Abstract
Microbial communities are pivotal in the biodegradation of xenobiotics including pesticides. In the case of atrazine, multiple studies have shown that its degradation involved a consortia rather than a single species, but little is known about how interdependency between the species composing the consortium is set up. The Black Queen Hypothesis (BQH) formalized theoretically the conditions leading to the evolution of dependency between species: members of the community called ‘helpers’ provide publicly common goods obtained from the costly degradation of a compound, while others called ‘beneficiaries’ take advantage of the public goods, but lose access to the primary resource through adaptive degrading gene loss. Here, we test whether liquid media supplemented with the herbicide atrazine could support coexistence of bacterial species through BQH mechanisms. We observed the establishment of dependencies between species through atrazine degrading gene loss. Labour sharing between members of the consortium led to coexistence of multiple species on a single resource and improved atrazine degradation potential. Until now, pesticide degradation has not been approached from an evolutionary perspective under the BQH framework. We provide here an evolutionary explanation that might invite researchers to consider microbial consortia, rather than single isolated species, as an optimal strategy for isolation of xenobiotics degraders.
Similar content being viewed by others
Introduction
Microorganisms in nature usually co-exist as communities whose complexity is under the influence of local environmental conditions and interactions between individuals. Spatially structured environments, such as biofilms, are more prone to support coexistence of multiple species consuming the same resource because access to the resource and its metabolic by-products is conditioned by the structure of the environment1,2. Interactions of various natures between bacterial species that stabilize the community diversity can then arise and persist, but are most likely to disappear when the structure is disrupted3,4. In mixed liquid cultures, competition for a single resource is supposed to lead to competitive exclusion5, although very recent research has shown that cross-feeding may stabilize coexistence on a single carbon source6. If the breaking down of the resource includes both private and public goods, natural selection could favor the emergence of dependency between species, and therefore coexistence, through adaptive gene loss according to the Black Queen Hypothesis (BQH)7,8.
Morris et al. formulated three key characteristics for a function to follow the BQH. The function allowing consumption of the resource must be i) costly, ii) essential and iii) leaky. Also, Estrela et al.9 and Oliveira et al.10 recently specified the conditions that may favor the emergence of interdependency, which include low privatization and low cost of secretion. Atrazine is one of the most heavily applied herbicides worldwide which is relatively persistent and mobile in soil, and whose degradation products can be traced in soils decades after application11,12. Atrazine can be mineralized by microorganisms and is a source of nitrogen13,14,15,16. Its entire biodegradation pathway is known (Fig. 1) and can be split into two parts: the upper part consisting in the dechlorination of atrazine and the removal of the two lateral chains from the s-triazinic cycle, followed by the lower part consisting in the complete degradation of cyanuric acid. Genes (atz and trz families) involved in its mineralization are most often located on plasmids or on catabolic cassettes delimited by insertion sequences17,18. Because of the cost of maintaining plasmids, atrazine genes can be easily lost after a small number of generations in culture media without atrazine19. However, in nitrogen limited environments, atrazine degrading capacities must be conserved because mineralization of atrazine leads to the essential delivery of nitrogen20. Also, metabolites, such as desisopropylamine, aminoethanol and desethylamine, are produced and released in the environment during atrazine degradation21. Consequently, atrazine biodegradation is (i) conferred by genes costly to maintain, (ii) essential under N-limited conditions because it provides nitrogen supply, and (iii) benefits to other members of the community because metabolites leak in the environment. Atrazine biodegradation is therefore a good candidate function to test the BQH predictions, and in particular the emergence of dependency between species in a spatially unstructured environment.
Atrazine metabolic degradation pathway and strains used. (a) Intermediary products of the upper and lower parts of the metabolic degradation of atrazine, as well as genes associated with the different reactions are indicated. (b) The four ancestral strains, as well as their corresponding atrazine degrading gene repertoire are indicated. Genes encoding enzymes involved in the formation of N-containing by-products are coloured in blue while genes encoding enzymes involved in the formation of C-containing by-products are contoured.
