Abstract
Antibiotic-resistant Klebsiella pneumoniae is increasingly being implicated in invasive infections worldwide with high mortalities. Forty-two multidrug resistant (MDR) K. pneumoniae isolates were collected over a 4-month period. Antimicrobial susceptibility was determined using Microscan. The evolutionary epidemiology, resistome, virulome and mobilome of the isolates were characterised using whole-genome sequencing and bioinformatics analysis. All isolates contained the blaCTX-M gene, whilst 41/42(97%) contained blaTEM, 36/42(86%) contained blaOXA and 35/42(83%) harboured blaSHV genes. Other resistance genes found included blaLEN, aac(6′)-lb-cr, qnrA, qnrB, qnrS, oqxAB, aad, aph, dfr, sul1, sul2, fosA, and cat genes. Fluoroquinolone and colistin resistance-conferring mutations in parC, gyrAB, pmrAB, phoPQ and kpnEF were identified. The blaLEN gene, rarely described worldwide, was identified in four isolates. The isolates comprised diverse sequence types, the most common being ST152 in 7/42(17%) isolates; clone-specific O and K capsule types were identified. Diverse virulence genes that were not clone-specific were identified in all but one isolate. IncF, IncH and IncI plasmid replicons and two novel integrons were present. The blaCTX-M-15 and blaTEM-1 genes were bracketed by Tn3 transposons, ISEc9, a resolvase and IS91 insertion sequence. There were 20 gene cassettes in 14 different cassette arrays, with the dfrA and aadA gene cassettes being the most frequent. Phylogenetic analysis demonstrated that the isolates were evolutionarily associated with strains from both South Africa and abroad. These findings depict the rich resistome, mobilome and virulome repertoire in clinical K. pneumoniae strains, which are mainly transmitted by clonal, multiclonal and horizontal means in South Africa.
Similar content being viewed by others
Introduction
Antibiotic resistance (ABR) is a global phenomenon widely described in the literature1,2, and is associated with treatment failure, as well as increased morbidity and mortality3,4,5. Dissemination of resistance in bacteria, particularly among Enterobacteriaceae, is mainly due to the exchange of ABR genes (ARGs) between and within species, mediated by mobile genetic elements (MGEs) harbouring ARGs6,7,8. MGEs include plasmids, transposons and integrons that are known to transmit ABR in both Gram-positive and Gram-negative bacteria, including cephalosporin- and carbapenem-resistant Enterobacteriaceae that have been classified as critical priority pathogens by the WHO9. Integrons that capture cassettes i.e., single gene fragments, usually insert into transposons, enabling their movement between bacteria. Of the eight integron classes described, class 1, 2 and 3 are associated with antimicrobial resistance although class 1 is more frequently described in the literature6,10,11,12.
Among Enterobacteriaceae, Klebsiella pneumoniae is increasingly implicated as an invasive and virulent pathogen that harbours several ARGs13, including AmpCs, extended-spectrum β-lactamases (ESBLs) and carbapenemases. Cephalosporin- and carbapenem-resistant K. pneumoniae have been reported in South Africa to cause several mortalities in Johannesburg (Gauteng Province), Cape Town (Western Cape Province) and KwaZulu-Natal Province in South Africa7,14,15,16,17, including fatal outbreaks among infants18. We subsequently undertook a four-month molecular surveillance of a referral laboratory that serves two major tertiary hospitals in Pretoria, South Africa, to determine the evolutionary epidemiology, resistance mechanisms and associated MGEs in antibiotic-resistant K. pneumoniae isolates.
Methods
Study design
The study sample consisted of 42 multidrug-resistant (MDR) K. pneumoniae isolates that were collected as part of a larger study where consecutive ESBL-positive Enterobacteriaceae, co-resistant to fluoroquinolones and aminoglycosides, were surveilled8,12,19. The isolates were collected from a referral laboratory of the National Health Laboratory Services (NHLS), in Pretoria, South Africa.
Bacterial isolates
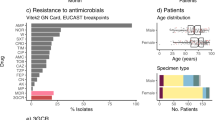
These 42 ESBL-producing K. pneumoniae clinical isolates were identified from urine (n = 14), blood (n = 13), pus/pus swab and other sources (n = 7), sputum (n = 5) and missing (n = 3) specimens of patients admitted to two referral hospitals over a four-month period from September to December in 2013 (Fig. 1). All the samples were collected from infected patients and plated on blood agar (Oxoid, Basingstoke, UK) for 24 hours at 37 °C. They were re-plated unto Mueller-Hinton agar for antibiotic sensitivity testing and ESBL screening using the disc diffusion method: cefoxitin, ceftazidime, and clavulanic acid antibiotic discs20.
Identification and antimicrobial susceptibility testing
The MicroScan WalkWay7465 (Beckman Coulter, Sacramento, USA) instrument was used to identify the species and establish the susceptibility of the isolates to 32 antibiotics: penicillins, cephems, carbapenems, polymyxins, fluoroquinolones, aminoglycosides, tetracyclines, tigecycline, sulphamethoxazole-trimethoprim, nitrofurantoin and fosfomycin (Table S1). The CLSI guidelines were used to interpret the minimum inhibitory concentrations (MICs) (CLSI M100) except for colistin and tigecycline for which the EUCAST breakpoints were used21,22. The taxonomic identity of the isolates was confirmed with the average nucleotide identity (ANI) database of NCBI.
Whole-genome sequencing
Genomic DNA was extracted from the isolates using the ZR Fungal/Bacterial DNA Mini-Prep kit (Zymo Research, Epigenetics, USA). Whole-genome sequencing was performed on the Ion torrent using already described methods23,24. Briefly, the genomic DNA were sheared to 200-bp libraries; 280-bp fragments were selected using 2% agarose gels and Pippen prep (Sage Science, Beverly, MA, USA). Individual libraries were pooled and sequenced on the Ion Proton (ThermoFisher, Waltham, MA, USA). The generated raw reads were de novo assembled using the SPAdes assembler.
Analysis of whole genome sequence data
Assembled sequences were annotated using the ResFinder web server and PGAP to detect resistance genes25,26. The MLST sequence types were identified on the MLST database hosted by the CGE (http://cge.cbs.dtu.dk/services/MLST/). The integrons and gene cassettes within the genomes were identified according to the INTEGRALL database (http://integrall.bio.ua.pt/). Plasmid types were identified using PlasmidFinder on the CGE (http://cge.cbs.dtu.dk/services/PlasmidFinder/) website. The capsule types were identified using Kaptive Web whilst the virulome was characterised using BacWGSTdb27,28. All sequences were deposited at the GenBank data-base and have been allocated accession numbers (Supplementary Tables S1 and S2) under Bioproject PRJNA355910.
Phylogenomic analysis
Whole genome sequences (WGS) from this study were analysed alongside whole-genome sequences of K. pneumoniae isolates curated at the PATRIC website (https.www.patric.org), from where genomes sequenced between 2000 and 2019 were downloaded to enable a current epidemiological and evolutionary analysis; WGS sequences of isolates from Durban, KwaZulu Natal Province, at Genbank (PRJNA287968) were also included. The downloaded sequences (n = 700 isolates) were analysed by RAxML and Parsnp, which was used for the phylogenetic analysis, using the “-C 1000 –c” flag to enable alignment over collinear regions29. The engendered trees were viewed with Gingr (https://harvest.readthedocs.io/en/latest/content/gingr.html) and edited with Figtree (http://tree.bio.ed.ac.uk/software/figtree/).
