Abstract
Feline chronic enteropathy (CE) is a common gastrointestinal disorder in cats and mainly comprises inflammatory bowel disease (IBD) and small cell lymphoma (SCL). Both IBD and SCL in cats share features with chronic enteropathies such as IBD and monomorphic epitheliotropic intestinal T-cell lymphoma in humans. The aim of this study was to characterize the fecal microbiome of 38 healthy cats and 27 cats with CE (13 cats with IBD and 14 cats with SCL). Alpha diversity indices were significantly decreased in cats with CE (OTU p = 0.003, Shannon Index p = 0.008, Phylogenetic Diversity p = 0.019). ANOSIM showed a significant difference in bacterial communities, albeit with a small effect size (P = 0.023, R = 0.073). Univariate analysis and LEfSE showed a lower abundance of facultative anaerobic taxa of the phyla Firmicutes (families Ruminococcaceae and Turicibacteraceae), Actinobacteria (genus Bifidobacterium) and Bacteroidetes (i.a. Bacteroides plebeius) in cats with CE. The facultative anaerobic taxa Enterobacteriaceae and Streptococcaceae were increased in cats with CE. No significant difference between the microbiome of cats with IBD and those with SCL was found. Cats with CE showed patterns of dysbiosis similar to those in found people with IBD.
Similar content being viewed by others
Introduction
Feline chronic enteropathy (CE) is common in elderly cats and is defined as the presence of clinical signs of gastrointestinal disease for more than 3 weeks in the absence of infectious intestinal diseases (e.g., parasites) and extraintestinal causes (e.g., renal disease, hyperthyroidism)1.
Feline CE mainly comprises inflammatory bowel disease (IBD) and small cell lymphoma (SCL)2,3,4.
Diagnosis and differentiation can be challenging as clinical signs might be virtually the same in both disease entities2,5. A clinical diagnosis of IBD in cats is based on the presence of chronic gastrointestinal signs of at least 3 weeks duration, the absence of known enteropathogens or other causes of signs of gastrointestinal disease, and the histopathologic confirmation of intestinal inflammation6. Thus, the diagnosis of IBD and the differentiation from SCL requires histopathologic examination of tissue biopsies collected under general anesthesia7,8. In cases where histopathology results are ambiguous, additional diagnostic modalities such as immunohistochemistry and clonality testing are indicated3,4,9,10. Hence, the diagnosis of IBD and SCL is elaborate, expensive, time- and resource consuming and invasive. Treatment of IBD and SCL is usually based on immunosuppression using various steroids and cytotoxic medication such as chlorambucil or cyclosporine2,6,11. Therefore, less invasive diagnostic and treatment modalities would be highly desirable.
The intestinal microbiome plays a substantial role in modulating the host’s immune system within and beyond the gastrointestinal tract. Studies in people12,13,14 and dogs15,16,17 with IBD have found alterations in the composition of the intestinal microbiome that might impair the host’s health status. These changes are commonly referred to as dysbiosis14. Despite variations among studies, species, and individuals, common themes characterize intestinal dysbiosis. Across different species, three main hallmarks of dysbiosis have been described: a reduction in overall bacterial diversity (alpha diversity)15,18,19,20, a decreased stability of microbial communities and thus a higher fluctuation rate over time21, and a reduction in obligately anaerobic taxa of the phyla Firmicutes and Bacteroidetes at the expense of an increase in facultative anaerobes, including members of the family Enterobacteriaceae15,17,18,19,21,22,23,24. Dysbiosis has been described in people with various forms of enteropathy, including ulcerative colitis, Crohn’s disease, and colorectal cancer, and it might be a driver or consequence of chronic inflammation and malignant transformation25. Feline IBD and SCL share some features with chronic enteropathies in people. Both human and feline IBD is characterized by chronic inflammatory changes in the gastrointestinal tract3,26. Feline SCL is characterized by a monomorphic infiltration of the intestinal mucosa with small to medium lymphocytes, mostly in the small intestine and is often associated with epitheliotropism4. This histologic appearance resembles that of monomorphic epitheliotropic intestinal T-cell lymphoma (MEITL) in people, formerly known as enteropathy associated T-cell lymphoma Type 2 (EATL Type 2)27,28. Whereas SCL is quite common in cats, MEITL is rare in people, and thus large intersectional studies are scarce28. Both IBD and SCL occur spontaneously and frequently in cats, and thus the cat could be an interesting model for IBD or MEITL in people.
Several studies have characterized the fecal microbiome in cats with acute and chronic diarrhea18,29. However, in previous studies the cats were categorized based on clinical signs without confirmation of the underlying disease process. This study aimed to characterize and compare the fecal microbiome in healthy cats and cats with histopathologically confirmed CE (IBD or SCL).
Results
Animal demographics and clinical activity index
A total of 65 cats were enrolled into this study, 38 healthy cats and 27 cats with chronic enteropathy (13 with IBD and 14 with SCL). A fecal sample was collected from all cats. Demographic characteristics are shown in Table 1.
Age did not differ significantly between healthy cats (median age: 9 years, range: 1–15 years) and cats with CE (median age: 10 years, range: 2–16 years; P = 0.052). Cats with CE had a significantly lower body weight (median: 4.6 kg, range: 2.9–10.5 kg) and body condition score (BCS; median: 4, range: 1–9) than healthy cats (median body weight: 5.4 kg, range: 2.5–8.6 kg, median BCS: 5, range: 4–9; P = 0.035 and P < 0.001, respectively). Cats with SCL were significantly older (median age: 11.5 years, range: 7–16 years) than cats with IBD (median age: 8 years, range: 2–16; P = 0.028). Cats with IBD and cats with SCL did not show statistically significant differences with regard to sex (p = 0.816), body weight (p = 0.454), and BCS (p = 0.529). Cats with CE had a median feline chronic enteropathy activity index (FCEAI)1 score of 5 (range: 2–11). FCEAI did not differ between cats with IBD (median: 6, range: 3–11) and cats with SCL (median: 5, range: 2–10; p = 0.838).
