Abstract
Several mud volcanoes are active in the Canadian Beaufort Sea. In this study, we investigated vertical variations in methanotrophic communities in sediments of the mud volcano MV420 (420 m water depth) by analyzing geochemical properties, microbial lipids, and nucleic acid signatures. Three push cores were collected with a remotely operated vehicle from visually discriminative habitats that were devoid of megafauna and/microbial mats (DM) to the naked eye, covered with bacterial mats (BM), or colonized by siboglinid tubeworms (ST). All MV420 sites showed the presence of aerobic methane oxidation (MOx)- and anaerobic methane oxidation (AOM)-related lipid biomarkers (4α-methyl sterols and sn-2-hydroxyarchaeol, respectively), which were distinctly different in comparison with a reference site at which these compounds were not detected. Lipid biomarker results were in close agreement with 16S rRNA analyses, which revealed the presence of MOx-related bacteria (Methylococcales) and AOM-related archaea (ANME-2 and ANME-3) at the MV420 sites. 4α-methyl sterols derived from Methylococcales predominated in the surface layer at the BM site, which showed a moderate methane flux (0.04 mmol cm−2 y−1), while their occurrence was limited at the DM (0.06 mmol cm−2 y−1) and ST (0.01 mmol cm−2 y−1) sites. On the other hand, 13C-depleted sn-2-hydroxyarchaeol potentially derived from ANME-2 and/or ANME-3 was abundant in down-core sediments at the ST site. Our study indicates that a niche diversification within this mud volcano system has shaped distinct methanotrophic communities due to availability of electron acceptors in association with varying degrees of methane flux and bioirrigation activity.
Similar content being viewed by others
Introduction
Submarine mud volcanoes (MVs) are characterized by seepage-related geomorphological features and act as windows for emission of deep subsurface hydrocarbon gases (mainly methane), fluids, and a complex mixture of sediments1,2,3. They are considered as significant geological sources of global atmospheric methane emissions4,5. In the atmosphere, methane is a strong greenhouse gas that is >30 times more potent than carbon dioxide6. Recently, a number of active MVs featuring gas hydrate deposits were discovered in the Canadian Beaufort Sea7,8. This marine province is expected to experience an increases of 1 to 2 °C in bottom water temperature in the near future9. Similar to other Arctic continental margin sediments10,11,12, bottom water temperature increase may lead to the dissociation of gas hydrate deposits13,14 and elevated methane emissions to the water column and, potentially, to the atmosphere where it further contributes to global warming8,15,16.
Submarine MVs often host dense and diverse microbial and faunal communities fueled by gases and fluids from the subsurface17,18,19,20. The richness of such chemosynthetic communities depends on long-term availability of electron acceptors (e.g. oxygen and sulfate) and donors (e.g. methane) in the sediments21,22. In this regard, microbial processes are typically dominated by aerobic and anaerobic oxidation of methane (MOx and AOM, respectively), which are performed by different clades of aerobic methanotrophic bacteria (MOB)23, or anaerobic methanotrophic archaea (ANMEs)24. ANMEs are often accompanied by sulfate-reducing bacteria (SRB)25,26,27. At highly dynamic MVs, the capacity for methane removal is often reduced because upwelling geofluids, devoid of electron acceptors, hinder influx of electron acceptors (e.g., oxygen and sulfate) from the water column15,19. Physical disturbances by upwelling mud and fluids at active MVs also can give rise to heterogeneous distributions of methanotrophic communities at small scales <1 m of a single MV28,29.
Since the discovery of active MVs across the slope of the Canadian Beaufort Sea, the biogeochemical signatures of archaeal communities involved in AOM have shown spatially discriminative distributions among three MVs (MV282, MV420, and MV740), at water depths of 282 m, 420 m and 740 m, respectively8,30. In this study, we collected sediment push cores from visually discriminative chemosynthetic fields at an active MV (MV420): (i) devoid of megafauna and/microbial mats (DM) to the naked eye, (ii) covered with bacterial mats (BM), or (iii) colonized by siboglinid tubeworms (ST), using a small remotely operated vehicle (ROV) (Fig. 1). In addition, a sediment core was retrieved in a reference site, using a box corer (Fig. 1). Gas, porewater, bulk element, lipid, and nucleic acid analyses were performed to obtain a comprehensive picture of spatial variation in bacterial and archaeal methanotrophic communities. Our study of the spatial and vertical distributions of methanotrophic communities at MV420 sheds light on a key biogeochemical factor shaping discriminative communities.
(a) Map showing the study area (red box) with an inset of the regional map of Alaska and northwestern Canada modified from Paull et al. (2015). (b) Map showing the mud volcano at a water depth of 420 m (MV420) and the reference site on the upper slope of the Canadian Beaufort Sea. The map was generated in Generic mapping tools version 5.4.5 (http://gmt.soest.hawaii.edu/projects/gmt). The high-resolution (1 m) bathymetry data was acquired using AUV in MBARI, processed in MB System (https://www3.mbari.org/data/mbsystem/index_ldeo.html). (c) Video photographs showing discriminative microbial habitats at MV420, i.e., devoid of megafauna and/microbial mats (DM), bacterial mats (BM), and siboglinid tubeworms (ST). The video photographs were taken by the MBARI ROV team during the Araon cruise in 2017.
Results
Geochemical properties
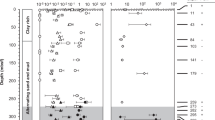
Methane concentration varied at three MV420 sites: DM, BM, and ST (Fig. 2). The highest methane concentration was observed at the DM site (6.6 ± 4.0 mM), followed by the BM (4.0 ± 3.6 mM) and ST (2.4 ± 1.2 mM) sites (Fig. 2). Methane concentration was below the detection limit at the reference site (Fig. 2). The carbon isotopic composition of methane (δ13CCH4) was −64.8‰ to −45.5‰, and generally showed 13C-enrichment toward the surface sediment layers, in accordance with the decrease of methane concentration (Fig. 2). Sulfate concentration ranged from 27.4 mM (sediment surface) to 1.1 mM (bottom section of the core), but penetration depths ranged from 1.5 cm to 7 cm, while the reference site showed a value of 27.8 ± 0.2 mM throughout the entire core (Table 1 and Fig. 2). Porewater chloride concentration varied between 495 mM and 550 mM, co-varying with sulfate concentration along the sediment depth (Fig. 2). Porewater dissolved inorganic carbon (DIC) concentration increased from surface sediments (1.8 mM) toward the bottom of the core (24.7 mM; Fig. 2). δ13CDIC values were −11.6 ± 3.2‰ at the DM site, while those of the BM and ST site were more 13C-depleted (Fig. 2). In contrast, at the reference site, porewater DIC concentration was relatively constant along the sediment depth at 2.7 ± 0.3 mM, and the δ13CDIC values were 13C-enriched (−4.8 ± 1.6‰). Diffusive methane flux was highest at the DM site (–648.5 mmol m−2 y−1), followed by the BM (−427.5 mmol m−2 y−1) and ST (−146.6 mmol m−2 y−1) sites (Table 1). The corresponding sulfate fluxes were 671.0 mmol m−2 y−1, 449.1 mmol m−2 y−1, and 191.4 mmol m−2 y−1, respectively.
