Abstract
Bacteriophage T7 is an intracellular parasite that recognizes its host via its tail and tail fiber proteins, known as receptor-binding proteins (RBPs). The RBPs attach to specific lipopolysaccharide (LPS) features on the host. Various studies have shown expansion of the phage’s host range via mutations in the genes encoding the RBPs, whereas only a few have shown contraction of its host range. Furthermore, most experimental systems have not monitored the alteration of host range in the presence of several hosts simultaneously. Here we show that T7 phage grown in the presence of five restrictive strains and one permissive host, each with a different LPS form, gradually avoids recognition of the restrictive strains. Remarkably, avoidance of the restrictive strains was repeated in different experiments using six different permissive hosts. The evolved phages carried mutations that changed their specificity, as determined by sequencing of the genes encoding the RBPs. This system demonstrates a major role for RBPs in narrowing the range of futile infections. The system can be harnessed for host-range contraction in applications such as detection or elimination of a specific bacterial serotype by bacteriophages.
Similar content being viewed by others
Introduction
Bacteriophages (phages) are ubiquitous biological entities that are found in habitats occupied by bacteria. As such, they influence the ecosystem of their habitat. Phages and bacteria exert mutual selection pressures in a never-ending molecular arms race. Therefore, phages must evolve and adapt to changing conditions. One of the ways in which phages can adapt is through changes in their host range.
A bacteriophage’s host range is defined as the span of hosts that it is capable of infecting1. This range is dependent on host factors (e.g., defense mechanisms such as CRISPR-Cas2 and restriction–modification systems3, presence of phage receptors4), environmental factors (e.g., temperature and pH5,6), and features encoded by the phage (e.g., receptor-binding proteins (RBPs)7). Specialist bacteriophages commonly display a narrow host range, i.e., they infect a limited number of bacterial strains of the same species. In contrast, generalist bacteriophages inherently display a broad host range8,9. It has been shown that the host range can be artificially expanded7. Similarly, host-range contraction is possible; it was shown, for example, that bacteriophage lambda can evolve to improve its binding to a specific receptor while losing the ability to bind another previously recognized receptor10.
From an evolutionary point of view, the host range of a phage can be paralleled to animals that choose one food source over another, or specialize in foraging and digesting a specific food source. Bull and colleagues11 built a mathematical model of phage–bacterium interactions based on the optimal foraging theory. They used phage T7 to validate their mathematical predictions by demonstrating its ability to discriminate between two bacterial strains. It could evolve to infect one strain of Escherichia coli while avoiding another strain that differed only by its surface molecules. They found that a single phage gene, 17, was responsible for the discrimination between the hosts.
Gene 17 encodes the tail fiber proteins that are essential for absorption because they recognize the outer membrane lipopolysaccharide (LPS) of E. coli. LPS molecules are abundant on the bacteria’s outer membrane; they are composed of lipid A, which is embedded in the outer membrane, and an appended core region containing keto-deoxy-d-manno-8-octanoic acid, heptose, galactose, and glucose. The biosynthetic pathway of the core region is partly catalyzed by enzymes encoded by the genes waaC, waaF, waaG, waaO, waaR, and waaU. E. coli hosts lacking these genes can be constructed, with the exception of waaU which is presumed to be an essential gene12,13. Deletion of any one of the other genes results in mutant forms of the LPS presented on the outer membrane of E. coli, as depicted in Fig. 1.
A simplified diagram of the LPS forms displayed on the outer membrane of mutants used in this study. The indicated genes encode enzymes required for the respective LPS biosynthesis. The indicated EOP of phage T7 on the different mutant hosts compared to K-12 is an average of six independent experiments.
