Abstract
Interactions between plants and the soil’s microbial & fungal flora are crucial for the health of soil ecosystems and food production. Microbe-plant interactions are difficult to investigate in situ due to their intertwined relationship involving morphology and metabolism. Here, we describe an approach to overcome this challenge by elucidating morphology and the metabolic profile of Medicago truncatula root nodules using Magnetic Resonance (MR) Microscopy, at the highest magnetic field strength (22.3 T) currently available for imaging. A home-built solenoid RF coil with an inner diameter of 1.5 mm was used to study individual root nodules. A 3D imaging sequence with an isotropic resolution of (7 μm)3 was able to resolve individual cells, and distinguish between cells infected with rhizobia and uninfected cells. Furthermore, we studied the metabolic profile of cells in different sections of the root nodule using localised MR spectroscopy and showed that several metabolites, including betaine, asparagine/aspartate and choline, have different concentrations across nodule zones. The metabolite spatial distribution was visualised using chemical shift imaging. Finally, we describe the technical challenges and outlook towards future in vivo MR microscopy of nodules and the plant root system.
Similar content being viewed by others
Introduction
Interactions between plants and microbes are considered to be important for the health of the soil ecosystem as a whole1. Understanding the metabolic interactions between plants and microbes, both commensal and parasitic, could help address many of the challenges we face today, related to agriculture and food security. One such interaction is the microbiome-mediated uptake of nitrogen by plants. Availability of biologically-active forms of nitrogen in the soil is an important factor determining crop yield. Current agricultural practice, therefore, relies strongly on nitrogen fertiliser to supplement soil nitrogen to ensure high crop yield2. The Haber-Bosch nitrogen fixation process, used to produce the ammonia needed for these fertilisers, currently consumes 1% of the world energy sources; making it the most energy consuming process in the chemical industry3. In contrast, alternative processes (e.g. symbiotic nitrogen fixation, SNF) achieve the same result of fixing nitrogen without the need for high pressure and temperature required by the Haber-Bosch process.
More precisely, plants have solved the problem of biological nitrogen fixation through commensal processes, involving bacterial infection of plant roots4. Mutualistic infections are omnipresent in nature, with a wide range of nitrogen-fixing bacteria – such as those that are collectively named rhizobium – invading not just plants but also the phycosphere of green algae5. The mutualistic symbiosis involves almost all parts of the plant cell machinery, including plastids and mitochondria. Of particular interest are the interactions between rhizobial bacteria and leguminous plants, which form dedicated organs - root nodules - to accommodate the bacteria. Medicago. truncatula infected with Sinorhizobium meliloti (S. meliloti) has long since been used as a model plant system to study plant-rhizobia interactions6. The root nodules facilitate intracellular hosting and nutrient exchange7. The result is the efficient formation of fixed nitrogen (NH4+), which provides considerable growth advantages to leguminous plants8. Recently, a detailed fate map of root nodule formation at a genetic level has been developed9. It has also been proposed that rhizobial metabolic activity may confer resistance to drought and salt stress10,11,12. More detailed understanding of key metabolites in plant-nodule metabolism and importantly, their localisation within root nodule tissues could, therefore, shed light on the mechanism by which Symbiotic Nitrogen Fixation (SNF) confers advantages to host plants.
Tracking metabolite exchange non-invasively within intact nodule systems is difficult with most imaging modalities, since they require a form of sample fixation, for example in the case of Matrix Assisted Laser Desorption/Ionisation Mass Spectrometry (MALDI-MS) imaging13. The current knowledge of the metabolic profile in the root nodules is based on analysis with destructive extraction procedures. For example, Gas Chromatography and Liquid Chromatography in combination with Mass Spectrometry (GC/LC-MS) has provided rich metabolic information but required destructive extraction procedures14. These in vitro methods may not faithfully reflect the native structural and molecular information. Examining and mapping the levels of metabolites directly in the intact root nodule would be important to understand the functional framework of metabolism in the native state. Magnetic resonance techniques may offer advantages in terms of spatially resolved spectroscopic information on a single nodule, which is difficult to access with alternative techniques. Magnetic resonance imaging (MRI) has been previously used to visualize belowground root structures15,16,17.
Advances in magnetic resonance technology have resulted in steady increase in sensitivity, making Magnetic Resonance Microscopy (MRM) a reality where resolution below 100 μM can be achieved18. A combination of advances in gradients strengths, magnetic field strengths and RF coil designs, have made it possible to attain ever higher resolutions19. Specifically, microcoils have played an important role in increasing sensitivity, by optimally matching the RF coil to a sample of interest20,21,22. The highest possible resolution on phantom has steadily increased over the years, and thus far maximum nominal resolutions of around 3 μm has been attained on phantoms23,24,25,26. Biological applications have been diverse, including imaging of neuronal tissue, where cellular resolution were obtained on individual neurons27,28,29,30,31. Moreover, highly resolved Nuclear Magnetic Resonance spectroscopy has also been demonstrated using the same type of hardware32.
In this work, we applied state-of-the-art magnetic resonance microscopy (MRM) in conjunction with localised spectroscopy at ultra-high magnetic field (22.3 T), using a home-built solenoid RF coil in order to image root nodules of Medicago truncatula. We obtained not only cellular level structural information but also mapped some of the metabolic information in localised zones and how they vary across different tissues. Lastly, we describe some of the technical challenges that were encountered and an outlook towards future in vivo imaging of nodules and plant root systems.
Results
Root nodule morphology resolved by MR microscopy in cellular detail
A representative M. truncatula specimen with a single root nodule shown in Fig. 1a, was imaged using a custom home-built 1.5 mm diameter solenoid coil as depicted in Fig. 1b.
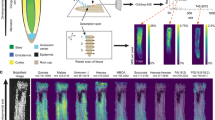
Photograph of M. truncatula root system and custom-designed home-built microcoil. (a) Photograph of a representative M. truncatula plant (five weeks old, three weeks post inoculation). Typical root nodule indicated with a red square. Some white perlite beads are still visible along the root system. (b) Photograph of a home-built solenoid coil insert mounted on a Bruker insert holder. The solenoid inner diameter is 1.5 mm; further details are described in the materials and methods.