Chelatobacter sp. SR3822, Pseudomonas sp. ADPe19, Arthrobacter sp. TES23 and Variovorax sp. 38R24 are all originally soil bacteria isolated from arable soils exposed to atrazine. They are all able to at least partly degrade either atrazine or its intermediary products (Fig. 1), and represent four different genetic repertoires for atrazine degradation. They have all been modified to resist to a different combination of two antibiotics in order to evaluate their frequency in mixed culture on double-antibiotics plates. Based on their genetic repertoire for atrazine degradation, it is quite straightforward to predict that in a liquid culture medium with atrazine as the unique nitrogen source, Arthrobacter sp. TES and Pseudomonas sp. ADPe must coexist because Arthrobacter sp. TES will degrade atrazine to supply its nitrogen needs and produce cyanuric acid that will be used by Pseudomonas sp. ADPe as its nitrogen reservoir. However, predicting what would happen in a more complex scenario where multiple competing strains evolve in environments containing various nitrogen sources is not trivial. Here, we experimentally questioned whether spatially unstructured atrazine-containing media could support coexistence of multiple atrazine degrading species through BQH mechanisms.
We propagated for ~100 generations four-species consortia, in triplicate, in seven minimal media supplemented with citrate in excess, as the carbon source, and either one among three nitrogen sources: atrazine, cyanuric acid: the metabolic product of the upper part of the atrazine degradation pathway, or ammonium sulfate; or 2-way and 3-way combinations of the above mentioned nitrogen sources, keeping the N molarity constant. We found that out of the seven media, all species coexisted in the four ones containing atrazine, whereas Arthrobacter sp. TES went extinct in the three atrazine-free media (Fig. 2a). The extinction of Arthrobacter sp. TES in cyanuric acid supplemented media was expected as it does not possess the metabolic pathway to degrade cyanuric acid, however its extinction also in ammonium sulfatesupplemented media, is more surprising since it can use ammonia as sole nitrogen source. Intriguingly, the equilibrium frequencies of the three other species, besides Arthrobacter sp. TES, are quite similar after ~100 generations in six out of the seven media, with Pseudomonas sp. ADPe dominant at ~107 CFU.mL−1 and the two remaining species at ~106 CFU.mL−1, the exception being the ammonium sulfate – cyanuric acid supplemented medium where Variovorax sp. 38R appeared to be dominant at ~107 CFU.mL−1.
Community composition of the ancestral and evolved consortia across the seven selection media. (a) Species composition of the ancestral and evolved consortia across the seven selection media. Abundances are expressed in log10 CFU.mL-1. Black, red, green and blue bars correspond respectively to Variovorax sp. 38R, Pseudomonas sp. ADPe, Chelatobacter sp. SR38 and Arthrobacter sp. TES. Atz, NH4 and Cya respectively stands for nitrogen source used either atrazine, ammonium or cyanuric acid. (b) atrazine-degrading genetic potential of the ancestral and evolved consortia across the seven selection medium evaluated via multiplex PCRs. Presence/absence of the corresponding genes is indicated with the presence/absence of the corresponding coloured or hatched rectangle on the model scheme.
We then assessed using multiplex PCRs the presence of the atrazine degradation genes in the evolved consortia across the seven different media. We found that in the four media containing atrazine, all initially present atz or trz genes were retained in the evolved consortia, preluding the conservation of the atrazine mineralisation potential in these environments (Fig. 2b). In the atrazine-free media, only atzA, responsible for the dechlorination of atrazine, a step that releases chlorine but does not provide any nitrogen input to cells, and atzD, responsible for the first step of the degradation of cyanuric acid, were conserved in the three environments while atzC and trzD were specifically kept in the ammonium sulfate – cyanuric acid supplemented medium. Since in that specific medium, atzC does not confer any advantage to its carrier, Chelatobacter sp. SR38, it has likely been carried along because of its colocalization with trzD on the same plasmid25, responsible for the degradation of cyanuric acid. Rapid evolution, probably through the loss of atz genes containing plasmids, therefore led to the loss of the atrazine degradation function in atrazine-free environments, confirming the genetic load of maintaining atrazine genes in atrazine uncontaminated environments.
