Abstract
Vascular smooth muscle cell (VSMC) dysfunction is a hallmark of small vessel disease, a common cause of stroke and dementia. Two of the most frequently mutated genes in familial small vessel disease are HTRA1 and NOTCH3. The protease HTRA1 cleaves the NOTCH3 ligand JAG1 implying a mechanistic link between HTRA1 and Notch signaling. Here we report that HTRA1 is essential for VSMC differentiation into the contractile phenotype. Mechanistically, loss of HTRA1 increased JAG1 protein levels and NOTCH3 signaling activity in VSMC. In addition, the loss of HTRA1 enhanced TGFβ-SMAD2/3 signaling activity. Activation of either NOTCH3 or TGFβ signaling resulted in increased transcription of the HES and HEY transcriptional repressors and promoted the contractile VSMC phenotype. However, their combined over-activation led to an additive accumulation of HES and HEY proteins, which repressed the expression of contractile VSMC marker genes. As a result, VSMC adopted an immature phenotype with impaired arterial vasoconstriction in Htra1-deficient mice. These data demonstrate an essential role of HTRA1 in vascular maturation and homeostasis by controlling Notch and TGFβ signaling.
Similar content being viewed by others
Introduction
Familial small vessel disease is a major cause of stroke and dementia under the age of 60 with NOTCH3 and HTRA1 being two of the most frequently mutated genes1. In addition, several loss-of-function mutations in HTRA1 are causative for cerebral autosomal recessive arteriopathy with subcortical infarcts and leukoencephalopathy (CARASIL syndrome)2, which is similar to dominantly inherited CADASIL syndrome caused by neomorphic NOTCH3 mutations3. A common feature of these diseases is vascular smooth muscle cell (VSMC) dysfunction on small arterial blood vessels leading to episodes of impaired blood perfusion in certain brain regions.
Since VSMC are critical regulators to maintain vascular homeostasis they show high phenotypic plasticity, where contractile and synthetic VSMC represent the two ends of a spectrum with intermediate phenotypes, which have different morphologies and functions. While naïve VSMC display a synthetic phenotype and are unable to contract but important for maintenance, contractile VSMC control blood flow and pressure. During development, vascular remodeling and injury, synthetic VSMC secrete extracellular matrix proteins and exhibit higher growth rates and migratory activity than contractile VSMC4.
Notch signaling is a juxtacrine signaling mode, which controls numerous cell differentiation processes. The signal sending cell expresses Notch ligands of the Delta-like (DLL) and Jagged (JAG) families which activate Notch receptors on adjacent signal receiving cells. The interaction induces receptor cleavage and translocation of the Notch intracellular domain (ICD) to the nucleus, where it interacts with RBP-Jκ and promotes cell type-specific gene expression and induction of the HES and HEY genes. These encode basic helix-loop-helix (bHLH) transcription factors, which repress gene expression through either binding other bHLH factors or through interacting directly with DNA at promoter regions5. In muscle stem cells, HeyL interacts with Hes1 to bind DNA sites with high affinity causing anti-myogenic effects6. In VSMC, HES and HEY proteins can inhibit transcription of contractile VSMC marker proteins7,8. As such, the effect of Notch signaling on promoting the contractile VSMC phenotype can be counteracted by HES and HEY bHLH factors. This indicates that the outcome of Notch signaling activity is strictly dose-dependent.
Similar to the Notch pathway, TGFβ signaling has also been shown to promote VSCM differentiation9. Interestingly, TGFβ signaling can also activate HEY and HES gene expression in certain cell types10,11. Provided that this also occurs in VSMC, HTRA1 might function through controlling expression levels of the HES and HEY transcriptional repressors via Notch and TGFβ signaling.
Here we aimed at better understanding how the serine protease HTRA1 controls Notch and TGFβ signaling in VSMC and how this affects the VSCM phenotype. HTRA1 is strongly expressed in VSMC and endothelial cells12,13 and is known to cleave several intracellular14,15,16,17 and extracellular substrates13,18. Loss of HTRA1 leads to increased levels of TGFβ1 availability and TGFβ1 signaling, potentially caused by the ability of HTRA1 to cleave either pro-TGFβ1 or GFD62,13,19,20,21. Recently, we have shown that the Notch ligand JAG1 is a substrate for HTRA1. After cleavage of JAG1 by HTRA1 in the cytosol the remaining JAG1 protein was rapidly degraded22. NOTCH3 and JAG1 are both abundantly expressed on VSMC7,8. In arterial blood vessels, JAG1/NOTCH3 signaling is required for differentiation, maintenance and contractility of VSMC23,24,25,26,27, which is crucial for vasoconstriction and proper organ perfusion. Such blood vessel functions are impaired in familial small vessel disease. Thus, we hypothesized that HTRA1 functions not only to control TGFβ signaling but also to fine-tune NOTCH3 activity in VSMC by regulating the abundance of its ligand JAG1. As both signaling pathways are critically involved in controlling VSMC differentiation7,8,9,23,26,28,29, loss of HTRA1 may lead to impaired VSMC function and vessel contraction capacity.