Results
Patient details
Most of the isolates (n = 36, 86%) were from patients admitted at the Tshwane Academic referral hospital and the rest were from Kalafong. Twenty of the patients were males whilst 22 were females, with ages ranging between 2 and 89 years old and a mean age of 39 years. Most of the samples were either from blood or urine, with five being from sputum (Fig. 1; Supplementary Table S1).
Genomic characteristics
The draft genome sizes of the isolates ranged from 5.2 to 5.7 Mb, with very diverse L50, N50 and contig numbers. The CRISPR arrays in the isolates ranged from zero (n = 21 isolates) to four (n = 1 isolate) (Supplementary Table S1).
Antibiotic susceptibility of isolates
The isolates obtained from the samples were identified phenotypically and genomically as K. pneumoniae. All the isolates were phenotypically positive for extended-spectrum β-lactamase (ESBLs) production. They were mostly resistant or non-susceptible to all the β-lactams except the carbapenems and cephem-β-lactamase inhibitors viz., sulbactam, tazobactam and clavulanic acid. Specifically, all isolates were resistant to aztreonam, cephalothin, cefuroxime, cefotaxime, ceftazidime and cefepime. Notably, resistance to amoxicillin-clavulanate (AUG) and piperacillin-tazobactam was very common compared to cephem-β-lactamase inhibitor combinations. Sixteen isolates had cefoxitin MICs ≥ 16 mg/L. There was variable susceptibility to carbapenems with all isolates demonstrating MICs ≤ 1 mg/L for imipenem and meropenem. Overall, 41 isolates had a doripenem MIC ≤ 1 mg/L, and four isolates, namely K181, K145, K059 and K091, had ertapenem MICs of >1 mg/L (Supplementary Table S2). The phenotypic resistance data tallied largely with the genomic results in that there was no carbapenemase gene found, confirming the absence of carbapenem resistance. As well, the phenotypic ESBL results were confirmed by the presence of blaTEM, blaCTX-M-15, blaSHV, blaOXA and blaLEN genes (Tables 1–2, Figs. 2–7 and supplementary Table S1).
Genetic environment of β-lactamases found in the Klebsiella pneumoniae strains. The genetic environment of the ESBL genes viz., blaCTX-M-15, blaTEM-1B, blaOXA-1/9, and blaSHV were determined using the annotated GFF files from GenBank. The blaSHV genes were mostly found on chromosomes whilst the blaCTX-M-15 and blaTEM-1B, genes were mostly found on Tn3 transposons, ISEc9 and IS91.
Genetic environment of β-lactamases found in the Klebsiella pneumoniae strains. The genetic environment of the ESBL genes viz., blaCTX-M-15, blaTEM-1B, blaOXA-1/9, and blaSHV were determined using the annotated GFF files from GenBank. The blaSHV genes were mostly found on chromosomes whilst the blaCTX-M-15 and blaTEM-1B, genes were mostly found on Tn3 transposons, ISEc9 and IS91.
Genetic environment of β-lactamases found in the Klebsiella pneumoniae strains. The genetic environment of the ESBL genes viz., blaCTX-M-15, blaTEM-1B, blaOXA-1/9, and blaSHV were determined using the annotated GFF files from GenBank. The blaSHV genes were mostly found on chromosomes whilst the blaCTX-M-15 and blaTEM-1B, genes were mostly found on Tn3 transposons, ISEc9 and IS91.
Genetic environment of β-lactamases found in the Klebsiella pneumoniae strains. The genetic environment of the ESBL genes viz., blaCTX-M-15, blaTEM-1B, blaOXA-1/9, and blaSHV were determined using the annotated GFF files from GenBank. The blaSHV genes were mostly found on chromosomes whilst the blaCTX-M-15 and blaTEM-1B, genes were mostly found on Tn3 transposons, ISEc9 and IS91.
Genetic environment of β-lactamases found in the Klebsiella pneumoniae strains. The genetic environment of the ESBL genes viz., blaCTX-M-15, blaTEM-1B, blaOXA-1/9, and blaSHV were determined using the annotated GFF files from GenBank. The blaSHV genes were mostly found on chromosomes whilst the blaCTX-M-15 and blaTEM-1B, genes were mostly found on Tn3 transposons, ISEc9 and IS91.
Genetic environment of β-lactamases found in the Klebsiella pneumoniae strains. The genetic environment of the ESBL genes viz., blaCTX-M-15, blaTEM-1B, blaOXA-1/9, and blaSHV were determined using the annotated GFF files from GenBank. The blaSHV genes were mostly found on chromosomes whilst the blaCTX-M-15 and blaTEM-1B, genes were mostly found on Tn3 transposons, ISEc9 and IS91.
Except for amikacin, the isolates were resistant to all aminoglycosides. Further, all the isolates were resistant to ciprofloxacin but susceptible to norfloxacin; twenty and two were respectively resistant to levofloxacin and nalidixic acid (Supplementary Table S2). Resistance to minocycline (n = 18 isolates) and tetracycline (n = 25 isolates) was common than tigecycline (n = 7 isolates). Resistance to chloramphenicol (n = 33), colistin (n = 10), nitrofurantoin (n = 16) and sulphamethoxazole-trimethoprim (SXT) (n = 42) was identified, with none being resistant to Fosfomycin (Supplementary Table S2). Contrary to the β-lactams, there were substantial discrepancies between the phenotypic and genomic results for the non-β-lactam antibiotics. For instance, the isolates were mostly susceptible to Fosfomycin although fosA genes were ubiquitous among the isolates. Further, the presence of aac(6′)-Ib, OqxAB, Qnr and aac(3′)-Ib genes and mutations in parC and gyrAB only conferred resistance to ciprofloxacin and levofloxacin but not to norfloxacin. A similar observation was made with regards to amikacin and gentamicin/tobramycin (aadA4, aph(3′), strAB, aac(6′)-Ib-cr) and between minocycline, tetracycline and tigecycline (tet). However, the presence of chloramphenicol and SXT resistance was mostly corroborated by the presence of the appropriate resistance genes i.e., cml/Cat and sul/dfrA. For some strains and antibiotics, no known resistance determinant was found to explain the observed phenotypic resistance (Supplementary Tables S1 and S2).
ARGs in Klebsiella pneumoniae
Several resistance genes were present in the isolates (Table 2 and Supplementary Table S1). Many isolates had at least two β-lactamase genes, with blaCTX-M-15 (n = 42), blaOXA and blaTEM being commonest (Figs. 2–7). The blaTEM gene was present in 41/42 (97.6%) isolates, blaOXA was present in 36/42 (85.7%) isolates and blaSHV in 35/42 (83.3%). We identified three blaTEM genes, of which the most frequent was blaTEM-1B. Isolate K021 did not have a TEM β-lactamase but had the blaOXA and blaCTX-M-15 genes. WGS revealed several blaSHV and blaOXA genes, with the most diverse being found in the blaSHV-containing isolates. The blaLEN gene, first identified by Arakawa et al. in 198630, was detected in four isolates, namely K053, K126, K137 and K146.
