Abstract
Pathological staging and histological grading systems are useful, but imperfect, predictors of recurrence in head and neck squamous cell carcinoma (HNSCC). Aberrant promoter methylation is the main type of epigenetic modification that plays a role in the inactivation of tumor suppressor genes. To identify new potential prognostic markers, we investigated the promoter methylation status of five neuropeptide receptor genes. The methylation status of the target genes was compared with clinical characteristics in 278 cases; 72 hypopharyngeal cancers, 54 laryngeal cancers, 75 oropharyngeal cancers, and 77 oral cavity cancers were studied. We found that the NTSR1, NTSR2, GHSR, MLNR, and NMUR1 promoters were methylated in 47.8%, 46.8%, 54.3%, 39.2%, and 43.5% of the samples, respectively. GHSR and NMUR1 promoter methylation independently predicted recurrence in HNSCC. In patients with oropharyngeal cancer (n = 75), GHSR and NMUR1 promoter methylation significantly correlates with survival in surgically treated patients. We classified our patients as having a low, intermediate, or high-risk of death based on three factors: HPV status, and GHSR and NMUR1 promoter methylation. The disease-free survival (DFS) rates were 87.1%, 42.7%, and 17.0%, respectively. Combined data analysis of the methylation status of ten-eleven translocation (TET) family genes indicated a trend toward greater methylation indices as the number of TET methylation events increased. In the current study, we presented the relationship between the methylation status of the GHSR and NMUR1 genes and recurrence in HNSCC, specifically in risk classification of oropharyngeal carcinomas cases with HPV status.
Similar content being viewed by others
Introduction
Head and neck squamous cell carcinoma (HNSCC) includes cancers of the pharynx, larynx, and oral cavity, and constitute approximately 4% of all cancers worldwide, with approximately 500,000 deaths annually1. Alcohol and tobacco consumption are the two most important risk factors for HNSCC, especially for cancers of the larynx, hypopharynx, and oral cavity2. Infection with various types of cancer-causing human papillomaviruses (HPV), primarily type 16, are risk factors for oropharyngeal cancers3. The 5-year overall survival rate of patients with HNSCC is approximately 40–50%4. Although HNSCC is a heterogeneous disease, the based on molecular studies have served to distinguish HPV-positive from HPV-negative HNSCC; however, validated molecular characterizations have not been established5.
G protein-coupled receptors (GPCRs) are the largest class of cell surface receptors involved in the development and progression of many cancers, including HNSCC6. GPCRs have highly druggable sites and comprise the largest class of pharmaceutical targets; at present, over 30% of FDA-approved drugs target GPCRs or their related pathways7. However, there are currently no anticancer drugs that specifically target GPCRs8. Frequent mutations of novel druggable oncogenes are not detected in HNSCC9,10. Epigenetic repression of GPCR genes correlates with worse prognosis and potential therapeutic targets9,10.
Whether patients with head and neck cancer who are regarded to be in the low-risk group could be saved the long-term complications of intensive, multimodal treatment without compromising their survival is now an extremely distinctive clinical question11,12. The initiation, progression, and resistance of cancer, traditionally considered a genetic disease, is now known to involve global epigenetic abnormalities, in addition to genetic alterations13. Epigenetic events are also potential drivers of acquired drug resistance in cancer14. Aberrant promoter methylation, an authentication of cancer cells, accounts for the inactivation of many tumor suppressor genes15. In a previous study, we found that ten-eleven translocation (TET) family genes of promoter region were aberrantly methylated in patients with HNSCC16. The five members of this family, TET1, TET2, and TET3, are enzymes that play a role of 5-methylcytosine oxidase and DNA demethylase17.
The principal aim of this study was to determine the methylation status of five GPCR-encoding genes in HNSCC and its association with survival and clinical parameters (e.g., tumor location and HPV status). All five genes, namely neurotensin receptor 1 (NTSR1), neurotensin receptor 2 (NTSR2), growth hormone secretagogue receptor (GHSR), motilin receptor (MLNR), and neuromedin U receptor 1 (NMUR1), encode neuropeptide receptors and belong to the Class Aβ subgroup clade 4. These five neuropeptide receptors have been implicated in the development of multiple types of cancer, but this study is the first to investigate their roles in the prognosis of HNSCC.