Results
Loss of HTRA1 in VSMC increases NOTCH3 signaling
The similarities between CARASIL and CADASIL syndromes3, as well as our recent finding that HTRA1 cleaves the Notch ligand JAG122, prompted us to investigate the potential interplay between HTRA1 and NOTCH3 signaling. Therefore, HTRA1 was silenced in primary human umbilical artery SMC (HUASMC) using established siRNAs22 (Fig. 1a). We observed that silencing HTRA1 increased mRNA levels of the Notch target genes HES1 and HEYL (Fig. 1b). Higher Notch signaling activity was further evidenced by increased NOTCH3-ICD protein levels and increased JAG1 protein levels (Fig. 1c).
Increased Notch3 signaling activity in Htra1-silenced VSMC. (a) HTRA1 was silenced with siRNA. Representative Western blot of HUASMC protein lysates probed with HTRA1 antibody. (b) Quantitative real-time PCR analysis of Notch target gene transcripts in HUASMC after silencing HTRA1 (n = 3). (c) Representative Western blot of HUASMC protein lysates probed with anti-JAG1 and anti-NOTCH3-ICD and quantification of band intensities (n = 3). (d) Immunoblot of protein lysates derived from mesenteric arteries of HtrA1+/+ and HtrA1−/− mice probed with anti-Notch3-ICD and anti-JAG1 antibody and quantification of band intensities (n = 3). (e) Quantitative real-time PCR analysis of Notch target gene transcripts in mesenteric arteries derived from HtrA1+/+ and HtrA1−/− mice (n = 3). Statistical significance was determined by an unpaired student’s t-test (b–e). *p < 0.05. Bar graphs show mean values, error bars indicate SD.
We next isolated arteries from adult Htra1−/− mice to verify these data in an in vivo model. Compared to wild-type littermate controls, there was an increase in NOTCH3-ICD and JAG1 protein levels in isolated mesenteric resistance arteries from Htra1−/− mice (Fig. 1d), indicating augmented NOTCH3 signaling activity. Consistently, we observed a substantial increase in Hes1 and HeyL mRNA levels in mesenteric arteries isolated from Htra1−/− mice (Fig. 1e).
Loss of HTRA1 in VSMC increases TGFβ signaling
Several reports have shown increased TGFβ1 availability and TGFβ signaling upon loss of HTRA12,13,19,20. In HUASMC, silencing of HTRA1 led to an induction of TGFβ signaling activity as evidenced by elevated levels of phosphorylated SMAD2/3 proteins (Fig. 2a,b). In addition, Smad2/3 phosphorylation levels were higher in isolated arteries from Htra1−/− mice when compared to wild-type littermate controls (Fig. 2c,d). Furthermore, when co-cultured with 3T3 reporter cells, silencing of HTRA1 in HUASMC increased SMAD-dependent luciferase activity (Fig. 2e), indicating that loss of Htra1 in VSMC results in increased TGFβ signaling in adjacent cells.
Loss of HTRA1 in VSMC enforces TGFβ signaling and induces dual induction of HES and HEY transcriptional repressors synergistically with NOTCH3. (a) Immunoblot of phosphorylated SMAD2/3 protein expression after siRNA-mediated silencing of HTRA1 in HUASMC. (b) Graphical representation of fold change in phosphorylated SMAD2/3 protein level (n = 3). (c) Immunoblot of pSMAD2/3 protein level in mesenteric arteries obtained from HtrA1+/+ and HtrA1−/− mice. (d) Graphical representation of fold change in pSMAD2/3 protein level (n = 3). (e) Co-culture experiment with HUASMC and 3T3 TGFß reporter cells expressing a luciferase cassette with a SMAD responsive promotor (n = 3). (f) Western blot of HUASMC protein lysates probed with anti-pSMAD2/3 antibody after stimulation with TGFß1 (5 ng/ml). (g) Quantitative real-time PCR analysis of Notch target gene transcripts in HUASMC after TGFß1 stimulation, NOTCH3 stimulation via immobilized JAG1 ligands or combination of TGFß1 and NOTCH3 stimulation (n = 3). (h) Quantitative real-time PCR analysis of Notch target gene transcripts in control and HTRA1-silenced HUASMC after two hours of TGFß inhibition using 10 µM ALK5 inhibitor (ALK5i, CAS 446859-33-2, Merck) or NOTCH3 inhibition for 16 hours using 25 µM DAPT (Merck). The dashed line denotes gene expression level of control cells (sh-ctrl) treated with DMSO (n = 3). Statistical significance was determined by an unpaired student’s t-test (b,d,e) or by 1-way ANOVA followed by Dunnett post-hoc test (g,h). *p < 0.05; **p < 0.01; ***p < 0.001. Bar graphs show mean values, error bars indicate SD.