The Qnr, Oqx and aac(6′)-lb-cr plasmid-mediated quinolone resistance (PMQR) genes were detected in fluoroquinolone-resistant isolates. Of the isolates with the qnr gene, the qnrB fluoroquinolone resistance gene was the most frequently identified (20/42, 47.6%). Both the qnrB6 and qnrB66 genes were identified in these isolates. Six out of 42 (14.3%) and 5/42 (11.9%) isolates had the qnrA and qnrS genes, respectively. The oqxA and oqxB genes were both identified in 36/42 (85.7%) isolates. The most frequently identified PMQR gene was the aac(6′)-lb-cr gene, which was detected in 38/42 (90%) isolates. Six other aac(6′)-lb genes were identified. Twenty-one (50%) isolates had mutations in the chromosomally encoded parC, gyrA and gyrB quinolone resistance-determining region (QRDR) genes (Table 3).
The aac, aad and aph aminoglycoside-modifying enzyme genes were detected in all isolates: several aac genes such as aac(3′)−11a (35/42, 83.3%), aac(3)−11d (4/42, 9.5%), aac (6′)−1b (6/42, 14.3%) and aacA4 (10/42, 23.8%) were found. We identified aad modifying enzymes, including aadA1 in 26/42 (61.9%) isolates, aadA2 in 3/42(7.1%) isolates, aadA5 in 4/42 (9.5%) isolates and aadA16 in 3/42 (7.1%) isolates. The aph(3′)−1a gene was detected in 7/42 (16.7%) isolates. Streptomycin resistance genes, strA and strB were found in 35/42 (83%) and 34/42 (81%) isolates, respectively (Table 1).
The fosA gene, which codes for fosfomycin resistance was identified in 40/42 (95%) isolates. No other fos gene was identified. Trimethoprim–sulfamethoxazole resistance was encoded by sul and dfr genes. We identified sul1 in 33/42 (78%) isolates and sul2 in 36/42 (86%) isolates. We identified several dfr genes, including dfrA4 in 15/42 (36%) isolates, dfrA1 in 14/42 (33%) isolates, dfrA27 in 8/42 (19%) isolates, dfrA15 in 6/42 (14%) isolates, dfrA12 in 3/42 (7%) isolates, dfrA7 in 1/42 (2%) isolates and dfrA30 in 1/42 (2%) isolate. Both the fosfomycin and trimethoprim-sulfamethoxazole genes were co-carried together with the β-lactamase, aminoglycoside and fluoroquinolone resistance genes (Figs. 2–7; Table 1; Supplementary Table S2).
No plasmid-mediated colistin resistance gene was identified in the isolates having increased colistin MICs. Chromosomally encoded mutations in the pmr, pho and kpn genes were, however, identified in 9/11 (82%) of these isolates (Table 1). No novel putative colistin ARGs were identified on any of the available databases. Isolates with ccrB mutations, however, were susceptible to colistin and no truncation were observed in the mgrB in all the isolates (Table 3).
Two isolates contained two different exporter, efflux pump genes for chloramphenicol. Isolate K120 had the floR gene and isolate K021 had the cmlA1 gene. Several chloramphenicol acetyltransferase genes were also identified: catB3 in 34/42 (81%) isolates, catA1 in 23/42 (55%) isolates and catA2 in 8/42 (19%) isolates. Isolates with chloramphenicol resistance had one or more transferase genes. The rifampicin ADP ribosylating transferase ARR−3 gene was detected in 12/42 (29%) isolates. Moreover, the ARGs were not influenced by the isolation source of the isolates as isolates from blood, urine; in fact, in some cases isolates from sputum had more ARGs than those from urine or blood (Supplementary Table S1).
Sequence types and the genetic environment of the ARGs
We detected 11 different sequence types in the isolates (Table 3). The most prevalent sequence types were the ST152 (n = 9, 33%) and ST1552 (n = 6, 22%). Both sequence types were associated with integron ln369 (Table 1). The globally distributed ST15 was only detected in four isolates (Tables 1–2).
As well, we identified the IncF, IncN and IncH incompatibility plasmid replicons (Supplementary Table 1), with nine isolates carrying multiple plasmid replicons simultaneously. The IncF plasmid group was most frequently identified. Nine of the isolates with this IncF incompatibility group also harboured the IncH group. Several unknown plasmid sequence types were also identified (Table 3). The IncF group was also associated with most of the STs.
All isolates contained only class 1 integrons. Several isolates had multiple integrons; K058, K025, K031, K051, K059, K061, K078, K086, K080, K145, K118, K038 K054 and K117. These integrons circulated between both tertiary hospitals (Supplementary Table S2). Two novel class 1 integrons were identified in two isolates, K021 and K145, which were given new numbers, In1481 and ln1482, respectively. The isolates were registered in the GeneBank database with specific accession numbers NXIU000108 (In1481) and NXKB01000066 (ln1482).
The most frequent integron on the IncF plasmid was In369, which captured the dfrA1b – aadA1b cassette array and has been described in other Enterobacteriaceae. Two isolates also harboured the aacA4cr cassette gene. This integron was more frequently associated with ST1552 and ST152 but was also found in two ST101 isolates and one ST234 isolate. In27, a narrow spectrum integron that has been described in the literature was only identified in ST234 isolates capturing the dfrA12-gcuF–aadA2 cassette array, confirming its narrow spectrum status. In191 was, however, identified in four different sequence types, indicating that it is not a narrow spectrum integron.
Most isolates had gene cassettes associated with trimethoprim and aminoglycoside resistance (Table 3). The frequency, in descending order, was dfrA in 38/42 (90%) isolates, aadA in 25/42 (59%) isolates, aacA and arr in 4/42 (9%) isolates and gcu in 3/42 (7%) isolates. Ten different dfr gene cassettes were identified, the most common being dfrA1b (n = 20), dfrA14b (n = 9) and dfrA15 (n = 6). Of the six aadA cassettes identified, the most frequent were aadA1b (n = 8) and aadA1a (n = 6). Only two aacA cassettes were identified, aacA4cr (n = 3) and aacA4 (n = 2). A gcu gene cassette of unknown function was identified in three isolates. The most frequent gene cassette array was the dfrA1b-aadA1b cassette array, responsible for trimethoprim and streptomycin resistance. No β–lactamase cassette was captured in any of the isolates.
Except for blaSHV genes, which were mostly chromosomal and not associated with any transposon, integron (resolvase/recombinase) and insertion sequence (IS), the blaTEM, blaCTX-M-15 and blaOXA genes were mostly bracketed by mobile genetic elements. In particular, blaTEM and blaCTX-M-15 genes co-existed within composite Tn3 transposons, with the blaCTX-M-15 being directly joined to an ISEc9 IS; blaTEM was mostly bracketed by a resolvase, an IS91 and aph(6)-Id, aph(3′)-Ib, and sul2. The genetic context of blaCTX-M-15 in K085 strongly suggests its presence on the chromosome (Fig. 4f). The blaOXA genes were also mostly associated with aac(6′)-Ib-cr and cat genes as well as a resolvase/recombinase (integrase). Notably, these associations were not clone specific as they occurred across different STs (Figs. 2–7).