Sequence analysis and rarefaction
In total, the sequence analysis of the 65 fecal samples yielded 2,837,900 quality sequences (median per sample: 73,741; range: 43,660–145,373).
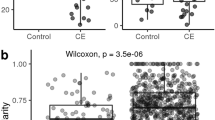
Alpha diversity at a depth of 43,660 sequences, as described by observed OTUs, Shannon Diversity Index, and Faith Phylogenetic Diversity Index, was significantly lower in cats with CE than in healthy cats (P = 0.003, P = 0.008, and P = 0.019, respectively; Fig. 1). In addition, alpha diversity appeared to continuously decrease from healthy cats, over cats with IBD to cats with SCL (Results for Kruskal Wallis ANOVA: observed OTUs P = 0.015, Shannon index P = 0.030, Phylogenetic Diversity Index P = 0.049). However, alpha diversity indices did not differ significantly between cats with IBD and cats with SCL. Detailed results for alpha diversity indices are shown in Supplementary Table 1.
Alpha diversity indices at a depth of 43.660 sequences. (A) Observed OTUs. (B) Shannon Diversity Index. (C) Faith Phylogenetic Diversity Index. Figures on the left show the comparison between healthy cats and cats with chronic enteropathy (FCE). Figures on the right show the comparison between different the CE subgroups, inflammatory bowel disease (IBD) and small cell lymphoma (SCL). The top dashed line represents the Kruskal Wallis ANOVA comparison, solid parentheses represent post hoc multiple comparisons using Dunn’s test.
Microbial communities
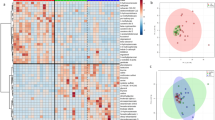
Although cats with CE showed no obvious visible clustering on Principal Component Analysis (PCoA) compared to healthy cats, a statistically significant difference between the two groups was found based on ANOSIM of unweighted Unifrac distances (P = 0.023, R = 0.073: Fig. 2A). Individual bacterial groups were analyzed using a Kruskal Wallis test. Taxa found to be significantly different before correction for the false discovery rate (FDR) are listed in Table 2. Within the phylum Firmicutes, bacterial taxa belonging to the family Ruminococcaceae (unclassified species of the genus Oscillospira) and members of the genus Turicibacter (class Bacilli, order Turicibacterales, family Turicibacteraceae) were significantly less abundant in cats with CE than in healthy cats. In addition, cats with CE had significantly decreased bacterial populations belonging to members of the phyla Bacteroidetes (one undetermined species and Bacteroides plebeius) and Actinobacteria (genus Bifidobacterium). In contrast, members of the families Enterobacteriaceae and Streptococcaceae were significantly more abundant in feces from cats with CE. However, although trends were noted, no statistically significant differences were found after correction for FDR. Figure 2B–F depicts results of the statistical analysis for some selected taxa.
(A) Beta diversity. Principal coordinate analysis (PCoA) of unweighted UniFrac distances of 16S rRNA genes. Analysis of similarity (ANOSIM) revealed significantly different between healthy cats and cats with chronic enteropathy (P = 0.023, R = 0.073), although with a small effect size and no visible clustering. Healthy cats are depicted as blue, cats with inflammatory bowel disease (IBD) as yellow, and cats with small cell lymphoma (SCL) as red dots. (B–F) Univariate analysis of relative percent abundance of selected taxa. Members of the family Ruminococcaceae, of the genus Bifidobacterium and the species Bacteroides plebeius were found to be lower in cats with chronic enteropathy (CE) compared to healthy cats. Members of the genus Streptococcus and the family Enterobacteriaceae were found to be higher in cats with CE compared to healthy cats.
Conversely, based on Linear Discriminant Analysis Effect Size (LEfSe), bacteria of the families Bifidobacteriaceae, Ruminococcaceae, Turicibacteriaceae, and Paraprevotellaceae were associated with feces from healthy cats, while Enterobacteriaceae and Streptococcaceae were associated with those of cats with CE. A detailed summary of the LEfSe results can be found in Table 3.
Effect of disease subtype on the feline fecal microbiota
Similar to alpha-diversity, beta-diversity showed continuous changes when comparing healthy cats, cats with IBD, and cats with SCL, with sequential increases or decreases of relative percentages between groups. According to Kruskal Wallis tests, the abundance of members of the genus Bifidobacterium (class Actinobacteria) differed significantly among the three groups, with highest numbers in healthy cats and lowest numbers in cats with SCL. In contrast, bacteria within the families Enterobacteriaceae (phylum Proteobacteria) and Streptococcaceae (phylum Firmicutes) serially increased from healthy cats, to cats with IBD to cats with SCL. However, after correction for the false discovery rate (FDR), none of the differences remained statistically significant. In addition, no significant differences in microbial communities between cats with IBD and cats with SCL were observed. A detailed summary of relative percentages of the most abundant bacterial groups appears in Supplementary Table 2.