Materials and Methods
Tumor samples
Tissues were sampled (n = 278) from patients undergoing major surgical resection for HNSCC at the Department of Otolaryngology, Hamamatsu University School of Medicine (Hamamatsu, Shizuoka, Japan). All patients gave their written informed consent. Ethical clearance was received by the ethical committee of the Hamamatsu University School of Medicine (date of board approval: October 2, 2015, ethic code: 25–149), and informed consent was obtained from the participants. All methods were performed in accordance with the Declaration of Helsinki. The ratio of males to females was 233:45. The mean age was 65.4 years (age 32 to 92 years). Primary tumors were composed of 72 hypopharyngeal carcinomas, 54 laryngeal carcinomas, 75 oropharyngeal carcinomas, and 77 oral cavity carcinomas. Detailed clinical information was obtained from the patients’ medical records.
DNA extraction and bisulfite modification
DNA was extracted from fresh specimens with a QIAamp DNA Mini Kit (Qiagen, Hilden, Germany) on the day of surgery. Purified genomic DNA was bisulfite-converted using the MethylEasy Xceed Rapid DNA Bisulfite Modification Kit (TaKaRa, Tokyo, Japan) following the manufacturer’s protocol.
Quantitative methylation-specific PCR analysis (Q-MSP)
DNA methylation at CpG sites near promoter regions of the target genes was defined via quantitative methylation-specific PCR analysis (Q-MSP) using the Thermal Cycler Dice Real Time System TP800 (TaKaRa). The sequences of the primers used in this study are presented in Additional File 1: Table S1. Exon one and CpG sites within views of the promoter region relative to the transcription start site are presented in Additional File 2: Figure S118. A standard curve for Q-MSP was constructed by plotting five serially diluted standard solutions of EpiScope Methylated HeLa gDNA (TaKaRa). The normalized methylation value (NMV) was defined as follows: NMV = (GPCRs gene-S/ GPCRs gene-FM)/(ACTB-S/ACTB-FM), where GPCRs gene-S and GPCRs gene-FM represent target gene methylation levels in the tumor sample and universal methylated DNA control, respectively. ACTB-S and ACTB-FM correspond to β-actin (ACTB) in the sample and the universally methylated DNA, respectively.
Detection of high-risk HPV DNA by PCR
For high-risk HPV DNA detection, samples were assessed by PCR using specific primers for HPV types 16, 18, 31, 33, 35, 52, and 58. The prevalence of HPV DNA was analyzed with the PCR HPV Typing Set (TaKaRa). The PCR products were separated by electrophoresis on 9% polyacrylamide gels followed by staining with 0.5 g/mol ethidium bromide.
Data mining in the Cancer Genome Atlas (TCGA)
The MethHC (http://methhc.mbc.nctu.edu.tw/php/index.php) was used to extract data from TCGA (available in April 2019)19. DNA methylation of GPCR genes was measured by Illumina Infinium Human Methylation 450 K BeadChip. The methylation score for each CpG site is represented as β values and ranges from 0 to 1, corresponding to unmethylated and completely methylated DNA, respectively.
Data analysis and statistics
The receiver-operator characteristic (ROC) curve analysis was used to evaluate the NMVs for 36 matched paired HNSCC and normal mucosal samples and the Stata/SE 13.0 system (Stata Corporation, TX, USA). In an area under the ROC curve, the true positive rate (Sensitivity) is plotted as a function of the false positive rate (1-Specificity) for different cutoff points, and the NMV thresholds were estimated for each target gene. Cutoff values showing the greatest accuracy were determined based on sensitivity/specificity, as indicated in Additional File 3: Table S2. We used the cutoff values to determine the methylation frequencies of the target genes. Calculation of the methylation index (MI) was defined as the ratio between the number of methylated genes and the number of tested genes in all the samples16.
Associations between the clinical variables were analyzed using the Student’s t-test. Disease-free survival (DFS) probabilities were estimated using the Kaplan-Meier method, and the log-rank test was applied to assess the significant differences among actuarial survival curves. Cox’s proportional hazards regression analysis that included age (≥65 vs. <65 years), sex, alcohol intake, smoking status, and tumor stage (I–II vs. III–IV), and the methylation status was used to identify the multivariate predictive value of the prognostic factors20. P < 0.05 was considered statistically significant.
Ethics approval and consent to participate
The research methodology employed in this study was approved by the Institutional Review Board of the Hamamatsu University School of Medicine. All study subjects provided written informed consent.
