Abstract
The Genji firefly, Luciola cruciata, is widely distributed throughout the major Japanese islands (Honshu, Shikoku, and Kyushu) and distinguished into two ecological types on the basis of the flash interval of the mate-seeking males (4-sec slow-flash or 2-sec fast-flash intervals). The boundary of the ecological types corresponds to the Fossa Magna, a great rupture zone that separates eastern and western Japan. Although the degree of genetic differentiation of the two types has been evaluated using allozyme and mitochondrial DNA sequence data, it has not been evaluated using genome-wide data. Based on the genome-wide data obtained using single-end restriction-site-associated DNA (RAD-Seq), principal component, gene-level phylogenetic tree, admixture, and Wright’s fixation index analyses, we identified three phylogenetic groups in L. cruciata: East-Honshu, West-Honshu, and Kyushu. This grouping corresponds to the ecological types: East-Honshu to the slow-flash type and West-Honshu and Kyushu to the fast-flash type. Although introgression was exceptionally observed around adjacent or artificially transplanted areas, gene flow among the groups was almost absent in the natural populations. The phylogenetic tree under the coalescent model also evaluated differentiation among the East-Honshu, West-Honshu and Kyushu groups. Furthermore, because the distribution patterns of the three groups are consistent with the geological history of Japanese islands, a vicariant speciation scenario of L. cruciata is concluded. In addition, we identified genetic markers that can be used to distinguish the three genetic groups for genetic management of firefly transplantation in nature conservation and regeneration.
Similar content being viewed by others
Introduction
The fantastic light of fireflies has been attractive to nearly all peoples. This attraction is particularly strong in Japan, where the firefly has appeared in the oldest Japanese history book, “Nihon Shoki,” compiled during the Nara period of the 8th century.
Light emission in adult fireflies is a mating adaptation1,2,3, and species-specific and mating-related flash signals play a major role in reproductive isolation4. Therefore, it is presumed that the flash signal differentiation and speciation processes are closely correlated. The Genji firefly, Luciola cruciata, is a species that shows synchronous flashes, and its flash communication and mating behaviour have been intensively studied5,6. The communication system is described in several phases: after sunset, the males begin to fly and seek females with synchronous flashing lights; the females emit single-pulsed flashes of light (not synchronized) on grasses; when a male finds the female’s flashed light, the male approaches the female; the male emits flashes with various patterns while approaching and walking around the female; thereafter, they copulate5,6,7.
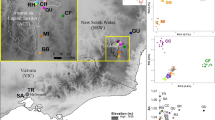
This species is distributed throughout the three major islands of Japan, i.e., Honshu, Shikoku, and Kyushu, and two ecological types have been recognized based on the synchronous flashing behaviours of males (Fig. 1)7. In the slow-flash type, mate-seeking males have a flash interval of approximately 4 sec while in the fast-flash type, they have a flash interval of approximately 2 sec. Furthermore, females of the fast-flash type aggregate at one site for oviposition, whereas those of the slow-flash type do not gregariously assemble. The slow-flash type is distributed in the eastern areas of Japan (East Japan), whereas the fast-flash type firefly occurs in the western areas (West Japan). The habitation boundary corresponds to a great rupture zone termed the Fossa Magna, which divides Honshu island into eastern (East-Honshu) and western (West-Honshu) areas. However, the two types are not morphologically distinguished.
Locality map of L. cruciata used in this study. L. cruciata is distributed on the three major islands of Japan, i.e., Honshu, Shikoku, and Kyushu. Honshu is divided into eastern and western areas geologically by the great rupture zone, Fossa Magna, shown in grey shading on the map. Collection sites are indicated by the sample (accession) numbers. The superscript letter “a” following the number indicates a purchased sample from distributors, and it can only be identified as an approximate location in Nos. 42, 43, 46, and 47 (Table 1). The sample number in the parentheses is the technical replicate of the previous number (Table 1). The slow-flash type firefly inhabits the eastern area of Honshu (red area), and the fast-flash type the western area of Honshu (blue area) as well as Shikoku and Kyushu (green area).
To evaluate the degree of genetic differentiation between the two ecological types, allozyme analysis was performed8. The phylogenetic tree showed two major clades. The first was the East-Japan (consistent with the East-Honshu areas) populations, and the other was the West-Japan (including the West-Honshu, Shikoku, and Kyushu areas) populations; these populations corresponded to the slow- and fast-flash types, respectively. The two types were genetically differentiated at the subspecies level (Nei’s genetic distance by allozymes was 0.09)9.
