Abstract
Addressing the growing problem of antibiotic resistance requires the development of new drugs with novel antibacterial targets. FtsZ has been identified as an appealing new target for antibacterial agents. Here, we describe the structure-guided design of a new fluorescent probe (BOFP) in which a BODIPY fluorophore has been conjugated to an oxazole-benzamide FtsZ inhibitor. Crystallographic studies have enabled us to identify the optimal position for tethering the fluorophore that facilitates the high-affinity FtsZ binding of BOFP. Fluorescence anisotropy studies demonstrate that BOFP binds the FtsZ proteins from the Gram-positive pathogens Staphylococcus aureus, Enterococcus faecalis, Enterococcus faecium, Streptococcus pyogenes, Streptococcus agalactiae, and Streptococcus pneumoniae with Kd values of 0.6–4.6 µM. Significantly, BOFP binds the FtsZ proteins from the Gram-negative pathogens Escherichia coli, Klebsiella pneumoniae, Pseudomonas aeruginosa, and Acinetobacter baumannii with an even higher affinity (Kd = 0.2–0.8 µM). Fluorescence microscopy studies reveal that BOFP can effectively label FtsZ in all the above Gram-positive and Gram-negative pathogens. In addition, BOFP is effective at monitoring the impact of non-fluorescent inhibitors on FtsZ localization in these target pathogens. Viewed as a whole, our results highlight the utility of BOFP as a powerful tool for identifying new broad-spectrum FtsZ inhibitors and understanding their mechanisms of action.
Similar content being viewed by others
Introduction
The discovery and development of antibiotics have saved millions of lives and revolutionized modern medicine1. However, the alarming rise in multidrug-resistant (MDR) bacteria has threatened the usefulness of our current arsenal of antibiotics, leading the World Health Organization (WHO) to suggest an impending post-antibiotic era, where minor infections would become lethal2. A recent WHO report highlights the global magnitude of an ever-worsening crisis, exacerbated by a chronic shortage of antibiotics capable of treating infections cause by MDR pathogens3. Currently, the majority of drug candidates in the antibiotic pipeline are derivatives of known antibiotics that will provide only a short-term solution to the problem3. Furthermore, many of these drug candidates have limited or no activity against MDR Gram-negative bacterial pathogens, which are particularly problematic to treat3.
Addressing the global antibiotic resistance problem requires the development of new drug chemotypes and the identification of new antibacterial drug targets. The filamentous temperature-sensitive Z (FtsZ) protein has been identified as a promising new target for the development of novel antibiotics4,5,6,7,8,9,10,11,12,13,14. FtsZ has several properties that make it an appealing antibacterial drug target. It is an essential protein required for bacterial division15,16,17. Inhibition of FtsZ has a bactericidal rather than bacteriostatic effect8,11, a property that reduces the potential for emergence of future resistance18. FtsZ also has no functional human homolog, offering the potential to target this protein specifically and with minimal toxicity12,19. It is one of the most abundant and highly conserved cytoskeleton proteins among eubacteria20, offering FtsZ inhibitors the potential for broad-spectrum antibacterial activity. Lastly, FtsZ is a “druggable” target whose function can be disrupted by small molecule targeting of a single site on the protein. While this latter property is appealing for a target protein, it also introduces the potential for the development of resistance via single mutations7,8,11,12,13,14,21,22.
We have previously developed prodrugs of benzamide FtsZ inhibitors (PC190723 and TXA707) that are highly efficacious against infections caused by methicillin-resistant Staphylococcus aureus (MRSA)9,10,11. One of these prodrugs (TXA709) is currently in phase I clinical trials6. To date, the bulk of the compounds that have been validated as FtsZ inhibitors both in vitro with purified FtsZ and in bacterial cells are associated with potent activity against staphylococci, Mycobacterium tuberculosis, and select other Gram-positive bacterial strains, but weaker or no activity against Gram-negative species8,11,12,13,23,24. Demonstration of in vivo efficacy among these FtsZ inhibitors has been limited almost exclusively to the treatment of S. aureus and M. tuberculosis infections7,8,9,10,11,12,14,24,25.
Advancing the development of new FtsZ inhibitors that can target a more expansive array of both Gram-positive and Gram-negative bacterial pathogens requires tools that allow us to screen for FtsZ inhibition in a broad range of bacterial species. Fluorescent antibiotics are useful tools for delineating the mechanisms underlying the antibacterial activities of compounds as well as the resistance phenotypes of bacteria26. In addition, such tools can be used to screen for new antibiotic candidates with desired mechanisms of action26. Early efforts aimed at developing fluorescent FtsZ inhibitors were centered on analogs of the benzamide inhibitor PC19072327. Several of these fluorescent analogs were shown to bind FtsZ from both S. aureus and Bacillus subtilis (SaFtsZ and BsFtsZ, respectively), though the interactions were weak (with estimated Kd values in the range of 11 to 29 µM for BsFtsZ at 25 °C), and none of the analogs were able to bind FtsZ from Escherichia coli (EcFtsZ) to a significant degree27. One analog was used to visualize FtsZ in S. aureus and B. subtilis cells27. However, visualization required prolonged (1- to 3-hour) treatment with large concentrations of the analog (25 to 200 µM) and was lost upon pre-treatment with the parent inhibitor PC190723, limiting the usefulness of the analog as a screening tool for FtsZ inhibitors.
Here we report the structure-guided design and characterization of a next-generation fluorescent FtsZ probe (BOFP) that overcomes the limitations associated with the early-generation analogs. Our design incorporates an oxazole-benzamide FtsZ inhibitor (1)13 (shown in Fig. 1a), whose crystal structure in complex with SaFtsZ we have previously determined (PDB entry: 5XDU)21, conjugated to a boron-dipyrromethene (BODIPY) fluorophore at the linker joining the oxazole and benzamide rings. Fluorescence anisotropy studies demonstrate that BOFP can target the FtsZ proteins from a broad range of Gram-positive pathogens (including S. aureus, Enterococcus faecalis, Enterococcus faecium, Streptococcus pyogenes, Streptococcus agalactiae, and Streptococcus pneumoniae) with high affinity (Kd values in the range of 1.0 to 3.5 µM at 25 °C). Significantly, BOFP targets the FtsZ proteins from clinically important Gram-negative pathogens (including E. coli, Klebsiella pneumoniae, Pseudomonas aeruginosa, and Acinetobacter baumannii) with an even higher affinity (Kd values in the range of 0.3 to 0.6 µM at 25 °C). The crystal structure of BOFP in complex with SaFtsZ confirms that the probe targets the same site on the protein as the parent compound, while revealing contributions from the BODIPY functionality itself to the stability of the complex. Most importantly, fluorescence microscopy studies demonstrate that brief (5-minute) exposure to BOFP (at a concentration of only 1.3 µM) can be used to visualize FtsZ in all the Gram-positive and Gram-negative bacterial pathogens listed above, even when pre-treated with other non-fluorescent FtsZ inhibitors. Taken together, our results indicate that BOFP can serve as a powerful tool for identifying new broad-spectrum FtsZ inhibitors and understanding their mechanisms of action.