Results
Analysis of the methylation status of HNSCC tissue samples
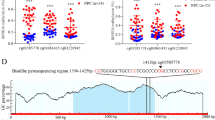
Q-MSP analysis was used to determine the methylation status of five genes encoding GPCR neuropeptide receptors in 278 primary HNSCC samples and it was a valuable test. The methylation frequencies are as follows: NTSR1 (47.8%), NTSR2 (46.8%), GHSR (54.3%), MLNR (39.2%), and NMUR1 (43.5%) (Fig. 1a). The average number of methylated genes per sample was 2.32 ± 1.61(range: 0–5) (Fig. 1b). Primary tumors were located in the hypopharynx (n = 72), larynx (n = 54), oropharynx (n = 75), or oral cavity (n = 77) (Fig. 1c). NTSR1, NTSR2, GHSR, MLNR, and NMUR1 promoter hypermethylation presented discriminative ROC curve profiles, which clearly differentiate cancer tissues from normal tissues [Area Under Curve (AUC) = 0.6220, 0.5736, 0.8103, 0.6049, and 0.5631, respectively] (Additional File 4: Fig. S2). A specimen was classified as methylated when its NMV exceeded 0.045, 0.009, 0.563, 0.700, and 0.735 for NTSR1, NTSR2, GHSR, MLNR, and NMUR1, respectively (Additional File 3: Table S2). NTSR1, NTSR2, GHSR, MLNR, and NMUR1 methylation levels in primary HNSCCs were significantly higher than those in matched paired normal mucosal tissues (Additional File 5: Fig. S3).
Methylation of the neuropeptide receptor gene promoters in 278 HNSCC samples. (a) Bar graph showing the methylation frequencies of the five genes. (b) Bar graph showing the percentage of tumors that express zero to five methylated target genes. (c) Comparison of the methylation status of the promoters of the five genes in patients with hypopharyngeal, laryngeal, oropharyngeal, or oral cancer. Filled boxes indicate the presence of methylation, and open boxes indicate the absence of methylation (d) Bar graph showing the methylation indices (MIs) according to selected clinical parameters. The mean MI for each parameter was determined by the Student’s t-test.
Clinicopathological characteristics of primary HNSCC samples
The MI was determined as the number of methylated genes to the number of tested genes in each sample. No significant differences in MI were observed regarding the age at disease onset, sex, alcohol consumption, smoking status, tumor size, lymph node status, clinical stage, or HPV status (Fig. 1d). Associations between the methylation status of the target genes and the clinical characteristics of the patients are shown in Additional File 6: Table S3. Except for HPV status, there is no significant association between neuropeptide receptor gene promoter hypermethylation and clinicopathological parameters. Methylation of the NTSR1 and GHSR promoters are significantly correlated with HPV status (P = 0.004 and P = 0.038, respectively) (Additional File 6: Table S3). Correlations between HPV status and primary sites are shown in the Additional File 7: Table S4.
Comparison of methylation frequencies for five neuropeptide receptor genes and TET family genes
Mean differences in the methylation index of the five GPCR neuropeptide receptors determined based on TET gene methylation events are illustrated in Fig. 2a. The MI was significantly higher in patients with full TET genes methylation events (3.55 ± 1.43), two TET genes methylation events (3.03 ± 1.26), and one TET gene methylation events (2.53 ± 1.38) than in patients with no TET gene methylation events (1.02 ± 1.26, P < 0.001 for all comparisons) (Fig. 2b).
Correlation between promoter methylation levels of the five neuropeptide receptor genes and TET family genes in cancer tissues. (a) Distribution of promoter methylation in TET family genes and the five neuropeptide receptor genes. Filled boxes indicate the presence of methylation, and open boxes indicate the absence of methylation. (b) Combined analysis of the MIs and methylation status of TET family genes. The number of methylation events is indicated for hypermethylated TET family genes. The mean MIs for the different groups were compared using the Student’s t-test. **P < 0.001. The data are shown as mean ± SD.
Kaplan-meier estimate
The Kaplan-Meier analysis of the DFS is shown in Fig. 3. DFS did not differ between in patients with methylated and unmethylated genes (Fig. 3a,b,d,f), with a few notable exceptions: DFS was significantly shorter when the GHSR (log-rank test, P = 0.009) and NMUR1 (log-rank test, P = 0.003) promoters were methylated (Fig. 3c,e). Among 135 cases with T1 and T2 tumor sizes, the DFS rates in those with GHSR and NMUR1 methylated genes were compared to the unmethylated group (log-rank test, P = 0.418 and P = 0.031, respectively) (Additional File 8: Fig. S4a,b). It was found that patients with T1 and T2 tumor sizes and methylated NMUR1 promoters had shorter DFS. Additional analysis that included only patients with oropharyngeal cancer (n = 75) revealed shorter DFS for methylated vs. unmethylated GHSR and NMUR1 (log-rank test, P = 0.004 and P = 0.008, respectively), but no differences for the other three genes (Additional File 8: Fig. S4c,d). Furthermore, in HPV-related oropharyngeal cancer (n = 37), GHSR and NMUR1 hypermethylation was significantly associated with shorter DFS (log-rank test, P = 0.003 and P = 0.026, respectively) (Additional File 8: Fig. S4e,f).
