Abstract
Alveolar echinococcosis (AE) is a zoonotic disease that is deadly if left untreated. AE is caused by the larval metacestode stage of the cestode Echinococcus multilocularis. Better knowledge on the host-parasite interface could yield novel targets for improvement of the treatment against AE. We analyzed culture media incubated with in vitro grown E. multilocularis metacestodes by 1H nuclear magnetic resonance spectroscopy to identify the unknown metabolic footprint of the parasite. Moreover, we quantitatively analyzed all amino acids, acetate, glucose, lactate, and succinate in time-course experiments using liquid chromatography and enzymatic assays. The E. multilocularis metacestodes consumed glucose and, surprisingly, threonine and produced succinate, acetate, and alanine as major fermentation products. The metabolic composition of vesicle fluid (VF) from in vitro grown E. multilocularis metacestodes was different from parasite-incubated culture medium with respect to the abundance, but not the spectrum, of metabolites, and some metabolites, in particular amino acids, accumulated in the VF. Overall, this study presents the first characterization of the in vitro metabolic footprint of E. multilocularis metacestodes and VF composition, and it provides the basis for analyses of potentially targetable pathways for future drug development.
Similar content being viewed by others
Introduction
Platyhelminths (flatworms), including cestodes (tapeworms) and trematodes (flukes) cause a variety of human and animal infections. Currently, the highest ranked foodborne parasite in Europe is Echinococcus multilocularis, the small fox tapeworm1. Infection with this cestode causes the disease alveolar echinococcosis (AE) in humans. E. multilocularis is endemic to the Northern hemisphere including areas in Europe, Asia, and North America2. Over the recent years, E. multilocularis has been emerging in Europe, North America and Asia3,4,5. The life cycle of E. multilocularis includes canids as definitive hosts, and voles as intermediate hosts. Humans, and other mammals such as dogs, captive monkeys, and beavers, can be infected as accidental intermediate hosts by ingesting parasite eggs shed within the feces of definitive hosts. Thereby, these accidental hosts can acquire the disease AE. In humans, E. multilocularis grows as larval metacestodes, which primarily affect the liver and cause the disease AE, but they can also form metastases in other organs, especially at the late stage of infection6. Human AE is a chronic disease with extensive morbidity and mortality if remained untreated. Curative treatment is based on radical surgical resection in combination with temporal chemotherapy, applied in 20 to 50% of all cases6. Lifelong chemotherapeutical treatment based on benzimidazole-carbamate derivatives is applied if radical surgery is not possible. This treatment, however, may fail, and has a limited potential for complete cure. In addition, it can induce severe, life-threatening side-effects7. With increasing numbers of patients and no alternative to benzimidazoles developed so far, new and better treatment options are urgently needed8.
Morphologically, metacestodes are multivesicular larval stages surrounded by an acellular, carbohydrate-rich laminated layer8. Inside the vesicles, the parasite tissue is comprised of the cellular germinal layer, which covers the inner surface of the laminated layer. The germinal layer consists of muscle cells, nerve cells, glycogen storage cells, connective tissue, and undifferentiated stem cells9,10. The syncytial tegument, as the most outer part of the germinal layer, constitutes the interface between the germinal and the laminated layer, and it forms microtriches protruding from the germinal layer into the laminated layer11. Release of small vesicles from these microtriches has been described, which could be involved in release or uptake of metabolites12.
Metabolomics provides an excellent tool to characterize host-parasite interactions13,14,15. Metabolites such as amino acids, lipids, or sugars directly reflect the biochemical activity and the metabolic state of cells or tissues16,17. Various studies applied metabolic profiling to study diseases caused by flatworms in vivo, namely Echinostoma caproni in mice18,19, Fasciola hepatica in rats20, Onchocerca volvulus in humans21, Opisthorchis felineus in hamsters22 and humans23, and Schistosoma mansoni in mice24,25,26 and humans27. Only few studies directly assessed the metabolic composition of flatworms like S. mansoni28 and F. hepatica29. Cestodes were largely neglected in modern metabolomic studies, and no in-depth analyses have been performed to investigate the metabolic behavior of flatworms or the metabolic relationship between hosts and flatworms. Such metabolic information, however, could promote the development of therapeutic treatments, for example by blocking the nutrient uptake by the parasites. 1H Nuclear magnetic resonance (NMR) provides a, highly reproducible tool for untargeted footprinting of known and especially unknown metabolites with minimal interference due to sample preparation. Mass spectrometry (MS) is frequently used in metabolomics as well, and it is more sensitive than NMR. However, MS has a lower reproducibility and requires targeted pre-processing of samples, which narrows down the group of metabolites identified. Thus, NMR is the method of choice for untargeted metabolomic footprinting30. The metabolism of E. multilocularis is of particular interest, as genomic and transcriptomic analyses revealed that the parasite is deficient in large parts of amino acid, nucleotide and lipid synthesis, but has developed several protein families for nutrient uptake31,32,33.
E. multilocularis metacestode vesicles are filled with vesicle fluid (VF), which contains parasite and host components, and it is assumed to be involved in nutrition storage. The analogous hydatid fluid of E. granulosus, a close relative to E. multilocularis, contains pools of amino acids, sugars, lipids, fermentation end products such as acetate, alanine, lactate, and succinate, in metacestodes harvested from naturally infected intermediate hosts or human patients34,35,36,37,38,39, or from experimentally infected mice40,41. In vitro uptake of cholesterol into E. granulosus cysts has been demonstrated by the use of radiolabeled lipids42. One earlier study addressed changes in metabolite composition of E. multilocularis VF in vitro upon drug treatment, identifying four major fermentation end products, namely acetate, alanine, lactate, and succinate43. In another study, the metabolites of E. multilocularis cysts grown in treated versus non-treated animals were compared, and most pronounced differences were found in the levels of acetate, alanine, glycerolphosphatidylcholine, glycine, glycogen, and succinate44. Thus far, no metabolomic study has been performed applying the defined E. multilocularis metacestode in vitro culture system45,46, which allows for further molecular analyses of interesting pathways.