Individual cells of a root nodule with two lobes and meristems (m) could be distinguished on the MRI scans. A high-resolution 3D Fast Low Angle SHot (FLASH) image of the root nodule, fixed with formaldehyde and submerged in perfluorodecalin (PFD), is shown in Fig. 2a. All image slices through the entire root nodule are shown in the Supplementary Material (Supplementary Video 1). Root nodule morphology in M. truncatula nodules are of the indeterminate type, i.e., four distinct regions (zones) occur, from the apex of the root nodule to the root attachment point4. Nodule tissue - from outward to inward - starting with the nodule cortex (nc), delineated as a high-intensity ring on the surface of the nodule, contrasted with the darker region of the meristem (m). Cells in the meristem, responsible for the growth of the nodule, were significantly smaller than mature bacteroid containing cells. In the next region, the rhizobial infection zone (if), young cells infected by rhizobia bacteria could be seen as a region of alternating high- and low-intensity patches. Not all cells in the rhizobial infection zone were infected, which was apparent due to the presence of variation in image intensities, i.e., non-infected plant cells had a higher image intensity as compared to cells infected with the bacterium. Lastly, in the active nitrogen fixation zone (fx), large cells (40 to 50 µm) could be distinguished by their grey rings, caused by the presence of bacteroids surrounding the vacuole33. The fixation zone is where rhizobia make nitrogenase, the enzyme required for nitrogen fixation. In the same area, uninfected interstitial cells were visible, which did not exhibit the rings. These interstitial cells perform an important role in the regulation of nitrogen fixation activity34. Lastly, connective tissues (vascular bundles) could be seen extending from the root towards the periphery of the nodules, providing a route for nutrient exchange with the rest of the plant.
High-resolution MR imaging and optical microscopy of root nodules. (a) FLASH image of fixed root nodule at 7 × 7 × 7 µm3 resolution with an acquisition time of 34 h24 m. Individual cells containing bacteroids (ba) can be discerned by the dark rings inside the cells, where S. meliloti cells accumulate. The nodule exhibits a double meristem (m) on opposed directions with cells becoming smaller towards the meristem until individual cells can no longer be resolved. Air pockets appear as hypo-intense regions marked with an arrowhead. (b) Optical transmission microscopy of nodule section (thickness 5 µm) stained with Toluidine blue. (c) FLASH image of in situ root nodule at 7 × 7 × 7 µm3 resolution with an acquisition time of 34 h24 m. Though lower in signal-to-noise, both uninfected cells (black arrow) and infected cells (ba) can be discerned. Abbreviations: rc, root cortex; xy, xylem; ph, phloem; nc, nodule cortex; vb, vascular bundle; fx, fixation zone; if, infection zone; ba, bacteroid containing cells; m, meristem. Scale bars 200 µm.
Figure 2b shows a light microscopy image of a histological slice stained with Toluidine blue, from the same root nodule at approximately the same plane as the MR image (co-registration). Toluidine blue stained the nuclei and lignin of the cell wall35. Common features in Fig. 2a,b are seen, including the bacteroid ring as observed on the MR image, occurring in the staining of cells in the nitrogen fixation zone. Especially visible were the vascular bundles extending from the central root, which due to their smaller cells appeared more densely stained in the optical microscopy image. In addition, air pockets could also be imaged as they appeared as hypo-intense regions in the FLASH image (Fig. 2a, arrowhead). Air pockets also introduced differences in magnetic susceptibility within the sample, which influenced image quality. Air pockets of different sizes could be seen within the fixation zone using multiple gradient echo (MGE) imaging with echoes acquired at increasing echo times (Supplementary Fig. 1). Because of the abundance of air pockets in the apical zone, the susceptibility-induced hypointense regions also increased in size in the apical direction.
To evaluate the possibilities of high-resolution MR imaging of root nodule without any fixation or treatment, measurements were performed on freshly excised nodules (Fig. 2c). While the image contrast and signal-to-noise ratio (SNR) was noticeably reduced, individual cells in the nitrogen fixation zone could still be distinguished. The uninfected cells appeared bright while infected cells appeared dark in the active infection zone. Cells in the meristem appear slightly brighter in freshly excised nodule (in situ) than in the fixed nodule, possibly due to the native contrast in freshly excised nodule, while in fixed nodule the contrast was possibly affected by formaldehyde fixation36,37.
Localised spectroscopy revealed the spatial distribution of nodule metabolites
Since the root nodule is an active plant organ whose primary function is to fix nitrogen and to facilitate nutrient exchange, SNF-linked metabolite distribution over the various nodule zones is of interest. According to earlier literature, a nodule may be divided into four zones, based on SNF activity: (I) meristem zone, (II) infection zone, (III) nitrogen fixation zone, and (IV) senescence zone, the latter observed mostly in older nodules. In the infection zone, bacteroids accumulate and mature, while in the fixation zone active SNF takes place38. Localised Magnetic Resonance Spectroscopy (MRS) was utilised to distinguish metabolites in a freshly excised and PFD submerged root nodule, based on their frequency differences (chemical shift) and location. Localised MR spectra were recorded from five different regions of interest (ROI), localised in the four different zones in an intact freshly excised root nodule, in situ (Fig. 3a). The presence of bacteroids was verified with optical microscopy after the MRI measurements (Fig. 3c). The first ROI was placed on the meristem (ROI 1) in Fig. 3a, where little MR signal was recorded, possibly due to the small volumes of individual cells which may cause T2-shortening, reducing the SNR. The second voxel (ROI 2) was placed in the infection/early fixation zone. The third (ROI 3) and fourth voxels (ROI 4) were placed in the largest zone: the active nitrogen fixation zone. The last voxel (ROI 5) was placed in the basal region of the root nodule.
Localised spectroscopy in in-situ root nodule reveals sugar differences in pre- and post-granule onset regions. (a) FLASH reference with PRESS Region of Interest (ROI), numbered one through five. Roman numerals indicate nodule zones: Meristem (I), Infection zone (II), Nitrogen fixation zone (III). Grey dotted boxes show ROI shift of Betaine due to Chemical Shift Displacement Error (CDSE). Pulse excitation was centred around 4.0 ppm. (b) PRESS spectra captured from Regions of Interest (ROI) shown in a, for Nodule Meristem (1), (early) Nitrogen fixation zone (2), Nitrogen fixation zone (3 & 4), Basal region (5). PRESS voxel sizes were 200 × 350 × 350 µm3. Spectra were aligned to the betaine peak at 3.3 ppm. Water peak region at 4.7 ppm has been omitted for ease of viewing. Intensities of the spectra have not been normalised, reflecting the strength of the signal recorded. This means that for the meristem little information can be discerned. Line broadening 10 Hz. (c) Co-registration of Optical Microscopy confirms the presence of bacteroids in the active nitrogen fixation region. Toluidine blue staining 10x magnification. Abbreviations: nc, nodule cortex; vb, vascular bundle; m, meristem; ra, root attachment area. Scale bars 200 µm.