Depth profiles of gas (methane), porewater (sulfate, chloride, and dissolved inorganic carbon [DIC]), and bulk elements (total organic carbon and total sulfur) from sediment cores of (a) devoid of megafauna and/microbial mats (DM), (b) bacterial mats (BM), (c) siboglinid tubeworms (ST), and (d) reference sites. “ND” denotes “not determined.”
Total organic carbon (TOC) content ranged from 0.8 wt.% to 1.9 wt.% at all MV420 sites and from 1.4 wt.% to 1.9 wt.% at the reference site (Fig. 2). δ13CTOC values (−28.0 ± 0.6‰) were lower in the upper 4 cm depth at the BM site, while other MV420 and reference sites showed relatively 13C-enriched values (−25.5 ± 0.1‰) (Fig. 2). Total sulfur (TS) content varied between 0.1 wt.% and 1.0 wt.%, with the highest value at the ST site (Fig. 2). δ34STS values were in the range of −18.6‰ to 9.2‰, with the largest sulfur isotopic variation (∆δ34S; ~20) at the ST site (Fig. 2).
Lipid signatures of methanotrophic communities
The 4α-methyl sterols that are putatively specific for bacterial groups involved in MOx31 were abundant at the surface of all MV420 sites but not the reference site. Concentrations were highest at the surface of the BM site, with a range of 0.03–0.37 μg g−1 (Fig. 3 and Supplementary Information Table S1). Specific fatty acids (FAs) such as C16:1ω8 and C16:1ω5, which were inferred as markers for aerobic methanotrophic bacteria and SRB, respectively,23,32 were detected in a range of 0.01 to 0.14 μg g−1 and 0.01 to 0.09 μg g−1, respectively (Fig. 3 and Supplementary Information Table S1). These FAs predominated near the surface of each MV420 site and decreased along the sediment depth.
In contrast to bacterial lipids, isoprenoidal dialkyl glycerol diethers (DGDs) (e.g., archaeol and sn-2-hydroxyarchaeol), which are specific AOM-related biomarkers33, were identified in deeper sediment depths (4 to 16 cm) at each MV420 site but not at the reference site (Fig. 3 and Supplementary Information Table S1). Isoprenoid glycerol dialkyl glycerol tetraethers (GDGTs) containing zero to three cyclopentane moieties (GDGT-0 to GDGT-3) and crenarchaeol were detected in all samples investigated (Fig. 3). Isoprenoid GDGTs were dominated by GDGT-0 and crenarchaeol, at concentrations of 0.05 to 0.72 μg g−1 and 0.03 to 0.77 μg g−1, respectively, while GDGT-1, GDGT-2, and GDGT-3 were much less abundant (<0.03 μg g−1) in all sediments analyzed (Fig. 3 and Supplementary Information Table S1).
The δ13C values of 4α-methyl sterols were −104.9‰ to −30. 5‰, and these compounds were strongly 13C-depleted near the surface layer of each site. The δ13C values of FAs ranged from −63.8‰ to −41.2‰ in all sediments investigated. The δ13C values of isoprenoid DGDs varied between −109.6‰ and −38.6‰, with higher depletion in sn-2-hydroxyarchaeol (−91.4 ± 16.4‰) than in archaeol (−47.6 ± 6.5‰) (Fig. 4 and Supplementary Information Table S1). The δ13C values of biphytanes (BPs) derived from isoprenoid GDGTs were in the range of −79.5‰ to −19.0‰, showing more 13C-depleted BP-1 values at all MV420 sites.
Dominant methanotrophic taxa
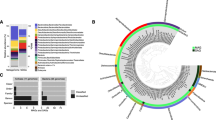
The number of 16S rRNA gene reads of bacteria and archaea ranged from 8711 to 122,357 and from 8070 to 63,065 sequences per sample, respectively (Supplementary Information Table S2). We identified 25 operational taxonomic units (OTUs) belonging to the order Methylococcales of Gammaproteobacteria, which are aerobic methanotrophic bacteria23,34 (Supplementary Information Table S3 and Fig. S1). Among them, B_OTU004 dominated near the surface of the DM site (8.0 ± 6.6%), followed by the BM site (3.3 ± 4.2%) and the ST site (1.1 ± 1.2%), while it was not detected at the reference site (Fig. 5 and Supplementary Information Tables S3 and S4 and Fig. S2). With the exception of B_OTU004, the proportions of other OTUs of Methylococcales were less than 0.2 ± 0.3% in the samples investigated (Supplementary Information Table S3).
Vertical profiles of relative abundance of major OTUs (Methylococcales; B_OTU004 and Methanomicrobia; A_OTU001, A_OTU008, A_OTU014, and A_OTU015) obtained from sediment cores of (a) devoid of megafauna and/microbial mats (DM), (b) bacteria mats (BM), (c) siboglinid tubeworms (ST) sites, and (d) reference sites. “ND” denotes “not determined.”
The relative abundance of A_OTU001, which belongs to ANME-3, was relatively higher (up to 97.1% of archaeal sequences) along sediment depths of MV420 sites, while it was less than 0.08% in the reference site (Fig. 5 and Supplementary Information Table S4 and Fig. S2). Another archaeal OTU (A_OTU008) that belongs to ANME-2 comprised 1.4% to 8.7% of archaeal sequences in the deeper sediment depths (4 to 11 cm) of the ST site. But this OTU was rare or not detected in the sediments of the DM and BM sites (Fig. 5 and Supplementary Information Table S4 and Fig. S2). A_OTU014 and A_OTU015 predominated near the surface of the DM site (up to 13.3% and 6.6% of archaeal sequences, respectively), while the proportion of these OTUs was less than 0.4% at the BM and ST sites (Fig. 5 and Supplementary Information Table S4 and Fig. S2).
