Abstract
Complex interplay between the intestinal environment and the host has attracted considerable attention and has been well studied with respect to the gut microbiome and metabolome. Oxygen free radicals such as superoxide and the hydroxyl radical (•OH) are generated during normal cellular metabolism. They are toxic to both eukaryotic and prokaryotic cells and might thus affect intestinal homeostasis. However, the effect of oxygen free radicals on the intestinal environment has not been widely studied. Herein, we applied electron spin resonance spectroscopy with spin trapping reagents to evaluate oxygen free radical production capacity in the intestinal lumen and the faeces of mice. •OH was generated in faeces and lumens of the small and large intestines. There were no remarkable differences in •OH levels between faeces and the large intestine, suggesting that faeces can be used as alternative samples to estimate the •OH production capacity in the colonic contents. We then compared free radical levels in faecal samples among five different mouse strains (ddY, ICR, C57BL/6, C3H/HeJ, and BALB/c) and found that strain ddY had considerably higher levels than the other four strains. In addition, strain ddY was more susceptible to dextran sulphate sodium-induced colitis. These differences were possibly related to the relative abundance of the gut bacterial group Candidatus Arthromitus, which is known to modulate the host immune response. From these results, we suggest that the production capacity of oxygen free radicals in mouse faeces is associated with intestinal homeostasis.
Similar content being viewed by others
Introduction
The intestinal environment is complex and dynamic and is composed of constituents of the diet, digestion products, secretions from the host, the microbiome, and microbial metabolites. Recent human cohort studies have revealed a significant influence of the intestinal microbiome on the host’s health. Inflammatory bowel disease (IBD)1, obesity2, type 2 diabetes3, and atherosclerosis4 have been all associated with changes in the intestinal microbiome. An increase in the Firmicutes/Bacteroidetes5 ratio has been implicated in obesity, whereas dysbiosis might contribute to the pathogenesis of both intestinal and extra-intestinal disorders6,7,8. However, there are significant inter-individual differences in the intestinal microbiome, even in patients with the same disease9, making it difficult to draw conclusions. It is possible that beneficial or toxic substances produced by specific bacteria or by the host might have a critical role in modulating host health.
Oxygen free radicals, such as superoxide (O2•−) and the hydroxyl radical (•OH), are generated by aerobic respiration in bacteria10 and are toxic to both eukaryotic and prokaryotic cells. As the lumen of the large intestine is devoid of oxygen, obligate anaerobes, such as Bacteroides, Clostridium, and Ruminococcus, are major inhabitants of this niche. However, the numbers of oxygen-tolerant facultative anaerobes such as members of Enterobacteriaceae and Enterococcaceae increase during dysbiosis11,12. Moreover, the existence of an intraluminal oxygen gradient has been recently reported13. Based on these observations, we considered the possibility that free radicals are generated within the lumen of the large intestine and that they influence host health. In fact, Huycke et al.14 have reported that •OH and O2•− can be observed in the colon contents of rats inoculated with specific strains of Enterococcus faecalis. These findings warrant further investigation to elucidate the interaction between the intestinal environment and host health based on the role of oxygen free radicals.
Electron spin resonance (ESR) spectroscopy is a useful technique that can detect, identify, and quantify various free radicals and has been successfully used to analyse oxygen free radicals in a variety of biological samples. O2•− and •OH are highly oxidizing species with a reduction potential of ~1.6 V and ~2.6–2.7 V, respectively15. These oxygen free radicals instantly react with almost all biomolecules, and therefore they can only be trapped by a spin trap, provided that the spin trap is in close proximity to the radicals at the exact moment when they are generated. In this study, we applied ESR with spin trapping reagents to assess free radical production capacity in the intestinal lumen and faeces of mice. We found significant differences in faecal free radical production capability among mouse strains with different genetic backgrounds. A possible relationship between this faecal free radical production capacity and the intestinal microbiome and/or intestinal homeostasis is also discussed.
Results
Free radical production capacity in the small and large intestines and faeces
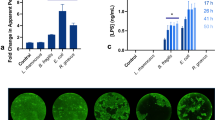
To evaluate the capacity of oxygen free radical production in the lumens of the large and small intestines, we performed a ligated intestinal loop experiment (Fig. 1a) and oxygen free radicals were detected by the ESR spin-trapping technique (Fig. 1b). At the same time, we analysed free radicals in faecal samples. Alpha-(4-pyridyl N-oxide)-N-tert-butyl nitrone (POBN) was used in this study, as the spin adduct from this spin trap is stable in biological environments16, and a sharp signal was observed. As expected, a signal from the POBN spin adduct was observed in all three sample sets. From the hyperfine coupling constant of the spectra (aN = 1.52 mT, aH = 0.22 mT), this POBN adduct was identified as POBN-•OH with reference to a previous report17, demonstrating the presence of •OH production capacity in both the intestinal lumen and faeces. •OH levels were similar in the large intestine and faeces (Fig. 1c), suggesting that •OH production capacity in faeces could be used to predict that in the large intestinal lumen.
