Abstract
Bovine tuberculosis (BTB) caused by Mycobacterium bovis remains a major problem in both the developed and developing countries. Control of BTB in the UK is carried out by test and slaughter of infected animals, based primarily on the tuberculin skin test (PPD). Vaccination with the attenuated strain of the M. bovis pathogen, BCG, is not used to control bovine tuberculosis in cattle at present, due to its variable efficacy and because it interferes with the PPD test. Diagnostic tests capable of Differentiating Infected from Vaccinated Animals (DIVA) have been developed that detect immune responses to M. bovis antigens absent in BCG; but these are too expensive and insufficiently sensitive to be used for BTB control worldwide. To address these problems we aimed to generate a synergistic vaccine and diagnostic approach that would permit the vaccination of cattle without interfering with the conventional PPD-based surveillance. The approach was to widen the pool of M. bovis antigens that could be used as DIVA targets, by identifying antigenic proteins that could be deleted from BCG without affecting the persistence and protective efficacy of the vaccine in cattle. Using transposon mutagenesis we identified genes that were essential and those that were non-essential for persistence in bovine lymph nodes. We then inactivated selected immunogenic, but non-essential genes in BCG Danish to create a diagnostic-compatible triple knock-out ΔBCG TK strain. The protective efficacy of the ΔBCG TK was tested in guinea pigs experimentally infected with M. bovis by aerosol and found to be equivalent to wild-type BCG. A complementary diagnostic skin test was developed with the antigenic proteins encoded by the deleted genes which did not cross-react in vaccinated or in uninfected guinea pigs. This study demonstrates the functionality of a new and improved BCG strain which retains its protective efficacy but is diagnostically compatible with a novel DIVA skin test that could be implemented in control programmes.
Similar content being viewed by others
Introduction
Bovine TB (BTB) is one of the most complex animal health problems that the world’s cattle industry faces today and which causes significant economic losses1,2. BTB also causes zoonotic TB, prompting, in 2017, the first road map to combat zoonotic TB to be rolled out3,4,5. The active surveillance and control of BTB in most parts of the world depends on animal screening by tuberculin skin testing and the removal of test-positive animals by culling6. The tuberculin skin test is performed by injection of small quantities of M. bovis purified protein derivative (PPD-B), which is a crude and complex antigenic mixture that provokes a visible immune reaction on the skin7. To mitigate cross-reactivity with environmental mycobacteria and to optimise specificity, in the so-called comparative tuberculin test, PPD-B is sometimes supplemented by injection of PPD derived from the related Mycobacterium avium (PPD-A). Many countries also use the so-called single intradermal test relying only on the injection of PPD-B, a test format optimised for sensitivity, also often used as international trade test. Tuberculin skin testing is compulsory in many countries. Yet control by test-and-cull is expensive and societally and/or economically unacceptable in many countries. It also fails to achieve TB eradication in some epidemiological circumstances8.
Vaccination is nearly always the most cost-effective means to control infectious disease. The BCG vaccine was developed for control of human tuberculosis by Albert Calmette and Camille Guerin by repeated sub-culturing of the bovine TB bacillus, Mycobacterium bovis until it lost virulence for guinea pigs yet protected them from challenge with live virulent TB bacilli. The BCG vaccine was first used in humans in 1921 and has since become the most widely used vaccine for humans9. Experimental approaches to developing an improved vaccine against TB have included the use of attenuated mycobacteria, subunit vaccines, and DNA vaccines10,11. Several live attenuated mycobacterial vaccines have shown promising efficacy against M. tuberculosis challenge in animal models11,12,13,14,15 and have progressed to clinical trials16,17. There is also evidence that deletion of genes from BCG can result in unaltered or even superior protective efficacy compared to the corresponding parental BCG18. BCG is also effective in protecting cattle against TB and has been shown to be capable of reducing the number, duration and severity of herd breakdowns19,20,21,22. However, it is not used to control BTB since BCG shares many antigens with M. bovis23,24 resulting in the failure of the PPD skin test to distinguish between a BCG-vaccinated and BTB-infected cow25.
The discovery that the development of BCG from M. bovis involved the concomitant deletion of large regions of the BCG chromosome (the RD regions)26,27 stimulated the search for antigens that have been lost in the BCG genome and may therefore be used in a novel test capable of Differentiating Infected from Vaccinated Animals (DIVA). Antigens such as ESAT-6 and CFP10, both encoded on the RD1 region have been used as DIVA antigens28,29 in humans and cattle blood tests (interferon-gamma release assays, IGRA)28,29,30. However, a drawback of the IGRA tests is that they are laboratory-based requiring blood samples to be taken from infected animals, transported to the laboratory within a tight timeline and under temperature-controlled transport conditions. The test result must then be reported back to the field. These requirements make the IGRA DIVA tests expensive and inappropriate for developing countries with limited resources and technological infrastructure31. In contrast, a DIVA skin test would overcome these limitations28.
Recently a defined diagnostic skin test was developed based on a cocktail of DIVA antigens, ESAT-6, CFP-10 and Rv3615c, that allow the differential diagnosis of M. bovis-infected from BCG-vaccinated animals29,32,33. The specificity of this DIVA test is high but its sensitivity is less than the SIT test32,34,35. Although adding additional M. bovis antigens to the cocktail would likely increase its sensitivity, this is not a viable option since no other potential DIVA antigens have been identified within the RD regions of BCG; and adding non-DIVA M. bovis antigens to the cocktail will likely decrease the specificity of the test.
Our aim was to develop a new BCG vaccine that is compatible with a sensitive and affordable DIVA skin test. The approach we used is based on the observation that at least three M. bovis antigens (ESAT-6, CFP-10 and Rv3615c) are dispensable for the protective efficacy of BCG. This led us to hypothesize that the programme of accidental deletion of antigens from M. bovis initiated by Calmette and Guerin to develop the BCG vaccine, could be continued by rational design. We therefore initiated a project to construct a novel strain of BCG, ΔBCG, deleted in additional antigens dispensable for protection. Vaccination with ΔBCG would thereby provide us with the opportunity of adding additional antigens, deleted from the ΔBCG genome, to the current DIVA cocktail that might provide an enhanced sensitivity skin test reagent capable of detecting BTB infection in ΔBCG inoculated animals.