Twelve OTUs of Deltaproteobacteria with a greater than 5% relative abundance of bacterial sequences were separately affiliated with the family Desulfobacteraceae or Desulfobulbaceae (Supplementary Information Table S4 and Fig. S2). Among eight OTUs belonging to the family Desulfobacteraceae, most (B_OTU002, B_OTU009, B_OTU011, B_OTU014, B_OTU024, B_OTU029, and B_OTU076) showed relatively higher abundances at deeper sediment depths at all MV420 sites but not the reference site. B_OTU014 showed the highest abundance (57.5%) at the 16 cm sediment depth of the DM site but did not exceed 2.4% at the BM and ST sites. In contrast, B_OTU018 showed the highest abundance (7.4%) near the sediment surface of the ST site. Among four OTUs belonging to the family Desulfobulbaceae, B_OTU010 showed relatively high abundance (2.8 to 14.4%) in the upper 3 cm of all the MV420 sites but not the reference site. In contrast, B_OTU064 showed high relative abundance only in the upper 4 cm of the ST site (2.1 to 5.5%), and the proportions of B_OTU119 was higher at the 0 to 1 cm depth of the BM site.
Co-occurrence of ANMEs with sulfate-reducing Deltaproteobacteria
To infer the putative sulfate-reducing partners of ANMEs, co-occurrence between two archaeal OTUs (ANMEs) and 12 bacterial OTUs (Deltaproteobacteria) was investigated by calculating Pearson correlation coefficients (Supplementary Information Table S5). At the DM and BM sites, no OTUs showed significant correlation (p < 0.01) between ANMEs and SRB. At the ST site, both archaeal OTUs showed significantly positive correlation (r = 0.71 to 0.92, p < 0.01) with a few bacterial OTUs (B_OTU002, B_OTU011, B_OTU018, and B_OTU029).
Discussion
Biogeochemical characteristics along chemosynthetic habitats
Gas bubbles were observed in the ROV video rising within the water column during sampling in the chemosynthetic habitats of MV420. Previous studies using an autonomous underwater vehicle (AUV) and an ROV reported that the fresh and warm subsurface muds exposed during eruptions were mostly saturated with methane (δ13CCH4 = −64‰ and δDCH4 = −222‰), indicating primarily microbial sources8,35. Consistent with those studies, the overall geochemical signatures observed at the MV420 sites were clearly different in comparison with those of the reference site (Fig. 2).
Methane and sulfate fluxes calculated based on concentration gradient varied among the three MV420 sites (Table 1). The DM site, which was devoid of both sessile and mobile taxa such as polychaeta and bivalves, had the highest methane and sulfate fluxes (648.5 mmol m−2 y−1 and 671.0 mmol m−2 y−1, respectively). At the DM site, strong methane fluxes may cause an imbalance in hydraulic pressure, especially in shallow layers where sediments are not yet consolidated, resulting in active invasion of bottom seawater36. Moreover, the DM sites have been interpreted to be on mud flows which are too young to have yet recruited sessile organisms and in place not even been invaded by mobile taxa8,37,38. However, in accordance with decreasing fluid flux (<450 mmol m−2 y−1), free-living bacteria communities and ST patches had developed differently on the surface of MV420. Thiotrophic bacterial communities were commonly found in seeps with moderate fluid flow rates (~1 m yr−1), while STs dominated at seeping systems with a fluid flow rate less than 0.5 m yr−1 15,22. Such biological succession has been reported at various seeping sites, which are differentiated by water depths and methane origins — for instance, at the Håkon Mosby mud volcano (HMMV)19, the Congo deep-sea fan39, the Gulf of Mexico40, and the deep Eastern Mediterranean Sea41. Varying fluid fluxes therefore appear to be a plausible control factor for distinguishing the near seafloor chemosynthetic habitats within MV420.
Corresponding with the depleted chloride concentrations along sediment depth profiles, the penetration depth of sulfate supplied from seawater varied among the three MV420 sites, possibly leading to isotopic variations of the total sulfur pool (Table 1 and Fig. 2). The largest isotopic variation (>20) investigated at the ST site may be attributed to the higher contributions of δ34S values (ca. + 20‰) from the bottom seawater sources42. The sulfate-methane transition zone was formed at the deeper sediment horizons of the ST site compared with other sites. In this regard, δ13CCH4 isotopic signatures were more enriched at the ST site, indicating that the remaining methane pool became progressively enriched in 13C35,43,44. These results are in close agreement with 13C-depleted DIC values, as a result of isotopic fractionation through active AOM under lower methane fluxes43,44,45,46. Considering that endosymbiotic bacteria inhabiting STs exploit some substrates (e.g., methane and by-products involved in AOM) in subsurface sediments47,48, bioirrigation activities by STs may enhance the availability of electron acceptors in deeper sediments17,49,50,51. These activities allow niche separation between chemosynthetic organisms competing for common energy sources over time29,52,53. Under varying methane fluxes investigated at MV420, both chemosynthetic organisms (e.g., free-living bacteria and STs) inhabiting seeping sediments appear to have been distinctly separated based on energy source. These results imply that different habitat types mirroring the discriminative intensities of ascending gas-fluids may influence the spatial distribution of methanotrophic communities by altering the availability of electron acceptors.
Occurrence of aerobic methanotrophs
MOx- and AOM-related microbial communities assessed according to lipid biomarkers and 16S rRNA gene sequences showed a clear down-core shift within each site and variance among the MV420 sites. For the first time in Beaufort MV, we found abundant and 13C-depleted 4α-methyl sterols in concert with high proportions of OTUs belonging to the order Methylococcales near the sediment surface, strongly indicating a high abundance of aerobic methanotrophs in these sediments (Figs. 3–5), as has been reported from various other seepage sites19,28,53,54. The predominance of aerobic methanotrophs at the surface appears to result from the intensities of oxidant-depleted subsurface fluids that limit downward diffusion of sulfate and oxygen from seawater to the uppermost centimeters of the sediment28,53,54,55. In such situations, which do not easily allow AOM reactions in deeper sediment layers, a preferential niche of MOx-related methanotrophs is formed in the surface sediment where oxygen is available. Notably, 13C-enriched 4α-methyl sterols in deeper sediment layers indicate that these compounds originate from processes other than methanotrophy (e.g. organisms using methanol)56. However, with the current data set, it is challenging to elucidate exact processes involved, and thus further research is needed.
With discriminative distributions of MOx-related methanotrophs among the three MV420 sites, 13C-depleted 4α-methyl sterols were much more prevalent at the BM site than at other sites (Figs. 3–5). Moreover, 13C-depleted TOC was abundant at the surface of the BM site, indicating that MOx-related biomass was incorporated and preserved in the organic carbon pool for this site57,58,59. Such distribution patterns investigated at MV420 appear to differ slightly from those of similar MV systems (i.e., HMMV) that showed predominant MOx-related methanotrophs under the highest methane flux19,28. One possible reason for this difference might be that the mud flows migrating to the tops of the DM site were too active and too young for colonization of MOx-related methanotrophs8, in contrast to the BM and ST sites. Moreover, previous studies of various seeping sites have suggested that seafloor temperature and oxygen availability are the key factors shaping seep microbial communities28,54. In this study, in situ bottom temperature in the surface did not vary across the MV420 sites (0.37 ± 0.1 °C), indicating that temperature is likely not the key factor explaining the discriminative distribution of MOx-related methanotrophic Methylococcales within a single MV.