Oxygen free radical production capacity in faeces and the small and large intestines. (a) Schematic diagram of ligated intestinal loop assay. ddY mice were anesthetized with isoflurane gas. After opening the abdomen, both ends of the large or small intestine were bound with surgical thread, and POBN solution was injected. The contents were collected and centrifuged, and the supernatants were analysed by electron spin resonance (ESR). Faeces were reacted with POBN solution and analysed similarly. (b) ESR spectra from faeces and the small and large intestines. The center line (from the bottom to the top of the peak) marked with an asterisk (*) was used to calculate the relative amounts of •OH. (c) Relative amounts of •OH. Data are shown as mean values ± SD. N = 6–7 per group. One-way ANOVA showed no significant differences among •OH levels in the faeces, small intestine, and large intestine. (d) A typical ESR spectrum from faeces obtained with the spin-trapping agent CYPMPO. The peak-to-peak intensities of the line marked with an asterisk (*) and double asterisk (**) were used to calculate relative amounts of •OH and O2•−, respectively. (e) Relative amounts of •OH in different mouse strains. Data are shown as mean values ± SD (n = 6 per group). There were significant differences for ddY vs. ICR, C57BL/6, C3H/HeJ, and BALB/c. *p < 0.001 (vs. ddY). (f) Relative levels of •OH in germ free (GF) and ex-germ-free (Ex-GF) mice Ex-GF mice were inoculated with faeces obtained from SPF C57BL/6 mice. Data are shown as mean values ± SD (n = 10 per group). *p < 0.001.
Oxygen free radical production capacity in mouse faeces from different strains
The genetic background of the host may affect free radical production capacity in the intestinal environment. Thus, we analysed this in the faeces of five different mouse strains from the same breeder. In this analysis, we used 5-(2,2-dimethyl-1,3-propoxy cyclophosphoryl)-5-Methyl-1-pyrroline N-oxide (CYPMPO) as a spin-trap agent, because it is more suitable for faecal samples, and more importantly, it enables precise discrimination between •OH and O2•−. A typical ESR spectrum obtained with CYPMPO is shown Fig. 1d. By calculation of the hyperfine coupling constant, •OH (aN = 1.37, 1.35 mT; aH = 1.36, 1.23 mT; aP = 4.87, 4.68 mT) and O2•− (aN = 1.27, 1.27 mT; aH = 1.11, 1.06 mT; aP = 5.24, 5.09 mT) were unambiguously identified with reference to a previous report18. However, O2•− levels were often very low, and thus, we used only •OH for further analysis. Surprisingly, ddY mice had significantly higher •OH levels than the other four strains (p < 0.001). In contrast, BALB/c mice exhibited the lowest free radical levels, although there were no statistically significant differences (Fig. 1e).
Comparison of faecal oxygen free radical levels in germ-free and ex-germ-free mice
It is possible that free radical levels in the intestinal environment are influenced by the gut microbiome or host genetic background. To explore this possibility, we evaluated oxygen free radical production capacity in the faeces of germ-free (GF) and ex-germ-free (Ex-GF) mice (Fig. 1f). The •OH levels in GF-C57BL/6 mice were higher than those in Ex-GF C57BL/6 mice inoculated with faeces from SPF-C57BL/6, indicating that the intestinal microbiome negatively affects •OH levels in this mouse strain. Furthermore, •OH levels in GF-ICR mice were higher than those in GF-C57BL/6 mice. Therefore, the host genetic background might also determine faecal •OH production capability.