Discussion
To our knowledge, this is the first study comparing the fecal microbiome in a cohort of cats with histopathologically confirmed spontaneous CE to that of clinically healthy cats using untargeted Illumina sequencing analysis. Alpha diversity was significantly lower in cats with CE than in healthy cats. In particular, cats with CE tended to show a lower abundance of obligately anaerobic members of the phyla Firmicutes (family Ruminococcaceae and Turicibacteraceae), Bacteroidetes (e.g. Bacteroides plebeius and unclassified species), and Actinobacteria (genus Bifidobacterium). In contrast, facultative anaerobes such as Enterobacteriaceae and Streptococcaceae tended to be more abundant in cats with CE than in healthy cats. However, although we found differences in abundance of various bacterial taxa between different groups of cats, there were no statistically significant differences after correction for FDR. Nevertheless, the trends found in our cohort of cats mirror common patterns of fecal and mucosal dysbiosis described in other species such as humans19,21,22,23,24 and dogs15,17,30, i.e. decreased bacterial diversity, decreased members of obligate anaerobes (Firmicutes and Bacteroidetes) and increased facultative anaerobes (especially of the family of Enterobacteriaceae). Dysbiosis has been documented in humans with various enteropathies, such as IBD (ulcerative colitis and Crohn’s disease), antibiotic-associated diarrhea, necrotizing enterocolitis, and colorectal cancer14,31 and also in dogs with chronic enteropathies16,30,32. Dysbiotic patterns appear to be similar across different forms of enteropathy; however, because the rarity of the disease, there are no studies published on the microbiomes of people with MEITL. A recent study analyzing the microbiome in feces from dogs with IBD or intestinal lymphoma compared to healthy dogs found dysbiosis to be associated with both disease entities32. Similar to the current study, the authors used an untargeted Illumina sequencing approach to characterize the fecal microbiome. The authors reported an increased abundance of Coprococcus, Oscillospira, and Eubateria in dogs with intestinal lymphoma compared to healthy dogs. These findings might indicate species differences in disease phenotype and etiopathogenesis.
One universal activity of the intestinal microbiome is the metabolism and fermentation of carbohydrates into short chain fatty acids, such as acetate, butyrate, and propionate. Members of the phylum Firmicutes are mostly anaerobic and have been shown to exert indirect anti-inflammatory and immune-modulatory effects by producing short chain fatty acids, particularly butyrate14. Butyrate is the major energy source of colonocytes33, thereby contributing to epithelial cell proliferation and repair and to intestinal barrier integrity34. In addition, evidence exists that butyrate may exhibit anti-inflammatory and anti-carcinogenic properties35,36. The main butyrate-producing bacteria in the human colon are members of the families Lachnospiraceae and Ruminococcaceae (phylum Firmicutes), and their fecal abundance in people is often depleted in dysbiotic states, including UC and CD21,37. Our report appears to be the first to show a depletion of butyrate-producing bacteria in cats with CE. Determining the association between the fecal abundance of Ruminococcaceae and butyrate concentration in cats with CE might further elucidate the function of this taxon in cats with CE.
Another major hallmark of dysbiosis involves members of the phylum Bacteroidetes. Bacteroidetes are highly abundant in the healthy microbiome but are decreased in the mucosal or fecal microbiome from humans and dogs with various forms of enteropathy17,19,22. Interestingly, Bacteroides plebeius has been found to be strongly associated with remission in people with CD38. In our cohort, Bacteroidetes and specifically Bacteroides plebeius tended to be decreased in cats with CE. Investigating the association between disease outcome and abundance of Bacteroides plebeius might be of interest for future studies in cats with CE.
Our study showed a trend towards a decreased abundance of Bifidobacteria in cats with IBD and SCL. Bifidobacteria are also commonly decreased in fecal and mucosal samples from human patients with IBD14,39. Various Bifidobacteria strains have been shown to exert anti-inflammatory properties by regulating immune cells and cytokine networks40 and by directly and indirectly enhancing intestinal barrier function41,42. For instance, Bifidobacterium strains have been shown to induce IL-10 producing regulatory T-cells43,44 and to exert immunoinhibitory effects by interacting with Toll-like receptor-245. Therefore, Bifidobacteria have become an attractive therapeutic target and are often used in probiotic formulations46. Our study confirms findings of a previous study utilizing fluorescence in situ hybridization, where Bifidobacteriaceae were reported to be decreased in fecal samples in cats with IBD. To the authors’ knowledge, such a decrease has not been documented in dogs. This highlights the difference between species and might point toward Bifidobacteria as potential therapeutic targets in cats with CE.
Besides a decrease in obligate anaerobic bacteria, dysbiosis is commonly characterized by an increase in facultative anaerobic bacteria, specifically members of the family of Enterobacteriaceae14,47,48. This phenomenon might be explained by the oxygen gradient model49,50. In this model it is hypothesized that during a steady state, the mucosal microbiome is controlled by mucosal immune responses, the intestinal barrier, and competition with luminal bacteria. The intestinal mucosa shows an oxygen gradient, in which the mucosal interface is mostly aerobic while the lumen is mostly anaerobic. It is thought that during inflammation, the intestinal barrier breaks downs, thereby increasing the luminal oxygen tension. This leads to a translocation and expansion of aerotolerant taxa usually located close to the mucosa centripetal into the lumen and centrifugal across the epithelial barrier into the lamina propria, contributing to the inflammatory response49,50. Our cohort of cats followed this pattern, with a trend of higher abundance of facultative anaerobic taxa, specifically Enterobacteriaceae and Streptococcaceae. Both taxa typically consist of facultative anaerobic members and thus might thrive with increased luminal oxygen tension. An increased abundance of Streptococcaceae has previously been documented in feces from people and dogs with IBD51,52.
Our study has several limitations. With regards to enrollment criteria, of 38 control cats enrolled into this study, 22 cats underwent a physical examination and laboratory testing to verify their health status, while 16 cats were classified as healthy based on owner responses to a questionnaire alone. In addition, a clinical diagnosis of IBD was based on the presence of chronic gastrointestinal signs of at least 3 weeks duration, the absence of known enteropathogens or other causes of signs of gastrointestinal disease, and the histopathologic confirmation of intestinal inflammation6. Thus, food-responsive enteropathy was not excluded in all of the enrolled cats. However, many human patients with IBD show complete or partial responses to dietary interventions without being reclassified as having food-responsive enteropathy53,54,55,56. Our study aimed to characterize clinically relevant differences between the fecal microbiome of cats with CE when compared to healthy control cats. We controlled for demographic characteristics such as age, sex, and breed. Environmental factors, such as housing, diet, previous parasitic or systemic infections etc. were not controlled for in this study, as it would have affected the clinical relevance of the results. This concept is supported by recent studies showed that standardization in is a major source of poor reproducibility of preclinical trials57,58.