It is well established that persistence of BCG in the host is crucial for its protective efficacy36. In a previous paper we describe the development of a high throughput transposon mutagenesis approach that was used to identify BCG genes that influence the ability of the vaccine to persist in the bovine host37. In this paper we describe the use of that dataset to identify BCG genes encoding antigens that do not impair the ability of BCG to persist in the bovine host and are therefore candidate genes that could be deleted. We go on to describe the construction of a ΔBCG antigen-depleted strain that it is safe and induces protective immunity comparable to wild-type BCG against M. bovis infection in guinea pigs. In parallel we also describe the development of a skin test diagnostic based on detection of antigens deleted from the ΔBCG strain. This study provides a novel approach to development of rationally-designed live vaccines.
Results
The starting point for our experiments was the identification of genes that influence the survival of BCG in the bovine lymph node. The details of these experiments are fully described elsewhere37, with the method based on the original BCG lymph node challenge model38. Briefly, a BCG Danish transposon (Tn) library was constructed and inoculated into the prescapular lymph nodes of three calves. The library was recovered from lymph nodes after 3 weeks and the input and output library pools were compared by Tn-seq and TRANSIT’s Resampling method analysis39 to identify genes that, when inactivated by the transposon, influenced persistence in bovine lymph nodes40. Genes that did not influence persistence were deemed dispensable and were therefore considered candidate for deletion in the engineered ΔBCG strain. Genes in the list of dispensable loci that encoded antigens were identified by cross-checking against a list of 500 proteins whose immunoreactivity in TB-infected cows has been already characterized41,42 to identify dispensable antigenic proteins.
Five genes encoding antigens were identified as Tn mutants in the library whose fold changes during in vivo passage in cattle was between 0.5 to 2-fold (Fig. 1a), indicating that they were not essential for persistence in the bovine lymph condition, and so could be eliminated from the genome via three genome deletions without compromising the ability to persist in cattle. The genes selected were BCG3043, BCG2897, BCG2895, BCG3679 and BCG3680 which are orthologs of M.tb genes Rv3020c, MPT70, MPT83, Rv3615c and Rv3616c, respectively. Rv3020c is a member of the ESX family of virulence factors known to induce a potent cellular immune responses43. Rv3615c [Esx-1 substrate protein C (EspC)], encoded outside of RD1, is an antigen that is already part of the DIVA skin test prototyope described by Whelan et al.33. The MPB70 and MPB83 antigens were reported to be highly specific and sensitive for the detection of M. bovis infection in animals without any cross-reaction with Mycobacterium spp. found in the environment44,45,46. Lastly, Rv3616c is a CD8 antigen in mice with characteristics of the Esx family of bacterial proteins47. It is also strongly recognised by T cells from infected cattle48. We therefore chose to delete these genes from BCG Danish.
(a) Bean plot of fold changes during in vivo passage in cattle for selected gene groups and (b) Schematic representation of creating △BCG TK. (a) Black lines show the medians; white lines represent individual antigenic genes; polygons represent the estimated density of the data. The grey regions is a density plot of the data’s distribution. Values are normalised to the median value of the ‘All gene’ group. The plots were created with BoxPlotR (Spitzer, Wildenhain et al. 2014) (b) △BCG TK was created by sequential gene knock out using specialized Transduction method (Bardov et al. 2002). The Phagemid used was Phae159 and the cosmids used were p0004S, pYUB854 and pANEE001 to create △BCG 3043, ΔBCG3043/BCG2897/BCG2895 (Double knock out) and ΔBCG3043/2897/2895/3679/3680 (Triple knock out) respectively.
Construction of modified ΔBCG TK vaccine
All 5 antigen genes were removed using specialized transduction method49 by three sequential deletion steps each using vectors with different antibiotic cassettes for the selection of mutants at each stage (Fig. 1b) to make a ΔBCG TK mutant. PCR analysis of the genetic constructs50 created to make the deletion mutants were all of the expected size (Sup. Fig. 2).
Growth analysis of ΔBCG TK mutants in standard growth medium and in bovine macrophages
To confirm that the deletion of the genes did not have any growth defect, we first tested the in vitro growth kinetics of the mutant strain compared to WT BCG in a competition assay. When co-cultured with wild type BCG in 7H9 media the TK mutant did not show any loss of fitness when compared to WT (Fig. 2a). We next sought to determine whether the deletion of BCG antigenic genes influenced the survival of BCG in bovine macrophages. PBMC-derived bovine macrophages were infected with the mixture (1:1) of WT BCG and the ΔBCG TK mutant. The ΔBCG triple mutant survived in the macrophages as well as WT BCG (Fig. 2b). This confirms that the removal of antigens does not alter the in vitro or ex vivo growth characteristics of the ΔBCG TK strain.
Competitive survival of selected mutants in vitro and ex vivo conditions. Competitive in vitro survival of selected mutants in (a) media and (b) bovine macrophages. Approximately equal number of WT BCG Danish and BCG knockout were mixed and used to inoculate media, infect PBMC derived bovine macrophages. Inoculants and recovered BCG were enumerated on selective media. Error bars represent standard errors.
Protective efficacy of ΔBCG TK in guinea Pigs
The aerosol-infection guinea pig model of human TB and bovine TB is commonly used as a screening tool to assess the protective efficacy of vaccines51,52. M. bovis challenge of guinea pigs has also proven useful to test the potency of vaccines against bovine TB53. Dunkin Hartley guinea pigs were randomly divided into four groups. Group 1 and Group 2 were immunised subcutaneously on the nape with ΔBCG TK (5 × 104 cfu) and the wild-type BCG respectively. Group 3 and Group 4 were unvaccinated controls. Protective immunity was assessed as the ability to reduce disease progression following challenge at 42 days post-vaccination (Fig. 3a) with virulent M. bovis (10–20 cfu delivered by aerosol). Disease progression was assessed by weight loss, a sensitive indicator of TB in the guinea pig model. Disease burden was quantified by measurement of viable M. bovis in lungs and spleens.