We wanted to explore whether bacteriophage T7 can adapt and evolve to distinguish between altered forms of its natural LPS receptor. Specifically, we aimed to evolve T7 specialists that can recognize hosts with an altered LPS form, while avoiding hosts with other LPS forms and even with the wild-type (WT) form. Unlike Bull and colleagues11 who propagated T7 phage on a mixture of two hosts at a time, we evolved T7 in a mixture of up to six hosts—each with a different LPS form. Furthermore, we used a continuous evolution system that enabled us to propagate ~100 phage generations in as little as 95 h. In any given experiment, only one host in the mixture was permissive for the propagation of T7, while the other strains were restrictive due to deletion of trxA, an essential gene for T7 phage replication. trxA encodes thioredoxin that serves as a subunit of the phage DNA polymerase14. While its absence consequently halts phage replication and propagation, it does not affect adsorption and DNA injection. This setup directed the evolution of phage T7 toward recognizing a specific LPS form while avoiding the other forms. Using this method, we evolved six different specialist T7 phages, having mutations in tail genes 11 and 12, and in tail fiber gene 17. The evolved phages all showed decreased recognition of LPS forms other than the one they evolved to recognize. These experiments suggest that a major role of RBPs is to avoid restrictive strains, thereby preventing futile injection of the phage DNA.
Results
Bacteriophage T7 Evolves to Avoid Its Natural Receptor
We hypothesized that if we use T7 phage to infect two types of host: (1) a strain displaying a poorly recognized LPS but permissive for phage propagation and (2) a strain displaying the natural LPS receptor but restrictive for phage propagation, the phage will evolve to avoid its natural receptor and concomitantly increase its recognition of the permissive host. To test our hypothesis, we used two E. coli K-12 BW25113 strains (henceforth K-12) lacking either trxA (K-12ΔtrxA) or waaC (K-12ΔwaaC). E. coli lacking waaC displays only a partial LPS host receptor, resulting in an efficiency of plating (EOP) of only 2.5 × 10−4 of that of the K-12 strain (Fig. 1). In contrast, E. coli K-12 hosts lacking trxA enable efficient recognition and subsequent absorption, but the infection is abortive because the phage cannot propagate inside the host.
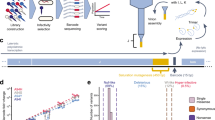
E. coli strains ΔtrxA and ΔwaaC were grown in separate flasks to a similar optical density at 600 nm (OD600) of ∼0.2, then mixed in equal volumes and transferred to a continuous evolution system based on a simple chemostat. In this system, E. coli are continuously grown in the chemostat with constant influx of fresh Luria-Bertani (LB) medium. They then flow to a second vessel called ‘lagoon’ (Fig. 2). The flow rate from the chemostat is fixed so that the E. coli are in the logarithmic growth phase when they enter the lagoon. The stay time of the bacteria in the lagoon is less than the time needed for E. coli to replicate, to avoid the accumulation of bacterial mutants. The lagoon contains bacteriophage T7, resulting in phage propagation. The replication time of the phage is shorter than the dilution rate of the lagoon, to avoid phage washout and thus enable selection of the fittest phages.
Illustration of the system used for the experiments. LB medium is flown by a pump to a chemostat. Bacteria are flown by a pump into a lagoon in a rate that maintains logarithmic growth. T7 bacteriophage are inoculated initially in the lagoon where they continuously infect bacteria. Both the chemostat and lagoon are diluted at a constant rate by a pump that flows media into a waste container.
The system was run for 95 h, allowing replication for ~100 generations of phage. Samples were taken from the lagoon upon addition of the phage (T0) and every 6–24 h thereafter. Phages were extracted from each sample and their EOP was tested on (1) the strain on which they were selected, i.e., the permissive host (in this case K-12ΔwaaC) and (2) a host displaying similar LPS receptors as the restrictive strain, but nonrestrictive as it encodes trxA. The latter strain contained a deletion of a control gene, ydhQ, which for simplicity will henceforth be omitted from the strains’ description in the main text (but not in Methods) as it does not change the LPS or any known parameter of T7 phage infection13. Figure 3 shows the changes in EOP over the course of 95 h. The permissive host served as the reference strain, and hence its value was fixed as 1. The EOP of the restrictive strain was calculated with respect to the permissive host. We used EOP measurements as they are technically easier to carry out in large scale compared to alternatives such as adsorption measurements. The EOP provides a relative number of plaque-forming units compared to a control, and in the absence of propagation barriers it also reflects on the actual adsorption of the phage.