Faecal microbiome analysis
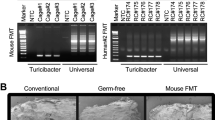
The gut microbiota is likely a key determinant of oxygen free radical production capacity in the mouse intestinal environment. To gain insight into this process, we carried out faecal microbiome analysis. A total of 1,055,363 reads (23,142–57,527 reads per sample) were obtained by sequencing and selection using the QIIME pipeline. Initially, the overall structure of the microbiome was evaluated using unweighted and weighted unifrac analysis (Fig. 2a,b). The unweighted unifrac analysis revealed differences between mouse strains. The microbiomes of ddY or BALB/c mice particularly differed from those of the other three mouse strains. Weighted unifrac analysis showed differences in microbial structure between BLAB/c and other mouse strains. Subsequently, differences in microbial composition were taxonomically evaluated at the phylum and genus levels (Fig. 2c). A total of 10 phyla and 73 genera were taxonomically identified. Although there were no clear differences in the microbial composition between the mouse strains at the phylum level, large differences were observed at the genus level (Fig. 2c). Compared to that in the other four mouse strains, BALB/c mice showed a lower relative abundance of Lactobacillus (Figs. 2c and S1). The genera in which significant differences were observed are shown in Fig. S1. The genera that correlated with •OH production were Anaerostipes, Bilophila, Candidatus Arthromitus, Odoribacter, and Rikenella (p < 0.0021; Fig. 3).
Structure of intestinal microbiome in different mouse strains. Amplicon sequencing was performed using the V1/V2 regions of the 16S rRNA gene, followed by analysis using the QIIME pipeline. (a) Unweighted unifrac analysis of intestinal microbiome. (b) Weighted unifrac analysis of intestinal microbiome. (c) Relative abundance of intestinal microbiota at the genus level. Genera for which the abundance was greater than 2% are shown.
Correlation between relative abundance of genera and relative peak height of •OH. Correlation between relative abundance of 24 genera (Fig. S1) and relative peak heights of •OH were evaluated by Spearman’s rank correlation coefficient. A probability value of less than 0.0021 (0.05/24) was considered statistically significant. The genera for which the probability value was less than 0.0021 are shown.
Susceptibility to dextran sulphate sodium (DSS)-induced colitis
The experiment described previously revealed that the mouse strain ddY exhibited significantly higher levels of faecal free radicals. This might reflect perturbations to intestinal homeostasis. To explore this possibility, we compared susceptibilities to DSS-induced colitis among five mouse strains used previously. On day 4, the body weight of ddY mice was significantly lower than that of ICR and C57BL/6 mice (Fig. 4a). Further, on day 7, the body weight of ddY mice was significantly lower than that of the other four strains, indicating greater susceptibility of ddY mice to DSS-induced colitis. Furthermore, the survival rate was significantly lower in ddY mice, again demonstrating the marked susceptibility of this strain (Fig. 4b). Interestingly, BALB/c mice, which had the lowest radical levels, were highly resistant to DSS-induced colitis.
Discussion
Here, we developed a simple method to evaluate oxygen free radical production capacity in the intestinal environment using the ESR spin-trapping technique. Babbs19 and Erhardt et al.20 have reported methods to quantify •OH production capacity in faeces using DMSO. The reaction times of faeces and DMSO were 5 h and 16 h, respectively. Besides, methane sulfinic acid, which is a byproduct of the reaction between DMSO and •OH, have been routinely analyzed as a proxy of the •OH production capacity in faeces. In comparison with these methods, ESR spin-trapping technique is capable of detecting free radicals in a short time and in a more immediate way. In this study, we firstly compare the •OH production capacities in faeces and colonic contents. Our results suggest that faeces can be used as alternative samples to estimate the •OH production capacity in the colonic contents.
Oxidative stress in intestinal epithelial cells has been implicated in the initiation of colonic diseases such as IBD21 and colon cancer22. Many studies have focused on the host inflammatory response, which results in increased oxidative stress23,24; however, little attention has been paid to oxygen free radicals in the gastrointestinal tract, probably because of the low O2 concentration in the intestinal lumen and separation of epithelial cells and colonic contents by mucus layer. Nonetheless, •OH could be generated from H2O2 by the Fenton reaction, and it has been reported that certain intestinal bacteria such as Lactobacillus, Enterococcus, and Streptococcus have the ability to produce H2O225. In addition, a report that no mucus layer separates colonic contents from epithelium in the proximal colon has been reported26. We also observed similar results (Fig. S2). Thus, it is not unreasonable to consider the oxygen exposure of colonic contents and free radical generation in the colon. Oxygen free radicals in the intestinal environment can have a negative influence on intestinal homeostasis; specifically, O2•− is reportedly produced in the colon of rats inoculated with a specific strain of Enterococcus faecalis, and this radical has been implicated in colon cancer14. In this regard, quantitative analysis of both •OH and O2•− production capacity in the intestinal environment would be valuable; in this study, both were found to be generated in faeces, whereas •OH was the predominant species in normal faecal samples.
