Abstract
The intestinal microbiota contributes to host metabolism and health. This study aimed to assess the effects of biochar on cecal microbiome-related metabolic changes in rats. Rats were orally administered rice straw biochar (RSB) at 1120 mg/kg body weight for 5 weeks. Cecal samples were analyzed to perform metabolic and microbial profiling via a combination of 16S rRNA gene sequencing and LC/MS techniques. We observed a significant influence of RSB in shaping the cecal bacterial community, including some potentially beneficial members of phylum Firmicutes belonging to unclassified Lachnospiraceae, Oscillibacter, and Clostridium XlVa and IV, as well as the depletion of some opportunistic pathogens belonging to Prevotella, Bacteroides and Paraprevotella. The metabolomic analysis revealed distinct changes in the cecal metabolic phenotype, including lower levels of L-isoleucine, indole-3-acetic acid, benzoic acid, and tetradecanoic acid as well as higher levels of L-phenylalanine, L-glutamate, 3-phenylpropanoic acid, chenodeoxycholic acid, cholic acid, 7-dehydrocholesterol, (5Z, 8Z, 11Z, 14Z, 17Z)-eicosapentaenoic acid, 11-deoxycorticosterone and retinol, which are mainly involved in the metabolic pathways of linoleic acid, amino acid and steroid hormone biosynthesis. Correlation analysis revealed a positive association of unclassified Lachnospiraceae, Oscillibacter and Clostridium IV with 3-phenylpropanoic acid, L-phenylalanine, L-glutamate, 11-deoxycorticosterone and 7-dehydrocholesterol. These results confirm that the gut microbiome is altered and may be critical for good performance under RSB application by interacting with metabolism.
Similar content being viewed by others
Introduction
The intestinal microbiota is composed of an extremely large number of different bacteria that produce a variety of metabolites, are exclusively responsible for microbe selection and are involved in important metabolic functions, such as the biosynthesis and biotransformation of amino acids and bile acid1. Factors such as changes in nutritional interventions, host condition, radiation and toxicological insult can induce microbial regulation2 and affect host health. Therefore, the integrative analysis of gut microbiota and metabolite profiles can help to understand the changes in the host’s physiological condition under the aforementioned types of interference.
Carbonaceous adsorbents have been employed worldwide to improve animal performance as dietary interventions3,4 and lead to profound microbial modulation5,6. These adsorbents, with inhomogeneous microporous structures, can supply appropriate habitats for microorganisms7. Biochar is one kind of carbonaceous adsorbent, produced by the degradation of organic matter from agricultural waste in an oxygen-limited, anaerobic environment8. This productive process makes biochar rich in micronutrients and forms a large surface area and more optimal macropores. All of these characteristics contribute to the supply of more spacious sites and nutrition for microorganisms, contributing to their proliferation or providing “refuge”9,10. In China, biochar possesses the outstanding advantages of territorially unlimited sources of plant-derived raw material, a reasonable price and environmentally friendly production process. The regulation of microorganism activity and abundance by biochar has been well documented11,12,13. China is a large agricultural country where there is considerable livestock breeding, in which the utilization of biochar as a dietary supplement could be highly significant. The use of biochar could not only help to realize new feed resource development but could also improve the rational utilization of agricultural waste. However, an understanding of the effect of biochar on the metabolite profile related to the gut microflora is lacking. Moreover, considering the numerous unculturable gut bacteria, the cultural approach adopted in previous studies has suffered from the underestimation of the actual microbial community14. Hence, there is the need to obtain an overall view of the changes in metabolite profiles related to the gut microbiota under the inclusion of biochar in the diet, which will contribute to better understanding the beneficial effect of biochar on animal well-being15.
High-throughput sequencing is an accurate and powerful approach for obtaining detailed information on the microbiome16. It can transcend the limitations of traditional culture strategies for some of the unculturable gut microflora in vitro. Metabolomics represents an unprecedented approach for detecting the often-subtle differences in a broad range of metabolites in tissues or biofluids. The metabolomics technique enables the identification of the global metabolic response between the host and its gut commensal microbiota17. The application of metabolomics for assessing the effect of nutrition interventions is effective, especially when metabolic changes easily escape detection by traditional hypothesis-led approaches18. Regarding the relationship between the gut microbiota and the host metabolic phenotype, a nontargeted metabolomic study provides an effective approach for resolving the metabolic phenotypic variations related to gut microbiota perturbations, about which many research questions remain19.
Recently, the changes in growth performance, intestinal mucosal conditions, and the cecal microbial community induced by RSB application in rats were examined20. In this study, we probe the cecal microbiome-related metabolic changes in rats following the dietary inclusion of rice straw biochar (RSB) by employing high-throughput sequencing and liquid chromatography coupled to mass spectrometry (LC-MS).
