Abstract
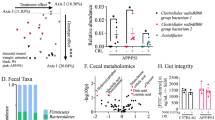
Alzheimer’s disease (AD) affects an estimated 5.8 million Americans, and advanced age is the greatest risk factor. AD patients have altered intestinal microbiota. Accordingly, depleting intestinal microbiota in AD animal models reduces amyloid-beta (Aβ) plaque deposition. Age-related changes in the microbiota contribute to immunologic and physiologic decline. Translationally relevant dietary manipulations may be an effective approach to slow microbiota changes during aging. We previously showed that calorie restriction (CR) reduced brain Aβ deposition in the well-established Tg2576 mouse model of AD. Presently, we investigated whether CR alters the microbiome during aging. We found that female Tg2576 mice have more substantial age-related microbiome changes compared to wildtype (WT) mice, including an increase in Bacteroides, which were normalized by CR. Specific gut microbiota changes were linked to Aβ levels, with greater effects in females than in males. In the gut, Tg2576 female mice had an enhanced intestinal inflammatory transcriptional profile, which was reversed by CR. Furthermore, we demonstrate that Bacteroides colonization exacerbates Aβ deposition, which may be a mechanism whereby the gut impacts AD pathogenesis. These results suggest that long-term CR may alter the gut environment and prevent the expansion of microbes that contribute to age-related cognitive decline.
Similar content being viewed by others
Introduction
The gut-brain axis is an integrated network in which the microbiota and central nervous system communicate via endocrine, immune, and neural signaling pathways1. Several translational studies show that transferring microbiota from patients with neurodevelopmental and neurological disorders including autism, multiple sclerosis, and Parkinson’s disease can influence behavior, motor dysfunction, and immune responses in relevant animal models2,3,4,5. These studies provide evidence that intestinal microbiota may play an etiologic role in diseases that emerge at differing points during the lifespan. Consistent with this notion, Alzheimer’s disease (AD) patients have altered gut microbiota compared to age-matched healthy subjects6,7. In established animal models of AD, depleting the microbiota either in germ-free or antibiotic-treated mice served as protection against the pathological hallmark amyloid-beta (Aβ) plaque deposition8,9.
While host genotypes influence AD risk, the most important risk factor is advanced age10. In older adults, the microbiota is less diverse11, and immunosenescence and age-related changes in host physiology can destabilize the microbiota12,13. An ‘aged’ microbiota promotes immune dysfunction, including increased systemic inflammation and impaired macrophage phagocytosis, which can be partially restored by transferring microbiota from young to aged mice14. Thus, understanding how to slow or reverse age-related changes in the gut microbiota has therapeutic implications for age-related brain diseases, including AD.
Diet is a major environmental factor that modulates the microbiota and has been proposed to prevent age-related changes of the microbiota11. Calorie restriction (CR), characterized by 20–40% reduction of total calorie intake without malnutrition, increases the healthspan and lifespan in multiple model organisms15. A 30% reduction in calories from carbohydrates activates neuroprotective signatures and suppresses age-related transcriptional changes in the hippocampus in wildtype (WT) mice16. In the context of AD, we found that CR prevents Aβ plaque accumulation and modulates the expression of the gamma-secretase complex, the amyloid-beta precursor protein (APP) processing enzymes, in a sex-dependent manner in Tg2576 mice17. In addition to effects on host physiology, CR modulates the microbiota and increases abundances of bacteria that positively correlate with lifespan18. However, the association between CR, the microbiome, and AD pathogenesis has not been established.
In this study, we investigated the effect of long-term 30% CR compared with ad libitum (AL) feeding on the microbiome in aging. We studied the Tg2576 model, where a mutant variant of the human APP originally identified in a Swedish family with early-onset AD (APPswe) is expressed in transgenic mice19,20. This transgene results in cerebral amyloid accumulation, synaptic loss, and cognitive impairment by 12 months of age (MO)20,21. We now interrogate whether the CR effects on the intestinal microbiome in aging can be related to the susceptibility to pathologic lesions in the brain. We also investigated how the microbiota changed with age in WT littermates which do not develop Aβ pathology. This study demonstrates for the first time that female Tg2576 mice show enhancement of age-related microbiota changes compared to WT littermates, and that CR reverses age- and Aβ-related changes in the gut microbiota.
Results
Diet, APPswe, and sex shapes the microbiota in a model of AD
We administered a 30% CR diet (with reduction in carbohydrates only) to male and female Tg2576 and nontransgenic WT littermates, initiating the diet at 2.5–3 MO and randomly assigning the diet without prior knowledge of microbiota composition. We sequenced the microbial 16S rRNA gene from longitudinally collected fecal samples at 11 different time points until mice reached 15 MO (n = 15–17 per group, with n = 7–9 per group followed until 15 MO). Differences in overall microbiota communities (β-diversity) were determined by comparing unweighted UniFrac distances. By Permanova testing, we found that microbiota communities differed significantly (p < 0.05) in mice by genotype, diet, and biologic sex (Fig. 1a–e and Supplementary Fig. 1). In addition to three-dimensional visualization of community structure, pair-wise UniFrac distances were used to examine the overall dissimilarity of microbiota composition (Fig. 1f–h). Differences between the groups (gray bars, e.g. WT mice versus Tg2576 mice stratified by sex and diet) were significantly larger than within-group differences (e.g., WT mice compared to other WT mice), indicating significant effects of genotype, diet, and sex on microbiota composition (Fig. 1f–h). In addition, while both genotypes showed similar intra-group UniFrac distances (Fig. 1f), CR-fed mice had reduced microbiota variation compared to AL-fed WT and Tg2576 mice, indicating that CR selects for a more homogeneous microbiota regardless of genotype.
Genotype, diet, and sex shape microbiota structure. Differences between microbiota were visualized by PCoA of unweighted UniFrac distances (panels a–e). (a) Overall stratification by diet and mouse genotype. (b) Tg2576 mice show altered microbiota from littermate WT mice, regardless of diet. (c) Diet shifts the microbiota in both WT and Tg2576 mice. (d,e) Male and female mice show differences in microbiota. (b–e) Clusters were significantly different by Permanova test, p < 0.05. (f) Unweighted UniFrac distances between WT and Tg2576 female mice (gray bars) were larger than within genotype distances. (g) CR reduces intragroup microbiota variation compared to AL-fed mice. (h) UniFrac distances between males and females were larger than intragroup distances within each sex. Bonferroni adjusted t-test, *p < 0.05, ***p < 0.001.
Examining α-diversity over the course of the study, Tg2576 mice showed consistently lower phylogenetic diversity than WT mice at multiple time points (Supplemental Fig. 2), although overall changes were not statistically significant. Conversely, CR-fed mice showed slightly higher α-diversity. High α-diversity has been associated with health, whereas lower α-diversity has been associated with obesity22 and inflammatory bowel disease. Thus, while the effect sizes are small, CR appears to restore the α-diversity losses observed in Tg2576 female mice.
The effect of CR on the microbiota in aging
Next, we examined whether there were any consistent age-related changes in the microbiota (Fig. 2). Microbiota composition in females from 3 to 15 MO is shown in Fig. 2a. Mice were individually housed throughout the course of the study, eliminating cage effects, and thus we could measure microbiota drift in a single animal over time. To assess how the microbiota in a single mouse diverges as they age (Fig. 2b), we calculated the unweighted UniFrac distances between the microbiota at study day 59 (5 MO) and later microbiota samples (7–15 MO) on a per-mouse basis. We selected study day 59 because there is a substantial weight loss during the initial 30 days of CR, followed by stabilization in weight. We found increased microbiota divergence in aging AL-fed Tg2576 mice (positive slope, significantly non-zero, linear regression), whereas CR-fed Tg2576 mice and all other groups did not show a significant increase in divergence over time.