Virulome and capsular characteristics
A total of 62 virulence genes were identified in all the strains, with EC588_3547, ecpABCR, entB, fepC, fimABCDEFGHK, mrkABC, pulBCED and rpoS occurring in almost all the isolates except for K094, which had no virulence gene (Fig. 8). Among the isolates, EC0103_3368, EC55989_3335, APEC01_3698, ECP_2822 and cah were the least occurring, with cah being only present in K090. As shown in Fig. 8c, the virulome was not clone-specific in that isolates of the same clone had different virulence genes. Although most of these virulence genes occurred in isolates obtained from urine (n = 653) and blood (n = 605), their distribution does not suggest their association with these sources (Fig. 8). The highest number of virulence genes to occur in a single isolate was 57 (K031) whilst all but K094 had more than 30 virulence genes in a single isolate (Fig. 8c). hypervirulence genes were however absent.
(a–d) Virulence genes distribution frequency and their association with specific K. pneumoniae clones and specimen sources. The virulence genes were not clone specific as same clones had different virulence genes (a). The virulence genes distribution and frequency were also not determined the specimen sources (a). K094 had no virulence gene, capping the highest frequency of virulence genes per isolate to 41 with the lowest (cah) being 1 (b). The highest number of virulence gene per clone was 57, with K094 having none (c). Strains from urine and blood had the highest number of virulence genes, which could obviously be due to the relatively larger number of strains isolated from urine and blood (d).
The O and K capsules types in the strains were highly clone specific, with same clones having the same O and K capsule types. However, minor discrepancies were observed as some strains of the same clone had O and K capsule types that were different from those of members of the same clone. A case in point is that of K031 of ST152, which had OL102 and KL107 while other members of the same clone had O4 and KL149 (Figs. 9–12; Supplementary data S3). As can be observed, the O1v1 capsule type was more dominant across the clones whilst KL149 was the most dominant K capsule type. In all, there were eight different O capsule types whilst 12K capsule types were identified, evincing the higher diversity of the K capsule types. It is worthy of mention that isolate K094 had no O or K capsule, just as it also had no virulence genes (Figs. 9–12; Supplementary data 3), which is a very interesting finding
Evolutionary phylogenomics and epidemiology
The K. pneumoniae isolates showed significant phylogenetic diversity (Fig. 13), with the whole-genome phylogenetics showing higher resolution than the MLST typing scheme. For instance, K080 (ST234) was phylogenetically closer to K038 (ST1552) than other ST1552 strains, which were themselves found on different branches, albeit of the same clade. Further, K129 (ST643) and K161 (ST25) were of very close evolutionary distance, albeit of the different clones and capsular types. However, strains of the same MLST clustered within the same clade, with some single STs clusteing closely. Examples include K118 (ST14) within the ST15 clade, K137 (ST182) within the ST101 clade, K014 (ST607) within the ST152 clade, K123 within the ST234 clade, and K090 (ST323) within the ST1414 clade. Interestingly, K120 (ST17), K001 (ST179) and K094 were phylogenetically distant from every strain within the collection (Fig. 13A).
(A–E) Phylogenomic characterisation of the K. pneumoniae strains and their evolutionary relationship with African and International strains. Isolates of the same clone (ST) clustered together, although some strains of different STs were found within or close to strains of same STs (A). Isolates from South Africa are coloured in blue letters while those from this study have red branches. Strains from South Africa were largely clustering together albeit the isolates also clustered with strains from Nigeria, Cameroon, Uganda and Sudan that had the same or closely related ST. Notably, strains from Durban (South Africa) were more closely related to our strains (B). Globally, the strains were related to clones from Belgium, Brazil, China, Ghana, India, Lebanon, Thailand, UK, and USA (C–E). RAXmL and Parsnp were used to draw the trees, which were subsequently annotated with Figtree.
Within the African context, the isolates (coloured with red branches and labelled in blue) were largely phylogenetically related to other K. pneumoniae strains of the same clones (STs) and clades such that strains of the same clones were clustered together within the same clade. This can be seen with K120 (ST17), which clustered with same clinical clones from Nigeria and South Africa; K0179 and K145 (ST39) with a ST38 strain from Nigeria, a ST39 strain from Cameroon and a strain from Uganda. This pattern is observed around the tree under the respective STs (Fig. 13B). However, as observed above, strains of different STs were also found clustered together; this can be seen with K137 (ST182) and K129 (ST643) and PR042E3 (ST31) from pigs in Cameroon (Fig. 13B; Supplementary data S4).
Globally, the STs clustered with same or closely related STs from Austria, Brazil, China, India, Lebanon, Thailand, UK, and USA (Fig. 13C–E; Supplementary data S4).
Discussion
The molecular mechanisms of resistance and virulence dissemination in clinical K. pneumoniae circulating in two referral hospitals in South Africa were characterised and found to be richly endowed with diverse determinants of resistance, virulence and mobile-genetic elements. Notably, the isolates were MDR to several clinically important antibiotics except for reserved ones such as the carbapenems, colistin and tigecycline. The presence of these MDR strains in specimens from this diverse patient demographics in two important referral hospitals make this a very worrying finding. Particularly, same STs were identified in both referral hospitals, suggesting their circulation in both health centres. As expected, the strains’ phenotypic resistance characteristics correlated with known genetic mediators of resistance except for amikacin, nalidixic acid, minocycline, and fosfomycin for which the presence of resistance genes such as aac, aad and aph, tet(A/D/J) and fosA led to no phenotypic resistance. Whereas we could not undertake expression analyses to determine the expression state of these genes, we suspect that the lack of resistance in their presence could be due to little or no expression.
The presence of the blaCTX-M, blaTEM, blaOXA, and blaSHV ESBL genes in similar genetic contexts have been described previously in same and different species in South African and international isolates7,8,12,19,20,31,32,33,34,35,36. The presence of these ARGs within the same genetic context and on the same plasmid replicons across same and different species around the globe strongly suggests the clonal and plasmid-mediated spread of these ARGs. Specifically, ISEc9 and IncF plasmids have been shown to mobilize and facilitate the global spread of blaCTX-M-15, alongside aac(6′)Ib-cr, blaOXA-10 and blaTEM, across species6,7,12,37. Thus, it is not surprising to have the ISEc9 and IncF plasmid replicons dominating in these strains that all harboured blaCTX-M-15 alongside aac(6′)Ib-cr, blaOXA-10 and blaTEM. Interestingly, the same genetic context around the aac(6′)Ib-cr, blaOXA-10 and blaTEM genes were also observed in E. coli strains from the same hospitals, suggesting plasmid-mediated circulation of these genes within these academic hospitals12.
blaCTX-M-15 being present in all the isolates is worth noting, but is not new as an earlier study also found this gene in all the Enterobacteriaceae species studied7. Other studies from South Africa have described the presence of the blaCTX-M-15 gene in E. coli isolates and more recently, in K. pneumoniae7,12,33. We also report a higher prevalence (87.5%) of OXA β–lactamase genes in these K. pneumoniae isolates as well as blaSHV in 83.3% of isolates, confirming that this supposedly chromosomally encoded gene, is not universally found in K. pneumoniae species38. Interestingly, four isolates also contained the narrow spectrum, chromosomally encoded blaLEN gene, comprising of blaLEN9 and blaLEN12, which are rare in South Africa and Africa although it has been previously described in Kenya39. The blaLEN β-lactamase gene was first identified by Arakawa30.
