Abstract
To compare different deep learning architectures for predicting the risk of readmission within 30 days of discharge from the intensive care unit (ICU). The interpretability of attention-based models is leveraged to describe patients-at-risk. Several deep learning architectures making use of attention mechanisms, recurrent layers, neural ordinary differential equations (ODEs), and medical concept embeddings with time-aware attention were trained using publicly available electronic medical record data (MIMIC-III) associated with 45,298 ICU stays for 33,150 patients. Bayesian inference was used to compute the posterior over weights of an attention-based model. Odds ratios associated with an increased risk of readmission were computed for static variables. Diagnoses, procedures, medications, and vital signs were ranked according to the associated risk of readmission. A recurrent neural network, with time dynamics of code embeddings computed by neural ODEs, achieved the highest average precision of 0.331 (AUROC: 0.739, F1-Score: 0.372). Predictive accuracy was comparable across neural network architectures. Groups of patients at risk included those suffering from infectious complications, with chronic or progressive conditions, and for whom standard medical care was not suitable. Attention-based networks may be preferable to recurrent networks if an interpretable model is required, at only marginal cost in predictive accuracy.
Similar content being viewed by others
Introduction
Modern machine learning algorithms such as artificial neural networks with multiple hidden (“deep”) layers can extract relevant features from medical data and make predictions for previously unseen patients. Examples of successful applications of deep learning techniques in the medical domain include the classification of skin cancer images with accuracy comparable to dermatologists1, the prediction of cardiovascular risk factors from retinal fundus photographs2, and the prediction of diseases including severe diabetes, schizophrenia, and various cancers based on information contained in electronic medical records (EMR)3. Nonetheless, integration of machine learning assistants into clinical processes remains scarce and actual benefits for patient care are yet to be demonstrated4.
Attention layers in deep neural networks enable them to focus on a subset of inputs (features) and respond accordingly, increasing model performance. They also offer much-needed model interpretability, by identifying which components of the input are attended to at any point in time. Current deep learning architectures used for risk prediction based on EMR data generally employ attention layers on top of recurrent layers5,6,7 (e.g. Long Short-Term Memory, LSTM8, or Gated Recurrent Units, GRU9). While these models yield state of the art results in prediction accuracy, the use of recurrent layers is associated with several drawbacks: interpretation of results is hampered by outputs being a nonlinear combination of current input and current memory state; lack of set-invariance (i.e. outputs differ based on the specific sequence of timestamped variables within the EMR, even if these variables were recorded at the same time)10; and long training times due to difficulties in parallelizing these sequential algorithms. Neural networks relying entirely on attention mechanisms have been proposed as an alternative to overcome the limitations of recurrent neural networks, with comparable or improved accuracy on several language processing tasks11 and when used for risk predictions based on EMR data12.
In their basic form, neither recurrent layers nor attention mechanisms are tailored to process time-series sampled at irregular time intervals, such as the diagnosis and procedure codes contained in EMRs. A wide variety of approaches to address this issue have been proposed in recent literature, ranging from simply adding or appending time-related information to the numerical vectors (“embeddings”) used to represent timestamped codes5, to modifying the internal workings of recurrent cells using exponential time-decay functions13,14,15 or ordinary differential equations (ODEs)16,17,18. Similarly, code embeddings may be determined end-to-end while optimising the other parameters of the network, or in an unsupervised pre-processing step that considers time-related information, e.g. through medical concept embeddings (MCEs) with time-aware attention19. An additional option may be the use of neural ODEs to describe how the embedding (intuitively, the “meaning”) of a medical code changes over time: codes related to a chronic condition, such as diabetes, will often maintain their relevance over years, whereas others may quickly become unimportant for prediction purposes. Amidst all these options, comprehensive work comparing neural network architectures for risk prediction based on EMR data is currently limited.
Predicting a patient’s risk of readmission within 30 days of discharge from the intensive care unit (ICU) presents an exemplar application for machine learning models using longitudinal EMR data. Readmission to the ICU is an adverse outcome experienced by approximately 10% of critically ill patients following discharge20,21 and may be an indicator of poor or incomplete medical care21,22,23,24. If patients at high risk of readmission can be identified, appropriate interventions, such as careful patient evaluation before discharge, planning for proper placement of the patient after discharge, and a safe and thorough handover of patient care between healthcare providers, can be implemented25. Readmission to the ICU also represents a major source of avoidable costs for the health care system, as up to 30% of total hospital costs and 1% of the US gross national product are directly linked to ICU expenses26. Points-based scores such as the Acute Physiology and Chronic Health (APACHE) score27, the Simplified Acute Physiology Score (SAPS)28, and the Oxford Acute Severity of Illness Score (OASIS)29 are routinely used in ICUs to evaluate severity of illness and predict mortality risk; they may also be useful in predicting the risk of readmission30. However, a recent study comparing several scores used to predict the risk of readmission within 48 hours from discharge determined only moderate discrimination power (area under the receiver operating characteristic curve, AUROC, between 0.65 and 0.67)31. It is plausible that the application of novel machine learning algorithms to EMR data could lead to more accurate predictions.
