Abstract
Sundaland constitutes one of the largest and most threatened biodiversity hotspots; however, our understanding of its biodiversity is afflicted by knowledge gaps in taxonomy and distribution patterns. The subfamily Rasborinae is the most diversified group of freshwater fishes in Sundaland. Uncertainties in their taxonomy and systematics have constrained its use as a model in evolutionary studies. Here, we established a DNA barcode reference library of the Rasborinae in Sundaland to examine species boundaries and range distributions through DNA-based species delimitation methods. A checklist of the Rasborinae of Sundaland was compiled based on online catalogs and used to estimate the taxonomic coverage of the present study. We generated a total of 991 DNA barcodes from 189 sampling sites in Sundaland. Together with 106 previously published sequences, we subsequently assembled a reference library of 1097 sequences that covers 65 taxa, including 61 of the 79 known Rasborinae species of Sundaland. Our library indicates that Rasborinae species are defined by distinct molecular lineages that are captured by species delimitation methods. A large overlap between intraspecific and interspecific genetic distance is observed that can be explained by the large amounts of cryptic diversity as evidenced by the 166 Operational Taxonomic Units detected. Implications for the evolutionary dynamics of species diversification are discussed.
Similar content being viewed by others
Introduction
Over the past two decades, the spectacular aggregation of species in biodiversity hotspots has attracted attention by scientists and stakeholders alike1,2,3,4. However, this exceptional concentration of often-endemic species at small spatial scales is threatened by the rise of anthropogenic disturbances. Of the 26 initially identified terrestrial biodiversity hotspots1, the ones located in Southeast Asia (SEA) (Indo-Burma, Philippines, Sundaland and Wallacea) currently rank among the most important both in terms of species richness and the extend of endemism but also rank as the most threatened by human activities3. Sundaland is currently the most diverse terrestrial biodiversity hotspot of SEA and is the most threatened5. Sundaland comprises Peninsular Malaysia and the islands of Java, Sumatra, Borneo, and Bali and its diversity originates from the complex geological history of the region, linked to major tectonic changes in the distribution of land and sea during the last 50 Million years (My)6, but also from eustatic fluctuations that have sporadically connected and disconnected Sundaland landmasses during glacial-interglacial cycles in the Pleistocene7,8,9. Therefore, Sundaland biogeography has received increased attention over the past decade resulting in the detection of contrasting spatial and temporal patterns in various groups9,10,11,12,13,14.
Species richness within vertebrate groups is high in Sundaland1 and freshwater fishes are no exceptions to that. With more than 900 species reported to date, and with nearly 45 percent of endemism, Sundaland’s ichthyofauna is the largest in SEA and accounts for nearly 75 percent of the entire ichthyodiversity of the Indonesian archipelago15. The inventory of Sundaland’s freshwater fishes started more than two centuries ago and despite the acceleration of species discovery over the last three decades, it is still a work in progress15. The complexity of this inventory was partly exacerbated by the abundance of minute species i.e. less than 5 cm length15, but also by the inconsistent use of species names through time for old descriptions due to the loss of type specimens or uncertainties in the location of type localities16,17. The family Cyprinidae sensu lato is a particularly good example for the complexity of Sundaland freshwater fishes taxonomy and systematics. The systematics of this large family of Cypriniformes, with over 3,000 species, has been controversial for more than a century18. Based on recent molecular phylogenetic studies19,20,21, Tan and Armbruster22 proposed a new classification dividing the Cyprinidae sensu lato into 12 families. The subfamily Rasborinae (Cypriniformes, Cyprinoidei, Danionidae) comprises roughly 140 species in 11 genera: Amblypharyngodon, Boraras, Brevibora, Horadandia, Kottelatia, Pectenocypris, Rasbora, Rasboroides, Rasbosoma, Trigonopoma, and Trigonostigma22. In Sundaland the subfamily Rasborinae is represented by 79 species in 7 genera. The genera Amblypharyngodon, Horadandia, Rasboroides, and Rasbosoma do not occur in Sundaland. By far the most species rich rasborine genus is Rasbora with over 100 species in total and 65 species in Sundaland. Long considered a catch-all group, several attempts have been made to provide a classification of the genus Rasbora that reflects phylogeny. In a comprehensive revision, Brittan23 recognized 3 subgenera (Rasbora, Rasboroides, and Megarasbora) and divided Rasbora into 8 species complexes, now regarded as species groups24 (Fig. 1). Subsequent authors have erected several new genera or suggested new species composition for the various Rasbora species groups19,24,25,26. Clearly, to better understand the evolutionary history of this unique group, the taxonomy and systematic of the Rasborinae needs to be better understood.
Selected species of Rasborinae that illustrate the diversity of the subfamily in Sundaland. All pictures, except 1, 6.1, 7.1 and 7.2, originate from the Barcode of Life Datasystem (dx.doi.org/110.5883/DS-BIFRA, Creative Commons Attribution - Non Commercial - Share Alike), pictures 1, 6.1 and 7.2 originate from FFish.asia (https://ffish.asia, Creative Commons Attribution - Non Commercial - Share Alike).
The use of standardized DNA-based approaches to the inventory of Sundaland’s ichthyofauna resulted in the detection of considerable knowledge gaps16,17,27. In addition, substantial levels of cryptic diversity (i.e. morphologically unrecognized diversity) were repeatedly reported for a wide range of Sundaland freshwater fish taxa10,27,28,29,30,31,32,33 including the Rasborinae16. The taxonomy of most Rasborinae species, particularly so for the genus Rasbora, remains challenging due to the diversity of the group and the morphological similarity of many closely related species. As a consequence, the actual distribution ranges of many species of Rasborinae are not well known.
This study aims to re-examine Rasborinae diversity in Sundaland. We generated a DNA barcode reference library to (1) explore biological species boundaries with DNA-based species delimitation methods, (2) validate species identity, taxonomy and precise range distribution by producing DNA barcodes from type localities or neighboring watersheds, (3) validate or revise of the previously published DNA barcodes records for the subfamily Rasborinae available on GenBank.