Results
RSB-induced changes in the cecal bacterial microbiome
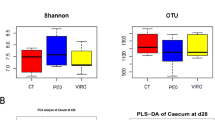
The high-quality sequences were clustered into OTUs according to a cut-off of 97% sequence similarity. The OTU rank abundance in the RSB group exhibited a gentler slope and wider distribution on the horizontal axis, indicating that both more diversity and a more uniform distribution were induced in the cecal microbiota in response to RSB addition (Fig. 1A). The Chao1 estimation reflects the ecological species richness of the bacterial community and primarily highlights species numbers, and the Shannon index represents and correlates positively with the species distribution. The Chao1 index (average values of 1561 vs 1364, P < 0.05) (Fig. 1B) and the Shannon index (average values of 8.01 vs 7.31, P < 0.05) (Fig. 1C) in the RSB group were higher than those in the control group, indicating that RSB addition supported significantly more richness and microbial diversity in the cecum of the rats.
Changes in the richness and diversity of the cecal microbiota as a result of RSB treatment. (A) The OTU rank abundance of the cecal microbiota in rats. (B) Bacterial richness estimated from the Chao 1 value; (C) bacterial diversity estimated from the Shannon index. C2~C7 indicate rats from the control group; R1~R7 indicate rats from the RSB group.
The taxon summary revealed distinct changes in the gut microbial composition in response to RSB administration. In terms of the assignment at the phylum level, 15 phyla were detected in all samples, and the dominant phyla were Firmicutes, Bacteroidetes, Proteobacteria, and unclassified Bacteria, with average relative abundances in the RSB group of 68.61%, 21.01%, 7.46% and 1.79%, respectively (Fig. 2A). More importantly, the abundance of phylum Firmicutes in the RSB group was markedly increased by 64.18% (P < 0.05), whereas the proportion of Bacteroidetes decreased by 57.03% (P < 0.05) compared to those in the control group. At the genus level, the taxa whose relative abundance values were significantly affected by RSB included unclassified Lachnospiraceae, unclassified Ruminococcaceae, Prevotella, and unclassified Porphyromonadaceae, with proportions in the RSB group of 39.92%, 10.90%, 5.14%, and 5.11%, respectively, among the 122 genera identified (Fig. 2B). The percentages of unclassified Lachnospiraceae (34.78% of all reads, 1079 OTUs), Oscillibacter (1.19% of all reads, 37 OTUs), Clostridium IV (0.84 of all reads, 26 OTUs) and Clostridium XlVa (0.42% of all reads, 13 OTUs) in the RSB group were markedly increased by 172.68% (P < 0.05), 67.48% (P < 0.05), 117.5% (P < 0.05) and 436.36% (P < 0.05), respectively. Conversely, the proportions of Prevotella (7.58% of all reads, 235 OTUs), Paraprevotella (0.77% of all reads, 24 OTUs) and Bacteroides (0.55% of all reads, 17 OTUs) were decreased by 81.23% (P < 0.05), 46.60% (P < 0.05), and 69.54% (P < 0.05), respectively, compared to the control group.
RSB-induced gut metabolic profiling analysis
Multivariate analysis methods including PCA and PLS-DA were subsequently employed to reveal the clustering trends of each group. Each point in the PCA score plot represents an individual sample, and the samples are divided into blocks suggesting that the two clusters are clearly divided, representing the two groups with different metabolic profiles (Fig. 3A), thus allowing the visualization of the general changes in the cecal metabolic profile after RSB application. The supervised pattern recognition of PLS-DA focuses on the actual class-discriminating variations and shows better separation compared to the unsupervised PCA. Every dot in this score plot represents an observed sample, and the distance between two dots reflects the similarity of the metabolite composition. The score plot of PLS-DA in Fig. 3B reveals good fitness and high predictability of this model with high statistical values of Q2 = 0.916 and R2 = 0.998, indicating significant changes in the cecal metabolomes in response to RSB treatment with good clustering of the RSB group on the right and the control group on the left.
Multivariate statistical analysis, heat map and summary of metabolome pathways. (A) PCoA score plot of metabolite profiling between the control and RSB groups. (B) PLS-DA score plots of cecal samples from the control and RSB groups. (C) Heat map summarizing the fold changes in significantly altered metabolites in the cecal samples. Red and blue represent higher and lower concentrations of metabolites, respectively. (D) The impact of pathways between the control and RSB groups. Mapping of the relative concentrations of metabolites to the metabolome indicating the contributions of the impact of metabolic pathways. Node color indicates significance based on p-values, and node size indicates the significance of pathway impact. Significantly impacted pathways include (1) linoleic acid metabolism, (2) steroid hormone biosynthesis, (3) D-glutamine and D-glutamate metabolism, (4) valine, leucine and isoleucine biosynthesis, (5) alpha-linolenic acid metabolism, and (6) phenylalanine metabolism.