Furthermore, the mitochondrial cytochrome oxidase (CO) II gene was also analysed10. The phylogenetic tree showed three major clades (I, II, and III). Clade I was composed of haplotypes distributed in East-Honshu, Clade II was in West-Honshu and Shikoku, and the Clade III was in Kyushu. Namely, Clade I corresponded to the slow-flash type, and Clades II and III corresponded to the fast-flash type. Additionally, the phylogenetic tree showed a clade branching; Clade III branched first, followed by branching of Clades II and I. Therefore, the slow-flash type character of Clade I was considered derivative. Moreover, as the branching order nearly corresponded to the formation of the Japanese islands11,12, a vicariant scenario concluded that the genetic and ecological diversity in L. cruciata would have arisen from phylogenetic separations subsequent to the Japanese island formation process.
In addition to the case of L. cruciata, other examples of unique genetic diversity of insects due to division of habitation areas by the formation of the Japanese islands with the Fossa Magna are also known13,14,15,16,17. Including them, however, these phylogeographic studies were performed using a small number of genes at the mitochondrial and nuclear DNA level but not using genome-wide DNA data. For genome-wide polymorphism analyses, restriction-site-associated DNA sequencing (RAD-Seq) is a very efficient and cost-effective method18. It produces a reduced representation of a genome via high-throughput sequencing for thousands of genomic regions flanked by targeted restriction sites and can provide a huge number of genome-wide polymorphisms across many individuals. The RAD-Seq approach can be used for population genetic studies in wild specimens and in the American firefly19,20. In this study, we used genome-wide polymorphism data obtained using the traditional single-end RAD-Seq techniques with the restriction enzyme EcoRI, which recognizes a 6-base pair sequence to evaluate the population structure and phylogeography of L. cruciata throughout its entire distribution range. Furthermore, we identified genetic markers that can be used to distinguish L. cruciata populations.
Methods
Sample collection and genomic DNA extraction
Adult L. cruciata firefly samples used in this study were 39 individuals collected from 17 localities and 10 individuals purchased by distributors from 5 localities (Table 1); the sample Nos. and localities are mapped in Fig. 1. Most of the samples were males; only two samples, Nos. 26 and 38, were females. Sample Nos. 50, 51, and 52 were technical replicates of Nos. 8, 16, and 22, respectively. The samples were stored in ethanol at room temperature or at −20 °C. Before extraction of genomic DNA, individual samples were air dried for 30 minutes, centrifuged for 1 minute, and air dried for an additional 1 hour to completely remove the ethanol. Then, the entire body excluding the wings was homogenized by freeze grinding using a ball mill. Genomic DNA extraction was performed using the Blood & Cell Culture DNA Mini Kit (Qiagen, Japan) according to the manufacturer’s instructions. The concentration of the isolated genomic DNA was assessed using a QuantiFluor ONE dsDNA System (Promega, Japan).
RAD-Seq analysis
The library for RAD-Seq was created according to the established method21. EcoRI was used as a single restriction enzyme. The library was sequenced by Macrogen (Seoul, South Korea) with 50 bp single-end reads in one lane of an Illumina HiSeq 2000 (Illumina, San Diego, CA, USA). Sequence data were deposited at the DDBJ Sequence Read Archive (DRA) with accession No. DRA007766. Because all firefly samples that were analysed in this study are potentially cross-compatible species, the Hardy–Weinberg principle was considered. The Stacks program is useful for analysing the RAD-Seq data of closely related samples by applying the Hardy–Weinberg principle22,23. Therefore, we analysed the RAD-Seq data using the Stacks program. The data were quality filtered using the process_radtags program in the Stacks package (v. 1.46). In addition, adapter sequences and low‐quality ends were trimmed using cutadapt (v. 1.14) and trimmomatic (v. 0.36), respectively24,25. The numbers of quality-filtered reads are shown in Supplementary Table S1. The data were aligned with the reference genome of L. cruciata (NCBI/DDBJ BioProject number: PRJDB7197) by bowtie2 with the -q–no-unal -L 15 options26. The data of the reference genome were used only for information on the nuclear sequences and not used for mitochondrial DNA. Aligned data were analysed using the ref_map.pl script (v. 1.47) of the Stacks package with default parameters.
Principal component analysis (PCA) and multidimensional scaling (MDS) analysis
The populations program of the Stacks package was used to create a variant call format (VCF) file with the -p 26 option27. This option indicates that the minimum number of populations in which a locus must be present to process that locus is 26. In this analysis, because of a reduction in the effect of accidental reads and increased reproducibility, the -p option was set to half of the total 52 samples within the cluster, and locus common to more than one-half of the samples were extracted. The mean sequencing depth for each sample was calculated using vcftools. Principal component analysis (PCA) and multidimensional scaling (MDS) analyses were conducted based on this VCF file using the SNPrelate program (v.1.18.1)28.