E. multilocularis protoscoleces use glucose as a major energy source, accumulate glycogen in vitro47 and express all enzymes of glycolysis, fermentation, and the tricarboxylic acid cycle48. Decreased glucose levels are detected in all organs of E. multilocularis infected jirds, and accordingly also reduced glycogen levels in the livers of these infected animals49. This further strengthens the importance of the energy source glucose also in an in vivo setting. The metabolic end products of glucose as an energy source of E. multilocularis protoscoleces are acetate, lactate, and succinate47. In contrast to protoscoleces, information on energy and intermediate metabolisms of the disease-causing stage of E. multilocularis, the metacestode, is scarce. Here, we present a study identifying the metabolites consumed and released by E. multilocularis metacestodes under anaerobic growth conditions using 1H-NMR for the identification, and liquid chromatography as well as enzymatic assays for the subsequent quantification of selected metabolites. Our study shows that metacestodes secrete acetate and succinate at the expense of glucose, and that threonine is the major amino acid consumed from the medium.
Methods
If not stated otherwise, all chemicals and materials were purchased from Sigma-Aldrich (Buchs, Switzerland). Cell culture media and fetal bovine serum were from Bioswisstec (Schaffhausen, Switzerland). Statistical analyses were performed using the software package R (V 3.4.1). Figure plotting was done with the R package ggplot2 (V 3.0.0) and figure layouts were finalized using Adobe Illustrator CC (V 22.0.1).
Experimental design
To identify metabolic changes at the interface of the E. multilocularis metacestode with its growth medium, the in vitro setup depicted in Fig. 1 was applied. Reuber rat hepatoma cells (RH) were used to precondition culture medium for subsequent E. multilocularis metacestode vesicle incubation. This preconditioned medium was incubated in vitro with E. multilocularis metacestode vesicles. The metabolites in the corresponding samples of parasite-interacted medium (vcDMEM) and control incubated medium (ccDMEM) were analyzed by NMR spectroscopy. Metacestode vesicle fluid (VF) was also analyzed. To verify the NMR results and get a quantitative time-course measurement of selected metabolites, the same in vitro interaction setup was repeated independently, and samples analyzed by enzymatic acetate, glucose, lactate, and succinate measurements, as well as by free amino acid quantification with HPLC for different timepoints.
Graphical representation of the experimental setup. The experimental setup for the metabolic foot printing of E. multilocularis metacestodes in vitro included (a) medium preconditioning with rat hepatoma cells for four days in DMEM including 0.2% FCS to enrich the medium with host cell metabolites (cDMEM). (b) incubation of cDMEM with in vitro cultured E. multilocularis metacestode vesicles (vcDMEM) or mock incubation for control (ccDMEM) over 72 h (for 1H NMR analysis) or for the time-points 0, 2, 6, 10, 24, 48, 72 h (for quantitative measurements). During this time, metacestode vesicles consumed and released metabolites (indicated by arrows). Samples of vcDMEM, ccDMEM, and vesicle fluid (VF) were harvested for subsequent metabolite identification by 1H NMR, HPLC, or enzymatic assays.
Echinococcus multilocularis metacestode culture
Echinococcus multilocularis metacestodes vesicles (isolate H95) were cultured as described before50.
Preparation of host cell conditioned medium
107 RH feeder cells were seeded in a T175 cell culture bottle in 50 ml DMEM containing 10% FBS, 100 U/ml penicillin, 100 μg/ml streptomycin, and 5 μg/ml tetracycline, and incubated for one day at 37 °C, 5% CO2, humid atmosphere. Subsequently, RH cells were washed three times with PBS and 50 ml DMEM without phenol red. For the next four days, cells were incubated in 50 ml DMEM without phenol red and 0.2% heat inactivated FBS at 37 °C in 5% CO2, humid atmosphere. This host cell conditioned medium (cDMEM) was sterile filtered and directly used for incubation with E. multilocularis metacestode vesicles.
Metabolite exchange between E. multilocularis metacestodes and host cell conditioned medium
To identify the metabolic footprint of E. multilocularis metacestodes, i.e. metabolites released or consumed by E. multilocularis metacestodes into or from the surrounding culture medium in vitro, the following setup was performed: Metacestodes were cultured for 12 weeks to reach a metacestode vesicle size of 2 to 4 mm in diameter. Metacestode vesicles were purified with 2% sucrose and PBS, and washed once in cDMEM. Subsequently, metacestode vesicles were incubated at a ratio of 1:2 in cDMEM for 72 h at 37 °C in a closed tube, resulting in the E. multilocularis metacestode vesicle interaction medium (vcDMEM). Integrity of metacestode vesicles was visually confirmed during the course of the experiment. Correspondingly, cDMEM only was incubated as control (ccDMEM) at 37 °C in a closed tube. For both media, ten biological replicates were analyzed. Medium supernatants were then stored at −80 °C for subsequent metabolite analysis. From the residual metacestodes, VF was extracted by washing the metacestodes 3 x in PBS and breaking them with a 1 ml pipette tip. Metacestodes were centrifuged at 9’000 × g for 20 min, 4 °C. The supernatant (VF) was centrifuged at 12’000 × g for 20 min, 4 °C and stored at −80 °C for subsequent metabolite analysis. VF was analyzed in five biological replicates.
NMR sample preparation and data acquisition
Medium and VF samples from the above-described interaction were further prepared and analyzed by NMR at the Imperial College, London, UK. Details on sample preparation and data acquisition are found in Supplementary Methods 1.
Metabolite identification
Metabolites were identified using Chenomx NMR Suite (V 8.2; May-01-2016 with Java 1.8.0_74 (x86)), Human Metabolite Database (HMDB)51, 2D NMR spectra and the published 2D NMR data51,52, and statistical total correlation spectroscopy (STOCSY) with the script IMPaCTS (v 1.0.0) in Matlab (V R2015b 8.6.0.267245)53.
Data processing and statistical analysis of NMR spectra
Spectra were digitized into 20,000 data points with a resolution of 0.0005 ppm and peak regions between δ1H -0.01 to 0.01 for TSP and δ1H 4.68 to 5.04 for water were removed. Peaks were aligned using recursive segment-wise peak alignment (RSPA)54 and probabilistic quotient normalization was performed using IMPaCTS. To investigate changes in the relative concentrations of metabolites between the experimental groups, principal component analysis (PCA) and orthogonal projection to latent structure-discriminant analysis (OPLS-DA) were performed using the IMPaCTS script55,56. For OPLS-DA, the PLS and orthogonal components were calculated based on the mean-centered data scaled to unit variance. A seven-fold cross validation was used, and the corresponding cross-validation parameter was expressed as Q2Y. The total explained variation of the X matrix was indicated by the goodness of the fit (R2X). Figures were plotted as vector graphics in Matlab.