Occurrence of anaerobic methanotrophs
The predominance of sn-2-hydroxyarchaeol over archaeol and OTUs belonging to ANME-2 and ANME-3 clades in sediment depths (4 to 16 cm) indicates the presence of AOM-related methanotrophic communities in these sediments, in contrast to the reference site (Figs. 3–5). The overall 13C-depleted sn-2-hydroxyarchaeol in all MV420 sites indicated persistent occurrence of AOM in sediments containing sulfate. Based on 16S rRNA results, ANME-3 was more dominant at the BM and ST sites than the DM site. On the other hand, ANME-2 was most frequent at the ST site, indicating a preference for a niche with lower methane flux. Notably, the low abundances of isoprenoid GDGTs (GDGT-1, GDGT-2, and GDGT-3) and 16S rRNA gene sequencing results indicate that the ANME-1 contribution was negligible for AOM in the sediments of MV42030,33,60,61,62. It is worthwhile to note that all surface sediments including that of the reference site showed a dominant crenarchaeol contribution to the total isoprenoid GDGTs pool, indicating a potential contribution of marine Thaumarchaeota63. The enriched δ13C values of archaeol and BPs suggest that archaeal communities are mixed due to processes other than AOM (e.g., assimilation of AOM-derived inorganic carbon)64. These δ13C signatures are similar to methane-related archaeal distributions investigated at other MVs (at water depths of 290 m and 740 m) in the Canadian Beaufort Sea30. In this regard, niche stratifications among ANMEs appear to be related to differences in environmental preferences within the sampled habitats with varying methane fluxes53,54.
At the three investigated MV420 sites, specific archaeal lipids and 16S rRNA signatures clearly reveal the discriminative presence of ANME-2 and ANME-3 involved in AOM between the BM and ST sites (Figs. 3–5). In contrast to ANME-1, which was observed at a wide range of temperatures (4 to 70 °C) and typically lower methane fluxes65,66,67, ANME-2 and ANME-3 have been found in samples with temperatures below 20 °C, vigorously emitting MVs such as HMMV19,28,53,54 and other seepages52,66. Among them, ANME-2 (particularly ANME-2c) was most frequently found in sediments bioturbated by benthic communities such as vesicomyid clams inhabiting low-fluid flux regimes29,54,68,69. Considering that vesicomyid clams have so far not been found in the western Arctic, our results indicate a requirement of ANME-2 for lower methane flux. The overall dominance of ANME-3 at the BM site is consistent with previous findings, prevailing across settings in Arctic seeping sites19,28,53,54. Meanwhile, at the ST site, ANME-2 can be regarded as the additional methane oxidizer under environmental conditions such as bioturbation and lower methane seepage60,70,71. Given that the produced sulfide is quickly oxidized by thiotrophic bioturbators, the presence of ANME-2 below tubeworm patches may imply that they seem to form a syntrophic relationship with SRB during AOM19,54. To some degree, ANME-3 appears to tolerate high sulfide concentrations produced during active AOM processes72, although the sulfide sensitivity of ANMEs needs to be investigated further. AOM activities performed by both ANMEs seem to support the metabolism of sulfur-oxidizing endosymbionts inhabiting frenulata Oligobrachia haakonmosbiensis73,74. The presence of diverse methanotrophs at the ST site can therefore be explained by environmental conditions under steady replenishment of sulfate and oxygen levels into the sediments, in accordance with the development of ST patches.
Niche adaptation between ANMEs and SRBs
With respect to the consortium of ANMEs and SRBs, the presence of sulfate-reducing Deltaproteobacteria, including the partner SRB of ANMEs, was assessed by analyzing specific FAs and deltaproteobacterial OTUs (Desulfobacteraceae and Desulfobulbaceae) at the three MV420 sites. The 13C-depleted, SRB-specific lipid (C16:1ω5) probably derived from Desulfobacteraceae was relatively abundant near the sediment surface, while another specific FA (C17:1ω6) derived most likely from Desulfobulbaceae was not identified in any sediments60. Most deltaproteobacterial OTUs showed non-significant correlations with major OTUs of the ANME-2 and ANME-3 clade. Only a few deltaproteobacterial OTUs had a positive correlation with major ANMEs at the ST site. This suggests a weak coupling between ANMEs and Desulfobacteraceae at the MV420 sites, which is inconsistent with reported couplings of SRBs (Desulfosarcina/Desulfococcus and Desulfobulbus) and ANMEs (ANME-2 and ANME-3) at HMMV19,28. Desulfobacteraceae often occur as single cells in surface sediments of bacterial mat- or clam-covered seeps75,76 and decrease with increasing sediment depths75. Therefore, this SRB seems to have a metabolic advantage over other SRBs in oxygenated sediments. Accordingly, at MV420, the occurrence of overall ANMEs without partner SRBs may imply that ANMEs were either associated with other SRBs, occurred as loose aggregates or a single cell without direct contact with SRBs24, or performed AOM without a partner SRB77. More studies are necessary to further investigate ANME-SRB consortia by, for example, applying the technique of fluorescence in situ hybridization.
Summary
In this study, we provide the first evidence for MOx- and AOM-related methanotrophic communities in connection with chemosynthetic communities at an active MV in the Canadian Beaufort Sea. Our results revealed that niche diversification of methanotrophic communities (i.e., Methylococcales and ANME-2 and ANME-3 groups, respectively) may have been shaped by environmental factors such as availability of electron acceptors and bioirrigation activities by chemosynthetic organisms. In comparison with highly local diversification of methanotrophic communities at heterogenous seeping sites54, our finding confirms that the supply of electron acceptors is a key factor determining local diversification of methanotrophic communities. In particular, our study is one of the few that show both methanotrophs (i.e., Methylococcales and ANME-3) distributed separately under seeping fluids and high methane fluxes19,24,28,66. Our study of methanotrophic signals provides valuable information on the effective removal pathways involved in methane release from marine sediments.