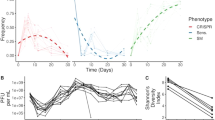
Evolution of phage T7 in the presence of two strains. (A) schematic illustration of the evolution. At the starting point of the experiment, most phages recognize the non-productive strain efficiently, while recognition of the permissive host is minimal. After evolving, most of the phage recognize the permissive strain most efficiently, while avoiding the non-productive host. (B) T7 phage was propagated in a continuous evolution experiment in the presence of K-12ΔwaaC permissive host and K-12ΔtrxA restrictive strain. Samples were collected at the indicated time points (dots) and the T7 plaque formation was determined on permissive isogenic strains encoding trxA. EOPs were normalized to the EOP of the permissive host.
This calculation expresses the difference between the EOP on the two host strains and how it changes over time as the phage evolves. Indeed, as hypothesized, the evolved phage reduced its recognition of the WT LPS form as determined by its EOP on the K-12 strain. While at T0 the phage infected K-12 hosts ∼3 orders of magnitude more efficiently than K-12ΔwaaC hosts, at T95 the evolved phage changed its differentiation of the two hosts by ∼5 orders of magnitude overall (Fig. 3). It recognized the permissive host by ∼2 orders of magnitude more efficiently than the WT phage, whereas it reduced its recognition of the K-12 strain by ~3 orders of magnitude compared to the WT phage (Dataset S1).
Bacteriophage T7 Evolves to Avoid Recognition of Restrictive Strains
We thus demonstrated that the continuous evolution system can select phages that avoid a restrictive strain displaying the natural receptor while recognizing permissive hosts having a poorly recognized receptor. We hypothesized that we could further evolve bacteriophage T7 to avoid multiple LPS forms simultaneously. To this end, we designed subsequent experiments that were similar to the first experiment. However, each experiment had six strains in total: five restrictive strains, and one permissive host. Each of these six strains displayed a different form of LPS, including the WT form. For example, to positively select for phages that specialize in infecting the Rc LPS form, which was found on a host lacking waaO, we used a permissive host lacking waaO (K-12ΔwaaO) and mixed it in the lagoon with equal volumes of additional E. coli hosts presenting all other LPS forms: the WT lacking trxA (K-12ΔtrxA), and another four hosts lacking trxA and waaC, waaF, waaG, or waaR (K-12ΔtrxAΔwaaC, K-12ΔtrxAΔwaaF, K-12ΔtrxAΔwaaG, and K-12ΔtrxAΔwaaR, respectively). The system was run for 95 h with samples taken every 4–19 h. The EOPs of the phage samples were determined on all six strains, all encoding trxA to enable plaque formation (K-12ΔwaaO, K-12ΔwaaC, K-12ΔwaaF, K-12ΔwaaG, K-12ΔwaaR, K-12). Remarkably, after 94 h, the evolved phage infected all of the restrictive strains at least 1000 times less efficiently than the permissive host (Fig. 4A). As expected, the evolved phage exhibited substitutions in gene product (Gp) 17: R200C, G479R, and R542S; Gp12: P694H; Gp11: S15P, all of which were likely responsible for the specialization (Table 1).
Evolution of phage T7 in the presence of six strains. T7 phage was propagated in a continuous evolution experiment in the presence of one permissive host (solid black line highlighted in yellow in each panel) and the other five restrictive strains (normal lines). Samples were collected at the indicated time points (dots) and the T7 plaque formation was determined on permissive isogenic strains encoding trxA. EOPs were normalized to the EOP of the permissive host. An arrow in panels B, D, and F indicates time of treatment with the EMS mutagen.
An additional five similar experiments were carried out by changing the permissive strain to display a different LPS form (Fig. 4B–F). Genes 11, 12, and 17 in the evolved phages were sequenced in all experiments, and the amino acid substitutions are summarized in Table 1. Many amino acid substitutions in Gp17 (residues 479, 480, 499, 520, 521, 540, 542) cluster to the very top of the tip domain, which forms a surface that likely directly interacts with and recognizes the LPS of E. coli (Fig. 5). These experiments showed that the WT T7 phage can evolve to better recognize a specific LPS form and simultaneously avoid other LPS forms.
Top view of the crystallographic structure of the carboxy-terminal domain of bacteriophage T7 tail fiber Gp17 containing residues 371–553 (PDB: 4A0T16). Side chains of residues G479, G480, C499, D520, G521, D540, and R542 are depicted in space-filled representation (labeled only in one of the three monomers). These substitutions cluster at the top of the tip domain which likely forms the LPS recognition surface.