The potential identified compounds were generated using the program metaX according to the following procedure: comparison of accurate MS and MSE fragments with the metabolites searched in the freely accessible HMDB database by detecting their molecular weights and elemental compositions. Compounds without information for the given mass fragment were removed from the candidate list according to the possible fragment mechanisms, and only the most likely compounds were reserved. To select the potential biomarkers responsible for this difference, the variable importance for projection (VIP) parameter combined with the p-value and fold change was used to select the variables with the most significant contribution in discriminating cecal compounds between the two groups. Fifteen metabolites with a critical VIP > 1.0, p value < 0.05 and fold change ≥1.2 or ≤0.833 were identified in the cecal samples of rats treated with RSB, and the results are listed in Table 1. The variation tendencies of these metabolites are shown in a heat map in Fig. 3C. Four metabolites were significantly decreased, including tetradecanoic acid, L-isoleucine, benzoic acid and indole-3-acetic acid. Nine metabolites were significantly enriched in the RSB addition group, including L-phenylalanine, L-glutamate, 3-phenylpropanoic acid, cholic acid, (5Z, 8Z, 11Z, 14Z, 17Z)-eicosapentaenoic acid, chenodeoxycholic acid, 11-deoxycorticosterone, 7-dehydrocholesterol and retinol. These metabolites influenced by RSB addition were mainly involved in pathways such as steroid hormone biosynthesis and amino acid transport and protein synthesis (phenylalanine metabolism, arginine biosynthesis and valine, leucine and isoleucine biosynthesis) (Fig. 3D).
Correlation between the gut microbiome and metabolome
Figure 4 shows the functional relationships of the altered gut microbiota and disturbed cecal metabolites following RSB application based on the Spearman correlation coefficients (|r| > 0.5, p < 0.05). Notably, unclassified Lachnospiraceae, Oscillibacter, Clostridium IV and Clostridium XlVa of phylum Firmicutes identified upon RSB application showed similar trends of correlation. The relative abundance of the predominant RSB-enriched gut bacterial genus of unclassified Lachnospiraceae was positively associated with the concentrations of nine increased metabolites, including cholic acid, 7-dehydrocholesterol, 11-deoxycorticosterone, retinol, chenodeoxycholic acid, L-phenylalanine, L-glutamate, 3-phenylpropanoic acid and (5Z, 8Z, 11Z, 14Z, 17Z)-eicosapentaenoic acid, and negatively correlated with the concentrations of four metabolites showing decreasing trends, including L-isoleucine, tetradecanoic acid, indole-3-acetic acid, and benzoic acid. when the increase in abundance was smaller, the only metabolites presenting negative associations were L-isoleucine and indole-3-acetic acid with the genus Oscillibacter. Clostridium IV and Clostridium XlVa possessed the same positive correlations with cholic acid and retinol. Interestingly, the relative abundances of several RSB-depleted opportunistic pathogens in phylum Bacteroidetes belonging to the genera Prevotella, Bacteroides and Paraprevotella showed opposite trends to those of the genera of phylum Firmicutes. Prevotella and Paraprevotella presented negative correlations with the same nine metabolites 93-phenylpropanoic acid, cholic acid, retinol, L-glutamate, L-phenylalanine, 7-dehydrocholesterol, chenodeoxycholic acid, (5Z, 8Z, 11Z, 14Z, 17Z)-eicosapentaenoic acid and 11-deoxycorticosterone) and positive correlations with four metabolites (tetradecanoic acid, L-isoleucine, benzoic acid, and indole-3-acetic acid). Our results also revealed clear correlations among some metabolites. For example, retinol was strongly positively correlated with chenodeoxycholic acid and cholic acid; chenodeoxycholic acid was correlated with cholic acid; and 11-deoxycorticosterone was associated with L-phenylalanine.