Methods
Sample collection
Sediment samples were obtained during the 2017 ARA08C cruise of the Korean icebreaker RV Araon at the Beaufort MV (water depth of 420 m) from different chemosynthetic provinces (ARA08C-DIVE105-8 and -9: BO site, ARA08C-DIVE104-7 and-13: BM site, and ARA08C_DIVE105-12 and -14: ST site) using a push core operated by an ROV system and at the reference site (ARA08C-21BC, water depth of 420 m) using a box corer (Fig. 1). Upon recovery, push- and box-core samples were subsampled horizontally with 20 mL serum vials, which were then sealed immediately with butyl rubber stoppers to prevent gas exchange and stored at 4 °C. Porewater was extracted from holes drilled at intervals of 2 to 3 cm down the core liner using rhizon samplers connected to 20 mL syringes. Extracted porewater was filtered through a 0.20 μm disposable polytetrafluorethylene in-line filter and stored at 4 °C until analysis. After porewater extraction, core sediments were sliced into 1cm sections and subsampled for analysis of bulk elements, lipid biomarkers, and 16S rRNA gene sequences. Subsamples were stored at −80 °C until analysis.
Gas and porewater analyses
Headspace methane concentration was measured with a gas chromatograph (GC) (7890 A, Agilent Technologies, CA, USA) with flame ionization detector. The precision of repeated standard analyses exceeded 5%. Isotopic compositions of methane were obtained using an isotope ratio mass spectrometer (Finnigan MAT 252; Thermo Fisher Scientific, Waltham, MA, USA) with a Combustion III interface (Thermo Fisher Scientific) at Nagoya University. Isotopic values were expressed as δ-notation (per mil) relative to Vienna Pee-Dee Belemnite (VPDB). The δ13C values were calibrated using analyses of a methane standard with δ13C value of −35.2‰, as certified by the National Institute of Standards and Technology (NIST, USA). Precision for δ13CCH4 was greater than ±0.2‰.
Chloride (Cl–) and DIC were analyzed by titration using silver nitrate (AgNO3) and 0.1 N HCl, respectively. The precision determined by repeated titrations of International Association of Physical Sciences of the Oceans standard seawater was ±0.5% for Cl– and <2% for DIC. Sulfate was measured using ion chromatography (Metrohm 761). The δ13C values of DIC were obtained using an isotope ratio mass spectrometer (Finnigan DELTAplusXL; Thermo Fisher Scientific, Waltham, MA, USA) with a Finnigan GasBench-II headspace autosampler at Oregon State University (Corvallis, OR, USA). Isotopic values were expressed as δ-notation (per mil) relative to VPDB. The δ13C values were calibrated through analyses of a CaCO3 standard with δ13C value of –0.4‰, certified by Oregon State University. The precision for δ13CDIC was greater than ±0.02‰.
Calculation of methane and sulfate fluxes
Assuming that related flux over the linear range of the sulfate profile was proportional to methane flux at a particular location78, we determined an AOM depth where the linear sulfate and methane concentration gradients overlapped in each core sediment. Diffusive fluxes of solutes (methane and sulfate) into the AOM layers were determined using Fick’s first law, and the slope of methane and sulfate concentration was plotted against depth over a linear range of data79,80 using the equation
where J represents the diffusive methane and sulfate fluxes (mmol m–2 yr–1), φ is the sediment porosity, Ds is the sediment diffusion coefficient, C is the range of methane and sulfate concentrations, and z is the range of depth for the linear section of the methane and sulfate porewater profiles. Ds values were calculated from free diffusion coefficients in seawater (D0, cm2 yr–1) by correcting for sediment tortuosity81:
Downward sulfate diffusion into the sediment is expressed as negative, and upward methane flux out of the sediment is expressed as positive.
Bulk element analysis
Sediment samples were freeze-dried and homogenized using an agate mortar prior to bulk geochemical analyses. For TOC analysis, sediment samples (~1 g) were treated with 8 mL of 1 N HCl to remove carbonates. The TOC content and its isotopic composition were measured using an elemental analyzer (EuroEA3028, Eurovector, Milan, Italy) connected to an isotope ratio mass spectrometer (Isoprime, GV instruments, Manchester, UK) according to Lee et al. (2018). Isotopic values are expressed as δ13C values in per mil relative to VPDB. The δ13C values were calibrated through a CH3 standard with a δ13C value of −24.7‰, certified by the International Atomic Energy Agency (IAEA, Austria). The precision for TOC and δ13CTOC was better than ±0.2 wt.%, and ±0.1‰, respectively. The total sulfur (TS) content and its isotopic composition were measured using an elemental analyzer (EA1110, Thermo) connected to an isotope ratio mass spectrometer (Dual pumped 20–20 S, Secon). Isotopic values are expressed as δ34S values in per mil relative to Vienna Canyon Diablo Troilite (VCDT). The δ34S values were calibrated using S-2 and S-3 standards with δ34S values of 22.7‰ and −32.3‰, respectively, certified by the IAEA. The precision for TS and δ34S was better than ±0.4 wt.% and ±0.4‰, respectively.
Lipid biomarker analysis
Lipid extraction and purification for different compound classes and analyses by gas chromatography, gas chromatography–mass spectrometry, gas chromatography–combustion–isotope ratio mass spectrometry, and high-performance liquid chromatography–atmospheric pressure positive–ion chemical ionization–mass spectrometry were conducted according to previously reported methods30,61,82. Double-bond positions in fatty acid methyl esters were determined by analysis as their dimethyl disulfide adducts according to the method of Nichols et al. (1986). All isotopic values were reported using δ-notation (per mil) with respect to VPDB. The precision for δ13C values was greater than ±0.5‰, corrected for introduction of additional carbon atoms during sylilation and methylation.
Genomic DNA extraction, amplification, and sequencing
Genomic DNA was extracted using an Exgene Soil SV mini kit (Cambio, UK) at GeneAll (Seoul, South Korea). The V4-V5 regions of bacterial 16S rRNA genes and V6-V8 regions of archaeal 16S rRNA genes were amplified by polymerase chain reaction using primer pairs 515F/926R and A956F/A1401R, respectively, at the Integrated Microbiome Resource (IMR) at Dalhousie University, Canada (http://cgeb-imr.ca)83. Sequencing of the amplicons was carried out at IMR using the paired-end (2 × 300 bp) Illumina MiSeq system (Illumina, USA).
16S rRNA gene sequences processing
Pair end sequences were trimmed based on Sickle84 quality scores followed by BayesHammer error corrections85. The resulting quality-trimmed and error-corrected paired end sequences were assembled using PANDAseq.86. Further sequence-processing steps were performed according to the mothur Pipeline package87. The assembled bacterial sequences were aligned against a SILVA alignment (http://www.arb-silva.de), and archaeal sequences were aligned against a SINA alignment88 and subsequently denoised using the “pre.cluster” command. Chimeric sequences were removed using the “chimera.uchime” command in de novo mode89. The sequences were further clustered to OTUs at a 97% sequence similarity level using the opticlust clustering algorithm. Taxonomic assignments of each OTU were determined by sequence similarity searches against the EzBioCloud database90. All sequence data used in this study were deposited in the Sequence Read Archive at the National Center for Biotechnology Information under the accession number PRJNA559003.