Results
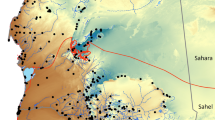
Sequencing of the DNA barcode marker Cytochrome Oxidase 1 (COI) yielded a total of 991 new sequences (Table S2) from 189 sampling sites distributed across Sundaland (Fig. 2). Together with 106 DNA barcodes mined from GenBank and BOLD (Table S3), we assembled a DNA barcode reference library of 1,097 sequences from 65 taxa of Rasborinae and 1 taxon of Sundadanionidae (Sundadanio retiarius). The number of specimens analyzed per species ranged from 1 to 143, with an average of 14.6 sequences per species and only six species represented by a single sequence. The sequences ranged from 459 bp to 651 bp long, with 99 percent of the sequences being above 500 bp length, and no stop codons were detected, suggesting that all the sequences correspond to functional mitochondrial COI sequences. DNA barcodes for 61 of the 79 nominal species of Rasborinae reported from Sundaland were recovered (approximately 78%) corresponding to the 7 Rasborinae genera currently recognized (Table S1). The present study achieved a complete coverage at the species level for the genera Boraras (2 species), Brevibora (3 species), Kottelatia (1 species), Trigonopoma (2 species) and Trigonostigma (3 species). In turn, two out of the three Pectenocypris species (66%) and 48 out of the 65 Rasbora species (74%) currently recognized in Sundaland were collected (Table S1). Geographically, our dataset includes 86% of the Rasborinae of Borneo (38 out of 44), all the Rasbora species of Java (4 species) and 68% of the Rasborinae species of Sumatra (26 out of 38) were collected (Table S1). Finally, four undescribed taxa are highlighted, two taxa of Rasbora in Java, one taxon of Trigonostigma in Borneo (Table S2) and an additional Rasbora taxon, previously assigned to R. paucisqualis in the literature (Table S3).
Collection sites for the newly generated 991 samples analyzed here. Each dot may represent several collection sites. Map data: Google, DigitalGlobe. Modified using Adobe Illustrator CS5 v 15.0.2. http://www.adobe.com/products/illustrator.html.
Species delimitation analyses provided varying numbers of Operational Taxonomic Units (OTUs) among methods (Fig. 3): 129 for PTP, 95 for mPTP, 178 for GMYC, 191 for mGMYC, 175 for ABGD and 146 for RESL (Table S3). Our consensus delimitation scheme yielded 166 OTUs, including 165 OTUs for the 65 Rasborinae taxa, 2.5-fold more than by using morphological characters. The number of OTUs observed within species ranged from two for 22 species to 11 for Trigonostigma pauciperforata (Table 1). Based on the results of the species delimitation analyses, a re-examination of the original species identity associated with 105 DNA barcodes mined from BOLD and GenBank revealed 13 cases of conflicts that likely originated from mis-identifications (Table S4). These concerned the genera Boraras (four records, two species), Brevibora (two records, two species) and Rasbora (seven records, six species). Along the same line, 12 uncertain identifications were revised for the genera Rasbora (10 records, five taxa) and Trigonostigma (two records, one taxa).
The examination of the maximum K2P genetic distances for species with multiple OTUs and within OTUs revealed large differences with maximum K2P distances ranging between 0.26 and 13.64 within species and between 0.00 and 2.37 within OTUs (Table 1). This trend was largely confirmed by the distribution of the genetic distances at both species and OTUs levels (Fig. 4). At the species level, the distribution of the maximum intraspecific K2P genetic distance broadly overlap with the distribution of the K2P distances to the nearest neighbor (Fig. 4A,B, Table S5) and no barcoding gap is observed. On average, the nearest neighbor K2P genetic distances are only 3.5-fold higher than the maximum intraspecific K2P distances. Plotting genetic distance for each species provides little improvement as a substantial number of species display maximum intraspecific K2P genetic distances higher than the minimum distance to the nearest neighbor (Fig. 4C). At the OTU level, the overlap is drastically reduced peaking between 0 and 0.99 for maximum intraspecific K2P distances and ranging between 1.0 and 1.99 for the K2P distance to the nearest neighbor (Fig. 4D,E). The distribution width of the maximum intraspecific K2P distance is much more restricted for OTUs than species and fewer cases of maximum intraspecific distance higher than the minimum distance to the nearest neighbor are observed (Fig. 4F). At the OTU level, the nearest neighbor K2P genetic distances were 7.2-fold higher on average than the maximum intraspecific K2P genetic distances.
Summary of the distribution of the K2P distances. (A,D) Maximum intraspecific K2P distances; (B,E) Minimum nearest-neighbor K2P distances; (C,F) Individual plotting of maximum intraspecific K2P distances and minimum nearest neighbor K2P distances. (A–C) Distributions of K2P distances for species delimitated using morphological characters. (D–F) Distributions of K2P distances for OTUs delimitated by the 50% consensus among species delimitation methods.
Range distributions inferred from the new records generated for this study indicate that most type localities are embedded in the observed species range (Fig. 5). The degree of overlap between species range, however, largely varies among genera with little or no overlap observed for Boraras, Pectenocypris, Trigonostigma and most Rasbora species while a substantial amount of overlap is observed for Brevibora and Trigonopoma species (Fig. 5).