Comparative analysis of body weight and survival ability of ΔBCG TK vaccinated Guinea pigs after M. bovis challenge. (a) Schematic representation of study schedule (b) Group mean body weight profiles recorded for each group of guinea pigs during the vaccination and challenge phase of the study. Solid red vertical bar on x-axis indicates the time point of vaccination and dashed-black vertical bar on x-axis indicates time point of challenge with M. bovis. (c) Survival data of unvaccinated and vaccinated (WT BCG, ΔBCG TK) guinea pigs up on M. bovis infection.
The uninfected controls (Group 4) together with all animals immunized with either ΔBCG TK or wild-type BCG gained weight normally after challenge (Fig. 3b). Indeed, there were no significant differences in weight gain between these two groups. After 21 days post-infection, however, non-immunized, infected animals (Group 3) exhibited a substantial weight loss and were euthanised at a pre-defined humane end-point; whilst the weight of vaccinated guinea pigs continued to increase steadily (Fig. 3b).
Although this study was not powered to measure survival, the notable difference between disease progression in vaccinated and unvaccinated animals permitted an analysis of survival (based upon time to humane end-point). The Kaplan Meier plot (Fig. 3c) shows that the vaccinated and challenged guinea pigs all survived until the end of the 4-week post-infection observation period, whereas all of the unvaccinated-challenged group had been euthanised by this time. In addition, none of the animals vaccinated with either wild-type or ΔBCG strains showed any pathological signs of clinical disease thus confirming the protective efficacies of the strain.
To assess the capacity of the recombinant vaccine ΔBCG TK to restrict the growth of M. bovis in tissues of challenged guinea pigs, the number of viable bacteria (colony forming units, cfu) in the lung, the primary site of infection, and spleen, a major site of bacterial dissemination was quantified at necropsy. The cfu data from lungs (Fig. 4a) and spleens (Fig. 4b) of challenged animals showed statistically significantly lower cfu burden in the lungs (p < 0.001) and spleens (p = 0.0004) of animals vaccinated with either vaccine compared to unvaccinated controls. The reductions in bacterial burden in either organ imparted by vaccination with wild-type BCG and ΔBCG TK was indistinguishable (Fig. 4). Together with the data demonstrating prevention of disease progression, these data on reduced bacterial burden in lungs and spleens following vaccination with either vaccine demonstrates that the deletion of the target genes from ΔBCG TK has not reduced its protective efficacy.
Protective efficacy of ΔBCG TK vaccination in Guinea pigs upon M. bovis challenge. (a) The bacterial load in the lungs of WT BCG (P = 0.0021) and ΔBCG TK (P = 0.0006) vaccinated guinea pigs is significantly low compared to the with the unvaccinated control group. (b) The bacterial load in spleen of WT BCG (P = 0.0015) and ΔBCG TK (P = 0.0006) vaccinated guinea pigs is significantly low compared to the unvaccinated control group. Values for each individual are shown and the horizontal bar denotes the mean for each group. *P = ≤ 0.05, P = ≤ 0.01, ***P = ≤ 0.001.
Skin test immune response against extended DIVA antigens in guinea pigs
To test whether the antigens deleted from ΔBCG TK could induce skin test responses in M. bovis infected guinea pigs, but not in vaccinated animals prior to infection, orthologs of the genes deleted from ΔBCG TK: Rv3020c (BCG3043), MPB70 (BCG2897), MPB83 (BCG2895), Rv3615c were prepared as three different fusion proteins (ESAT-6-CFP-10, MPB70-MPB83, Rv3615c-Rv3020c). These were tested alone, or in combination, as synthetic antigen cocktails.
Groups of guinea pigs were vaccinated as above with either WT BCG, or ΔBCG TK, or left unvaccinated controls, and subsequently challenged with M. bovis. Skin tests were performed on all animals post-vaccination to determine specificities, and also performed again post-infection to determine the sensitivities of the test reagents. The following antigen preparations were injected in a Latin Square arrangement54 in the sites shown in Fig. 5A: PPD-B, fusion proteins of ESAT6-CFP10, MPB70 and MPB83, Rv3615c and Rv3020c. A cocktail of all of these three fusion proteins (Triple antigen cocktail) was also tested. The site of injections on both sides of guinea pig’s trunk was marked “a, b, c” and “d, e, f (Fig. 5A) and the skin-testing injection regimes were showed in Sup. Table 2.
Pre and post-challenge skin test reaction in ΔBCG TK vaccinated guinea pigs. (A) Diagram to show the injection layout on animal. At least 2.5 cm between each inoculation of antigen preparations were injected on the same flank in a Latin Square arrangement54. The site of injections on both sides of guinea pig’s trunk was marked “a, b, c” and “d, e, f”. (B,C) Pre-challenge skin test: The group mean size of the diameter of erythema at 24 (B) and at 48 hours (C) after injection. (D,E) Post-challenge skin test: the mean size of the diameter of erythema at 24 (D) and 48 (E) hours after injection. The red line denotes the minimum skin test response (STR) threshold (2 mm) for DIVA antigens to consider it as positive. The x-axis denotes the vaccine group. The error bars indicate standard deviation for each vaccine group.
The groups of animals vaccinated with WT BCG or ΔBCG TK gave no skin reactions with any of the DIVA antigen cocktails either 24 h or 48 h post vaccination before challenge with M. bovis. As expected, injection of the standard PPD-B skin test reagent gave rise to reactions in both groups of vaccinated guinea pigs, Unvaccinated animals did not respond to either the DIVA cocktails or to PPD-B (Fig. 5B,C).