We then evaluated the atrazine mineralization potential, using 14C-atrazine either labelled on the ethylamino chain or on the s-triazinic cycle, of the consortia that have evolved in the four atrazine containing environments (Fig. 3) and compared them to the ancestral ones. Interestingly, we observed a significant increase of the mineralization potential of the ethylamino chain in evolved consortia compared to ancestors in all but the medium which contained all three nitrogen sources. The observed gain is however much stronger in the atrazine only supplemented medium (from 48%; CI95% = [41.9–54.1] to 71.4%; CI95% = [69.4–73.4]). When evaluating the mineralization potential of the s-triazinic cycle, results are quite contrasted with a significant increase of the mineralization potential in the atrazine only supplemented medium (from 48.3%; CI95% = [36.5–60] to 86.9%; CI95% = [81.6–92.2]), and significant decreases or no change in media supplemented with combinations of two or all three nitrogen sources. Altogether, these results indicate that the upper part of the atrazine degradation pathway (atzA, trzN, atzB and atzC) has been improved in the evolved consortia, presumably leading to increased intermediary metabolite production, such as aminoethanol, ethylamine or hypoxanthine21. While the upper pathway is constitutively expressed, studies have shown that the lower pathway was repressed by the presence of ammonium in the environment26. It was therefore quite likely that the stronger accumulation of intermediary metabolites and their consumption by consortium members led to an increased concentration of ammonium in the environment explaining at least partly the decreased mineralization potential of the s-triazinic cycle in evolved consortia compared to ancestors.
Atrazine mineralization potential of the ancestral and evolved consortia. The mineralization potentials of the ethylamino chain (a) and the s-triazinic cycle (b) were evaluated with 14C-labelled atrazine across the four atrazine-containing selection media. 14C-labelled atoms are indicated in red on the atrazine molecule. Atz, NH4 and Cya respectively stands for atrazine, ammonium and cyanuric acid. Significant differences between ancestral (dark grey bars) and evolved (light grey bars) consortia performances are indicated with a star (Tukey test; p < 0.05).
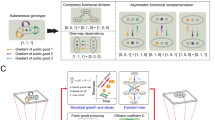
The impressive boost of atrazine mineralization potential observed in the medium supplemented only with atrazine led us to focus on phenotypic and genotypic changes that occurred in this specific environment. We therefore isolated members of the evolved consortia, evaluated individually their genetic repertoire using multiplex PCRs, and assessed their corresponding growth characteristics using BioscreenTM assays. We found that only Arthrobacter sp. TES was able to grow in isolated culture, while the three other members of the evolved consortia depend on Arthrobacter sp. TES to grow in this medium (Fig. 4a). This acquired dependency for Chelatobacter sp. SR38 is explained by loss of part of its atz genes repertoire, namely the upper pathway genes atzA and atzB (Fig. 4b). We then wanted to characterize the interactions between members of the evolved consortia. We therefore reconstructed all possible duos and trios, and compared growth performances of each member to their performance in the evolved four-species consortia and in monoculture over 3.5 days (Fig. 4c). A relatively simple and straightforward interaction scheme can be drawn from these results (Fig. 4d): Variovorax sp. 38R, Chelatobacter sp. SR38 and Pseudomonas sp. ADPe growths are clearly supported by Arthrobacter sp. TES. However, Arthrobacter sp. TES is penalized in duos or trios with Chelatobacter sp. SR38 and Pseudomonas sp. ADPe, in the absence of Variovorax sp. 38R. Interestingly, Variovorax sp. 38R by itself has no positive effect on Arthrobacter sp. TES, but it counteracts the negative impacts of the two other consortium members on Arthrobacter sp. TES. Therefore, coexistence of the four members might be favoured, even though two of the three direct beneficiaries exert antagonistic relationships against the public goods provider, because the third one acts as a buffering intermediary.