The effect of CR on the aging microbiota in WT and Tg2576 mice. (a) Composition of the fecal microbiota from study day 0 (~3 MO) until study day 369 (~15 MO). Mice were individually housed throughout the experiment, eliminating cage effects. (b) Age-related microbiota drift. Divergence from microbiota at study day 59 was measured using unweighted UniFrac distances on a per-mouse basis; slopes were calculated and tested by linear regression whether significantly non-zero. *p < 0.05. (c–f) Microbial taxa that are increased in 5 MO or 15 MO female mice, LEfSe p < 0.05. Each cladogram represents all taxa detected at >0.1%, shown at the Kingdom phylogenetic level through the genus level. A yellow circle depicts taxa present, but not enriched. Red circles are enriched in aging, and green enriched in young animals. The size of the circle corresponds to the population of each taxon.
To determine the basis for this variation, we identified specific taxa (at the phylum to genus level) that changed with aging (Fig. 2c–f). In AL-fed females, we found that age increased the order Bacteroidales and genus Alistipes and decreased the order Bacillales compared to young mice (Fig. 2c,d). These changes occurred in both the WT and Tg2576 mice, indicating that these specific age-related microbiota changes are independent of APPswe expression. Similar increases in Bacteroides were found in humans in association with both aging and AD6,13. In AL-fed Tg2576 female mice, additional age-related changes not observed in WT littermates included decreases in the phylum Firmicutes and an increase in lower order taxa in the phylum Bacteroidetes. Genera within Firmicutes including Faecalibaculum, Turicibacter, Lachnospiraceae, Lactococcus, Staphylococcus, and Sporosarcina and genera within Bacteroidetes including Bacteroides, Odoribacter and Parabacteroides appeared to drive overall differences (Fig. 2d). CR-fed mice had less substantial change in their microbiota than AL-fed mice (Fig. 2e,f). With the exception of Planococcaceae and genus Staphylococcus, nearly all of the age-related microbiota differences observed in AL-fed females were rescued in the aged CR-fed mice. Both CR-fed WT and Tg2576 mice showed increased Clostridium sensu stricto 1 and Lachnospiraceae NK4B4 group in aging. In summary, these data indicate that in female mice, CR substantially and consistently reduces the age-related changes in the microbiota, especially in Tg2576 mice. We also identified specific taxa that changed during aging in male mice (Supplementary Fig. 3). With aging, all males showed an increase in Ruminococcaceae UCG_004 and Lachnospiraceae NK4B4 group, and CR-fed males showed a decrease in Bacteroides and Eubacterium xylanophilum. However, all other changes were inconsistent between the 4 different groups, suggesting that the effect of aging and CR on the microbiota in male mice is less uniform than in females.
Protective effects of CR are sex-specific17. Moreover, sex-specific microbiota differences can influence chronic disease23. Therefore, we examined microbiota differences between males and females. Female mice have higher Gammaproteobacteria than males regardless of age, genotype, or diet (Supplementary Fig. 4). With aging, there are broad phylum level changes in AL-fed mice, with higher Proteobacteria and lower Firmicutes in 15 MO WT and Tg2576 females compared to males. Aged AL-fed WT and Tg2576 females have higher Bacteroides and lower Faecalibaculum compared to males, but these sex-specific differences were not observed with CR. These data indicate that the age-related depletion in Faecalibaculum and increase in Bacteroides observed in females are not present in males and that CR can restore the levels of these microbes to those seen in WT aged males (Supplementary Fig. 4c).
Sex-specific associations between AD pathology and the microbiota in aging
Investigating the presence of Aβ plaques in Tg2576 mice by histology and ELISA showed that CR significantly reduced Aβ deposition in the brains of female but not male mice (Fig. 3a–c)17. Because microbiota composition can drive sex-specific disease susceptibility23,24, we asked whether the protective effect of CR in female mice was related to the microbiota. We used a random forest test with the Boruta algorithm25 to identify microbiota operational taxonomic units (OTUs) associated with brain Aβ40 and Aβ42 levels. In female mice, Faecalibaculum, Bacteroides, Lactobacillus, Enterococcus, Lactococcus, Parabacteroides goldsteinii, and Desulfovibrionaceae were associated with Aβ40 or Aβ42 levels in the hippocampus (Fig. 3c). Both males and females shared significantly associated OTUs belonging to Faecalibaculum, Bacteroides, Romboutsia, and Parabacteroides. In male mice, there were additional OTUs associated with Aβ, including those identified as Ruminococcaceae, Alloprevotella, Staphylococcus, Lachnclostridium, Muribaculaceae, and Clostridium. Several taxa from samples collected before the expected development of amyloid plaques, including Faecalibaculum, Bacteroides, Enterococcus, Desufovibronaceae and Parabacteroides were associated with Aβ, consistent with the hypothesis that changes in the microbiota that precede disease development may influence neuropathology.
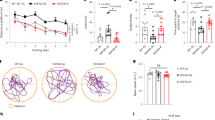
CR modulates plaque load and microbiota in a sex-specific manner. (a) Aβ40 and Aβ42 levels are reduced in the entorhinal cortex by CR (blue) in female Tg2576 mice, n = 7–8/group, T-test, as measured by ELISA (AL, black). (b) CR decreases Aβ plaque load in the hippocampus in females, as determined by immunohistochemistry. (c) Aβ40 and Aβ42 levels in the entorhinal cortex. Panels a–c modified from Schafer et al., Neurobiology of Aging 2015 with permission. (d) OTUs from younger (Early, 5–10 MO) or older (Late, 12–15 MO) mice, that were predictive of Aβ40 and Aβ 42 levels in the brain. (e) Microbiota that correlate with Aβ40 and Aβ42 levels differ by sex. *p < 0.05, ** < 0.01, *** < 0.001, Spearman correlation. (f) AL-fed Tg2576 female mice (red) show accelerated age-related changes in Faecalibaculum and Bacteroides compared to WT (black) and are reversed by CR (orange). (g) Males show minimal age-related bacterial patterns that correlate with Aβ plaque burden.
To determine whether the direction of these relationships was direct or inverse, we correlated bacterial populations with Aβ40 and Aβ42 levels in the brain (Fig. 3d). Results indicated several taxa were specifically linked with either low Aβ (green ellipse) or high Aβ (red ellipse). In females, Faecalibaculum and Lactobacillus had the strongest association with Aβ protection, and Bacteroides and Lachnospiraceae had the strongest association with elevated pathogenic Aβ levels.
We next asked whether bacteria associated with Aβ changed in abundance over time and whether this was affected by long-term CR. Faecalibaculum and Bacteroides were the two most abundant taxa in this study. Following stabilization on the control diet (Supplementary Fig. 5), Faecalibaculum decreased in WT-AL mice from 13–15 MO (Fig. 3e). Tg2576 mice had an accelerated age-related decrease in Faecalibaculum, starting at 9 MO with levels dropping 10-fold. However, in CR-fed Tg2576 female mice, the diet restored Faecalibaculum levels in aging above those observed in young WT mice. We also found that Bacteroides levels nearly doubled during aging in AL-fed WT females and more than doubled with age in Tg2576 mice. However, CR reduced Bacteroides in Tg2576 mice to levels observed in young WT mice. In humans, Bacteroides increase with aging13 and in AD6, and their toxins have been hypothesized to play a role in AD pathogenesis26. In male mice, the strongest correlations with Aβ levels were with Staphylococcus and Erysipelotrichaceae. These bacteria represented minor populations (<1% in Tg2576 mice) and did not show altered population levels with aging or with respect to diet (Fig. 3f). In summary, CR suppresses age-related changes in the taxa that were linked with AD pathology in female mice.