Using the example of predicting the risk of readmission within 30 days of discharge from the ICU, the aims of the present study are: (1) to evaluate the feasibility of using neural ODEs to model how the predictive relevance of recorded medical codes changes over time; (2) to perform a comprehensive comparison of deep learning models that have been proposed for processing time-series sampled at irregular intervals, including MCEs, neural ODEs, attention mechanisms, and recurrent layers; and (3) to leverage the interpretability of attention layers combined with Bayesian deep learning to gain a better understanding of intensive-care patients at increased risk of readmission.
Methods
Study population
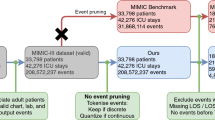
The algorithms were evaluated on the publicly available MIMIC-III data set (ethics approval was not required)32. This data set comprises deidentified health data associated with 61,532 ICU stays and 46,476 critical care patients at Beth Israel Deaconess Medical Center in Boston, Massachusetts between 2001 and 2012.
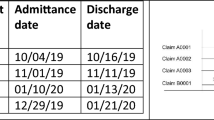
The supervised learning task consists of predicting, for a given ICU stay, whether the patient will be readmitted to the ICU within 30 days from discharge. Patients were excluded if they died during the ICU stay (N = 4,787 ICU stays), were not adults (18 years old or older) at the time of discharge (N = 8,129 ICU stays) or died within 30 days from discharge without being readmitted to the ICU (N = 3,318 ICU stays). The final data set comprised 45,298 ICU stays for 33,150 patients, labelled as either positive (N = 5,495) or negative (N = 39,803) depending on whether a patient did or did not experience readmission within 30 days from discharge. To develop and evaluate the algorithms, patients were subdivided randomly into training and validation (90%) and test sets (10%). This subdivision was based on patient identifiers and not on ICU stay identifiers to prevent information leaks between data sets (since the prediction is based on the entire clinical history of a patient).
Model variables
The EMR of a patient can be represented as a set of static variables and timestamped codes. In the present study, static variables included the patient’s gender, age, ethnicity, insurance type, marital status, the previous location of the patient prior to arriving at the hospital (admission location), and whether the patient was admitted for elective surgery. Both length of ICU stay and length of hospital stay prior to ICU admission were recorded. An additional static variable was given by the number of ICU admissions in the year preceding the considered index ICU stay.
Data types of timestamped codes included international classification of diseases and related health problems (ICD-9) diagnosis and procedure codes, prescribed medications, and patient vital signs. All diagnosis and procedure codes in the clinical history of a patient were considered for predictive purposes; however, prescribed medications and recorded vital signs were restricted to the ICU stay of interest. Following the OASIS severity of illness score29, assessed vital signs comprised the Glasgow Coma Scale score (sum of eye response, verbal response, motor response components), heart rate, mean arterial pressure, respiratory rate, body temperature, urine output, and whether the patient necessitated ventilation. Continuous measurements of vital signs were categorised in the same manner as in OASIS and assigned corresponding codes29. To reduce redundant information, whenever the same vital sign-related code was recorded consecutively more than once, only the latest observation was kept in the data.
Elapsed times, measured in days, associated with diagnosis and procedure codes were based on the date and time of discharge from the corresponding hospital admission. Elapsed times, measured in hours, associated with medications and vital signs were based on the date and time of prescription start and measurement, respectively. In the present study, the simplifying assumption is made that diagnosis and procedure codes are available immediately at the time of discharge from the ICU. Categorical values of static variables or timestamped codes associated with less than 100 ICU stays were re-labelled as “other”.
Artificial neural network architectures
Several “deep” neural network architectures for predicting a patient’s risk of readmission to the ICU were implemented and compared. To make this comparison as fair as possible, all architectures shared a similar high level structure: (1) timestamped codes were mapped to vector embeddings; (2) numerical scores associated with diagnosis and procedure codes, and with medication and vital sign codes, were computed using attention mechanisms and/or recurrent layers; (3) these scores were concatenated with the static variables and passed on to a “logistic regression layer” (i.e. a fully connected layer with a sigmoid activation function). Further details about individual network components are reported in the following sections.
Embeddings
Diagnosis and procedure codes, as well as medication and vital sign codes, were mapped to corresponding “embeddings” (real-valued vectors). The size of these embeddings was set proportional to the fourth root of the total number of codes in the dictionary (diagnoses/procedures and medications/vital signs were processed separately since they were measured on different time scales)7. Time-aware code embeddings were computed in three different manners. A first approach used MCEs with time-aware attention19. MCEs are based on the continuous bag-of-words model33, but instead of using fixed-sized temporal windows to determine a code’s context, attention mechanisms learn the temporal scope of a code together with its embedding. A second approach optimised an embedding matrix at the same time as the other parameters of the network and, optionally, concatenated the elapsed times to the resulting vectors. A third approach optimised an embedding matrix at the same time as the other parameters of the network and modelled the dynamics in time of the computed embeddings using neural ODEs16,17,18. More in detail, the embedding of a code at time zero (i.e. at the time of discharge from the ICU) was stored in the embedding matrix whereas the embedding of a code recorded before discharge was computed by solving an initial value problem where derivatives with respect to time were approximated by a multilayer perceptron.