Unexpectedly, when we positively selected for a phage recognizing the LPS form of K-12ΔwaaC, we found that after 95 h, the evolved specialist phage did not avoid the host K-12ΔwaaF (Fig. 4B). The LPS of K-12ΔwaaF contains one heptose group, which is lacking in K-12ΔwaaC (Fig. 1). We wanted to obtain a specialist phage that differentiates between these two hosts, in addition to the other hosts, which it discriminated against by at least 1000-fold. To this end, we facilitated mutations by treating the phages that evolved for 95 h with the mutagenic chemical ethyl methanesulfonate (EMS). Next, the treated phages were reintroduced to the hosts in the lagoon of the continuous evolution system for a second time, for an additional 94 h. During that time, the EOP on all strains was measured. Remarkably, the EOP for K-12ΔwaaF was now reduced by ∼2 orders of magnitude, demonstrating that the facilitated evolution enabled stricter avoidance of futile hosts. Moreover, we found that phages sampled after 4 h (99 h total in both rounds of the system) completely lost recognition of the restrictive K-12ΔwaaR, and after 21 h (116 h total), the phages lost recognition of the restrictive K-12. Similarly, when the permissive host was K-12ΔwaaG, we found that the evolved phage did not adequately avoid the restrictive strains K-12ΔwaaO and K-12ΔwaaF after 95 h (Fig. 4D), and when the permissive host was K-12ΔwaaR, the evolved phage did not adequately avoid the restrictive strains K-12ΔwaaO and K-12ΔwaaG after 95 h (Fig. 4F). EMS treatment in these cases also reduced recognition of the restrictive strains by accumulating mutations in genes 11, 12, and 17 (Fig. 4D,F; Table 1). These experiments demonstrated that with facilitated mutation rates or presumably, with longer evolution time, the phages evolve to further de-recognize and discriminate against restrictive strains.
Discussion
We showed simultaneous contraction of phage T7’s host range to exclude five different hosts in six different experimental systems. In all cases, we observed that the phages evolved to discriminate against restrictive strains. However, the evolved phages did not always improve their recognition of the permissive host. These findings highlight the notion that a significant role for RBPs is in avoiding restrictive strains, in addition to recognizing permissive hosts. This concept can be envisioned by imagining that the first common ancestor of all current phages developed a simple mechanism to recognize bacterial membranes, which did not require a complex protein structure. Presumably, merely displaying a hydrophobic surface would suffice for such recognition. However, due to the non-selective recognition, some of the progeny of this ancestor infected bacteria that did not support their growth, thus becoming extinct. We speculate that a RBP gradually evolved to avoid recognition of these nonproductive bacterial strains, thus providing a competitive advantage over those phages with the simpler recognition elements. The common perception that RBPs evolved to positively signal the phage as to which host to infect probably stems from observations on host range-expansion observed in the evolution of certain phages15. In our system, nevertheless, the RBP is more of a negative signal, indicating primarily where the phage should not attach, rather than being a positive signal for host recognition.
Further support for RBPs evolving to restrict recognition of futile hosts stems from our experiments using K-12 as the permissive host. Arguably, the T7 phage RBPs are fully adapted to recognizing K-12 LPS. When K-12 was used as the permissive host, we observed loss of the T7 phage’s ability to recognize other LPS forms. For example, in less than 20 h of continuous evolution, it completely avoided K-12ΔwaaC, and markedly avoided K-12ΔwaaG. We thus demonstrated that the RBP has the potential to avoid other forms of LPS, but does so only if those forms are displayed by restrictive strains. In this case too, the RBP did not evolve to better recognize the K-12 LPS, but only to avoid other hosts. These results reiterate the major role of the RBPs in avoiding restrictive strains.
The latter experiment raises the question of why T7 phage reserves residual recognition of these mutant hosts in the first place, when it can evolve to avoid them efficiently? The likely answer is that these hosts are permissive in nature, and the phage retains the option to infect them, should the host display such mutant receptors. If the phage evolved to avoid residual recognition of those receptors, the bacterial host would evolve to display only those receptors and the phage population would become extinct. We therefore postulate that, in nature, the RBPs of most phages also recognize the derivatives of the major receptor that may be displayed by mutated hosts, but avoid any other receptors that are displayed by restrictive strains. In this respect, it is noteworthy that host restriction in nature can occur in various ways such as restriction-modification systems, CRISPR-Cas systems, and abortive infection systems.