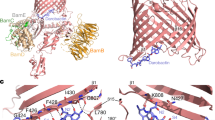
(a) Chemical structures of the oxazole-benzamide FtsZ inhibitors 1 and 2. The methyl group in 2 that differentiates this compound from 1 is highlighted in red. 2 was prepared as a racemic mixture of R and S enantiomers. (b) Expanded view of the binding site for the R enantiomer of 2 [(R)-2] in complex with SaFtsZ, with the F0 – Fc omit map (cyan) being contoured at 3.0σ. The anomalous difference map (purple) is contoured at 4.0σ. (c) Superposition of the SaFtsZ−(R)-2 complex (orange) with the corresponding SaFtsZ−1 complex (blue). The methyl group in 2 shown in red in (a) is highlighted by the red arrows in (b,c).
Results and Discussion
Structure-guided identification of a suitable site on the oxazole-benzamide FtsZ inhibitor for conjugation of a fluorophore
A clue to a potential site on the oxazole-benzamide FtsZ inhibitor 1 for conjugation of a fluorophore came from the crystal structure we determined for the complex of SaFtsZ12–316 (the enzymatic domain of SaFtsZ, residues 12–316) with 213, a methyl analog of 1 whose chemical structure is shown in Fig. 1a. The only difference between the two compounds is the presence of a methyl group (shown in red in Fig. 1a) on the linker connecting the oxazole and difluorobenzamide rings of 2 that is absent in 1. This difference makes 2 a chiral molecule, while 1 is achiral. A racemic mixture of the R and S enantiomeric forms of 2 was dissolved in DMSO and introduced into crystals of SaFtsZ12–316 by soaking. The structure of the R enantiomer of 2 [(R)-2] in complex with SaFtsZ12–316 was determined at 1.4 Å resolution. An extra electron density was clearly observed in the cleft between the N- and C-terminus domains, which enabled us to determine the position and orientation of (R)-2 with an occupancy of 1.0 (Fig. 1b). We have previously reported the structure of 1 in complex with SaFtsZ12–31621. (R)-2 binds the same protein cleft as 1 in a similar orientation to that observed in the structure of the 1-SaFtsZ12–316 complex (see the overlay of 1 and (R)-2 depicted in Fig. 1c). Note that no SaFtsZ complexes were observed with the S enantiomer of 2, suggesting that SaFtsZ is selective for the R over the S enantiomer.
Inspection of the crystal structure of (R)-2 in complex with SaFtsZ12–316 reveals that the methyl group distinguishing (R)-2 from 1 does not engage in significant interactions with SaFtsZ, but rather is oriented away from the cleft of the FtsZ molecule (as highlight by the red arrows in Fig. 1b,c). This observation suggested to us that a bulky fluorescent moiety could be conjugated to the same site on the linker in 1 as the methyl group in 2 without disrupting the FtsZ binding interaction to a significant degree.
Design of a BODIPY-conjugated fluorescent derivative of 1 (BOFP) that binds SaFtsZ with affinity in the Kd range of 0.9 to 3.1 µM
Armed with the structural results described above, we designed and synthesized BOFP, a fluorescent analog of 1 in which a BODIPY fluorophore is conjugated to the desired site on 1 by reaction of BODIPY FL carboxylate (BODIPY FL-COOH) with 312,13 (a hydroxymethyl analog of 1 prepared as a racemic mixture of R and S enantiomers) as schematically depicted in Fig. 2a. This reaction resulted in a racemic mixture of BOFP as well. In our initial characterizations, we sought to determine whether BOFP could still bind SaFtsZ, as hypothesized in our structure-guided design approach. Toward this end, we used fluorescence anisotropy to monitor the interaction of BOFP with SaFtsZ at three different temperatures (15, 25, and 37 °C). Significantly, the fluorescence anisotropy (r) of BOFP increases markedly with added SaFtsZ (Fig. 2b), indicating the presence of a binding interaction. No such anisotropy change was observed in control studies with unreacted BODIPY FL-COOH (Fig. S1). This important control observation not only demonstrates that BODIPY FL-COOH does not interact with the target FtsZ protein but also that the ester linkage connecting the BODIPY fluorophore in the BOFP conjugate is not hydrolyzed during the binding reactions, as BODIPY FL-COOH would be the fluorescent product of such a hydrolysis.
(a) Scheme for the synthesis of BOFP by reacting 3 with BODIPY FL-COOH in CH2Cl2 containing 1-ethyl-3-(3-dimethylaminopropyl)carbodiimide (EDC) and 4-dimethylaminopyridine (DMAP). Both 3 and BOFP were prepared as racemic mixtures of the R and S enantiomers. The tethered hydroxymethyl functionality on 3 and BODIPY functionality on BOFP are both highlighted in red. (b) Fluorescence anisotropy profiles of 0.1 µM BOFP as a function of increasing concentrations of SaFtsZ. The titration experiments were conducted at 15 °C (red), 25 °C (black), or 37 °C (cyan) in solution containing 50 mM Tris-HCl (pH 7.6) and 50 mM KCl. The solid lines reflect non-linear least squares fits of the experimental data points with Eq. 1. (c) Expanded view of the binding site for the R enantiomer of BOFP [(R)-BOFP] in complex with SaFtsZ, with the F0 – Fc omit map (cyan) being contoured at 2.0σ. (d) Superposition of the SaFtsZ−(R)-BOFP complex (green) with the corresponding SaFtsZ−1 complex (blue). (e) Hydrophobic interactions at the surface interface between the BODIPY moiety of BOFP and residues Ile228, Val230, and Val307 of SaFtsZ.
Analysis of the r-based binding isotherms of BOFP with the 1:1 binding formalism embodied by Eq. 1 yielded excellent fits of the experimental data points (depicted by the solid curves in Fig. 2b) and the corresponding Kd values listed in Table 1, which ranged from 0.88 ± 0.08 µM at 15 °C to 3.14 ± 0.13 µM at 37 °C. These gratifying results indicate that BOFP can bind SaFtsZ with a robust affinity in the sub- to low-micromolar Kd range.
The binding of BOFP to SaFtsZ does not require the presence of GTP or magnesium
The filamentation of SaFtsZ requires the presence of both GTP and magnesium28. Note that neither of these reagents was present in the fluorescence anisotropy binding studies depicted in Fig. 2b, indicating that the binding of BOFP to SaFtsZ does not require the presence of GTP or magnesium. This observation markedly contrasts the fluorescence anisotropy studies previously reported by Artola et al.27, which demonstrated that the fluorescent analogs of the benzamide FtsZ inhibitor PC190723 required both GTP and magnesium in order to bind SaFtsZ. For comparative purposes, we sought to determine whether the presence of a non-hydrolyzable analog of GTP (GMPCPP) and magnesium exerted an impact on the binding of BOFP to SaFtsZ, as reflected by a change in fluorescence anisotropy. At identical concentrations of BOFP (0.1 µM) and SaFtsZ (10 µM), the presence of neither GMPCPP alone (at 0.1 mM) nor MgCl2 alone (at 10 mM) has a significant effect on the anisotropy of SaFtsZ-bound BOFP (Fig. S2), confirming that the binding of BOFP to SaFtsZ is independent of either GTP or magnesium. The presence of both GMPCPP and MgCl2 results in a modest increase in the anisotropy of bound BOFP (Fig. S2), which likely reflects the filamentation of BOFP-bound SaFtsZ induced by the combination of GMPCPP and magnesium.