NOTCH3 and TGFβ signaling induces HES and HEY expression
We and others have shown that TGFβ1 and BMPs can activate HEY and HES gene expression via SMAD proteins in endothelial cells10,11. To examine whether this also applies to VSMC, we treated HUASMC with recombinant TGFβ1 and observed not only increased phosphorylation of SMAD2/3 (Fig. 2f), but also increased transcription of HES1 and HEYL (Fig. 2g). To test this further, we stimulated NOTCH3 signaling in HUASMC using immobilized JAG1 ligands. While stimulation with JAG1 did not significantly affect HEY and HES gene transcription, combined treatment with JAG1 and TGFβ1 led to an additive induction of HES1, HES5 and HEYL expression (Fig. 2g). Of note, the induction of HES and HEY genes upon silencing of HTRA1 was fully reverted by inhibiting Notch signaling with the gamma-secretase inhibitor DAPT as well as by inhibiting TGFβ signaling using the ALK5 inhibitor CAS 446859-33-2 (Fig. 2h).
Taken together, these experiments revealed that loss of HTRA1 leads to over-activation of NOTCH3 and TGFβ signaling, two pathways that synergistically stimulate expression of HES and HEY transcriptional repressors.
Loss of HTRA1 leads to decreased contractile protein expression
Notch signaling is essential for the differentiation of VSMC from precursor cells and for maintenance of the contractile phenotype. JAG1 and NOTCH3 are the most abundantly expressed ligands and receptors respectively on VSMC. Several studies have shown that NOTCH3-ICD activates the transcription of contractile proteins, whereas Notch target genes of HES and HEY families, encoding transcriptional repressor proteins, inhibit the expression of these contractile VSMC markers7,8.
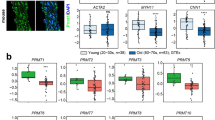
Therefore, we analyzed the effect of HTRA1 on expression of contractile proteins. Western blotting revealed a strong reduction of the contractile VSMC phenotype marker proteins α-SMA and SM22α upon silencing of HTRA1 in HUASMC (Fig. 3a). Similar to cultured VSMC, the protein expression levels of SM22α and α-SMA were diminished in the arterial wall of Htra1−/− mice compared to wild-type littermate controls (Fig. 3b,c). Furthermore, the mRNA expression levels of classical contractile VSMC phenotype marker genes SM22α, α-SMA and Smoothelin were diminished after silencing HTRA1 in HUASCM (Fig. 3d).
Htra1 silencing inhibits contractile protein expression. (a) Immunoblot and quantification of contractile proteins in control or HTRA1-silenced HUASMC (n = 3). (b) Immunoblot and graphical representation of fold change in SM22α protein expression in mesenteric arteries (n = 3). (c) Immunoblot and graphical representation of fold change in α-SMA protein expression in mesenteric arteries (n = 3). (d) Quantitative real-time PCR analysis of contractile gene transcripts in HTRA1-silenced HUASMC (n = 3). (e) Representative immunoblot of HUASMC protein lysates probed with anti-NOTCH3-ICD and anti-SM22α antibodies. (f) Quantitative real-time PCR analysis of contractile gene transcripts in HUASMC overexpressing HEYL and HES1 (n = 3). Statistical significance was determined by an unpaired student’s t-test (a,b,c,d,f). *p < 0.05; **p < 0.01. Bar graphs show mean values, error bars indicate SD.
To further analyze how hyperactive NOTCH3 signaling contributed to the observed changes seen after HTRA1 knockdown, we expressed increasing amounts of NOTCH3-ICD in HUASMC. As expected based on previous reports7,8,23,24,26,28, moderate over-expression of NOTCH3-ICD led to substantially increased SM22α protein levels (Fig. 3e). This result was however contradictory to what we observed after HTRA1 silencing (Fig. 3a–d) questioning the relevance of Notch signaling downstream of HTRA1. Interestingly, with higher NOTCH3-ICD expression levels the expression of SM22α decreased again (Fig. 3e), indicating hyper-activation of a negative feedback loop. Indeed, combined over-expression of HES1 and HEYL repressed transcription of α-SMA and SM22α in HUASMC (Fig. 3f).
Taken together, these data suggest that the strong upregulation of HES and HEY transcriptional repressors by increased Notch and TGFβ signaling upon loss of HTRA1 leads to repression of genes encoding contractile proteins in VSMC.
Loss of HTRA1 in VSMC impairs contractility
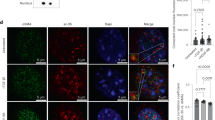
Impaired vascular contractility contributes to the development of small vessel disease30. Therefore, we investigated whether loss of HTRA1 affects VSMC differentiation and function. Immunofluorescence microscopy revealed substantially less actin and myosin fibers in cultured VSMC after HTRA1 knockdown (Fig. 4a). The impairment of F-actin formation upon HTRA1 silencing was restored by inhibition of Notch signaling using DAPT or by inhibition of TGFβ signaling using the ALK5 inhibitor CAS 446859-33-2 (ALK5i) (Fig. 4b).