Interspecies dependency for atrazine degradation. (a) Axenical growth performances on atrazine supplemented medium of the ancestral (grey bars) strains and strains isolated from evolved community (black bars) estimated as the final OD600nm reached after a 3.5 days cycle Strains are abbreviated by their strain names, for instance Variovorax sp. 38R is named 38R. Strains were considered as non-growing if their final OD600nm was <0.1 Arbitrary Units (AU). Significant differences between ancestral (dark grey bars) and evolved (light grey bars) consortia performances are indicated with a star (Tukey test; p < 0.05). (b) Atrazine-degrading genetic repertoire of the community-evolved strains and corresponding metabolic reactions. (c) Evaluating interactions between the four species evolved in the atrazine only supplemented medium. Growth performances of evolved Pseudomonas sp. ADPe (a), Chelatobacter sp. SR38 (b), Variovorax sp. 38R (c) and Arthrobacter sp. TES (d) at the end of a 3.5 days cycle in monoculture, as well as in reconstructed duos, trios and in the 4-species consortium were evaluated using CFU plating on TY medium supplemented with the adequate antibiotics combinations. Bars represent mean values with s.e.m (n = 3). Non-overlapping 95% confidence interval were considered as significant (p < 0,05). (d) Interactions scheme drawn from mono- and co-cultures of the four community-evolved strains in the atrazine only supplemented medium. Interactions types were defined based on growth performances in co-cultures experiments. Positive (+) or negative (−) interactions are represented by arrows. Arrows point towards the impacted strain. The dotted arrow expresses the conditional positive effect of Variovorax sp. 38R on Arthrobacter sp. TES in the presence of the two other strains.
Here, we have witnessed the establishment of dependencies in a four species artificial consortium after evolution in an unstructured, atrazine only supplemented medium during ~100 generations. According to the BQH, dependency was set up via adaptive gene loss, and one specific member of the consortium, Arthrobacter sp. TES ensuring the costly part of the function, became the provider of public goods to the three other beneficiaries, Variovorax sp. 38R, Chelatobacter sp. SR38 and Pseudomonas sp. ADPe. In many cases, pesticide degradation in nature is thought to occur through the involvement of microbial consortia, rather than being the corollary of a single species14. Our study provides strong experimental evidences, using the herbicide atrazine as a case study, that the division of labour, sensu Giri et al.27, between populations in microbial communities could also be considered for the biodegradation of xenobiotics. Also, we anticipate that future research should focus on the bioaugmentation with stabilized and tightly structured microbial degrading consortia as an effective solution for in situ bioremediation of sites polluted with recalcitrant compounds.
Methods
Experimental evolution
Four strains with variable atrazine degrading abilities (Pseudomonas sp. ADPe, Chelatobacter sp. SR38, Arthrobacter sp. TES and Variovorax sp. 38R, Fig. 1) were cultivated in seven media in four species consortia (n = 3) constituting “community” evolution lines. Mineral salt (MS) media (K2HPO4 9.2 mM, KH2PO4 3.0 mM, CaCl2 0.2 mM, MgSO4-7H2O 0.8 mM, NaCl 1.7 mM, H3BO3 32.3 µM, FeSO4-6 H2O 19.2 µM, MnSO4-H2O 10.6 µM, NaMo 2.1 µM, ZnSO4 1.2 µM, CuSO4 0.63 µM, biotin 0.4 µM, thiamin 0.15 µM,) containing citrate as carbon source (5 mM) were supplemented with varied nitrogen sources combinations (atrazine, cyanuric acid, (NH4)2SO4, or 2 and 3 sources combinations) with a final nitrogen equimolarity of 1.4 mM (7 media). The evolution experiment was realized in 96-wells 2 mL Deepwell microplate at 28 °C without agitation during 23 growing cycles of 3.5 days each. For each new cycle, 35 µL of previous culture was used as inoculum for the next cycle into 700 µL fresh medium (1/20 dilution, (log2(20) x 23 cycles ~ 100 generations)). An aliquot of each cycle culture was mixed with glycerol (30% final concentration) and preserved at −80 °C.