Enzymatic assays for detection of acetate, glucose, lactate, and succinate
The levels of the metabolites acetate, glucose, lactate, and succinate were measured in the time-course samples (0, 2, 6, 10, 24, 48, 72 h) of vcDMEM and ccDMEM, as well as the endpoint sample VF (72 h) using the Acetate Colorimetric Assay Kit (MAK086, Sigma, USA), the Glucose-Glo TM Assay kit (J6021, Promega, USA), the Lactate-Glo TM Assay kit (J5021, Promega, USA), and the Succinic Acid Assay kit (K-SUCC, Megazyme, Germany) according to the manufacturers’ protocols. Measurements were made on an EnSpire 2300 plate reader (PerkinElmer Life Sciences, Schwerzenbach, Switzerland). Acetate, glucose, lactate, and succinate concentrations were calculated by linear regression based on the standard (0.00 to 10.00 nmol for acetate, 4.39 to 50.00 mM for glucose, 0.20 to 50.00 mM for lactate, and 0.13 µg to 4.00 µg total succinate) after subtraction of the respective enzyme blanks. Biological triplicates were analyzed for each sample and median and range (min, max) were calculated to represent the measured values. Linear reduction/accumulation rates were calculated using linear regression analysis in R and expressed as concentration changes in mM per hour.
Amino acid quantification
To analyze the free amino acid content in the time-course samples (0, 2, 6, 10, 24, 48, 72 h) of vcDMEM and ccDMEM, the endpoint samples of VF (72 h), as well as germinal layer cell extracts of metacestodes at 72 h (for purification see Supplementary Methods 2), high-performance liquid chromatography (HPLC) was performed as described in Supplementary Methods 3. Biological triplicates were analyzed for each sample. For each amino acid and timepoint, median and range were calculated to represent the measured values. For metabolites with a linear change over time, reduction/accumulation rates were calculated using linear regression analysis in R and expressed as concentration changes in mM per hour. Other metabolites were analyzed by polynomial regression analysis in R and expressed as half-maximal or half-minimal times in hours.
Amino acid frequencies in proteins of germinal layer cells (cell preparation and hydrolysis in Supplementary Methods 2) were analyzed by HPLC accordingly. The amino acid content of metacestode proteins was calculated by subtracting the free amino acids from the hydrolyzed amino acids of the germinal layer cell extracts.
To analyze the relationship between the consumption and the concentration of essential amino acids in VF, germinal layer cell extracts, and metacestode proteins, linear regression and Cook’s distance outlier analysis were performed in R. Consumption of essential amino acids by metacestodes was defined as the difference between the pool sizes in ccDMEM and vcDMEM.
Comparative statistical analyses of metabolite measurements
Integral intensities of the selected peaks from each metabolite identified by 1D 1H NMR were calculated for vcDMEM, ccDMEM and VF samples in Matlab (V R2015b 8.6.0.267245) and compared with the metabolite concentrations measured by enzymatic assays for acetate, glucose, lactate, and succinate and by free amino acid quantification using t-distribution of Person’s product-moment correlation in R. Tow sample t-test (two-tailed) was performed between metabolites of vcDMEM and VF as measured by free amino acid quantification and enzymatic assays. P values were Bonferroni adjusted with a significance level of α = 0.05.
To determine the percentage of glucose used for production of acetate, alanine, lactate, and succinate, pyruvate equivalents were calculated from the concentrations derived from enzymatic assays and free amino acid quantification. Thereby percentages were calculated by setting the glucose pyruvate equivalent concentration to 100%.
Results
The metabolic footprint of the Echinococcus multilocularis metacestode in culture medium
Culture media containing E. multilocularis metacestode vesicles and control media (see Fig. 1) were analyzed using 1D 1H NMR spectroscopy to identify the metabolic footprint of E. multilocularis metacestodes in vitro. Representative 1D 1H NMR spectra are shown in Supplementary Fig. 1a,b. Multivariate statistical analyses including PCA and OPLS-DA were performed on the 1H NMR spectral data to investigate the metabolic differences between vcDMEM and ccDMEM. Clear group separations between vcDMEM and ccDMEM were observed in the PCA scores plot along the first principal component with an explained variation (R2X) of 98.39% (Fig. 2a). OPLS-DA scores plot also showed a clear separation between the two groups with a total explained variance of R2X = 0.98 and the corresponding cross validation value Q2Y = 0.99, which indicates a high goodness of fit (R2X) and predictability (Q2Y) (Fig. 2b). Thus, substantial differences between vcDMEM and ccDMEM were detectable by 1H NMR, reflecting the parasite-induced changes in the culture media.
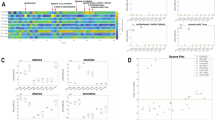
Multivariate statistical analysis of 1H NMR spectra. Multivariate comparison between vcDMEM (red, n = 10) and ccDMEM (blue, n = 10) acquired from 1D 1H NMR spectra after normalization. (a) Principal component analysis (PCA) scores plot of vcDMEM and ccDMEM. Groups are separated in the first principal component (PC1) with an explained variance of 98.39%. (b) Respective orthogonal projection to latent structure discriminant analysis (OPLS-DA) scores plot of vcDMEM and ccDMEM also clearly separated the two groups. Tcv depicts the inter-group variation, Tosc the intra-group variation. (c) Annotated PCA loadings plot of vcDMEM and ccDMEM. Annotated metabolites are indicated by various colors. (d) Annotated OPLS-DA coefficient plot of vcDMEM and ccDMEM. The x-axis depicts the chemical shifts in ppm. Peaks orientating upwards form the baseline show higher metabolite abundance in vcDMEM, from the baseline downwards pointing peaks show higher abundance in ccDMEM. The significance that each peak contributes to the difference between the two groups is indicated color-coded from blue low to red (scale on the right). The total explained variance (R2X) was 0.98 and the corresponding cross-validation (Q2Y) was 0.99. Amino acids are annotated using the three-letter code, for all other metabolites, full names are given.