BOFP binds in the same cleft of SaFtsZ as 1 and 2
We next sought to determine the crystal structure of BOFP in complex with SaFtsZ12–316. Toward this end, a racemic mixture of the R and S enantiomeric forms of BOFP in DMSO was introduced into crystals of SaFtsZ12–316 by soaking. The structure of the R enantiomer of BOFP [(R)-BOFP] in complex with SaFtsZ12–316 was determined at 1.6 Å resolution. Although the electron density of (R)-BOFP was not quite as robust as that of (R)-2 due to a lower occupancy of 0.6, we were able to confirm that (R)-BOFP does indeed bind in the same cleft (compare Figs. 1b and 2c). The conjugated BODIPY extends toward the outside of the FtsZ molecule, away from the binding cleft. As hypothesized, interactions between the 1 portion of (R)-BOFP and SaFtsZ are similar to those exhibited by 1 (see the overlay of 1 and (R)-BOFP depicted in Fig. 2d). In addition to these interactions, the complex of (R)-BOFP with SaFtsZ is further stabilized by hydrophobic interactions between the BODIPY moiety and the hydrophobic surface formed by residues Ile228, Val230, and Val307 (Fig. 2e), with these interactions being unlikely to hamper the ability of SaFtsZ to self-associate into filaments (Fig. S3).
As seen with 2, no SaFtsZ complexes were observed with the S enantiomer of BOFP, further suggestive of the selectivity of SaFtsZ for the R versus the S enantiomeric form. Previous studies by Stokes et al. have shown that the R enantiomeric form of 3 has significantly greater activity against methicillin-sensitive S. aureus (MSSA) than the S enantiomeric form13. This enhanced antistaphylococcal activity of the R enantiomer likely reflects the corresponding selectivity of SaFtsZ for the R enantiomeric form.
Note that the FtsZ targeting of BOFP confers the compound with antistaphylococcal activity, though this activity is somewhat reduced relative to the parent compounds 1 and 3 (MIC versus MRSA NRS705 = 0.25, 0.5, and 1.0 µg/mL for 1, 3, and BOFP, respectively). The reduced activities of both 3 and BOFP relative to 1 may be due in part to 3 and BOFP being racemic mixtures of active R and weakly active S enantiomers. In the aggregate, our collective fluorescence anisotropy, crystallographic, and antibacterial results for BOFP serve to validate our structure-guided design approach.
BOFP can target the FtsZ proteins from a broad range of clinically important Gram-positive bacterial pathogens, including enterococcal and streptococcal species
In addition to SaFtsZ, we also sought to determine whether BOFP can target the FtsZ proteins from other Gram-positive bacterial pathogens, including E. faecalis (EfsFtsZ) E. faecium (EfmFtsZ), S. pyogenes (SpyFtsZ), S. agalactiae (SagFtsZ), and S. pneumoniae (SpnFtsZ). Fluorescence anisotropy studies conducted at 15, 25, and 37 °C reveal that addition of each of the five target FtsZ proteins increases the anisotropy of BOFP significantly (Fig. 3a–e), indicative of a binding interaction between the probe and each of the host proteins. Significantly, no such binding interactions were observable with BODIPY FL-COOH (as exemplified by the SpyFtsZ results shown in Fig. S1). Thus, BOFP can target not only SaFtsZ, but also the FtsZ proteins from a broad range of other clinically important Gram-positive pathogens.
Fluorescence anisotropy profiles of 0.1 µM BOFP as a function of increasing concentrations of EfsFtsZ (a), EfmFtsZ (b), SpyFtsZ (c), SagFtsZ (d), or SpnFtsZ (e). Acquisition and display parameters are as described in the legend to Fig. 2b. Panel (f) shows plots of ln(1/Kd) vs. 1/T for the interaction of BOFP with SaFtsZ (filled circles), EfsFtsZ (filled triangles), EfmFtsZ (open triangles), SpyFtsZ (open diamonds), SagFtsZ (filled diamonds), and SpnFtsZ (open squares). The solid lines reflect linear fits of the experimental data points with Eq. 3.
Analysis of the anisotropy isotherms in Fig. 3 with Eq. 1 yielded outstanding fits of the experimental data points (as depicted by the solid curves), with the Kd values derived from these fits being listed in Table 1. Kd values for BOFP binding to the Gram-positive FtsZ proteins ranged from 1.72 ± 0.06 µM at 15 °C to 4.62 ± 0.12 µM at 37 °C for EfsFtsZ, 2.50 ± 0.14 µM at 15 °C to 3.14 ± 0.22 µM at 37 °C for EfmFtsZ, 0.91 ± 0.06 µM at 15 °C to 1.55 ± 0.08 µM at 37 °C for SpyFtsZ, 0.62 ± 0.05 µM at 15 °C to 1.31 ± 0.04 µM at 37 °C for SagFtsZ, and 3.02 ± 0.30 µM at 15 °C to 3.81 ± 0.69 µM at 37 °C for SpnFtsZ. These Kd ranges are similar in magnitude to that observed for SaFtsZ. Recall that Artola et al. demonstrated the binding of their early-generation fluorescent FtsZ inhibitors only to SaFtsZ and BsFtsZ, while reporting Kd values in the range of 11 to 29 µM for BsFtsZ at 25 °C27. Significantly, the lower range of Kd values we observe for the binding of BOFP to all six Gram-positive FtsZ proteins at 25 °C (1.02 ± 0.09 to 3.49 ± 0.25 µM) indicates a much broader spectrum of FtsZ targeting as well as a binding affinity that is approximately 3- to 29-fold higher.
Brief exposure to a low concentration of BOFP effectively labels FtsZ in live S. aureus, E. faecalis, E. faecium, S. pyogenes, S. agalactiae, and S. pneumoniae cells
Having demonstrated the high-affinity binding of BOFP to the Gram-positive FtsZ proteins SaFtsZ, EfsFtsZ, EfmFtsZ, SpyFtsZ, SagFtsZ, and SpnFtsZ, we explored the potential of the probe to label FtsZ in live cells of the corresponding pathogens themselves. Figure 4 shows differential interference contrast (DIC) and fluorescence micrographs of S. aureus, E. faecalis, E. faecium, S. pyogenes, S. agalactiae, and S. pneumoniae cells labeled for 5 minutes with 1 µg/mL (1.3 µM) BOFP. In each of the six pathogens, bright bands of fluorescence staining are clearly visible at midcell (as highlighted by the arrows in Fig. 4b,d,f,h,j,l), consistent with the labeling of FtsZ Z-rings. Additional fluorescence staining (though weaker than that at midcell) is also evident along the periphery of each cell, suggesting that FtsZ is also localized throughout the cell membrane. These results indicate that brief exposure to a low concentration of BOFP affords outstanding visualization of FtsZ and its localization patterns in live Gram-positive bacterial cells.