Attention and/or recurrent layers
The sequence of code embeddings associated with a patient is usually of arbitrary length and needs to be integrated into a fixed-size vector for further processing. Attention mechanisms, such as dot-product attention34, compute a weighted average of the code embeddings, where a higher weight is assigned to the most relevant codes. Alternatively, recurrent layers iteratively process an input sequence of codes and, at each iteration, update an internal memory state and generate an output vector8,9. Information may be integrated for further processing by using either the final memory state of the recurrent cell or by applying an attention mechanism to the set of output vectors5,6,7. Specifically, in this work, recurrent cells were implemented using bi-directional gated recurrent units9. Time-related information was taken into account by concatenating the time differences between observations to the embedding vectors, by applying an exponential decay proportional to the time differences between observations to the internal memory state of the recurrent cell13,14,15 or by modelling the dynamics in time of the internal memory state using neural ODEs16,17,18. To aid subsequent interpretation without altering network capacity, the fixed-size vectors produced by attention mechanisms and/or recurrent layers were reduced to two scalar-valued scores (one related to diagnoses/procedures and one related to medications/vital signs) using fully connected layers with a linear activation function.
Logistic regression layer
The computed diagnoses/procedures and medications/vital signs scores were concatenated to the vector of static variables and passed to a fully connected layer with a sigmoid activation function. The output of the network corresponds to the risk of readmission to the ICU within 30 days from discharge.
Architectures
The following neural network architectures were compared for predicting readmission to the ICU:
ODE + RNN + Attention: dynamics in time of embeddings are modelled using neural ODEs, embeddings are passed to RNN layers, dot-product attention is applied to RNN outputs.
ODE + RNN: dynamics in time of embeddings are modelled using neural ODEs, embeddings are passed to RNN layers, the final memory states are used for further processing.
RNN (ODE time decay) + Attention: embeddings are passed to RNN layers with dynamics in time of the internal memory states modelled using neural ODEs, dot-product attention is applied to RNN outputs.
RNN (ODE time decay): embeddings are passed to RNN layers with dynamics in time of the internal memory states modelled using neural ODEs, the final memory states are used for further processing.
RNN (exp time decay) + Attention: embeddings are passed to RNN layers with internal memory states decaying exponentially over time, dot-product attention is applied to RNN outputs.
RNN (exp time decay): embeddings are passed to RNN layers with internal memory states decaying exponentially over time, the final memory states are used for further processing.
RNN (concatenated Δtime) + Attention: embeddings are concatenated with time differences between observations and passed to RNN layers, dot-product attention is applied to RNN outputs.
RNN (concatenated Δtime): embeddings are concatenated with time differences between observations and passed to RNN layers, the final memory states are used for further processing.
ODE + Attention: dynamics in time of embeddings are modelled using neural ODEs, dot-product attention is applied to the embeddings.
Attention (concatenated time): embeddings are concatenated with elapsed times, dot-product attention is applied to the embeddings.
MCE + RNN + Attention: MCE is used to compute the embeddings, embeddings are passed to RNN layers, dot-product attention is applied to RNN outputs.
MCE + RNN: MCE is used to compute the embeddings, embeddings are passed to RNN layers, the final memory states are used for further processing.
MCE + Attention: MCE is used to compute the embeddings, dot-product attention is applied to the embeddings.
The dimension of the internal memory state of RNN cells was set equal to the dimension of the input embeddings. Similarly, the dimension of the hidden representation of embeddings when computing dot-product attention was left unchanged. Derivatives with respect to time used to implement neural ODEs were approximated by a multilayer perceptron with three hidden layers of constant width equal to the size of the input. The Euler method was used as ODE solver.