Species composition
We were able to easily identify the four strains in the ancestor and evolved “community” lines because each of them was double-resistant to a different combination of two of the following antibiotics: rifampicin 100 mg/L, kanamycin 50 mg/L, spectinomycin 100 mg/l, streptomycin 100 mg/L. To do so, each bacterial strain was grown in MS medium containing either atrazine or cyanuric acid as the nitrogen source and then spread on solid rich medium (TY) containing the antibiotic for which it was to be resistant. Resistant growing colonies were collected and cultivated in their specific liquid MS medium to ensure that they have kept their metabolic ability towards atrazine. This procedure was repeated to confer a second resistance to each strain. Pseudomonas sp. ADPe is Strep+/Rif+, Chelatobacter sp. SR38 is Kan+/Strep+, Arthrobacter sp. TES is Spec+/Rif+ and Variovorax sp. 38R is Spec+/Kan+.
Ten-fold successive dilutions of T1 and T23 “community” lines were done and inoculated onto LB agar plates supplemented with different antibiotic combinations. After 3-days of growth at 28 °C, CFU were counted across 3 dilution levels.
Atrazine-degrading genetic repertoire
Multiplex PCR targeting atzA/atzD/trzD and atzC/trzN/atzB genes were used to determine the atrazine degradation repertoire of each strain of the ancestral and evolved “community” lines. 3 or 4 isolated colonies from the ancestral and evolved “community” lines were resuspended in water and served as DNA template for PCR reactions. PCR reactions were conducted using the following primers [atzA (5′-TGA AGC GTC CAC ATT ACC-3′, 5′-CCA TGT GAA CCA GAT CCT-3′), atzD (5′-GGG TCT CGA GGA TTT GAT TG-3′, 5′-TCC CAC CTG ACA TCA CAA AC-3′) trzD5′-CCT CGC GTT CAA GGT CTA CT-3′, 5′-TCG AAG CGA TAA CTG CAT TG-3′] or [trzN (5′-CAC CAG CAC CTG TAC GAA GG-3′, 5′-GAT TCG AAC CAT TCC AAA CG-3′), atzB (5′-CAC CAC TGT GCT GTG GTA GA-3′, 5′-AGG GTG TTG AGG TGG TGA AC-3′), atzC (5′-GTACCATATCACCGTTGCCA-3′, 5′-GCTCACATGCAGGTACTCCA-3′)] at a final concentration of 10 µM each, the temperature of primers annealing being of 57 °C and the elongation time of 1 minute.
Atrazine mineralisation activity
Ancestors and “community” lines that have evolved on the four media containing atrazine were inoculated at ~0.001 OD600nm in 200 µL fresh medium supplemented with 4000 dpm final of 14C atrazine labelled either on the ethylamino chain or on the s-triazine cycle in triplicate. Cultures were then placed at 28 °C. A whatman® 3 mm Chr paper soaked with barite (saturated solution 0.37 M) was placed and sealed on the top of the 96-wells plates and replaced periodically during 3.5 days. Dried papers were fluorography printed on Storage PhosphorScreen (Molecular Dynamics®) for 2 days. Screens were scanned by a phosphorimager (Storm 860 Molecular Imager). Indirect reading of released radioactivity intensity was done by ImageQuant 5.2 software, and referred to a standard curve. Mineralisation percentages of lines were determinate with respect to initial radioactivity dosage realized on 200 µL of each culture medium with a Beckman® liquid scintillation counter. Significant differences between ancetral and evolved mineralization percentages were assessed using Tukey’s test (p < 0.05).