FtsZ visualization in the Gram-positive bacterial pathogens S. aureus NRS705 (a,b), E. faecalis ATCC 29212 (c,d), E. faecium ATCC 19434 (e,f), S. pyogenes ATCC 19615 (g,h), S. agalactiae ATCC 12386 (i,j), and S. pneumoniae ATCC 49619 (k,l). Differential interference contrast (DIC) and fluorescence micrographs of the indicated bacterial cells treated for 5 minutes with 1 µg/mL BOFP just prior to visualization. The arrows in panels (b,d,f,h,j,l) highlight representative FtsZ Z-rings at midcell labeled by BOFP.
To ensure that the ester linkage connecting the BODIPY fluorophore in the BOFP conjugate was not being hydrolyzed by the bacterial cells, we examined the potential, if any, of BODIPY FL-COOH to label FtsZ in S. aureus cells. Significantly, no fluorescence staining of the S. aureus cells was detectable upon treatment with 1 µg/mL BODIPY FL-COOH for 5 minutes (Fig. S4). This behavior markedly contrasts the clear staining patterns observed when the cells are treated similarly with BOFP (Fig. S4). These results confirm that BOFP is not hydrolyzed by the cells during the treatment regimen and that the staining patterns observed were not the result of nonspecific interactions with a fluorescent product of hydrolysis.
BOFP targets FtsZ proteins from Gram-negative bacterial pathogens with an even higher affinity than FtsZ proteins from Gram-positive pathogens
In addition to targeting Gram-positive FtsZ proteins, we sought to determine whether BOFP could also target Gram-negative FtsZ proteins. To this end, we used fluorescence anisotropy to explore the interactions of BOFP with the FtsZ proteins from the four Gram-negative pathogens, E. coli (EcFtsZ), K. pneumoniae (KpFtsZ), P. aeruginosa (PaFtsZ), and A. baumannii (AbFtsZ), with the resulting anisotropy profiles acquired at 15, 25, and 37 °C being depicted in Fig. 5a–d. Inspection and analysis of these anisotropy profiles reveals that BOFP binds to all four Gram-negative FtsZ proteins with sub-micromolar affinity. At 25 °C, the Kd values for EcFtsZ, KpFtsZ, PaFtsZ, and AbFtsZ are 0.28 ± 0.02, 0.58 ± 0.04, 0.36 ± 0.06, and 0.55 ± 0.02 µM, respectively (Table 1). A comparison of the Kd values at 25 °C listed in Table 1 indicates that BOFP binds the Gram-negative FtsZ proteins with an approximately 2- to 12-fold higher affinity than the Gram-positive FtsZ proteins. Thus, in striking contrast to the early-generation fluorescent FtsZ inhibitors reported by Artola et al.27, BOFP can target a broad range of Gram-negative FtsZ proteins with a high degree of affinity. As observed with the Gram-positive FtsZ proteins, no binding interactions with BODIPY FL-COOH were detectable with the Gram-negative FtsZ proteins (as exemplified by the KpFtsZ and PaFtsZ results shown in Fig. S1).
Fluorescence anisotropy profiles of 0.1 µM BOFP as a function of increasing concentrations of EcFtsZ (a), KpFtsZ (b), PaFtsZ (c), or AbFtsZ (d). Acquisition and display parameters are as described in the legend to Fig. 2b. Panel (e) shows plots of ln(1/Kd) vs. 1/T for the interaction of BOFP with EcFtsZ (open circles), KpFtsZ (open squares), PaFtsZ (filled diamonds), and AbFtsZ (filled circles). The solid lines reflect linear fits of the experimental data points with Eq. 3.
For the majority of target FtsZ proteins studied, enthalpy provides a significant driving force for the binding of BOFP
We used the temperature dependence of the Kd values for the binding of BOFP to the Gram-positive and Gram-negative FtsZ proteins to derive the thermodynamic parameters associated with the binding reactions. Free energy changes (ΔG) at 37 °C (310 K) were derived from the corresponding Kd values using Eq. 2, while enthalpy and entropy changes (ΔH and ΔS, respectively) were determined from linear fits of the ln(1/Kd) vs. 1/T plots shown in Figs. 3f and 5e with Eq. 3. The resulting thermodynamic parameters are listed in Table 1. For seven of the ten FtsZ proteins studied (SaFtsZ, EfsFtsZ, SpyFtsZ, SagFtsZ, EcFtsZ, KpFtsZ, and PaFtsZ), ΔH contributes >50% to the observed ΔG of binding, with the enthalpic contribution to binding being 100% for two of those seven FtsZ proteins (SaFtsZ and EfsFtsZ). As suggested by our crystal structure of BOFP in complex with SaFtsZ, these favorable enthalpic contributions to binding likely stem from the extensive array of favorable van der Waals contacts between the host protein and both the 1 and BODIPY portions of the probe (Fig. 2c–e). For the remaining three FtsZ proteins (EfmFtsZ, SpnFtsZ, and AbFtsZ), ΔS contributes >50% to the observed ΔG of binding. These favorable entropic contributions to binding may reflect favorable binding-induced changes in hydration and/or conformation of the host proteins.
In addition to Gram-positive bacterial cells, BOFP also labels FtsZ effectively in live Gram-negative cells of E. coli, K. pneumoniae, P. aeruginosa, and A. baumannii
Armed with knowledge that BOFP binds to the Gram-negative FtsZ proteins EcFtsZ, KpFtsZ, PaFtsZ, and AbFtsZ with an even higher affinity than any of the Gram-positive FtsZ proteins, we further explored the potential of the probe to label FtsZ in live Gram-negative bacterial cells. When exposed to BOFP in a similar manner (1 µg/mL for 5 minutes) to that described above for the Gram-positive bacteria, little or no FtsZ labeling in the Gram-negative cells was observable by fluorescence microscopy (as exemplified by the K. pneumoniae results shown in Fig. S5b). Previous studies have indicated that the large size of fluorescent antibiotics resulting from the conjugation of bulky fluorophores restricts the passage of the agents across the outer membrane of Gram-negative cells29. Stokes et al. have shown that pentamidine can effectively permeabilize the outer membrane of Gram-negative bacterial cells to large antibiotics that would normally be unable to cross the membrane30. When co-treating K. pneumoniae cells with 1 µg/mL BOFP and 3.5 mg/mL pentamidine isethionate for 5 minutes, bright fluorescence staining becomes visible both at midcell and along the cell periphery (compare Fig. S5b,d). Co-treatment of E. coli, P. aeruginosa, and A. baumannii cells results in a similar fluorescence staining pattern to that observed in K. pneumoniae cells (Fig. 6b,d,f,h). The fluorescent bands visible at midcell are consistent with the labeling of FtsZ Z-rings (highlighted by the arrows in Figs. S5d and 6b,d,f,h), with the peripheral staining reflecting the presence of FtsZ in the cell membrane as well. Note that no such FtsZ staining was detectable in K. pneumoniae cells co-treated for 5 minutes with 1 µg/mL BODIPY FL-COOH and 3.5 mg/mL pentamidine isethionate (Fig. S6), thus confirming that BOFP is not being hydrolyzed by the Gram-negative cells during the labeling procedure. Viewed as a whole, our results indicate that BOFP is useful for visualizing FtsZ not only in live Gram-positive cells, but also in live Gram-negative cells.