The spectra obtained from ROIs 2–5 showed the presence of several metabolites including various amino acids, sugars, choline and betaine. Betaine was present in a high amount in all four ROIs (Fig. 3b). The signal from sucrose and asparagine/aspartate was also clearly detected. Assignment of various metabolites was confirmed with solution-state NMR performed on an extract obtained from the root nodules, using two-dimensional 1H-1H correlation spectroscopy (2D-COSY) (see Supplementary Fig. 2). It should be noted that, due to the chemical shift displacement error, which increases with magnetic field strength, the regions of interest chosen for the localised spectra were shifted slightly for each metabolite, as the spatial selection relied on the frequency of the metabolite relative to the chosen centre frequency of 4.0 ppm39. This shift meant that the indicated regions of interest in Fig. 3a were correct for signals at δ = 4 ppm, while for example the strongest signal (at δ = 3.3 ppm) originated from an ROI which was shifted by 35 µm along x, 97 µm along y and 97 µm along z (see the dotted region of interest in Fig. 3a).
The strongest peak observed in all spectra was attributed to a resonance of glycine betaine (δ = 3.3 ppm) (Fig. 3b). A complementary spectroscopic imaging method, Chemical shift imaging (CSI) confirmed the presence of betaine throughout the nodule (Supplementary Fig. 3). Betaine serves as an important osmoprotectant produced by S. meliloti40. Furthermore choline – a precursor to glycine betaine in both plants and bacteria, was visible in ROI 3 as a shoulder on the low frequency side of the betaine peak, and was even more pronounced in ROIs 4 and 541. In uninfected root tissue, choline, but not glycine betaine was detected as seen in solution NMR measurements (Supplementary Fig. 4).
In root nodule, another strong peak from asparagine and aspartate was seen primarily in ROIs 2–5 (Fig. 3b). Notably, glutamine - though present in ROI 4 - appeared much lower in concentration in all ROIs than asparagine, suggesting a smaller pool or higher turnover rate in the nitrogen assimilation process.
Intermediates from the tricarboxylic acid cycle such as acetate (δ = 1.9 ppm), pyruvate (δ = 2.3 ppm) and succinic acid (δ = 2.4 ppm) were also seen in the nitrogen fixation zone (region 3 & 4). Lactate (δ = 1.3 ppm) was visible only in the active nitrogen fixation zone (region 3 & 4). Alanine (δ = 1.5 ppm) has been suggested by metabolic network modelling to be the primary metabolite for the uptake of assimilated nitrogen by plants42. However, our results indicated that the pool of alanine was small compared to asparagine.
Sucrose, the most important resource for growth and root nodule activity, was present in all regions with a concentration gradient towards the basal area, starting with the early fixation zone (Fig. 3b, ROI 2). Several peaks were assigned to sucrose (e.g., δ = 5.3 ppm), with some overlap with choline (δ = 3.5 ppm) and fructose (δ = 3.8 ppm). In the basal region (region 5), the sucrose peak at δ = 5.3 ppm was hard to distinguish, possibly due to baseline distortions. Overall, the metabolic pattern exhibited in regions 3–4 appeared to be maintained in the basal region (region 5), though attenuated and with a larger linewidth, possibly reflecting a more disorganised structure of the tissue. At the same time active SNF still appeared to take place, indicating this region is not (yet) the senescent zone IV in which the activity would be reduced.
Optical microscopic detection of starch correlates with MR based sucrose profile
Starch is a carbohydrate storage polymer that is unique to plants. Indeterminate root nodules are known to form amyloplasts in certain cells, detectable by optical microscopy, especially in the youngest cells of the nitrogen-fixation zone of an actively growing nodule43. Accumulation of amyloplasts has also been found in cells located behind the division zone in shoot meristems44. Zeeman et al. speculated that this might be caused by ‘a temporary imbalance between carbon import and utilisation as cells move from division to expansion and differentiation’45. If correct, this marker for high metabolic activity should be reflected in changes to the metabolic profile of bacteroid containing cells. Therefore, we aimed to detect pre- and post-starch granule formation zones in an intact root nodule, and correlate them with sugar metabolites as measured by localised spectroscopy.
Lugol’s solution was used to stain amyloplasts (Fig. 4a) on an optical section immediately adjacent to that of Fig. 3c, to relate the distribution of sucrose and other metabolites to starch. Starch distribution was seen as dark spots, concentrated in a band located in the first layer of the nitrogen fixation zone as well as in smaller amyloplasts throughout the fixation zone (Fig. 4b,d). In the nitrogen fixation zone, amyloplasts were small and dispersed over infected cells (black arrowhead) as well as uninfected interstitial cells (white arrowhead)(Fig. 4d). In contrast, the cells present in the first layer on the border of the infection zone (II) and fixation zone (III) exhibited a different distribution of starch (Fig. 4b). At higher magnification, large amyloplasts were visible at the periphery of the infected cells, which were almost completely filled with bacteroids (Fig. 4c). Thus, sucrose and starch appear to co-localise with SNF activity. Taken together, starch distribution patterns could provide a possible biomarker for changes in cellular metabolism during nodule development.
Optical microscopy of root nodules reveals multiple starch distribution patterns. Coupes were stained with both Lugol’s solution and Toluidine blue. (a) A section immediately adjacent to that of Fig. 3c reveals starch distribution throughout the nodule. 10x magnification, scale bar 200 µm. A black dotted rectangle indicates the zone shown in (b) on an alternate section. A white rectangle indicates an enlargement seen in (d). 10x magnification, zoomed; scale bar 50 µm. (b) At the first layer of cells in zone III, starch is present as a band (s). A white rectangle indicates the area shown in (c). Alternate section, 20x Magnification, scale bar 100 µm. (c) Starch granules (s) are located near the periphery of infected cells. The vacuole (v) reappears in the fixation zone after disorganisation in the infection zone. (b). 40x Magnification, scale bar 20 µm. (d) A fine distribution of starch granules can be seen in Zone III contained in nitrogen fixating cells (black arrowhead) and uninfected cells (white arrowhead). Abbreviations: ra, root attachment site; nc, nodule cortex; vb, vascular bundles; m, meristem; s, starch granules; v, vacuole; ba, bacteroids.