Kaplan-Meier survival curves for the 278 patients with HNSCC according to the methylation status of the five target genes. DFS for (a) NTSR1, (b) NTSR2, (c) GHSR, (d) MLNR, and (e) NMUR1 in the case of methylated (red lines) and unmethylated (blue lines) genes. (f) Combined analysis of the five genes. Blueline: patients with 0–3 methylated genes; red line: patients with 4–5 methylated genes. A probability of <0.05 (*P < 0.05) was considered a statistically significant difference.
Stratification analysis
The relation between methylation and risk of recurrence was analyzed through multivariate analysis using a Cox proportional hazards regression model adjusted for age, sex, smoking status, alcohol consumption, and clinical stage. In patients with GHSR promoter methylation, the adjusted odds ratio (OR) for recurrence was 1.656 [95% confidence interval (CI) 1.116–2.459, P = 0.012]. NMUR1 promoter methylation had a significant association with the OR for recurrence (OR = 1.670, 95% CI 1.133–2.458, P = 0.009) (Fig. 4). The OR for recurrence according to original tumor sites, was also determined. Methylation of the GHSR and NMUR1 promoters correlated positively with recurrence in oropharyngeal cancer patients, both individually (OR, 3.853; 95% CI, 1.510–9.832; P = 0.005 and OR, 2.872; 95% CI, 1.172–7.037; P = 0.036, respectively) and together (OR, 3.272; 95% CI, 1.216–8.801; P = 0.019) (Fig. 4).
In patients with T1 and T2 tumor sizes, NMUR1 promoter methylation has revealed a significant association with the OR for recurrence (OR = 2.14, 95% CI: 1.08–4.24, P = 0.028). For patients with T3 and T4 tumor sizes with a methylated GHSR promoter, the OR was 1.95 (95% CI: 1.17–3.24; P = 0.010) (Fig. 5a). Notably, the OR was significantly higher in HPV-positive oropharyngeal cancer patients (n = 37) in whom the GHSR promoter was methylated (OR, 19.00; 95% CI, 1.87–193.01; P = 0.013) (Fig. 5b).
Odds ratios for recurrence based on the Cox proportional hazards model. Multivariate Cox regression analyses were performed to assess the correlations between (a) recurrence and patients with T1–2 (n = 135) and T3–4 tumor sizes (n = 143) and between (b) recurrence and patients with HPV-positive (n = 37) and HPV-negative oropharyngeal cancer (n = 38).
Multivariate analysis including HPV status and methylation status in oropharyngeal cancer patients
For correlation analysis of the association between tumor HPV status, GHSR methylation status, and NMUR1 methylation status with survival, we combined data for all patients with oropharyngeal carcinoma. The DFS was correlated to better outcomes for patients with HPV-positive cancers than those with HPV-negative cancers (log-rank test, P = 0.017) (Fig. 6a). The study patients were classified into three categories with respect to the risk of recurrence: low-risk (Group 1 and 2), HPV-positive with both GHSR and NMUR1 unmethylated, or either GHSR or NMUR1 methylated; intermediate-risk (Group 4 and 5), HPV-negative with both GHSR and NMUR1 unmethylated, or either GHSR or NMUR1 methylated; and high-risk (Group 3 and 6), any HPV status with both GHSR and NMUR1 methylated. HPV-positive cancer patients were regarded to be at low-risk, with the exception of patients with both GHSR and NMUR1 methylated (Fig. 6a). As shown in Fig. 6b, DFS rates in the patients were 87.1% (95% CI, 73.4–100%), 42.7% (95% CI, 5.4–80.1%), and 17.0% (95% CI, 0–43.4%), for the low-risk, intermediate-risk, and high-risk groups, respectively. DFS was statistically significant different across risk groups (P < 0.001 for low- vs. high-risk and P = 0.019 for low- vs. intermediate-risk) (Fig. 6b).