FtsZ visualization in the Gram-negative bacterial pathogens E. coli ATCC 25922 (a,b), K. pneumoniae ATCC 13883 (c,d), P. aeruginosa ATCC 27853 (e,f), and A. baumannii ATCC 19606 (g,h). Differential interference contrast (DIC) and fluorescence micrographs of the indicated bacterial cells treated for 5 minutes with 1 µg/mL BOFP in the presence of pentamidine isethionate (at 0.875 mg/mL for E. coli and 3.5 mg/mL for the other three strains) just prior to visualization. The arrows in panels (b,d,f,h) highlight representative FtsZ Z-rings at midcell labeled by BOFP.
BOFP can also be used to visualize the impact of non-fluorescent FtsZ inhibitors on the localization of FtsZ in both Gram-positive and Gram-negative bacterial cells
For BOFP to be useful as a tool for identifying new FtsZ inhibitors in a live cell-based assay, it must facilitate the detection of changes in FtsZ localization induced by non-fluorescent test compounds with the potential for FtsZ inhibition. Toward this end, we tested the ability of BOFP to visualize the impact of the oxazole-benzamide FtsZ inhibitor 1 on FtsZ localization in live S. aureus, E. coli, and K. pneumoniae cells, with the results being shown in Fig. 7. As expected with a known FtsZ inhibitor, treatment with 1 (at 4x MIC for 3 hours) induces a significant change in cell morphology consistent with the impairment of cell division (compare Fig. 7a,c,e with Fig. 7g,i,k). This morphological change takes the form of cell enlargement in cocci like S. aureus and filamentation in rods like E. coli and K. pneumoniae. Significantly, BOFP effectively labels FtsZ in the cells treated with 1, showing clear mislocalization of FtsZ and an absence of Z-rings in any of the treated cells (Fig. 7h,j,l). In addition, the presence of FtsZ in the membranes of cells treated with 1 is reduced relative to that in the membranes of untreated cells. In S. aureus, we observed similar results for cells treated with the benzamide FtsZ inhibitors PC190723 and TXA707 (Fig. S7).
Visualization of the impact of treatment with 1 on FtsZ localization in S. aureus NRS705, E. coli N43, and K. pneumoniae ATCC 10031. Differential interference contrast (DIC) and fluorescence micrographs of the indicated bacterial cells treated for 3 hours with either DMSO vehicle (a–f) or 1 (g–l) at 4× MIC (1 µg/mL for S. aureus and 4 µg/mL for E. coli and K. pneumoniae). Just prior to visualization, cells were labeled for 5 minutes with 1 µg/mL BOFP in the absence (for S. aureus) or presence of pentamidine isethionate (at 0.875 mg/mL for E. coli and 3.5 mg/mL for K. pneumoniae). The arrows in panels (b,d,f) highlight representative FtsZ Z-rings at midcell labeled by BOFP.
We sought to verify that the BOFP staining observed in S. aureus cells treated with non-fluorescent FtsZ inhibitors still reflected specific interaction with FtsZ and not off-target interactions. To this end, we used BOFP to monitor the impact of treatment with 1 on FtsZ localization in live MRSA LAC cells that expresses a FtsZ-mCherry fusion protein (MRSA LAC FtsZ-mCherry). As shown in Fig. 8, the green fluorescence associated with BOFP labeling colocalizes with the red fluorescence associated with the induced expression of FtsZ-mCherry in both vehicle-treated and 1-treated cells. Thus, the observed BOFP labeling reflects specific interactions with FtsZ even in cells treated with non-fluorescent FtsZ inhibitors.
Comparison of BOFP and induced expression of the FtsZ-mCherry fusion protein for visualization of the impact of treatment with 1 on FtsZ localization in the MRSA LAC FtsZ-mCherry strain. Differential interference contrast (DIC) and fluorescence micrographs of the bacterial cells treated for 3 hours with either DMSO vehicle (a–d) or 1 (e–h) at 4× MIC (0.25 µg/mL). Just prior to visualization, cells were labeled for 5 minutes with 1 µg/mL BOFP. The arrows in panel panels (b,c,d) highlight representative FtsZ Z-rings at midcell as visualized either by induced expression of FtsZ-mCherry or by labeling with BOFP.
Our collective results highlight the usefulness of BOFP as a screening tool for identifying novel FtsZ inhibitors in both Gram-positive and Gram-negative pathogens. The fluorescent FtsZ inhibitors previously reported by Artola et al. lacked this utility, as their ability to label FtsZ was significantly diminished in live cells treated with known FtsZ inhibitors like PC19072327.
Conclusions
This work describes the structure-guided design of a fluorescent FtsZ-targeting probe (BOFP) consisting of an oxazole-benzamide FtsZ inhibitor conjugated to a BODIPY fluorophore. Crystallographic studies demonstrate that BOFP targets the same FtsZ site as inhibitors like PC190723 and TXA707, while also highlighting favorable contributions from the conjugated BODIPY moiety itself to the FtsZ binding reaction. Fluorescence anisotropy studies with BOFP provide the first demonstration of a fluorescent probe capable of targeting the FtsZ proteins from a broad range of clinically relevant Gram-positive and Gram-negative bacterial pathogens with a high degree of affinity. In addition, fluorescence microscopy studies highlight the utility of BOFP for visualizing FtsZ in these pathogenic bacteria as well as for visualizing the impact of non-fluorescent inhibitors on FtsZ localization. These properties make BOFP a robust tool for identifying new broad-spectrum FtsZ inhibitors and understanding their mechanisms of action.
Materials and Methods
Bacterial strains and other reagents
S. aureus NRS705 (a USA100 MRSA strain isolated from the blood of a 14-day-old male in New York) was provided by the Network on Antimicrobial Resistance in Staphylococcus aureus (NARSA) for distribution by BEI Resources, NIAID, NIH. E. coli N43 acrA− was a generous gift from Dr. Lynn Zechiedrich (Baylor College of Medicine, Houston, TX). MRSA LAC FtsZ-mCherry was generated as detailed in the Supplementary Information. All other bacterial strains were obtained from the American Type Culture Collection (ATCC). MRS, cation-adjusted Mueller-Hinton (CAMH), and tryptic soy media were from Becton-Dickinson. Todd Hewitt (TH) media was made from components specified by Becton-Dickinson. Lysed horse blood (LHB) was from Hardy Diagnostics, the sodium salt of 2′-deoxyguanosine-5′-[(α,β)-methyleno]triphosphate (GMPCPP) was from Jena Bioscience, and phosphate-buffered saline (PBS) was from Lonza. Pentamidine isethionate, isopropyl β-d-1-thiogalactopyranoside (IPTG), and PC190723 were from Sigma. 3-(5,5-difluoro-7,9-dimethyl-5H-5λ4,6λ4-dipyrrolo[1,2-c:2′,1′-f][1,3,2]diazaborinin-3-yl)propanoic acid (BODIPY FL-COOH) was from Lumiprobe Corp.