With regards to results of this study, the following limitations have to be acknowledged. Although, we found differences in the abundance of various bacterial taxa between different groups of cats, there were no statistically significant differences after correction for FDR, and thus differences did not appear to be as strong as they have been reported for humans and dogs. This phenomenon might be explained by the different disease phenotypes. Whereas diarrhea is the predominant clinical sign of chronic enteropathy in humans59 and dogs60,61, it is less common in cats, in which weight loss, hyporexia, and vomiting are the dominant clinical signs of CE1,2. In our cohort of cats, only 5 out of 27 cats showed diarrhea. The reason for the different disease phenotypes in humans, dogs, and cats is not entirely clear, but factors may include different disease localization (i.e. small vs. large bowel) and different disease pathophysiology. Cats often show involvement of the small intestinal tract1,2,4, but the fecal microbiome represents predominantly bacterial communities present in the distal part of the intestinal tract62. In addition, the fecal microbiome might not accurately represent the mucosa-associated microbiome62. Therefore, investigating the luminal content within the small intestine or the mucosal microbiome might reveal more distinct differences in microbial communities in cats than this study of fecal samples. However, a recent study comparing the fecal and mucosal microbiomes in human patients with and without IBD found large overlaps between the two microbial habitats in each group49. Although it is important to point out that IBD in humans is predominantly a large bowel disease. A recent study, comparing the mucosa associated microbiome in cats with CE using targeted analysis by fluorescence in situ hybridization found significantly increased numbers of Bacteroides spp. in ileal biopsies from cats with SCL compared to biopsies from cats with IBD63. These findings further underline possible differences between intestinal compartments (small vs. large bowel) and media (e.g. fecal vs mucosa-associated microbiome). We also cannot exclude that a larger sample size would have revealed statistically significant differences after FDR correction. Finally, the mode and speed of fecal collection and processing was slightly different between groups and individuals and might have had an effect on our results.
Differentiation of IBD from SCL in cats can be challenging as clinical signs as well as intestinal mucosal changes can be overlapping3. In addition, mucosal changes can have an uneven distribution, inflammation and neoplastic lesions can coexist might and it has been suggested that inflammatory lesions might progress to lymphoma over time3,4. Hence, every classification into groups of IBD and SCL is associated with a degree of uncertainty. However, all cats in the CE group underwent gastoduodenoscopy and ileoscopy. In addition, immunohistochemistry and clonality testing were applied, where indicated, to ensure the maximum accuracy ion class assignment.
Today, histopathology, immunohistochemistry and clonality testing on intestinal samples is considered to be the gold standard for the differentiation of inflammatory from neoplastic lesions10. However, histopathological examination of intestinal tissue biopsies from dogs and cats is associated with a high inter-observer variability64. In addition, a recent study showed that healthy elderly cats can also have abnormal histopathological findings without any apparent clinical significance65. Therefore, alternative methods for the diagnosis and differentiation of feline IBD and SCL would be desirable. We did not find any bacterial taxa in which abundance differed significantly between cats with IBD and SCL. However, as a striking pattern many bacterial taxa appeared to either serially increase (e.g., Enterobacteriaceae) or decrease (e.g., Actinobacteria, Bifidobacteria, Prevotella) when comparing to healthy cats, cats with IBD, and cats with SCL. Similarly, progression of IBD to SCL over months to years has long been suspected, and inflammatory lesions frequently coexist with SCL4,9. Our findings might lend support to the hypothesis that IBD and SCL are not two different diseases but rather a continuum.
In summary, we found that cats with CE show patterns of dysbiosis that have previously been described in people with IBD. Obligately anaerobic taxa in the phyla Firmicutes, Bacteroidetes, and Actinobacteria were depleted in cats with CE, while facultative anaerobes such as Enterobacteriaceae and Streptococcaceae were more abundant. Bacteroides plebeius, a species shown to be associated with positive outcome in people with CD, was decreased in our cohort of cats.
Methods
Animals
This prospective study was conducted at the Veterinary Medical Teaching Hospital at Texas A&M University between May 2015 and September 2017. The study protocol was approved by the Texas A&M University Institutional Animal Care and Use Committee and all methods were performed in accordance with relevant guidelines and regulations.
The health status of cats in the group considered healthy was verified by an owner questionnaire on general and gastrointestinal health. The questionnaire covered the following areas: attitude/activity, appetite, drinking, urination, chronic illnesses, weight loss, vomiting, diarrhea, and treatment with antibiotics, antacids, anti-inflammatory drugs, or steroids. In 22 cats, physical examination was performed by a single board-certified internist (SM). The body condition score was assessed using a previously established a nine-point condition scoring system66. Blood was collected from a peripheral vein or the jugular vein and the following tests were performed: complete blood count, serum chemistry profile, total T4, cobalamin, folate, feline pancreatic lipase immunoreactivity (fPLI), and feline trypsin-like immunoreactivity (fTLI). Cats with gastrointestinal signs (weight loss, hyporexia, vomiting > 2x/ month, diarrhea) within 6 months prior to enrollment were excluded. In addition, cats with systemic diseases, chronic illnesses or clinically significant laboratory abnormalities were excluded from the study. Finally, cats that had received any antibiotics, antacids, anti-inflammatory drugs, or corticosteroids within the past 6 months were excluded.