The lower •OH levels in Ex-GF-mice compared to those in GF-mice implied the role of the intestinal microbiome in the elimination of oxygen free radicals. In addition, the host genetic background was also found to affect •OH levels. These results suggest that a complex mechanism involving both the intestinal microbiome and host genetic background determines the levels of oxygen free radicals. Sex- and strain-dependent differences in the intestinal microbiome in mice have been reported27. In addition, the association between host genetics and heritable bacterial taxa in the human intestinal microbiome has been reported28. Therefore, it is possible that the host genetic background affects •OH levels in the intestinal environment, which in turn influences the composition of the intestinal microbiome.
DSS-induced colitis in mice has widely been used as an animal model of IBD29,30, in which the intestinal microbiome might play a role in the induction of colitis. Recent studies have shown the interaction between specific bacteria and the host immune response31,32,33,34. Our results show that susceptibility to DSS-induced colitis varies significantly between mouse strains. Moreover, mouse strains showing higher •OH levels tended to be more susceptible to IBD. Our genus-level classification of the intestinal microbiome showed that •OH levels were correlated with the relative abundance of several genera, and particularly Candidatus Arthromitus, a segmented filamentous bacterial group that is oxygen-tolerant35 and known to activate Th17 cells34. Together, these findings suggest that certain bacteria modulate intestinal homeostasis via free radical production, or that free radicals generated by the host metabolism influence the survival of certain bacteria. However, it is necessary to verify whether the free radical production capacity is linked to the onset of colitis, and test if radical scavengers could influence the intestinal microbiome or the outcomes of colitis.
BALB/c mice, which showed the lowest faecal •OH levels and a marked resistance to DSS-induced colitis, exhibited a low relative abundance of the genus Lactobacillus compared to that in the other four mouse strains. Intestinal inflammation can lead to dysbiosis, which is characterized by a marked decrease in the occupancy of obligate anaerobes and an increased relative abundance of facultative anaerobes36. Although the beneficial effects of specific strains of Lactobacillus have been suggested37,38,39, an increase in commensal facultative anaerobes including Lactobacillus might reflect enhanced oxidative stress in the lumen. The reported increase in the abundance of facultative anaerobes such as Lactobacillus during aging in humans40 is probably related to this notion. Similar changes have been observed in an accelerated mouse model of aging41. Thus, understanding the complex interplay between free radicals and facultative anaerobes in the intestinal environment, intestinal homeostasis, and aging is important.
In conclusion, rapid and straightforward experimental techniques to evaluate oxygen free radical production capacity in mouse faeces were established, and we propose their possible application to evaluating the intestinal environment. Although this study did not show direct evidence of free radical production in the intestinal environment, the association between free radical production capacity in faeces and the intestinal microbiome or physiology was significant. Further, it provides a novel perspective in the field of intestinal microbiology. In the future, research combining our free radical analysis with microbiome, metagenome, and metabolome analyses to evaluate the relationships between the intestinal environment and host health status might lead to a better understanding of the complex interplay between the intestinal microbiome and the host.
Materials and Methods
Mice
Six- to seven-week-old male ddY, ICR, C57BL/6, C3H/HeJ, and BALB/c mice were purchased from Japan SLC Inc. (Shizuoka, Japan). The mice were bred in Frontier Laboratories for Value Creation, Sapporo Holdings Ltd., Shizuoka, Japan. They were housed at 4–5 per cage (depth 26, width 18, and height 13 cm), and given water and commercial CRF-1 pellets (CLEA Japan, Inc.) ad libitum. After >1 week of preliminary breeding, the mice were used for experiments. In addition, 6–7-week-old male GF-ICR and GF-C57BL/6 mice were purchased from CLEA Japan, Inc. (Tokyo, Japan). GF-mice were housed in flexible film plastic isolators (CLEA Japan, Inc.) and given sterilized water and sterilized CRF-1 pellets ad libitum. The diet was sterilized using an autoclave (121 °C, 60 min). After 1 week of preliminary breeding, Ex-GF mice were inoculated with 0.2 mL of a 10−1 suspension of faeces obtained from SPF C57BL/6 mice in the stomach using a metal catheter. The protocols were approved by the Sapporo Holdings Animal Use Committee (permit number: 2015–016), and all the experimental procedures were performed according to the guidelines of the Animal Care Committee of Sapporo Holdings Ltd. and were in accordance with the Guide for the Care and Use of Laboratory Animals published by the National Academies Press.