Many of the substitutions we identified clustered at the tip of Gp17. Previously identified substitutions in Gp17 involved in host avoidance were also located on this surface11. Intriguingly, the other large concentration of substitutions occurs at the phage-proximal rod (residues 150–260), that is located 10–15 nm away from the tip domain16,17. Possibly the proximal rod can also interact with bacterial receptors at some stage of viral infection or alternatively, mutations in the rod could modify the orientation of the tip domain with respect to the bacterial surface, indirectly regulating interactions with LPS. Several substitutions appear also in Gp11 and Gp12 suggesting that the nozzle of the viral DNA injection machinery can also adapt to varying LPS compositions. It is also noteworthy that no substitutions were found in the pyramid domain between the phage-proximal rod and the tip, suggesting that this domain is not involved in avoidance of undesired LPS receptors.
In addition to the theoretical implications of this study, we demonstrate how a continuous evolution system can be used for host-range contraction of bacteriophages. This system may be useful for other research areas that utilize phages. For example, a narrow host range is desirable in bacteriophage therapy. A phage that is limited to a specific species, and even a strain within that species, is sometimes sought because it is likely to minimally interact with other bacteria, and it will therefore not affect the host’s microbiome. It might also help produce phages that are more likely to overcome regulatory demands due to their specific host range. Furthermore, phages are used to identify specific bacteria18. Contracting the phage host range to desired bacteria may provide specific identification of certain serotypes. Thus, this study provides both conceptual and translational implications for phage biology and phage applications.
Materials and Methods
Reagents
LB medium (10 g/L tryptone, 5 g/L yeast extract, and 5 g/L NaCl) and agar were from Acumedia. Antibiotics were from Calbiochem. EMS was from Sigma-Aldrich. Kappa® High-Fidelity Polymerase was from Kappa Biosystems.
Strains
The bacterial strains used in this study are listed in Table S1. Single knock-out mutants, BW25113ΔwaaC, BW25113ΔwaaF, BW25113ΔwaaG, BW25113ΔwaaR, BW25113ΔwaaO, BW25113ΔtrxA, and BW25113ΔydhQ were acquired from the Keio collection12. IYB5709 is a strain lacking trxA from the Keio collection12 that was flipped for its resistance cassette using pCP20 as previously described19. It was then used as an ancestor strain in homologous recombination to produce double mutants as described below. The WT bacteriophage T7 used in this study has a Gp12 S694P substitution compared to the published sequence (GenBank accession no. CAA24430.1), but is otherwise similar to GenBank AY264774. Phage-generation time during continuous evolution was calculated as previously described20.
Homologous Recombination
Homologous recombination to produce double mutants was performed as previously described21. Briefly, an overnight culture of IYB5709 harboring pSIM6 was diluted 70-fold in 35 mL LB medium supplemented with appropriate antibiotics and grown at 32 °C with shaking to an OD600 of ∼0.5. The culture was heat-induced at 42 °C for 15 min to express the lambda Red proteins. Then the culture was placed on ice, and pelleted at 3,600 g at 0 °C for 10 min. The pellet was washed twice in ice-cold double-distilled H2O (ddH2O) and then resuspended in 200 µL ice-cold ddH2O and kept on ice until electroporation with 500 ng of a gel-purified PCR product. The different PCR products were amplified from the Keio collection strains listed in Table S1. Each PCR product encoded a kanamycin resistance gene with a minimum of 50-bp flanking ends homologous to the target gene. Primers used to produce the different cassettes are listed in Table S2. A 50-µL aliquot of electrocompetent cells was used for each electroporation in a 0.1-cm cuvette. After electroporation, the bacteria were recovered in 1 mL LB medium for 2 h at 32 °C in a shaking water bath and inoculated on selection plates supplemented with 25 µg/mL kanamycin. Homologous recombination into the resulting strains BW25113ΔtrxA, BW25113ΔtrxAΔwaaF, BW25113ΔtrxAΔwaaO, BW25113ΔtrxAΔwaaR, BW25113ΔtrxAΔwaaG, and BW25113ΔtrxAΔydhQ, was confirmed by PCR using the primers IY21F and 21 R to ensure the presence of the KanR cassette. In addition, to ensure that the insert entered in the desired location, another PCR was carried out using IY21F and the respective forward or reverse primer that was used to produce the PCR cassette (Table S2).