Maps depicting species distribution ranges as established based on the present sampling sites (black margin) and type localities (white margin) following the checklist generated for this study (Table S1). 1 Sampling sites and type localities of Boraras maculatus and B. merah. 2 Sampling sites, type localities and distribution ranges of Brevibora cheeya, B. dorsiocellata and B. exilis. 3 Type locality of Kottelatia brittani, sampling sites unknown, sequences originating from GenBank. 4 Sampling sites, type localities and distribution ranges of Pectenocypris korthausea and P. balaena. 5 Sampling sites, type localities and distribution ranges of Trigonopoma gracile, T. pauciperforatum and T. sp16. 6 Sampling sites, type localities and distribution ranges of Trigonostigma heteromorpha, T. hengeli (sampling sites outside the map); T. espei not displayed, type locality outside the map and sampling sites unknown, sequences originating from GenBank. 7 Sampling sites, type localities and distribution ranges of Rasbora argyrotaenia, R. sumatrana and R. borapetensis, multiple type localities for Rasbora argyrotaenia as detailed in16, Type locality of R. borapetensis outside the map. 8 Sampling sites, type localities and distribution ranges of Rasbora caudimaculata, R. trilineata and R. meinkeni, sampling sites of R. meinkeni unknown, sequence originating from GenBank. 9 Sampling sites, type localities and distribution ranges of Rasbora vulcanus, R. maninjau, R. subtilis and R. jacobsoni. 10 Sampling sites, type localities and distribution ranges of Rasbora tawarensis, R. paviana, R. sp. 116, R. volzii, R. tubbi, R. laticlavia and R. vaillanti, sampling sites corresponding to the type locality for Rasbora tawarensis. 11 Sampling sites, type localities and distribution ranges of Rasbora vulgaris, R. arundinata and R. chrysotaenia, type locality of Rasbora chrysotaenia located in Sumatra with no further details (Table S1). 12 Sampling sites, type localities and distribution ranges of Rasbora aprotaenia, R. elegans and R. spilotaenia. 13 Sampling sites, type localities and distribution ranges of Rasbora sarawakensis, R. rutteni, R. lacrimula, R. hubbsi, R. semilineata and R. kalbarensis. 14 Sampling sites, type localities and distribution ranges of Rasbora sp216, R. lateristriata and R. baliensis. 15 Sampling sites, type localities and distribution ranges of Rasbora beauforti, R. johannae, R. trifasciata and R. paucisqualis. 16 Sampling sites, type localities and distribution ranges of Rasbora tuberculata, R. ennealepis and R. bankanensis. 17 Sampling sites, type localities and distribution ranges of Rasbora tornieri and R. dusonensis. 18 Sampling sites, type localities and distribution ranges of Rasbora cephalotaenia, R. kottelati and R. kalochroma. 19 Sampling sites, type localities and distribution ranges of Rasbora patrickyapi and R. einthovenii. 20 Sampling sites, type localities and distribution ranges of Rasbora borneensis, R. myersi, R. rheophila. Sampling sites and type locality of R. daniconius not displayed, outside the map. Each locality may represent several sampling sites. Map data: https://maps-for-free.com/. Modified using Adobe Illustrator CS5 v 15.0.2. http://www.adobe.com/products/illustrator.html.
Discussion
This study represents the most comprehensive molecular survey conducted for the subfamily Rasborinae19,34. Our DNA barcode reference library consists of 65 Rasborinae species distributed across 7 genera and covering 78% of the Rasborinae diversity reported from Sundaland. DNA barcoding delivers reliable species-level identifications when taxa possess unique COI sequence clusters characterized by multiple private mutations. This condition was met for all the Rasborinae species examined here and no cases of retention of ancestral polymorphism were detected35. However, this clearly contrasts with multiples discrepancies observed within the set of previously published COI sequences obtained on GenBank and BOLD. About 25 percent of these records were either misidentified or associated with uncertain identifications. Such mis-identifications were expected considering the morphological uniformity within some Rasborinae genera, particularly in the genus Rasbora where multiple cases of taxonomic conflicts have been highlighted already16,36,37,38,39. Unexpectedly, most of the conflicts we detected were within the larger species of Rasbora, particularly those of the Rasbora argyrotaenia group and the R. sumatra group, and not within closely related smaller species such as members of the R. trifasciata group (Fig. 1). In facts, conflicts in species level population assignments have been previously reported for the R. argyrotaenia group in Java and Bali where R. lateristriata and R. baliensis have been confounded for decades as recently revealed by re-examination of species boundaries and distribution through DNA barcodes16. Other morphologically similar species of the Rasbora argyrotaenia group have been previously confused with R. lateristriata, such as R. elegans, R. spilotaenia and R. chrysotaenia. These species are difficult to separate due to overlapping meristic counts and coloration patterns40. Our study, however, highlights that these species have disjunct range distributions (Fig. 5) and cluster into well-differentiated mitochondrial lineages (Fig. S1, Table S3). Several of the detected misidentifications also involve species from different Rasbora species groups24 such as Rasbora dusonensis, from the R. argyrotaenia group, that has been previously mistaken for R. sumatrana from the sumatrana group and R. myersi, from the R. sumatrana group, that has been confounded with R. dusonensis from the argyrotaenia group. Despite being distantly related (Fig. S1), these species show overlapping meristic counts and similar coloration patterns with no dark spots on the body40. This result further calls for a broader assessment of the monophyly of the different Rasbora groups, previously identified by Liao24, as they are poorly supported by our study (Fig. S1).
The observed average ratio of 3.5 between intraspecific and interspecific distances is very low compared to earlier values found for the Javanese ichtyofauna, where minimum nearest neighbor genetic distances are on average 28-fold higher than the maximum intraspecific genetic distances27. This value is also very low in comparison to previous large-scale fish DNA barcode surveys41,42,43,44,45,46. This deviation can be attributed to a substantial amount of cryptic diversity revealed by our species delimitation analyses. For 61 species, delimitated on the basis of morphological characters, and validated by a match between species range distributions and type localities, we recovered a total of 166 OTUs. When accounting for this cryptic diversity the ratio between the minimum nearest neighbor and maximum intraspecific distances rose to 7.5. Earlier large scale surveys in Sundaland already pointed to substantial levels of cryptic diversity28,29,30,31,33 and it has also been demonstrated that small-size species are more sensitive to fragmentation, experience faster genetic drift and as such accumulate cryptic diversity at a faster rate than large-size species45,47. Along the same line, small-size species are more frequently confounded and lumped together, a bias that tend to inflate the proportion of hidden diversity48.