Contractile gene expression in HTRA1-deficient VSMC is affected by Notch and TGFß signaling. (a) Representative photomicrographs of F-actin and immunolabeled myosin in HUASMC and quantification of F-actin and myosin positive areas (n = 4). (b) Representative photomicrographs of F-actin in HUASMC after four hours of TGFß inhibition using 10 µM ALK5 inhibitor (ALK5i, CAS 446859-33-2, Merck) or NOTCH3 inhibition for 16 hours using 25 µM DAPT (Merck) (n = 3). (c–e) Primary aortic VSMC derived from HtrA1−/− or wildtype littermate controls were cultured until confluence was reached. Quantitative real-time PCR analysis of alpha smooth muscle actin (α-SMA). (d,e) VSMC were treated with DAPT (25 µM) for 16 hours or Alk5i (10 µM) for two hours before RNA isolation. Treatment with the solvent DMSO was used as control (n = 3). Statistical significance was determined by an unpaired student’s t-test (a,c,d,e). *p < 0.05; **p < 0.01. Bar graphs show mean values, error bars indicate SD. Scale bar equals 10 µm.
Next primary aortic VSMC were isolated from HtrA1−/− mice. As expected these had lower α-SMA mRNA expression levels compared to aortic VSMC isolated from their wildtype littermate controls (Fig. 4c). While inhibiting Notch activity with DAPT or TGFβ signaling using ALK5i lowered α-SMA mRNA expression in wildtype VSMC (Fig. 4d), they partially restored α-SMA expression in HtrA1−/− aortic VSMC (Fig. 4e).
To further analyze whether HTRA1 silencing deteriorates cell contractility, we measured the extent of collagen gel contraction by HUASMC. Indeed, cells silenced for HTRA1 expression were severely impaired in their ability to contract collagen gels when compared to control cells (Fig. 5a).
Loss of HTRA1 in VSMC impairs contractility. (a) Cell contraction of control and HTRA1-silenced HUASMC at indicated time-points (n = 3). (b) Relative increase in diameter of mesenteric artery segments from HtrA1+/+ and HtrA1−/− mice challenged with increasing intraluminal pressure. Diameter of unchallenged arteries was set to 100%. Statistical significance was determined by an unpaired student’s t-test (a,b). *p < 0.05; **p < 0.01; ***p < 0.001. Bar graphs show mean values, error bars indicate SD.
SM22α is needed to bundle actin filaments that interact with myosin fibers31. Although SM22α−/− mice are viable, they show impaired vasoconstriction32,33. Based on this, we tested how 3rd order branches of mesenteric resistance arteries isolated from Htra1−/− mice respond to an increase in blood pressure. Consistent with the in vitro data, we observed a less pronounced pressure response in mesenteric arteries from Htra1−/− mice compared to control mice, suggesting a switch from contractile to synthetic phenotype (Fig. 5b). We therefore conclude that HTRA1 is required for VSMC maturation and for appropriate contraction of small resistance arteries.
Discussion
HTRA1 mutations or altered expression levels of this gene are associated with several diseases, whose pathogenesis is mainly driven by impaired blood vessel function. In humans, NOTCH3 and HTRA1 are two of the most frequently mutated genes in familial small vessel disease which is a major cause of stroke and vascular dementia1. Moreover, disturbed VSMC function and integrity are main pathogenic factors in small vessel diseases, which impair white matter perfusion leading to dementia30. For example, attenuated myogenic responses, reduced caliber of brain arteries and impaired cerebrovascular autoregulation were detected in a mouse model of CADASIL34. In autopsies of CARASIL patients, degeneration and loss of VSMC and extracellular matrix proteins, and in addition also severe adventitial fibrosis in small and medium-sized arteries were observed3.
Our study shows that the serine protease HTRA1 controls not only TGFβ signaling but also Notch3 signaling, a central regulator of VSMC function. The data suggest that HTRA1 is needed for full VSMC differentiation into the contractile phenotype. In agreement with this, it was very recently reported that VSMC in aged Htra1−/− mice adopt the synthetic phenotype and are prone to cell death35. Similar to loss of NOTCH323,26,28 or overexpression of CADASIL-related NOTCH3 mutations36, loss of HTRA1 is compatible with normal development of the knockout mouse37. However, our data demonstrate that under stressful situations, reduced expression levels of contractile proteins may impair proper blood perfusion.