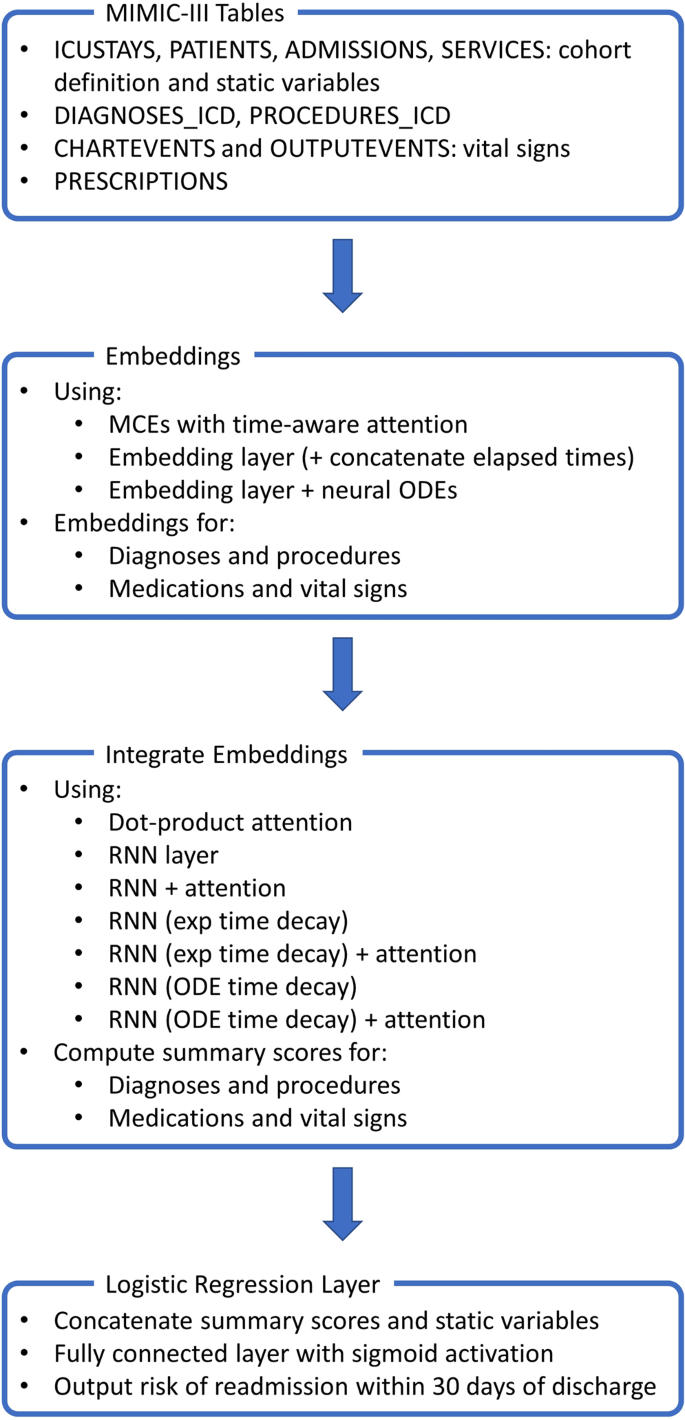
An overview of the considered neural network architectures is presented in Fig. 1. For completeness, the deep learning approaches were also compared with a logistic regression model using all static variables and the most recent vital signs for each patient as covariates.
Interpretation of attention-based models
For the proposed neural network architectures, the weights of the final fully connected layer can be used to determine the impact of static variables and timestamped codes on estimated risk. As in traditional logistic regression, these weights can be interpreted as increases in log-odds for unplanned early ICU readmission if the corresponding static variables or scores are increased by one unit.
It is also of interest to determine which codes (i.e. diagnoses, procedures, medications, vital signs) are associated with a prediction of high risk. Dot-product attention computes a weighted average of embedded codes; fully connected layers are then used to output scores associated with diagnoses/procedures and medications/vital signs. By passing single codes (i.e. the rows of the embedding matrix) to the fully connected layers computing these scores, it is possible to associate each code with a score. The higher the score, the higher the risk of ICU readmission when a patient’s EMR contains that code.
To estimate Bayesian credible intervals around network weights and computed risk scores, the posterior distribution of weights was approximated using stochastic variational inference with mean-field approximation35,36. In the present study, the variational posterior is assumed to be a diagonal Gaussian distribution and is estimated using the Bayes by Backprop algorithm37. Following the original paper, a priori sparsity of the network weights is encouraged by formulating the prior distribution as a scale mixture of two zero-mean Gaussian densities with standard deviations of σ1 = 1 and σ2 = e−6, respectively, and mixture weight π = 0.537. After the posterior distribution has been computed, 95% credible intervals around network weights (or combinations thereof) can be estimated by repeated sampling. Sampling of network weights may also be used to compute credible intervals around the risk prediction for a given patient.
Training
To compare the classification accuracy of the considered neural network architectures, maximum likelihood estimates of network parameters were obtained using a log-loss cost function on the training data, extensive use of dropout with 50% probability after each embedding, RNN, and attention layers38, and stochastic gradient descent with an Adam optimizer (batch size of 128 and learning rate of 0.001)39. Class imbalance was taken into consideration by assigning a proportionally higher cost of misclassification to the minority class40. Training was terminated after 80 epochs since overfitting of the training data started to become apparent with additional training epochs (based on average precision on the validation data). For interpretation purposes, Bayes by Backprop was used to train the “Attention (concatenated time)” neural network architecture on the entire data set, terminating if the loss function (the expected lower bound) did not decrease for 10 consecutive epochs.
Statistical analysis
Baseline characteristics were determined for the analysed patient population. The prediction accuracy of each considered algorithm was evaluated based on average precision, AUROC, F1-Score, sensitivity, and specificity. Average precision may reflect algorithmic performance on imbalanced data sets better than AUROC as it does not reward true negatives41,42. The F1-Score was maximised over different threshold values on risk predictions. Sensitivity and specificity were computed by maximising Youden’s J statistic43. 95% confidence intervals associated with each metric were computed by bootstrapping, i.e. by sampling the test set with replacement 100 times and re-evaluating the models each time44. Since the bootstrap estimator assumes the resampling of independent events, sampling was based on patient identifiers rather than on ICU stay identifiers.