Gene-level phylogenetic tree analysis
A multiple alignment was created by aligning the data to the reference genome. The Stacks package populations program was used to create multiple alignments within the cluster using the options -phylip and -phylip_var. The phylogenetic tree based on the maximum likelihood (ML) method was constructed using the RAxML program (version 8.2.12)29 (-f = a, -x = 12,345, -p = 12,345, -N (bootstrap value) = 1,000, and -m = GTRGAMMA). Another phylogenetic tree based on Bayesian inference was constructed using the MrBayes program (version 3.2.6)30 (lset nst = 6, rates = invgamma, mcmc ngen = 500,000, samplefreq = 1,000, nchains = 4, and savebrlens = yes). In each analysis, the midpoint was used as a root.
Phylogenetic tree analysis under the coalescent model
Phylogenetic tree analysis was performed using SVDquartets integrated in the software PAUP ver. 4.0a166 using multiple alignments31,32,33. The parameters used for SVDquartets were as follows (Quartet evaluation; Evaluate all possible quartets, Tree inference; Select trees using the QFM quartet assembly, Tree model; Multi-species coalescent; Handling of ambiguities; and Distribute). The number of bootstrap analyses conducted was 1,000 replicates. The same position used in the RAxML and MrBayes analyses was used as a root.
Admixture analysis
For the admixture analysis, the Stacks population program was used to create a PLINK file with the -p 26 option34. The admixture analysis was conducted based on this PLINK file using ADMIXTURE (v. 1.3.0) under default conditions35.
Wright’s fixation indices
To compare the genetic diversity of firefly accessions within and among populations, pairwise Fst values were calculated using the Stacks populations program with the –fstats option.
Genetic marker search and distribution of SNPs
SNPs for grouping L. cruciata into two (East- and West-Japan types) or three (East-Honshu, West-Honshu, and Kyushu types) populations in Japan were determined by running two types of our original Ruby scripts. The list of all loci including SNPs was output by the allsnps.rb program using the VCF file. Towards the output file of this calculation, a second program popdiff.rb was run to group the populations. Custom scripts are available on the GitHub web site (https://github.com/yoshinobuhayashi/RADseq_Firefly_2019).
Mitochondrial CO II gene analysis
The mitochondrial CO II gene was amplified from the genomic DNA using PCR primers (TL2-J-3037: ATGGCAGATTAGTGCAATGG, TK-N-3785: GTTTAAGAGACCAGTACTTG)36. Sequence analysis was performed using the Sanger method with BigDye 3.1, and a 3500xL Genetic Analyzer (Applied Biosystems) was used as a DNA sequencer. Sequence data were deposited at BioProject accession No. PRJDB8812. The haplotype tree construction was conducted using MEGA X (ver. 10.0.5)37. The nucleotide substitution model best fitted for maximum likelihood (ML) analyses was selected; the ML tree was constructed using the selected parameters. Bootstrap analysis was performed with 1,000 replicates. The closely related species Luciola owadai was used as an outgroup.
Results
RAD-Seq loci
A comparison of the genome-wide SNPs of the firefly L. cruciata, consisting of 52 accessions including 49 individual samples and three technical replicates (Nos. 50–52), was performed. From over 224 million raw single-end reads obtained from single-digest RAD-Seq using the EcoRI, 187 million reads were retained after quality filtering with an average of 3.6 million reads per pooled sample (Supplementary Table S1). This number of reads corresponds to 1.12% of the predicted targeted restriction sites on the firefly genome. By catalogue building and SNP calling on reference genome data using Stacks, 435,165 RAD loci that were shared between two or more samples (p-option = 26 of the cut-off value) and 19 loci found in all fifty-two samples (p-option = 52) were obtained. These data were used for population genetics and phylogenetic analyses.
PCA and MDS analysis
To explore the relationship among the 52 accessions and examine the population structure, PCA was performed; graphs were created using the first three components (Fig. 2). The contribution rate of the three components reached up to 25.2% of the analysed data, broken down as follows: first principle comportment (PC1) = 14.7% of the data, PC2 = 6.4%, and PC3 = 4.1%. Each plot of the accessions was drawn using three colours, i.e., green, blue, and red, based on their collected or purchased area, i.e., Kyushu, West-Honshu, and East-Honshu, respectively. On the graph, firefly accessions were first divided into two groups (the cluster plotted in red vs. the other two colours) on the basis of the values of PC1. The analysis of PC2 indicated a second separation into two other groups (green vs. the other two colours). Consequently, the two-dimensional plots of PC1 vs. PC2 showed a clear separation of three large clusters, and all 52 accessions were clustered into approximately three groups according to the aforementioned habitation areas (Fig. 2A). There were, however, three exceptions in that accession No. 4 (red) was in the blue West-Honshu group, No. 9 (blue) in the red East-Honshu group, and No. 14 (blue) in the green Kyushu group. More detailed information regarding the residential area of each firefly was obtained from the three-dimensional data of MDS analysis; the distribution of the plots of the East-Honshu fireflies (red) tended to be narrower than that of other two clusters and concentrated in a restricted location (Supplementary Fig. S1).