Discussion
Here, we investigated the effect of RSB addition on cecal microbiota-related metabolic phenotype alterations in rats. Our findings showed that RSB treatment improved cecal microbial richness, which was proven by the higher OTU rank abundance and the values of the Chao 1 and Shannon indexes. Similarly, the microbial diversity of the cecal contents was improved according to the notable abundance changes in phylum Firmicutes in the genera of unclassified Lachnospiraceae, Oscillibacter, Clostridium IV, and Clostridium XlVa as well as phylum Bacteroidetes in the genera Prevotella, Paraprevotella, and Bacteroides. These components were verified in our previous report20. These results indicated that RSB inclusion induced more specific flora OTUs and a positive impact on the diversity of the ecosystem of the gut bacterial community. Additionally, PLS-DA revealed 13 cecal compounds that were the most important in differentiating the RSB and control groups, including L-phenylalanine, L-glutamate, L-isoleucine, 3-phenylpropanoic acid, benzoic acid, tetradecanoic acid, chenodeoxycholic acid, cholic acid, 7-dehydrocholesterol, 11-deoxycorticosterone, retinol, (5Z, 8Z, 11Z, 14Z, 17Z)-eicosapentaenoic acid and indole-3-acetic acid.
Increasing evidence suggests that metabolite profiles related to the gut microbiome that change under nutritional interventions are important factors in the modulation of tissue functions and achieving health improvement1. Accordingly, positive associations between the compounds 3-phenylpropanoic acid and benzoic acid and the microbiota components of Clostridium, Oscillibacter, and unclassified Lachnospiraceae were observed in our work. 3-Phenylpropionic acid and benzoic acid are major polyphenol compounds identified in the digestive tract21. It is not only the interactions between the gut microbiota and polyphenols but also the mechanisms of their interactions and the effects on host health that have been investigated22. Phylum Firmicutes produces a smaller number of glycan-degrading enzymes involved in polyphenol degradation than Bacteroidetes23; thus, the gut microbiota might be reshaped due to the intake of different polyphenols. Conversely, phenolic compounds have the ability to alter the gut microbiota balance of Bacteroides/Firmicutes24, which was supported by our observation of an increased ratio of Firmicutes to Bacteroidetes among the cecal bacteria phyla under RSB inclusion, and this change in the bacterial ratio has been considered to contribute to the improvement of health status in many studies25,26. Moreover, genus Clostridium is mainly involved in the metabolism of many phenolics27. Its positive associations with 3-phenylpropanoic acid and benzoic acid demonstrated that this microbial population may be closely associated with the generation and bioconversion of these polyphenols under RSB application. This suggested that RSB exerted beneficial effects on animal well-being, which is consistent with our previous report demonstrating the beneficial effects of the application of RSB on growth and intestinal health20.
Amino acids are the major building blocks of proteins and are important in the regulation of energy and protein metabolism in organisms28. Some amino acids, including the essential amino acids L-isoleucine and L-phenylalanine as well as the nonessential amino acid L-glutamate, were affected after RSB application in our work. The intimate relationships between amino acids and amino acid-fermenting bacteria belonging to the Clostridium clusters have been identified this group as the predominant amino acid-fermenting microbiota along the digestive tract of humans and animals28. In line with the above research, the current study showed a clear positive association of L-glutamate and L-phenylalanine with the abundance of genus Clostridium IV under RSB inclusion, which suggested that oral RSB supplementation might particularly impact amino acid metabolism and concomitant host nutrition and health. Previous research has uniformly supported the conclusion that animal dietary supplementation of biochar can improve intestinal villus histology20,29 and enhance nutrient digestibility15 and feed conversion30, which may be partly related to improvement of AA metabolism by biochar.
RSB application also changed the concentrations of some compounds participating in fat metabolism, such as tetradecanoic acid, chenodeoxycholic acid, cholic acid, 7-dehydrocholesterol, 11-deoxycorticosterone and (5Z, 8Z, 11Z, 14Z, 17Z)-eicosapentaenoic acid. Tetradecanoic acid belongs to the saturated fatty acids and is derived from the microbial hydrogenation of nonconjugated dienoic acids31. It has been identified in the animal digestive tract32 and in general triggered inflammatory bowel disease33. However, (5Z, 8Z, 11Z, 14Z, 17Z)-eicosapentaenoic acid is a long-chain poly-unsaturated fatty acid that contributes to regulating the triad of adiposity, inflammation, and fatty acid metabolism34. The positive associations of tetradecanoic acid with Prevotella and Paraprevotella demonstrate that microbial hydrogenation to synthesize tetradecanoic acid by hydrogen-producing Prevotella and Paraprevotella is suppressed and consequently decreases the risk of a gut inflammatory reaction in response to RSB treatment.