Following M. bovis challenge of these animals, both vaccine groups, as well as the unvaccinated control group, showed consistently positive responses to PPD-B with no significant difference in response between the vaccinated and unvaccinated animals (Fig. 5D,E). Challenged animals also reacted to the DIVA antigen cocktails, particularly against the triple fusion protein cocktail injection (MPB70-MPB83, ESAT-6-CFP10 and Rv3615-Rv3020) at the 24 h and 48 h measuring time point (Fig. 5D,E). The response to the MPB70/83 DIVA antigen was however not consistently observed in all animals. Unvaccinated animals showed response to PPD-B and the antigen cocktails only after M.bovis challenge. Only 24 h skin-test reactions are shown for animals in this group as they had reached the humane end point. Importantly, the antigen cocktails distinguished between BCG and M. bovis exposure with skin test reactions occurring only in challenged animals irrespective of their immunization status (Sup. Fig. 3).
Discussion
Bovine TB is a serious animal health problem55 and is one of the biggest challenges facing the cattle farming industry today56. It has been estimated that >50 million cattle are infected worldwide, costing an estimated US$3 billion annually57. Even though vaccination with M. bovis BCG showed widely diverging efficacies in cattle and humans58,59,60, it is still the only realistic vaccine candidate with potential benefits in reducing prevalence and spread of bovine TB in the cattle population40,57,61 and also reduce the severity of a herd breakdown20,22. The differential diagnostic tests will be required in countries where the test and slaughter control strategies are in operation62,63, particularly when vaccination involves BCG that will compromise the specificity of standard tuberculin-based diagnostic tools64. In order to control the spread of tuberculosis, effective vaccination and accurate early diagnosis of the disease are critical. Development of new and improved cattle vaccines, and associated diagnostic reagents, could contribute to improved disease control65.
This study is the first step in a novel strategy to engineer a diagnostically compatible BCG vaccine that has similar protective efficacy to the current commercially available BCG vaccines. We constructed a ΔBCG TK strain that gave indistinguishable protection against BTB challenge as WT BCG. We developed a compatible extended DIVA skin test that proved to be specific in not provoking skin reactions in vaccinated guinea pigs before challenge but provoking reactions post-challenge. The test, although specific, was not as sensitive as PPD-B in the Guinea pig model. Adding additional antigens in a cocktail of 6 antigens, including the prototype DIVA antigens ESAT-6, CFP-10 and Rv361566, alongside our triple antigen proteins (MPB70, MPB83 and Rv3020c) led to significant increases in skin responses in all groups post-challenge whilst retaining the absence of skin test responses post-BCG vaccination prior to challenge.
We note that, as expected, the synthetic antigen cocktail did not provoke a reaction in guinea pigs inoculated with the ΔBCG TK strain, but somewhat surprisingly neither did guinea pigs inoculated with WT BCG, which still contains the antigens present in the antigen cocktail. This outcome is likely because the guinea pig study was not powered sufficiently to demonstrate the relatively small, but economically highly significant, immunoreactivity to these antigens in animals vaccinated with WT BCG. In support of this interpretation, addition of just one of these antigens, Rv3020c, to the standard DIVA cocktail of ESAT-6, CFP-10 and Rv3615c, reduced the specificity of the skin test in BCG vaccinated calves from 100% to 96% (100% specificity of ESAT-6, CFP-10, Rv3615c combination in BCG vaccinated calves in 187 tested animals vs 96% specificity ESAT-6, CFP-10, Rv3615c plus Rv3020c in 125 tested animals) (Jones and Vordermeier, unpublished observations). Although this may appear to be a relatively minor reduction in specificity, the economic and societal impact of a 4% false discovery rate when its consequence may be the culling of healthy animals is likely to prevent implementation of such a test, particularly when compared to the false-discovery rate of the standard comparative tuberculin test with of 0.02%67. Demonstrating a 4% false-positivity with WT BCG and loss of this false-positivity with the ΔBCG TK, advanced DIVA test combination would have required large number of guinea pigs and was beyond the scope of this project. Further, larger studies, preferentially in the cattle as target host, will be required to demonstrate this aspect of our strategy.
In summary, in this study we demonstrate, for the first time, a new strategy for engineering a live bacterial vaccine that has been rationally-designed to optimize both protection and diagnostic compatibility. The DIVA cocktail described here is specific and is not affected by vaccination. Although further experiments are needed to confirm protective efficacy and diagnostic efficacy in the bovine host, the development of combination of effective vaccine and skin test regents could transform bovine TB control programmes worldwide. Similar strategies may also be of value for control of human TB, and perhaps other infectious diseases.
Materials and Methods
BCG culture preparation
M. bovis BCG Danish 1331 (Staten’s Serum Institute, batch 111013B) was grown on Middlebrook 7H11 solid media or in Middlebrook 7H9 supplemented with 0.2% glycerol, 0.05% Tween-80 and 10% OADC at 37 °C shaking at 150 rpm in an orbital shaker37. When selection was required antibiotics were used at 50 µg ml−1 for apramycin, 50 µg ml−1 for hygromycin and 25 µg ml−1 for zeocin.
Construction of the recombinant cosmids containing allelic exchange substrates (AESs)
Cosmid pANE001 zeomycin (Sup. Fig. 1) was derived from pYUB854 (Hyg). The original pYUB854 cosmid was modified by replacing the hygromycin resistance cassette with zeomycin cassettes. An inverse PCR of pYUB854 with primers designed to amplify plasmid backbone less the hygromycin cassette and add Nde1 and Mfe1 restriction ends37. The zeomycin cassettes were amplified from pNCMTB plasmid and Nde1 and Mfe1 restriction sites added to the ends37. The antibiotic cassette was then cloned into pYUB854, and confirmed by Sanger sequencing. The modified res–MfeI-zeo–NdeI-res gene cassette flanked by multiple cloning sites (MCSs) was thus created.