Training the “Attention (concatenated time)” network using Bayes by Backprop allowed computation of odds ratios (OR) associated with static variables and ranking of the timestamped codes (diagnoses, procedures, medications, and vital signs) according to their associated average scores (a high positive score corresponds to increased risk of readmission to the ICU); corresponding 95% credible intervals were determined using 10,000 network samples.
Software was implemented in Python using Scikit-learn45 and PyTorch46; the developed algorithms are publicly available at https://github.com/sebbarb/time_aware_attention.
Results
Baseline characteristics
Baseline characteristics of the analysed patient population are reported in Supplementary Table S1. In total, the models were trained using 23 static variables, 992 unique ICD-9 diagnosis codes, 298 unique ICD-9 procedure codes, 586 unique medications, and 32 codes related to vital signs. The embedding dimension was 12 for diagnoses and procedures and 10 for medications and vital signs. Each patient’s EMR contained at most 552 ICD-9 diagnosis and procedure codes and 392 medications and vital sign codes associated with the current ICU stay.
Model comparison
Average precision, AUROC, F1-score, sensitivity, and specificity for the considered deep learning architectures and the logistic regression model are reported in Table 1. The highest average precision of 0.331 was obtained by the ODE + RNN model. In general, the predictive accuracy of neural networks was considerably higher than for the logistic regression baseline model (average precision of 0.257). Models with a recurrent component (average precision range: 0.298–0.331) performed slightly better than models based solely on attention layers (average precision range: 0.269–0.294). Applying an attention layer to the outputs of RNNs at each time step instead of directly using the final memory state of the RNN, and similarly learning code embeddings end-to-end instead of using pre-trained MCE, increased network capacity but only lead to occasional, and marginal, improvements in predictive accuracy.
Interpretation of attention-based models
Table 2 reports the exponentiated coefficients of the last fully connected layer of the “Attention (concatenated time)” model. They can be interpreted as odds ratios for readmission within 30 days from ICU discharge. Length of stay in the ICU was not associated with readmission risk; however, a longer length of stay before admission to the ICU had a minor protective effect on the odds of experiencing readmission, with an expected OR of 0.994 (95% credible interval of [0.993, 0.996]) between patients hospitalised for one additional day and other patients with the same values for other covariates. Male gender, a higher number of recent admissions, and older age were all associated with higher odds of readmission (OR: 1.114 [1.092, 1.136], 1.187 [1.170, 1.205], and 1.009 [1.009, 1.010], respectively) whereas admission for elective surgery was associated with lower odds of readmission (OR: 0.941 [0.891, 0.993]). Patients with admission location physician referral/normal delivery (OR: 0.882 [0.844, 0.922]) had lower odds of readmission compared with patients admitted through the emergency department. Patients insured through other government programs, with private health insurance, or who self-paid the hospitalization had lower odds of readmission compared with Medicare patients (OR: 0.775 [0.694, 0.865], 0.820 [0.798, 0.843], 0.559 [0.447, 0.700], respectively); however, the odds were similar for Medicare and Medicaid patients (OR: 0.997 [0.992, 1.002]). Marital status was not associated with a difference in odds. Black/African American patients were more likely to experience readmission than white patients (OR: 1.165 [1.118, 1.215]).
Scores for individual diagnosis, procedure, and medication codes associated with an increased risk of readmission within 30 days of discharge from the ICU are reported in Table 3. Patients-at-risk included subjects suffering from infectious complications, e.g. following the insertion of cardiac devices or venous catheters. Similarly, patients diagnosed with specific chronic (uncontrolled type I diabetes, liver disease) or progressive (hepatorenal syndrome) conditions were also at increased risk of early readmission. Another group of interest included patients for whom standard medical care is not possible, e.g. due to contraindications to surgery or medications, such as hydantoin derivatives (a class of anticonvulsants), and requiring desensitization to allergens. The high scores assigned by the model to diagnosed dysphagia and gastrostomy procedures may be associated with critically ill patients. Vital signs associated with increased risk of readmission included a recorded body temperature in the range 33.22–35.93 °C (score: 1.7 [0.2, 3.2]), mean arterial pressure in the range 51–61.32 mmHg (score: 1.2 [0.7, 1.7]), and a respiratory rate in the range 31–44 breaths per minute (score: 1.1 [0.2, 1.9]).
Discussion
Principal results
This study benchmarked several deep learning architectures for processing time-series sampled at irregular intervals, including a novel application of neural ODEs to model the dynamics in time of medical code embeddings. Models were trained to predict the risk of readmission within 30 days of discharge from the ICU using the MIMIC-III data set. In general, the predictive accuracy of the different deep learning models was similar, but considerably better than logistic regression. It is possible that diagnosis and procedure codes associated with ICU admissions prior to the index admission, as well as medications and vital signs, provide limited additional value when predicting readmission, therefore limiting the impact of different network architectures on predictive accuracy. Neural ODEs applied to code embeddings did generally result in improved performance, suggesting that they may constitute a building block of interest for neural networks processing not only continuous time series, but also timestamped codes.