Representation of the L. cruciata accessions via principal component analysis (PCA). The three major component data are shown by the three two-dimensional data sets: (A) first and second PCA axes, (B) first and third PCA axes, and (C) second and third PCA axes. The contribution rate of each component is shown in parentheses. The colour of each plotted accession indicates the sample collection area (Fig. 1); the eastern area of Honshu is red, the western area of Honshu is blue, and Kyushu is green. The three clusters are distinguished by the coloured rope hook.
Gene-level phylogenetic tree analysis
We constructed two types of gene-level phylogenetic trees at the nuclear DNA level based on the ML method and Bayesian inference analysis (Fig. 3 and Supplementary Fig. S2), in which it was assumed that within-gene recombination was absent. For these analyses, 69,314 phylogenetically informative sites, which were detected by multiple alignments using the reference genome, were used. In addition, the midpoint was set as a root. Notably, this point is estimated as a root using the UPGMA method, in which root estimation is possible. A comparison of the two phylogenetic trees showed no differences in the topology of the trees, and both trees showed two major clades, A and B-C, with the latter clade divided into B and C. The inferred clades were supported by a bootstrap score of 100. The accessions belonging to each clade were consistent with their habitation areas, East-Honshu (Clade A), West-Honshu (Clade B), and Kyushu (Clade C), respectively. This result was the same with the grouping via PCA and MDS analyses with the exception of three accessions; No. 4 collected in East-Honshu was included in Clade B, No. 9 collected in West-Honshu was included in Clade A, and No. 14 collected in West-Honshu was included in Clade C. Based on the PCA results and branch lengths in the nuclear DNA trees, the genome similarity between the West-Honshu and Kyushu fireflies was higher than that between the West- and East-Honshu fireflies.
Phylogenetic tree of L. cruciata accessions using nuclear DNA data based on maximum likelihood (ML) reconstruction. The numbers at the nodes indicate bootstrap values (% over 1,000 replicates). The scale bar shows the number of substitutions per site. The colours of each accession indicate the sample collection area (Fig. 1); the eastern area of Honshu is red, the western area of Honshu is blue, and Kyushu is green. The right sidebar shows the three major clades divided by the branching pattern of the phylogenetic tree: (A) East-Honshu, (B) West-Honshu, and (C) Kyushu. The midpoint of the longest branch is used as a root.
Another phylogenetic tree using the mitochondrial CO II gene was constructed using the ML method (Fig. 4). The Tamura 3-parameter nucleotide substitution model was selected using the ML method with a discrete Gamma distribution (parameter = 0.1358)38. This tree showed two major clades, I-II and III, with the former clade divided into I and II. The haplotypes belonging to each clade were also consistent with their habitation areas: East-Honshu (Clade I), West-Honshu (Clade II), and Kyushu (Clade III). In addition to the same three exceptions of the nuclear tree (Nos. 4, 9, and 14), the haplotypes Nos. 38 and 39 were also exceptional in that they were included in Clade I, although they were collected in the West-Honshu area. The branching order of the clades was different from the nuclear trees, and the mitochondrial genome similarity between the West- and East-Honshu fireflies was higher than that between the West-Honshu and Kyushu fireflies as also found in our previous paper10.
Phylogenetic tree of the haplotypes of L. cruciata using the mitochondrial CO II gene based on maximum likelihood (ML) reconstruction. The numbers at the nodes indicate bootstrap values (% over 1,000 replicates). The scale bar shows the number of substitutions per site. The colours of each haplotype indicate the sample collection area (Fig. 1); the eastern area of Honshu is red, the western area of Honshu is blue, and Kyushu is green. The right sidebar shows three major clades: (I) East-Honshu, (II) West-Honshu, and (III) Kyushu. The haplotype of L. owadai is used as outgroup.
Phylogenetic tree analysis under the coalescent model
In contrast to the aforementioned two methods, which were used to analyse nuclear DNA, in the method under the coalescent model such as SVDquartets, it is assumed that within-gene recombination is present. In the SVDquartets analysis shown in Fig. 5, L. cruciata was resolved into two major groups, East-Honshu and West Japan, and the latter group divided into two subgroups, West-Honshu and Kyushu, which is the same as the gene-level phylogenetic trees of ML and Bayesian models (Fig. 3 and Supplementary Fig. S2). The inferred groups were supported by bootstrap scores from 97.6 to 100. The introgression samples (sample No. 9 in the East-Honshu group, No. 4 in West-Honshu, and No. 14 in Kyushu) were also recognized.
Phylogenetic tree constructed using the SVDquartets method with PAUP using nuclear DNA data of L. cruciata. The numbers at the nodes indicate bootstrap values (% over 1,000 replicates). The colours of each accession indicate the sample collection area (Fig. 1): the eastern area of Honshu is red, the western area of Honshu is blue, and Kyushu is green. The right sidebar shows the three groups inferred: (A) East-Honshu, (B) West-Honshu, and (C) Kyushu. The midpoint is used as a root.