Compound synthesis
1, 2, 3, and TXA707, were synthesized as previously described11,12,13,31. BOFP was synthesized as detailed in the Supplementary Information.
Crystallization and structure determination of SaFtsZ in complex with 2 and BOFP
SaFtsZ12–316 was cloned, expressed, and purified as described previously21. After SaFtsZ12–316 was crystallized by the sitting-drop vapor-diffusion technique at 20 °C in reservoir conditions based on the JBScreen pentaerythritol 2 (Jena Bioscience), racemic mixtures of the R and S enantiomeric forms of 2 or BOFP were introduced by soaking. For the complex of 2 with SaFtsZ, the protein was crystallized at 9.5 mg/mL under conditions of 100 mM Tris-HCl (pH 7.8), 45% (w/v) pentaerythritol propoxylate 629 (PEP629), and 300 mM KCl. After 3 weeks, the crystal was soaked in the same reservoir supplemented with 5 mM 2 for 3 days. For the BOFP complex, the protein was crystallized at 20 mg/mL under conditions of 100 mM HEPES (pH 8.0) and 39% (w/v) PEP629. After 7 days, the crystal was soaked in the same reservoir supplemented with 1.4 mM BOFP for 3 days.
The crystal of the 2-SaFtsZ complex was flash-frozen in a nitrogen gas stream at −180 °C without cryoprotectants. The crystal of the BOFP-SaFtsZ complex was briefly soaked in a cryoprotectant solution containing 100 mM HEPES (pH 8.0), 39% (w/v) PEP629, and 25% (v/v) glycerol and then flash-frozen in the same manner as described above. X-ray diffraction data from the crystals of the 2-SaFtsZ and BOFP-SaFtsZ complexes were collected at SPring-8 BL44XU (Hyogo, Japan) under a cryogenic nitrogen gas stream at 100 K. Diffraction data were processed and scaled with HKL200032 and XDS33, respectively. The phases for both complexes were determined by molecular replacement with Phaser in the CCP4 suite34 using the previously determined structure of the SaFtsZ-GDP complex (PDB entry: 3VOA)35 as a search model. Both models were refined with REFMAC536 and PHENIX37, with manual modification using Coot38. The refined structures were validated with MolProbity39. Data collection and refinement statistics are summarized in Table 2. The final atomic coordinates and structure factor amplitudes have been deposited in the RCSB Protein Data Bank (PDB entries: 6KVP and 6KVQ). Figures were prepared with PyMOL (Schrödinger).
Fluorescence anisotropy assays for the binding of BOFP to FtsZ proteins
Fluorescence anisotropy experiments were performed using an AVIV model ATF105 spectrofluorometer at 15, 25, or 37 °C. In these experiments, bandwidths were set to 4 nm in both the excitation and emission directions, with the excitation and emission wavelengths being set at 488 nm and 510 nm, respectively. BOFP (0.1 µM) was titrated with increasing concentrations (ranging from 0 to 12 µM) of FtsZ in 120 µL of buffer containing 50 mM Tris-HCl (pH 7.6) and 50 mM KCl. After each protein addition, the samples were equilibrated for 3 minutes, whereupon the fluorescence anisotropy was measured.
Plots of the fluorescence anisotropy (r) of BOFP as a function of FtsZ concentration (as shown in Figs. 2, 3, and 5) were analyzed by non-linear least squares regression using the following 1:1 binding formalism:
In this equation, r0 is the anisotropy of the protein-free compound, r∞ is the anisotropy of the compound in the presence of an infinite concentration of FtsZ, [C]tot is the total concentration of the compound, and [P]tot is the total concentration of protein with each addition. These analyses yielded the equilibrium dissociation constant (Kd) for each binding reaction.
The binding free energy (ΔG) at temperature T was derived from the corresponding Kd value determined at T using the following relationship:
The binding enthalpy (ΔH) and entropy (ΔS) were derived from linear fits of the ln(1/Kd) vs. 1/T plots shown in Figs. 3f and 5e using the following relationship:
The impact of guanosine nucleotide and magnesium on the binding of BOFP to SaFtsZ was assessed at 37 °C in the same buffer described above. In these studies, the anisotropy of 0.1 µM BOFP alone or in the presence of 10 µM SaFtsZ was measured, with the latter also being measured in the presence of 0.1 mM GMPCPP (a non-hydrolyzable GTP analog), 10 mM MgCl2, or both.
In comparative control experiments, the anisotropy of 0.1 µM BODIPY FL-COOH or BOFP alone or in the presence of SaFtsZ (10 µM), SpyFtsZ (10 µM), KpFtsZ (2 µM), or PaFtsZ (2 µM) was measured at 37 °C in the same buffer described above.
A quartz ultra-micro cell (Hellma) with a 2 × 5 mm aperture and a 15 mm center height was used for all measurements. The pathlengths in the excitation and emissions directions were 1 and 0.2 cm, respectively. All steady-state anisotropy experiments were conducted in at least triplicate, with the reported anisotropies reflecting the average values.
Minimum inhibitory concentration (MIC) Assays
MIC assays of 1, 3, and BOFP were conducted by standard broth microdilution in TH media. Briefly, log-phase S. aureus NRS705 (MRSA) cells were added to 96-well microtiter plates (at 5 × 105 CFU/mL) containing 2-fold serial dilutions of each test compound in 0.1 mL of TH broth, with each compound concentration being present in duplicate. The MIC is defined as the lowest compound concentration at which growth is ≥90% inhibited after 18-24 hours of aerobic growth.
Differential interference contrast and fluorescence microscopy
All differential interference contrast (DIC) and fluorescence microscopy experiments were conducted using an Olympus BX50 microscope equipped with an X-cite Exacte 200 W mercury lamp, a 100× Olympus UPLSAPO oil immersion objective (1.40 aperture), as well as both Chroma ET-EGFP (FITC/Cy2) and ET mCherry, Texas Red filters. Images were captured using a QImaging Retiga R3 charge-coupled device (CCD) camera and the Ocular-Version 2.0 software package (QImaging).