Plaque Assay and Lysate Preparation
Plaque assay for quantification of the number of infectious phage particles was performed as previously described22. Briefly, chloroform was added to a 5-mL culture of exponentially growing cells mixed with T7 phage extracted from the lagoon of the continuous evolution system at different time points. The lysate was cleared by centrifugation. Then, it was serially diluted and spotted on LB-agar plates with a layer of soft agar containing the appropriate host cells. Plaques were counted and normalized to values obtained by plating on the permissive host.
EMS Treatment of Phages
Overnight cultures of E. coli BW25113ΔydhQ were diluted 1:50 in 5 mL of LB medium supplemented with 25 ug/ml of Kanamycin. The culture was aerated at 37 °C for several hours. Upon reaching mid log-phase, cells were infected with the evolved T7 extracted from the end of each continuous evolution experiment as described above. EMS (Sigma-Aldrich) was then added at 1% to the culture infected with the evolved phage and aerated at 37 °C until lysis appeared. Then, phages were precipitated with PEG 8000 (Promega) and washed twice with 5 mL PBS as previously described23.
Continuous Evolution
A schematic of the system can be found in Fig. 2. The system was composed of two connected 500 mL vessels each contained 125 mL culture, stirred continuously. The first vessel, “chemostat”, was used for E. coli growth, and the second vessel, “lagoon”, for viral evolution. Before entering the system, each host strain was grown separately with shaking at 37 °C in 125 mL flask containing 25 mL LB with appropriate antibiotics until reaching ∼0.5 OD600. Then, all hosts were mixed in equal parts to total volume of 125 mL and were introduced aseptically into the chemostat. Fresh LB media was added continuously to the chemostat with a flow rate of 2 volumes per hour (2 V/h) until reaching a steady stare of ∼1.0 OD600. Then, the chemostat culture was flown into the lagoon with a flow rate of 2 V/h, which was also the flow rate used for fluid removal from the lagoon to the waste container. At T0, T7 bacteriophage was aseptically injected to the lagoon with a MOI of 1. Samples of 15 mL were taken aseptically from the chemostat and from the lagoon via sampling ports, from which, 5 mL were used to prepare a lysate as described above. In order to maintain bacterial strain ratio in the chemostat, 300 mL of freshly prepared bacterial mix with ~1 OD600 was aseptically added to the chemostat culture every 24 h. At that time, media supplement to the chemostat vessel was stopped until reaching the initial volume of 125 mL.
References
Hyman, P. & Abedon, S. T. Bacteriophage host range and bacterial resistance. Adv. Appl. Microbiol. 70, 217–248 (2010).
Barrangou, R. et al. CRISPR provides acquired resistance against viruses in prokaryotes. Science 315, 1709–1712 (2007).
Bertani, G. & Weigle, J. J. Host controlled variation in bacterial viruses. J. Bacteriol. 65, 113–121 (1953).
Smith, S. G., Mahon, V., Lambert, M. A. & Fagan, R. P. A molecular Swiss army knife: OmpA structure, function and expression. FEMS Microbiol. Lett. 273, 1–11 (2007).
Fister, S. et al. Influence of Environmental Factors on Phage-Bacteria Interaction and on the Efficacy and Infectivity of Phage P100. Front. Microbiol. 7, 1152 (2016).
Kerby, G. P. & Gowdy, R. A. et al. Purification pH stability and sedimentation properties of the T7 bacteriophage of Escherichia coli. J. Immunol. 63, 93–107 (1949).
Yosef, I., Goren, M. G., Globus, R., Molshanski-Mor, S. & Qimron, U. Extending the Host Range of Bacteriophage Particles for DNA Transduction. Mol. Cell 66, 721–728.e723 (2017).
Yu, P., Mathieu, J., Li, M., Dai, Z. & Alvarez, P. J. Isolation of Polyvalent Bacteriophages by Sequential Multiple-Host Approaches. Appl. Env. Microbiol. 82, 808–815 (2016).