Models with a recurrent component performed marginally better in terms of accuracy than models based solely on attention mechanisms; these findings are consistent with a previous study which predicted clinical outcomes using the MIMIC-III data12. However, by formulating an attention-based model in a Bayesian setting it was possible to evaluate not only predictive performance12,47 but also to derive credible intervals on network weights and risk predictions; as well as to provide a high degree of interpretability for all model coefficients including those encoding the longitudinal aspect of EMR data. Since training attention-based networks is more efficient than training RNNs, it should be possible to scale the proposed architecture to larger data sets and frequently update network weights as new data is collected within hospitals. Accuracy could be improved further by constructing a Bayesian ensemble of classifiers37.
Comparison with prior work
The interpretation of the attention-based model supports several previous studies which identified associations between increased risk of readmission and male gender, older age, and admission location25,48,49,50. Whereas previous work found that length of ICU stay was higher among readmitted patients, the present study was inconclusive in this regard25,49,50. Similarly to the OASIS severity of illness score, a higher risk of readmission was predicted for patients with a very short hospital length of stay before admission to the ICU and for patients who were not admitted for elective surgery29.
This study also identified patients suffering from infectious complications51,52 or chronic conditions, such as diabetes53 or cirrhosis25,31, as being at increased risk of readmission. Further, this study emphasizes that patients for whom standard medical care is not possible, e.g. due to contraindications to surgery or medications, may also be at increased risk of readmission, a finding which should be examined further by future studies.
Compared with previous work on predicting general hospital readmission, this study also identified a significant risk for patients belonging to minority groups54,55 but not for patients living alone55,56. The latter finding may reflect the fact that patients are usually discharged from the ICU to a hospital ward, rather than directly to home. Finally, Medicaid and Medicare patients may be at increased risk of ICU readmission compared with privately insured patients56,57.
Limitations
The present study has several limitations. Since all dates in the MIMIC-III data set were shifted to protect patient confidentiality, it was not possible to ascertain which patients were admitted after 2001 and had at least 12 months of prior data, possibly leading to some incorrect values for the number of ICU admissions in the year preceding discharge. The use of attention mechanisms with multiple keys/values or multiple attention heads11 was not assessed; however, these algorithms might be more relevant for networks with a decoder component or multi-class/multi-label classification tasks. The proposed model does not address interactions between static variables or non-linear associations between static variables and predicted risk. It also doesn’t account for within-patient clustering of multiple ICU admissions. Further, information from clinical notes58 was not included and the simplifying assumption was made that various diagnosis- and procedure-related codes were available immediately at the time of discharge. Including prior medical knowledge in the model is currently not possible (however, as expected, normal vital signs were assigned a very low risk score by the attention-based model). A larger prospective study using Australian hospital data will be used to address these limitations.
Conclusions
In conclusion, this study compared several deep-learning models for predicting the risk of readmission within 30 days of discharge from the ICU based on the full clinical history of a patient. The considered models included a novel application of neural ODEs to model how the predictive relevance of medical codes changes over time. Training an attention-based network in a Bayesian setting allowed insights into intensive-care patients at increased risk of readmission. Overall, attention-based networks may be preferable to recurrent networks if an interpretable model is required, at only marginal cost in predictive accuracy. The development of interpretable machine learning techniques such as proposed here is necessary to allow the integration of predictive models in clinical processes.
Data availability
The MIMIC-III data set used in this study is publicly available32.
References
Esteva, A. et al. Dermatologist-level classification of skin cancer with deep neural networks. Nature 542, 115–118, https://doi.org/10.1038/nature21056 (2017).
Poplin, R. et al. Prediction of cardiovascular risk factors from retinal fundus photographs via deep learning. Nature Biomedical Engineering 2, 158–164, https://doi.org/10.1038/s41551-018-0195-0 (2018).
Miotto, R., Li, L., Kidd, B. A. & Dudley, J. T. Deep patient: an unsupervised representation to predict the future of patients from the electronic health records. Scientific reports 6, 26094, https://doi.org/10.1038/srep26094 (2016).
Chen, J. H. & Asch, S. M. Machine learning and prediction in medicine—beyond the peak of inflated expectations. The New England journal of medicine 376, 2507, https://doi.org/10.1056/NEJMp1702071 (2017).
Choi, E. et al. In Advances in Neural Information Processing Systems. 3504–3512 (2016).
Zhang, J., Kowsari, K., Harrison, J. H., Lobo, J. M. & Barnes, L. E. Patient2Vec: A Personalized Interpretable Deep Representation of the Longitudinal Electronic Health Record. IEEE Access, https://doi.org/10.1109/ACCESS.2018.2875677 (2018).
Rajkomar, A. et al. Scalable and accurate deep learning with electronic health records. npj Digital Medicine 1, 18, https://doi.org/10.1038/s41746-018-0029-1 (2018).
Hochreiter, S. & Schmidhuber, J. Long short-term memory. Neural computation 9, 1735–1780, https://doi.org/10.1162/neco.1997.9.8.1735 (1997).