Classification of study patients with oropharyngeal carcinoma into the risk of recurrence categories and Kaplan-Meier estimates of DFS according to the categories. (a) Kaplan-Meier estimates of DFS among oropharyngeal cancer patients to classify patients into categories of low-, intermediate-, or high-risk of recurrence, according to GHSR methylation, NMUR1 methylation, and HPV status. (b) Patients with oropharyngeal carcinoma were classified into three categories with respect to the risk of recurrence. Group 1 and 2: low-risk group; Group 4 and 5: intermediate-risk group; Group 3 and 6: high-risk group.
External validation of our results using methylation data from the TCGA database
The methylation status of GPCR neuropeptide receptor gene promoters was estimated in an additional 516 HNSCC samples and 50 normal samples from the database. The average β values of promoter methylation for the five genes were significantly higher in the HNSCC samples than in the normal samples (P < 0.001) (Additional File 9: Fig. S5a). The validation of TCGA data was used to assess the methylation status of the five genes in tumors from the hypopharynx, larynx, oropharynx, or oral cavity (Additional File 9: Fig. S5b; Additional File 9: Fig. S5c). The mRNA expression status of the five neuropeptide receptor genes in HNSCC and normal samples were obtained from the TCGA database (Additional File 10: Fig. S6).
Discussion
GPCRs belong to a superfamily of cell surface signaling proteins that have an important role in many physiological functions and multiple diseases, including the development of malignant neoplastic disease21. We found that aberrant methylation of the GHSR and NMUR1 promoters correlates with survival and recurrence in patients with HNSCC. In addition, the site-specific analysis revealed that abnormal CpG island hypermethylation in the GHSR and NMUR1 promoters was independently associated with aggressive clinical behavior in oropharyngeal cancer. It is worth noting that the GHSR and NMUR1 methylation status is a strong predictor of poorer survival among patients with HPV-positive oropharyngeal cancer.
The GPCR family members include the neurotensin receptors, of which there are two subtypes, NTSR1 and NTSR2, which are associated with carcinogenesis, cancer progression, and prognosis22,23,24. Neurotensin is a 13-amino acid neuropeptide that is localized principally in the central nervous system25. The actions of neurotensins are mediated through a high-affinity receptor (NTSR1) and a low-affinity receptor (NTSR2)26. In neuroendocrine tumors, a lack of NTSR1 promoter methylation with overexpression and dense of NTSR2 promoter methylation are observed24. On the contrary, NTSR1 methylation is related to lateral and noninvasive tumor growth of colorectal tumors27.
GHSR is also known as the ghrelin receptor, and the hormone ghrelin is its endogenous ligand. Strikingly, our study identified a single locus within the promoter region of the GHSR gene that is hypermethylated in 54.3% (151 of 278) of HNSCC, independently of patient age or tumor stage. GHSR promoter methylation was the most accurately detected (with AUROC of 0.81 obtained from the ROC) in tumor samples and matched paired normal mucosal samples. Loss of GHSR expression correlates with hypermethylation of GHSR in breast, cervical, prostate, pancreatic, colorectal, and pharyngeal cancers, and glioblastoma28,29,30,31. GHSR methylation in cervical tissue and scrapes is associated with 3q gain for the detection of HPV-induced cervical precancer29. These findings indicate that GHSR methylation may represent a potential pancancer marker for the detection of multiple tumor types including HNSCC (Additional File 11: Table S5).
Motilin is a gastrointestinal hormone released from the duodenum32. The MLNR shares significant amino acid sequence identity with GHSR33. Recently, it was reported that variants in the MLNR gene (rs9568169) are specific genetic risk factors for bile duct cancer34. Neuromedin U and its structurally related peptide, neuromedin S, are reported to regulate multiple physiological processes, and their functions are mediated by two receptors, NMUR1 and NMUR2. In a DNA methylation analysis to elucidate the potential molecular mechanisms underlying osteosarcoma, neuromedin U and NMUR1 methylation exhibited the highest degrees selected from the protein-protein interaction network35. The NMUR2 promoter region is C+ G-rich; however, the level of condensation of CpG sequences in this region is too low to design primers for a methylation assay36.
CpG hypermethylation is a major epigenetic DNA modification that tumor suppresses gene expression in cancer tumorigenesis and progression37. Analysis of genomic structure showed a CpG island in a region containing translational start sites that extends to the first exon of NTSR1, NTSR2, GHSR, MLNR, and NMUR1. Proper DNA methylation depends on the underlying mechanisms that regulate the writing, reading, and erasing of methylation marks38. Interestingly, the activity of TET enzymes is involved in removing epigenetic methylation marks39. TET proteins prevent unwanted DNA methyltransferase activity by binding to CpG-rich regions40. Recently, we reported that TET inactivation through promoter methylation occurs in HNSCC and promotes the inactivation of tumor suppressor genes16.
