Abstract
Asthma and hypertension are complex diseases coinciding more frequently than expected by chance. Unraveling the mechanisms of comorbidity of asthma and hypertension is necessary for choosing the most appropriate treatment plan for patients with this comorbidity. Since both diseases have a strong genetic component in this article we aimed to find and study genes simultaneously associated with asthma and hypertension. We identified 330 shared genes and found that they form six modules on the interaction network. A strong overlap between genes associated with asthma and hypertension was found on the level of eQTL regulated genes and between targets of drugs relevant for asthma and hypertension. This suggests that the phenomenon of comorbidity of asthma and hypertension may be explained by altered genetic regulation or result from drug side effects. In this work we also demonstrate that not only drug indications but also contraindications provide an important source of molecular evidence helpful to uncover disease mechanisms. These findings give a clue to the possible mechanisms of comorbidity and highlight the direction for future research.
Similar content being viewed by others
Introduction
Asthma and hypertension affect hundreds of millions of people worldwide1,2 and coincide in adults more frequently than expected by chance. Patients with asthma are more likely to have high blood pressure3 and, in turn, the presence of hypertension is associated with the increased severity of asthma4. This association between asthma and hypertension was confirmed in multiple studies of different patient cohorts3,4,5,6,7 but its underlying causes remain unknown. The coexistence of two or more diseases, called comorbidity, was shown for many human disorders8,9,10. Moreover, not only complex disorders demonstrate comorbidity but also several Mendelian diseases coincide with complex disorders10,11. Comorbidity can be direct or inverse8,12, when the presence of one disease protects from the development of another one. For example, the coincidence of neoplasms with several nervous system disorders is lower than expected13,14.
Comorbidity may point to causal relationships between two diseases. For instance, hypertension may cause cardiovascular damage, leading to consequences such as heart failure, stroke, and kidney problems2. Alternatively, comorbidity may result from confounder effects, e.g. lifestyle or environmental factors, predisposing to multiple health problems. For instance, smoking is a risk factor for multiple diseases15, including hypertension2 and asthma1. Also, stress may trigger asthma attack1, and long-term stress exposure is associated with the risk of hypertension2. Finally, comorbidity of two diseases may be an effect of a third disease. For instance, obesity is another common risk factor for asthma and hypertension and decreasing of the body-mass index is helpful for management of both diseases16,17.
In addition to these risk factors, some anti-asthmatic drugs may worsen hypertension and vice versa, and several anti-hypertensive drugs are contraindicated in asthma. For example, beta-blockers used to control blood pressure can cause asthma attacks18,19 and therefore are contraindicated for asthma patients. At the same time, beta-agonists, used for treatment of asthma may increase heart rate20 and should be used with caution in patients with hypertension. Corticosteroids are aimed at suppressing the immune system and are used against asthma1, but can at the same time elevate blood pressure due to their effect on kidneys, leading to enhanced liquid retention21.
Nevertheless, comorbidity between asthma and hypertension can only in part be explained by excessive weight, smoking and the use of specific drugs and persists after consideration of these variables, although becomes weaker4. Besides environmental risk factors, including other diseases and drugs, comorbidity may arise as the result of shared molecular genetic basis22, predisposing the patient to the development of both diseases. Comorbid diseases may share associated genes themselves23,24,25 or demonstrate strong connectivity between two sets of associated genes in protein interaction11,26,27,28,29, gene coexpression26 or metabolic networks30.
Recent studies have shown that many epidemiologically correlated diseases and traits share risk loci identified in genome-wide association studies (GWAS)31,32, but this is not true for hypertension and asthma31. However, GWAS results are not the only information source of gene-disease associations. Many different approaches are used to establish associations between genes and diseases. For example, genes altering expression in disease and genes encoding drug targets are likely to be also involved in its pathogenesis.
Despite the fact that many genes are associated with isolated asthma and hypertension, molecular mechanisms underlying their comorbidity remain unclear. To investigate the genetic basis of this comorbidity, we constructed comprehensive lists of asthma and hypertension associations of different nature and identified genes implicated in both diseases. We analyzed the relationships between these genes, revealed six tightly interconnected gene modules and characterized these modules by enriched GO terms, pathways and tissue specificity. Identification of the genes and gene modules potentially involved in both disorders may advance the unraveling the mechanisms of their comorbidity, and help get further insight into the pathogenesis of asthma and hypertension comorbidity.