Population dynamics characteristics of the ancestral and evolved line
The population dynamics of ancestral and evolved “community” lines were measured using BioscreenTM. All lines were inoculated at ~0.001 OD600nm in 400 µL of their evolution medium (n = 3 for each replicate of each line) in Bioscreen TM plates. Plates were incubated 3.5 days at 28 °C and the evolution of OD600nm was monitiored every 10 minutes after a 10-seconds gentle shake. Population dynamics characteristics of each strain that has evolved in “community” lines was also measured the same way starting from 10 CFU isolated from the evolved “community” lines as inoculums and inoculated at ~0,001 OD600nm. Those data were used to determine the maximal OD600nm. Cultures that did not reach 0.1 OD600nm at the end of the 3.5 days were considered as non-growing. Significant differences between ancetral and evolved mineralization percentages were assessed using Tukey’s test (p < 0.05).
Evaluating interactions between the four evolved species in the atrazine only supplemented medium
For each strain, ten CFU isolated from each community-evolved lines (n = 3) in MS-Citrate Atrazine were used as inoculum pools. 4-species consortia, as well as all possible duos and trios were reconstructed and compared to monocultures. Mixtures and monocultures were inoculated at ~0,001 OD600nm and final population densities, after a 3.5 days cycle, were measured for each strain by CFU plating on TY medium supplemented with the appropriate antibiotics combinations. Non-overlapping 95% confidence interval were considered as significant (p < 0,05).
References
Chao, L. & Levin, B. R. Structured habitats and the evolution of anticompetitor toxins in bacteria. Proc. Natl. Acad. Sci. USA 78, 6324–6328 (1981).
Travisano, M. & Velicer, G. J. Strategies of microbial cheater control. Trends Microbiol. 12, 72–78 (2004).
Kerr, B., Neuhauser, C., Bohannan, B. J. M. & Dean, A. M. Local migration promotes competitive restraint in a host-pathogen ‘tragedy of the commons’. Nature. 442, 75–78 (2006).
Hassell, M. P., Comins, H. N. & May, R. M. Species coexistence and self-organizing spatial dynamics. Nature. 370, 290–292 (1994).
Hardin, G. The competitive exclusion principle. Science. 131, 1292–1297 (1960).
Godford, J. E. et al. Emergent simplicity in microbial community assembly. Science. 361, 469–474 (2018).
Morris, J. J., Lenski, R. E. & Zinser, E. R. The Black Queen Hypothesis: evolution of dependencies through adaptive gene loss. mBio. 3, e00036–00012 (2012).
Mas, A., Jamshidi, S., Lagadeuc, Y., Eveillard, D. & Vandenkoornhuyse, P. Beyond the Black Queen Hypothesis. The ISME J. 10, 2085–2091 (2016).
Estrela, S., Morris, J. J. & Kerr, B. Private benefits and metabolic conflicts shape the emergence of microbial interdependencies. Environ. Microbiol. 18, 1415–1427 (2016).
Oliveira, N. M., Niehus, R. & Foster, K. R. Evolutionary limits to cooperation in microbial communities. Proc. Natl. Acad. Sci. USA 111, 17941–17946 (2014).
Jablonowski, N. D., Köppchen, S., Hofmann, D., Schäffer, A. & Burauel, P. Persistence of 14C-labelled atrazine and its residues in a field lysimeter soil after 22 years. Environ Pollut. 157, 2126–2131 (2009).
Udikovic-Kolic, N. et al. Genetic potential, diversity and activity of an atrazine-degrading community enriched from a herbicide factory effluent. J. Appl. Microbiol. 105, 1334–134 (2008).
de Souza, M. L. et al. Molecular basis of a bacterial consortium: interspecies catabolism of atrazine. Appl. Environ. Microbiol. 64, 178–184 (1998).
Smith, D., Alvey, S. & Crowley, D. E. Cooperative catabolic pathways within an atrazine-degrading enrichment culture isolated from soil. FEMS Microbiol. Ecol. 53, 265–275 (2005).