In addition, Clostridium IV and XIVa, which are members of phylum Firmicutes, have been demonstrated to be significantly less abundant in inflammatory bowel disease patients than in healthy individuals. These microbiota have proven to be important in maintaining the diversity of the clostridia and preventing intestinal inflammation35. The negative correlation of Clostridium IV and XIVa with tetradecanoic acid potentially suggests that these microbiota may contribute to the resistance of gut inflammation through the modulation of tetradecanoic acid. Cholesterol can be metabolized for the synthesis of cholic acid, chenodeoxycholic acid, 11-deoxycorticosterone and 7-dehydrocholesterol through various metabolic processes34. Cholic acid and chenodeoxycholic acid are the primary bile acids in most species. They play an important physiological role in boosting the absorption of vitamins and fats and regulating bile acid biosynthesis and cholesterol homeostasis36. 11-Deoxycorticosterone is a steroid hormone that is mainly involved in the maintenance of the blood salt balance37. The elevated proportions of chenodeoxycholic acid, cholic acid, 7-dehydrocholesterol, and 11-deoxycorticosterone identified in our study suggested that RSB particularly participated in the regulation of fat metabolism and sterol biosynthesis. Retinyl phosphate plays an important role in the synthesis of cell membrane proteins and participates in the maintenance of cell membrane integrity. Retinyl phosphate also participates in the synthesis of steroid hormones and affects the growth and development of the host38. Our observation of increased concentrations of retinol combined with elevated proportions of 11-deoxycorticosterone and 7-dehydrocholesterol in the steroid biosynthesis pathways after RSB application further indicated that RSB may mainly affect the synthesis of steroid hormones to modulate host growth and development.
In summary, our results from 16S rRNA gene sequencing showed that RSB application significantly changed the gut microbiota community in terms of its diversity and composition. Additionally, RSB altered the metabolic phenotype according to LC-MS metabolomic analysis, which revealed major roles in linoleic acid metabolism, steroid hormone biosynthesis, amino acid transport and protein synthesis. In addition, the correlation analysis revealed relationships between some differentially abundant gut microbes and the candidate metabolites. In general, our results confirmed that the gut microbiome is altered and contributes to performance by interacting with metabolism under RSB application.
Materials and Methods
Rice straw biochar preparation and characterization
The details of the preparation and the characteristics of RSB have been described previously20.
Animals and experimental design
The animals and experimental design used in our study were consistent with our previous report20. Specific pathogen-free grade female Wistar rats at 5 weeks of age weighing 190–210 g that were commercially obtained from the Liaoning Experimental Animal Resources Center (Benxi, China), were used in the current study. All the animals were housed under standard laboratory conditions (22 ± 2 °C, 53% ± 2% relative humidity, 12:12 light/dark cycle) with ad libitum access to chow and water. All the rats were randomly divided into control and RSB groups of 10 rats per group after acclimation for one week. An RSB solution with a concentration of 112 mg/mL was prepared using ultrapure deionized water and administered to the rats via oral gavage at a dose of 1120 mg/kg body weight (BW) daily for 5 weeks. The control group received an equivalent amount of ultrapure deionized water instead. The gavage volume of 1 mL/100 g BW was adjusted according to BW once per week. The oral dose of RSB was selected in reference to the results obtained from our previous study15 by using a dose conversion formula for rats39. Following anesthesia and cervical dislocation, samples of the cecal contents of the rats were immediately collected and snap-frozen in liquid nitrogen to halt metabolism, then stored at −80 °C prior to the metabolic profiling and microbial community analyses. The Animal Ethics Committee of Shenyang Agricultural University approved this study, and all management and experimental procedures were conducted according to the Guidelines for the Care and Use of Animals of Shenyang Agricultural University.
Bacterial sequencing and analysis
The bacterial sequencing analysis performed in our study was consistent with our previous report20. Total DNA was extracted from the cecal contents by utilizing the E.Z.N.A. Stool DNA kit (D4015, Omega, Inc., USA). The V3-V4 region of the 16S rRNA gene was selected for subsequent pyro-sequencing. PCR amplification was performed with slightly modified versions of the primers 338F (5′-ACTCCTACGGGAGGCAGCAG-3′) and 806R (5′-GGACTACHVGGGTWTCTAAT-3′)40. PCR amplification reactions were performed in a total volume of 25 μL, and the reaction mixtures contained 25 ng of template DNA, 12.5 μL of PCR premix, 2.5 μL of each primer, and PCR-grade water to adjust the volume. The PCR conditions for the amplification of the prokaryotic 16 S fragments consisted of initial denaturation at 98 °C for 30 s, followed by 35 cycles of denaturation at 98 °C for 10 s, primer annealing at 54 °C for 30 s, and extension at 72 °C for 45 s, with a final extension at 72 °C for 10 min. Then, the PCR products were visualized on a 2% agarose gel and cleaned by using AMPure XP beads (Beckman Coulter Genomics, Danvers, MA, USA) and quantified with a Qubit fluorometer (Invitrogen, Carlsbad, California, USA). The amplicon library was assessed for size and quantity by using an Agilent 2100 Bioanalyzer (Agilent, USA) with a Library Quantification Kit for Illumina (Kapa Biosciences, Woburn, MA, USA) and sequenced on the Illumina 300PE MiSeq platform. Chimeric sequences were filtered using Verseach software (V 2.3.4). The filtered sequences were clustered into operational taxonomic units (OTUs) based on a threshold of 97% sequence similarity. An analysis of UniFrac dissimilarities was calculated according to the levels of changes in the gut microbiota using R (version 3.2.1) with the vegan package.