Phylogenetic analysis
Phylogenetic trees of major archaeal OTUs of Methanomicrobia, bacterial OTUs of Deltaproteobacteria and Methylococcales of Gammaprotoebacteria with greater than 5% relative abundance (Supplementary Information Sequence 1–3) were constructed using the maximum-likelihood algorithm91 with the GTR evolutionary model using IQ-TREE92.
References
Ivanov, M. K., Limonov, A. F. & Woodside, J. M. Extensive deep fluid flux through the sea floor on the Crimean continental margin (Black Sea). Geol. Soc. London, Spec. Publ. 137, 195–213 (1998).
Shnukov, E. F., Starostenko, V. I., Rusakov, O. M., Kobolev, V. P. & Maslakov, N. A. In International workshop on methane in sediments and water column of the Black Sea: Formation, transport, pathways and the role within the carbon cycle. In Gas volcanism in the Black Sea 50–51 (2005).
Mazzini, A. Mud volcanism: Processes and implications. Mar. Pet. Geol. 26, 1677–1680 (2009).
Kopf, A. J. Significance of mud volcanism. Rev. Geophys. 40, 2–46 (2002).
Milkov, A. V., Sassen, R., Apanasovich, T. V. & Dadashev, F. G. Global gas flux from mud volcanoes: A significant source of fossil methane in the atmosphere and the ocean. Geophys. Res. Lett. 30, 17–20 (2003).
Etminan, M., Myhre, G., Highwood, E. J. & Shine, K. P. Radiative forcing of carbon dioxide, methane, and nitrous oxide: A significant revision of the methane radiative forcing. Geophys. Res. Lett. 43(12), 614–12,623 (2016).
Paull, C. K. et al. Tracking the decomposition of submarine permafrost and gas hydrate under the shelf and slope of the Beaufort Sea. in Proceedings of the 7th International Conference on Gas Hydrates 1, 1689–1699 (2011).
Paull, C. K. et al. Active mud volcanoes on the continental slope of the Canadian Beaufort Sea. Geochemistry Geophys. Geosystems 16, 1541–1576 (2015).
Biastoch, A. et al. Rising Arctic Ocean temperatures cause gas hydrate destabilization and ocean acidification. Geophys. Res. Lett. 38, 1–6 (2011).
Levitus, S., Antonov, J. I., Boyer, T. P. & Stephens, C. Warming of the world ocean. Science. 287, 2225–2229 (2000).
Westbrook, G. K. et al. Escape of methane gas from the seabed along the West Spitsbergen continental margin. Geophys. Res. Lett. 36, 1–5 (2009).
Polyakov, I. V. et al. Arctic Ocean warming contributes to reduced polar ice cap. J. Phys. Oceanogr. 40, 2743–2756 (2010).
Post, E. et al. Ecological dynamics across the arctic associated with recent climate change. Science. 325, 1355–1358 (2009).
Whiteman, G., Hope, C. & Wadhams, P. Vast costs of Arctic change. Nature 499, 401–403 (2013).
Felden, J., Wenzhöfer, F., Feseker, T. & Boetius, A. Transport and consumption of oxygen and methane in different habitats of the Håkon Mosby Mud Volcano (HMMV). Limnol. Oceanogr. 55, 2366–2380 (2010).
James, R. H. et al. Effects of climate change on methane emissions from seafloor sediments in the Arctic Ocean: A review. Limnol. Oceanogr. 61, S283–S299 (2016).
Treude, T., Boetius, A., Knittel, K., Wallmann, K. & Jørgensen, B. B. Anaerobic oxidation of methane above gas hydrates at Hydrate Ridge, NE Pacific Ocean. Mar. Ecol. Prog. Ser. 264, 1–14 (2003).
Levin, L. A. Ecology of cold seep sediments: Interactions of fauna with flow, chemistry and microbes. Oceanogr. Mar. Biol. 43, 1–46 (2005).
Niemann, H. et al. Novel microbial communities of the Håkon Mosby mud volcano and their role as a methane sink. Nature 443, 854–858 (2006).
Omoregie, E. O. et al. Biogeochemistry and community composition of iron- and sulfur-precipitating microbial mats at the Chefren mud volcano (Nile Deep Sea Fan, Eastern Mediterranean). Appl. Env. Microbiol. 74, 3198–3215 (2008).
Boetius, A. & Suess, E. Hydrate Ridge: A natural laboratory for the study of microbial life fueled by methane from near-surface gas hydrates. Chem. Geol. 205, 291–310 (2004).
Boetius, A. & Wenzhöfer, F. Seafloor oxygen consumption fuelled by methane from cold seeps. Nat. Geosci. 6, 725–734 (2013).
Hanson, R. S. & Hanson, T. E. Methanotrophic bacteria. Microbiol. Rev. 60, 439–71 (1996).
Knittel, K. & Boetius, A. Anaerobic oxidation of methane: progress with an unknown process. Annu. Rev. Microbiol. 63, 311–334 (2009).
Knittel, K. et al. Diversity and distribution of methanotrophic archaea at cold seeps diversity and distribution of methanotrophic archaea at cold seeps. Appl. Environ. Microbiol. 71, 467–479 (2005).
Reeburgh, W. Oceanic methane biogeochemistry. Am. Chem. Soc. 107, 486–513 (2007).
Timmers, P. H. A. et al. Reverse methanogenesis and respiration in methanotrophic archaea. Archaea 2017 (2017).
Lösekann, T. et al. Diversity and abundance of aerobic and anaerobic methane oxidizers at the Haakon Mosby Mud Volcano, Barents Sea. Appl. Environ. Microbiol. 73, 3348–3362 (2007).
Felden, J., Ruff, S. E., Ertefai, T., Inagaki, F. & Hinrichs, K. U. Anaerobic methanotrophic community of a 5346-m-deep vesicomyid clam colony in the Japan Trench. Geobiology 12, 183–199 (2014).
Lee, D.-H. et al. Biogeochemical evidence of anaerobic methane oxidation on active submarine mud volcanoes on the continental slope of the Canadian Beaufort Sea. Biogeosciences 15, 7419–7433 (2018).
Elvert, M. & Niemann, H. Occurrence of unusual steroids and hopanoids derived from aerobic methanotrophs at an active marine mud volcano. Org. Geochem. 39, 167–177 (2008).
Elvert, M., Boetius, A., Knittel, K. & Jørgensen, B. B. Characterization of specific membrane fatty acids as chemotaxonomic markers for sulfate-reducing bacteria involved in anaerobic oxidation of methane. Geomicrobiol. J. 20, 403–419 (2003).