Cho, K. et al. Learning phrase representations using RNN encoder-decoder for statistical machine translation. arXiv preprint arXiv:1406.1078 (2014).
Nguyen, P., Tran, T. & Venkatesh, S. Resset: A Recurrent Model for Sequence of Sets with Applications to Electronic Medical Records. arXiv preprint arXiv:1802.00948 (2018).
Vaswani, A. et al. In Advances in Neural Information Processing Systems. 5998–6008 (2017).
Song, H., Rajan, D., Thiagarajan, J. J. & Spanias, A. Attend and Diagnose: Clinical Time Series Analysis using Attention Models. arXiv preprint arXiv:1711.03905 (2017).
Che, Z., Purushotham, S., Cho, K., Sontag, D. & Liu, Y. Recurrent neural networks for multivariate time series with missing values. Scientific reports 8, 6085 (2018).
Cao, W. et al. In Advances in Neural Information Processing Systems. 6775–6785 (2018).
Mozer, M. C., Kazakov, D. & Lindsey, R. V. Discrete event, continuous time rnns. arXiv preprint arXiv:1710.04110 (2017).
Chen, T. Q., Rubanova, Y., Bettencourt, J. & Duvenaud, D. K. In Advances in neural information processing systems. 6571–6583 (2018).
Rubanova, Y., Chen, R. T. & Duvenaud, D. Latent odes for irregularly-sampled time series. arXiv preprint arXiv:1907.03907 (2019).
Dupont, E., Doucet, A. & Teh, Y. W. Augmented neural odes. arXiv preprint arXiv:1904.01681 (2019).
Cai, X. et al. Medical concept embedding with time-aware attention. arXiv preprint arXiv:1806.02873 (2018).
Garland, A., Olafson, K., Ramsey, C. D., Yogendran, M. & Fransoo, R. Epidemiology of critically ill patients in intensive care units: a population-based observational study. Critical Care 17, R212, https://doi.org/10.1186/cc13026 (2013).
Rosenberg, A. L. & Watts, C. Patients readmitted to ICUs: a systematic review of risk factors and outcomes. Chest 118, 492–502 (2000).
Li, P., Stelfox, H. T. & Ghali, W. A. A prospective observational study of physician handoff for intensive-care-unit-to-ward patient transfers. The American journal of medicine 124, 860–867, https://doi.org/10.1016/j.amjmed.2011.04.027 (2011).
Kramer, A. A., Higgins, T. L. & Zimmerman, J. E. The association between ICU readmission rate and patient outcomes. Critical care medicine 41, 24–33, https://doi.org/10.1097/CCM.0b013e3182657b8a (2013).
Rosenberg, A. L., Hofer, T. P., Hayward, R. A., Strachan, C. & Watts, C. M. Who bounces back? Physiologic and other predictors of intensive care unit readmission. Critical care medicine 29, 511–518 (2001).
Ponzoni, C. R. et al. Readmission to the intensive care unit: incidence, risk factors, resource use, and outcomes. A retrospective cohort study. Annals of the American Thoracic Society 14, 1312–1319, https://doi.org/10.1513/AnnalsATS.201611-851OC (2017).
Oye, R. K. & Bellamy, P. E. Patterns of resource consumption in medical intensive care. Chest 99, 685–689 (1991).
Knaus, W. A., Draper, E. A., Wagner, D. P. & Zimmerman, J. E. APACHE II: a severity of disease classification system. Critical care medicine 13, 818–829 (1985).
Le Gall, J.-R., Lemeshow, S. & Saulnier, F. A new simplified acute physiology score (SAPS II) based on a European/North American multicenter study. Jama 270, 2957–2963 (1993).
Johnson, A. E., Kramer, A. A. & Clifford, G. D. A new severity of illness scale using a subset of acute physiology and chronic health evaluation data elements shows comparable predictive accuracy. Critical care medicine 41, 1711–1718 (2013).
Frost, S. A. et al. Severity of illness and risk of readmission to intensive care: a meta-analysis. Resuscitation 80, 505–510, https://doi.org/10.1016/j.resuscitation.2009.02.015 (2009).
Rosa, R. G. et al. Comparison of unplanned intensive care unit readmission scores: a prospective cohort study. PloS one 10, e0143127, https://doi.org/10.1371/journal.pone.0143127 (2015).
Johnson, A. E. et al. MIMIC-III, a freely accessible critical care database. Scientific data 3, 160035, https://doi.org/10.1038/sdata.2016.35 (2016).
Mikolov, T., Chen, K., Corrado, G. & Dean, J. Efficient estimation of word representations in vector space. arXiv preprint arXiv:1301.3781 (2013).
Yang, Z. et al. In Proceedings of the 2016 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies. 1480–1489 (2016).
Hinton, G. & Van Camp, D. In Proc. of the 6th Ann. ACM Conf. on Computational Learning Theory. (1993).
Graves, A. In Advances in neural information processing systems. 2348–2356 (2011).
Blundell, C., Cornebise, J., Kavukcuoglu, K. & Wierstra, D. Weight uncertainty in neural networks. arXiv preprint arXiv:1505.05424 (2015).