For visualizing FtsZ in Gram-positive bacteria using BOFP, the bacterial cells were grown to log-phase in media suitable for each individual pathogen. Specifically, S. aureus NRS705 (MRSA) was grown in tryptic soy broth (TSB), E. faecalis ATCC 29212 and E. faecium ATCC 19434 were grown in lactobacilli MRS broth, S. agalactiae ATCC 12386 and S. pneumoniae ATCC 49619 were grown in TH broth, and S. pyogenes ATCC 19615 was grown in CAMH broth supplemented with 3% (v/v) LHB. For each Gram-positive bacterial strain, a total of 1 mL of cell culture was centrifuged at 15,000 × g for 1 minute and washed 2–3 times with 1 mL of PBS. After the final wash, the pelleted cells were resuspended in 500 µL of PBS containing 1 µg/mL of BOFP and incubated in the dark for 5 minutes at room temperature. The cells were then centrifuged at 15,000 × g for 1 minute, washed twice with 1 mL of PBS, and subsequently resuspended in 200 µL of PBS. 8 µL of this final cell suspension was then spread on a 0.25 mm layer of 1.5% high-resolution agarose (Sigma) in PBS, which was mounted on a standard 75 × 25 × 1 mm microscope slide (Azer Scientific) using a 1.7 × 2.8 × 0.025 cm Gene Frame (ThermoFisher). A 24 × 40 mm cover slip (Azer Scientific) was then applied to the agarose pad to prepare the slide for microscopic visualization. Comparative control experiments with 1 µg/mL BODIPY FL-COOH were conducted in S. aureus NRS705 cells as described above for BOFP.
For visualizing FtsZ in Gram-negative bacteria using BOFP, E. coli ATCC 25922, K. pneumoniae ATCC 13883, P. aeruginosa ATCC 27853, and A. baumannii ATCC 19606 were grown to log-phase in CAMH broth. For each Gram-negative bacterial strain, a total of 1 mL of cell culture was centrifuged at 15,000 × g for 1 minute and washed twice with 1 mL of Tris-buffered saline (TBS) composed of 50 mM Tris-HCl (pH 7.6) and 150 mM NaCl. After the final wash, the pelleted cells were resuspended in 500 µL of TBS containing 1 µg/mL of BOFP and pentamidine isethionate (at 0.875 mg/mL for E. coli and 3.5 mg/mL for the other three strains). The resuspended cells were then incubated in the dark for 5 minutes at room temperature, centrifuged at 15,000 × g for 1 minute, washed twice with 1 mL of TBS, and subsequently resuspended in 200 µL of TBS. This final cell suspension was then prepared for microscopy as described for the Gram-positive bacterial strains. Comparative control experiments with 1 µg/mL BODIPY FL-COOH were conducted in K. pneumoniae ATCC 13883 cells as described above for BOFP.
To visualize the impact of treatment with the FtsZ inhibitor 1 using BOFP, S. aureus NRS705 (MRSA), E. coli N43, and K. pneumoniae ATCC 10031 were grown to log-phase in CAMH broth and diluted to an OD600 of 0.1. Each cell culture was then treated with either DMSO vehicle or 1 at 4× MIC (1 µg/mL for S. aureus or 4 µg/mL for E. coli and K. pneumoniae) for 3 hours at 37 °C. Following this treatment, 1–5 mL of each culture was centrifuged at 15,000 × g for 1 minute and washed twice with 1 mL of PBS (for S. aureus) or TBS (for E. coli and K. pneumoniae). The resulting S. aureus cell pellets were further processed and labeled with BOFP as described above for the Gram-positive bacterial strains and the resulting E. coli and K. pneumoniae cell pellets were further processed and labeled with BOFP as described above for the Gram-negative bacterial strains. The impact of treating S. aureus NRS705 cells with the FtsZ inhibitors PC190723 and TXA707 at 4× MIC (2 µg/mL for PC190723 and 4 µg/mL for TXA707) was also examined using BOFP as described above.
To visualize the impact of treating MRSA LAC FtsZ-mCherry with 1, cells were grown to log-phase in CAMH broth and diluted to an OD600 of 0.1. The cells were then treated with 10 µM IPTG and either DMSO vehicle or 1 at 4× MIC (0.25 µg/mL) for 3 hours at 37 °C. Following this treatment, 1–5 mL of each culture was centrifuged at 15,000 × g for 1 minute and washed twice with 1 mL of PBS. The resulting cell pellets were further processed and labeled with BOFP as described above for the Gram-positive bacterial strains.
References
Ventola, C. L. The Antibiotic Resistance Crisis Part 1: Causes and Threats. P. T. 40, 277–283 (2015).
World Health Organization. Antimicrobial resistance: global report on surveillance. (World Health Organization, 2014).
World Health Organization. Antibacterial Agents in Clinical Development: An Analysis of the Antibacterial Clinical Development Pipeline, Including Tuberculosis. 48 (Geneva, Switzerland, 2017).
Lock, R. L. & Harry, E. J. Cell-division Inhibitors: New Insights for Future Antibiotics. Nat. Rev. Drug. Discov. 7, 324–338 (2008).
Sass, P. & Brotz-Oesterhelt, H. Bacterial Cell Division as a Target for New Antibiotics. Curr. Opin. Microbiol. 16, 522–530 (2013).
Kusuma, K. D., Payne, M., Ung, A. T., Bottomley, A. L. & Harry, E. J. FtsZ as an Antibacterial Target: Status and Guidelines for Progressing This Avenue. ACS Infect. Dis. 5, 1279–1294 (2019).
Haydon, D. J. et al. Creating an Antibacterial with in Vivo Efficacy: Synthesis and Characterization of Potent Inhibitors of the Bacterial Cell Division Protein FtsZ with Improved Pharmaceutical Properties. J. Med. Chem. 53, 3927–3936 (2010).
Haydon, D. J. et al. An Inhibitor of FtsZ with Potent and Selective Anti-Staphylococcal Activity. Science 321, 1673–1675 (2008).
Kaul, M. et al. An FtsZ-Targeting Prodrug with Oral Antistaphylococcal Efficacy In Vivo. Antimicrob. Agents Chemother. 57, 5860–5869 (2013).
Kaul, M. et al. Pharmacokinetics and In Vivo Antistaphylococcal Efficacy of TXY541, a 1-methylpiperidine-4-carboxamide Prodrug of PC190723. Biochem. Pharmacol. 86, 1699–1707 (2013).
Kaul, M. et al. TXA709, an FtsZ-Targeting Benzamide Prodrug with Improved Pharmacokinetics and Enhanced In Vivo Efficacy against Methicillin-Resistant Staphylococcus aureus. Antimicrob. Agents Chemother. 59, 4845–4855 (2015).
Stokes, N. R. et al. An Improved Small-Molecule Inhibitor of FtsZ with Superior in Vitro Potency, Drug-Like Properties, and in Vivo Efficacy. Antimicrob. Agents Chemother. 57, 317–325 (2013).
Stokes, N. R. et al. Design, Synthesis and Structure-Activity Relationships of Substituted Oxazole-Benzamide Antibacterial Inhibitors of FtsZ. Bioorg. Med. Chem. Lett. 24, 353–359 (2014).
Tan, C. M. et al. Restoring Methicillin-Resistant Staphylococcus aureus Susceptibility to β-lactam Antibiotics. Sci. Transl. Med. 4, 126ra135 (2012).
Beall, B. & Lutkenhaus, J. FtsZ in Bacillus subtilis is Required for Vegetative Septation and for Asymmetric Septation During Sporulation. Genes. Dev. 5, 447–455 (1991).
Dai, K. & Lutkenhaus, J. ftsZ is an Essential Cell Division Gene in. Escherichia coli. J. Bacteriol. 173, 3500–3506 (1991).