Discussion
This paper showed the feasibility of attaining cellular resolution (7 × 7 × 7) µm3 in M. truncatula root nodules over a relatively large volume (1.8 × 1.4 × 1.4 mm3), opening up new applications for MR microscopy in plant studies (Fig. 2). Individual cells, especially young and mature bacteroids containing cells, were resolved with high contrast in fixed root nodule, exhibiting a unique ‘grey ring’ pattern where S. meliloti cells accumulated near the cell walls around a central vacuole. Smaller cells, such as those present in the meristem, could not be resolved individually. A low signal to noise in meristem may be attributed to the shortening of T1 and T2 relaxation times as a result of formaldehyde-based fixation. Formaldehyde is known to alter relaxation properties of tissues by cross-linking proteins, which may lead to a gel-like cytoplasm. This alters the rates of chemical exchange between water protons and proteins, thus leading to a reduced T1 and T236,37,46. However, vacuoles are less affected by this, potentially contributing to the strong contrast between bacteroids and vacuoles. Looking towards in situ imaging in Fig. 2c, although nodules suffered from lower image contrast and SNR, the meristem appears brighter as compared to the fixed nodule (Fig. 2a). These results also support the notion that lower signal in meristem in fixed nodule could be due to shortening of relaxation parameters. A possible reason for the overall lower image quality in freshly excised nodule may be due to the abundance of air spaces, since no vacuum treatment was applied for in situ imaging. More quantitative measurements are required to determine the causes of reduced SNR and contrast.
Although the hypo-intense regions caused by small and large air pockets can be detrimental to overall image quality (Supplementary Fig. 1), they also contain useful information. Air pockets in root nodules of M. truncatula are of particular interest, because of the delicate balance of oxygen levels required for the proper functioning of the nitrogen fixation. The pockets may thus have a regulatory role, as nitrogenase is paradoxically both dependent on oxygen for carrying out its function but can also be poisoned by oxygen under high concentrations47. Previous MR investigations into root nodules of soybean (Glycine max) have focused on investigating the oxygen diffusion barrier, believed to regulate nodule functioning48,49. Furthermore, Glycine max root nodules are known to be sensitive to changes in gas perfusion50. Thus, MRM could be a useful tool to study oxygen regulation mechanisms, due to its ability to detect air spaces through enhanced signal loss.
PRESS localised spectroscopy results show that betaine was distributed throughout the nodule (Fig. 3b). The relatively high concentration of betaine could be due to maintaining a high degree of osmoregulation, an indicator of stress, or both41. High asparagine levels seen especially in the nitrogen fixation zone indicate significant SNF activity. The presence of asparagine, as well as betaine, was also confirmed by high-resolution 2D correlation spectroscopy on extracts prepared from nodules (Supplementary Fig. 2). Asparagine, together with glutamine, is known to be a major export product of assimilated nitrogen51. Our results show that asparagine accumulates mostly in active SNF areas (regions 3–5). As such, asparagine was absent in normal root tissue (Supplementary Fig. 4).
Metabolite levels of betaine and asparagine might therefore potentially provide useful markers for stress and SNF activity, respectively. The sucrose concentration gradient, decreasing from the basal to the apical end of the nodule, may reflect either consumption rates or limitations in transport through the nodule. In conjunction, starch granules appeared to be localised to two specific regions, a band in the transition from infection to fixation zone, and the fixation zone (III) itself (Fig. 4). The band seemed to provide support for Zeeman’s prediction of amyloplast formation due to a ‘temporary metabolic imbalance’, though it is located at some distance from the meristem (Fig. 4b)45. As for the fixation zone, it has been observed in the literature that starch amyloplast accumulation in uninfected cells is related to the expression of SNF-related (nif) genes (Fig. 4d)52,53. Considering that sucrose was present throughout the nodule, albeit in a diminishing concentration gradient, it is likely that cell-specific metabolism and environmental conditions determine whether and in what quantity starch amyloplasts are formed38,54. For instance, sucrose must be metabolised by the host cell in order to provide carboxylic acids that are preferentially taken up by the bacteroids55,56. The observed distribution in size and location of amyloplasts could be affected in this way as well.
As an explorative study of ultra-high field MRM for plant imaging, several recommendations might be of interest to the reader. Sample handling and preparation are essential for high-resolution imaging. More generally, special attention needs to be given to managing sample susceptibility at ultra-high fields, since the inclusion of even small air bubbles in the vicinity of samples can distort image quality significantly, as well as cause line broadening in localised spectroscopic experiments. Perfluorodecalin (PFD) was useful as a submerging fluid as its magnetic susceptibility is close to that of tissues and it gives no observable 1H signal. It is also efficient in displacing and dissolving air pockets, which are present in root nodule tissue. The absence of extraneous signal allowed for a manual shimming strategy that was sufficient for imaging and provided a good starting point for shimming voxels of interest for localised spectroscopy. Still, maximum shim currents on a particular system may prove to be a limiting factor for micro imaging applications. Another source of sample susceptibility could be the presence of iron in the form of legheamogloblin which regulates oxygen levels. In bacteroids of mature nodules, up to 25% of the total soluble protein consists of legheamoglobin7. The high concentrations of this iron-heme containing protein might have implications for the values of longitudinal relaxation (T1) and transverse relaxation (T2) and their interpretation, as these properties may depend strongly on the physiological state of the root nodule when measured. Thus, with further characterisation, susceptibility-weighted imaging may provide a useful method for evaluating the physiological state of root nodules. Although MRS was done on freshly excised root nodules, the use of PFD as a submerging fluid may have some influence on the vitality of the nodule, since PFD displaces air pockets within the nodule thus, altering oxygen diffusion to the cells.
Rather than a higher resolution per se, the future applications for MRM at ultrahigh field for – in vivo – imaging are also enticing, yet challenging. To achieve in vivo imaging, novel coil-insert designs with the necessary life support systems would be required. The solenoid coil type used here could be modified to allow for a relatively simple setup that would support whole plants. Vertical-bore MRI systems allow top access to supply light and fresh medium. Additionally, controlled nutrient supply would be desirable, e.g. a continuous medium perfusion setup57. Overall, an imaging setup optimised for longitudinal studies would allow testing of stress conditions for host-symbionts, non-destructively.
In conclusion, our results demonstrate that MRM at ultra-high fields in conjunction with microcoils provides a promising technique to determine physiology and metabolic profiling non-invasively in plant root nodules. Further research and development on coil design are required to exploit this application to its full potential.