Srivastava, N., Hinton, G., Krizhevsky, A., Sutskever, I. & Salakhutdinov, R. Dropout: a simple way to prevent neural networks from overfitting. The Journal of Machine Learning Research 15, 1929–1958 (2014).
Kingma, D. P. & Ba, J. Adam: A method for stochastic optimization. arXiv preprint arXiv:1412.6980 (2014).
Weiss, G. M., McCarthy, K. & Zabar, B. Cost-sensitive learning vs. sampling: Which is best for handling unbalanced classes with unequal error costs? Dmin 7, 24 (2007).
Davis, J. & Goadrich, M. In Proceedings of the 23rd international conference on Machine learning. 233–240 (2006).
Saito, T. & Rehmsmeier, M. The precision-recall plot is more informative than the ROC plot when evaluating binary classifiers on imbalanced datasets. PloS one 10, e0118432 (2015).
Youden, W. J. Index for rating diagnostic tests. Cancer 3, 32–35 (1950).
Efron, B. & Tibshirani, R. J. An introduction to the bootstrap. (CRC press, 1994).
Pedregosa, F. et al. Scikit-learn: Machine learning in Python. Journal of machine learning research 12, 2825–2830 (2011).
Paszke, A. et al. In NIPS 2017 Workshop. (2017).
Cheung, B. L. P. & Dahl, D. In Biomedical & Health Informatics (BHI), 2018 IEEE EMBS International Conference on. 222–225 (IEEE, 2018).
Wong, E. G., Parker, A. M., Leung, D. G., Brigham, E. P. & Arbaje, A. I. Association of severity of illness and intensive care unit readmission: A systematic review. Heart & Lung: The Journal of Acute and Critical Care 45, 3-9. e2, https://doi.org/10.1016/j.hrtlng.2015.10.040 (2016).
Chen, L. M., Martin, C. M., Keenan, S. P. & Sibbald, W. J. Patients readmitted to the intensive care unit during the same hospitalization: clinical features and outcomes. Critical care medicine 26, 1834–1841 (1998).
Santamaria, J. D. et al. Readmissions to intensive care: a prospective multicenter study in Australia and New Zealand. Critical care medicine 45, 290–297, https://doi.org/10.1097/CCM.0000000000002066 (2017).
Kareliusson, F., De Geer, L. & Tibblin, A. O. Risk prediction of ICU readmission in a mixed surgical and medical population. Journal of intensive care 3, 30 (2015).
Woldhek, A. L., Rijkenberg, S., Bosman, R. J. & van der Voort, P. H. Readmission of ICU patients: A quality indicator? Journal of critical care 38, 328–334 (2017).
Ostling, S. et al. The relationship between diabetes mellitus and 30-day readmission rates. Clinical diabetes and endocrinology 3, 3 (2017).
Bottle, A., Aylin, P. & Majeed, A. Identifying patients at high risk of emergency hospital admissions: a logistic regression analysis. Journal of the Royal Society of Medicine 99, 406–414, https://doi.org/10.1177/014107680609900818 (2006).
He, D., Mathews, S. C., Kalloo, A. N. & Hutfless, S. Mining high-dimensional administrative claims data to predict early hospital readmissions. Journal of the American Medical Informatics Association 21, 272–279, https://doi.org/10.1136/amiajnl-2013-002151 (2014).
Hasan, O. et al. Hospital readmission in general medicine patients: a prediction model. Journal of general internal medicine 25, 211–219, https://doi.org/10.1007/s11606-009-1196-1 (2010).
Silverstein, M. D., Qin, H., Mercer, S. Q., Fong, J. & Haydar, Z. In Baylor University Medical Center Proceedings. 363–372 (Taylor & Francis, 2008).
Liu, J., Zhang, Z. & Razavian, N. Deep ehr: Chronic disease prediction using medical notes. arXiv preprint arXiv:1808.04928 (2018).
Author information
Authors and Affiliations
Contributions
S.B. conceived and designed the analysis. S.B. and J.K. performed the analysis. S.K., M.G., A.R. provided clinical input. O.P.-C. and L.J. provided input on the analytical methods. S.B. wrote the manuscript with the help of all other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Barbieri, S., Kemp, J., Perez-Concha, O. et al. Benchmarking Deep Learning Architectures for Predicting Readmission to the ICU and Describing Patients-at-Risk. Sci Rep 10, 1111 (2020). https://doi.org/10.1038/s41598-020-58053-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-58053-z
This article is cited by
-
Prediction of emergency department revisits among child and youth mental health outpatients using deep learning techniques
BMC Medical Informatics and Decision Making (2024)
-
Automated screening of potential organ donors using a temporal machine learning model
Scientific Reports (2023)
-
Predicting unplanned readmissions in the intensive care unit: a multimodality evaluation
Scientific Reports (2023)
-
Assessment of Prediction Tasks and Time Window Selection in Temporal Modeling of Electronic Health Record Data: a Systematic Review
Journal of Healthcare Informatics Research (2023)
-
Current Trends in Readmission Prediction: An Overview of Approaches
Arabian Journal for Science and Engineering (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.