The global dominance of the IncF plasmid, a MGE associated with HGT, is thus herein confirmed6,7,40. As well, an association between IncF replicons and multi-drug resistance (MDR) was observed, as reported globally6,10,11. The presence of several plasmids in which integrons that capture cassette genes are located has been shown to cause MDR6,10,11. The rich repertoire of ARGs in these isolates suggest the presence of one or multiple plasmids, corroborated by the plasmid replicons (Supplementary Table 1).
The integrons identified herein contained diverse gene cassettes, which are novel in the South African context. Particularly, the dfrA and aadA gene cassettes corroborates the global spread of these MGEs41,42. Moreover, aadA1 types, aadA1b and aadA1a, which are different from results reported in K. pneumoniae from Korea where aadA2 types were more frequently identified, were observed. These differences signify subtle changes in genetic composition at a local level11,13,31. Overall, 20 gene cassettes in 14 different cassette arrays, which have not been described previously, were observed. Significantly, the most prevalent cassette arrays viz., dfrA1b-aadA1a and dfrA1b-aadA1b, differed from those described by Partridge et al., which included aadA1a, aadA2 and aadB cassettes11,13.
Integron In369, which captured the dfrA1b – aadA1b cassette array was identified for the first time in South African isolates. This integron and cassette array were also reported in a Portuguese environmental study43. In the current study, this integron was more frequently associated with ST1552 and ST152 but was also found in two ST101 isolates, suggesting its broad host range and promiscuity. Interestingly, integron 27, capturing the dfrA12-gcuF–aadA2 cassette array was only identified in ST234 isolates. Integron 191 was however identified in four different STs suggesting a diversity of clones with this MGE. The dominance of the class 1 integron in this study is also consistent with genomes found in Africa, Europe and South America8,31,41,44. Two novel integrons identified had the aadA16 cassette variant in K021 as well as dfr30b and dfrA14b cassette variants in K145, confirming the ongoing evolutionary processes in these genomes resulting in the diversity of gene cassettes in K. pneumoniae.
Tn3 transposons, which bracketed blaCTX-M-15, blaTEM-1B, aac(6′)-Ib-cr, 6′)-r, aph(6)-Id, aph(3′) and sul2 with myriad ISs in the isolates (Figs. 2–7), commonly encode resistance to β-lactams45,46,47,48; this synteny and genetic environment were also observed in E. coli from the same hospitals12. This rich diversity of transposons and ISs obviously contribute to the genome plasticity, ARGs composition and HGT of ARGs within and across the strains6,8,10. None of the isolates carried Tn21, which was shown to carry multiple resistance genes in a Kenyan study, and was associated with the transfer of antimicrobial resistance in these isolates44.
Our findings suggest plasmid and chromosomally mediated quinolone resistance genes in the clinical K. pneumoniae isolates. The simultaneous presence of qnr, oqx and aac(6″)-lb-cr PMQR genes in these quinolone-resistant isolates and the dominance of the latter is consistent with the literature49,50,51,52,53. This is the second report of the oqxAB gene being found in Africa, although previously described in carbapenemase-producing Enterobacteriaceae51,54. Significantly, we did not detect the qep efflux pump gene which is not frequently reported55,56. In this study, 90% of isolates contained the aac(6′)-lb-cr gene, which is of higher prevalence than studies reported from Spain, Uruguay and Sweden53,57,58,59, which could be explained by the localised spread of related sequence types carrying this gene. Mutations such as S83A and D87A/G in gyrA, D553V and Q/L657G/M in gyrB and S80I and N304S in parC were also found in K. pneumoniae and other Enterobacteriaceae in Durban, South Africa51. Coupled with the PMQR genes, these mutations underlies the resistance to fluoroquinolones to some of the strains (Table 3), albeit no resistance was expressed towards norfloxacin60.
Hospital-acquired Gram-negative infections are usually caused by multi-drug resistant organisms, limiting options available for treating such patients1,2,3,61. Herein, these isolates from hospitalised patients simultaneously contained ESBL β-lactamase genes and PMQR genes, as well as co-resistance to other antibiotic classes, implying that significant antibiotic use causes resistance, co-selection of resistance genes and more significantly, HGT (horizontal gene transfer)3,54,62. These genes have been shown to be co-transmitted on plasmids and other MGEs6,11.
All the isolates in this study were trimethoprim-resistant, although trimethoprim-sulfamethoxazole is no longer recommended for treatment of outpatient conditions such as urinary tract infections in South Africa and internationally because of the high prevalence of resistance63,64,65. Similarly, fosfomycin, used in the outpatient setting for urinary tract infections may not be efficacious since most isolates had the fosA gene, a finding only recently described in Enterobacteriaceae in South Africa7,64,65. The presence of chromosomal colistin resistance in nine out of the 11 ESBL-containing isolates, with an MIC ≥ 4 g/mL, is of concern (Table 3). However, we were unable to confirm these with the broth microdilution, which is the recommended method for colistin resistance determination66. No mcr gene was found, suggesting that these were vertically acquired or engendered de novo.
The diversity and complexity of the virulome and capsule types identified among the strains are concerning as they are implicated in virulence. Fortunately, no hypervirulence genes were found albeit capsule type K2, identified in some strains (Figs. 9–12), are associated with increased virulence and resistance67,68,69. It is interesting to note that the K2 serotype was also identified in a ST14 strain (K118) in this study as was reported in China recently69. The highly clone-specific nature of the O and K capsule types suggest their conserved nature within the genome, contrary to the virulence genes, which differed even within clones (Figs. 3 and 4). The diversity of the virulence genes within clones from the same hospital setting suggest that they were mostly acquired horizontally rather than vertically and could be associated with plasmids. We were unfortunately unable to determine their mobility and presence on plasmids. Further, no association could be established between the specimen sources and the virulome or capsule types. K094 presents a very interesting observation in that it contained no virulence gene or capsules, had no mutations in genes conferring resistance to fluoroquinolones and colistin, contained very few resistance genes (n = 8) and was phylogenetically distant from all STs and clades (Figs. 3–5).
The diversity of sequence types identified in these MDR K. pneumoniae isolates is consistent with findings reported from South, Central and North America, Europe, Asia and North Africa70,71,72,73,74. While ST152 and ST1552 were dominant in this study, and related to Chinese, Ghanaian and Thai genomes, they are not often described in the literature, compared to ST238, an ST responsible for many CRE outbreaks globally6,75,76. The only globally reported ST identified was ST15, confirming the global diversity of this clonal group that was identified in 15% of K. pneumoniae CRE isolates in an international multicentre study encompassing Morocco, Cameroon, Senegal, Madagascar and Vietnam77. ST152 has also been reported in Durban14, as a major clone in Cuba77 and as a major career of NDM-1 in Saudi Arabia78. However, ST147 and ST258, which are global STs associated with antimicrobial resistance79,80,81, were not detected in these isolates. Most sequence types had global spread, being more frequently related to Ghanaian, Thai and Chinese isolates. Four sequence types, viz., ST1552, ST234, ST1414 and ST152 also demonstrated local spread. Local and international outbreaks, in both South Africa and abroad, were also observed in the trees under distinct clades.