Cats with clinical signs of chronic enteropathy (weight loss, hyporexia, vomiting, diarrhea) of at least 3 weeks duration were eligible for enrollment into the CE group. Cats in this group were either presented to the Small Animal Hospital at Texas A&M University, College Station, Texas, or the Veterinary Specialty Hospital, San Diego, California. Extra-gastrointestinal disease as well as possible infectious intestinal diseases were excluded based on a complete blood count, serum chemistry profile, total T4 and fecal examination. All cats in this group underwent gastro-duodenoscopy and ileo-colonoscopy for diagnostic purposes. Histopathologic examination of H&E stained endoscopic formalin-fixed, paraffin-embedded (FFPE) tissue sections was performed by board-certified pathologists (MA or JSE) blinded to the clinical status of the cats.
Cases with a histopathological diagnosis of SCL or in which the pathologist was suspicious of an underlying SCL underwent additional diagnostic testing with immunohistochemistry and PCR for antigen receptor rearrangement testing for diagnostic confirmation. A final diagnosis of IBD or SCL was reached upon integration of results from histopathology, immunohistochemistry, and PARR based on the current EuroClonality/BIOMED-2 guidelines for interpretation and reporting of Ig/TCR clonality testing in suspected lymphoproliferations10,67,68. Cats that had received antibiotics within 4 weeks or corticosteroids within the past 2 weeks prior to fecal sampling were excluded from the study.
Spontaneously passed fecal samples were collected from healthy cats, refrigerated and shipped to the Texas A&M Gastrointestinal Laboratory within 24 hours of passing. Fecal samples from cats with CE were either collected after spontaneous void or digitally while the cat was under anesthesia for endoscopy. All samples were shipped on cold packs or on dry ice. Upon arrival, fecal samples were immediately transferred to a lysis buffer and DNA was extracted using the Mobio Power Soil DNA Extraction kit (MoBio Laboratories, Inc., CA) following manufacturer’s instructions.
Amplification and sequencing of the V4 variable region of the 16 S rRNA gene were performed utilizing the Illumina MiSeq Sequencing platform. Sequencing was performed at MR DNA (Shallowater, TX, USA) following the manufacturer’s guidelines using forward and reverse primers: 515 F (5′-GTGCCAGCMGCCGCGGTAA-3′) and 806 R (5′- GGACTACVSGGGTATCTAAT-3′). Briefly, the PCR reaction was performed in a single-step 28 cycle PCR using the HotStarTaq Plus Master Mix Kit (Qiagen, USA) under the following conditions: 94 °C for 3 minutes, followed by 28 cycles (5 cycles used on PCR products) of 94 °C for 30 seconds, 53 °C for 40 seconds and 72 °C for 1 minute, after which a final elongation step at 72 °C for 5 minutes was performed. After sequencing, barcodes and primers were removed from the sequences; then short (<150 bp), ambiguous, homopolymeric, and chimeric sequences were depleted from the dataset. Operational Taxonomic Units (OTUs) were assigned based on at least 97% sequence similarity using the QIIME 2.0 pipeline69. Sequences assigned as chloroplast, mitochondria and Unassigned were removed before downstream analysis. Additionally, OTUs assigned to the phylum cyanobacteria were considered to be potential plant chloroplast contaminants and excluded from the analysis. Sequences were rarefied to an equal depth of 12,000 sequences per sample. The sequences were deposited in SRA under accession number SRP168128.
Statistical analysis
All datasets were tested for normality using the Shapiro-Wilk test (JMP 10, SAS software Inc.). Differences in bacterial communities between healthy cats and cats with CE were analyzed using the phylogeny-based unweighted UniFrac distance metric, and PCoA plots and rarefaction curves were generated within QIIME69. ANOSIM (Analysis of Similarity) within the software package PRIMER 6 (PRIMER-E Ltd., Luton, UK) was used to determine significant differences in microbial communities between healthy cats and diseased cats. Because most datasets did not meet the assumptions of normal distribution, statistical testing between healthy and diseased cats were performed using non-parametric Kruskal-Wallis tests or a Mann-Whitney U test where applicable. The resulting p-values were adjusted for multiple comparisons using the Benjamini & Hochberg’s False Discovery Rate (FDR), and an adjusted q < 0.05 was considered statistically significant70. A Dunn’s post-test was used to determine which disease types differed significantly. Linear discriminant analysis Effect Size (LEfSe) was used to elucidate bacterial taxa (16 S rRNA genes) associated with healthy or diseased cats. LEfSe was used online in the Galaxy workflow framework.
The data that support the findings of this study are available from the corresponding author (SM) upon reasonable request.
Ethical approval and informed consent
The study protocol was approved by the Texas A&M University Animal Care and Use Committee (IACUC 2015-0276 CA and IACUC 2014-0369 CA).
References
Jergens, A. E. et al. A clinical index for disease activity in cats with chronic enteropathy. J. Vet. Intern. Med. 24, 1027–1033, https://doi.org/10.1111/j.1939-1676.2010.0549.x (2010).
Jergens, A. E. Feline idiopathic inflammatory bowel disease: what we know and what remains to be unraveled. J. Feline Med. Surg. 14, 445–458, https://doi.org/10.1177/1098612X12451548 (2012).
Sabattini, S. et al. Differentiating feline inflammatory bowel disease from alimentary lymphoma in duodenal endoscopic biopsies. J. Small Anim. Pract. 57, 396–401, https://doi.org/10.1111/jsap.12494 (2016).
Moore, P. F., Rodriguez-Bertos, A. & Kass, P. H. Feline gastrointestinal lymphoma: mucosal architecture, immunophenotype, and molecular clonality. Vet. Pathol. 49, 658–668, https://doi.org/10.1177/0300985811404712 (2012).
Barrs, V. R. & Beatty, J. A. Feline alimentary lymphoma: 1. Classification, risk factors, clinical signs and non-invasive diagnostics. J. Feline Med. Surg. 14, 182–190, https://doi.org/10.1177/1098612x12439265 (2012).
Jergens, A. E. & Simpson, K. W. Inflammatory bowel disease in veterinary medicine. Front. Biosci. E4, 1404–1419 (2012).