Blumenberg, M., Seifert, R., Reitner, J., Pape, T. & Michaelis, W. Membrane lipid patterns typify distinct anaerobic methanotrophic consortia. Proc. Natl. Acad. Sci. 101, 11111–11116 (2004).
Parte, A. C. LPSN - List of prokaryotic names with standing in nomenclature. Nucleic Acids Res. 42, 613–616 (2014).
Whiticar, M. J. Carbon and hydrogen isotope systematics of bacterial formation and oxidation of methane. Chem. Geol. 161, 291–314 (1999).
Tryon, M. D. et al. Measurements of transience and downward fluid flow near episodic methane gas vents, Hydrate Ridge, Cascadia. Geology 27, 1075–1078 (1999).
Wenzhöfer, F. & Glud, R. N. Benthic carbon mineralization in the Atlantic: A synthesis based on in situ data from the last decade. Deep. Res. Part I 49, 1255–1279 (2002).
Luff, R. & Wallmann, K. Fluid flow, methane fluxes, carbonate precipitation and biogeochemical turnover in gas hydrate-bearing sediments at Hydrate Ridge, Cascadia Margin: Numerical modeling and mass balances. Geochim. Cosmochim. Acta 67, 3403–3421 (2003).
Pastor, L. et al. Early diagenesis in the sediments of the Congo deep-sea fan dominated by massive terrigenous deposits: Part III – Sulfate- and methane- based microbial processes. Deep Sea Res. II 142, 139–150 (2017).
Sassen, R. et al. Free hydrocarbon gas, gas hydrate, and authigenic minerals in chemosynthetic communities of the northern Gulf of Mexico continental slope: Relation to microbial processes. Chem. Geol. 205, 195–217 (2004).
Omoregie, E. O. et al. Microbial methane oxidation and sulfate reduction at cold seeps of the deep Eastern Mediterranean Sea. Mar. Geol. 261, 114–127 (2009).
Hoefs, J. Stable isotope geochemistry. Supringer-Verlag (2007).
Yoshinaga, M. Y. et al. Carbon isotope equilibration during sulphate-limited anaerobic oxidation of methane. Nat. Geosci. 7, 190–194 (2014).
Chuang, P. C. et al. Carbon isotope exchange during anaerobic oxidation of methane (AOM) in sediments of the northeastern South China Sea. Geochim. Cosmochim. Acta 246, 138–155 (2019).
Coffin, R. B. et al. Spatial variation in shallow sediment methane sources and cycling on the Alaskan Beaufort Sea Shelf/Slope. Mar. Pet. Geol. 45, 186–197 (2013).
Wilson, R. M. et al. Subsurface methane sources and migration pathways within a gas hydrate mound system, Gulf of Mexico. Geochemistry, Geophys. Geosystems 15, 89–107 (2014).
Dubilier, N., Bergin, C. & Lott, C. Symbiotic diversity in marine animals: The art of harnessing chemosynthesis. Nat. Rev. Microbiol. 6, 725–740 (2008).
Hilário, A. et al. New perspectives on the ecology and evolution of siboglinid tubeworms. PLoS One 6, e16309 (2011).
Wallmann, K. et al. Quantifying fluid flow, solute mixing, and biogeochemical turnover at cold vents of the eastern Aleutian subduction zone. Geochim. Cosmochim. Acta 61, 5209–5219 (2002).
Levin, L. A. et al. Spatial heterogeneity of macrofauna at northern California methane seeps: Influence of sulfide concentration and fluid flow. Mar. Ecol. Prog. Ser. 265, 123–139 (2003).
Ruff, S. E. et al. Microbial communities of deep-sea methane seeps at Hikurangi continental margin (New Zealand). PLoS One 8, e72627 (2013).
Pop Ristova, P. et al. Bacterial diversity and biogeochemistry of different chemosynthetic habitats of the REGAB cold seep (West African margin, 3160 m water depth). Biogeosciences 9, 5031–5048 (2012).
Ruff, S. E. et al. In situ development of a methanotrophic microbiome in deep-sea sediments. ISME J. 13, 197 (2018).
Ruff, S. E. et al. Global dispersion and local diversification of the methane seep microbiome. Proc. Natl. Acad. Sci. USA 112, 4015–20 (2015).
de Beer, D. et al. In situ fluxes and zonation of microbial activity in surface sediments of the Håkon Mosby mud volcano. Limnol. Ocean. 51, 1315–1331 (2006).
Coolen, M. J. L., Muyzer, G., Schouten, S., Volkman, J. K. & Sinninghe Damsté, J. S. In Past and Present Water Column Anoxia 41–65 (Springer, 2006).
Canuel, E. A. & Martens, C. S. Seasonal variations in the sources and alteration of organic matter associated with recently-deposited sediments. Org. Geochem. 20, 563–577 (1993).
Gong, C. & Hollander, D. J. Differential contribution of bacteria to sedimentary organic matter in oxic and anoxic environments, Santa Monica Basin, California. Org. Geochem. 26, 545–563 (1997).
Coffin, R. B. et al. Deep sediment-sourced methane contribution to shallow sediment organic carbon: Atwater Valley, Texas-Louisiana Shelf, Gulf of Mexico. Energies 8, 1561–1583 (2015).
Niemann, H. & Elvert, M. Diagnostic lipid biomarker and stable carbon isotope signatures of microbial communities mediating the anaerobic oxidation of methane with sulphate. Org. Geochem. 39, 1668–1677 (2008).
Stadnitskaia, A., Ivanov, M. K. & Sinninghe Damsté, J. S. Application of lipid biomarkers to detect sources of organic matter in mud volcano deposits and post-eruptional methanotrophic processes in the Gulf of Cadiz, NE. Atlantic. 255, 1–14 (2008).
Chevalier, N. et al. Authigenic carbonates at cold seeps in the Marmara Sea (Turkey): A lipid biomarker and stable carbon and oxygen isotope investigation. Mar. Geol. 288, 112–121 (2011).
Schouten, S., Hopmans, E. C. & Sinninghe Damsté, J. S. The organic geochemistry of glycerol dialkyl glycerol tetraether lipids: A review. Org. Geochem. 54, 19–61 (2013).
Wegener, G., Krukenberg, V., Ruff, S. E., Kellermann, M. Y. & Knittel, K. Metabolic capabilities of microorganisms involved in and associated with the anaerobic oxidation of methane. Front. Microbiol. 7, 1–16 (2016).
Holler, T. et al. Thermophilic anaerobic oxidation of methane by marine microbial consortia. ISME J. 5, 1946–1956 (2011).
Vigneron, A. et al. Archaeal and anaerobic methane oxidizer communities in the Sonora Margin cold seeps, Guaymas Basin (Gulf of California). ISME J. 7, 1595–1608 (2013).