Methods
Growing conditions medicago truncatula
Medicago truncatula accession R108 was grown in perlite saturated with Färhaeus medium without nitrate in a growth chamber at a temperature of 21 °C and 16/8-h light/dark cycle58. Plants were inoculated with Sinorhizobium meliloti Rm41 (OD600 = 0.1, 2 mL per plant)59. Root nodules were collected for analysis 21 days after inoculation. Perlite was used as a soil replacement to facilitate the extraction of intact root systems.
Sample preparation
For MRI measurements, a single M. truncatula plant was carefully extracted from the perlite substrate. Healthy nodules that exhibited a light-pinkish colour were selected and used for MRI imaging. For in situ MRI and MRS measurements, a nodule was excised and was directly used for measurements without fixation. For ex vivo MRI and optical microscopy, root nodules were fixed with paraformaldehyde (4% w/v) and glutaraldehyde (5% v/v) in 50 mM phosphate buffer (pH 7.2) at 4 °C overnight, followed by vacuum treatment for 30 minutes. Samples were then stored at a temperature of 4 °C until use.
Using an SZ40 stereomicroscope (Olympus, Tokyo, Japan), individual nodules were selected that could fit within capillaries with an inner diameter of (1050 ± 50) µm (Hilgenberg GmbH, Malsfeld, Germany). To avoid artefacts during MRI acquisition from air bubbles both within and outside the sample, a Perfluorochemical solution (Perfluorodecalin, PFD) was used to submerge the sample. The use of a Perfluorochemical solution not only prevents sample dehydration but also minimizes susceptibility artefacts. PFD was chosen because it has several advantageous properties, making it suitable as an infiltration agent. Importantly, it is a non-toxic compound, capable of dissolving both CO2 and O260. Furthermore, PFD exhibits a low surface tension (1.9 × 10−2 N m−1), below the limit for stomatal penetration (2.5–3.5 × 10−2 N m−1) in Arabidopsis thaliana leaves60,61, making it useful for reducing air bubbles within the sample without entering the cells. Care was taken to minimise the required amount of PFD as it is known to function as a potent greenhouse gas62.
During insertion of the nodule in the centre position of the capillary, both nodule and capillary were kept submerged in PFD. Capillary wax (Hampton Research, California, USA) was then applied to both ends of the open capillary with a MaxWax Pen (Hampton Research, California, USA). Considerable effort was spent on minimising the inclusion of air bubbles within the capillary.
For solution NMR measurements, metabolites were extracted from root and root nodules as described in detail by Kim et al. (2010)63. Key steps included manual excision of root and root nodule tissue and consequent crushing with mortar and pestle in the presence of liquid nitrogen. Samples were then freeze-dried and lastly extracted with an extraction buffer consisting of 50% (v/v) deuterated methanol and 50% D2O phosphate buffer (pH 6)63.
MRI measurements
A custom solenoid microcoil was built in order to achieve the necessary sensitivity required for high-resolution imaging. For the microcoil assembly, a glass-fibre circuit board was used as a base to hold all the necessary components in place. The radiofrequency (RF) coil consisted of coated 28 AWG (Ø 0.4 mm) copper wire, wound into a solenoid with six windings around a glass capillary with an outer diameter of (1500 ± 50) µm. A fixed 1.5 picoFarad (pF) capacitor was placed in series with the RF coil. To complete the resonant circuit, a single 1.5–6 pF variable capacitor was placed parallel to the RF coil and 1.5 pF capacitor, which allowed for fine-tuning the resonant frequency to 950 MHz. The circuit board was then attached to a Micro5 probe (Bruker Biospin, Ettlingen, Germany) compatible circuit holder.
MRI measurements were performed on a Bruker 22.3 T (950 MHz) spectrometer (Bruker Biospin, Ettlingen, Germany), at the uNMR-nl national facility (Utrecht, The Netherlands). The magnet had a vertical bore of 54 mm in diameter and was connected to an Avance III HD console. A Bruker Micro5 probe with exchangeable RF inserts allowed operation of the 1500 µm custom-built solenoid resonator. The Micro5 probe contained a built-in 48 mT/m/A (2.88 T/m at 60 A) triple axial gradient system coupled to GREAT 60 A amplifiers and cooled with a BCU 20 water cooler. Control of spectrometer operations was performed with Paravision 6.0.1 and Topspin 3.1PV running on a CentOS workstation. Sample temperature was maintained at (298 ± 1) K using a BCU II cooler.
High-resolution 3D Fast Low Angle Shot (FLASH) images on ex vivo and in situ root nodules were acquired with the following parameters: matrix size 256 × 192 × 192; read direction along the largest matrix direction; the field of view was (1.8 × 1.4 × 1.4) mm3. Resolution (7 × 7 × 7) µm3 isotropic, number of averages 28, acquisition time 34 h 24 m, repetition time 120 ms, flip angle 30° and echo time 2.9 ms. Receiver bandwidth was set to 100 kHz. To allow for co-registration with coupes from optical microscopy, the JIVE tool was utilised to generate oblique slices.
Localised Spectroscopy
Point Resolved Spectroscopy (PRESS) was used for localised magnetic resonance spectroscopy (MRS) on root nodules64,65. PRESS employs 90°-180°-180° orthogonal pulses with concurrent slice selective gradients. The PRESS sequence was preceded by a VAPOR (Variable Pulse Power and Optimized Relaxation Delays) sequence for global water suppression, which consists of seven variable power RF pulses with an optimised relaxation delay66. Prior to MRS, a reference FLASH image was acquired with the following parameters: echo time 3.3 ms; repetition time 80 ms; matrix 192 × 128; field of view 1.8 × 1.2 mm and receiver bandwidth 50 kHz; in plane resolution 9 × 9 µm; slice thickness 52 µm; number of averages 256; and acquisition time 43 m. Appropriate volumes of interest (voxel) for PRESS were selected from the reference FLASH image. Correct positioning of the voxel was also verified after all spectroscopic measurements, by high-resolution 2D FLASH. Five individual MRS voxel (200 × 350 × 350 µm3) were localised covering various zones in the nodule. The local field homogeneity was optimised over the voxel by MAPSHIM. The field homogeneity was further improved by manual shimming of up to second order shims (Z-Z2-Z-X-Y-Z-Z2-Z-XY-XZ-YZ-Z). The time needed for shimming varied between 5–10 min. The field homogeneity resulted in a water line width of 10 Hz. For PRESS measurement, repetition time was set to 1000 ms, echo time was 7.2 ms, and 2048 averages were acquired for a total acquisition time of 34 m 8 s. Spectra were manually phased and automatically baseline corrected with a 3rd or 6th order Bernstein-polynomial fit. Chemical shift alignment was adjusted to the largest peak (3.3 ppm).