Microbiota predicted functions associated with AD pathology
To investigate microbial functional changes, we used PICRUSt27 to reconstruct a predicted microbial metagenome and we classified genes into KEGG orthologous groups (KOs) and pathways28. We found pathways involved in amino acid metabolism and tRNA biosynthesis linked with Aβ40 and Aβ42 levels in female Tg2576 mice (Fig. 4). Specific amino acid metabolites produced by the intestinal microbiota can modulate immunity in the central nervous system29,30, thus the effect of CR on these putative functions may be important for AD. We also found base excision repair, pantothenate biosynthesis, peptidoglycan synthesis, and terpenoid biosynthesis linked with Aβ levels, implicating microbial DNA damage, vitamin production, immune signaling, and antioxidant production in the APPswe-related Aβ pathology. Both male and female mice showed associations between the insulin signaling pathway, apoptosis, photosynthesis, and dibasic amino acid metabolism and Aβ pathology. In male mice, we found an association with sulfur, terpenoid, and sphingolipid metabolism, the peroxisome pathway, and Vibrio cholerae pathogenic cycle, suggesting that proinflammatory mediators, or changes in lipid or antioxidant metabolism could shape the Aβ response, as opposed to protein metabolism which was detected in females. Paralleling the changes in specific microbes, we found that Tg2576 mice show accelerated age-related changes in microbiota functions, which were reversed by CR.
CR modulates intestinal transcriptional signatures in aged female mice
We sought to characterize the global effects of CR on gut immune responses, as this could be an intermediary between the microbiota and the brain. We measured gene expression in the ileum of 15 MO WT and Tg2576 female mice using the nCounter Nanostring Immunology Panel of 547 genes related to inflammation and immune function. Results revealed that CR disproportionally led to downregulation in intestinal expression of those genes in both WT (35 genes) and Tg2576 female mice (51 genes), with only 3 genes upregulated (Fig. 5a–c). In both WT and Tg2576 mice, the pro-apoptotic genes BH3 interacting-domain death agonist (Bid) and cullin-9 (Cul9) were downregulated, as well as the membrane attack complex component: complement factor 7 (C7), which promotes programmed cell death via a Bid-dependent pathway31 (Fig. 5d). CR also upregulated retinoic alpha-related orphan receptor c (Rorc), the transcription factor for Th17 cells, which contributes to defense against intestinal pathogens32. Examining the effect of APPswe expression, there were multiple genes that were significantly altered between AL-fed Tg2576 mice and WT mice (37 upregulated and 11 downregulated), whereas only 6 genes were altered in parallel between CR-fed Tg576 and WT mice (1 upregulated and 5 downregulated), consistent with the hypothesis that CR reverses APP-driven changes within the intestine in aged mice (Fig. 5e,f). Several genes increased in AL-fed Tg2576 mice compared to AL-fed WT mice were reduced by CR, including antiviral responsive genes interferon alpha1 (Ifna1), interleukin-28a (IL28a, also known as interferon lambda 2), and Myxovirus resistance 1 (Mx1) (Fig. 5g). Ecto-5′-nucleotidase (NT5E), which converts adenosine-monophosphate into adenosine33, was also elevated in AL-fed Tg2576 mice, and reduced by CR. In summary, we demonstrate for the first time that an animal model of AD shows an inflammatory intestinal transcriptomic profile compared to WT mice, and that CR can reverse this effect.
CR reverses APP-associated gene expression in the intestine. Ileal gene expression was measured by Nanostring nCounter analysis in 15 MO female mice. (a) Enumeration of significantly upregulated or downregulated genes modulated by diet in WT or Tg2576 mice and modulated by genotype in AL- or CR-fed mice. (b,c) Expression levels of genes that are modulated by diet in WT (b) and in Tg2576 (c) mice. (d) Selection of genes altered by diet in both WT and Tg2576 mice. (e,f) Expression levels of genes that differ between WT and Tg2576 littermates in AL (e) and in CR (f) mice. (g) Selection of genes that show altered expression in AL-fed Tg2576 mice compared to AL-fed WT mice, which are reduced to WT levels with a CR diet. *p < 0.05, **p < 0.01, ***p < 0.001 t-test.
We also examined transcriptional changes in male mice (Supplementary Fig. 6). Consistent with the effects observed in females, CR upregulated T cell transcription factor Rorc and downregulated the pro-apoptotic gene Bid. Unlike CR-fed female mice who showed a global decrease in intestinal expression of immune genes, CR-fed males had a similar number of upregulated and downregulated genes. Examining differences related to APP, Tg2576 males had many genes that were downregulated compared to WT, including T-cell markers, Cd3e, and Ccr9, which plays a role in T and plasma cell homing to the gut. Unlike females, CR in males did not reverse AD-related transcriptional changes in the gut.
B. fragilis increases Aβ plaque deposition in female mice
Because Bacteroides showed the greatest abundance increase in aging in the AL-fed Tg2576 females, which we linked with Aβ plaques, we tested whether B. fragilis could drive plaque deposition in vivo. We administered the Type strain of B. fragilis DSM2151 or pre-reduced anaerobically sterilized (PRAS) saline as control to 2.5 MO APP/PS1 female mice. The 2.5 MO point was selected to accelerate Aβ pathogenesis at the time of disease onset but before cognitive signs would become apparent. We selected the APP/PS1 construct in order to accelerate the histopathological endpoints for AD. These APP/PS1 mice have the same APPswe mutation (KM670/671 NL) as Tg2576 mice, and in addition have the L166P mutation in PSEN1 (encoding a subunit of γ-secretase) under the control of the endogenous Thy1 promoter. These mice begin to show the first signs of Aβ pathology in the cortex at 2 MO and in the hippocampus by 4 MO. We assessed plaque deposition by immunohistochemistry with fluorescent antibodies and found that the mice that received B. fragilis had increased plaque size in the cortex (Fig. 6). This provides evidence that B. fragilis, which is enriched in aging and AD, may contribute to disease pathogenesis.
Bacteroides fragilis promotes Aβ deposition. (a) Merged fluorescent image of Aβ (red) and nuclei (blue). (b,c) Cortical (b) and hippocampal (c) regions were outlined in 3 brain sections per mouse and plaque area and number were quantified in ImageJ. Aβ-labeled plaques are shown in red, region of interest and automatically detected plaques shown in yellow. p-value listed for Student t-test, n = 3–4 mice/group.
Discussion
The composition of the microbiota changes with age in humans12 and in relevant animal models14,34, consistent with the hypothesis that dysfunction in the microbiome could contribute to age-related diseases, including AD. Lifestyle modifications that reduce the risk of AD, including diet and exercise, can modulate the aging microbiota7, which may prove to be major public health prevention strategies. We demonstrate that female mice with hallmark AD lesions develop substantially larger changes in their microbiota with aging, and that age-related changes in the microbiota could be rescued by a CR diet.
We found that Bacteroides abundance correlates with aging and plaque pathology. In humans, Bacteroides increase with both aging13 and AD6,35, and their toxins have been hypothesized to play a role in AD26. We demonstrate that the Type strain of Bacteroides fragilis can increase Aβ plaques in the brain. This is consistent with our hypothesis that beneficial effects female mice receive from CR are partially mediated by controlling age-related changes in the microbiome. Bacteroides species have highly varied carbohydrate and polysaccharide utilization genes which are important in the establishment and maintenance of colonization36,37. The CR diet limited calories only in carbohydrates, which likely directly impacted Bacteroides colonization.