Pinho, M. G. & Errington, J. Dispersed Mode of Staphylococcus aureus Cell Wall Synthesis in the Absence of the Division Machinery. Mol. Microbiol. 50, 871–881 (2003).
Stratton, C. W. Dead Bugs Don’t Mutate: Susceptibility Issues in the Emergence of Bacterial Resistance. Emerg. Infect. Dis. 9, 10–16 (2003).
Tripathy, S. & Sahu, S. K. FtsZ Inhibitors as a New Genera of Antibacterial Agents. Bioorg. Chem. 91, 103169 (2019).
Hurley, K. A. et al. Targeting the Bacterial Division Protein FtsZ. J. Med. Chem. 59, 6975–6998 (2016).
Fujita, J. et al. Structural Flexibility of an Inhibitor Overcomes Drug Resistance Mutations in Staphylococcus aureus FtsZ. ACS Chem. Biol. 12, 1947–1955 (2017).
Kaul, M., Mark, L., Parhi, A. K., LaVoie, E. J. & Pilch, D. S. Combining the FtsZ-Targeting Prodrug TXA709 and the Cephalosporin Cefdinir Confers Synergy and Reduces the Frequency of Resistance in Methicillin-Resistant Staphylococcus aureus. Antimicrob. Agents Chemother. 60, 4290–4296 (2016).
Margalit, D. N. et al. Targeting Cell Division: Small-Molecule Inhibitors of FtsZ GTPase Perturb Cytokinetic Ring Assembly and Induce Bacterial Lethality. Proc. Natl. Acad. Sci. USA 101, 11821–11826 (2004).
Knudson, S. E. et al. A Trisubstituted Benzimidazole Cell Division Inhibitor with Efficacy Against Mycobacterium tuberculosis. PLoS One 9, e93953 (2014).
Knudson, S. E. et al. Cell Division Inhibitors With Efficacy Equivalent to Isoniazid in the Acute Murine Mycobacterium tuberculosis Infection Model. J. Antimicrob. Chemother. 70, 3070–3073 (2015).
Stone, M. R. L., Butler, M. S., Phetsang, W., Cooper, M. A. & Blaskovich, M. A. T. Fluorescent Antibiotics: New Research Tools to Fight Antibiotic Resistance. Trends Biotechnol. 36, 523–536 (2018).
Artola, M. et al. The Structural Assembly Switch of Cell Division Protein FtsZ Probed with Fluorescent Allosteric Inhibitors. Chem. Sci. 8, 1525–1534 (2017).
Rivas, G. et al. Magnesium-induced Linear Self-association of the FtsZ Bacterial Cell Division Protein Monomer. The Primary Steps for FtsZ Assembly. J. Biol. Chem. 275, 11740–11749 (2000).
Matijašić, M. et al. Fluorescently Labeled Macrolides as a Tool for Monitoring Cellular and Tissue Distribution of Azithromycin. Pharmacol. Res. 66, 332–342 (2012).
Stokes, J. M. et al. Pentamidine Sensitizes Gram-negative Pathogens to Antibiotics and Overcomes Acquired colistin Resistance. Nat. Microbiol. 2, 17028 (2017).
Haydon, D. J. et al. Aromatic Amides and Uses Thereof. US Patent 9,511,073 B2 (2016).
Otwinowski, Z. & Minor, W. Processing of X-Ray Diffraction Data Collected in Oscillation Mode. Methods Enzymol. 276, 307–326 (1997).
Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
McCoy, A. J. et al. Phaser Crystallographic Software. J. Appl. Crystallogr. 40, 658–674 (2007).
Matsui, T. et al. Structural Reorganization of the Bacterial Cell-Division Protein FtsZ from Staphylococcus aureus. Acta Crystallogr. D Biol. Crystallogr. D68, 1175–1188 (2012).
Murshudov, G. N. et al. REFMAC5 for the Refinement of Macromolecular Crystal Structures. Acta Crystallogr. D Biol. Crystallogr. 67, 355–367 (2011).
Adams, P. D. et al. PHENIX: A Comprehensive Python-Based System for Macromolecular Structure Solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Emsley, P. & Cowtan, K. Coot: Model-Building Tools for Molecular Graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Chen, V. B. et al. MolProbity: All-Atom Structure Validation for Macromolecular Crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Acknowledgements
We are indebted to Dr. Lynn Zechiedrich for providing the E. coli N43 strain and to Mr. Jeffrey Yang for his assistance with the 13C NMR characterizations. This work was supported by National Institutes of Health grant RO1 AI118874 (to D.S.P.); Grant-in-Aid for the Japan Society for the Promotion of Science (JSPS) Fellows (15J00589 to J.F.); Grants-in-Aid for Scientific Research (19K16060 to T.Y.; 17H05732, 18K06094, 19H04735, and 19K07582 to H.M.); the Takeda Science Foundation; the Science Research Promotion Fund from the Promotion and Mutual Aid Corporation for Private Schools of Japan; the Cooperative Research Program of the Institute for Protein Research, Osaka University (CR-18-05 and CR-19-05); and the Platform Project for Supporting Drug Discovery and Life Science Research (Basis for Supporting Innovative Drug Discovery and Life Science Research (BINDS)) from AMED under Grant Number JP19am0101070. The crystallographic work was performed under the approval of the Photon Factory and SPring-8 Program Advisory Committee (Proposal Nos. 2017A6748, 2017A2570, 2017B6748, 2018A2719, 2018A6859 and 2017G702).
Author information
Authors and Affiliations
Contributions
E.F.-G. contributed to the conceptualization and performance of experiments, interpretation of results, and writing of the paper. J.F. and T.Y. contributed to the performance of experiments, interpretation of results, and writing of the paper. J.M.N. contributed to the performance of experiments, interpretation of results, and writing of the paper. A.J.P. contributed to the conceptualization and performance of experiments and interpretation of results. E.H., M.O., N.K., and H.M.A.-T. contributed to the performance of experiments and interpretation of results. J.M.B. and E.J.L. contributed to the conceptualization of experiments, interpretation of results, and writing of the paper. H.M. contributed to the conceptualization of experiments, interpretation of results, writing and editing of the paper, and funding acquisition. D.S.P. designed BOFP and contributed to the conceptualization of experiments, interpretation of results, writing and editing of the paper, and funding acquisition.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ferrer-González, E., Fujita, J., Yoshizawa, T. et al. Structure-Guided Design of a Fluorescent Probe for the Visualization of FtsZ in Clinically Important Gram-Positive and Gram-Negative Bacterial Pathogens. Sci Rep 9, 20092 (2019). https://doi.org/10.1038/s41598-019-56557-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-56557-x
This article is cited by
-
Molecular dynamics simulations reveal differences in the conformational stability of FtsZs derived from Staphylococcus aureus and Bacillus subtilis
Scientific Reports (2024)
-
An in silico approach for identification of lead compound as FtsZ inhibitor
Molecular Diversity (2024)
-
Evaluation of 2,6-difluoro-3-(oxazol-2-ylmethoxy)benzamide chemotypes as Gram-negative FtsZ inhibitors
The Journal of Antibiotics (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.