Sample preparation for metabolic profiling and LC-MS analysis
After the gut contents were thawed on ice, precooled 50% methanol was added to 25 mg of the samples, followed by vortexing for 1 min and incubation at room temperature for 10 min. The extraction mixture held overnight at −20 °C and centrifuged (25,000 × g) at 4 °C for 15 min to obtain the supernatant. Then, the supernatants were transferred to new 96-well plates and stored at −80 °C for subsequent LC-MS analysis.
Chromatographic separation was performed in an ultra-performance liquid chromatography system (UPLC) (SCIEX, California, USA) using an ACQUITY UPLC BEH C18 reversed-phase column (2.1 mm × 100 mm, 1.7 μm) (Waters, Massachusetts, USA) with a constant column temperature of 35 °C. After equilibration for 2 min, the column flow rate remained at 400 μL/min, and the injection volume of each sample was 4 μL. The mobile phase consisted of solvent A (25 mM ammonium acetate + 25 mM NH3·H2O) and solvent B (isopropanol:acetonitrile = 9:1 + 0.1% formic acid). The UPLC elution conditions were optimized and set as follows: 0~0.5 min, 95% B; 0.5~9.5 min, 95% to 65% B; 9.5~10.5 min, 65~40% B; 10.5~12 min, 40% B; 12~12.2 min, 40~95% B; 12.2~15 min, 95% B.
A Triple TOF5600+ high-resolution tandem mass spectrometer (SCIEX California, USA) was used to detect the eluted metabolites. Electrospray ionization in both positive and negative modes was employed to introduce the sample to the mass spectrometer. The curtain gas was set at 30 PSI with ion source gas 1 and gas 2 both at 60 PSI. The capillary voltage and cone voltages were set at 2000 V and 40 V for positive ion mode and 1000 V and 40 V for negative ion mode, respectively. The desolation gas was set at 450 L/h at 350 °C; the cone gas was set at 50 L/h; and the source temperature was set at 120 °C. The TOF mass range was from 60 to 1200 Da. The survey scans were acquired over 0.15 s, and as many as 12 production scans were collected if they exceeded a threshold of 100 counts per second (counts/s). Four time bins were summed for each scan at a impulsator frequency value of 11 kHz with four-anode/channel detection.
Data processing and statistical analysis
LC/MS raw data were converted into mzXML format and processed using XCMS software to detect and align features. The XCMS analysis provided a matrix including peak picking, peak grouping, retention time (RT) correction, second peak grouping, and annotation of isotopes for every sample. Quality control samples (QCs) were injected at the beginning of the analysis and detected after every 5 samples to evaluate the stability of the LC/MS throughout acquisition. The features detected in less than 50% of QCs or 80% of biological samples were removed; the combination of the RT and m/z data was used to identify each ion. The intensity data of each peak were recorded and further preprocessed with the in-house software metaX. Outlier detection and batch effect evaluation were performed through principal component analysis (PCA) by using the preprocessed dataset. Quality control-based robust LOESS signal correction was fitted to the QC data with respect to the order of injection to minimize signal intensity drift over time. The relative standard deviations of the metabolic features exceeding 30% were removed, and the P value was adjusted for multiple tests using a false discovery rate (Benjamini–Hochberg). Supervised PLS-DA was conducted with metaX to discriminate the different variables between groups. The VIP value was calculated, and a cut-off value of 1.5 was used to select important features. The online HMDB database (http://www.hmdb.ca) was used to annotate the metabolites by matching the exact molecular mass data (m/z) of the samples with those from the database. The correlations between the cecal compounds and bacterial profiles were assessed with Spearman’s correlation test using R (version 3.2.1) with the vegan package.
Statistical analysis
The results were statistically analyzed using IBM SPSS statistical software (version 22.0). The significant differences between the control and RSB supplementation groups were analyzed by one-way ANOVA, followed by a two-tailed Student’s t-test. P < 0.05 was considered to indicate a statistically significant difference.