HPV-related oropharyngeal carcinomas belong to an independent tumor type with regard to cellular, biological, and clinical features41. HPV status, smoking status, tumor stage, and lymph node status are important factors that can be used to classify patients with low-, intermediate-, and high-risk of death11,42,43,44. However, the optimal classification for patients with low-, intermediate-, and high-risk disease remains to be determined. Our findings suggest that it is important to integrate HPV status and methylation status as determinants of recurrence risk for patients with oropharyngeal carcinoma.
Conclusion
We have shown that high-throughput methylation profiles can be correlated with recurrence and survival in HNSCC. Our results indicate that the methylation status of the GHSR and NMUR1 genes is an independent prognostic indicator for patients with oropharyngeal cancers. Furthermore, the function of TET genes as a methylation eraser should be considered in future studies of HNSCC carcinogenesis and its potential biomarkers and therapeutic targets. Our findings support the use of methylation markers in patient selection for adjuvant therapy following primary treatment with surgery and oropharyngeal cancer surveillance programs. Since our study is preliminary, it needs to be validated in larger cohorts of patients with more homogeneous oropharyngeal cancer patients.
References
Bray, F. et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: a cancer journal for clinicians 68, 394–424, https://doi.org/10.3322/caac.21492 (2018).
Chaturvedi, A. K. et al. Worldwide trends in incidence rates for oral cavity and oropharyngeal cancers. Journal of clinical oncology: official journal of the American Society of Clinical Oncology 31, 4550–4559, https://doi.org/10.1200/JCO.2013.50.3870 (2013).
Hashibe, M. et al. Interaction between tobacco and alcohol use and the risk of head and neck cancer: pooled analysis in the International Head and Neck Cancer Epidemiology Consortium. Cancer epidemiology, biomarkers & prevention: a publication of the American Association for Cancer Research, cosponsored by the American Society of Preventive. Oncology 18, 541–550, https://doi.org/10.1158/1055-9965.EPI-08-0347 (2009).
Leemans, C. R., Braakhuis, B. J. & Brakenhoff, R. H. The molecular biology of head and neck cancer. Nature reviews. Cancer 11, 9–22, https://doi.org/10.1038/nrc2982 (2011).
Kimple, R. J. et al. Development and characterization of HPV-positive and HPV-negative head and neck squamous cell carcinoma tumorgrafts. Clinical cancer research: an official journal of the American Association for. Cancer Research 19, 855–864, https://doi.org/10.1158/1078-0432.CCR-12-2746 (2013).
Dorsam, R. T. & Gutkind, J. S. G-protein-coupled receptors and cancer. Nature reviews. Cancer 7, 79–94, https://doi.org/10.1038/nrc2069 (2007).
Saikia, S., Bordoloi, M. & Sarmah, R. Established and In-trial GPCR Families in Clinical Trials: A Review for Target Selection. Current drug targets 20, 522–539, https://doi.org/10.2174/1389450120666181105152439 (2019).
Arakaki, A. K. S., Pan, W. A. & Trejo, J. GPCRs in Cancer: Protease-Activated Receptors, Endocytic Adaptors and Signaling. International journal of molecular sciences 19, https://doi.org/10.3390/ijms19071886 (2018).
Leemans, C. R., Snijders, P. J. F. & Brakenhoff, R. H. The molecular landscape of head and neck cancer. Nature reviews. Cancer 18, 269–282, https://doi.org/10.1038/nrc.2018.11 (2018).
Kanazawa, T., Misawa, K. & Carey, T. E. Galanin receptor subtypes 1 and 2 as therapeutic targets in head and neck squamous cell carcinoma. Expert opinion on therapeutic targets 14, 289–302, https://doi.org/10.1517/14728221003598922 (2010).
Ang, K. K. et al. Human papillomavirus and survival of patients with oropharyngeal cancer. The New England journal of medicine 363, 24–35, https://doi.org/10.1056/NEJMoa0912217 (2010).
Mehanna, H. et al. PET-CT Surveillance versus Neck Dissection in Advanced Head and Neck Cancer. The New England journal of medicine 374, 1444–1454, https://doi.org/10.1056/NEJMoa1514493 (2016).
You, J. S. & Jones, P. A. Cancer genetics and epigenetics: two sides of the same coin? Cancer cell 22, 9–20, https://doi.org/10.1016/j.ccr.2012.06.008 (2012).