Gut microbiome alterations have been observed in patients with AD6,7,38. Levels of Bacteroides are elevated in AD and correlate with blood levels of Aβ levels6,35. This is consistent with our finding that Bacteroides was increased in aging associated with higher plaque burden in females. Paradoxically, some studies have found lower levels of Bacteroides in AD patients7,32. Bacteroides can have differential effects in health and disease, based on strain variation. For example, B. fragilis strains producing polysaccharide A induce regulatory T cells, reduce inflammation, and improve disease severity in experimental autoimmune encephalomyelitis, the animal model for multiple sclerosis39. In contrast, enterotoxigenic B. fragilis strains promote inflammation40. Further studies to examine the role of different strains of Bacteroides in the pathogenesis of AD are warranted.
Altered microbiota has been observed in several AD animal models41,42,43,44,45,46,47. Treating APP/PS1 mice with broad spectrum low-dose antibiotics led to decreased Aβ plaque deposition in male but not female mice with the protective effect associated with increased Allobaculum, Akkermansia, and Lachnospiraceae9. In another study, WT mice had higher levels of Allobaculum compared to APP/PS1 mice and transfer of WT microbiota decreased Aβ plaques in APP/PS1 mice8. The genus Faecalibaculum, which we identified as potentially protective, was only discovered from culturing the mouse microbiota in 201548. There is little known about Faecalibaculum and it is missing from the commonly used taxonomic database, GreenGenes49. When GreenGenes is used for taxonomic assignment, this bacterium is classified as Allobaculum, a bacterial taxon originally isolated from a canine50. Thus, it is possible that other studies reporting findings on Allobaculum could in fact be identifying Faecalibaculum. Allobaculum has been associated with both increased lifespan in response to CR and improved metabolic health in a model of antibiotic-induced obesity18,51, and was depleted in other animal models of AD9, consistent with the hypothesis that these are potentially protective bacteria. Taken together, these studies indicate microbiota alterations drive Aβ pathogenesis and suggest that Faecalibaculum or closely related species could be key players, which have translational implications.
Altered microbiota composition in aging and AD implies altered microbiota function, which could contribute to AD. We identified microbiota pathways involved in amino acid metabolism, vitamin production, and antioxidant production linked with Aβ levels in the brain that showed enhanced age-related changes in AD mice. This suggests that the AD brain may be more vulnerable to pathogenic fluctuations in the gut microbiota associated with aging, providing a clear role for peripheral changes mediating brain-level alterations in AD. Microbial amino acid metabolism modulates inflammation in multiple sclerosis animal models30, and the gut microbiota can alter oxidative stress, known to enhance neurodegeneration in AD52. Specifically, we observed an age-related increase in predicted microbial genes involved in isoleucine biosynthesis in Tg2576 mice, which was reversed by CR. It has recently been shown that the sodium oligomannate prevents Aβ accumulation, tau phosphorylation, an cognitive decline in an animal model of AD, which was linked with a reduction in microbially produced isoleucine53. Thus, altered microbiota homeostatic metabolism in aging may be one mechanism by which the microbiota contributes to AD.
Another mechanism by which the aging gut microbiota may contribute to AD is by modulating intestinal and systemic immunity. Microbiota from aged mice elevate circulating inflammatory cytokines and impair macrophage phagocytosis when transferred to young mice14. Systemic inflammation can contribute to cognitive decline in AD54, and depleting the microbiota can modulate immunity in the central nervous system in animal models of AD8,9,55. We establish a mechanistic link between AD pathology and intestinal inflammation. Specifically, we find an inflammatory intestinal gene signature in aged Tg2576 female mice compared to WT littermates, which was abolished by CR. In particular, we found increased expression of Ifna1, IL-28a and their downstream target Mx1, which are involved in Th1 inflammatory responses, as well as Nt5e (also known as CD73), which substantially regulates extracellular adenosine. Since cell injury releases pro-inflammatory ATP, which is then converted by CD39 to AMP and to (anti-inflammatory) adenosine by Nt5e, elevated Nt5e levels are a marker of increased tissue damage and inflammation33. In the brain, Nt5e can activate, rather than repress, microglia in a Parkinson’s disease model and contribute to neuronal toxicity31. Centenarian humans have lower Nt5e levels compared to octogenarians56, suggesting that a physiologic decrease is associated with normal, healthy aging.
Aging leads to a loss of gut barrier function and dysregulation at the mucosal surfaces57, which could contribute to AD. Aged microbiota increases both intestinal inflammation including complement activation34 and intestinal permeability, which lead to circulating microbial cell wall components and systemic inflammation14. CR increases gut barrier function58, and reduces serum lipopolysaccharide binding protein in aged mice18. Decreased intestinal expression of pro-apoptotic genes Bid, Cul931, and complement factor C7 that we observed in both aged WT and Tg2576 mice, suggestsing that CR may lower inflammation in female mice and increase intestinal cell survival. CR also increased the expression of Rorc, the Th17 cell transcription factor, which plays a critical defensive role against intestinal pathogens32. Thus, CR appears to modulate intestinal physiology to limit inflammation and maintain defenses in females. In both males and females, CR reduced Bid and increased Rorc. However, beyond these genes, few changes were conserved. A major sex-specific difference is that AL-fed female Tg2576 mice show increased expression of inflammatory genes compared in WT, which is reversed by CR. In contrast, AL-fed Tg2576 male mice show decreased expression of inflammatory genes compared to WT, which is not altered by CR.
Age-related microbiota changes were more pronounced in Tg2576 females than in males, despite being littermates. AD is more prevalent in females in both humans59,60 and in animal AD models61. This study of sex-specific microbiota associations in AD models reinforces the importance of studying sex differences, which may have therapeutic implications24. CR improves gut barrier function, reduces inflammation, alters the microbiota18, and extends lifespan in females62. Sexually-dimorphic gastrointestinal tract aging critically mediates the benefits that females derive from this diet62. CR also reduces Aβ plaque in Tg2576 females but not males16, which we now link to the microbiota. Sex-specific differences in the microbiota can contribute to disease. In the NOD mouse model of type 1 diabetes, females have a higher disease incidence than males, but can be protected when colonized with male microbiota, indicating that sex-dependent changes in the microbiota can drive disease pathology23. Furthermore, broad-spectrum antibiotics protect male, but not female mice from Aβ accumulation9, and the microbiota modulate transcription in microglia, the key immune cells of the brain, in a sex-dependent manner63. We found that Bacteroides levels are lower in male than in female mice. Whether this microbiota difference could account for lower plaque load in males or whether B. fragilis could drive AD pathogenesis in in males remains to be determined.
A major unanswered question is whether other bacterial species selected by CR or by male sex could have a beneficial effect, thus it would be useful to assess whether transfer of whole microbiota from CR-fed mice or from aged males would decrease plaque load in aged AL- fed females. Long-term exposure to the microbiota in young animals has been achieved by co-housing an aged germ-free mouse with a young conventionally raised mouse14. The approach of co-housing males with females or co-housing CR is not feasible in the context of male to female microbiota transfer (due to risk of pregnancy), or in the CR paradigm in which the calories are controlled by providing a daily allotment of food to singly housed mice. However, an alternate microbiota could be transferred between males and females via exposure to bedding materials, or long-term administration of microbiota via oral gavage.
Although CR may be a translationally relevant, nontoxic, and inexpensive modality to prevent AD-like pathology throughout life, there are concerns with nutritive status and frailty in aging that complicate later-life interventions. Evidence from our group and others suggests that CR can prevent or reduce Aβ burden when initiated before overt plaque deposition17,64,65 or after plaque pathology has developed66. In a translational context, it would be more practical to apply CR at the onset of AD-related pathology to prevent further decline. However, we anticipate that introduction of restricted feeding at late stages of pathology and/or old age may not be universally well-tolerated. When CR is introduced at to mice 17 MO, which is commensurate with the age of emergence of memory deficits in our animal models, it leads to higher mortality, in contrast to lengthening lifespan when initiated in younger mice67.