Chemical shift imaging
Chemical Shift Imaging (CSI) was used to acquire a multi-voxel spatial map of metabolic distribution. A large volume of interest was selected using a PRESS approach, consisting of three mutually orthogonal slice selective pulses. Then phase encoding occurred in two directions by applying incremental pulse gradients in each direction, resulting in a matrix of spectroscopic spin echoes.
The following basic parameters were used: Repetition time = 760 ms; echo time = 3 ms; Matrix = 24 × 16; FOV = 1.8 × 1.2 mm2; slice thickness = 400 µm; resolution = 150 × 150 × 400 µm3; number of averages = 1024. Spectra were acquired with 2048 points; dwell time 26.4 µs; spectral width 19.93 ppm (18,939 Hz). Magnetic field homogeneity in the selected volume was optimized by shimming the water resonance. A VAPOR suppression scheme was applied for efficient water signal saturation before CSI acquisition.
Reference FLASH images were acquired with the following parameters: matrix size was 96 × 64; Read direction along the largest matrix direction; field of view was 1.8 × 1.2 mm2. Slice thickness was 0.1 mm; resolution (19 × 19 × 100) µm3 isotropic. Number of slices 4; number of averages 128; acquisition time 28 m 40 s. Repetition time was 210 ms; flip angle 30° and echo time 3.26 ms. Receiver bandwidth was set to 50 kHz.
For the processing of the CSI data, the integration of selected signal of specific metabolite areas in magnitude mode was overlaid on reference FLASH images using the Bruker CSI Visualisation Tool.
NMR measurements
NMR measurements were performed on a Bruker 20.0 T (850 MHz) spectrometer with a vertical bore 54 mm in diameter connected to an Avance III HD console. A triple-tuned broadband cryoprobe was used. Topspin 3.2 was used to control the spectrometer and for processing of the acquired data. The sample temperature was maintained at 298 K.
1H NMR spectra were collected using a zgpr30 sequence with pre-saturation to suppress water efficiently. Data was acquired into 65k points; dwell time 29.3 µs, spectral width 20 ppm or 17,045 Hz; number of averages, 512; dummy scans 4; pre-scan delay 10 µs. 2D homonuclear 1H-1H experiments were performed using chemical shift correlated spectroscopic sequence (COSY). The following parameters were used: 3400 points in the direct dimension, 400 points in the indirect dimension in Digital Quadrature Mode. Both dimensions zero-filled to 4096 points. Number of averages, 96; dummy scans, 16; Spectral width 12.0 ppm, 10,000 Hz; Temperature 298 K.
Assignments were cross-checked against metabolite databases and fitting was performed using Chenomx deconvolution software (Chenomx, Edmonton, Canada)67,68. 1D reference spectra were acquired with parameters as follows: data was acquired into 65k points; dwell time 41.6 µs, spectral width 20 ppm or 12,019 Hz; number of averages, 64; dummy scans 4; pre-scan delay 10 µs. COSY reference spectra were acquired with the following parameters: 4096 points in the direct dimension, 256 points in the indirect dimension in Digital Quadrature Mode. Indirect dimension zero-filled to 4096 points. Number of averages, 4; dummy scans, 16; Spectral width 12.3 ppm, 7200 Hz; Temperature 298 K.
All experimental data were acquired and processed using Paravision 6.0.1 (Bruker Biospin, Ettlingen, Germany) and Topspin 3.1 running on Linux. The default bi-cubic interpolation was applied to enhance the details of the image prior to exporting images. Figures were prepared in Adobe Photoshop CC 2019 and Adobe Illustrator CC 2019 (Adobe Systems Incorporated, Mountain View, California, USA). Stacked spectra were produced in MestReNova (MestreLab Research S.L., Santiago de Compostela, Spain).
Light microscopy
After MR imaging, the sample material was dehydrated in an ethanol series and subsequently embedded in Technovit 7100 resin (Heraeus Kulzer, Hanau, Germany) according to the manufacturer’s protocol. Sections (5 µm) were made by using a microtome (Reichert-Jung, Leica Microsystems, Netherlands), stained with 0.05% Toluidine blue O for 1 min and additionally with Lugol’s iodine solution for a few seconds (if applicable). Sections were analysed by using a DM5500B microscope equipped with a DFC425C camera (Leica Microsystems, Wetzlar, Germany).
References
Doran, J. W. & Zeiss, M. R. Soil health and sustainability: managing the biotic component of soil quality. Appl. Soil Ecol. 15, 3–11 (2000).
Smil, V. Nitrogen cycle and world food production. World Agric. 9–13 (2001).
Smith, B. E. Nitrogenase reveals its inner secrets. Science 297, 1654–1655 (2002).
Suzaki, T., Yoro, E. & Kawaguchi, M. Leguminous Plants: Inventors of Root Nodules to Accommodate Symbiotic Bacteria. International Review of Cell and Molecular Biology 316, (Elsevier Ltd, 2015).
Kim, B. H., Ramanan, R., Cho, D. H., Oh, H. M. & Kim, H. S. Role of Rhizobium, a plant growth promoting bacterium, in enhancing algal biomass through mutualistic interaction. Biomass and Bioenergy 69, 95–105 (2014).
Barker, D. G. et al. Medicago truncatula, a model plant for studying the molecular genetics of the Rhizobium-legume symbiosis. Plant Mol. Biol. Report. 8, 40–49 (1990).
Nap, J. P. & Bisseling, T. Developmental biology of a plant-prokaryote symbiosis: The legume root nodule. Science 250, 948–954 (1990).
Udvardi, M. K. & Day, D. A. Metabolite transport across symbiotic membranes of legume nodules. Annu. Rev. Plant Physiol. Plant Mol. Biol. 48, 493–523 (1997).
Xiao, T. T. et al. Fate map of Medicago truncatula root nodules. Development 141, 3517–3528 (2014).
Kunert, K. J. et al. Drought Stress Responses in Soybean Roots and Nodules. Front. Plant Sci. 7, 1–7 (2016).
Staudinger, C. et al. Evidence for a rhizobia-induced drought stress response strategy in Medicago truncatula. J. Proteomics 136, 202–213 (2016).
López, M., Tejera, N. A. & Lluch, C. Validamycin A improves the response of Medicago truncatula plants to salt stress by inducing trehalose accumulation in the root nodules. J. Plant Physiol. 166, 1218–1222 (2009).