Brown, R., Curry, E., Magnani, L., Wilhelm-Benartzi, C. S. & Borley, J. Poised epigenetic states and acquired drug resistance in cancer. Nature reviews. Cancer 14, 747–753, https://doi.org/10.1038/nrc3819 (2014).
Easwaran, H. P. et al. Aberrant silencing of cancer-related genes by CpG hypermethylation occurs independently of their spatial organization in the nucleus. Cancer research 70, 8015–8024, https://doi.org/10.1158/0008-5472.CAN-10-0765 (2010).
Misawa, K. et al. The neuropeptide genes SST, TAC1, HCRT, NPY, and GAL are powerful epigenetic biomarkers in head and neck cancer: a site-specific analysis. Clinical epigenetics 10, 52, https://doi.org/10.1186/s13148-018-0485-0 (2018).
Li, L. et al. Epigenetic inactivation of the CpG demethylase TET1 as a DNA methylation feedback loop in human cancers. Scientific reports 6, 26591, https://doi.org/10.1038/srep26591 (2016).
Brenet, F. et al. DNA methylation of the first exon is tightly linked to transcriptional silencing. PloS one 6, e14524, https://doi.org/10.1371/journal.pone.0014524 (2011).
Huang, W. Y. et al. MethHC: a database of DNA methylation and gene expression in human cancer. Nucleic acids research 43, D856–861, https://doi.org/10.1093/nar/gku1151 (2015).
Misawa, K. et al. Genes Located on 18q23 Are Epigenetic Markers and Have Prognostic Significance for Patients with Head and Neck Cancer. Cancers 11, https://doi.org/10.3390/cancers11030401 (2019).
Lappano, R. & Maggiolini, M. G protein-coupled receptors: novel targets for drug discovery in cancer. Nature reviews. Drug discovery 10, 47–60, https://doi.org/10.1038/nrd3320 (2011).
Guo, S. et al. Identification and validation of the methylation biomarkers of non-small cell lung cancer (NSCLC). Clinical epigenetics 7, 3, https://doi.org/10.1186/s13148-014-0035-3 (2015).
Kim, J. T. et al. Differential expression and tumorigenic function of neurotensin receptor 1 in neuroendocrine tumor cells. Oncotarget 6, 26960–26970, https://doi.org/10.18632/oncotarget.4745 (2015).
Kim, J. T., Weiss, H. L. & Evers, B. M. Diverse expression patterns and tumorigenic role of neurotensin signaling components in colorectal cancer cells. International journal of oncology 50, 2200–2206, https://doi.org/10.3892/ijo.2017.3990 (2017).
Levitas-Djerbi, T., Yelin-Bekerman, L., Lerer-Goldshtein, T. & Appelbaum, L. Hypothalamic leptin-neurotensin-hypocretin neuronal networks in zebrafish. The Journal of comparative neurology 523, 831–848, https://doi.org/10.1002/cne.23716 (2015).
Wu, Z., Martinez-Fong, D., Tredaniel, J. & Forgez, P. Neurotensin and its high affinity receptor 1 as a potential pharmacological target in cancer therapy. Frontiers in endocrinology 3, 184, https://doi.org/10.3389/fendo.2012.00184 (2012).
Kamimae, S. et al. Epigenetic silencing of NTSR1 is associated with lateral and noninvasive growth of colorectal tumors. Oncotarget 6, 29975–29990, https://doi.org/10.18632/oncotarget.5034 (2015).
Botla, S. K. et al. Diagnostic values of GHSR DNA methylation pattern in breast cancer. Breast cancer research and treatment 135, 705–713, https://doi.org/10.1007/s10549-012-2197-z (2012).
Verlaat, W. et al. Genome-wide DNA Methylation Profiling Reveals Methylation Markers Associated with 3q Gain for Detection of Cervical Precancer and Cancer. Clinical cancer research: an official journal of the American Association for. Cancer Research 23, 3813–3822, https://doi.org/10.1158/1078-0432.CCR-16-2641 (2017).
Moskalev, E. A. et al. GHSR DNA hypermethylation is a common epigenetic alteration of high diagnostic value in a broad spectrum of cancers. Oncotarget 6, 4418–4427, https://doi.org/10.18632/oncotarget.2759 (2015).
Nakagawa, T. et al. Frequent promoter hypermethylation associated with human papillomavirus infection in pharyngeal cancer. Cancer letters 407, 21–31, https://doi.org/10.1016/j.canlet.2017.08.008 (2017).
Brown, J. C., Cook, M. A. & Dryburgh, J. R. Motilin, a gastric motor activity stimulating polypeptide: the complete amino acid sequence. Canadian journal of biochemistry 51, 533–537 (1973).