Construction of BCG null mutants
Mutants were generated sequentially using the mycobacteriophage-based method of specialized transduction37,49, and cosmids pANE001 or p0004S. Upstream (LF) and downstream (RF) sequences flanking the genes to be mutated were PCR amplified from BCG Danish genomic DNA using Qiagen High Fidelity Taq polymerase according to manufacturer’s instructions, cloned into the appropriate cosmids and confirmed by Sanger sequencing to generate the knock-out plasmids p0004S3043 (Apra), pANE3679/80 (Zeo) and pYUB2897/95 (Hyg). Primer sequences used in this study are listed in Sup. Table 4.
Confirmation of mutant construction
As described in the earlier publication37, knockout genotypes were confirmed by PCR using primers outside the upstream and downstream flanking regions both alone, and in combination with antibiotic cassette specific primers (Sup. Table 4), such that PCR products would be obtained only if the antibiotic cassette was inserted in the required genomic location50.
Growth analysis of BCG strains
BCG wild-type and ΔBCG TK mutant were grown to mid-log phase (OD 0.8). The cells were pelleted down and washed twice with PBS, then resuspended in 7H9 medium. This resuspended bacterium was used to inoculate fresh to a starting OD of 0.05. Growth was measured by taking OD readings. All experiments were performed in triplicate.
In vitro competition assays
A mix of strains containing approximately equal amounts of the ΔBCG TK mutant and WT BCG Danish were inoculated into broth and cultured for 14 days. At selected time points the numbers of each mutant were determined by serially diluting onto selective media. Numbers of WT BCG were estimated by subtracting the antibiotic resistant colony numbers from counts from plates without antibiotics37. The assays were repeated three times.
Bovine Macrophage preparations and infections
Peripheral blood mononuclear cells (PBMCs) obtained from the heparin-anticoagulated blood collected from the adult cows. The PBMCs isolated using Ficoll-Histopaque density gradient centrifugation. Monocytes from the pool of PBMCs were selected using CD14 MicroBeads (Miltenyi Biotec). The monocytes were differentiated into macrophages in 24 well plates containing complete RPMI supplemented with 1% sodium pyruvate, 1% penicillin/streptomycin and 20 ng ml-1 macrophage colony-stimulating factor (Miltenyi Biotec)37. Fresh medium was added on day 3 and infection on day 6 at a MOI of 1 with a mixed BCG culture containing approximately equal amounts of WT BCG and ΔBCG TK mutant. Control macrophages were incubated with culture medium only. After 4 h, the infected cells were washed three times with PBS. The intracellular bacilli were harvested by lysing the cells with 0.1% Triton X-100 at different time points. The mixed culture used for infection, and harvested intracellular bacilli were enumerated as described previously37. The assays were repeated three times.
Cloning and expression of recombinant proteins
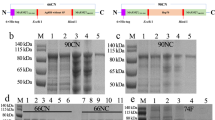
The fusion gene construct (GenScript, USA) of coding sequences of ESAT-6 and CFP-10, and of Rv3615c and Rv3020 of M. tuberculosis H37Rv were synthesized. These gene constructs were then cloned into prokaryotic expression vector pET28a (Novagen) and transformed into E. coli BL21 DE3 cells (Invitrogen). The protein expression was induced with 1 mM IPTG overnight at 25 °C. The His6 tagged ESAT-6::CFP10 and Rv3615c::Rv3020 fusion proteins were purified from the soluble fraction of the bacterial lysate using Ni–NTA agarose (immobilized metal affinity chromatography)68. The pooled protein fractions were dialyzed against PBS (pH 7.4) and SDS-PAGE was used to assess the purity of the protein. The protein was detected in a Western blot using anti-His6 antibody and the LPS from recombinant fusion proteins were removed using Triton X-100 as per the procedure68,69.
Guinea pig experiments
The Guinea pig experiments were conducted according to the United Kingdom Home Office Legislation for animal experimentation and approved by a local ethical committee at Public Health England (Porton Down, United Kingdom). Dunkin Hartley guinea pigs free from pathogen-specific infection were randomly assigned to vaccine groups and identified using subcutaneously implanted microchips (Plexx, the Netherlands) to enable blinding of the analyses wherever possible70. Group sizes were determined by statistical power calculations (Minitab, version 16) performed using previous data (SD, approximately 0.5) to reliably detect a difference of 1.0 log10 in the median number of colony-forming units (cfu) per millilitre70. The guinea pigs were housed in groups of up to eight during vaccination and in pairs post-challenge. Animals were monitored daily for behavioural changes.
The 32 animals were divided into 4 groups (n = 8). Groups 1 and 2 were vaccinated subcutaneously on the nape with 5 × 104 cfu of either ΔBCG TK (Group 1) or wild type BCG (Group 2) at day 0. Groups 3 and 4 remained unvaccinated. All groups received the pre-challenge skin tests at 34 days post-vaccination. Skin test responses (STR) were measured at 24- and 48-hours following inoculation with the antigens. Groups 1, 2 and 3 were challenged with M. bovis (AF2122/97) at 42 days (6 weeks) post-vaccination and received a post-challenge skin test before the scheduled cull and necropsy at 70 days (4 weeks post-challenge). Group 4 was not challenged as this was a control group to test for non-specific skin test responses. Guinea pigs in groups 1–3 were challenged by the aerosol delivery with a target estimated dose of 10–20 cfu of M. bovis using a contained Henderson apparatus in conjunction with an AeroMP control unit71,72,73. Fine particle aerosols of M. bovis, with a mean diameter of 2μm, were generated in a Collison nebulizer and delivered directly to the snout of each animal. The AeroMP is a platform system designed to manage the aerosol generation, characterization and sampling processes via a dashboard software laptop system. Throughout the study, the body weight of each animal was measured and recorded at least weekly. The frequency of checks was increased on appearance of any clinical signs or weight loss. The humane endpoint was reached when 20% loss of maximal body weight was recorded and/or observation of defined clinical signs such as laboured breathing.
The determination of bacterial load was scheduled at 4 weeks post-challenge. Guinea pigs from each group were killed and the lungs and spleens were aseptically removed and stored at −20 °C on the day of necropsy until they were processed in a single batch. On the day of tissue processing, each tissue was homogenized in 10 ml (lung) or 5 ml (spleen) sterile phosphate buffered saline (PBS). Each tissue homogenate was serially diluted in sterile PBS and 100 μl of each dilution plated, in duplicate onto Middlebrook 7H11 + OADC + pyruvate selective agar. Following incubation, colonies were enumerated to determine the colony forming units (cfu).