Gründger, F. et al. Methane-fuelled biofilms predominantly composed of methanotrophic ANME-1 in Arctic gas hydrate-related sediments. Sci. Rep. 9, 9725 (2019).
Elvert, M., Hopmans, E. C., Treude, T., Boetius, A. & Suess, E. Spatial variations of methanotrophic consortia at cold methane seeps: Implications from a high-resolution molecular and isotopic approach. Geobiology 3, 195–209 (2005).
Wegener, G., Niemann, H., Elvert, M., Hinrichs, K. & Boetius, A. Assimilation of methane and inorganic carbon by microbial communities mediating the anaerobic oxidation of methane. Environ. Microbiol. 10, 2287–2298 (2008).
Rossel, P. E. et al. Intact polar lipids of anaerobic methanotrophic archaea and associated bacteria. Org. Geochem. 39, 992–999 (2008).
Rossel, P. E., Elvert, M., Ramette, A., Boetius, A. & Hinrichs, K. U. Factors controlling the distribution of anaerobic methanotrophic communities in marine environments: Evidence from intact polar membrane lipids. Geochim. Cosmochim. Acta 75, 164–184 (2011).
Gonzalez-Gil, G., Meulepas, R. J. W. & Lens, P. N. L. Biotechnological aspects of the use of methane as electron donor for sulfate reduction. (Academic Press, Burlington, 2011).
Lösekann, T. et al. Endosymbioses between bacteria and deep-sea siboglinid tubeworms from an Arctic Cold Seep (Håkon Mosby mud volcano, Barents Sea). Environ. Microbiol. 10, 3237–3254 (2008).
Lee, D.-H. et al. Chemosynthetic bacterial signatures in frenulata tubeworm (Oligobrachia sp.) in an active mud volcano of the Canadian Beaufort Sea. Mar. Ecol. Prog. Ser. 628, 95–104 (2019).
Knittel, K. et al. Activity, distribution, and diversity of sulfate reducers and other bacteria in sediments above gas hydrate (Cascadia margin, Oregon). Geomicrobiol. J. 20, 269–294 (2003).
Kleindienst, S., Ramette, A., Amann, R. & Knittel, K. Distribution and in situ abundance of sulfate-reducing bacteria in diverse marine hydrocarbon seep sediments. Environ. Microbiol. 14, 2689–2710 (2012).
Milucka, J. et al. Zero-valent sulphur is a key intermediate in marine methane oxidation. Nature 491, 541–6 (2012).
Borowski, W. S., Paull, C. K. & Ussier, W. Carbon cycling within the upper methanogenic zone of continental rise sediments: An example from the methane-rich sediments overlying the blake ridge gas hydrate deposits. Mar. Chem. 57, 299–311 (1997).
Berner, R. A. An idealized model of dissolved sulfate distribution in recent sediments. Geochim. Cosmochim. Acta 28, 1497–1503 (1964).
Berner, R. A. Sulfate reduction and the rate of deposition of marine sediments. Earth Planet. Sci. Lett. 37, 492–498 (1978).
Boudreau, B. P. Diagenetic models and their implementation, Modelling transport and reactions in aquatic sediments. (Springer, Germany., 1997). https://doi.org/10.1016/S0264-8172(98)80005-6
Kim, J.-H. et al. Contribution of petroleum-derived organic carbon to sedimentary organic carbon pool in the eastern Yellow Sea (the northwestern Pacific). Chemosphere 168, 1389–1399 (2017).
Walters, W. et al. Improved Bacterial 16S rRNA Gene (V4 and V4-5) and Fungal Internal Transcribed Spacer Marker Gene Primers for Microbial Community Surveys. mSystems 1, e00009–15 (2016).
Schirmer, M. et al. Insight into biases and sequencing errors for amplicon sequencing with the Illumina MiSeq platform. Nucleic Acids Res. 43 (2015).
Nikolenko, S. I., Korobeynikov, A. I. & Alekseyev, M. A. BayesHammer: Bayesian clustering for error correction in single-cell sequencing arXiv: 1211. 2756v1 [q-bio. QM] 12 Nov 2012. BMC Genomics 14, 1–11 (2013).
Masella, A. P., Bartram, A. K., Truszkowski, J. M., Brown, D. G. & Neufeld, J. D. PANDAseq: Paired-end assembler for illumina sequences. BMC Bioinformatics 13, 1–7 (2012).
Schloss, P. D. et al. Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 75, 7537–7541 (2009).
Pruesse, E., Peplies, J. & Glöckner, F. O. SINA: Accurate high-throughput multiple sequence alignment of ribosomal RNA genes. Bioinformatics 28, (1823–1829 (2012).
Edgar, R. C., Haas, B. J., Clemente, J. C., Quince, C. & Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27, 2194–2200 (2011).
Yoon, S. H. et al. Introducing EzBioCloud: A taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int. J. Syst. Evol. Microbiol. 67, 1613–1617 (2017).
Felsenstein, J. Evolutionary trees from DNA sequences: A maximum likelihood approach. J. Mol. Evol. 17, 368–376 (1981).
Trifinopoulos, J., Nguyen, L. T., von Haeseler, A. & Minh, B. Q. W-IQ-TREE: a fast online phylogenetic tool for maximum likelihood analysis. Nucleic Acids Res. 44, W232–W235 (2016).
Acknowledgements
We would like to thank the captain and crew of the RV Araon for their help at sea. We are grateful to Dale Graves, Frank Flores, and Roberto Gwiazda at Monterey Bay Aquarium Research Institute for remotely operated vehicle operation and sample collection. We also thank Sujin Kang, Dahae Kim, and Solbin Kim for analytical assistance in the laboratory at Hanyang University. This study was supported by the Korea Institute of Marine Science and Technology Promotion (KIMST Grant 20160247; KOPRI-PM18050) and by the David and Lucile Packard Foundation.
Author information
Authors and Affiliations
Contributions
D.-H.L., Y.M.L., and J.-H.K. designed and coordinated the study and led the writing. Y.K.J., C.P., and K.-H.S. contributed to sample acquisition. D.-H.L. conducted experiments for gas, porewater, and lipid biomarkers, and Y.M.L. analyzed microbial diversity. Ji.-H.K. provided methane and DIC concentration data. D.-H.L., Y.M.L., J.-H.K., and H.N. led most data interpretation. All authors commented on the manuscript and contributed to the writing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lee, DH., Lee, Y.M., Kim, JH. et al. Discriminative biogeochemical signatures of methanotrophs in different chemosynthetic habitats at an active mud volcano in the Canadian Beaufort Sea. Sci Rep 9, 17592 (2019). https://doi.org/10.1038/s41598-019-53950-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-53950-4
This article is cited by
-
Methane hydrate dissociation across the Oligocene–Miocene boundary
Nature Geoscience (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.