Day, M. J. et al. Histopathological standards for the diagnosis of gastrointestinal inflammation in endoscopic biopsy samples from the dog and cat: a report from the World Small Animal Veterinary Association Gastrointestinal Standardization Group. J. Comp. Pathol. 138(Suppl 1), S1–43, https://doi.org/10.1016/j.jcpa.2008.01.001 (2008).
Washabau, R. J. et al. Endoscopic, biopsy, and histopathologic guidelines for the evaluation of gastrointestinal inflammation in companion animals. J. Vet. Intern. Med. 24, 10–26 (2010).
Moore, P. F., Woo, J. C., Vernau, W., Kosten, S. & Graham, P. S. Characterization of feline T cell receptor gamma (TCRG) variable region genes for the molecular diagnosis of feline intestinal T cell lymphoma. Vet. Immunol. Immunopathol. 106, 167–178, https://doi.org/10.1016/j.vetimm.2005.02.014 (2005).
Keller, S. M., Vernau, W. & Moore, P. F. Clonality Testing in Veterinary Medicine: A Review With Diagnostic Guidelines. Vet. Pathol. 53, 711–725, https://doi.org/10.1177/0300985815626576 (2016).
Stein, T. J., Pellin, M., Steinberg, H. & Chun, R. Treatment of feline gastrointestinal small-cell lymphoma with chlorambucil and glucocorticoids. J. Am. Anim. Hosp. Assoc. 46, 413–417, https://doi.org/10.5326/0460413 (2010).
Frank, D. N., Zhu, W., Sartor, R. B. & Li, E. Investigating the biological and clinical significance of human dysbioses. Trends Microbiol. 19, 427–434, https://doi.org/10.1016/j.tim.2011.06.005 (2011).
Tamboli, C. P., Neut, C., Desreumaux, P. & Colombel, J. F. Dysbiosis in inflammatory bowel disease. Gut 53, 1–4 (2004).
Walker, A. W. & Lawley, T. D. Therapeutic modulation of intestinal dysbiosis. Pharmacol. Res. 69, 75–86, https://doi.org/10.1016/j.phrs.2012.09.008 (2013).
Minamoto, Y. et al. Alteration of the fecal microbiota and serum metabolite profiles in dogs with idiopathic inflammatory bowel disease. Gut Microbes 6, 33–47, https://doi.org/10.1080/19490976.2014.997612 (2015).
AlShawaqfeh, M. K. et al. A dysbiosis index to assess microbial changes in fecal samples of dogs with chronic inflammatory enteropathy. FEMS Microbiol. Ecol. 93 https://doi.org/10.1093/femsec/fix136 (2017).
Suchodolski, J. S. et al. The fecal microbiome in dogs with acute diarrhea and idiopathic inflammatory bowel disease. PloS one 7, e51907, https://doi.org/10.1371/journal.pone.0051907 (2012).
Suchodolski, J. S. et al. The fecal microbiome in cats with diarrhea. PLoS One 10, e0127378, https://doi.org/10.1371/journal.pone.0127378 (2015).
McLaughlin, S. D. et al. The bacteriology of pouchitis: a molecular phylogenetic analysis using 16S rRNA gene cloning and sequencing. Ann. Surg. 252, 90–98, https://doi.org/10.1097/SLA.0b013e3181e3dc8b (2010).
Lepage, P. et al. Twin study indicates loss of interaction between microbiota and mucosa of patients with ulcerative colitis. Gastroenterology 141, 227–236, https://doi.org/10.1053/j.gastro.2011.04.011 (2011).
Halfvarson, J. et al. Dynamics of the human gut microbiome in inflammatory bowel disease. Nature microbiology 2, 17004, https://doi.org/10.1038/nmicrobiol.2017.4 (2017).
Frank, D. N. et al. Molecular-phylogenetic characterization of microbial community imbalances in human inflammatory bowel diseases. Proc. Natl. Acad. Sci. USA 104, 13780–13785, https://doi.org/10.1073/pnas.0706625104 (2007).
Willing, B. P. et al. A pyrosequencing study in twins shows that gastrointestinal microbial profiles vary with inflammatory bowel disease phenotypes. Gastroenterology 139, 1844–1854 e1841, https://doi.org/10.1053/j.gastro.2010.08.049 (2010).
Pascal, V. et al. A microbial signature for Crohn’s disease. Gut 66, 813–822, https://doi.org/10.1136/gutjnl-2016-313235 (2017).
Jahani-Sherafat, S. Role of gut microbiota in the pathogenesis of colorectal cancer; a review article. 11, 101–109 (2018).
Magro, F. et al. European consensus on the histopathology of inflammatory bowel disease. J. Crohns Colitis 7, 827–851, https://doi.org/10.1016/j.crohns.2013.06.001 (2013).
Swerdlow, S. H. et al. The 2016 revision of the World Health Organization classification of lymphoid neoplasms. Blood 127, 2375–2390, https://doi.org/10.1182/blood-2016-01-643569 (2016).
Delabie, J. et al. Enteropathy-associated T-cell lymphoma: clinical and histological findings from the international peripheral T-cell lymphoma project. Blood 118, 148–155, https://doi.org/10.1182/blood-2011-02-335216 (2011).
Ramadan, Z. et al. Fecal microbiota of cats with naturally occurring chronic diarrhea assessed using 16S rRNA gene 454-pyrosequencing before and after dietary treatment. J. Vet. Intern. Med. 28, 59–65 (2014).
Suchodolski, J. S. Diagnosis and interpretation of intestinal dysbiosis in dogs and cats. Vet. J. 215, 30–37, https://doi.org/10.1016/j.tvjl.2016.04.011 (2016).
Jahani-Sherafat, S., Alebouyeh, M., Moghim, S., Ahmadi Amoli, H. & Ghasemian-Safaei, H. Role of gut microbiota in the pathogenesis of colorectal cancer; a review article. Gastroenterology and Hepatology From Bed to Bench 11, 101–109 (2018).