Upon metabolite annotation of the 1H NMR spectra of vcDMEM and ccDMEM, a total of 21 metabolites were identified (Table 1, Supplementary Fig. 1a,b). To identify differences in the metabolite levels between vcDMEM and ccDMEM, the PCA loadings plot and the OPLS-DA coefficient loadings plot were derived from 1D 1H NMR spectra. The PCA loadings plot indicates that acetate, glucose, and succinate contribute most to the variation between vcDMEM and ccDMEM (Fig. 2c). Metabolites with a negative OPLS-DA discrimination score (Fig. 2d, Table 2), and thereby present at higher levels in ccDMEM than vcDMEM, i.e. being consumed by the parasite, were glucose, isoleucine, leucine, methionine, phenylalanine, threonine, tryptophan, tyrosine, and valine. Accordingly, several metabolites were released by the parasite with a positive OPLS-DA correlation coefficient (Fig. 2d, Table 2), exhibiting higher levels in vcDMEM than ccDMEM: acetate, acetone, alanine, aspartate, ethanol, formate, fumarate, glycine, lactate, lysine, malate, and succinate. Note that acetone, ethanol, lysine, phenylalanine, tryptophan, and tyrosine showed only a low correlation coefficient.
The amino acid threonine was found exclusively in ccDMEM, and only baseline levels were found in vcDMEM (Table 2). The amino acids alanine and aspartate, and the dicarboxylic acids fumarate, malate, and succinate were identified in vcDMEM with baseline levels in ccDMEM (Table 2). All other identified metabolites (acetate, acetone, ethanol, formate, glucose, glycine, isoleucine, lactate, leucine, lysine, methionine, phenylalanine, tryptophan, tyrosine, and valine) were found in both vcDMEM and ccDMEM (Table 2).
Quantification of selected metabolites in culture media
To confirm the 1H-NMR results and to quantify selected metabolites, concentrations of acetate, glucose, lactate, and succinate, as well as all amino acids, were determined by enzymatic assays or HPLC. Moreover, the changes in metabolite composition were analyzed in time-course experiments. Hereby, five amino acids not detectable by NMR were identified, namely arginine, glutamate, histidine, proline and serine (Fig. 3). The correlation of the concentrations determined by 1H-NMR and by quantitative methods were highly significant for both the samples of ccDMEM (Pearson: r = 0.99, p = 2.20 * 10−16) and of vcDMEM (Pearson: r = 0.91, p = 2.84 * 10−6). Amongst all quantitatively measured metabolites, glucose showed the largest concentration reduction after 72 h of culture with a stable median concentration of 24.71 mM in ccDMEM and a linear reduction to 16.73 mM in vcDMEM (linear reduction rate = 0.10 mM glucose/hour, Fig. 3). Acetate had a stable median concentration of 0.64 mM in ccDMEM. In vcDMEM acetate accumulated over the first 10 h with a linear rate of 0.23 mM acetate/hour. Between 10 and 72 h of in vitro culture, this rate dropped to 0.03 mM acetate/hour (Fig. 3). Lactate was at a concentration of 3.01 mM in ccDMEM and 4.04 mM in vcDMEM, with an increase mostly during the first 10 h (Fig. 3). Succinate showed a stable median concentration of 0.19 mM in ccDMEM that increased to 9.01 mM in vcDMEM (linear accumulation rate = 0.13 mM/hour). Alanine was stable at 0.02 mM in ccDMEM, but reached 0.71 mM in vcDMEM, plateauing after 48 h (half-maximal concentration reached after 13.46 h). Aspartate, glutamate, and glycine concentrations increased slightly over time in vcDMEM (Fig. 3). Conversely, the median concentrations of arginine, isoleucine, leucine, lysine, methionine, phenylalanine, threonine, tyrosine, and valine were decreased in vcDMEM after 72 h of incubation (Fig. 3). In particular threonine concentrations were rapidly diminished within 72 h of in vitro culture (half time = 9.23 h). The concentrations of histidine, proline, serine, and tryptophan did not change substantially between ccDMEM and vcDMEM during the incubation time (Fig. 3).
Quantitative time-course measurements of acetate, glucose, lactate, succinate, and amino acids in vcDMEM and ccDMEM. Metabolite concentrations were determined by (a) enzymatic assays (acetate, glucose, lactate, and succinate) and (b) free amino acid quantification by HPLC. Biological triplicates were assessed for each timepoint, and median concentrations and ranges are given for each metabolite for the timepoints 0, 2, 6, 10, 24, 48, and 72 h for vcDMEM and ccDMEM.
Metabolomic comparison between vesicle fluid and culture medium incubated with E. multilocularis metacestodes
In addition to the culture medium, we analyzed the pool sizes of metabolites in VF by 1H-NMR and by quantitative methods. A representative 1D 1H NMR spectrum for VF and analysis of metabolite abundance (peak intensities) is given in Supplementary Fig. 1c and Supplementary Table 1. A priori, the 1H-NMR-based metabolite profiles cannot be used for a quantitative comparison between culture media and VF, as they constitute different biofluids with different TSP values. Nevertheless, as for the culture medium, the correlation between the peak integral values obtained by 1H-NMR and by the values obtained by quantitative measurements (Fig. 4) was highly significant (Pearson: r = 0.97, p = 2.92 * 10−9). Glucose showed significantly lower concentrations in VF than in vcDMEM and threonine was not detectable at all in VF. Conversely, alanine, arginine, glycine, histidine, isoleucine, leucine, lysine, methionine, phenylalanine, serine, tryptophan, tyrosine, and valine showed higher concentrations in VF than in vcDMEM. Aspartate, glutamate, lactate, proline, and succinate did not significantly differ between vcDMEM and VF (Fig. 4). This difference in metabolite abundance and profiles between vcDMEM and VF indicates for selective metabolite transport through the metacestode wall.
Concentrations of amino acids and selected metabolites in vcDMEM and VF. (a) Quantitative enzymatic assays of acetate, glucose, lactate, and succinate and (b) free amino acid quantification by HPLC were applied to measure metabolite concentrations in the samples vcDMEM and VF. Metabolite concentrations from three biological replicates are depicted as median and range in mM for the 72 h timepoint. The numbers above each bar pair represent Bonferroni adjusted p values derived from two sample two-sided T-test with a significance level of α = 0.05. ND = not detected.
Amino acid composition of metacestodes: Is threonine overrepresented in vesicle fluid, in germinal layer cells, or in germinal layer proteins?
The fact that threonine as the only amino acid showed an unexpectedly high difference between ccDMEM and vcDMEM prompted us to determine the concentrations of free amino acids in VF and germinal layer cells, as well as the amino acid composition of the proteome of germinal layer cells (Supplementary Fig. 2). We compared the concentration differences for the amino acids with lower contents in vcDMEM than in ccDMEM, i.e. consumed metabolites, with their corresponding concentrations in the samples VF, germinal layer cells and germinal layer proteome. All three regressions were significant, when threonine – identified by Cook’s distance outlier analysis - was removed (Fig. 5). Taken together, this confirms that the high threonine consumption by E. multilocularis metacestodes can neither be explained by protein synthesis, nor by storage in germinal layer cells or VF.