Ye, H. et al. MALDI mass spectrometry-assisted molecular imaging of metabolites during nitrogen fixation in the Medicago truncatula-Sinorhizobium meliloti symbiosis. Plant J. 75, 130–145 (2013).
Ogden, A. J., Gargouri, M., Park, J. J., Gang, D. R. & Kahn, M. L. Integrated analysis of zone-specific protein and metabolite profiles within nitrogen-fixing Medicago truncatula-Sinorhizobium medicae nodules. PLoS One 12, 1–17 (2017).
Schmittgen, S. et al. Magnetic resonance imaging of sugar beet taproots in soil reveals growth reduction and morphological changes during foliar Cercospora beticola infestation. J. Exp. Bot. 66, 5543–5553 (2015).
van Dusschoten, D. et al. Quantitative 3D analysis of plant roots growing in soil using magnetic resonance imaging. Plant Physiol. 170, 01388.2015 (2016).
Metzner, R., van Dusschoten, D., Bühler, J., Schurr, U. & Jahnke, S. Belowground plant development measured with magnetic resonance imaging (MRI): exploiting the potential for non-invasive trait quantification using sugar beet as a proxy. Front. Plant Sci. 5, 469 (2014).
Glover, P. & Mansfield, S. P. Limits to magnetic resonance microscopy. Reports Prog. Phys. 65, 1489–1511 (2002).
Vegh, V., Gläser, P., Maillet, D., Cowin, G. J. & Reutens, D. C. High-field magnetic resonance imaging using solenoid radiofrequency coils. Magn. Reson. Imaging 30, 1177–85 (2012).
Peck, T. L., Magin, R. L. & Lauterbur, P. C. Design and analysis of microcoils for NMR microscopy. Journal of magnetic resonance. Series B 108, 114–124 (1995).
Minard, K. R. & Wind, R. A. Solenoidal microcoil design. Part I: Optimizing RF homogeneity and coil dimensions. Concepts Magn. Reson. 13, 128–142 (2001).
Minard, K. R. & Wind, R. A. Solenoidal microcoil design Part II: Optimizing winding parameters for maximum signal-to-noise performance. Concepts Magn. Reson. 13, 190–210 (2001).
Chen, H.-Y. & Tycko, R. Low-temperature magnetic resonance imaging with 2.8 μm isotropic resolution. J. Magn. Reson. 287, 47–55 (2018).
Weiger, M. et al. NMR microscopy with isotropic resolution of 3.0 μm using dedicated hardware and optimized methods. Concepts Magn. Reson. Part B Magn. Reson. Eng. 33B, 84–93 (2008).
Mattle, E., Weiger, M., Schmidig, D., Boesiger, P. & Fey, M. MRI of human hair. Magn. Reson. Mater. Physics, Biol. Med. 22, 181–186 (2009).
Ciobanu, L., Seeber, D. A. & Pennington, C. H. 3D MR microscopy with resolution 3. 7 µm by 3.3 µm by 3.3 µm. J. Magn. Reson. 158, 178–182 (2002).
Lee, C. H., Flint, J. J., Hansen, B. & Blackband, S. J. Investigation of the subcellular architecture of L7 neurons of Aplysia californica using magnetic resonance microscopy (MRM) at 7.8 microns. Sci. Rep. 5, 11147 (2015).
Flint, J. J. et al. Magnetic resonance microscopy of mammalian neurons. Neuroimage 46, 1037–40 (2009).
Lee, C. H. et al. Magnetic Resonance Microscopy (MRM) of Single Mammalian Myofibers and Myonuclei. Nat. Publ. Gr. 1–9, https://doi.org/10.1038/srep39496 (2016).
Grant, S. C., Buckley, D. L., Gibbs, S., Webb, A. G. & Blackband, S. J. MR microscopy of multicomponent diffusion in single neurons. Magn. Reson. Med. 46, 1107–1112 (2001).
Ciobanu, L. & Pennington, C. H. 3D micron-scale MRI of single biological cells. Solid State Nucl. Magn. Reson. 25, 138–141 (2004).
Olson, D. L. et al. High-Resolution Microcoil 1H-NMR for Mass-Limited, Nanoliter-Volume Samples. Science (80-.). 270, 1967–1970 (1995).
Gavrin, A. et al. Adjustment of Host Cells for Accommodation of Symbiotic Bacteria: Vacuole Defunctionalization, HOPS Suppression, and TIP1g Retargeting in Medicago. Plant Cell 26, 3809–3822 (2014).
Dakora, F. D. & Atkins, C. A. Adaptation of Nodulated Soybean (Glycine max L. Merr.) to Growth in Rhizospheres Containing Nonambient pO2. PLANT Physiol. 96, 728–736 (1991).
O’Brien, T. P., Feder, N. & McCully, M. E. Polychromatic staining of plant cell walls by toluidine blue O. Protoplasma 59, 368–373 (1964).
Rivlin, M., Eliav, U. & Navon, G. NMR studies of proton exchange kinetics in aqueous formaldehyde solutions. J. Magn. Reson. 242, 107–112 (2014).
Rivlin, M., Eliav, U. & Navon, G. NMR studies of the equilibria and reaction rates in aqueous solutions of formaldehyde. J. Phys. Chem. B 119, 4479–4487 (2015).
Dupont, L. et al. The Legume Root Nodule: From Symbiotic Nitrogen Fixation to Senescence. In Senescence 2, 64 (InTech, 2012).
van der Graaf, M. In vivo magnetic resonance spectroscopy: basic methodology and clinical applications. Eur. Biophys. J. 39, 527–40 (2010).
Smith, L. T., Pocard, J. A., Bernard, T. & Le Rudulier, D. Osmotic control of glycine betaine biosynthesis and degradation in Rhizobium meliloti. J. Bacteriol. 170, 3142–9 (1988).
Ashraf, M. & Foolad, M. R. Roles of glycine betaine and proline in improving plant abiotic stress resistance. Environ. Exp. Bot. 59, 206–216 (2007).
Pfau, T. et al. The intertwined metabolism of Medicago truncatula and its nitrogen fixing symbiont Sinorhizobium meliloti elucidated by genome-scale metabolic models. bioRxiv, https://doi.org/10.1101/067348 (2016).
López, M., Herrera-Cervera, J. A., Iribarne, C., Tejera, N. A. & Lluch, C. Growth and nitrogen fixation in Lotus japonicus and Medicago truncatula under NaCl stress: Nodule carbon metabolism. J. Plant Physiol. 165, 641–650 (2008).