Free radical production capacity in faeces and the intestine
Faeces were collected from 8-week-old-mice and stored on crushed ice until use (approximately 1–3 h). POBN (Tokyo Chemical Industry Co., Ltd., Tokyo, Japan) was used for spin trapping. Faecal samples (approximately 200 mg) were diluted nine-fold with 100 mM POBN solution (Dulbecco’s phosphate buffered saline, 50 mM EDTA, pH 7.4) and mixed for 1 min with a spatula. After 15 s of weak vortexing, samples were left to sit at room temperature (approximately 25 °C) for 30 min. The samples were centrifuged at 12,000 × g for 2 min at 25 °C, and 500 μL of the supernatant was transferred into a flat quartz ESR cell and measured at 25 °C (Keycom X10SA X-band spectrometer; Keycom Corp., Tokyo, Japan). The measurement conditions were as follows: field modulation, 100 kHz; microwave power, 16 mW; field scan width and rate, 16 mT/60 s; time constant, 0.03 s.
To analyse intestinal free radicals, the ligated intestinal loop technique was applied. Mice were anesthetized with isoflurane gas. After opening the abdomen, both ends of the large intestine (2–7 cm from cecum) or small intestine (5–15 cm from cecum) were bound with surgical thread, and 700 μL (for large intestine) or 800 μL (for small intestine) of 100 mM POBN solution was injected using a 22 G intravenous cannula. Thirty minutes after injection, intestinal contents were collected and centrifuged at 12,000 × g for 2 min at 25 °C. The supernatants were analysed by the aforementioned method, whereas precipitates were freeze-dried and weighed. The peak height was normalized by the signal intensity of Mn marker, and corrected by the weight of precipitate. Mice used for the analysis of the large intestine were the same populations as those used for the analysis of faeces. In contrast, mice used for the analysis of the small intestine belonged to separate populations.
Analysis of free radicals in faeces of different mouse strains, GF-mice, and Ex-GF mice
Faeces were collected from 8-week-old male ddY, ICR, C57BL/6, C3H/HeJ, and BALB/c mice, which were used for both free radical and microbiome analyses. Faeces were collected from 7–8-week old male GF-ICR and GF-C57BL/6 mice. Faeces from Ex-GF C57BL/6 mice were collected after 10 days of faecal transplantation. The samples were stored on crushed ice until use. CYPMPO (Shidai System, Tokyo, Japan) was used for spin trapping. Samples (approximately 100 mg) were diluted nine-fold with 100 mM CYPMPO solution (100 mM tris, 50 mM EDTA, pH 8.0) and mixed for 1 min with a spatula. After 15 s of weak vortexing, the samples were left to sit at room temperature (approximately 25 °C) for 30 min. The samples were centrifuged at 3,000 × g for 1 min at 25 °C, and approximately 300 μL of the supernatant was transferred to a flat disposable ESR cell (Radical Research Inc., Hino, Japan). Measurements were performed at 25 °C using a Bruker EMX-plus X-band spectrometer (Bruker Biospin, Billerica, Massachusetts, USA). The measurement conditions were as follows: field modulation, 100 kHz; resonance field, 342–362 mT; field modulation width, 0.1 mT; microwave power, 6 mW; time constant, 0.1 s; scan time, 10 times. The peak height was normalized by the signal intensity of Mn marker.
Microbiome analysis
Fresh faecal samples (approximately 100 mg) were washed twice with 1 mL Dulbecco’s phosphate buffered saline, stored at −80 °C, and used for the analysis of the intestinal microbiome. Bacterial DNA was isolated using the method described by Matsuki et al.42, with some modifications. Briefly, the bacterial suspension was treated with lysis buffer at 70 °C for 10 min in a water bath and vortexed vigorously for 40 s at 6.0 m/s.
Primers for the amplification of the V1 and V2 regions of the 16S rRNA gene, reported by Kim et al.43, were used with some modifications44 to allow for the use of the Ion PGM sequencer (Life Technologies, Carlsbad, CA, USA). PCR was performed in a reaction volume of 25 μL. Each reaction mixture contained 22.5 μL of platinum PCR mix, 2 μL of template DNA (~4 ng), and 0.5 μL of 10 μM primer mix. The amplification reaction was carried out in the Veriti Thermal Cycler (Applied Biosystems, Foster City, CA, USA) using the following program: 3 min at 94 °C, followed by 25 cycles of 30 s, each at 94 °C, 45 s at 55 °C, and 1 min at 68 °C. After each reaction, the mixture was purified using a PureLink Quick PCR Purification Kit (Invitrogen, Carlsbad, CA, USA); the concentration of each purified sample was measured using a Qubit 2.0 fluorometer (Life Technologies). Purified samples were mixed at equal concentrations. The mixed samples were visualized by electrophoresis on 2% agarose gels and purified by gel extraction using the FastGene Gel/PCR Extraction Kit (Nippon Gene Co. Ltd., Tokyo, Japan). Subsequently, emulsion PCR and sequencing were carried out using the Ion PGM sequencing system (Life Technologies).