Zborowsky, S. & Lindell, D. Resistance in marine cyanobacteria differs against specialist and generalist cyanophages. Proc. Natl Acad. Sci. U S A 116, 16899–16908 (2019).
Meyer, J. R. et al. Ecological speciation of bacteriophage lambda in allopatry and sympatry. Science 354, 1301–1304 (2016).
Heineman, R. H., Springman, R. & Bull, J. J. Optimal foraging by bacteriophages through host avoidance. Am. Nat. 171, E149–157 (2008).
Baba, T. et al. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol. Syst. Biol. 2, 2006.0008 (2006).
Qimron, U., Marintcheva, B., Tabor, S. & Richardson, C. C. Genomewide screens for Escherichia coli genes affecting growth of T7 bacteriophage. Proc. Natl Acad. Sci. U S A 103, 19039–19044 (2006).
Mark, D. F. & Richardson, C. C. Escherichia coli thioredoxin: a subunit of bacteriophage T7 DNA polymerase. Proc. Natl Acad. Sci. USA 73, 780–784 (1976).
Hall, J. P., Harrison, E. & Brockhurst, M. A. Viral host-adaptation: insights from evolution experiments with phages. Curr. Opin. virology 3, 572–577 (2013).
Garcia-Doval, C. & van Raaij, M. J. Structure of the receptor-binding carboxy-terminal domain of bacteriophage T7 tail fibers. Proc. Natl Acad. Sci. U S A 109, 9390–9395 (2012).
Steven, A. C. et al. Molecular substructure of a viral receptor-recognition protein. The gp17 tail-fiber of bacteriophage T7. J. Mol. Biol. 200, 351–365 (1988).
Ripp, S. Bacteriophage-based pathogen detection. Adv. Biochem. Eng. Biotechnol. 118, 65–83 (2010).
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl Acad. Sci. U S A 97, 6640–6645 (2000).
Esvelt, K. M., Carlson, J. C. & Liu, D. R. A system for the continuous directed evolution of biomolecules. Nature 472, 499–503 (2011).
Sharan, S. K., Thomason, L. C., Kuznetsov, S. G. & Court, D. L. Recombineering: a homologous recombination-based method of genetic engineering. Nat. Protoc. 4, 206–223 (2009).
Kropinski, A. M., Mazzocco, A., Waddell, T. E., Lingohr, E. & Johnson, R. P. Enumeration of bacteriophages by double agar overlay plaque assay. Methods Mol. Biol. 501, 69–76 (2009).
Manor, M. & Qimron, U. Selection of Genetically Modified Bacteriophages Using the CRISPR-Cas System. Bio Protoc 7, (2017).
Acknowledgements
This project has received funding from the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation program (grant no. 818878), the ERC PoC program (grant no. 811322), the Israel Science Foundation (grant no. 268/14), and the Israeli Ministry of Science (grant no. 3-14351).
Author information
Authors and Affiliations
Contributions
Conceptualization, T.H., R.G., U.Q.; Methodology, T.H., R.G., S.M.M., and I.Y.; Writing –Original Draft, R.G. and U.Q.; Writing – Review & Editing, T.H., R.G., I.Y., A.B.S., and U.Q.; Funding Acquisition, U.Q.; Supervision, U.Q.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Holtzman, T., Globus, R., Molshanski-Mor, S. et al. A continuous evolution system for contracting the host range of bacteriophage T7. Sci Rep 10, 307 (2020). https://doi.org/10.1038/s41598-019-57221-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-57221-0
This article is cited by
-
Biochemical characterisation and production kinetics of high molecular-weight (HMW) putative antibacterial proteins of insect pathogenic Brevibacillus laterosporus isolates
BMC Microbiology (2024)
-
PHEIGES: all-cell-free phage synthesis and selection from engineered genomes
Nature Communications (2024)
-
A global atlas of soil viruses reveals unexplored biodiversity and potential biogeochemical impacts
Nature Microbiology (2024)
-
Hi-C metagenome sequencing reveals soil phage–host interactions
Nature Communications (2023)
-
Biological control of the fire blight pathogen Erwinia amylovora using bacteriophages
Journal of Plant Pathology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.