Furthermore, the data indicate that HTRA1 is required to limit the activity of NOTCH3 and TGFβ signaling in VSMC. TGFβ and NOTCH3 signaling induce VSMC differentiation7 and synergistically activate expression of contractile marker proteins38 whereupon the Notch transducer RBP-Jκ interacts with Smad2/3 and increases its transcriptional activity. Many promoters of VSMC marker genes contain RBP-Jκ and Smad consensus binding sites in close proximity, suggesting combined action as a transcriptional activator complex38. Our data highlight that the outcome of NOTCH3 and TGFβ signaling in VSMC is highly dependent on the signaling strength. While both pathways activate the expression of contractile genes, they also activate HES and HEY transcriptional repressors. These counteract the Notch-induced up-regulation of contractile proteins like smooth muscle actin39. We therefore conclude that the combined over-activation of Notch and TGFβ signaling in Htra1-deficient VSMC may counteract the expression of contractile proteins by inappropriately high HES and HEY gene expression.
In summary, we demonstrated the first functional link between HTRA1 and NOTCH3 signaling. The similar clinical presentation of CARASIL and CADASIL syndromes3 and a recent report about aberrant HTRA1 protein levels in a CADASIL mouse model3 already suggested such a potential connection. Future work will address how aberrant JAG1 expression influences VSMC biology and how this may be involved in the pathogenesis of small vessel diseases.
Material and Methods
Animals and procedures
Htra1−/− mice on a C57/Bl6 background were described before37. Mice were kept under pathogen-free barrier conditions and animal procedures were performed in accordance with the institutional and national regulation. The local authorities approved all animal experiments.
Perfusion of isolated mouse arteries
Mice were sacrificed and second/third order mesenteric arteries were isolated and perfused with Tyrode buffer at a longitudinal pressure gradient of 20 mm Hg (70–110 mmHg at the inflow and 50–90 mmHg at the outflow) with a resulting flow of ~0.07 mL/min. Arteries which showed no myogenic response were excluded from the analyses. Pressure-induced changes in vessel diameter were measured using the VediView software (DMT, Copenhagen, Denmark).
Human vascular smooth muscle cell isolation and cultivation
VSMC were freshly isolated and cultured as previously described40. Transfection of cells was done as previously described22. Generation of lentivirus and transduction of cells with lentivirus was done as shown elsewhere41. To inhibit Notch signaling, cells were incubated with 25 µM DAPT (Merck) overnight. TGFβ signaling was inhibited by adding 10 µM ALK5 inhibitor (CAS 446859-33-2, Merck) for two hours to basal cell medium.
For the co-culture experiment, the same numbers of HUASMC were seeded with 3T3 cells expressing a TGFβ reporter (Luciferase cassette under the control of 12 SMAD binding sites (5′-CAGA-3′) and cultured overnight in basal medium. Luciferase activity was determined using the Luciferase Assay System (E1500, Promega) and a multiwell plate reader (Clariostar, BMG Labtech).
Adenovirus was obtained from abm Inc. (Vancouver, Canada): HES1 (096900 A), HEYL (096954 A). HUASMC were infected with MOI = 50 and cell lysate were prepared 24 hours later.
Mouse aortic VSMC isolation
Mice were sacrificed by cervical dislocation. After exposing the heart and the descending aorta, fat and adventitia were dissected. The aorta was removed and rinsed in a 60 mm dish with cold PBS containing penicilin and streptomycin and cut into small rings, approximately 1 mm long. Rings were transferred in new 35 mm dishes with 2 ml DMEM with 15% FBS and penicilin/streptomycin containing collagenase at a 10 mg/ml concentration. Rings were incubated overnight. On the next day aortic rings and cells in suspension were transferred to a 15 ml tube, centrifuged (1000 rpm, 5 minutes) and suspended in DMEM with 15% FBS. Cells were incubated until confluence was reached.