Skin testing
The skin testing was performed 34 days post-vaccination prior to M. bovis challenge (pre-challenge skin test) and at 62 days post-vaccination around 4 weeks after M. bovis challenge (post-challenge skin tests). All guinea pigs, regardless of vaccination and challenge status were given PPD-B (Group A) and four specific DIVA skin test antigen preparations (Group B-E) at six separate injection sites in a Latin square formation. A diagram of the six sites for each animal (three sites on each flank) is shown in Fig. 5A. The details of the group and skin test antigen are given in Sup. Table 1.
Each antigen cocktail was prepared prior to delivery. 100 μl of each antigen preparation (2 μg of PPD-B or 1 μg of antigen cocktail preparation) was given to the appropriate site by the intradermal route. Each guinea pig received each of the five types of antigen preparation and a repeat of one other (on opposite flank) as described in Sup. Table 2. The rationale to test one preparation on both flanks of each animal was to determine whether the flank side (left or right) influenced the magnitude of the inflammatory response. We didn’t observe any significant difference in the magnitude of the skin test response when the same test was given in different locations or on either flank which nullify the influence of the position or flank of the skin test location on the magnitude of the inflammatory response (Sup. Table 3).
Skin test responses were measured at 24 h and 48 h following antigen inoculation. As we expected, the skin test reaction to recombinant proteins were lower than to PPD. Based on the observations in cattle by33 we defined cut-off values for positivity for the recombinant proteins at both time points at >2 mm, and >4 mm for PPD. The size of the individual erythema reactions (if present) was measured in millimetres (mm) and the average of these values was used for analysis. Skin test data were initially analysed using an ANOVA general linear model (Latin square) statistical analysis. Group comparisons of the magnitude of skin test were performed using the non-parametric Mann-Whitney test (Minitab software version 16). A test for normality was applied to the bacterial load data and the data from each vaccine group were compared and ranked using the non-parametric Mann-Whitney test (Minitab software version 16).
Ethics approval and consent to participate
All animal experimentation was carried out within a project license granted by the United Kingdom Home Office Legislation for animal experimentation and approved by a local ethical committee at Public Health England (Porton Down, United Kingdom).
Data availability
Illumina and MiSeq sequence data are being submitted to the SRA database under submission SUB4615409.
Change history
01 October 2020
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Brown, C. Emerging zoonoses and pathogens of public health significance–an overview. Revue scientifique et technique-office international des epizooties 23, 435–442 (2004).
Perry, B. D. Investing in animal health research to alleviate poverty. (ILRI (aka ILCA and ILRAD), 2002).
Olea-Popelka, F. et al. Zoonotic tuberculosis in human beings caused by Mycobacterium bovis—a call for action. The Lancet Infectious Diseases 17, e21–e25 (2017).
Organization, W. H. Global Tuberculosis Report 2017. (World Health Organization, 2017).
Organization, W. H. Roadmap for zoonotic tuberculosis (2017).
Pritchard, D. A century of bovine tuberculosis 1888–1988: conquest and controversy. Journal of comparative pathology 99, 357–399 (1988).
Borsuk, S., Newcombe, J., Mendum, T. A., Dellagostin, O. A. & McFadden, J. Identification of proteins from tuberculin purified protein derivative (PPD) by LC-MS/MS. Tuberculosis 89, 423–430 (2009).
Donnelly, C. A. et al. Impact of localized badger culling on tuberculosis incidence in British cattle. Nature 426, 834 (2003).
Calmette, A. Preventive Vaccination Against Tuberculosis with BCG. Proceedings of the Royal Society of Medicine 24(11), 1481–1490 (1931).
Horwitz, M. A new TB. vaccine. Immunologist 5, 15–20 (1997).
Zhu, B., Dockrell, H. M., Ottenhoff, T. H., Evans, T. G. & Zhang, Y. Tuberculosis vaccines: Opportunities and challenges. Respirology 23, 359–368 (2018).
Horwitz, M. A., Harth, G., Dillon, B. J. & Masleša-Galić, S. A novel live recombinant mycobacterial vaccine against bovine tuberculosis more potent than BCG. Vaccine 24, 1593–1600 (2006).
Larsen, M. H. et al. Efficacy and safety of live attenuated persistent and rapidly cleared Mycobacterium tuberculosis vaccine candidates in non-human primates. Vaccine 27, 4709–4717 (2009).
Martin, C. et al. The live Mycobacterium tuberculosis phoP mutant strain is more attenuated than BCG and confers protective immunity against tuberculosis in mice and guinea pigs. Vaccine 24, 3408–3419 (2006).
Sayes, F. et al. CD4+ T cells recognizing PE/PPE antigens directly or via cross reactivity are protective against pulmonary Mycobacterium tuberculosis infection. PLoS pathogens 12, e1005770 (2016).
Arbues, A. et al. Construction, characterization and preclinical evaluation of MTBVAC, the first live-attenuated M. tuberculosis-based vaccine to enter clinical trials. Vaccine 31, 4867–4873 (2013).
Kaufmann, S. H. et al. Progress in tuberculosis vaccine development and host-directed therapies—a state of the art review. The Lancet Respiratory Medicine 2, 301–320 (2014).
Sander, P. et al. Deletion of zmp1 improves Mycobacterium bovis BCG-mediated protection in a guinea pig model of tuberculosis. Vaccine 33, 1353–1359 (2015).
Suazo, F. M., Escalera, A. M. A. A. & Torres, R. M. G. A review of M. bovis BCG protection against TB in cattle and other animals species. Preventive veterinary medicine 58, 1–13 (2003).
Chambers, M. et al. Vaccination against tuberculosis in badgers and cattle: an overview of the challenges, developments and current research priorities in Great Britain. Veterinary Record 175, 90–96 (2014).