Free amino acid concentrations in vesicle fluid, in germinal layer cells, and germinal layer cell proteins. Amino acid uptake by E. multilocularis metacestodes (y-axis) was identified by the difference in amino acid concentrations in vcDMEM (n = 3) and ccDMEM (n = 3) as measured by HPLC. Amino acid consumption is depicted in relation to (a) free amino acids in VF (n = 3, x-axis), (b) free amino acids in germinal layer cells (n = 3, x-axis), and (c) amino acid composition of germinal layer cell proteins (n = 3, x-axis) as analyzed by HPLC. Median values in mM are depicted. The respective linear regressions are given excluding threonine. P values were determined by F-test.
Discussion
In the present study, we analyzed the metabolic footprint and requirements of in vitro cultured E. multilocularis metacestodes by 1H-NMR, by HPLC and by enzymatic assays to identify essential pathways that may constitute druggable targets. Overall, the results obtained within these independent setups and with three analytical methods were in good agreement. Confirming previously published results on protoscoleces48, glucose was the metabolite with the highest consumption, and most likely serves as the major energy source for both, metacestodes and protoscoleces. Under anaerobic conditions, the energy metabolism of E. multilocularis metacestodes is fermentative yielding acetate, alanine, lactate, and succinate as major end products, which accumulate in the culture medium. Glucose was metabolized yielding acetate (26%), alanine (4%), lactate (6%), and succinate (55%, see Fig. 6) as end products. The residual 8% of glucose equivalents were not traceable. The generation of succinate and acetate – in our study the most abundant end products – is of particular interest. Under anaerobic conditions, succinate and acetate are produced in mitochondria by an additional pathway, the malate dismutation pathway. Malate dismutation is found in helminths, marine invertebrates, and euglenids57,58,59. This pathway can be considered as a partial inversion or a shunt of the citric acid cycle. All knowledge on this pathway in helminths is based on nematodes (Ascaris suum and filaria), trematodes (Schistosoma mansoni and Fasciola hepatica) as well as the cestode Hymenolepis diminuta58. Under anaerobic conditions, pyruvate is imported into the mitochondria, but not decarboxylated via pyruvate dehydrogenase. Rather, it is carboxylated to oxaloacetate via pyruvate carboxylase. The reduction of oxaloacetate to malate, the conversion of malate to fumarate and the reduction of fumarate to succinate is performed by the corresponding enzymes of the citrate cycle working in the reverse direction, thereby regenerating NAD from NADH. The reduction of fumarate to succinate is dependent on complexes I and II of the mitochondrial respiratory chain. Complex I transfers electrons from NADH to rhodoquinone, an electron carrier with a much lower redox potential than the mammalian carrier ubiquinone. Rhodoquinone transfers these electrons to complex II which operates in the opposite direction than in oxidative phosphorylation, namely as a fumarate reductase instead of succinate dehydrogenase58,59. Succinate as a final product of this fermentative pathway is then excreted to the medium (see Fig. 6 for an overview). Some parasitic helminths further metabolize succinate to propionate or volatile fatty acids. So far, there is no indication that these reactions also take place in Echinococcus, as these metabolites have never been detected in the parasite, nor are the responsible enzymes identified in the Echinococcus genomes. Thus, for helminths, malate dismutation is an important pathway to eliminate redox equivalents under anaerobic conditions. By generating a proton gradient via complex I, it may even contribute to mitochondrial ATP synthesis59,60. To date, Echinococcus species were largely excluded from studies of the malate dismutation pathway, even though accumulation of succinate was described early, mostly in E. granulosus protoscoleces47,61,62 and cysts37, but also in E. multilocularis protoscoleces47 and metacestodes63. Absent in mammalian cells, malate dismutation may constitute a highly interesting drug target, which has been addressed in one study that investigated this pathway in protoscoleces of E. multilocularis64. Moreover, the high release of metabolic end products could modulate the host response, as it was shown that succinate and fumarate can induce host cell apoptosis65, and succinate stimulates IL-1β secretion in T-cells and could therefore cause inflammation66.
Overview of the mitochondrial energy metabolism and released and consumed metabolites by E. multilocularis metacestodes in vitro. The mitochondrial energy metabolism of E. multilocularis is given based on information from previous studies32,33,47,48,62,75. Aerobic pathways are given in purple, anaerobic ones in blue. Outside the mitochondrial membrane (black lines), glycolysis (G) and fermentation pathways (F, alanine, ethanol, and lactic) are depicted. Inside the mitochondrial membrane, malate dismutation (MD), rhodoquinone (RQ), tricarboxylic acid cycle (TCA), and ubiquinone (UQ) are given. Metabolites consumed in vitro by E. multilocularis metacestodes, as found within this study, are depicted in yellow, released ones in green. Metabolites that were released more than five times are indicated with bold letters and a rectangle (acetate, alanine, fumarate, malate, succinate; aspartate is not depicted). The pathways for consumed metabolites need to be further investigated. Putative pathways for use in the energy metabolism were determined in KEGG (indicated by asterisks)76. The most strongly consumed metabolites are indicated with bold letters and a rectangle (glucose and threonine). Percentages of these metabolites as produced from consumed glucose equivalents are given.
The second most abundant fermentation product of metacestodes detected in this study is acetate which confirms the results described for E. multilocularis protoscoleces47. The generation of acetate as fermentative end product may be a consequence of the malate dismutation pathway which shunts the citrate cycle (see above). Lacking oxaloacetate as acceptor, acetyl-CoA resulting from imported pyruvate or from lipid degradation67,68 transiently accumulates and is hydrolyzed to acetate, thereby regenerating coenzyme A. Acetate is then secreted.
The third major fermentation end product, alanine, is generated from pyruvate by transamination in the cytosol. Conversely, pool sizes of lactate, the classical fermentation end product in mammalian cells, do not markedly differ between the three fluids investigated in this study.