RNA extraction and real-time PCR analysis
Total RNA was isolated using the innuPREP RNA Mini Kit (Analytik Jena). cDNA was synthesized from 1 µg of DNA-free total RNA using M-MLV Reverse Transcriptase (Life Technologies) and random hexamer primers (Life Technologies). Gene-specific transcription levels were determined using the POWER SYBR-Green Mastermix (Life Technologies) and an ABI StepOnePlus real-time cycler (Applied Biosystems). Ct-values were determined with LinReg software and normalization was done with the housekeeping genes OAZ1 (human) and Gapdh (mouse). Primers used were as follows: hOAZ (fw): 5′-gagccgaccatgtcttcatt-3′, hOAZ (rev): 5′-ctcctcctctcccgaagact-3′; mGapdh (fw): 5′-aactttggcattgtggaagg-3′, mGapdh (rev): 5′-acacattgggggtaggaaca-3′; hHES1 (fw): 5′-tcaacacgacaccggataaa-3′, hHES1 (rev): 5′-ccgcgagctatctttcttca-3′; mHes1 (fw): 5′-gtcacctcgttcatgcactc-3′, mHes1 (rev): 5′-tctggaaatgactgtgaagca-3′; hHES5 (fw): 5′-gcatggcccccagca-3′, hHES5 (rev): 5′-acgaaggctttgctgtgctt-3′; mHes5 (fw): 5′-caaggagaaaaaccgactgc-3′, mHes5 (rev): 5′-ggctttgctgtgtttcaggt-3′; hHEYL (fw): 5′-gcaagccaggaagaaacaca-3′, hHEYL (rev): 5′-agaatcctgtcccaccagtg-3′; mHeyL (fw): 5′-tatgatccctctgcgcttct-3′, mHeyL (rev): 5′-catcgatgtgggtcaagaga-3′; hα-SMA (fw): 5′-ctgttccagccatccttcat-3′, hα-SMA (rev): 5′-ccgtgatctccttctgcatt-3′; hSMTN (fw): 5′-cctggtgcacaacttcttcc-3′, hSMTN (rev): 5′-aagacacacttggggtcagg-3′; hCNN1 (fw): 5′-ggagctgagagagtggatcg-3′, hCNN1 (rev): 5′-gtgccaattttgggttgact-3′; hMYH11 (fw): 5′-gtgaacgcactcaagagcaa-3′, hMYH11 (rev): 5′-tattcactggccttggttcc-3′
Western Blot analysis
Proteins were separated by SDS-polyacrylamide gel electrophoresis and transferred to nitrocellulose membranes (Whatman). The membranes were blocked with 5% skim milk or 3% bovine serum albumin in 0.1% Tween-20 in TBS and incubated with primary antibodies at 4 °C over-night. The membranes were incubated with peroxidase-conjugated secondary antibody for 2 hours at room temperature followed by ECL detection of the antibody. Band intensities were analyzed by the Image lab software (Biorad).
Antibodies
Anti-JAG1 (Santa Cruz, sc-6011), anti-HTRA1 (R&D, MAB2916), anti-GAPDH (abcam, ab8245), anti-α-SMA (Sigma, C6198), anti-SM22α (abcam, ab137453), anti-NOTCH3 (Santa Cruz, sc-5593), anti-myosin (eBioscience, 14-6400), pSMAD2/3 (Cell Signaling, #8828) were used as primary antibodies. F-actin was stained with Alexa Fluor 568-conjugated phalloidin (Invitrogen).
Immunofluorescence staining
HUASMC were seeded on glass slides coated with 0.5% gelantine. Cells were washed twice with PBS, fixed with 4% PFA for 10 min, washed three times with PBS, permeabilized with PBS-T (containing 0.1% TritonX) and washed again three times with PBS. After blocking with 3% BSA in PBS, cells were incubated with the primary antibodies over night at 4 °C and secondary antibodies (Thermo Fisher Scientific, 1:400) for 1 hour at room temperature. Sections were counterstained with DAPI, washed three times with PBS and mounted with Fluoromount (S3023, Dako). Confocal images were obtained using an LSM 700 microscope (Carl Zeiss) and analyzed using Fiji software.
Statistical analysis
Statistical analysis was performed with the Prism 6.0 software (GraphPad Software). Statistical significance was determined by an unpaired student t-test for two samples or by 1-way ANOVA followed by Dunnett post-hoc test when more than 2 groups were compared. Statistical significance is indicated as *p < 0.05, **p < 0.01, ***p < 0.001.
Study approval
Human umbilical smooth muscle cells were freshly isolated from umbilical cords according to the declaration of Helsinki and with approval of the Heidelberg University ethics review board. Written informed consent from the donor was obtained prior to inclusion in the study. The Animal Care and Use Committee of the Bezirksregierung Baden-Württemberg, Germany, approved all procedures performed on mice.
References
Verdura, E. et al. Heterozygous HTRA1 mutations are associated with autosomal dominant cerebral small vessel disease. Brain 138, 2347–2358, https://doi.org/10.1093/brain/awv155 (2015).
Hara, K. et al. Association of HTRA1 mutations and familial ischemic cerebral small-vessel disease. N Engl J Med 360, 1729–1739, https://doi.org/10.1056/NEJMoa0801560 (2009).
Tikka, S. et al. CADASIL and CARASIL. Brain Pathol 24, 525–544, https://doi.org/10.1111/bpa.12181 (2014).
Owens, G. K. Regulation of differentiation of vascular smooth muscle cells. Physiol Rev 75, 487–517, https://doi.org/10.1152/physrev.1995.75.3.487 (1995).
Fischer, A. & Gessler, M. Delta-Notch–and then? Protein interactions and proposed modes of repression by Hes and Hey bHLH factors. Nucleic Acids Res 35, 4583–4596, https://doi.org/10.1093/nar/gkm477 (2007).
Noguchi, Y. T. et al. Cell-autonomous and redundant roles of Hey1 and HeyL in muscle stem cells: HeyL requires Hes1 to bind diverse DNA sites. Development 146, https://doi.org/10.1242/dev.163618 (2019).