Wilson, G. J., Carter, S. P. & Delahay, R. J. Advances and prospects for management of TB transmission between badgers and cattle. Veterinary microbiology 151, 43–50 (2011).
Vordermeier, H. M. et al. Correlation of ESAT-6-specific gamma interferon production with pathology in cattle following Mycobacterium bovis BCG vaccination against experimental bovine tuberculosis. Infection and immunity 70, 3026–3032 (2002).
Chaparas, S. D., Maloney, C. J. & Hedrick, S. R. Specificity of tuberculins and antigens from various species of mycobacteria. American Review of Respiratory Disease 101, 74–83 (1970).
Mahairas, G. G., Sabo, P. J., Hickey, M. J., Singh, D. C. & Stover, C. K. Molecular analysis of genetic differences between Mycobacterium bovis BCG and virulent M. bovis. Journal of bacteriology 178, 1274–1282 (1996).
Fogan, L. PPD antigens and the diagnosis of mycobacterial diseases: a study of atypical mycobacterial disease in Oklahoma. Archives of internal medicine 124, 49–54 (1969).
Frothingham, R., Hills, H. G. & Wilson, K. H. Extensive DNA sequence conservation throughout the Mycobacterium tuberculosis complex. Journal of Clinical Microbiology 32, 1639–1643 (1994).
Imaeda, T. Deoxyribonucleic acid relatedness among selected strains of Mycobacterium tuberculosis, Mycobacterium bovis, Mycobacterium bovis BCG, Mycobacterium microti, and Mycobacterium africanum. International Journal of Systematic and Evolutionary Microbiology 35, 147–150 (1985).
Vordermeier, M., Gordon, S. V. & Hewinson, R. G. Mycobacterium bovis antigens for the differential diagnosis of vaccinated and infected cattle. Veterinary microbiology 151, 8–13 (2011).
Vordermeier, M., Jones, G. J. & Whelan, A. O. DIVA reagents for bovine tuberculosis vaccines in cattle. Expert review of vaccines 10, 1083–1091 (2011).
Sester, M. et al. Interferon-γ release assays for the diagnosis of active tuberculosis: a systematic review and meta-analysis. European Respiratory Journal 37, 100–111 (2011).
Vordermeier, H. M., Jones, G. J., Buddle, B. M. & Hewinson, R. G. Development of immune-diagnostic reagents to diagnose bovine tuberculosis in cattle. Veterinary immunology and immunopathology 181, 10–14 (2016).
Vordermeier, H. M., Jones, G. J., Buddle, B. M., Hewinson, R. G. & Villarreal-Ramos, B. Bovine tuberculosis in cattle: vaccines, DIVA tests, and host biomarker discovery. Annual review of animal biosciences 4, 87–109 (2016).
Whelan, A. O. et al. Development of a skin test for bovine tuberculosis for differentiating infected from vaccinated animals. Journal of clinical microbiology 48, 3176–3181 (2010).
Vordermeier, H. et al. Development of diagnostic reagents to differentiate between Mycobacterium bovis BCG vaccination and M. bovis infection in cattle. Clinical and diagnostic laboratory immunology 6, 675–682 (1999).
Vordermeier, H. M., Sidders, B., Stoker, N. G. & Ewer, K. (Google Patents, 2014).
Weir, R. E. et al. Persistence of the immune response induced by BCG vaccination. BMC infectious diseases 8, 9 (2008).
Mendum, T. A. et al. Transposon libraries identify novel Mycobacterium bovis BCG genes involved in the dynamic interactions required for BCG to persist during in vivo passage in cattle. BMC genomics 20, 431 (2019).
Villarreal-Ramos, B. et al. Development of a BCG challenge model for the testing of vaccine candidates against tuberculosis in cattle. Vaccine 32, 5645–5649 (2014).
DeJesus, M. A., Ambadipudi, C., Baker, R., Sassetti, C. & Ioerger, T. R. TRANSIT-a software tool for Himar1 TnSeq analysis. PLoS computational biology 11, e1004401 (2015).
Hope, J. et al. Exposure to Mycobacterium avium induces low‐level protection from Mycobacterium bovis infection but compromises diagnosis of disease in cattle. Clinical & Experimental Immunology 141, 432–439 (2005).
Flower, D. R. & Perrie, Y. In Immunomic Discovery of Adjuvants and Candidate Subunit Vaccines (eds Darren, R. Flower & Yvonne Perrie) 1–11 (Springer New York, 2013).
Vordermeier, M., Jones, G. J., Sampson, S. & Gordon, S. V. In Immunomic Discovery of Adjuvants and Candidate Subunit Vaccines (eds Darren R. Flower & Yvonne Perrie) 73–90 (Springer New York, 2013).
Mahmood, A. & Arora, A. G. Characterization of RD1 related secretory protein (s) from Mycobacterium tuberculosis H37Rv (2015).
Harboe, M. & Nagai, S. MPB70, a unique antigen of Mycobacterium bovis BCG. American review of respiratory disease 129, 444–452 (1984).
Surujballi, O. et al. Sensitive diagnosis of bovine tuberculosis in a farmed cervid herd with use of an MPB70 protein fluorescence polarization assay. Canadian Journal of Veterinary Research 73, 161 (2009).
Harrington, N. P. Immune Responses of Deer (Cervus Elaphus) to Mycobacterium Bovis Infection: Potential for Immunodiagnostics. (ProQuest, 2006).
Nayak, K. et al. Identification of novel Mycobacterium tuberculosis CD4 T-cell antigens via high throughput proteome screening. Tuberculosis 95, 275–287 (2015).
Mustafa, A. et al. Immunogenicity of Mycobacterium tuberculosis antigens in Mycobacterium bovis BCG-vaccinated and M. bovis-infected cattle. Infection and immunity 74, 4566–4572 (2006).
Bardarov, S. et al. Specialized transduction: an efficient method for generating marked and unmarked targeted gene disruptions in Mycobacterium tuberculosis, M. bovis BCG and M. smegmatis. Microbiology 148, 3007–3017 (2002).