Omori, M. et al. Fecal microbiome in dogs with inflammatory bowel disease and intestinal lymphoma. J. Vet. Med. Sci. 79, 1840–1847, https://doi.org/10.1292/jvms.17-0045 (2017).
Roediger, W. E. Role of anaerobic bacteria in the metabolic welfare of the colonic mucosa in man. Gut 21, 793–798 (1980).
Ashida, H., Ogawa, M., Kim, M., Mimuro, H. & Sasakawa, C. Bacteria and host interactions in the gut epithelial barrier. Nat. Chem. Biol. 8, 36–45, https://doi.org/10.1038/nchembio.741 (2011).
Canani, R. B. et al. Potential beneficial effects of butyrate in intestinal and extraintestinal diseases. World J. Gastroenterol. 17, 1519–1528, https://doi.org/10.3748/wjg.v17.i12.1519 (2011).
Segain, J. P. et al. Butyrate inhibits inflammatory responses through NFkappaB inhibition: implications for Crohn’s disease. Gut 47, 397–403 (2000).
Nagao-Kitamoto, H. & Kamada, N. Host-microbial Cross-talk in Inflammatory Bowel Disease. Immune Netw. 17, 1–12, https://doi.org/10.4110/in.2017.17.1.1 (2017).
Mondot, S. et al. Structural robustness of the gut mucosal microbiota is associated with Crohn’s disease remission after surgery. Gut 65, 954–962, https://doi.org/10.1136/gutjnl-2015-309184 (2016).
Mylonaki, M., Rayment, N. B., Rampton, D. S., Hudspith, B. N. & Brostoff, J. Molecular characterization of rectal mucosa-associated bacterial flora in inflammatory bowel disease. Inflamm. Bowel Dis. 11, 481–487 (2005).
O’Neill, I., Schofield, Z. & Hall, L. J. Exploring the role of the microbiota member Bifidobacteriumin modulating immune-linked diseases. Emerging Topics in Life Sciences 1, 333–349 (2017).
Bergmann, K. R. et al. Bifidobacteria stabilize claudins at tight junctions and prevent intestinal barrier dysfunction in mouse necrotizing enterocolitis. Am. J. Pathol. 182, 1595–1606, https://doi.org/10.1016/j.ajpath.2013.01.013 (2013).
Hsieh, C. Y. et al. Strengthening of the intestinal epithelial tight junction by Bifidobacterium bifidum. Physiological reports, 3, https://doi.org/10.14814/phy2.12327 (2015).
Jeon, S. G. et al. Probiotic Bifidobacterium breve induces IL-10-producing Tr1 cells in the colon. PLoS Pathog. 8, e1002714, https://doi.org/10.1371/journal.ppat.1002714 (2012).
Zheng, B. et al. Bifidobacterium breve attenuates murine dextran sodium sulfate-induced colitis and increases regulatory T cell responses. PLoS One 9, e95441, https://doi.org/10.1371/journal.pone.0095441 (2014).
Zeuthen, L. H., Fink, L. N. & Frokiaer, H. Toll-like receptor 2 and nucleotide-binding oligomerization domain-2 play divergent roles in the recognition of gut-derived lactobacilli and bifidobacteria in dendritic cells. Immunology 124, 489–502, https://doi.org/10.1111/j.1365-2567.2007.02800.x (2008).
Saez-Lara, M. J., Gomez-Llorente, C., Plaza-Diaz, J. & Gil, A. The role of probiotic lactic acid bacteria and bifidobacteria in the prevention and treatment of inflammatory bowel disease and other related diseases: a systematic review of randomized human clinical trials. Biomed Res Int 2015, 505878, https://doi.org/10.1155/2015/505878 (2015).
Young, V. B. & Schmidt, T. M. Antibiotic-associated diarrhea accompanied by large-scale alterations in the composition of the fecal microbiota. J. Clin. Microbiol. 42, 1203–1206 (2004).
Lupp, C. et al. Host-mediated inflammation disrupts the intestinal microbiota and promotes the overgrowth of Enterobacteriaceae. Cell Host Microbe 2, 119–129, https://doi.org/10.1016/j.chom.2007.06.010 (2007).
Tang, M. S. et al. Inferred metagenomic comparison of mucosal and fecal microbiota from individuals undergoing routine screening colonoscopy reveals similar differences observed during active inflammation. Gut Microbes 6, 48–56, https://doi.org/10.1080/19490976.2014.1000080 (2015).
Albenberg, L. et al. Correlation between intraluminal oxygen gradient and radial partitioning of intestinal microbiota. Gastroenterology 147, 1055–1063.e1058, https://doi.org/10.1053/j.gastro.2014.07.020 (2014).
Xenoulis, P. G. et al. Molecular-phylogenetic characterization of microbial communities imbalances in the small intestine of dogs with inflammatory bowel disease. FEMS Microbiol. Ecol. 66, 579–589, https://doi.org/10.1111/j.1574-6941.2008.00556.x (2008).
Santoru, M. L. et al. Cross sectional evaluation of the gut-microbiome metabolome axis in an Italian cohort of IBD patients. Sci. Rep. 7, 9523, https://doi.org/10.1038/s41598-017-10034-5 (2017).
Riordan, A. M. et al. Treatment of active Crohn’s disease by exclusion diet: East Anglian multicentre controlled trial. Lancet 342, 1131–1134, https://doi.org/10.1016/0140-6736(93)92121-9 (1993).
Bartel, G. et al. Ingested matter affects intestinal lesions in Crohn’s disease. Inflamm. Bowel Dis. 14, 374–382, https://doi.org/10.1002/ibd.20295 (2008).