After sequencing, the obtained reads were analysed by the QIIME pipeline45 (http://qiime.org/) for an assignment of taxonomic classification. The reads that included precise primer sequences were selected, and those with an average quality value >20 were used for further analysis. The reads were grouped into operational taxonomic units (OTUs), with a sequence identity threshold of 97%, and chimeric OTUs were removed using ChimeraSlayer. Unifrac distance analysis was performed, and the proportions of intestinal microbiota at the genus level were determined based on the RDP classifier.
DSS-induced colitis
Seven-week-old ddY, ICR, C57BL/6, C3H/HeJ, and BALB/c mice were used (n = 10 per group). DSS, with a molecular weight of 25,000 (Tokyo Chemical Industry, Tokyo, Japan), was dissolved in tap water to obtain a 4% DSS solution. Mice were allowed free access to the 4% DSS solution. The weight and cumulative survival rate of the mice were then evaluated.
Statistical analysis
One-way analysis of variance with a post-hoc Turkey HSD test was performed for the statistical evaluation of the levels of free radicals and the intestinal microbiome composition. The Spearman’s rank correlation coefficient was used to analyse correlations between intestinal bacteria and free radical production. A probability value of less than 0.05 ÷ number of tests was considered statistically significant. For DSS-induced colitis experiments, the statistical evaluation of differences in survival rates was performed by the generalized Wilcoxon test with Holm-correction by 10 pairs. A probability value of less than 0.05 was considered statistically significant.
Data availability
All sequence data were deposited in the DDBJ sequence read archive database under the accession number DRA008158.
References
Qin, J. et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464, 59–65, https://doi.org/10.1038/nature08821 (2010).
Turnbaugh, P. J. et al. A core gut microbiome in obese and lean twins. Nature 457, 480–484, https://doi.org/10.1038/nature07540 (2009).
Qin, J. et al. A metagenome-wide association study of gut microbiota in type 2 diabetes. Nature 490, 55–60, https://doi.org/10.1038/nature11450 (2012).
Jonsson, A. L. & Backhed, F. Role of gut microbiota in atherosclerosis. Nat. Rev. Cardiol. 14, 79–87, https://doi.org/10.1038/nrcardio.2016.183 (2017).
Turnbaugh, P. J. et al. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444, 1027–1031, https://doi.org/10.1038/nature05414 (2006).
Carding, S., Verbeke, K., Vipond, D. T., Corfe, B. M. & Owen, L. J. Dysbiosis of the gut microbiota in disease. Microb. Ecol. Health Dis. 26, 26191, https://doi.org/10.3402/mehd.v26.26191 (2015).
Vangay, P., Ward, T., Gerber, J. S. & Knights, D. Antibiotics, pediatric dysbiosis, and disease. Cell host & microbe 17, 553–564, https://doi.org/10.1016/j.chom.2015.04.006 (2015).
Schulz, M. D. et al. High-fat-diet-mediated dysbiosis promotes intestinal carcinogenesis independently of obesity. Nature 514, 508–512, https://doi.org/10.1038/nature13398 (2014).
Lozupone, C. A., Stombaugh, J. I., Gordon, J. I., Jansson, J. K. & Knight, R. Diversity, stability and resilience of the human gut microbiota. Nature 489, 220–230, https://doi.org/10.1038/nature11550 (2012).
Farr, S. B. & Kogoma, T. Oxidative stress responses in Escherichia coli and Salmonella typhimurium. Microbiol. Rev. 55, 561–585 (1991).
Larsen, P. E. & Dai, Y. Metabolome of human gut microbiome is predictive of host dysbiosis. GigaScience 4, 42, https://doi.org/10.1186/s13742-015-0084-3 (2015).
Gutierrez-Orozco, F. et al. Intestinal microbial dysbiosis and colonic epithelial cell hyperproliferation by dietary alpha-mangostin is independent of mouse strain. Nutrients 7, 764–784, https://doi.org/10.3390/nu7020764 (2015).