Williams, K. J. et al. The Mycobacterium tuberculosis β-oxidation genes echA5 and fadB3 are dispensable for growth in vitro and in vivo. Tuberculosis 91, 549–555 (2011).
Clark, S., Hall, Y. & Williams, A. Animal models of tuberculosis: guinea pigs. Cold Spring Harbor perspectives in medicine 5, a018572 (2015).
McMurray, D. N. In Tuberculosis 135–147 (American Society of Microbiology, 1994).
Clark, S. et al. Assessment of different formulations of oral Mycobacterium bovis Bacille Calmette-Guérin (BCG) vaccine in rodent models for immunogenicity and protection against aerosol challenge with M. bovis. Vaccine 26, 5791–5797 (2008).
Bradley, J. V. Complete counterbalancing of immediate sequential effects in a Latin square design. Journal of the American Statistical Association 53, 525–528 (1958).
Enticott, G. Relational distance, neoliberalism and the regulation of animal health. Geoforum 52, 42–50 (2014).
Reynolds, D. A review of tuberculosis science and policy in Great Britain. Veterinary microbiology 112, 119–126 (2006).
Waters, W. R., Palmer, M. V., Buddle, B. M. & Vordermeier, H. M. Bovine tuberculosis vaccine research: historical perspectives and recent advances. Vaccine 30, 2611–2622 (2012).
Berggren, S. Field experiment with BCG vaccine in Malawi. British Veterinary Journal 137, 88–94 (1981).
Francis, J. Bovine Tuberculosis Including A Contras With Human Tuberculisis. (Staples Press Limited Cavendish Place, London W, 1947).
Rodrigues, L. C. & Smith, P. G. Tuberculosis in developing countries and methods for its control. Transactions of the Royal Society of Tropical Medicine and Hygiene 84, 739–744 (1990).
Lopez-Valencia, G. et al. Field evaluation of the protective efficacy of Mycobacterium bovis BCG vaccine against bovine tuberculosis. Research in veterinary science 88, 44–49 (2010).
Caffrey, J. Status of bovine tuberculosis eradication programmes in Europe. Veterinary Microbiology 40, 1–4 (1994).
Cousins, D. Mycobacterium bovis infection and control in domestic livestock. Revue Scientifique et Technique-Office International des Epizooties 20, 71–85 (2001).
Mukundan, H., Chambers, M., Waters, R. & Larsen, M. Tuberculosis, Leprosy and Mycobacterial Diseases of Man and Animals: The Many Hosts of Mycobacteria (CABI, 2015).
Vordermeier, H. M. et al. Vaccination of domestic animals against tuberculosis: review of progress and contributions to the field of the TBSTEP project. Research in veterinary science 97, S53–S60 (2014).
Bezos, J. et al. Evaluation of the immunogenicity and diagnostic interference caused by M. tuberculosis SO2 vaccination against tuberculosis in goats. Research in veterinary science 103, 73–79 (2015).
Goodchild, A., Downs, S., Upton, P., Wood, J. & De La Rua-Domenech, R. Specificity of the comparative skin test for bovine tuberculosis in Great Britain. The Veterinary Record 177, 258 (2015).
Veerasami, M. et al. Multi-antigen print immunoassay for seroepidemiological surveillance of bovine tuberculosis on Indian cattle farms. Veterinaria italiana 48, 253–267 (2012).
Veerasami, M. et al. Point of Care Tuberculosis Sero-Diagnosis Kit for Wild Animals: Combination of Proteins for Improving the Diagnostic Sensitivity and Specificity. Indian journal of microbiology 58, 81–92 (2018).
Clark, S. et al. Revaccination of guinea pigs with the live attenuated mycobacterium tuberculosis vaccine MTBVAC improves BCG’s protection against tuberculosis. The Journal of infectious diseases 216, 525–533 (2017).
Clark, S., Hall, Y., Kelly, D., Hatch, G. & Williams, A. Survival of Mycobacterium tuberculosis during experimental aerosolization and implications for aerosol challenge models. Journal of applied microbiology 111, 350–359 (2011).
Hartings, J. M. & Roy, C. J. The automated bioaerosol exposure system: preclinical platform development and a respiratory dosimetry application with nonhuman primates. Journal of pharmacological and toxicological methods 49, 39–55 (2004).
Williams, A., Davies, A., Marsh, P. D., Chambers, M. A. & Hewinson, R. G. Comparison of the protective efficacy of bacille calmette-Guerin vaccination against aerosol challenge with Mycobacterium tuberculosis and Mycobacterium bovis. Clinical infectious diseases 30, S299–S301 (2000).
Acknowledgements
We acknowledge Faye Lanni and the Porton Biological Investigations Group for the Guinea pig experiments. We acknowledge Gareth Jones, Animal and Plant Health Agency, UK for the supporting data on specificity of DIVA antigens in Cattle. This study was supported by the UK BBSRC & Department of Biotechnology, India (Grant Number BB/L004569/1) and the Gates Foundation (BMGF:OPP1106822).
Author information
Authors and Affiliations
Contributions
A.C. generated the B.C.G. double and triple knock out, carried out the competitions experiments, data analysis and wrote the manuscript; K.W. generated libraries, individual mutants and carried out data analysis; T.M. contributed to the experiments and reviewed the manuscript; GS conceived the project and reviewed the manuscript; S.C., S.Z., N.M., F.L. & A.W. carried out the Guinea pig experiment; B.V.R. reviewed the manuscript; M.V. conceived the project and reviewed the manuscript; V.M. prepared antigen cocktail for skin test; A.P., N.B., R.B., S.M.K. carried out TRANSIT analysis; JM conceived the project and reviewed the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chandran, A., Williams, K., Mendum, T. et al. Development of a diagnostic compatible BCG vaccine against Bovine tuberculosis. Sci Rep 9, 17791 (2019). https://doi.org/10.1038/s41598-019-54108-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-54108-y
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.