We found very high numbers of OTUs with deep genetic divergences (up to 13.64% in Trigonopoma gracile) in a number of species (ranging from 7 to 11) such as in Rasbora bankanensis, Rasbora einthovenii, Rasbora trilineata, Trigonopoma gracile and Trigonopoma pauciperforatum. These five species also display some of the widest range distributions in Sundaland with OTUs occurring in Borneo, Sumatra, Peninsular Malaysia and several small islands across the Java sea (R. bankanensis, Fig. 5(16); R. einthovenii, Fig. 5(19); R. trilineata, Fig. 5(8); T. gracile, Fig. 5(5); T. pauciperforatum, Fig. 5(4)). However, the scarcity of OTU range overlap for those species suggests ongoing population fragmentation across the species range distribution (Tables S2 and S3). This pattern is likely connected to the complex geological history of Sundaland which over the last 10 Million years was influenced by the subduction activity of the Asian and Australian plates and the resulting intense volcanic activity which produced multiple volcanic arches5. Furthermore, climatic fluctuations during the Pleistocene induced major sea levels changes leading to merging of Sundaland landmasses during glacial maxima and multiple fragmentations during glacial sea level low-stands7,8. In such dynamic landscapes, complex patterns of distribution and high lineage diversity are to be expected10. The influence of eustatic fluctuations in Sundaland is exemplified by Rasbora bankanensis, Rasbora einthovenii, Rasbora trilineata, Trigonopoma gracile and Trigonopoma pauciperforatum all of which display wide range distributions among watersheds neighboring the Java sea. Those have been repeatedly connected during glacial maxima (Fig. 5(5), 5(8), 5(16) and 5(19)). This pattern strongly contrasts with the lower genetic diversity and restricted range distribution of the species occurring in the Eastern part of Borneo such as Rasbora vaillantii (Fig. 5(10)), R. laticlavia (Fig. 5(10)), R. trifasciata (Fig. 5(15)) and R. reophila (Fig. 5(20)) or species occurring in the Western part of Sumatra such as Rasbora vulcanus (Fig. 5(9)), R. maninjau (Fig. 5(9)), R. jacobsoni (Fig. 5(9)), R. tawarensis (Fig. 5(10)); R. chrysotaenia (Fig. 5(11)) and R. arundinata (Fig. 5(11)) and species in Java and Bali such as Rasbora sp1 (Fig. 5(10)), R. sp2 (Fig. 5(14)), R. lateristriata (Fig. 5(14)) and R. baliensis (Fig. 5(14)). These parts of Borneo, Sumatra and partially Java were disconnected from the central region of Sundaland around the Java sea during the Pleistocene. This trend highlights the sensitive status of the endemic Rasborinae species in the peripheral areas of Sundaland due to their highly restricted distribution ranges. The present study also argues against translocation programs for the most widespread species, considering the high proportion of cryptic diversity, if species and OTUs identity are not determined through DNA barcodes16,31.
Conclusions
The subfamily Rasborinae is the most diverse freshwater fish group of Sundaland and therefore represents an excellent model to explore the evolutionary response of local freshwater biotas to a dynamic geological history and repeated eustatic fluctuations. Affected by taxonomic confusions for decades, the genus Rasbora has been left aside of recent large-scale molecular studies aimed at exploring the diversification of aquatic biotas in Sundaland. Our comprehensive DNA barcode reference library for the subfamily enables further evolutionary studies on the diversification of the group, in particular within the genus Rasbora, which allowed us to trace evolutionary dynamics at the local scale in Sundaland16. The contrasting patterns of molecular diversity and species range distributions between Rasborinae species inhabiting the watersheds neighboring the Java sea and the species located on the Eastern part of Borneo call for a larger assessment of their dynamics of species proliferation based on broader genomic analyses. Clearly, future studies will also have to address the systematics of the Rasborinae as no evidence supporting the monophyly of Rasbora nor the different Rasbora species groups are detected here.
Material and Methods
Sampling and collection management
Material used in the present study is the result of a collective effort to assemble a global Rasborinae DNA barcode reference library through various field sampling efforts conducted by several of the coauthors in Sundaland over the past decade. Specimens were captured using gears such as electrofishing, seine nets, cast nets and gill nets across sites that encompass the diversity of freshwater lentic and lotic habitats in Sundaland (Fig. 2). Specimens were identified following original descriptions where available, as well as monographs40,49. Species names were further validated using several online catalogs50,51. Specimens were photographed, individually labeled and voucher specimens were preserved in a 5% formalin solution. Prior to fixation a fin clip or a muscle biopsy was taken and fixed separately in a 96% ethanol solution for further genetic analyses. Both tissues and voucher specimens were deposited in the national collections at the Museum Zoologicum Bogoriense (MZB), Research Center for Biology (RCB), Indonesian Institute of Sciences (LIPI).
Assembling a checklist of the Sundaland Rasborinae
A checklist of the Rasborinae species occurring in Sundaland was assembled from available online catalogs including Fishbase51 and Eschmeyer’s Catalog of Fishes50 as detailed in Hubert et al.15. This checklist was used to estimate the taxonomic coverage of the present DNA barcoding campaign and to identify type localities for each species. The following information was included: (1) authors of the original description, (2) type locality, (3) latitude and longitude of the type locality, (4) holotype and paratypes catalog numbers, (5) distribution in Sundaland. This information is available as online Supplementary Material (Table S1).
Sequencing and international repositories
Genomic DNA was extracted using a Qiagen DNeasy 96 tissue extraction kit following manufacturer’s specifications. A 651-bp segment from the 5′ region of the cytochrome oxidase I gene (COI) was amplified using primers cocktails C_FishF1t1/C_FishR1t1 including M13 tails52. PCR amplifications were done on a Veriti 96-well Fast (ABI-AppliedBiosystems) thermocycler with a final volume of 10.0 μl containing 5.0 μl Buffer 2× 3.3 μl ultrapure water, 1.0 μl each primer (10 μM), 0.2 μl enzyme Phire Hot Start II DNA polymerase (5U) and 0.5 μl of DNA template (~50 ng). Amplifications were conducted as followed: initial denaturation at 98 °C for 5 min followed by 30 cycles denaturation at 98 °C for 5 s, annealing at 56 °C for 20 s and extension at 72 °C for 30 s, followed by a final extension step at 72 °C for 5 min. The PCR products were purified with ExoSap-IT (USB Corporation, Cleveland, OH, USA) and sequenced in both directions. Sequencing reactions were performed using the “BigDye Terminator v3.1 Cycle Sequencing Ready Reaction” and sequencing was performed on the automatic sequencer ABI 3130 DNA Analyzer (Applied Biosystems). DNA barcodes obtained at the Naturhistorisches Museum Bern were generated as previously described in Conte-Grand et al.33.