Yang, C. et al. Atrazine degradation by a simple consortium of Klebsiella sp. A1 and Comamonas sp. A2 in nitrogen enriched medium. Biodegradation. 21, 97–105 (2010).
Chiai-Hernandez, A. C. et al. Long-term persitence of pesticides and TPs in archived agricultural soil samples and comparison with pesticide application. Environ. Sci. Technol. 51, 10642–10651 (2017).
Devers, M., El Azhari, N., Kolic, N. U. & Martin-Laurent, F. Detection and organization of atrazine-degrading genetic potential of seventeen bacterial isolates belonging to divergent taxa indicate a recent common origin of their catabolic functions. FEMS Microbiol. Lett. 273, 78–86 (2007).
Esquirol, L. et al. An unexpected vestigial protein complex reveals the evolutionary origins of an s-triazine catabolic enzyme. J. Biol. Chem. 293, 7880–7891 (2018).
Changey, F., Devers-Lamrani, M., Rouard, N. & Martin-Laurent, F. In vitro evolution of an atrazine-degrading population under cyanuric acid selection pressure: evidence for the selective loss of a 47 kb region on the plasmid ADP1 containing the atzA, B and C genes. Gene. 490, 18–25 (2011).
Govantes, F. et al. Regulation of the atrazine-degradative genes in Pseudomonas sp. ADP. FEMS Microbiol Lett. 310, 1–8 (2010).
Xu, X. et al. Modeling microbial communities from atrazine contaminated soils promotes the development of biostimulation solutions. The ISME J. 13, 494–508 (2019).
Rousseaux, S., Hartmann, A. & Soulas, G. Isolation and characterization of new Gram-negative and Gram-positive atrazine degrading bacteria from different French soils. FEMS Microbiol. Ecol. 36, 211–222 (2001).
El Sebaï, T., Devers-Lamrani, M., Changey, F., Rouard, N. & Martin-Laurent, F. Evidence of atrazine mineralization in a soil from the Nile Delta: Isolation of Arthrobacter sp. TES6, an atrazine-degrading strain. Int. Biodeter. Bioderg. 65, 1249–1255 (2011).
Devers, M., Henry, S., Hartmann, A. & Martin-Laurent, F. Horizontal gene transfer of atrazine-degrading genes (atz) from Agrobacterium tumefaciens St96-4 pADP1::Tn5 to bacteria of maize-cultivated soil. Pest. Manag. Sci. 61, 870–880 (2005).
Rousseaux, S., Soulas, G. & Hartmann, A. Plasmid localisation of atrazine-degrading genes in newly described Chelatobacter and Arthrobacter strains. FEMS Microbiol. Ecol. 41, 69–75 (2002).
Garcia-Gonzàlez, V. et al. Distinct roles for NtrC and GlnK in nitrogen regulation of the Pseudomonas sp. strain ADP cyanuric acid utilization operon. FEMS Microbiol. Lett. 300, 222–229 (2009).
Giri, S., Waschina, S., Kaleta, C. & Kost, C. Defining division of labor in microbial communities. J. Mol. Biol. In Press (2019).
Acknowledgements
We thank L. Philippot for comments and discussion. Grants from FP7-PEOPLE-2012-IAPP (Industry-Academia Partnerships and Pathways) Marie-Curie project ‘Love-to-Hate’ funded by the European Commission (Grant Agreement number 389 324349) supported this work.
Author information
Authors and Affiliations
Contributions
L.B.: laboratory experiments, data analysis, manuscript writing. M.D.: study conception, laboratory experiments, manuscript editing. N.R.: laboratory experiments. F.M.-L.: study conception, manuscript editing. A.S.: study conception, data analysis, manuscript writing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Billet, L., Devers, M., Rouard, N. et al. Labour sharing promotes coexistence in atrazine degrading bacterial communities. Sci Rep 9, 18363 (2019). https://doi.org/10.1038/s41598-019-54978-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-54978-2
This article is cited by
-
The interplay between microbial communities and soil properties
Nature Reviews Microbiology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.