Admixture analysis
Based on the results of the admixture program, hypothetical ancestral population (K) value selection was performed from K = 1 to 7. It was found that the error rate was lowest when setting the value as 3 (Fig. 6 and Supplementary Fig. S3). The second lowest error rate was confirmed at K = 2. The sample accessions were horizontally aligned according to their habitation areas, East-Honshu, West-Honshu, and Kyushu; the vertical axis indicated the ancestral ratio of the three hypothetical ancestral populations, which were coloured in purple, orange, and brown, respectively. The distribution of the three hypothetical ancestral populations was consistent with the habitation area of the accessions, which was the same as the grouping via PCA and MDS analyses (Fig. 2 and Supplementary Fig. S1). The clustering via nuclear and mitochondrial DNA trees (Figs. 3 and 4 and Supplementary Fig. S2) had the same exceptions of accessions Nos. 4, 9, and 14. However, no gene mixing was observed in the No. 4 and 9 fireflies, though it was observed between the West-Honshu and Kyushu areas in the No. 10, 14, 15, 46, 47, and 48 accessions. The No. 10, 14, and 15 fireflies inhabited the border area between the West-Honshu and Kyushu areas; however, the original habitation area of Nos. 46 and 47 were not identified because they were purchased by distributors. Gene mixing between the East- and West-Honshu areas was also observed in Nos. 38 and 39; >90% of the genome was from a hypothetical population of West-Honshu, although their mitochondrial haplotypes were confirmed as the East-Honshu type (Fig. 4).
Admixture analysis of the L. cruciata accessions. The number of populations (K) is set to 3 for admixture analysis. The accessions are horizontally aligned according to the three clades (East-Honshu, West-Honshu, and Kyushu) components of the phylogenetic tree using nuclear DNA data (Fig. 3). The colour of each accession indicates the sample collection area; the eastern area of Honshu is red, the western area of Honshu is blue, and Kyushu is green (Fig. 1). The vertical axis indicates the ancestry ratio of the three hypothetical ancestral populations coloured purple, orange, and brown, respectively.
Wright’s fixation indices
According to the result of admixture and phylogenetic tree analyses, the firefly accessions in this study can be reasonably divided into 3 groups; East-Honshu, West-Honshu, and Kyushu fireflies. To estimate the degree of genetic diversity among and within the three groups, 2 types of pairwise Fst values were calculated.
First Fst is the value among the three groups, and was calculated as 0.347794 (East-Honshu vs. West-Honshu), 0.546587 (East-Honshu vs. Kyushu), and 0.199462 (West-Honshu vs. Kyushu), respectively (Table 2). The lowest value was observed in the West-Honshu vs. Kyushu firefly sets, and the highest value in the East-Honshu vs. Kyushu sets. These results indicate that the genetically closest pair is the West-Honshu and Kyushu groups and that the East-Honshu and Kyushu groups are the furthest. This result was the same as that of the PCA, MDS, and phylogenetic tree analyses using nuclear DNA but different from that of the mitochondrial DNA analysis.
Second Fst is the value among accessions within each group (Supplementary Table S2). These pairwise Fst values were plotted in Fig. 7 and average values were calculated as 0.73559 (in East-Honshu), 0.81016 (in West-Honshu), and 0.81076 (in Kyushu), respectively. The East-Honshu group significantly had lowest genetic diversity of the groups (p < 2.20 × 10−16 between the East- and West-Honshu groups; p < 2.20 × 10−9 between the East-Honshu and Kyushu groups),while the average value between the West-Honshu and Kyushu groups was not significantly different (p = 0.9398).
Plot of pairwise Fst values among accessions within the three groups of the phylogenetic tree using nuclear DNA data (Fig. 3): East-Honshu, West-Honshu, and Kyushu clades. Details of each Fst value are shown in Supplementary Table S2. The averaged value for each group is indicated by a cross mark. *: no significant difference, **: significantly different at the 5% level (p < 2.20 × 10−16 between East- and West-Honshu; p < 2.20 × 10−9 between East-Honshu and Kyushu).
Genetic marker search and distribution of SNPs
To distinguish the three groups, genetic markers were searched and two SNP sites were identified (Fig. 8). These two SNPs were positioned in the noncoding region of the L. cruciata genome. In addition, other genetic markers were also searched to divide them into East-Japan (consistent with East-Honshu) and West-Japan (including West-Honshu, Shikoku, and Kyushu) groups; 2,545 SNPs were predicted on the entire genome sequence of L. cruciata. The results are shown in Table 3 and Supplementary Table S3. Although 1,964 SNPs were positioned in the noncoding region, the remaining 581 SNPs were located in the gene regions. Among them, 215 SNPs were distributed on protein-coding sequences (CDS), and in particular, 77 SNPs were found to specifically cause amino acid substitutions at translated protein levels.