The sequences and associated information were deposited on BOLD53 and are available in the data set DS-BIFRA (Table S2, dx.doi.org/10.5883/DS-BIFRA). DNA sequences were submitted to GenBank (accession numbers are accessible directly at the individual records in BOLD). An additional set of 106 Rasborinae COI sequences were downloaded from GenBank (Table S3).
Genetic distances and species delimitation
Kimura 2-parameter (K2P)54 pairwise genetic distances were calculated using the R package Ape 4.155. Maximum intraspecific and nearest neighbor genetic distances were calculated from the matrice of pairwise K2P genetic distances using the R package Spider 1.556. We checked for the presence of a barcoding gap, i.e. the lack of overlap between the distributions of the maximum intraspecific and the nearest neighbor genetic distances57, by plotting both distances and examining their relationships on an individual basis instead of comparing both distributions independently58. A neighbor-joining (NJ) tree was built based on K2P distances using PAUP 4.0a59 in order to visually inspect genetic distances and DNA barcode clusters (Fig. S1). This NJ tree was rooted using Sundadanio retiarius.
Several alternative methods have been proposed for delimitating molecular lineages60,61,62,63. Each of these methods have pitfalls, particularly when it comes to singletons (i.e. delimitated lineages represented by a single sequence) and a combination of different approaches is increasingly used to overcome potential pitfalls arising from uneven sampling16,43,64,65,66. We used four different sequence-based methods of species delimitation. For the sake of clarity, we refer to species identified based on morphological characters as species while species delimited using DNA sequences are referred to as Operational Taxonomic Unit (OTU)67,68,69. OTUs were delimitated using the following algorithms: (1) Refined Single Linkage (RESL) as implemented in BOLD and used to generate Barcode Index Numbers (BIN)62, (2) Automatic Barcode Gap Discovery (ABGD)61, (3) Poisson Tree Process (PTP) in its multiple rates version (mPTP) as implemented in the stand-alone software mptp_0.2.363,70, (4) General Mixed Yule-Coalescent (GMYC) in its multiple rate version (mGMYC) as implemented in the R package Splits 1.0–1971. RESL and ABGD used DNA alignments as input files while a ML tree was used for mPTP and a Bayesian Chronogram based on a strict-clock model using a 1.2% of genetic distance per million year72 for mGMYC. The mPTP algorithm uses a phylogenetic tree as an input file, thus, a maximum likelihood (ML) tree was first reconstructed using RAxML73 based on a GTR + Γ substitution model. Then, an ultrametric and fully resolved tree was reconstructed using the Bayesian approach implemented in BEAST 2.4.874. Two Markov chains of 50 millions each were ran independently using the Yule pure birth model tree prior, a strict-clock model and a GTR + I + Γ substitution model. Trees were sampled every 10,000 states after an initial burnin period of 10 millions. Both runs were combined using LogCombiner 2.4.8 and the maximum credibility tree was constructed using TreeAnnotator 2.4.774. Identical haplotypes were pruned for further species delimitation analyses.
References
Myers, N., Mittermeier, R. A., Mittermeier, C. G., da Fonseca, G. A. B. & Kent, J. Biodiversity hotspots for conservation priorities. Nature 403, 853–858 (2000).
Lamoreux, J. F. et al. Global tests of biodiversity concordance and the importance of endemism. Nature 440, 212–214 (2006).
Hoffman, M. et al. The impact of Conservation on the status of the world’s vertebrates. Science (80-.) 330, 1503–1509 (2010).
Schipper, J. et al. The status of the world’s land and marine mammals: diversity, threat, and knowledge. Science (80-.) 322, 225–230 (2008).
Lohman, K. et al. Biogeography of the Indo-Australian archipelago. Annu. Rev. Ecol. Evol. Syst. 42, 205–226 (2011).
Hall, R. Late Jurassic-Cenozoic reconstructions of the Indonesian region and the Indian ocean. Tectonophysics 570–571, 1–41 (2012).
Woodruff, D. S. Biogeography and conservation in Southeast asia: how 2.7 million years of repeated environmental fluctuations affect today’s patterns and the future of the remaining refugium-phase biodiversity. Biodivers. Conserv. 19, 919–941 (2010).
Voris, H. K. Maps of Pleistocene sea levels in Southeast Asia: shorelines, river systems and time durations. J. Biogeogr. 27, 1153–1167 (2000).
De Bruyn, M. et al. Borneo and Indochina are major evolutionary hotspots for Southeast Asian biodiversity. Syst. Biol. 63, 879–901 (2014).
De Bruyn, M. et al. Paleo-drainage basin connectivity predicts evolutionary relationships across three Southeast Asian biodiversity hotspots. Syst. Biol 62, 398–410 (2013).
O’Connell, K. A. et al. Within-island diversification underlies parachuting frog (Rhacophorus) species accumulation on the Sunda shelf. J. Biogeogr. 45, 929–940 (2018).
O’Connell, K. A. et al. Diversification of bent-toed geckos (Cyrtodactylus) on Sumatra and west Java. Mol. Phylogenet. Evol. 134, 1–11 (2019).
Hendriks, K. P., Alciatore, G., Schilthuizen, M. & Etienne, R. S. Phylogeography of Bornean land snails suggests long-distance dispersal as a cause of endemism. J. Biogeogr. (2019).
Dong, J. et al. Biogeographic patterns and diversification dynamics of the genus Cardiodactylus Saussure (Orthoptera, Grylloidea, Eneopterinae) in Southeast Asia. Mol. Phylogenet. Evol. 129, 1–14 (2018).