An analysis of the molecular epidemiology of these isolates confirmed the international dissemination of specific STs between South Africa and the world. Clonal similarity of isolates from South Africa, Thailand, Nigeria and China was evident although there was a diversity of sequence types associated with other countries. There was also similarity in clones within South Africa; specifically, between Durban and Pretoria. The resolution power of whole-genome sequencing over MLST is demonstrated herein by the clustering of different STs on the same branch and clade, supporting the need to shift to genomic epidemiology for better epidemiological surveillance and infection control.
Conclusion
The burden of ARGs and virulence genes in these isolates from hospitalised patients in two major referral hospitals within Pretoria confirm the global threat of ABR, mediated by MGEs. The findings demonstrate the centrality of MGEs in defining the resistome of MDR strains. Phylogenetic analysis confirmed this global spread including evolutionary relationships of the different STs. IncF plasmid replicons and class1 integrons, both of which have been globally reported, were also dominant in these isolates. Significantly, two novel integrons were identified. The presence of the rare chromosomal blaLEN gene in four isolates is also notable.
Ethics
Ethical approval was provided by the Human Research Ethics Committee of the University of Witwatersrand (Ref M1710100). All protocols and consent forms were executed according to the agreed ethical approval terms and conditions. All clinical samples were obtained from a reference laboratory and not directly from patients, who agreed to our using their specimens for this research. The guidelines stated by the Declaration of Helsinki for involving human participants were followed in the study.
References
World Health Organization. WHO|Antimicrobial resistance. (2017).
WHO. Global Antimicrobial Resistance Surveillance System (GLASS) Report. https://doi.org/10.1016/S1473-3099(18)30060-4 (2017).
Asante, J. & Osei Sekyere, J. Understanding antimicrobial discovery and resistance from a metagenomic and metatranscriptomic perspective: Advances and applications. Environ. Microbiol. Rep. 1–25, https://doi.org/10.1111/1758-2229.12735 (2019).
Rule, R., Said, M., Mbelle, N. & Osei Sekyere, J. Genome Sequence of a Clinical Salmonella Enteritidis Sequence Type 11 Strain from South Africa. J. Glob. Antimicrob. Resist. https://doi.org/10.1016/j.jgar.2019.09.014 (2019).
Rule, R. et al. A rare case of Colistin-resistant Salmonella Enteritidis meningitis in an HIV-seropositive patient. BMC Infect. Dis. 19, 806 (2019).
Kopotsa, K., Osei Sekyere, J. & Mbelle, N. M. Plasmid Evolution in Carbapenemase-Producing Enterobacteriaceae: A Review. Ann. New Accepted (2019).
Pedersen, T. et al. Spread of Plasmid-Encoded NDM-1 and GES-5 Carbapenemases among Extensively Drug-Resistant and Pandrug-Resistant Clinical Enterobacteriaceae in Durban, South Africa. Antimicrob. Agents Chemother. AAC. 02178–17, https://doi.org/10.1128/AAC.02178-17 (2018).
Mbelle, N. M. et al. Genomic analysis of a multidrug-resistant clinical Providencia rettgeri (PR002) strain with the novel integron ln1483 and an A/C plasmid replicon. Ann. N. Y. Acad. Sci. Accepted (2019).
World Health Organization. Global Priority List Of Antibiotic-Resistant Bacteria To Guide Research, Discovery, And Development Of New Antibiotics. WHO (World Health Organization, 2017).
Stokes, H. W. & Gillings, M. R. Gene flow,mobile genetic elements and the recruitment of antibiotic resistance genes into Gram-negative pathogens. FEMS Microbiol. Rev. 35, 790–819 (2011).
Partridge, S. R., Kwong, S. M., Firth, N. & Jensen, S. O. Mobile Genetic Elements Associated with Antimicrobial Resistance. Clin. Microbiol. Rev. 31, 1–61 (2018).
Mbelle, N. M. et al. The Resistome, Mobilome, Virulome and Phylogenomics of Multidrug-Resistant Escherichia coli Clinical Isolates from Pretoria, South Africa. Sci. Rep. Accepted, 1–43 (2019).
Partridge, S. R. Analysis of antibiotic resistance regions in Gram-negative bacteria. FEMS Microbiol. Rev. 35, 820–855 (2011).
Founou, R. C., Founou, L. L., Allam, M., Ismail, A. & Essack, S. Y. Whole Genome Sequencing of Extended Spectrum β-lactamase (ESBL)-producing Klebsiella pneumoniae Isolated from Hospitalized Patients in KwaZulu-Natal, South Africa. Sci. Rep. 9, 6266 (2019).
Founou, L. L. et al. Genome Sequencing of Extended-Spectrum beta-Lactamase (ESBL)-Producing Klebsiella pneumoniae Isolated from Pigs and Abattoir Workers in Cameroon. Front. Microbiol. 9, 188 (2018).
NICD. Carbapenem-resistant Klebsiella pneumoniae outbreak at a hospital in the City of Cape Town Metropolitan Municipality, May 2018-February 2019. (2019).
Jacobson, R. K. et al. Molecular characterisation and epidemiological investigation of an outbreak of blaOXA-181 carbapenemase-producing isolates of Klebsiella pneumoniae in South Africa. S. Afr. Med. J. 105, 1030–5 (2015).
Anonymous. Sixth baby dies in Klebsiella outbreak. news24 (2018).
Mbelle, N. et al. Genomic Analysis of Two Drug‐Resistant Clinical Morganella morganii Strains Isolated from UTI Patients in Pretoria, South Africa. Lett. Appl. Microbiol. Accepted, lam.13237 (2019).
Chirindze, L. M. et al. Faecal colonization of E. coli and Klebsiella spp. producing extended-spectrum beta-lactamases and plasmid-mediated AmpC in Mozambican university students. BMC Infect. Dis. 18, 244 (2018).
Clinical and Laboratory Standards Institute (CLSI). Performance standards for Antimicrobial Susceptibility Testing; Twenty-Seventh Informational Supplement M100-S27. (CLSI, Wayne, PA, USA, 2019).
European Committee on Antimicrobial Susceptibility Testing. Breakpoint tables for interpretation of MICs and zone diameters. Version 9.0, 2019. http://www.eucast.org/fileadmin/src/media/PDFs/EUCAST_files/Breakpoint_tables/v_9.0_Breakpoint_Tables.pdf (2019).
Mbelle, N. M. et al. First Report of a Whole-Genome Shotgun Sequence of a Clinical Enterococcus faecalis Sequence Type 6 Strain from South Africa. Genome Announc. 5, e01382–17 (2017).
Mbelle, N. M. et al. Draft Genome Sequence of a Clinical Enterococcus faecium Sequence Type 18 Strain from South Africa. Genome Announc. 5, e01381–17 (2017).
Kleinheinz, K. A., Joensen, K. G. & Larsen, M. V. Applying the ResFinder and VirulenceFinder web-services for easy identification of acquired antibiotic resistance and E. coli virulence genes in bacteriophage and prophage nucleotide sequences. Bacteriophage 4, e27943 (2014).
Tatusova, T. et al. NCBI prokaryotic genome annotation pipeline. 44, 6614–6624 (2016).
Wyres, K. L. et al. Identification of Klebsiella capsule synthesis loci from whole genome data. Microb. genomics 2, e000102 (2016).
Ruan, Z. & Feng, Y. BacWGSTdb, a database for genotyping and source tracking bacterial pathogens. Nucleic Acids Res. 44, 682–687 (2016).
