.2010.04.006 (2010).
Acknowledgements
The authors thank Koji Igarashi PhD, Bioscience Division, TOSOH Corporation, Kanagawa, Japan, for his valuable support and technical assistance. This work was supported by JSPS KAKENHI Grant Numbers 15H05906 (to Y.Y.) and 16K09343 (to H.I.).
Author information
Authors and Affiliations
Contributions
B.U.: acquisition, analysis and interpretation of the data, and drafting the manuscript. M.K.: analysis and interpretation of the data. M.S.: statistical analysis, acquisition of the data. H.I.: study concept and design, analysis and interpretation of the data, and drafting the manuscript. T.I.: acquisition of the data. K.H. and N.K.: study supervision Y.Y.: study concept and design, and drafting the manuscript
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Uranbileg, B., Kurano, M., Sato, M. et al. Possible involvement of PS-PLA1 and lysophosphatidylserine receptor (LPS1) in hepatocellular carcinoma. Sci Rep 10, 2659 (2020). https://doi.org/10.1038/s41598-020-59590-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-59590-3
This article is cited by
-
Distinctly altered lipid components in hepatocellular carcinoma relate to impaired T cell-dependent antitumor immunity
Hepatology International (2024)
-
PLA1A expression as a diagnostic marker of BRAF-mutant metastasis in melanoma cancer
Scientific Reports (2021)
-
Current Knowledge on the Biology of Lysophosphatidylserine as an Emerging Bioactive Lipid
Cell Biochemistry and Biophysics (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.