Discussion
The results of PCA, MDS, gene-level phylogenetic tree, and admixture analyses using nuclear DNA data show that 52 accessions of L. cruciata are separated into three groups, East-Honshu, West-Honshu, and Kyushu types (Figs. 2, 3, and 6 and Supplementary Fig. S1). The grouping is also confirmed by the phylogenetic tree, in which the within-gene recombination is considered (Fig. 5). The groups geologically and geographically correspond to the eastern and western areas of Honshu divided by Fossa Magna and Kyushu. Gene flow among the groups is almost absent in natural populations. Exceptionally, accession Nos. 4 and 9 were collected in East- and West-Honshu, but were unequivocally placed in the West- and East-Honshu group, respectively, according to both their nuclear and mitochondrial DNA. Therefore, it is speculated that the two firefly specimens were artificially transplanted to the collection areas. Meanwhile, accession No. 14 collected from West-Honshu is characterized in the Kyushu group. The collection site is geographically very near Kyushu, and 65% of the nuclear DNA belongs to the Kyushu group. Therefore, it is conjected that the firefly specimen is a progeny of the crossing between West-Honshu and Kyushu fireflies. However, it is unknown whether the sample originates from artificial transplantation or natural migration. Although nuclear DNA introgression among the groups is rarely observed, there is a possibility of intercrossing in a close area naturally or by artificially transplanted specimens in the seven accessions Nos. 10, 15, 38, 39, 46, 47, and 48 (Figs. 3 and 6 and Supplementary Fig. S2). Remarkably, the Izu population (accession Nos. 38 and 39) is included in the West-Honshu group at a nuclear DNA level but in the East-Honshu type at the mitochondrial DNA level (Clade I in Fig. 4). This result means there is a possibility of backcrossing for paternal characteristics in the first filial generation (F1) between females from East-Honshu and males from West-Honshu39. The tree under the coalescent model shows two major groups, East-Honshu and West Japan, and the latter is divided into West-Honshu and Kyushu subgroups (Fig. 5). Because the East-Honshu and West Japan groups correspond to the ecological types, the slow- and fast-flash types, respectively, the major two groups are differentiated by the coalescent model analysis. Ecologically, it is recognized by our field observations that not only the flash interval of mate-seeking males but also their flying speed and activity hours and the gregarious oviposition behaviour of females were differentiated between the eastern and western areas of Japan7,40. Furthermore, preferential swarming activity of males for artificial flash light was also differentiated in that the males of the slow-flash population preferentially responded to a longer flash interval (4 and 5 sec),while those of the fast-flash type population to a shorter flash interval (2 and 4 sec); pre-mating isolation was suggested41. Chromosomal polymorphism is also observed in two populations in that the number of chromosomes in the East-Honshu (Kanagawa prefecture) and West-Japan (Ehime prefecture, Shikoku) is 2n = 20 and 18, respectively42,43. Further studies are necessary to clarify the chromosomal differentiation and fertility between the two chromosomal types. At least considering that the gene flow among the three groups is almost absent in natural populations and the grouping corresponds to the two ecological types (flash signal differentiation), L. cruciata appears to be in the process of speciation in the Japanese islands.
The consistency between the distribution pattern of the three groups and the geological structure of the Japanese islands implies a vicariant speciation scenario for this species. According to the geological history of Japanese island formation, the ancient landmass of the island was separated from the eastern periphery of the Eurasian Continent and was divided into east and west landmasses during the early phase of Fossa Magna formation approximately 16 million years (MY) ago11,12. A considerable part of the current eastern area of the Japanese islands was island-like, mostly under the sea and formed by subsequent crustal uplift (13 MY ago); an outline of the current Japanese islands (Hokkaido, Honshu, Shikoku and Kyusyu) was established from 0.1 to 0.5 MY ago. All trees via nuclear DNA analysis show the branching order of the three groups: the East-Honshu and West-Honshu–Kyushu branch first, followed by branching of West-Honshu and Kyushu (Figs. 3 and 5 and Supplementary Fig. S2). Thus, it seems more likely that the split occurred first between the populations of East and West Japan, and later between West-Honshu and Kyushu. The East–West first scenario is also supported by the PCA and MDS (Fig. 2 and Supplementary Fig. S1), the branch lengths (which lead to the midpoint root) of the nuclear ML tree (Fig. 3 and Supplementary Fig. S2) and the K = 2 admixture analysis (Supplementary Fig. S3). Furthermore, because the Fst values among the accession within each group were significantly lowest in the East Honshu population, it may have undergone a bottleneck effect due to island formation via subsidence and uplift (Fig. 7 and Supplementary Table S2). Thus, in conclusion, we argue the East-West divergence happening first is more likely and better supported by the data. On the other hand, the absence of a nuclear DNA outgroup means the branching order cannot be accurately reconstructed. The rooted mitochondrial tree even suggests a different branching order, which could hint at a different evolutionary history for the mitochondrial DNA (Fig. 4). Both vicariance scenarios (East–West or Honshu–Kyushu first) have been reported from other insects13. The evolutionary process of the ecological types is not parsimoniously conclusive, because both scenarios, slow- or fast flash type are ancestral, require the same number of evolutionary steps. In addition, it is ambiguous whether the flash type of L. owadai, a closely related species of L. cruciata and used as an outgroup in mitochondrial DNA tree preparation, is an Eastern slow- or a Western fast flash type.