Approaches to treat AD in aging that are microbiota-based may have profound translational utility. Our data show that CR can rescue age- and APP-related microbiome changes, implicating the microbiota as a potential therapeutic target in aging and AD. Sex-specific microbiota interactions that we and others observe with AD suggests that separate therapies may be needed for males and females. Whether or not the protective effects of CR are microbiota-mediated and whether there are similar relationships between the microbiota and tau pathology remains unknown. Although this study identifies key relationships between the microbiome and AD-related Aβ pathogenesis in aging that can be reversed by diet, applications to humans will require much further study and limitations may exist. Currently, most available probiotics contain Lactobacillus, Bifidobacterium, and Streptococcus, but administration of Lactobacillus and Bifidobacterium had no effect on cognitive function in AD patients68, suggesting that new generation probiotics with AD-specific indications remains a future approach.
Materials and Methods
Animal care
Animal protocols were approved by the Institutional Animal Care and Use Committee (IACUC) of the Nathan Kline Institute/NYU Langone Medical Center and by the IACUC at the Brigham & Women’s Hospital and were in full accordance with NIH guidelines. Tg2576 mice, which harbor a human variant of APP termed the “Swedish” mutation (APPswe), (Lys670/Asn and Met671/Leu), and nontransgenic littermates on a Swiss Webster DBA/C57BL6 F1 background were used for CR studies. APP/PS1 mice were used for the Bacteroides fragilis experiments. Mice were kept on a 12-hour light-dark cycle in temperature-controlled conditions.
CR feeding regimen
At approximately 2.5 MO, male and female, WT and Tg2576 littermates (n = 15–17 per group) were randomized to either an AL or CR diet. Mice were singly housed in order to ensure that the appropriate number of calories were consumed. For every 1 g consumed by an AL-fed animal, 0.71 g of the CR diet was administered to the CR cohort. The AL and CR diets were prepared by Research Diets (New Brunswick, NJ), and were designed with matched protein and fat levels, with a 30% reduction in carbohydrates only as previously described17. Approximately half of each group was sacrificed at 5.6 MO and the other half was sacrificed at 15 MO. For the aged-cohort we employed AL-fed WT mice (n = 9 females and 8 males), CR-fed WT mice (n = 9 females and 8 males), AL-fed Tg2576 mice n = 7 females and 8 males, CR-fed WT mice (n = 8 females and 8 males).
Assessment of Aβ plaque deposition in Tg2576 mice
At approximately 15 MO, mice were euthanized with an overdose of ketamine and xylazine and perfused transcardially with ice-cold 0.1M phosphate buffer. Brains were rapidly removed and 1 hemibrain was used for histopathologic analysis of human Aβ as previously described17. The entorhinal cortex and the hippocampus were dissected from the other hemibrain, and used for measurement of Aβ40 and Aβ42 protein levels quantified via colorimetric sandwich ELISA as previously described17. The data presented in this manuscript regarding the effect of CR on Aβ plaque load (Fig. 3a–c) is a new representation of findings originally published in Neurobiology of Aging, Schafer M. J. et al., Reduction of beta-amyloid and gamma-secretase by calorie restriction in female Tg2576 mice, 36:1293–1302, Copyright Elsevier (2015)17. Published with permission from Elsevier.
16S microbiota sequencing and analysis
Fecal microbiota samples were collected from WT and Tg2576 mice prior to dietary intervention (study day 0), then 1 day, 1 week, 1 month, and approximately every two months thereafter, and stored at −80 °C. DNA was extracted using the MoBio PowerLyzer kit. The V4 region of the 16S rRNA gene was amplified with barcoded-fusion primers and paired-end 151 base-pair reads were sequenced on the Illumina MiSeq instrument as previously described51,69. Quantitative insights for microbial ecology (QIIME) was used for quality filtering and downstream analysis for β-diversity, and compositional analysis70. Sequences were filtered for quality by trimming reads below a quality score of q20 and discarding reads shorter than 75% percent of the original length. OTUs were picked using the de-novo method and taxonomy was assigned using the Silva 132 database71. Distances between samples (β-diversity), were calculated using the phylogenetic based distance UniFrac72,73. Statistical testing for differential clustering of samples on the PCoA plots was performed using the Permanova test using 999 permutations. Significant differences in taxa modulated by genotype and diet was determined by linear discriminant analysis effect size (LEfSe)74. Metagenomic content of the microbiota samples was predicted from the 16S rRNA profiles and KEGG pathway functions were categorized at level 3 using the phylogenetic investigation of communities by reconstruction of unobserved states (PICRUSt) tool27. Operational taxonomic units (OTUs) or predicted KEGG metagenomic pathways associated with Aβ40 or Aβ42 levels in the hippocampus were identified using a random forest model and the Boruta algorithm25. Spearman correlations between microbial abundance and Aβ40 and Aβ42 levels was calculated in the R statistical framework75 using the cor.test in the stats package and plotted using the ellipse package76.
Intestinal transcriptomic profiling
Distal ileum samples were collected from 15 MO female mice, microbiota contents were removed by gentle scraping. Methods were taken to prevent RNA degradation, including using methods for inactivating RNAses on laboratory instruments and surfaces using RNAse-ZAP (ThermoFisher). Samples also were preserved in RNAlater to block the activity of tissue RNAses. RNA was extracted using the Qiagen RNeasy kit. We previously validated that our methods yielded intact RNA, as visualized as two distinct bands on a 3-[N-morpholino]propane sulfonic acid (MOPS) gel. In this study, we did not directly measure the RNA integrity number (RIN) on the bioanalyzer chip, however, our work in the past yielded high quality RNA with an average RIN of 916. We further assessed RNA quality using Nanodrop. Samples with a low 260/280 ratio indicating protein contamination, or a low 260/230 ratio indicating solvent contamination, underwent further purification using ethanol precipitation. 100 ng RNA from each sample was analyzed on the Nanostring Immunology Panel, consisting of 547 genes. Transcriptional profiles were normalized using the nSolver software using the default (i) background subtraction of the geometric mean of negative controls, (ii) normalization by positive controls, and (iii) housekeeping genes including on the Nanostring Panel. Low abundance genes with an average count less than 10 were excluded from analysis, and differential testing was performed using Student’s T-test. Heat-maps were made using R package gplots77.
Assessment of the effect of B. fragilis on Aβ plaque deposition
In order to accelerate the histopathological endpoints for AD, we used the APP/PS1 model, which have the same APPswe (KM670/671 NL) mutation in APP as the Tg2576 mice as well as the L166P mutation in PSEN1 (coding for a subunit of γ-secretase), under the control of the endogenous Thy1 promoter78. APP/PS1 mice begin to show the first signs of Aβ pathology at 2 MO in the cortex and in the hippocampus by 4 MO78,79,80,81. 2.5 MO female mice were randomly assigned to receive B. fragilis gavage (n = 4) or pre-reduced anaerobically sterilized (PRAS) saline as control (n = 3). This timepoint was selected to accelerate Aβ pathogenesis at the time of disease onset but before cognitive symptoms would become apparent. B. fragilis was given once a week for 10 weeks. Mice were housed in sterilized micro-isolator cages in a biosafety level 2 (BSL2) facility and AL-fed on standard chow. All animal experimental procedures were performed in accordance with the approved Animal Care and Use Protocols and began when the animals were approximately 10 weeks.