Jordy, M. N. Seasonal variation of organogenetic activity and reserves allocation in the shoot apex of Pinus pinaster ait. Ann. Bot. 93, 25–37 (2004).
Zeeman, S. C., Kossmann, J. & Smith, A. M. Starch: Its Metabolism, Evolution, and Biotechnological Modification in Plants. Annu. Rev. Plant Biol. 61, 209–234 (2010).
Thavarajah, R., Mudimbaimannar, V., Rao, U., Ranganathan, K. & Elizabeth, J. Chemical and physical basics of routine formaldehyde fixation. J. Oral Maxillofac. Pathol. 16, 400 (2012).
Lundquist, P. O. Nitrogenase activity in Alnus incana root nodules. Responses to O(2) and short-term N(2) deprivation. Plant Physiol. 122, 553–562 (2000).
Chudek, J. A., Hunter, G., Sprent, J. I. & Wurz, G. An application of NMR microimaging to investigate nitrogen fixing root nodules. Magn. Reson. Imaging 15, 361–368 (1997).
Brown, S. M. et al. Proton density and apoplastic domains within soybean nodules in relation to the oxygen diffusion barrier. Plant, Cell Environ. 20, 1019–1029 (1997).
Macfall, J. S., Pfeffer, P. E., Rolin, D. B., Macfall, J. R. & Johnson, G. A. Observation of the Oxygen Diffusion Barrier in Soybean (Glycine Max) Nodules with Magnetic-Resonance Microscopy. Plant Physiol. 100, 1691–1697 (1992).
Sulieman, S. & Tran, L. S. P. Asparagine: An amide of particular distinction in the regulation of symbiotic nitrogen fixation of legumes. Crit. Rev. Biotechnol. 33, 309–327 (2013).
Hirsch, A. M. & Smith, C. A. Effects of Rhizobium meliloti nif and fix mutants on alfalfa root nodule development. J. Bacteriol. 169, 1137–1146 (1987).
Hirsch, A. M., Bang, M. & Ausubel, F. M. Ultrastructural analysis of ineffective alfalfa nodules formed by nif::Tn5 mutants of Rhizobium meliloti. J. Bacteriol. 155, 367–380 (1983).
Prell, J. & Poole, P. Metabolic changes of rhizobia in legume nodules. Trends Microbiol. 14, 161–168 (2006).
Hohnjec, N., Perlick, A. M., Pühler, A. & Küster, H. The Medicago truncatula Sucrose Synthase Gene MtSucS1 Is Activated Both in the Infected Region of Root Nodules and in the Cortex of Roots Colonized by Arbuscular Mycorrhizal Fungi. Mol. Plant-Microbe Interact. 16, 903–915 (2007).
Yang, L. O., Udvardi, M. K. & Day, D. A. Specificity and regulation of the dicarboxylate carrier on the peribacteroid membrane of soybean nodules. Planta 182, 437–444 (1990).
Flint, J. J., Menon, K., Hansen, B., Forder, J. & Blackband, S. J. Metabolic support of excised, living brain tissues during magnetic resonance microscopy acquisition. J. Vis. Exp. 2017, 1–10 (2017).
Limpens, E. et al. RNA interference in Agrobacterium rhizogenes-transformed roots of Arabidopsis and Medicago truncatula. J. Exp. Bot. 55, 983–992 (2004).
Weidner, S. et al. Genome Sequence of Sinorhizobium meliloti Rm41. Genome Announc. 1, 12–13 (2013).
Littlejohn, G. R. & Love, J. A simple method for imaging Arabidopsis leaves using perfluorodecalin as an infiltrative imaging medium. J. Vis. Exp. 2, 3–6 (2012).
Schönherr, J. & Bukovac, M. J. Penetration of Stomata by Liquids. Plant Physiol. 49, 813–819 (1972).
Tsai, W. T. Environmental property modelling of perfluorodecalin and its implications for environmental fate and hazards. Aerosol Air Qual. Res. 11, 903–907 (2011).
Kim, H. K., Choi, Y. H. & Verpoorte, R. NMR-based metabolomic analysis of plants. Nat. Protoc. 5, 536–549 (2010).
Bottomley, P. A. Spatial Localization in NMR Spectroscopy in Vivo. Ann. N. Y. Acad. Sci. 508, 333–348 (1987).
Bottomley, P. A. Point resolved spectroscopy (PRESS). US Pat. 4, 228 (1984).
Tkác, I., Starcuk, Z., Choi, I. Y. & Gruetter, R. In vivo 1H NMR spectroscopy of rat brain at 1 ms echo time. Magn. Reson. Med. 41, 649–56 (1999).
Wishart, D. S. et al. HMDB: a knowledgebase for the human metabolome. Nucleic Acids Res. 37, D603–10 (2009).
Cui, Q. et al. Metabolite identification via the Madison Metabolomics Consortium Database. Nat. Biotechnol. 26, 162–4 (2008).
Acknowledgements
Experiments at the 950 MHz instrument were supported by uNMR-NL, an NWO-funded National Roadmap Large-Scale Facility of the Netherlands (project 184.032.207). R.S. was supported by the BioSolarCells consortium project U2.3. J.R.K. was supported by the NWO-funded graduate school Netherlands’ Magnetic Resonance Research School (NMARRS) graduate school [022.005.029]. D.S was supported by China Scholarship Council (201306040120). A.G.W. was partially supported by European Research Grant 670629 NOMA MRI. We thank Klaartje Houben, Marie Renault and Johan van der Zwan for technical support at the uNMR-NL facility. We would further like to thank Volker Lehmann, Henny Janssen and Pieter de Waard for help in troubleshooting.
Author information
Authors and Affiliations
Contributions
R.S., J.R.K., A.H.V., T.B., A.A. designed the experiments. R.S. and J.R.K. conducted MRI experiments. D.S. performed microscopy measurements. K.B.S.S.G. performed NMR measurements. R.S., J.R.K., K.B.S.S.G., F.J.V., A.A. analysed the results. H.J.M.G., A.G.W., H.V.A., A.H.V., A.A. supervised the project. R.S., J.R.K., A.A. wrote the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
van Schadewijk, R., Krug, J.R., Shen, D. et al. Magnetic Resonance Microscopy at Cellular Resolution and Localised Spectroscopy of Medicago truncatula at 22.3 Tesla. Sci Rep 10, 971 (2020). https://doi.org/10.1038/s41598-020-57861-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-57861-7
This article is cited by
-
NMR imaging and diffusion
Adsorption (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.