References
Putignani, L. et al. Gut microbiota dysbiosis as risk and premorbid factors of IBD and IBS along the childhood-adulthood transition. Inflamm Bowel Dis 22, 487–504 (2015).
David, L. A. et al. Diet rapidly and reproducibly alters the human gut microbiome. Nature 505, 559–563 (2014).
Chu, G. M. et al. Effects of bamboo charcoal on the growth performance, blood characteristics and noxious gas emission in fattening pigs. J Appl Anim Res 41, 48–55 (2013a).
Ruttanavut, J., Yamauchi, K., Goto, H. & Erikawa, T. Effects of dietary bamboo charcoal powder including vinegar liquid on growth performance and histological intestinal change in Aigamo ducks. In. J Poultry Sci 8, 229–236 (2009).
Chu, G. M. et al. Effects of bamboo charcoal and bamboo vinegar as antibiotic alternatives on growth performance, immune responses and fecal microflora population in fattening pigs. Anim Sci J 84, 113–120 (2013b).
Watarai, S. & Tana. Eliminating the carriage of Salmonella enterica serovar Enteritidis in domestic fowls by feeding activated charcoal from bark containing wood vinegar liquid (Nekka-Rich). Poultry Sci 84, 515–521 (2005).
Dubinin, M. M. & Plavnik, G. M. Microporous structures of carbonaceous adsorbents. Carbon 6, 321–324 (1981).
Lehmann, J., Gaunt, J. & Rondon, M. Bio-char sequestration in terrestrial ecosystems: A review. Mitig Adapt Strat Gl 11, 403–427 (2006).
Ogawa, M. Symbiosis of people and nature in the tropics. III. Tropical agriculture using charcoal. Farming Jpn 28, 21–35 (1994).
Samonin, V. V. & Elikova, E. E. A study of the adsorption of bacterial cells on porous materials. Microbiology 73, 696–701 (2004).
Steinbeiss, S., Gleixner, G. & Antonietti, M. Effect of biochar amendment on soil carbon balance and soli microbial activity. Soil Biol Biochem 41, 1301–1310 (2009).
Jindo, K. et al. Biochar influences the microbial community structure during manure composting with agricultural wastes. Sci Total Environ 416, 476–481 (2012).
Sun, D. Q., Lan, Y., Xu, E. G., Meng, J. & Chen, W. F. Biochar as a novel niche for culturing microbial communities in composting. Waste Manage 54, 93–100 (2016).
Zoetendal, E. G. et al. Mucosa-associated bacteria in the human gastrointestinal tract are uniformly distributed along the colon and differ from the community recovered from feces. Appl Environ Microb 68, 3401–3407 (2002).
Han, J., Zhang, F., Du, L., Chen, W. F. & Meng, J. Effects of dietary biochar including vinegar liquid on growth performance, nutrient digestibility, blood characteristics and fecal noxious gas emission in weaned piglets. J Anim Vet Adv 18, 1072–1079 (2014).
Loman, N. J. et al. High-throughput bacterial genome sequencing: an embarrassment of choice, a world of opportunity Nature Reviews. Nat Rev Microbiol 10, 599–606 (2012).
Swann, J. R. & Claus, S. P. Nutrimetabonomics: nutritional applications of metabolic profiling. Sci Prog 97, 41–47 (2014).
Rezzi, S., Ramadan, Z., Fay, L. B. & Kochhar, S. Nutritional metabonomics: applications and perspectives. J Proteome Res 6, 513–525 (2007).
Meng, Y. et al. Variations in gut microbiota and fecal metabolic phenotype associated with depression by 16S rRNA sequencing and LC/MS-based metabolomics. J Pharmaceut Biomed 138, 231–239 (2017).
Han, J., Meng, J., Chen, S. Y., Li, C. & Wang, S. Rice straw biochar as a novel niche for improved alterations to the cecal microbial community in rats. Sci Rep 8, 16426 (2018).
Scott, T. W., Ward, P. F. V. & Dawson, R. M. C. The formation and metabolism of phenyl-substituted fatty acids in the ruminant. Biochem J 90, 12–24 (1964).
Cardona, F. et al. Benefits of polyphenols on gut microbiota and implications in human health. J Nutr Biochem 24, 1415–1422 (2013).
Mahowald, M. A. et al. Characterizing a model human gut microbiota composed of members of its twodominant bacterial phyla. P Natl Acad Sci USA 106, 5859–5864 (2009).
Stoupi, S., Williamson, G., Drynan, J. W., Barron, D. & Clifford, M. N. A comparison of the in vitro biotransformation of (−)-epicatechin and procyanidin B2 by human faecal microbiota. Mol Nutr Food Res 54, 747–59 (2010).