Depoortere, I. Motilin and motilin receptors: characterization and functional significance. Verhandelingen - Koninklijke Academie voor Geneeskunde van Belgie 63, 511–529 (2001).
Xu, H. L. et al. Variants in motilin, somatostatin and their receptor genes and risk of biliary tract cancers and stones in Shanghai, China. Meta gene 2, 418–426, https://doi.org/10.1016/j.mgene.2014.04.012 (2014).
Chen, X. G., Ma, L. & Xu, J. X. Abnormal DNA methylation may contribute to the progression of osteosarcoma. Molecular medicine reports 17, 193–199, https://doi.org/10.3892/mmr.2017.7869 (2018).
Liu, J. J. et al. Discovery and pharmacological characterization of a small-molecule antagonist at neuromedin U receptor NMUR2. The Journal of pharmacology and experimental therapeutics 330, 268–275, https://doi.org/10.1124/jpet.109.152967 (2009).
Hug, M. et al. Transcriptional repression by methylation: cooperativity between a CpG cluster in the promoter and remote CpG-rich regions. FEBS letters 379, 251–254 (1996).
Parker, M. J., Weigele, P. R. & Saleh, L. Insights into the Biochemistry, Evolution, and Biotechnological Applications of the Ten-Eleven Translocation (TET) Enzymes. Biochemistry 58, 450–467, https://doi.org/10.1021/acs.biochem.8b01185 (2019).
Rasmussen, K. D. & Helin, K. Role of TET enzymes in DNA methylation, development, and cancer. Genes & development 30, 733–750, https://doi.org/10.1101/gad.276568.115 (2016).
Xu, Y. et al. Genome-wide regulation of 5hmC, 5mC, and gene expression by Tet1 hydroxylase in mouse embryonic stem cells. Molecular cell 42, 451–464, https://doi.org/10.1016/j.molcel.2011.04.005 (2011).
Harbison, R. A. et al. The mutational landscape of recurrent versus nonrecurrent human papillomavirus-related oropharyngeal cancer. JCI insight 3, https://doi.org/10.1172/jci.insight.99327 (2018).
Tinhofer, I. et al. Targeted next-generation sequencing identifies molecular subgroups in squamous cell carcinoma of the head and neck with distinct outcome after concurrent chemoradiation. Annals of oncology: official journal of the European Society for Medical Oncology 27, 2262–2268, https://doi.org/10.1093/annonc/mdw426 (2016).
Fakhry, C. et al. Validation of NRG oncology/RTOG-0129 risk groups for HPV-positive and HPV-negative oropharyngeal squamous cell cancer: Implications for risk-based therapeutic intensity trials. Cancer, https://doi.org/10.1002/cncr.32025 (2019).
Wagner, S. et al. Human papillomavirus association is the most important predictor for surgically treated patients with oropharyngeal cancer. British journal of cancer 116, 1604–1611, https://doi.org/10.1038/bjc.2017.132 (2017).
Acknowledgements
We would also like to thank the research, laboratory, and clinical staff who supported this study. This study was funded by a Grant-in-Aid for Scientific Research (No. 16K11228, No. 16K20239, 17K16904, 17K16903, 19K09866, 19K09906, and 19K18728) from the Ministry of Education, Culture, Sports, Science, and Technology of Japan.
Author information
Authors and Affiliations
Contributions
K.M., M.M., Y.S. and Y.M. conceived the study and designed the experiments. A.I., D.M., T.N., T.K., M.O., R.I., Y.Y. S.E., H.K., T.K. and H.M. analyzed the data and prepared figures and tables. All authors wrote the manuscript, reviewed its drafts, approved its final version and agreed with its submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Consent for publication Consent for publication was obtained from all patients.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Misawa, K., Mima, M., Satoshi, Y. et al. Neuropeptide receptor genes GHSR and NMUR1 are candidate epigenetic biomarkers and predictors for surgically treated patients with oropharyngeal cancer. Sci Rep 10, 1007 (2020). https://doi.org/10.1038/s41598-020-57920-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-57920-z
This article is cited by
-
Embryo-scale epithelial buckling forms a propagating furrow that initiates gastrulation
Nature Communications (2022)
-
Rare deleterious germline variants and risk of lung cancer
npj Precision Oncology (2021)
-
Identification of novel methylation markers in HPV-associated oropharyngeal cancer: genome-wide discovery, tissue verification and validation testing in ctDNA
Oncogene (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.