Hubert, N. et al. DNA barcoding Indonesian freshwater fishes: challenges and prospects. DNA Barcodes 3, 144–169 (2015).
Hubert, N. et al. Revisiting species boundaries and distribution ranges of Nemacheilus spp. (Cypriniformes: Nemacheildae) and Rasbora spp. (Cypriniformes: Cyprinidae) in Java, Bali and Lombok through DNA barcodes: implications for conservation in a biodiversity hotspot. Conserv. Genet. 20, 517–529 (2019).
Keith, P. et al. Schismatogobius (Gobiidae) from Indonesia, with description of four new species. Cybium 41, 195–211 (2017).
Conway, K. W., Hirt, M. V, Yang, L., Mayden, R. L. & Simons, A. M. Conway, K. W., Hirt, M. V., Yang, L., Mayden, R. L., & Simons, A. M. Cypriniformes: Systematics & Paleontology: Festschrift in honor of G. Arratia. In Origin and Phylogenetic Interrelationships of Teleosts 295–316 (2010).
Tang, K. et al. Systematics of the subfamily Danioniae (Teleostei: Cypriniformes: Cyprinidae). Mol. Phylogenet. Evol. 57, 189–214 (2010).
Stout, C. C., Tan, M., Lemmon, A. R., Lemmon, E. M. & Armbruster, J. W. Resolving Cypriniformes relationships using an anchored enrichment approach. BMC Evol. Biol. 16, 244 (2016).
Hirt, M. V. et al. Effects of gene choice, base composition and rate heterogeneity on inference and estimates of divergence times in cypriniform fishes. Biol. J. Linn. Soc 121, 319–339 (2017).
Tan, M. & Armbruster, J. W. Phylogenetic classification of extant genera of fishes of the order Cypriniformes (Teleostei: Ostariophysi). Zootaxa 4476, 6–39 (2018).
Brittan, M. R. A revision of the Indo-Malayan frash-water fish genus Rasbora. Monogr. Inst. Sci. Tech. Manila 3, 3–pls (1954).
Liao, T. Y., Kullander, S. O. & Fang, F. Phylogenetic analysis of the genus Rasbora (Teleostei: Cyprinidae). Zool. Scr 39, 155–176 (2010).
Kottelat, M. & Vidthayanon, C. Boraras micros, a new genus and species of minute freshwater fish from Thailand (Teleostei: Cyprinidae). Ichthyol. Explor. Freshwaters 4, 161–176 (1993).
Kottelat, M. & Witte, K.-E. Two new species of Microrasbora from Thailand and Myanmar, with two new generic names for small Southeast Asian cyprinid fishes (Teleostei: Cyprinidae). J. South Asian Nat. Hist 4, 49–56 (1999).
Dahruddin, H. et al. Revisiting the ichthyodiversity of Java and Bali through DNA barcodes: Taxonomic coverage, identification accuracy, cryptic diversity and identification of exotic species. Mol. Ecol. Resour. 17, 288–299 (2017).
Nurul Farhana, S. et al. Exploring hidden diversity in Southeast Asia’s Dermogenys spp. (Beloniformes: Zenarchopteridae) through DNA barcoding. Sci. Rep 8, 10787 (2018).
Beck, S. et al. Plio-Pleistocene phylogeography of the Southeast Asian Blue Panchax killifish, Aplocheilus panchax. PLoSONE 12, e0179557 (2017).
Lim, H.-C., Abidin, M. Z., Pulungan, C. P., De Bruyn, M. & Mohd Nor, S. A. DNA barcoding reveals high cryptic diversity of freshwater halfbeak genus Hemirhamphodon from Sundaland. PLoSONE 11, e0163596 (2016).
Hutama, A. et al. Identifying spatially concordant evolutionary significant units across multiple species through DNA barcodes: Application to the conservation genetics of the freshwater fishes of Java and Bali. Glob. Ecol. Conserv 12, 170–187 (2017).
Nguyen, T. T. T., Na-Nakorn, U., Sukmanomon, S. & ZiMing, C. A study on phylogeny and biogeography of mahseer species (Pisces: Cyprinidae) using sequences of three mitochondrial DNA gene regions. Mol. Phylogenet. Evol. 48, 1223–1231 (2008).
Conte-Grand, C. et al. Barcoding snakeheads (Teleostei, Channidae) revisited: Discovering greater species diversity and resolving perpetuated taxonomic confusions. PLoS One 12, e0184017 (2017).
Collins, R. A. et al. Barcoding and border biosecurity: identifying cyprinid fishes in the aquarium trade. PLoSONE 7, e28381 (2012).
Funk, D. J. & Omland, K. E. Species-level paraphyly and polyphyly: frequency, causes and consequences, with insights from animal mitochondrial DNA. Annu. Rev. Ecol. Syst. 34, 397–423 (2003).
Siebert, D. J. The identities of Rasbora paucisqualis Ahl in Schreitmuller, 1935, and Rasbora bankanensis (Bleeker, 1853), with the designation of a lectotype for R. paucisqualis (Teleostei: Cyprinidae). Raffles Bull. Zool. 45, 29–37 (1997).
Kottelat, M. Rasbora rheophila, a new species of fish from northern Borneo(Teleostei: Cyprinidae). Rev. Suisse Zool 119, 77–87 (2012).
Ng, H. H. & Kottelat, M. The identity of the cyprinid fishes Rasbora dusonensis and R. tornieri (Teleostei: Cyprinidae). Zootaxa 3635, 62–70 (2013).
Muchlisin, Z. A., Fadli, N. & Siti-Azizah, M. N. Genetic variation and taxonomy of Rasbora group (Cyprinidae) from Lake Laut Tawar, Indonesia. J. Ichthyol 52, 284–290 (2012).
Kottelat, M., Whitten, A. J., Kartikasari, S. R. & Wirjoatmodjo, S. Freshwater Fishes of Western Indonesia and Sulawesi . (Periplus editions, 1993).