Boucher, J., Gridley, T. & Liaw, L. Molecular pathways of notch signaling in vascular smooth muscle cells. Front Physiol 3, 81, https://doi.org/10.3389/fphys.2012.00081 (2012).
Fouillade, C., Monet-Lepretre, M., Baron-Menguy, C. & Joutel, A. Notch signalling in smooth muscle cells during development and disease. Cardiovasc Res 95, 138–146, https://doi.org/10.1093/cvr/cvs019 (2012).
Guo, X. & Chen, S. Y. Transforming growth factor-beta and smooth muscle differentiation. World J Biol Chem 3, 41–52, https://doi.org/10.4331/wjbc.v3.i3.41 (2012).
Itoh, F. et al. Synergy and antagonism between Notch and BMP receptor signaling pathways in endothelial cells. EMBO J 23, 541–551, https://doi.org/10.1038/sj.emboj.7600065 (2004).
Woltje, K., Jabs, M. & Fischer, A. Serum induces transcription of Hey1 and Hey2 genes by Alk1 but not Notch signaling in endothelial cells. PLoS One 10, e0120547, https://doi.org/10.1371/journal.pone.0120547 (2015).
De Luca, A. et al. Distribution of the serine protease HtrA1 in normal human tissues. J Histochem Cytochem 51, 1279–1284, https://doi.org/10.1177/002215540305101004 (2003).
Oka, C. et al. HtrA1 serine protease inhibits signaling mediated by Tgfbeta family proteins. Development 131, 1041–1053, https://doi.org/10.1242/dev.00999 (2004).
Campioni, M. et al. The serine protease HtrA1 specifically interacts and degrades the tuberous sclerosis complex 2 protein. Mol Cancer Res 8, 1248–1260, https://doi.org/10.1158/1541-7786.MCR-09-0473 (2010).
Chien, J. et al. Serine protease HtrA1 associates with microtubules and inhibits cell migration. Mol Cell Biol 29, 4177–4187, https://doi.org/10.1128/MCB.00035-09 (2009).
Grau, S. et al. Implications of the serine protease HtrA1 in amyloid precursor protein processing. Proc Natl Acad Sci USA 102, 6021–6026, https://doi.org/10.1073/pnas.0501823102 (2005).
Tennstaedt, A. et al. Human high temperature requirement serine protease A1 (HTRA1) degrades tau protein aggregates. J Biol Chem 287, 20931–20941, https://doi.org/10.1074/jbc.M111.316232 (2012).
Tiaden, A. N. & Richards, P. J. The emerging roles of HTRA1 in musculoskeletal disease. Am J Pathol 182, 1482–1488, https://doi.org/10.1016/j.ajpath.2013.02.003 (2013).
Friedrich, U. et al. Synonymous variants in HTRA1 implicated in AMD susceptibility impair its capacity to regulate TGF-beta signaling. Hum Mol Genet 24, 6361–6373, https://doi.org/10.1093/hmg/ddv346 (2015).
Shiga, A. et al. Cerebral small-vessel disease protein HTRA1 controls the amount of TGF-beta1 via cleavage of proTGF-beta1. Hum Mol Genet 20, 1800–1810, https://doi.org/10.1093/hmg/ddr063 (2011).
Zhang, L. et al. High temperature requirement factor A1 (HTRA1) gene regulates angiogenesis through transforming growth factor-beta family member growth differentiation factor 6. J Biol Chem 287, 1520–1526, https://doi.org/10.1074/jbc.M111.275990 (2012).
Klose, R. et al. Inactivation of the serine protease HTRA1 inhibits tumor growth by deregulating angiogenesis. Oncogene, https://doi.org/10.1038/s41388-018-0258-4 (2018).
Boulos, N. et al. Notch3 is essential for regulation of the renal vascular tone. Hypertension 57, 1176–1182, https://doi.org/10.1161/HYPERTENSIONAHA.111.170746 (2011).
Domenga, V. et al. Notch3 is required for arterial identity and maturation of vascular smooth muscle cells. Genes Dev 18, 2730–2735, https://doi.org/10.1101/gad.308904 (2004).
Henshall, T. L. et al. Notch3 is necessary for blood vessel integrity in the central nervous system. Arterioscler Thromb Vasc Biol 35, 409–420, https://doi.org/10.1161/ATVBAHA.114.304849 (2015).
Li, X. et al. Notch3 signaling promotes the development of pulmonary arterial hypertension. Nat Med 15, 1289–1297, https://doi.org/10.1038/nm.2021 (2009).
Liu, H., Kennard, S. & Lilly, B. NOTCH3 expression is induced in mural cells through an autoregulatory loop that requires endothelial-expressed JAGGED1. Circ Res 104, 466–475, https://doi.org/10.1161/CIRCRESAHA.108.184846 (2009).
Belin de Chantemele, E. J. et al. Notch3 is a major regulator of vascular tone in cerebral and tail resistance arteries. Arterioscler Thromb Vasc Biol 28, 2216–2224, https://doi.org/10.1161/ATVBAHA.108.171751 (2008).