The difference in abundance, but not spectrum, of metabolites between vcDMEM and VF could be explained by the fact that small metabolites (alanine, arginine, glycine, histidine, isoleucine, leucine, lysine, methionine, phenylalanine, serine, tryptophan, tyrosine, and valine) pass the metacestode wall but are selectively accumulated inside. Myo-inositol was detected in traces in VF, not in vcDMEM, but no further focus was laid on this metabolite in the present study. Overall, our results on VF composition are in agreement with previous studies on the composition of E. granulosus ex vivo hydatid fluid34,37,38,41. The only difference is that in E. multilocularis metacestode VF, alanine is the most abundant amino acid whereas in hydatid fluid, glycine is the most abundant amino acid34,41,69. E. multilocularis releases traces of aspartate, glutamate, and glycine into the medium. Thus, we consider that in addition to alanine, these three amino acids can be synthesized by E. multilocularis. Based on transcriptomic information, the synthesis pathways for all other amino acids are incomplete or lacking32. Therefore, they have to be taken up from the culture medium. Most striking is the almost complete exhaustion of the threonine pool (0.67 mM) from the culture medium. This confirms a previous study where threonine was, amongst other metabolites, taken up by E. granulosus metacestodes40, but this effect was not further investigated. The mucin-type glycoprotein Em2, synthesized in germinal layer cells and a major antigen of the laminated layer, is known to be rich in threonine and proline70. However, we could not detect any overuse of threonine for protein synthesis. Therefore, Em2 production cannot be the main cause of the high consumption of threonine by E. multilocularis metacestodes. Interestingly, threonine is known to be the only amino acid absolutely essential for the rapid division of mouse embryonic stem cells71. Threonine is hereby converted by threonine dehydrogenase and 2-amino-3-ketobutyrate coenzyme A ligase into glycine and acetyl-CoA, two metabolites that are then further used for purine synthesis and DNA-replication, and energy generation, respectively71. In the procyclic (insect stage) form of African trypanosomes, threonine is also converted via threonine dehydrogenase and 2-amino-3-ketobutyrate coenzyme A ligase, but finally feeding into fatty acid synthesis72,73,74. Whether the respective pathways are present and active in E. multilocularis should be further investigated. The release of glycine within our in vitro setup is in favor of according pathways being active. Other consumed amino acids, such as isoleucine, leucine, methionine, and valine, could also be metabolized into mitochondrial energy-generating pathways (Fig. 6). Glutamine was not detected by the methods used here, because the media used contained an altered, stabilized form of glutamine.
Taken together, this is the first characterization of the in vitro metabolomic footprint of E. multilocularis metacestodes and the respective composition of VF. The predominance of malate dismutation and the consumption of the amino acid threonine in E. multilocularis metacestodes in vitro lay the basis for further studies on potentially targetable pathways for targeting this most deadly of all helminth diseases.
Change history
17 February 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41598-021-83843-4
References
Bouwknegt, M. et al. Prioritisation of food-borne parasites in Europe, 2016. Euro Surveill. Bull. Eur. Sur Mal. Transm. Eur. Commun. Dis. Bull. 23 (2018).
Deplazes, P. et al. Global Distribution of Alveolar and Cystic Echinococcosis. Adv. Parasitol. 95, 315–493 (2017).
Bebezov, B. et al. Intense Focus of Alveolar Echinococcosis, South Kyrgyzstan. Emerg. Infect. Dis. 24, 1119–1122 (2018).
Gottstein, B. et al. Threat of alveolar echinococcosis to public health - a challenge for Europe. Trends Parasitol., https://doi.org/10.1016/j.pt.2015.06.001 (2015).
Robertson, L. J. Parasites in Food: From a Neglected Position to an Emerging Issue. Adv. Food Nutr. Res. 86, 71–113 (2018).
Kern, P. et al. The Echinococcoses: Diagnosis, Clinical Management and Burden of Disease. Adv. Parasitol. 96, 259–369 (2017).
Grüner, B. et al. Comprehensive diagnosis and treatment of alveolar echinococcosis: A single-center, long-term observational study of 312 patients in Germany. GMS Infect. Dis. 1–12, https://doi.org/10.3205/id000027 (2017).
Lundström-Stadelmann, B., Rufener, R., Ritler, D., Zurbriggen, R. & Hemphill, A. The importance of being parasiticidal… an update on drug development for the treatment of alveolar echinococcosis. Food Waterborne Parasitol., e00040, https://doi.org/10.1016/j.fawpar.2019.e00040 (2019).
Koziol, U., Rauschendorfer, T., Zanon Rodríguez, L., Krohne, G. & Brehm, K. The unique stem cell system of the immortal larva of the human parasite Echinococcus multilocularis. EvoDevo 5, 10 (2014).
Thompson, R. C. A. Biology and Systematics of Echinococcus. Adv. Parasitol. 95, 65–109 (2017).
Gottstein, B. & Hemphill, A. Echinococcus multilocularis: The parasite–host interplay. Exp. Parasitol. 119, 447–452 (2008).
Küster, T., Stadelmann, B., Aeschbacher, D. & Hemphill, A. Activities of fenbendazole in comparison with albendazole against Echinococcus multilocularis metacestodes in vitro and in a murine infection model. Int. J. Antimicrob. Agents 43, 335–342 (2014).
Nicholson, J. K. et al. Metabolic phenotyping in clinical and surgical environments. Nature 491, 384–392 (2012).
Sengupta, A. et al. Host metabolic responses to Plasmodium falciparum infections evaluated by (1)H NMR metabolomics. Mol. Biosyst. 12, 3324–3332 (2016).
Wang, Y. et al. Advances in metabolic profiling of experimental nematode and trematode infections. Adv. Parasitol. 73, 373–404 (2010).
Fernie, A. R. The future of metabolic phytochemistry: larger numbers of metabolites, higher resolution, greater understanding. Phytochemistry 68, 2861–2880 (2007).
Reaves, M. L. & Rabinowitz, J. D. Metabolomics in systems microbiology. Curr. Opin. Biotechnol. 22, 17–25 (2011).
Saric, J. et al. Panorganismal metabolic response modeling of an experimental Echinostoma caproni infection in the mouse. J. Proteome Res. 8, 3899–3911 (2009).
Saric, J. et al. Metabolic Profiling of an Echinostoma caproni Infection in the Mouse for Biomarker Discovery. PLoS Negl. Trop. Dis. 2, e254 (2008).
Saric, J. et al. Systems parasitology: effects of Fasciola hepatica on the neurochemical profile in the rat brain. Mol. Syst. Biol. 6, 396 (2010).
Denery, J. R., Nunes, A. A. K., Hixon, M. S., Dickerson, T. J. & Janda, K. D. Metabolomics-based discovery of diagnostic biomarkers for onchocerciasis. PLoS Negl. Trop. Dis. 4 (2010).