Treangen, T. J., Ondov, B. D., Koren, S. & Phillippy, A. M. The Harvest suite for rapid core-genome alignment and visualization of thousands of intraspecific microbial genomes. Genome Biol. 15, 524 (2014).
Siebor, E., Péchinot, A., Duez, J.-M. & Neuwirth, C. One new LEN enzyme and two new OKP enzymes in Klebsiella pneumoniae clinical isolates and proposed nomenclature for chromosomal beta-lactamases of this species. Antimicrob. Agents Chemother. 49, 3097–8 (2005).
Shin, J. H. et al. High Rates of Plasmid-Mediated Quinolone Resistance QnrB Variants Among Ciprofloxacin-Resistant Escherichia coli and Klebsiella pneumoniae from Urinary Tract Infections in Korea. Microb. Drug Resist. 14, 221–226 (2008).
Mhlongo, N., Essack, S. & Govinden, U. NDM-1, novel TEM-205, novel TEM-213 and other extended-spectrum β-lactamases co-expressed in isolates from cystic fibrosis patients from South Africa. South. African J. Infect. Dis. 30, 103–107 (2015).
Peirano, G., van Greune, C. H. J. & Pitout, J. D. D. Characteristics of infections caused by extended-spectrum β-lactamase–producing Escherichia coli from community hospitals in South Africa. Diagn. Microbiol. Infect. Dis. 69, 449–453 (2011).
Fallah, F. et al. Prevalence of bla NDM, bla PER, bla VEB, bla IMP, and bla VIM Genes among Acinetobacter baumannii Isolated from Two Hospitals of Tehran, Iran. Scientifica (Cairo). 2014, 245162 (2014).
Fernando, D. M. et al. Detection of Antibiotic Resistance Genes in Source and Drinking Water Samples from a First Nation Community in Canada. Appl. Environ. Microbiol. AEM. 00798-16, https://doi.org/10.1128/AEM.00798-16 (2016).
Cuzon, G., Bonnin, R. A. & Nordmann, P. First identification of novel NDM carbapenemase, NDM-7, in Escherichia coli in France. PLoS One 8, e61322 (2013).
Coque, T. M. et al. Dissemination of clonally related Escherichia coli strains expressing extended-spectrum β-lactamase CTX-M-15. Emerg Infect Dis 14, 195–200 (2008).
Paterson, D. L. Resistance in Gram-Negative Bacteria: Enterobacteriaceae. Am. J. Med. 119, S20–S28 (2006).
Henson, S. P. et al. Molecular epidemiology of Klebsiella pneumoniae invasive infections over a decade at Kilifi County Hospital in Kenya. Int. J. Med. Microbiol. 307, 422–429 (2017).
Carattoli, A. Resistance Plasmid Families in Enterobacteriaceae. Antimicrob. Agents Chemother. 53, 2227–2238 (2009).
Pallecchi, L. et al. Population structure and resistance genes in antibiotic-resistant bacteria from a remote community with minimal antibiotic exposure. Antimicrob Agents Chemother 51, 1179–1184 (2007).
Soliman, A. M. et al. First report in Africa of two clinical isolates of Proteus mirabilis carrying Salmonella genomic island (SGI1) variants, SGI1-PmABB and SGI1-W. Infect. Genet. Evol. 51, 132–137 (2017).
Jones-Dias, D. et al. Architecture of Class 1, 2, and 3 Integrons from Gram Negative Bacteria Recovered among Fruits and Vegetables. Front. Microbiol. 7, 13 (2016).
Kiiru, J., Butaye, P., Goddeeris, B. M. & Kariuki, S. Analysis for prevalence and physical linkages amongst integrons, ISEcp1, ISCR1, Tn21 and Tn7 encountered in Escherichia coli strains from hospitalized and non-hospitalized patients in Kenya during a 19-year period (1992–2011). BMC Microbiol. 13, 109 (2013).
Quiroga, M. P. et al. Distribution and functional identification of complex class 1 integrons. Infect. Genet. Evol. 19, 88–96 (2013).
Norman, A. et al. Nucleotide sequence of pOLA52: a conjugative IncX1 plasmid from Escherichia coli which enables biofilm formation and multidrug efflux. Plasmid 60, 59–74 (2008).
Manageiro, V. et al. Molecular Epidemiology and Risk Factors of Carbapenemase-Producing Enterobacteriaceae Isolates in Portuguese Hospitals: Results From European Survey on Carbapenemase-Producing Enterobacteriaceae (EuSCAPE). Front. Microbiol. 9, 2834 (2018).
Roh, K. H. et al. Isolation of a Klebsiella pneumoniae isolate of sequence type 258 producing KPC-2 carbapenemase in Korea. Korean J. Lab. Med. 31, 298–301 (2011).
Liu, X., Boothe, D. M., Thungrat, K. & Aly, S. Mechanisms accounting for fluoroquinolone multidrug resistance Escherichia coli isolated from companion animals. Vet. Microbiol. 161, 159–168 (2012).
Rodríguez-Martínez, J. M. et al. Plasmid-mediated quinolone resistance: an update. J Infect Chemother 17, 149–182 (2011).
Osei Sekyere, J. & Amoako, D. G. Genomic and Phenotypic Characterisation of Fluoroquinolone Resistance Mechanisms in Enterobacteriaceae in Durban, South Africa. PLoS One 12, 1–14 (2017).
Wachino, J.-i. & Arakawa, Y. Exogenously acquired 16S rRNA methyltransferases found in aminoglycoside-resistant pathogenic Gram-negative bacteria: An update. Drug Resist. Updat. 15, 133–148 (2012).
Karah, N. et al. Plasmid-mediated quinolone resistance determinants qnr and aac(6′)-Ib-cr in Escherichia coli and Klebsiella spp. from Norway and Sweden. Diagn. Microbiol. Infect. Dis. 66, 425–431 (2010).
Osei Sekyere, J. & Asante, J. Emerging mechanisms of antimicrobial resistance in bacteria and fungi: advances in the era of genomics. Future Microbiol. 13, 241–262 (2018).
Ruiz, J., Pons, M. J. & Gomes, C. Transferable mechanisms of quinolone resistance. Int J Antimicrob Agents 40, 196–203 (2012).
Heidary, M. et al. Detection of acrA, acrB, aac(6′)-Ib-cr, and qepA genes among clinical isolates of Escherichia coli and Klebsiella pneumoniae. Acta Microbiol. Immunol. Hung. 64, 63–69 (2017).
Vien, L. T. M. et al. The Co-Selection of Fluoroquinolone Resistance Genes in the Gut Flora of Vietnamese Children. PLoS One 7, e42919 (2012).
Briales, A. et al. Prevalence of plasmid-mediated quinolone resistance determinants qnr and aac(6′)-Ib-cr in Escherichia coli and Klebsiella pneumoniae producing extended-spectrum β-lactamases in Spain. Int. J. Antimicrob. Agents 39, 431–434 (2012).
Garcia-Fulgueiras, V. et al. Extended-spectrum -lactamases and plasmid-mediated quinolone resistance in enterobacterial clinical isolates in the paediatric hospital of Uruguay. J. Antimicrob. Chemother. 66, 1725–1729 (2011).