Low, E. L., Baker, A. H. & Bradshaw, A. C. TGFbeta, smooth muscle cells and coronary artery disease: a review. Cell Signal 53, 90–101, https://doi.org/10.1016/j.cellsig.2018.09.004 (2019).
Tan, R. Y. & Markus, H. S. Monogenic causes of stroke: now and the future. J Neurol 262, 2601–2616, https://doi.org/10.1007/s00415-015-7794-4 (2015).
Han, M. et al. Smooth muscle 22 alpha maintains the differentiated phenotype of vascular smooth muscle cells by inducing filamentous actin bundling. Life Sci 84, 394–401, https://doi.org/10.1016/j.lfs.2008.11.017 (2009).
Je, H. D. & Sohn, U. D. SM22alpha is required for agonist-induced regulation of contractility: evidence from SM22alpha knockout mice. Mol Cells 23, 175–181 (2007).
Zhang, J. C. et al. Analysis of SM22alpha-deficient mice reveals unanticipated insights into smooth muscle cell differentiation and function. Mol Cell Biol 21, 1336–1344, https://doi.org/10.1128/MCB.2001.21.4.1336-1344.2001 (2001).
Joutel, A. et al. Cerebrovascular dysfunction and microcirculation rarefaction precede white matter lesions in a mouse genetic model of cerebral ischemic small vessel disease. J Clin Invest 120, 433–445, https://doi.org/10.1172/JCI39733 (2010).
Ikawati, M., Kawaichi, M. & Oka, C. Loss of HtrA1 serine protease induces synthetic modulation of aortic vascular smooth muscle cells. PLoS One 13, e0196628, https://doi.org/10.1371/journal.pone.0196628 (2018).
Dubroca, C. et al. Impaired vascular mechanotransduction in a transgenic mouse model of CADASIL arteriopathy. Stroke 36, 113–117, https://doi.org/10.1161/01.STR.0000149949.92854.45 (2005).
Tsuchiya, A. et al. Expression of mouse HtrA1 serine protease in normal bone and cartilage and its upregulation in joint cartilage damaged by experimental arthritis. Bone 37, 323–336, https://doi.org/10.1016/j.bone.2005.03.015 (2005).
Tang, Y. et al. Notch and transforming growth factor-beta (TGFbeta) signaling pathways cooperatively regulate vascular smooth muscle cell differentiation. J Biol Chem 285, 17556–17563, https://doi.org/10.1074/jbc.M109.076414 (2010).
Tang, Y., Urs, S. & Liaw, L. Hairy-related transcription factors inhibit Notch-induced smooth muscle alpha-actin expression by interfering with Notch intracellular domain/CBF-1 complex interaction with the CBF-1-binding site. Circ Res 102, 661–668, https://doi.org/10.1161/CIRCRESAHA.107.165134 (2008).
Ghosh, S. et al. Loss of the mechanotransducer zyxin promotes a synthetic phenotype of vascular smooth muscle cells. J Am Heart Assoc 4, e001712, https://doi.org/10.1161/JAHA.114.001712 (2015).
Tetzlaff, F. et al. MPDZ promotes DLL4-induced Notch signaling during angiogenesis. Elife 7, https://doi.org/10.7554/eLife.32860 (2018).
Acknowledgements
We thank the members of the DKFZ laboratory animal facility for excellent support. This work was supported by the Fritz-Thyssen Foundation (10.13.1.160 to A.F.) and the Deutsche Forschungsgemeinschaft (DFG) - project number 394046768 - SFB1366 (to A.F. and T.K.).
Author information
Authors and Affiliations
Contributions
R.K., A.P., F.T., E.M.W., I.M., T.K., J.R.V. and A.F. conceived and designed experiments and analyzed the data; C.O. provided essential reagents and discussed data; R.K., F.T., E.M.W. and A.F. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Klose, R., Prinz, A., Tetzlaff, F. et al. Loss of the serine protease HTRA1 impairs smooth muscle cells maturation. Sci Rep 9, 18224 (2019). https://doi.org/10.1038/s41598-019-54807-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-54807-6
This article is cited by
-
Blood vessel organoids generated by base editing and harboring single nucleotide variation in Notch3 effectively recapitulate CADASIL-related pathogenesis
Molecular Neurobiology (2024)
-
Porphyromonas gingivalis-mediated disruption in spiral artery remodeling is associated with altered uterine NK cell populations and dysregulated IL-18 and Htra1
Scientific Reports (2022)
-
Cross-sectional associations among P3NP, HtrA, Hsp70, Apelin and sarcopenia in Taiwanese population
BMC Geriatrics (2021)
-
Notch-ing up knowledge on molecular mechanisms of skin fibrosis: focus on the multifaceted Notch signalling pathway
Journal of Biomedical Science (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.