Albenberg, L. et al. Correlation between intraluminal oxygen gradient and radial partitioning of intestinal microbiota. Gastroenterology 147, 1055–1063.e1058, https://doi.org/10.1053/j.gastro.2014.07.020 (2014).
Huycke, M. M., Abrams, V. & Moore, D. R. Enterococcus faecalis produces extracellular superoxide and hydrogen peroxide that damages colonic epithelial cell DNA. Carcinogenesis 23, 529–536 (2002).
Buxton, G. V., Greenstock, C. L., Helman, W. P. & Ross, A. B. Critical Review of rate constants for reactions of hydrated electrons, hydrogen atoms and hydroxyl radicals (·OH/·O− in Aqueous Solution. Journal of Physical and Chemical Reference Data 17, 513–886, https://doi.org/10.1063/1.555805 (1988).
Sato, K., Corbett, J., Mason, R. P. & Kadiiska, M. B. In vivo evidence of free radical generation in the mouse lung after exposure to Pseudomonas aeruginosa bacterium: an ESR spin-trapping investigation. Free Radic. Res. 46, 645–655, https://doi.org/10.3109/10715762.2012.667089 (2012).
Dikalova, A. E., Kadiiska, M. B. & Mason, R. P. An in vivo ESR spin-trapping study: free radical generation in rats from formate intoxication–role of the Fenton reaction. Proc. Natl. Acad. Sci. USA 98, 13549–13553, https://doi.org/10.1073/pnas.251091098 (2001).
Oowada, S., Endo, N., Kameya, H., Shimmei, M. & Kotake, Y. Multiple free-radical scavenging capacity in serum. J. Clin. Biochem. Nutr. 51, 117–121, https://doi.org/10.3164/jcbn.11-113 (2012).
Babbs, C. F. Free radicals and the etiology of colon cancer. Free Radic. Biol. Med. 8, 191–200 (1990).
Erhardt, J. G., Lim, S. S., Bode, J. C. & Bode, C. A diet rich in fat and poor in dietary fiber increases the in vitro formation of reactive oxygen species in human feces. J. Nutr. 127, 706–709 (1997).
McKenzie, S. J., Baker, M. S., Buffinton, G. D. & Doe, W. F. Evidence of oxidant-induced injury to epithelial cells during inflammatory bowel disease. J. Clin. Invest. 98, 136–141, https://doi.org/10.1172/jci118757 (1996).
Seril, D. N., Liao, J., Yang, G. Y. & Yang, C. S. Oxidative stress and ulcerative colitis-associated carcinogenesis: studies in humans and animal models. Carcinogenesis 24, 353–362 (2003).
Jones, R. M., Mercante, J. W. & Neish, A. S. Reactive oxygen production induced by the gut microbiota: pharmacotherapeutic implications. Curr. Med. Chem. 19, 1519–1529 (2012).
Lambeth, J. D. & Neish, A. S. Nox enzymes and new thinking on reactive oxygen: a double-edged sword revisited. Annu. Rev. Pathol. 9, 119–145, https://doi.org/10.1146/annurev-pathol-012513-104651 (2014).
Strus, M. et al. A role of hydrogen peroxide producing commensal bacteria present in colon of adolescents with inflammatory bowel disease in perpetuation of the inflammatory process. J. Physiol. Pharmacol. 60(Suppl 6), 49–54 (2009).
Kamphuis, J. B. J., Mercier-Bonin, M., Eutamene, H. & Theodorou, V. Mucus organisation is shaped by colonic content; a new view. Sci. Rep. 7, 8527, https://doi.org/10.1038/s41598-017-08938-3 (2017).
Elderman, M. et al. Sex and strain dependent differences in mucosal immunology and microbiota composition in mice. Biol. Sex Differ. 9, 26, https://doi.org/10.1186/s13293-018-0186-6 (2018).
Goodrich, J. K. et al. Genetic Determinants of the Gut Microbiome in UK Twins. Cell host & microbe 19, 731–743, https://doi.org/10.1016/j.chom.2016.04.017 (2016).
Okayasu, I. et al. A novel method in the induction of reliable experimental acute and chronic ulcerative colitis in mice. Gastroenterology 98, 694–702 (1990).
Krieglstein, C. F. et al. Regulation of murine intestinal inflammation by reactive metabolites of oxygen and nitrogen: divergent roles of superoxide and nitric oxide. J. Exp. Med. 194, 1207–1218 (2001).
Caprioli, F., Pallone, F. & Monteleone, G. Th17 immune response in IBD: A new pathogenic mechanism. Journal of Crohn’s & colitis 2, 291–295, https://doi.org/10.1016/j.crohns.2008.05.004 (2008).