Jones, V. A. et al. Crohn’s disease: maintenance of remission by diet. Lancet 2, 177–180, https://doi.org/10.1016/s0140-6736(85)91497-7 (1985).
Chiba, M. et al. Lifestyle-related disease in Crohn’s disease: relapse prevention by a semi-vegetarian diet. World J. Gastroenterol. 16, 2484–2495, https://doi.org/10.3748/wjg.v16.i20.2484 (2010).
Voelkl, B., Vogt, L., Sena, E. S. & Wurbel, H. Reproducibility of preclinical animal research improves with heterogeneity of study samples. PLoS Biol. 16, e2003693, https://doi.org/10.1371/journal.pbio.2003693 (2018).
Karp, N. A. J. P. B. Reproducible preclinical research—Is embracing variability the answer? 16, e2005413 (2018).
Nahon, S. et al. Evolution in clinical presentation of inflammatory bowel disease over time at diagnosis: a multicenter cohort study. Eur. J. Gastroenterol. Hepatol. 30, 1125–1129, https://doi.org/10.1097/meg.0000000000001201 (2018).
Allenspach, K., Wieland, B., Grone, A. & Gaschen, F. Chronic enteropathies in dogs: evaluation of risk factors for negative outcome. J. Vet. Intern. Med. 21, 700–708 (2007).
Jergens, A. E. et al. A scoring index for disease activity in canine inflammatory bowel disease. J. Vet. Intern. Med. 17, 291–297 (2003).
Donaldson, G. P., Lee, S. M. & Mazmanian, S. K. Gut biogeography of the bacterial microbiota. Nat. Rev. Microbiol. 14, 20–32, https://doi.org/10.1038/nrmicro3552 (2016).
Garraway, K. et al. Relationship of the mucosal microbiota to gastrointestinal inflammation and small cell intestinal lymphoma in cats. J. Vet. Intern. Med. 32, 1692–1702, https://doi.org/10.1111/jvim.15291 (2018).
Willard, M. D. et al. Interobserver variation among histopathologic evaluations of intestinal tissues from dogs and cats. J. Am. Vet. Med. Assoc. 220, 1177–1182 (2002).
Marsilio, S., Ackermann, M. R., Lidbury, J. A., Suchodolski, J. S. & Steiner, J. M. Results of histopathology, immunohistochemistry, and molecular clonality testing of small intestinal biopsy specimens from clinically healthy client-owned cats. J. Vet. Intern. Med. 33, 551–558, https://doi.org/10.1111/jvim.15455 (2019).
Bjornvad, C. R. et al. Evaluation of a nine-point body condition scoring system in physically inactive pet cats. Am. J. Vet. Res. 72, 433–437, https://doi.org/10.2460/ajvr.72.4.433 (2011).
Langerak, A. W. et al. EuroClonality/BIOMED-2 guidelines for interpretation and reporting of Ig/TCR clonality testing in suspected lymphoproliferations. Leukemia 26, 2159–2171, https://doi.org/10.1038/leu.2012.246 (2012).
Langerak, A. W. Toward Standardization of Clonality Testing in Veterinary Medicine. Vet. Pathol. 53, 705–706, https://doi.org/10.1177/0300985816638724 (2016).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7, 335–336, https://doi.org/10.1038/nmeth.f.303 (2010).
Benjamini, Y. & Hochberg, Y. Controlling the False Discovery Rate - a Practical and Powerful Approach to Multiple Testing. Journal of the Royal Statistical Society Series B-Methodological 57, 289–300 (1995).
Acknowledgements
The study was funded by the 2016 Miller’s Trust Fund of the WINN Feline Foundation. The authors thank SoYoung Park and Jacqueline Sedillo for their help with handling samples and sorting data.
Author information
Authors and Affiliations
Contributions
S.M. examined a subset of cats, collected and aliquoted the feces, performed data collection and analysis, prepared all figures and wrote the manuscript. R.P. performed data analysis, assisted with preparation of Fig. 2. A, as well as the outline of all other figures, and gave input into manuscript writing and review. B.S. performed data analysis and prepared Table 2 and Supplementary Table 2. B.C. and S.H. examined a subset of cats, collected feces, and gave input into manuscript writing and review. M.R.A. and J.S.E. reviewed the tissue biopsy sections and assisted in the disease classification of the cases. J.A.L., J.M.S., J.S.S. assisted with the data analysis and gave input into manuscript writing and review.
Corresponding author
Ethics declarations
Competing interests
Drs Marsilio, Pilla, Sarawichitr, Lidbury, Suchodolski, and Steiner are employed by the Gastrointestinal Laboratory at Texas A&M University, which offers laboratory tests, including histopathology services, on a fee-for-service basis. Dr. Ackermann is a consultant of the Gastrointestinal Laboratory at Texas A&M University. Dr. Estep is employed by Texas Veterinary Pathology, LLC., which offers tissue histopathology on a fee-for-service basis. Drs. Chow and Hill have nothing to disclose.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Marsilio, S., Pilla, R., Sarawichitr, B. et al. Characterization of the fecal microbiome in cats with inflammatory bowel disease or alimentary small cell lymphoma. Sci Rep 9, 19208 (2019). https://doi.org/10.1038/s41598-019-55691-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-55691-w
This article is cited by
-
Unbiased serum metabolomic analysis in cats with naturally occurring chronic enteropathies before and after medical intervention
Scientific Reports (2024)
-
Faecal microbiota and fatty acids in feline chronic enteropathy
BMC Veterinary Research (2023)
-
The effect of a hydrolyzed protein diet on the fecal microbiota in cats with chronic enteropathy
Scientific Reports (2022)
-
GWAS-associated bacteria and their metabolites appear to be causally related to the development of inflammatory bowel disease
European Journal of Clinical Nutrition (2022)
-
Untargeted metabolomic analysis in cats with naturally occurring inflammatory bowel disease and alimentary small cell lymphoma
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.