Byerley, L. et al. Walnut consumption changes the relative abundance of bacteroidetes and firmicutes in the gut. FASEB J 29(1006), 1 (2015).
Cui, C., Shen, C. J., Jia, G. & Wang, K. N. Effect of dietary Bacillus subtilis on proportion of Bacteroidetes and Firmicutes in swine intestine and lipid metabolism. Genet Mol Res 12, 1766–1776 (2013).
Selma, M. V., Espin, J. C. & Tomás-Barberán, F. A. Interaction between phenolics and gutmicrobiota: role in human health. J Agr Food Chem 57, 6485–6501 (2009).
Dai, Z. L., Wu, G. & Zhu, W. Y. Amino acid metabolism in intestinal bacteria: links between gut ecology and host health. Front Biosci 16, 1768–1786 (2011).
Ruttanavut, J., Yamauchi, K., Goto, H. & Erikawa, T. Effects of dietary bamboo charcoal powder including vinegar liquid on growth performance and histological intestinal change in Aigamo ducks. Int J Poult Sci 8, 229–236 (2009).
Leng, R. A., Preston, T. R. & Inthapanya, S. Biochar reduces enteric methane and improves growth and feed conversion in local “Yellow” cattle fed cassava root chips and fresh cassava foliage. Livest Res Rural Dev 24 (2012).
Sachan, D. S. & Davis, C. L. Hydrogenation of linoleic acid by a rumen spirochete. J Bacteriol 98, 300–301 (1969).
Ward, P. F., Scott, T. W. & Dawson, R. M. The hydrogenation of unsaturated fatty acids in the ovine digestive tract. Biochemistry J 92, 60–68 (1964).
Devkota, S. & Chang, E. B. Interactions between diet, bile acid metabolism, gut microbiota, and inflammatory bowel diseases. Digest Dis 33, 351–356 (2015).
Robinson, L. E., Buchholz, A. C. & Mazurak, V. C. Inflammation, obesity, and fatty acid metabolism: influence of n-3 polyunsaturated fatty acids on factors contributing to metabolic syndrome. Appl Physiol Nutr Me 32, 1008–1024 (2007).
Kanai, T., Mikami, Y. & Hayashi, A. A breakthrough in probiotics: Clostridium butyricum regulates gut homeostasis and anti-inflammatory response in inflammatory bowel disease. J Gastroenterol 50, 928–939 (2015).
Klaassen, C. D. & Aleksunes, L. M. Xenobiotic, bile acid, and cholesterol transporters: function and regulation. Pharmacol Rev 62, 1–96 (2010).
Shimizu, K. Dynamics of steroid hormone biosynthesis from cholesterol in adrenal tissue. Steroid Dynamics, 481–492 (1966).
Shidoji, Y., Sasak, W., Silverman-Jones, C. S. & De Luca, L. M. Recent studies on the involvement of retinyl phosphate as a carrier of mannose in biological membranes. Ann NY Acad Sci 359, 345–357 (1981).
Xu, S. Y., Bian, R. L. & Chen, X. Pharmacological experiment methodology 25–28 (People’s Medicine, 2002).
Peiffer, J. A. et al. Diversity and heritability of the maize rhizosphere microbiome under field conditions. Proc Natl Acad Sci USA 110, 6548–6553 (2013).
Acknowledgements
The authors are grateful for the financial support from the National Key Research and Development Projects (2017YFD0200800), the Earmarked Fund for Modern Agro-industry Technology Research System (CARS01-46), and the Shenyang Science and Technology Project (17-182-9-00).
Author information
Authors and Affiliations
Contributions
J.H. and J.M. designed the experiment; J.H. analyzed the data and wrote the paper; S.Y.-C. and C.L. conducted the experiment; all authors have carefully read and agree to be accountable for all aspects of the work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Han, J., Meng, J., Chen, S. et al. Integrative analysis of the gut microbiota and metabolome in rats treated with rice straw biochar by 16S rRNA gene sequencing and LC/MS-based metabolomics. Sci Rep 9, 17860 (2019). https://doi.org/10.1038/s41598-019-54467-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-54467-6
This article is cited by
-
Gut microbiome and plasma metabolome changes in rats after oral gavage of nanoparticles: sensitive indicators of possible adverse health effects
Particle and Fibre Toxicology (2022)
-
Serum metabolic traits reveal therapeutic toxicities and responses of neoadjuvant chemoradiotherapy in patients with rectal cancer
Nature Communications (2022)
-
Cage bedding modifies metabolic and gut microbiota profiles in mouse studies applying dietary restriction
Scientific Reports (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.