B. fragilis strain DSM2151 (corresponding to ATCC strain 25285) was obtained from the DSMZ-German Collection of Microorganisms. This strain was selected because it is the Type strain, the first strain of B. fragilis. This strain was originally isolated in 1898 from an appendix abscess82. Bacteria were grown for 48 h on Brucella agar plates with sheep blood, vitamin K and hemin in an anaerobic chamber, and sub-cultured in between to maintain a fresh culture with high viability. The gavage of the mice was performed 3 times during the 1st week and then once a week for 9 weeks. Each mouse received 0.2 mL of PBS or of B. fragilis inoculum with a turbidity of 0.70 ± 0.1 optical density (OD) units, which corresponds to 1 × 109 bacteria/mL.
At 5 MO, mice were euthanized with CO2 narcosis and perfused transcardially with ice-cold HBSS. Brains were rapidly removed and the left hemibrain was fixed for 24 h in 4% Paraformaldehyde (PFA), followed by an immersion in a 30% sucrose solution until they were no longer buoyant. The brain was then frozen in OCT in a dry-ice ethanol bath, and kept at −30 °C. The hemispheres were then sliced coronally with a cryostat with a width of 20 microns. After drying, the slices were kept at −30 °C. Just before staining, tissue sections were thawed and fixed with acetone. Sections were incubation with blocking buffer (10% Normal Horse Serum +2% Bovine Serum Albumin +1% glycerin +0.1% Triton X-100 in PBS), washed, and incubated overnight with at 1:300 dilution of Purified anti-β-Amyloid, 1–16 Antibody, clone 6E10 (Biolegend catalog 803001), which recognizes amino acids 3–8 of Aβ and APP. The next day, the slides were washed again, incubated with the secondary antibody Cy5 AffiniPure Goat Anti-Mouse IgG, Fcγ subclass 1 specific, (Jackson Immuno Research catalog 115-175-205) at a 1:300 dilution. Then washed, dried, and the coverslip was attached with VECTASHIELD Antifade Mounting Medium with DAPI (Vector Laboratories). After drying overnight at room temperature in the dark, slides were kept at 4 °C (then at −30 °C for long term storage). The images were taken with fluorescent Leica microscope, with 5x magnification, an exposure of 300 ms and a light intensity of 81% for DAPI and of 100% for the Cy5 channel. Using ImageJ, the hippocampus and the overlying cortex above it were analyzed separately. The background was subtracted (0.30) and the threshold was calculated automatically using the triangle algorithm. The percent area covered by the plaques within the traced region of interest (cortex or hippocampus) was calculated, and differences between B. fragilis and control were determined by t-test.
Data availability
Sequencing data from the microbiota 16S rRNA is submitted in the NCBI short-read archive under Bioproject number PRJNA573528.
References
Cox, L. M. & Weiner, H. L. Microbiota Signaling Pathways that Influence Neurologic Disease. Neurotherapeutics 15, 135–145, https://doi.org/10.1007/s13311-017-0598-8 (2018).
Sampson, T. R. et al. Gut Microbiota Regulate Motor Deficits and Neuroinflammation in a Model of Parkinson’s Disease. Cell 167, 1469–1480 e1412, https://doi.org/10.1016/j.cell.2016.11.018 (2016).
Berer, K. et al. Gut microbiota from multiple sclerosis patients enables spontaneous autoimmune encephalomyelitis in mice. Proc Natl Acad Sci USA 114, 10719–10724, https://doi.org/10.1073/pnas.1711233114 (2017).
Cekanaviciute, E. et al. Gut bacteria from multiple sclerosis patients modulate human T cells and exacerbate symptoms in mouse models. Proc Natl Acad Sci USA 114, 10713–10718, https://doi.org/10.1073/pnas.1711235114 (2017).
Sharon, G. et al. Human Gut Microbiota from Autism Spectrum Disorder Promote Behavioral Symptoms in Mice. Cell 177, 1600–1618 e1617, https://doi.org/10.1016/j.cell.2019.05.004 (2019).
Vogt, N. M. et al. Gut microbiome alterations in Alzheimer’s disease. Sci Rep 7, 13537, https://doi.org/10.1038/s41598-017-13601-y (2017).
Zhuang, Z.-Q. et al. Gut Microbiome is Altered in Patients with Alzheimer’s Disease. Journal of Alzheimer’s Disease, 1–10 (2018).
Harach, T. et al. Reduction of Abeta amyloid pathology in APPPS1 transgenic mice in the absence of gut microbiota. Sci Rep 7, 41802, https://doi.org/10.1038/srep41802 (2017).
Minter, M. R. et al. Antibiotic-induced perturbations in gut microbial diversity influences neuro-inflammation and amyloidosis in a murine model of Alzheimer’s disease. Sci Rep 6, 30028, https://doi.org/10.1038/srep30028 (2016).
Alzheimer’s, A. 2013 Alzheimer’s disease facts and figures. Alzheimers Dement 9, 208–245, https://doi.org/10.1016/j.jalz.2013.02.003 (2013).
Claesson, M. J. et al. Gut microbiota composition correlates with diet and health in the elderly. Nature 488, 178–184, https://doi.org/10.1038/nature11319 (2012).
Tiihonen, K., Ouwehand, A. C. & Rautonen, N. Human intestinal microbiota and healthy ageing. Ageing Res Rev 9, 107–116, https://doi.org/10.1016/j.arr.2009.10.004 (2010).
Odamaki, T. et al. Age-related changes in gut microbiota composition from newborn to centenarian: a cross-sectional study. BMC Microbiol 16, 90, https://doi.org/10.1186/s12866-016-0708-5 (2016).
Thevaranjan, N. et al. Age-Associated Microbial Dysbiosis Promotes Intestinal Permeability, Systemic Inflammation, and Macrophage Dysfunction. Cell Host Microbe 21, 455–466 e454, https://doi.org/10.1016/j.chom.2017.03.002 (2017).
Fontana, L., Partridge, L. & Longo, V. D. Extending healthy life span—from yeast to humans. science 328, 321–326 (2010).
Schafer, M. J., Dolgalev, I., Alldred, M. J., Heguy, A. & Ginsberg, S. D. Calorie Restriction Suppresses Age-Dependent Hippocampal Transcriptional Signatures. PLoS One 10, e0133923, https://doi.org/10.1371/journal.pone.0133923 (2015).
Schafer, M. J. et al. Reduction of beta-amyloid and gamma-secretase by calorie restriction in female Tg2576 mice. Neurobiol Aging 36, 1293–1302, https://doi.org/10.1016/j.neurobiolaging.2014.10.043 (2015).
Zhang, C. et al. Structural modulation of gut microbiota in life-long calorie-restricted mice. Nature communications 4, 2163, https://doi.org/10.1038/ncomms3163 (2013).
Citron, M. et al. Mutation of the beta-amyloid precursor protein in familial Alzheimer’s disease increases beta-protein production. Nature 360, 672–674, https://doi.org/10.1038/360672a0 (1992).
Hsiao, K. et al. Correlative memory deficits, Aβ elevation, and amyloid plaques in transgenic mice. Science 274, 99–103 (1996).
Lanz, T., Carter, D. & Merchant, K. Dendritic spine loss in the hippocampus of young PDAPP and Tg2576 mice and its prevention by the ApoE2 genotype. Neurobiology of disease 13, 246–253 (2003).
Le Chatelier, E. et al. Richness of human gut microbiome correlates with metabolic markers. Nature 500, 541–546, https://doi.org/10.1038/nature12506 (2013).