Kokova, D. A. et al. Exploratory metabolomics study of the experimental opisthorchiasis in a laboratory animal model (golden hamster, Mesocricetus auratus). PLoS Negl. Trop. Dis. 11, e0006044 (2017).
Kostidis, S., Addie, R. D., Morreau, H., Mayboroda, O. A. & Giera, M. Quantitative NMR analysis of intra- and extracellular metabolism of mammalian cells: A tutorial. Anal. Chim. Acta 980, 1–24 (2017).
Garcia-Perez, I. et al. Bidirectional correlation of NMR and capillary electrophoresis fingerprints: a new approach to investigating Schistosoma mansoni infection in a mouse model. Anal. Chem. 82, 203–210 (2010).
Li, J. V. et al. Metabolic profiling of a Schistosoma mansoni infection in mouse tissues using magic angle spinning-nuclear magnetic resonance spectroscopy. Int. J. Parasitol. 39, 547–558 (2009).
Li, J. V. et al. Global Metabolic Responses of NMRI Mice to an Experimental Plasmodium berghei Infection. J. Proteome Res. 7, 3948–3956 (2008).
Balog, C. I. A. et al. Metabonomic investigation of human Schistosoma mansoni infection. Mol. Biosyst. 7, 1473–1480 (2011).
Giera, M. et al. The Schistosoma mansoni lipidome: Leads for immunomodulation. Anal. Chim. Acta 1037, 107–118 (2018).
Saric, J. et al. Systematic evaluation of extraction methods for multiplatform-based metabotyping: application to the Fasciola hepatica metabolome. Anal. Chem. 84, 6963–6972 (2012).
Markley, J. L. et al. The future of NMR-based metabolomics. Curr. Opin. Biotechnol. 43, 34–40 (2017).
Bernthaler, P., Epping, K., Schmitz, G., Deplazes, P. & Brehm, K. Molecular characterization of EmABP, an apolipoprotein A-I binding protein secreted by the Echinococcus multilocularis metacestode. Infect. Immun. 77, 5564–5571 (2009).
Tsai, I. J. et al. The genomes of four tapeworm species reveal adaptations to parasitism. Nature 496, 57–63 (2013).
Zheng, H. et al. The genome of the hydatid tapeworm Echinococcus granulosus. Nat. Genet. 45, 1168–1175 (2013).
Celik, C., Amanvermez, R. & Ozkan, K. Free amino acid concentration in hydatid cyst fluids from fertile and infertile human and animal Echinococcus granulosus. Parasite Paris Fr. 8, 343–348 (2001).
Frayha, G. J. & Haddad, R. Comparative chemical composition of protoscolices and hydatid cyst fluid of Echinococcus granulosus (Cestoda). Int. J. Parasitol. 10, 359–364 (1980).
Garg, M. et al. Fertility assessment of hydatid cyst by proton MR spectroscopy. J. Surg. Res. 106, 196–201 (2002).
Hosch, W. et al. Metabolic viability assessment of cystic echinococcosis using high-field 1H MRS of cyst contents. NMR Biomed. 21, 734–754 (2008).
Hurd, H. Echinococcus granulosus: a comparison of free amino acid concentration in hydatid fluid from primary and secondary cysts and host plasma. Parasitology 98(Pt 1), 135–143 (1989).
Kohli, A., Gupta, R. K., Poptani, H. & Roy, R. In vivo proton magnetic resonance spectroscopy in a case of intracranial hydatid cyst. Neurology 45, 562–564 (1995).
Jeffs, S. A. & Arme, C. Echinococcus granulosus (Cestoda): uptake of L-amino acids by secondary hydatid cysts. Parasitology 96(Pt 1), 145–156 (1988).
Yao, M. Y., Xiao, S. H., Feng, J. J., Xue, C. L. & Shimada, M. Effect of mebendazole on free amino acid composition of cyst wall and cyst fluid of Echinococcus granulosus harbored in mice. Zhongguo Yao Li Xue Bao 15, 521–524 (1994).
Bahr, J. M., Frayha, G. J. & Hajjar, J.-J. Mechanism of cholesterol absorption by the hydatid cysts of Echinococcus granulosus (Cestoda). Comp. Biochem. Physiol. A Physiol. 62, 485–489 (1979).
Ingold, K. et al. Efficacies of albendazole sulfoxide and albendazole sulfone against In vitro-cultivated Echinococcus multilocularis metacestodes. Antimicrob. Agents Chemother. 43, 1052–1061 (1999).
Modha, A., Novak, M. & Blackburn, B. J. Treatment of experimental alveolar echinococcosis with albendazole: a 1H NMR spectroscopic study. Can. J. Zool. 75, 198–204 (1997).
Spiliotis, M., Tappe, D., Sesterhenn, L. & Brehm, K. Long-term in vitro cultivation of Echinococcus multilocularis metacestodes under axenic conditions. Parasitol. Res. 92, 430–432 (2004).
Spiliotis, M. et al. Transient transfection of Echinococcus multilocularis primary cells and complete in vitro regeneration of metacestode vesicles. Int. J. Parasitol. 38, 1025–1039 (2008).
McManus, D. P. & Smyth, J. D. Differences in the chemical composition and carbohydrate metabolism of Echinococcus granulosus (horse and sheep strains) and E. multilocularis. Parasitology 77, 103–109 (1978).
McManus, D. P. & Smyth, J. D. Intermediary carbohydrate metabolism in protoscoleces of <span class=“italic”>Echinococcus granulosus</span> (horse and sheep strains) and <span class=“italic”>E. multilocularis</span>. Parasitology 84, 351–366 (1982).
Novak, M., Modha, A. & Blackburn, B. J. Metabolic alterations in organs of Meriones unguiculatus infected with Echinococcus multilocularis. Comp. Biochem. Physiol. B 105, 517–521 (1993).
Rufener, R. et al. Repurposing of an old drug: In vitro and in vivo efficacies of buparvaquone against Echinococcus multilocularis. Int. J. Parasitol. Drugs Drug Resist. 8, 440–450 (2018).
Wishart, D. S. et al. HMDB 4.0: the human metabolome database for 2018. Nucleic Acids Res. 46, D608–D617 (2018).
Dona, A. C. et al. A guide to the identification of metabolites in NMR-based metabonomics/metabolomics experiments. Comput. Struct. Biotechnol. J. 14, 135–153 (2016).