Aldred, K. J., Kerns, R. J. & Osheroff, N. Mechanism of Quinolone Action and Resistance. Biochemistry 53, 1565–1574 (2014).
Bassetti, M., Russo, A., Carnelutti, A., La Rosa, A. & Righi, E. Antimicrobial resistance and treatment: an unmet clinical safety need. Expert Opin. Drug Saf. 17, 669–680 (2018).
Wang, Y. et al. Occurrence of Highly Conjugative IncX3 Epidemic Plasmid Carrying blaNDM in Enterobacteriaceae Isolates in Geographically Widespread Areas. Front. Microbiol. 9 (2018).
Lewis, D. A. et al. Antimicrobial susceptibility of organisms causing community-acquired urinary tract infections in Gauteng Province, South Africa. S. Afr. Med. J. 103, 377–81 (2013).
McMillan, E. A. et al. Antimicrobial Resistance Genes, Cassettes, and Plasmids Present in Salmonella enterica Associated With United States Food Animals. Front. Microbiol. 10, 1–18 (2019).
Osei Sekyere, J. Genomic insights into nitrofurantoin resistance mechanisms and epidemiology in clinical Enterobacteriaceae. Futur. Sci. OA 4, FSO293 (2018).
Osei Sekyere, J. Mcr colistin resistance gene: a systematic review of current diagnostics and detection methods. Microbiologyopen e00682, https://doi.org/10.1002/mbo3.682 (2018).
Giske, C. G. et al. Diverse Sequence Types of Klebsiella pneumoniae Contribute to the Dissemination of blaNDM-1 in India, Sweden, and the United Kingdom. Antimicrob. Agents Chemother. 56, 2735–2738 (2012).
Dorman, M. J., Feltwell, T., Goulding, D. A., Parkhill, J. & Short, F. L. The Capsule Regulatory Network of Klebsiella pneumoniae Defined by density-TraDISort. MBio 9 (2018).
Mei, Y. et al. Virulence and Genomic Feature of a Virulent Klebsiella pneumoniae Sequence Type 14 Strain of Serotype K2 Harboring blaNDM–5 in China. Front. Microbiol. 8 (2017).
Osei Sekyere, J., Govinden, U. & Essack, S. The Molecular Epidemiology and Genetic Environment of Carbapenemases Detected in Africa. Microb Drug Resist 22, 59–68 (2015).
Nordmann, P., Cuzon, G. & Naas, T. The real threat of Klebsiella pneumoniae carbapenemase-producing bacteria. Lancet Infect Dis 9, 228–236 (2009).
David, S. et al. Epidemic of carbapenem-resistant Klebsiella pneumoniae in Europe is driven by nosocomial spread. Nat. Microbiol. https://doi.org/10.1038/s41564-019-0492-8 (2019).
Kontopoulou, K. et al. Hospital outbreak caused by Klebsiella pneumoniae producing KPC-2 β-lactamase resistant to colistin. J Hosp Infect 76, 70–73 (2010).
Tian, G.-B. et al. MCR-1-producing Klebsiella pneumoniae outbreak in China. Lancet Infect. Dis. 17, 577 (2017).
Giuffrè, M. et al. Successful control of an outbreak of colonization by Klebsiella pneumoniae carbapenemase-producing K. pneumoniae sequence type 258 in a neonatal intensive care unit, Italy. J Hosp Infect In Press, (2013).
Samuelsen, Ø. et al. Emergence of clonally related Klebsiella pneumoniae isolates of sequence type 258 producing plasmid-mediated KPC carbapenemase in Norway and Sweden. J Antimicrob Chemother 63, 654–658 (2009).
Quiñones, D. et al. High Clonal Diversity in a Non-Outbreak Situation of Clinical ESBL-Producing Klebsiella pneumoniae Isolates in the First National Surveillance Program in Cuba. Microb. Drug Resist. 20, 45–51 (2014).
Zaman, T. U. et al. Clonal diversity and genetic profiling of antibiotic resistance among multidrug/carbapenem-resistant Klebsiella pneumoniae isolates from a tertiary care hospital in Saudi Arabia. BMC Infect. Dis. 18, 205 (2018).
Muggeo, A. et al. Spread of Klebsiella pneumoniae ST395 non-susceptible to carbapenems and resistant to fluoroquinolones in North-Eastern France. J. Glob. Antimicrob. Resist. 13, 98–103 (2018).
Sato, T. et al. Tigecycline Susceptibility of Klebsiella pneumoniae Complex and Escherichia coli Isolates from Companion Animals: The Prevalence of Tigecycline-Nonsusceptible K. pneumoniae Complex, Including Internationally Expanding Human Pathogenic Lineages. Microb. Drug Resist. 24, 860–867 (2018).
Carbapenemase, K. et al. Chromosomal Integration of the Klebsiella pneumoniae Carbapenemase Gene, blaKPC,in Klebsiella Species Is Elusive but Not Rare. Antimicrob Agents Chemother 61, 1–12 (2017).
Acknowledgements
This work was made possible by the National Health Laboratory Services (NHLS), who enabled access to the clinical isolates. Olesegun Atanda assisted with the collection of the isolates and Monwabisi Litshie phenotypic analysis. Part of this work was presented at the International Society for Infectious Diseases (ISID) Congress, Buenos Aires, Argentina, 2018 (Abstract #1732) and the American Society of Microbiology (ASM) congress, Atlanta, USA 2018 (Abstract #5351). We are grateful to Hlambani Shirinda and Chane of NHLS/TAD, Department of Medical Microbiology, University of Pretoria, for helping with the collation of the virulence genes. This study was supported by the National Health Laboratories Services Grant #94445, the University of Pretoria Grant #ADW0702 and the South African Medical Research Council.
Author information
Authors and Affiliations
Contributions
N.M.: co-conceptualised study and design. J.O.S.: conceptualised study and design, interpretation, data analysis, bioinformatics of MGEs (resistance genes environment), virulome, mobilome and capsules, phylogenomics, write-up, editing, design of images and formatting of manuscript. N.E.M. and L.M.: isolates’ phenotypic analysis, DNA extraction and curation. S.E.: Co-supervised study. C.F.: Supervised study
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mbelle, N.M., Feldman, C., Sekyere, J.O. et al. Pathogenomics and Evolutionary Epidemiology of Multi-Drug Resistant Clinical Klebsiella pneumoniae Isolated from Pretoria, South Africa. Sci Rep 10, 1232 (2020). https://doi.org/10.1038/s41598-020-58012-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-58012-8
This article is cited by
-
Exploring the resistome, virulome, and mobilome of multidrug-resistant Klebsiella pneumoniae isolates: deciphering the molecular basis of carbapenem resistance
BMC Genomics (2024)
-
Antibiotic resistance and its correlation with biofilm formation and virulence genes in Klebsiella pneumoniae isolated from wounds
Journal of Applied Genetics (2024)
-
Assessment of three antibiotic combination regimens against Gram-negative bacteria causing neonatal sepsis in low- and middle-income countries
Nature Communications (2024)
-
Investigating the evolution and predicting the future outlook of antimicrobial resistance in sub-saharan Africa using phenotypic data for Klebsiella pneumoniae: a 12-year analysis
BMC Microbiology (2023)
-
Third generation cephalosporin-resistant Klebsiella pneumoniae thriving in patients and in wastewater: what do they have in common?
BMC Genomics (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.