Markle, J. G. et al. Sex differences in the gut microbiome drive hormone-dependent regulation of autoimmunity. Science 339, 1084–1088, https://doi.org/10.1126/science.1233521 (2013).
Cox, L. M., Abou-El-Hassan, H., Maghzi, A. H., Vincentini, J. & Weiner, H. L. The sex-specific interaction of the microbiome in neurodegenerative diseases. Brain Res 1724, 146385, https://doi.org/10.1016/j.brainres.2019.146385 (2019).
Kursa, M. B. & Rudnicki, W. R. Feature Selection with the Boruta Package. J Stat Software 36, 1–13 (2010).
Zhao, Y. & Lukiw, W. J. Bacteroidetes Neurotoxins and Inflammatory Neurodegeneration. Mol Neurobiol 55, 9100–9107, https://doi.org/10.1007/s12035-018-1015-y (2018).
Langille, M. G. et al. Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotechnol 31, 814–821, https://doi.org/10.1038/nbt.2676 (2013).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28, 27–30 (2000).
Kennedy, P. J., Cryan, J. F., Dinan, T. G. & Clarke, G. Kynurenine pathway metabolism and the microbiota-gut-brain axis. Neuropharmacology 112, 399–412, https://doi.org/10.1016/j.neuropharm.2016.07.002 (2017).
Rothhammer, V. et al. Type I interferons and microbial metabolites of tryptophan modulate astrocyte activity and central nervous system inflammation via the aryl hydrocarbon receptor. Nat Med 22, 586–597, https://doi.org/10.1038/nm.4106 (2016).
Li, Z. et al. CUL9 mediates the functions of the 3M complex and ubiquitylates survivin to maintain genome integrity. Mol Cell 54, 805–819, https://doi.org/10.1016/j.molcel.2014.03.046 (2014).
Littman, D. R. & Rudensky, A. Y. Th17 and Regulatory T Cells in Mediating and Restraining Inflammation. Cell 140, 845–858, https://doi.org/10.1016/j.cell.2010.02.021 (2010).
Bono, M. R., Fernández, D., Flores-Santibáñez, F., Rosemblatt, M. & Sauma, D. CD73 and CD39 ectonucleotidases in T cell differentiation: Beyond immunosuppression. FEBS Letters 589, 3454–3460, https://doi.org/10.1016/j.febslet.2015.07.027 (2015).
Fransen, F. et al. Aged Gut Microbiota Contributes to Systemical Inflammaging after Transfer to Germ-Free Mice. Frontiers in immunology 8, 1385–1385, https://doi.org/10.3389/fimmu.2017.01385 (2017).
Haran, J. P. et al. Alzheimer’s Disease Microbiome Is Associated with Dysregulation of the Anti-Inflammatory P-Glycoprotein Pathway. MBio 10, https://doi.org/10.1128/mBio.00632-19 (2019).
Wexler, H. M. Bacteroides: the good, the bad, and the nitty-gritty. Clin Microbiol Rev 20, 593–621, https://doi.org/10.1128/CMR.00008-07 (2007).
Lee, S. M. et al. Bacterial colonization factors control specificity and stability of the gut microbiota. Nature 501, 426–429, https://doi.org/10.1038/nature12447 (2013).
Cattaneo, A. et al. Association of brain amyloidosis with pro-inflammatory gut bacterial taxa and peripheral inflammation markers in cognitively impaired elderly. Neurobiol Aging 49, 60–68, https://doi.org/10.1016/j.neurobiolaging.2016.08.019 (2017).
Shen, Y. et al. Outer membrane vesicles of a human commensal mediate immune regulation and disease protection. Cell Host Microbe 12, 509–520, https://doi.org/10.1016/j.chom.2012.08.004 (2012).
Sears, C. L. Enterotoxigenic Bacteroides fragilis: a rogue among symbiotes. Clin Microbiol Rev 22, 349–369, Table of Contents, https://doi.org/10.1128/CMR.00053-08 (2009).
Bauerl, C., Collado, M. C., Diaz Cuevas, A., Vina, J. & Perez Martinez, G. Shifts in gut microbiota composition in an APP/PSS1 transgenic mouse model of Alzheimer’s disease during lifespan. Lett Appl Microbiol 66, 464–471, https://doi.org/10.1111/lam.12882 (2018).
Wang, J. et al. The Effects of LW-AFC on Intestinal Microbiome in Senescence-Accelerated Mouse Prone 8 Strain, a Mouse Model of Alzheimer’s Disease. J Alzheimers Dis 53, 907–919, https://doi.org/10.3233/JAD-160138 (2016).
Shen, L., Liu, L. & Ji, H. F. Alzheimer’s Disease Histological and Behavioral Manifestations in Transgenic Mice Correlate with Specific Gut Microbiome State. J Alzheimers Dis 56, 385–390, https://doi.org/10.3233/JAD-160884 (2017).
Sanguinetti, E. et al. Microbiome-metabolome signatures in mice genetically prone to develop dementia, fed a normal or fatty diet. Sci Rep 8, 4907, https://doi.org/10.1038/s41598-018-23261-1 (2018).
Brandscheid, C. et al. Altered Gut Microbiome Composition and Tryptic Activity of the 5xFAD Alzheimer’s Mouse Model. J Alzheimers Dis 56, 775–788, https://doi.org/10.3233/JAD-160926 (2017).
Bonfili, L. et al. Microbiota modulation counteracts Alzheimer’s disease progression influencing neuronal proteolysis and gut hormones plasma levels. Sci Rep 7, 2426, https://doi.org/10.1038/s41598-017-02587-2 (2017).
Minter, M. R. et al. Antibiotic-induced perturbations in microbial diversity during post-natal development alters amyloid pathology in an aged APPSWE/PS1DeltaE9 murine model of Alzheimer’s disease. Sci Rep 7, 10411, https://doi.org/10.1038/s41598-017-11047-w (2017).
Chang, D. H. et al. Faecalibaculum rodentium gen. nov., sp. nov., isolated from the faeces of a laboratory mouse. Antonie Van Leeuwenhoek 108, 1309–1318, https://doi.org/10.1007/s10482-015-0583-3 (2015).
Mcdonald, D. et al. An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISME J 6, 610–618, https://doi.org/10.1038/ismej.2011.139 (2011).
Greetham, H. L. et al. Allobaculum stercoricanis gen. nov., sp. nov., isolated from canine feces. Anaerobe 10, 301–307, https://doi.org/10.1016/j.anaerobe.2004.06.004 (2004).
Cox, L. M. et al. Altering the intestinal microbiota during a critical developmental window has lasting metabolic consequences. Cell 158, 705–721, https://doi.org/10.1016/j.cell.2014.05.052 (2014).
Dumitrescu, L. et al. Oxidative Stress and the Microbiota-Gut-Brain Axis. Oxidative medicine and cellular longevity 2018, 2406594–2406594, https://doi.org/10.1155/2018/2406594 (2018).
Wang, X. et al. Sodium oligomannate therapeutically remodels gut microbiota and suppresses gut bacterial amino acids-shaped neuroinflammation to inhibit Alzheimer’s disease progression. Cell Res 29, 787–803, https://doi.org/10.1038/s41422-019-0216-x (2019).
Holmes, C. et al. Systemic inflammation and disease progression in Alzheimer disease. 73, 768–774, https://doi.org/10.1212/WNL.0b013e3181b6bb95%JNeurology (2009).
Dodiya, H. B. et al. Sex-specific effects of microbiome perturbations on cerebral Abeta amyloidosis and microglia phenotypes. J Exp Med 216, 1542–1560, https://doi.org/10.1084/jem.20182386 (2019).