Hubert, N. et al. Identifying Canadian freshwater fishes through DNA barcodes. PLoS One 3, e2490 (2008).
April, J., Mayden, L. R., Hanner, R. H. & Bernatchez, L. Genetic calibration of species diversity among North America’s freshwater fishes. Proc. Natl. Acad. Sci. USA 108, 10602–10607 (2011).
Shen, Y. et al. DNA barcoding the ichthyofauna of the Yangtze River: insights from the molecular inventory of a mega-diverse temperate fauna. Mol. Ecol. Resour. 19, 1278–1291 (2019).
Hubert, N. et al. Cryptic diversity in indo-pacific coral-reef fishes revealed by DNA-barcoding provides new support to the centre-of-overlap hypothesis. PLoS One 7, e28987 (2012).
Hubert, N. et al. Geography and life history traits account for the accumulation of cryptic diversity among Indo-West Pacific coral reef fishes. Mar. Ecol. Prog. Ser. 583, 179–193 (2017).
Pereira, L. H. G., Hanner, R., Foresti, F. & Oliveira, C. Can DNA barcoding accurately discriminate megadiverse Neotropical freshwater fish fauna? BMC Genet. 14, 20 (2013).
April, J., Hanner, R., Mayden, R. L. & Bernatchez, L. Metabolic rate and climatic fluctuations shape continental wide pattern of genetic divergence and biodiversity in fishes. PLoSONE 8, e70296 (2013).
Kottelat, M., Britz, R., Tan, H. H. & Witte, K.-E. Paedocypris, a new genus of Southeast Asian cyprinid fish with a remarkable sexual dimorphism, comprises the world’s smallest vertebrate. Proc. R. Soc. London, B 273, 895–899 (2006).
Kottelat, M. The fishes of the inland waters of Southeast Asia: A catalog and core bibliography of the fishes known to occur in freshwaters, mangroves and estuaries. Raffles Bull. Zool Supplement 27, 1–663 (2013).
Eschmeyer, W. N., Fricke, R. & van der Laan, R. Catalog of fishes electronic version. (2018).
Froese, R. & Pauly, D. FishBase. Available at, http://www.fishbase.org (2014).
Ivanova, N. V., Zemlak, T. S., Hanner, R. H. & Hebert, P. D. N. Universal primers cocktails for fish DNA barcoding. Mol. Ecol. Notes 7, 544–548 (2007).
Ratnasingham, S. & Hebert, P. D. N. BOLD: The Barcode of Life Data System, www.barcodinglife.org. Mol. Ecol. Notes 7, 355–364 (2007).
Kimura, M. A Simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide-sequences. J. Mol. Evol. 16, 111–120 (1980).
Paradis, E., Claude, J. & Strimmer, K. E. APE: Analyses of Phylogenetics and Evolution in R language. Bioinformatics 20, 289–290 (2004).
Brown, S. D. J. et al. Spider: An R package for the analysis of species identity and evolution, with particular reference to DNA barcoding. Mol. Ecol. Resour 12, 562–565 (2012).
Meyer, C. & Paulay, G. DNA barcoding: Error rates based on comprehensive sampling. Plos 3, 2229–2238 (2005).
Blagoev, G. A. et al. Untangling taxonomy: A DNA barcode reference library for Canadian spiders. Mol. Ecol. Resour. 16, 325–341 (2015).
Swofford, D. L. Version 4.0 b10. PAUP*. Phylogenetic Anal. Using Parsimony (*Other Methods) (2001).
Pons, J. et al. Sequence-based species delimitation for the DNA taxonomy of undescribed insects. Syst. Biol 55, 595–606 (2006).
Puillandre, N., Lambert, A., Brouillet, S. & Achaz, G. ABGD, Automatic Barcode Gap Discovery for primary species delimitation. Mol. Ecol 21, 1864–1877 (2012).
Ratnasingham, S. & Hebert, P. D. N. A DNA-based registry for all animal species: the barcode index number (BIN) system. PLoSONE 8, e66213 (2013).
Zhang, J., Kapli, P., Pavlidis, P. & Stamatakis, A. A general species delimitation method with applications to phylogenetic placements. Bioinformatics 29, 2869–2876 (2013).
Kekkonen, M. & Hebert, P. D. N. DNA barcode-based delineation of putative species: Efficient start for taxonomic workflows. Mol. Ecol. Resour. 14, 706–715 (2014).
Kekkonen, M., Mutanen, M., Kaila, L., Nieminen, M. & Hebert, P. D. N. Delineating species with DNA Barcodes: A case of taxon dependent method performance in moths. PLoS One 10, e0122481 (2015).
Blair, C. & Bryson, J. R. W. Cryptic diversity and discordance in single-locus species delimitation methods within horned lizards (Phrynosomatidae: Phrynosoma). Mol. Ecol. Resour. 17, 1168–1182 (2017).
Avise, J. C. Molecular Markers, Natural History and Evolution. (1989).
Moritz, C. Defining ‘Evolutionary significant units’ for conservation. Trends Ecol. Evol. 9, 373–375 (1994).
Vogler, A. P. & DeSalle, R. Diagnosing units of conservation management. Conserv. Biol. 6, 170–178 (1994).
Kapli, P. et al. Multi-rate Poisson Tree Processes for single-locus species delimitation under Maximum Likelihood and Markov Chain Monte Carlo. Bioinformatics 33, 1630–1638 (2017).
Fujisawa, T. & Barraclough, T. G. Delimiting species using single-locus data and the generalized mixed Yule coalescent approach: A revised method and evaluation on simulated data sets. Syst. Biol 62, 707–724 (2013).
Bermingham, E., McCafferty, S. & Martin, A. P. Fish biogeography and molecular clocks: Perspectives from the Panamanian Isthmus. In Molecular Systematics of Fishes (eds. Kocher, T. D. & Stepien, C. A.) 113–128 (CA Academic Press, 1997).
Stamatakis, A. RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Bouckaert, R. R. et al. BEAST 2: A software platform for Bayesian evolutionary analysis. PLoS Comput. Biol. 10, e1003537 (2014).