Results
Genes previously associated with asthma and hypertension
Although the phenomenon of comorbidity of asthma and hypertension has long been known, only a few works discussing genes potentially involved in both diseases or related pathophysiological processes have been published before 2018. We have found six publications which discussed ten genes in total potentially relevant to both asthma and hypertension and summarized the results of our literature search in Table 1. These studies were mostly focused on genes and their functions and did not aim to discover the mechanisms of comorbidity between the two diseases.
In our recent work33, we applied ANDSystem34,35 to perform the automatic reconstruction of the associative gene network for asthma and hypertension from scientific literature. The resultant associative network included 205 genes potentially responsible for comorbid asthma and hypertension. We proposed gene prioritization methodology based on ten criteria including relevance scores calculated by state-of-the-art gene prioritization tools36,37, association with biological processes, position in the associative network and evidence of genetic associations and regulations. The IL10, TLR4, and CAT genes had the highest ranks among all candidate genes, and ADRB2 was ranked sixth. Our later works demonstrated experimental evidence supporting possible roles of IL10 and TLR4 in comorbid asthma and hypertension. Drevytska et al. created an animal model of comorbid asthma and hypertension – SHR with OVA-induced asthma – and demonstrated that IL10 knock-down in this model improves cardiac and lung function parameters38. Bragina et al. identified expression quantitative trait loci (eQTL) associated with asthma and hypertension comorbidity for CAT, TLR4, ANG and RNASE4 genes on the cohort of 587 individuals from West Siberia39. More recently, Saik et al. reanalyzed associative asthma/hypertension network and, based on the cross-talk centrality, proposed ten more immune-related genes for further experimental validation40.
All the previous works investigated the genetic basis of asthma/hypertension comorbidity evaluated the relevance of single genes. In contrast with previous studies, in this work, among all the associations extracted from a variety of data sources, we identified and characterized groups of functionally related genes. Our approach aligns with current understanding of asthma and hypertension as complex disorders, which thought to be polygenic and heterogeneous41,42. Yet another advantage of this work is that we incorporated types of associations not considered in previous studies nor provided by large gene-disease association portals, such as Open Targets43 or DisGeNET44. In particular, we included into the analysis genes controlled by eQTLs associated with asthma and hypertension and target genes of drugs contraindicated in these diseases.
The overlap between genes associated with asthma and hypertension independently
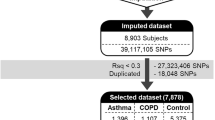
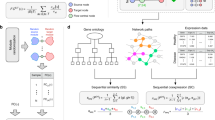
Aiming to compose comprehensive lists of genes associated with asthma and hypertension and investigate their overlap, we collected gene-disease associations supported by a variety of evidence types: genetic associations, differential expression, co-occurrence in texts, and targeting by drugs used against asthma and hypertension. Also, to expand these gene lists, we used GWAS hits mapped to known eQTLs and selected genes regulated by these eQTLs. Finally, we hypothesized, that genes targeted by drugs causing adverse effects similar to asthma and hypertension may also be involved in their pathogenesis. Therefore, in addition to drugs that can treat asthma and hypertension, we took into account drugs that exaggerate these diseases and added their targets in the lists of genes for asthma or hypertension.
After combination of gene-disease associations from various evidence sources (see Methods and Fig. 1), the lists of 980 genes associated with asthma and 1204 genes associated with hypertension were compiled. Out of them, 330 genes were associated with both diseases according to at least one data source (Supplementary Table S1). To facilitate further use of associations relevant for asthma and hypertension comorbidity, we added them to GenCoNet database45.
Identification and characterization of gene modules associated with asthma and hypertension. Network nodes represent genes and are colored according to membership in a module. Nodes not assigned to clusters are shown in grey. Size of each node is proportional to the number of evidence sources supporting the association of corresponding gene with asthma or hypertension. All gene set overlap analysis results are shown in Supplementary Tables S2A–C.
Network analysis of shared genes reveals six modules
In order to identify groups of functionally related genes, we built a functional network of shared asthma/hypertension genes using Cytoscape46 v3.6.1 stringApp47 v1.3.0 and performed module detection. In this network, two nodes were connected, if corresponding genes are co-expressed or if proteins physically interact or share biological pathways. Only edges representing gene co-expression, physical interactions of encoded proteins and biological pathway sharing, were included in the network. Overall, 257 out of 330 genes (78%) were connected with at least one of other genes. Six tightly interconnected modules were detected using ClusterViz App48 v1.0.3 EAGLE algorithm49 (with minimal Clique Size and Complex Size thresholds set to 10) and annotated with overrepresented pathways, GO terms, and tissue-specific gene sets (Supplementary Tables S2A–C provide all significant gene set overlap analysis results).