Crooke, A. et al. Low expression of CD39 and CD73 genes in centenarians compared with octogenarians. Immunity & ageing: I & A 14, 11–11, https://doi.org/10.1186/s12979-017-0094-3 (2017).
Sato, S., Kiyono, H. & Fujihashi, K. Mucosal immunosenescence in the gastrointestinal tract: a mini-review. Gerontology 61, 336–342 (2014).
Ma, T. Y., Hollander, D., Dadufalza, V. & Krugliak, P. Effect of aging and caloric restriction on intestinal permeability. Experimental gerontology 27, 321–333 (1992).
Mielke, M. M., Vemuri, P. & Rocca, W. A. Clinical epidemiology of Alzheimer’s disease: assessing sex and gender differences. Clin Epidemiol 6, 37–48, https://doi.org/10.2147/CLEP.S37929 (2014).
Snyder, H. M. et al. Sex biology contributions to vulnerability to Alzheimer's disease: A think tank convened by the Women’s Alzheimer’s Research Initiative. Alzheimers Dement 12, 1186–1196, https://doi.org/10.1016/j.jalz.2016.08.004 (2016).
Jiao, S. S. et al. Sex Dimorphism Profile of Alzheimer’s Disease-Type Pathologies in an APP/PS1 Mouse Model. Neurotox Res 29, 256–266, https://doi.org/10.1007/s12640-015-9589-x (2016).
Regan, J. C. et al. Sex difference in pathology of the ageing gut mediates the greater response of female lifespan to dietary restriction. Elife 5 (2016).
Thion, M. S. et al. Microbiome Influences Prenatal and Adult Microglia in a Sex-Specific Manner. Cell 172, 500–516 e516, https://doi.org/10.1016/j.cell.2017.11.042 (2018).
Wang, J. et al. Caloric restriction attenuates beta-amyloid neuropathology in a mouse model of Alzheimer’s disease. FASEB J 19, 659–661, https://doi.org/10.1096/fj.04-3182fje (2005).
Patel, N. V. et al. Caloric restriction attenuates Abeta-deposition in Alzheimer transgenic models. Neurobiol Aging 26, 995–1000, https://doi.org/10.1016/j.neurobiolaging.2004.09.014 (2005).
Mouton, P. R., Chachich, M. E., Quigley, C., Spangler, E. & Ingram, D. K. Caloric restriction attenuates amyloid deposition in middle-aged dtg APP/PS1 mice. Neurosci Lett 464, 184–187, https://doi.org/10.1016/j.neulet.2009.08.038 (2009).
Forster, M. J., Morris, P. & Sohal, R. S. Genotype and age influence the effect of caloric intake on mortality in mice. FASEB J 17, 690–692, https://doi.org/10.1096/fj.02-0533fje (2003).
Agahi, A. et al. Does Severity of Alzheimer’s Disease Contribute to Its Responsiveness to Modifying Gut Microbiota? A Double Blind Clinical Trial. Front Neurol 9, 662, https://doi.org/10.3389/fneur.2018.00662 (2018).
Caporaso, J. G. et al. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J 6, 1621–1624, https://doi.org/10.1038/ismej.2012.8 (2012).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7, 335–336, https://doi.org/10.1038/nmeth.f.303 (2010).
Yilmaz, P. et al. The SILVA and “All-species Living Tree Project (LTP)” taxonomic frameworks. Nucleic Acids Research 42, D643–D648, https://doi.org/10.1093/nar/gkt1209 (2013).
Lozupone, C. & Knight, R. UniFrac: a new phylogenetic method for comparing microbial communities. Appl Environ Microbiol 71, 8228–8235, https://doi.org/10.1128/AEM.71.12.8228-8235.2005 (2005).
Lozupone, C., Lladser, M. E., Knights, D., Stombaugh, J. & Knight, R. UniFrac: an effective distance metric for microbial community comparison. ISME J 5, 169–172 (2010).
Segata, N. et al. Metagenomic biomarker discovery and explanation. Genome Biol 12, R60, https://doi.org/10.1186/gb-2011-12-6-r60 (2011).
Team, R. C. A language and environment for statistical computing (2015).
Murdoch, D. E. D. C. ellipse: Functions for drawing ellipses and ellipse-like confidence regions (2013).
Warnes, G. R. et al. Various R Programming Tools for Plotting Data. R package version 3.0.1.1, https://CRAN.R-project.org/package=gplots (2019).
Radde, R. et al. Abeta42-driven cerebral amyloidosis in transgenic mice reveals early and robust pathology. EMBO Rep 7, 940–946, https://doi.org/10.1038/sj.embor.7400784 (2006).
Gengler, S., Hamilton, A. & Hölscher, C. Synaptic plasticity in the hippocampus of a APP/PS1 mouse model of Alzheimer’s disease is impaired in old but not young mice. PLoS one 5, e9764 (2010).
Maia, L. F. et al. Changes in amyloid-beta and Tau in the cerebrospinal fluid of transgenic mice overexpressing amyloid precursor protein. Sci Transl Med 5, 194re192, https://doi.org/10.1126/scitranslmed.3006446 (2013).
Webster, S. J., Bachstetter, A. D. & Van Eldik, L. J. Comprehensive behavioral characterization of an APP/PS-1 double knock-in mouse model of Alzheimer’s disease. Alzheimers Res Ther 5, 28, https://doi.org/10.1186/alzrt182 (2013).
Veillon, A. & Zuber, A. Recherches sur quelques microbes strictement anaérobies et leur rôle en pathologie. Arch Med Exp 10, 517–545 (1898).
Acknowledgements
This work was supported by NIH grants 1UL1RR029893, AG017617, AG043375, and AG014449, the Harvard NeuroDiscovery Pilot Award, the Women’s Alzheimer’s Movement Research Fund, the Women’s Brain Initiative Fellowship at Brigham & Women’s Hospital, and by the C&D and Zlinkoff funds. We acknowledge assistance with 16S sequencing from the NYULMC Genome Technology Center and with the acquisition of fluorescent immunohistochemistry images from Dr. Lai Ding at the NeuroTechnology Studio at Brigham and Women’s Hospital. We thank Melissa Alldred, Paul Mathews, and Arthur Saltzman for assistance with implementation of CR and analysis of tissue.
Author information
Authors and Affiliations
Contributions
L.M. Cox sequenced and analyzed the microbiota 16S rRNA data and performed the Nanostring nCounter profiling. M.J. Schafer carried out the CR feeding regimen and amyloid-beta analysis. J. Sohn extracted DNA and RNA and contributed to the bioinformatic analysis for the microbiome. J. Vincentini and L.M. Cox tested the effect of B. fragilis on plaques. L.M. Cox, M.J. Schafer, H.L. Weiner, S.D. Ginsberg, and M.J. Blaser designed the study, interpreted the data, and wrote the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cox, L.M., Schafer, M.J., Sohn, J. et al. Calorie restriction slows age-related microbiota changes in an Alzheimer’s disease model in female mice. Sci Rep 9, 17904 (2019). https://doi.org/10.1038/s41598-019-54187-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-54187-x
This article is cited by
-
Bacteroidota inhibit microglia clearance of amyloid-beta and promote plaque deposition in Alzheimer’s disease mouse models
Nature Communications (2024)
-
Cardiac cell senescence: molecular mechanisms, key proteins and therapeutic targets
Cell Death Discovery (2024)
-
Gut microbiota and healthy longevity
Science China Life Sciences (2024)
-
The connection between gut microbiota and its metabolites with neurodegenerative diseases in humans
Metabolic Brain Disease (2024)
-
Influence of complement protein C1q or complement receptor C5aR1 on gut microbiota composition in wildtype and Alzheimer’s mouse models
Journal of Neuroinflammation (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.