Acknowledgements
The authors wish to thank Siti Nuramaliati Prijono, Bambang Sunarko, Witjaksono, Mohammad Irham, Marlina Adriyani, Ruliyana Susanti, Rosichon Ubaidillah, the late Renny K. Hadiaty, Hari Sutrisno and Cahyo Rahmadi at Research Centre for Biology (RCB-LIPI) in Indonesia; Edmond Dounias, Jean-Paul Toutain, Robert Arfi and Valérie Verdier from the ‘Institut de Recherche pour le Développement’; Joel Le Bail and Nicolas Gascoin at the French embassy in Jakarta for their continuous support. We also would like to thank Eleanor Adamson, Hendry Budianto, Tob Chann Aun, Pak Epang, Herman Ganatpathy, Sébastien Lavoué, Michael Lo, Hendry Michael, Joshua Siow, Heok Hui Tan, Elango Velautham, Norsham S. Yaakob, and Denis Yong for their help in the field. We are also particularly thankful to Sumanta at IRD Jakarta for his help during the field sampling in Indonesia. Part of the present study was funded by the Institut de Recherche pour le Développement (UMR226 ISE-M and IRD through incentive funds) to N.H.), the MNHN (UMR BOREA) to P.K., the French Ichthyological Society (SFI) to P.K., the Foundation de France to P.K., the French embassy in Jakarta to N.H., the Natural Environmental Research Council (NERC, NE/F003749/1) to L.R. and Ralf Britz; National Geographic (8509-08) to L.R. and North of England Zoological Society‐Chester Zoo to L.R. The present study and all associated methods were carried out in accordance with relevant guidelines and regulation of the Indonesian Ministry of Research and Technology (Indonesia), the Economic Planning Unit, Prime Minister’s Department (Malaysia), the Forest Department Sarawak (Malaysia), the Vietnam National Museum of Nature (Vietnam) and the Inland Fisheries Research and Development Institute (Cambodia). Field sampling in Indonesia was conducted according to the research permits 097/SIP/FRP/SM/IV/2014 for Philippe Keith, 60/EXT/SIP/FRP/SM/XI/2014 for Frédéric Busson, 41/EXT/SIP/FRP/SM/VIII/2014 for Nicolas Hubert, 200/E5/E5.4/SIP/2019 for Erwan Delrieu-Trottin and, 1/TKPIPA/FRP/ SM/I/2011 and 3/TKPIPA/FRP/SM/III/2012 for Lukas Rüber. The Fieldwork in Peninsular Malaysia and Sarawak was conducted under permits issued by the Economic Planning Unit, Prime Minister’s Department, Malaysia (UPE 40/200/19/2417 and UPE 40/200/19/2534) and the Forest Department Sarawak (NCCD.970.4.4[V]-43) and were obtained with the help of Norsham S. Yaakob (Forest Research Institute Malaysia, Kepong, Kuala Lumpur, Malaysia). Luong Van Hao and Pham Van Luc (Vietnam National Museum of Nature) helped with arranging research permits in Vietnam and So Nam (Inland Fisheries Research and Development Institute, IFReDI) helped with arranging research permits in Cambodia. All experimental protocols were approved by the Indonesian Ministry of Research and Technology (Indonesia), the Indonesian Institute of Sciences (Indonesia), the Forest Department Sarawak (Malaysia), Economic Planning Unit of the Prime Minister’s Department (Malaysia), the Vietnam National Museum of Nature (Vietnam) and the Inland Fisheries Research and Development Institute (Cambodia). It is a great pleasure to thank Soraya Villalba for generating the DNA barcodes at the Naturhistorisches Museum Bern. Sequence analysis was aided by funding through the Canada First Research Excellence Fund as part of the University of Guelph Food from Thought program. We thank Paul Hebert, Alex Borisenko and Evgeny Zakharov as well as BOLD and CCDB staff at the Centre for Biodiversity Genomics, University of Guelph for their valuable support. This publication has the ISEM number 2019-293-SUD.
Author information
Authors and Affiliations
Contributions
L.R. and N.H. designed the study. A.S., T.S., H.D., R.R., R.E., A.W., K.K., F.B., S.S., U.N., E.D., I.V.U., Z.A.M., D.W., P.K., L.R. and N.H. conducted the field sampling. A.S., E.D.T., T.S., H.D., A.W., L.R. and N.H. performed the morphological identifications. H.D., S.S., U.N., F.B. and N.H. curated the specimen collection. A.S., E.D.T., H.D., M.S.A.Z., Y.F., I.V.U., R.H., D.S., L.R. and N.H. conducted the laboratory work. A.S., E.D.T., H.D., F.B., N.H., R.H. and D.S. curated the DNA barcode records in BOLD. A.S., E.D.T., H.D., J.F.A., L.R. and N.H. analyzed the data. A.S., E.D.T., H.D., J.F.A., D.S., L.R. and N.H. wrote the initial manuscript and all authors commented and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sholihah, A., Delrieu-Trottin, E., Sukmono, T. et al. Disentangling the taxonomy of the subfamily Rasborinae (Cypriniformes, Danionidae) in Sundaland using DNA barcodes. Sci Rep 10, 2818 (2020). https://doi.org/10.1038/s41598-020-59544-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-59544-9
This article is cited by
-
Molecular identification and phylogenetic relationship of fishes belonging to the Family Danionidae from Brahmaputra Basin, Assam, Northeast India
Molecular Biology Reports (2024)
-
Reassessing fish diversity of Penang Island’s freshwaters (northwest Peninsular Malaysia) through a molecular approach raises questions on its conservation status
Biodiversity and Conservation (2022)
-
Molecular phylogeny and phylogeography of the freshwater-fish genus Pethia (Teleostei: Cyprinidae) in Sri Lanka
BMC Ecology and Evolution (2021)
-
Genetic diversity and morphological stasis in the Ceylon Snakehead, Channa orientalis (Teleostei: Channidae)
Ichthyological Research (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.