Finally, SNP markers are presented in Fig. 8, Table 3, and Supplementary Table S3. L. cruciata is a popular firefly in Japan and is used as a symbolic insect for nature conservation and regeneration. In this effort, artificial transplantation among the three groups was observed via mitochondrial CO II gene analysis44. The SNP markers of the nuclear DNA would also be available for genetic management of firefly transplantation in nature conservation and regeneration.
References
Kastle, J. H. & McDermott, F. A. Some observations on the production of light by the firefly. Cunningham, Curtiss & Welch. 27, 122–151 (1910).
Mast, S. O. Behavior of the firefly (Photinus pyralis) with special reference to the problem of orientation. Journal of Animal Behavior 2, 256–272 (1912).
Barber, H. S. & McDermott, F. A. North American fireflies of the genus Photuris. Smithsonian Inst. Misc. Collections 117, 1–58 (1951).
Lloyd, J. E. Studies of the flash communication in Photinus fireflies. Miscellaneous Publications of the Museum of Zoology, University of Michigan 130, 1–95 (1966).
Ohba, N. Studies on the communication system of Japanese fireflies. Sci. Rep. Yokosuka City Mus. 30, 1–62 (1983).
Ohba, N. Flash Communication Systems of Japanese Fireflies. Integrative and Comparative Biology 44, 225–233 (2004).
Ohba, N. Synchronous flashing in the Japanese firefly. Luciola cruciata (Coleoptera: Lampyridae). Sci. Rep. Yokosuka City Mus. 32, 23–32 (1984).
Suzuki, H., Saito, Y., Fujiyama, S. & Ohba, N. Allozymic differentiation between two ecological types of flashing behavior in the Japanese firefly. Luciola cruciata. Jpn. J. Ent. 64, 682–691 (1996).
Nei, M. Genetic distance between populations. Am. Nat. 106, 283–292 (1972).
Suzuki, H., Saito, Y. & Ohba, N. Gene diversity and geographic differentiation in mitochondrial DNA of the Genji Firefly. Luciola cruciata (Coleoptera: Lampyridae). Mol. Phy. Evol. 22, 193–205 (2002).
Otsuki, K. Oblique subduction, collision of microcontinents and subduction of oceanic ridge: Their implications on the Cretaceous tectonics of Japan. Island Arc 1, 51–63 (1992).
Editorial group for computer graphics, geology of Japanese islands. Computer Graphics, Geology of Japanese Islands. Editorial group for computer graphics, geology of Japanese islands. ed. 150 pp. Maruzen, Tokyo (1996).
Suzuki, T., Kitano, T. & Tojo, K. Contrasting genetic structure of closely related giant water bugs: Phylogeography of Appasus japonicus and Appasus major (Insecta: Heteroptera, Belostomatidae). Mol. Phylogenet. Evol. 72, 7–16 (2014).
Takenaka, M. & Tojo, K. Ancient origin of a dipteromimid mayfly family endemic to the Japanese Islands and its genetic differentiation across tectonic faults. Biol. J. Linnean Soc. 126, 555–573 (2019).
Su, Z.-H. et al. Phylogenetic relationships and evolution of the Japanese Carabinae group beetles based on mitochondrial ND5 gene sequences. J. Mol. Evol. 42, 124–129 (1996).
Gamboa, M., Muranyi, D., Kanmori, S. & Watanabe, K. Molecular phylogeny and diversification timing of the Nemouridae family (Insecta, Plecoptera) in the Japanese Archipelago. PLoS ONE 14, e0210269 (2019).
Tojo., K. et al. Species diversity of insects in Japan: Their origins and diversification processes. Entomol. Sci. 20, 357–381 (2017).
Baird, N. A. et al. Rapid SNP discovery and genetic mapping using sequenced RAD markers. PLoS ONE 3, e3376 (2008).
Davey, J. W. & Blaxter, M. L. RADSeq: next-generation population genetics. Brief. Funct. Genomics 9, 416–423 (2010).
Lower, S. E., Stanger-Hall, K. F. & Hall, D. W. Molecular variation across populations of a widespread North American firefly, Photinus pyralis, reveals that coding changes do not underlie flash color variation or associated visual sensitivity. BMC Evol. Biol. 18, 129 (2018).