Farkas, A. M. et al. Induction of Th17 cells by segmented filamentous bacteria in the murine intestine. J. Immunol. Methods 421, 104–111, https://doi.org/10.1016/j.jim.2015.03.020 (2015).
Atarashi, K. et al. Th17 Cell Induction by Adhesion of Microbes to Intestinal Epithelial Cells. Cell 163, 367–380, https://doi.org/10.1016/j.cell.2015.08.058 (2015).
Ivanov, I. I. et al. Induction of intestinal Th17 cells by segmented filamentous bacteria. Cell 139, 485–498, https://doi.org/10.1016/j.cell.2009.09.033 (2009).
Schnupf, P. et al. Growth and host interaction of mouse segmented filamentous bacteria in vitro. Nature 520, 99–103, https://doi.org/10.1038/nature14027 (2015).
Winter, S. E., Lopez, C. A. & Baumler, A. J. The dynamics of gut-associated microbial communities during inflammation. EMBO reports 14, 319–327, https://doi.org/10.1038/embor.2013.27 (2013).
Isolauri, E. & Salminen, S. Probiotics: use in allergic disorders: a Nutrition, Allergy, Mucosal Immunology, and Intestinal Microbiota (NAMI) Research Group Report. J. Clin. Gastroenterol. 42(Suppl 2), S91–96, https://doi.org/10.1097/MCG.0b013e3181639a98 (2008).
Kalliomaki, M. et al. Probiotics in primary prevention of atopic disease: a randomised placebo-controlled trial. Lancet 357, 1076–1079, https://doi.org/10.1016/s0140-6736(00)04259-8 (2001).
Yan, F. et al. Soluble proteins produced by probiotic bacteria regulate intestinal epithelial cell survival and growth. Gastroenterology 132, 562–575, https://doi.org/10.1053/j.gastro.2006.11.022 (2007).
Odamaki, T. et al. Age-related changes in gut microbiota composition from newborn to centenarian: a cross-sectional study. BMC Microbiol. 16, 90, https://doi.org/10.1186/s12866-016-0708-5 (2016).
Houghton, D. et al. Impact of Age-Related Mitochondrial Dysfunction and Exercise on Intestinal Microbiota Composition. J. Gerontol. A Biol. Sci. Med. Sci. 73, 571–578, https://doi.org/10.1093/gerona/glx197 (2018).
Matsuki, T. et al. Quantitative PCR with 16S rRNA-gene-targeted species-specific primers for analysis of human intestinal bifidobacteria. Appl. Environ. Microbiol. 70, 167–173 (2004).
Kim, S. W. et al. Robustness of gut microbiota of healthy adults in response to probiotic intervention revealed by high-throughput pyrosequencing. DNA Res. 20, 241–253, https://doi.org/10.1093/dnares/dst006 (2013).
Wakita, Y. et al. Taxonomic classification for microbiome analysis, which correlates well with the metabolite milieu of the gut. BMC Microbiol. 18, 188, https://doi.org/10.1186/s12866-018-1311-8 (2018).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nature methods 7, 335–336, https://doi.org/10.1038/nmeth.f.303 (2010).
Acknowledgements
We would like to thank Dr. Fukuda Shinji (Institute for Advanced Biosciences, Keio University) for meaningful discussion. We thank Hiroko Kumeno and Takamoto Hino (SAPPORO HOLDINGS LTD.) for help with animal care. We thank Dr. Kiyoshi Takoi (SAPPORO BREWERIES LTD.) for support with ESR.
Author information
Authors and Affiliations
Contributions
Y.W., H. Kameya and S.O. designed this study. Y.W., A.S. and S.S. performed animal experiments. Y.W., A.S. and H. Kameya analysed the free radicals. Y.W. analysed the microbiome. Y.W., H. Kaneda, H. Kameya, Y.T. and S.O. analysed the data. Y.W. and S.O. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
This research did not receive specific grant from any funding agency in the public, commercial, or not-for-profit sectors. Y.W., A.S., H. Kaneda, S.S. and Y.T. are employees of SAPPORO HOLDINGS LTD. The remaining authors have no conflict of interest to disclose.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wakita, Y., Saiki, A., Kaneda, H. et al. Analysis of free radical production capacity in mouse faeces and its possible application in evaluating the intestinal environment: a pilot study. Sci Rep 9, 19533 (2019). https://doi.org/10.1038/s41598-019-56004-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-56004-x
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.