Ghaggart. Csmsoftware/Impacts: Version 1.0.3, https://doi.org/10.5281/zenodo.803330 (2017).
Veselkov, K. A. et al. Recursive segment-wise peak alignment of biological (1)h NMR spectra for improved metabolic biomarker recovery. Anal. Chem. 81, 56–66 (2009).
Worley, B. & Powers, R. Multivariate Analysis in Metabolomics. Curr. Metabolomics 1, 92–107 (2013).
Worley, B. & Powers, R. PCA as a Practical Indicator of OPLS-DA Model Reliability. Curr. Metabolomics 4, 97–103 (2016).
Kita, K., Nihei, C. & Tomitsuka, E. Parasite mitochondria as drug target: diversity and dynamic changes during the life cycle. Curr. Med. Chem. 10, 2535–2548 (2003).
Komuniecki, R. & Harris, B. G. Carbohydrate and Energy Metabolism in Helminths. in Biochemistry and Molecular Biology of Parasites (Academic Press Inc, San Diego, CA 92101, 1995).
Tielens, A. G. Energy generation in parasitic helminths. Parasitol. Today Pers. Ed 10, 346–352 (1994).
Bryant, C. Electron transport in parasitic helminths and protozoa. Adv. Parasitol. 8, 139–172 (1970).
Agosin, M. Studies on the metabolism of Echinococcus granulosus. II. Some observations on the carbohydrate metabolism of hydatid cyst scolices. Exp. Parasitol. 6, 586–593 (1957).
Agosin, M. & Repetto, Y. Studies on the metabolism of echinococcus granulosus. vii. reactions of the tricarboxylic acid cycle in e. granulosus scolices. Comp. Biochem. Physiol. 9, 245–261 (1963).
Novak, M., Hameed, N., Buist, R. & Blackburn, B. J. Metabolites of alveolar Echinococcus as determined by [31P]- and [1H]-nuclear magnetic resonance spectroscopy. Parasitol. Res. 78, 665–670 (1992).
Matsumoto, J. et al. Anaerobic NADH-fumarate reductase system is predominant in the respiratory chain of Echinococcus multilocularis, providing a novel target for the chemotherapy of alveolar echinococcosis. Antimicrob. Agents Chemother. 52, 164–170 (2008).
Wentzel, J. F. et al. Exposure to high levels of fumarate and succinate leads to apoptotic cytotoxicity and altered global DNA methylation profiles in vitro. Biochimie 135, 28–34 (2017).
Jiang, S. & Yan, W. Succinate in the cancer-immune cycle. Cancer Lett. 390, 45–47 (2017).
Houten, S. M., Violante, S., Ventura, F. V. & Wanders, R. J. A. The Biochemistry and Physiology of Mitochondrial Fatty Acid β-Oxidation and Its Genetic Disorders. Annu. Rev. Physiol. 78, 23–44 (2016).
Shi, L. & Tu, B. P. Acetyl-CoA and the regulation of metabolism: mechanisms and consequences. Curr. Opin. Cell Biol. 33, 125–131 (2015).
Garg, M. et al. Differentiation of hydatid cyst from cysticercus cyst by proton MR spectroscopy. NMR Biomed. 15, 320–326 (2002).
Hülsmeier, A. J. et al. A major Echinococcus multilocularis antigen is a mucin-type glycoprotein. J. Biol. Chem. 277, 5742–5748 (2002).
Wang, J. et al. Dependence of mouse embryonic stem cells on threonine catabolism. Science 325, 435–439 (2009).
Gilbert, R. J., Klein, R. A. & Miller, P. G. The role of threonine in the metabolism of acetyl coenzyme a by Trypanosoma brucei brucei. Comp. Biochem. Physiol. B 74, 277–281 (1983).
Millerioux, Y. et al. De novo biosynthesis of sterols and fatty acids in the Trypanosoma brucei procyclic form: Carbon source preferences and metabolic flux redistributions. PLoS Pathog. 14, e1007116 (2018).
Millerioux, Y. et al. The threonine degradation pathway of the Trypanosoma brucei procyclic form: the main carbon source for lipid biosynthesis is under metabolic control: Metabolic adaptation for acetate production. Mol. Microbiol., n/a-n/a, https://doi.org/10.1111/mmi.12351 (2013).
Agosin, M. & Aravena, L. Studies on the metabolism of Echinococcus granulosus III. Glycolysis, with special reference to hexokinases and related glycolytic enzymes. Biochim. Biophys. Acta 34, 90–102 (1959).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Acknowledgements
We thank Caroline J. Sands (Imperial College London, UK) for the bioinformatic support of the NMR post processing pipeline. Many thanks to Andrew Hemphill (Institute of Parasitology, University of Bern) for helpful comments on the manuscript. The study was funded by the Swiss National Science Foundation (SNSF, 179439). D.R. was a recipient of the Karl Enigk Stiftung, Hannover, Germany, for parts of the project.
Author information
Authors and Affiliations
Contributions
B.L., J.L., R.R. and D.R. designed the NMR setup. R.R., D.R., J.L. and B.L. prepared the NMR samples. D.R. and R.R. prepared the samples for quantitative analysis. D.R., J.L. and R.R. performed the NMR analysis. B.L., D.R., S.S., U.K. and C.B. designed the amino acid quantification by HPLC. U.K., C.B., S.S. and D.R. performed the amino acid analysis. D.R. and B.L. designed the quantification by enzymatic assays. D.R. performed the enzymatic assays. D.R., B.L., J.L., J.M. and S.S. interpreted the results. B.L., D.R. and J.M. wrote the manuscript text, which was approved by all co-authors. D.R. and B.L. prepared the figures.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ritler, D., Rufener, R., Li, J.V. et al. In vitro metabolomic footprint of the Echinococcus multilocularis metacestode. Sci Rep 9, 19438 (2019). https://doi.org/10.1038/s41598-019-56073-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-56073-y
This article is cited by
-
Worms and bugs of the gut: the search for diagnostic signatures using barcoding, and metagenomics–metabolomics
Parasites & Vectors (2022)
-
Cestodes in the genomic era
Parasitology Research (2022)
-
Metabolic profiling of liver and faeces in mice infected with echinococcosis
Parasites & Vectors (2021)
-
Structural changes and expression of hepatic fibrosis-related proteins in coculture of Echinococcus multilocularis protoscoleces and human hepatic stellate cells
Parasites & Vectors (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.