Etter, P. D., Bassham, S., Hohenlohe, P. A., Johnson, E. A. & Cresko, W. A. SNP Discovery and Genotyping for Evolutionary Genetics Using RAD Sequencing. Methods Mol. Biol. 772, 157–178 (2011).
Catchen, J. M., Amores, A., Hohenlohe, P., Cresko, W. & Postlethwait, J. H. Stacks: building and genotyping loci de novo from short-read sequences. G3 (Bethesda) 1, 171–182 (2011).
Catchen, J. et al. Stacks: an analysis tool set for population genomics. Mol. Ecol. 22, 3124–3140 (2013).
Martin, M. Cutadapt removes adapter sequences from high‐throughput sequencing reads. EMBnet. Journal 17, 10 (2011).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: A flexible trimmer for Illumina Sequence Data. Bioinformatics 30, 2114–2120 (2014).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Zheng, X. et al. A high-performance computing toolset for relatedness and principal component analysis of SNP data. Bioinformatics 28, 3326–3328 (2012).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Ronquist, F. & Huelsenbeck, J. P. MRBAYES 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19, 1572–1574 (2003).
Chifman, J. & Kubatko, L. Quartet inference from SNP data under the coalescent model. Bioinformatics 30, 3317–3324 (2014).
Swofford, D. L. PAUP* Phylogenetic Analysis Using Parsimony (* and Other methods) Version 4. (Sinauer Associates, 1998).
Swofford, D. L. PAUP* (*Phylogenetic Analysis Using PAUP), http://paup.phylosolutions.com/ (2019).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664 (2009).
Simon, C. et al. Evolution, weighting, and phylogenetic utility of mitochondrial gene sequences and a compilation of conserved polymerase chain reaction primers. Ann. ent. Soc. Am. 87, 651–701 (1994).
Kumar, S., Stecher, G., Li, M., Knyaz, C. & Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across computing platforms. Mol. Biol. Evol. 35, 1547–1549 (2018).
Tamura, K. Estimation of the number of nucleotide substitutions when there are strong transition-transversion and G + C-content biases. Mol. Biol. Evol. 9, 678–687 (1992).
Avise, J. C. Phylogeography: The History and Formation of Species. (Harvard University Press, 2000).
Ohba, N. Geographical variation, morphology and flash pattern of the firefly, Luciola cruciata (Coleoptera: lampyridae). Sci. Rep. Yokosuka City Mus. 48, 45–89 (2001).
Tamura, M., Yokoyama, J., Ohba, N. & Kawaa, M. Geographic differences in flash intervals and pre-mating isolation between populations of the Genji firefly, Lucola cruciate. Ecol. Entomol. 30, 241–245 (2005).
Inoue, M. & Yamamoto, H. Cytological studies of family Lampyridae I. Karyotypes of Luciola lateralis and L. cruciata. La Kromosomo II 45, 1440–1443 (1987).
Suzuki, H. & Satoh, T. Luciola cruciata, Lampyride. In Chromosome Calendar 2017. The society of chromosome research, Japan ed. Tokyo, (2016)
Suzuki, H. Studies on biological diversity of firefly in Japan. Int. J. Indust. Emtomol. 2, 91–105 (2001).
Acknowledgements
This work was partially supported by a KAKENHI Grant-in-Aid for Scientific Research(C) (18K05320) from the Japan Society for the Promotion of Science (JSPS) and the Public Foundation Yonemori-seishinikuseikai. Firefly specimens were partly provided by Dr. N. Ohba (Yokosuka City).
Author information
Authors and Affiliations
Contributions
D.K. designed, analysed and interpreted all of the experiments and wrote the manuscript. H.S. collected the fireflies, performed the haplotype analysis of the mitochondrial CO II gene, and wrote the manuscript. A.T. and J.M. planned the experiments and analysed the data. Y.H. helped to identify the genetic markers by providing a custom Ruby program. K.A. and Y.I. performed the biochemical experiments and analysed the data. Y.N. performed the RAD-Seq analysis and helped to write the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kato, Di., Suzuki, H., Tsuruta, A. et al. Evaluation of the population structure and phylogeography of the Japanese Genji firefly, Luciola cruciata, at the nuclear DNA level using RAD-Seq analysis. Sci Rep 10, 1533 (2020). https://doi.org/10.1038/s41598-020-58324-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-58324-9
This article is cited by
-
RAD-Seq analysis of wild Japanese garlic (Allium macrostemon Bunge) growing in Japan revealed that this neglected crop was previously actively utilized
Scientific Reports (2023)
-
Genetic diversity of loquat (Eriobotrya japonica) revealed using RAD-Seq SNP markers
Scientific Reports (2022)
-
Genetic diversity and population structure of razor clam Sinonovacula constricta in Ariake Bay, Japan, revealed using RAD-Seq SNP markers
Scientific Reports (2021)
-
Elucidation of Japanese pepper (Zanthoxylum piperitum De Candolle) domestication using RAD-Seq
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.