Module 1 comprised 77 genes was enriched with genes participating in cytokine signaling, in particular, in IL-4, IL-13, and IL-23, and Th17 differentiation pathways. IL-4 and IL-13 regulate inflammatory response to allergen exposure in asthma50. IL-23 and Th17 cells are elevated in hypertension51 and promote neutrophilic inflammation in asthma52. A very recent review on the treatment of comorbid asthma and hypertension summarizes implications of cytokine signaling and Th17 cells for both diseases53. Module 2 comprised 43 genes participating in the metabolism of xenobiotics and defending the cell from oxidative stress. Reactive oxygen species are generated by airway epithelium cells exposed to irritants, activate immune cells and thought to be implicated in asthma54. Hypertension is characterized by elevated oxidative stress biomarkers in blood and increased oxidative damage of vasculature55,56. A large body of evidence confirms roles of antioxidant defence enzymes in asthma57,58,59 and hypertension60,61. Interestingly, a key antioxidant enzyme, catalase (CAT), was also previously associated with comorbid asthma and hypertension33,39. Module 3 enriched with genes involved in extracellular matrix organization and platelet degranulation. Platelets are altered in hypertension62 and their release products participate in airway inflammation and remodeling in asthma63. Module 4 was enriched by genes encoding G protein-coupled receptors and responsible for regulation of blood vessel diameter. This module included adrenoreceptor genes, many of which are targeted by drugs against asthma and hypertension. Module 5 was enriched with genes participating in cytokine signaling. In contrast to module 1, genes from module 5 were associated with interferon gamma response and more specific for CD16-monocytes and sputum, than for neutrophils. Interferon-gamma inhibits Th2-induced inflammation but promotes cytotoxic response64. Interferon gamma signaling is implicated in asthma64 and angiotensin-II-induced hypertension65. Module 6 was composed of genes encoding histones, whose modifications define chromatin state and regulate gene expression. Changes of histone modifications are shown in many diseases including asthma66 and hypertension67. Inhibition of histone deacetylases decreases inflammations68 and demonstrate anti-hypertensive effects67,69.
Some modules overlapped in a few genes: MAPK1 and EDN1 belonged to modules 1 and 4, C3 and C5 to modules 2 and 4, STAT1 to modules 1 and 5, and SERPINA1 to modules 2 and 3. Such genes may be responsible for a cross-talk between biological processes represented by modules these genes connect. However, of these six genes, only MAPK1 and STAT1 were associated with asthma or hypertension through evidence other than co-occurrence in texts.
Analysis of evidence types supporting gene-disease associations
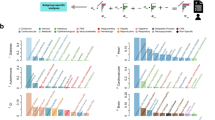
To investigate the contribution of various evidence types to the detection of shared genes, we labeled every gene with the kind of relationship, linking it with asthma or hypertension, and analyzed the distribution of labels among six modules. Figure 2 shows that the distribution of labels over the network does not seem to be random, and the representations of different evidence type labels among modules are not equal. The majority of shared genes (231 of 330) had an association established via text mining by HuGE Phenopedia70 v2.1 and were supported by at least two independent studies. Moreover, associations of 125 of them were established via text mining only and constituted essential parts of modules 1–4. Although frequent co-occurrence in texts does not seem to be the most confident evidence of association, we could not exclude it from consideration without losing 206 (62%) of shared genes.
Evidence sources supporting gene associations with asthma and hypertension. In this figure, we used node style similar to Fig. 1B in103. Here, nodes represent genes associated with both asthma and hypertension, edges correspond gene interactions (only 257 nodes connected with at least one other node are shown). Genes are colored according to evidence sources (see figure legend) from which associations came from. The size of each node is proportional to the number of evidence sources supporting its association with asthma and hypertension. All 330 genes associated with asthma and hypertension annotated with evidence types supporting associations are listed in Supplementary Table S1.
We also found that all members of module 6 and almost all of module 5 included genes that are controlled by GWAS-identified eQTLs associated with asthma or hypertension. In total, 107 out of 330 shared genes were regulated by such eQTLs. Since this number significantly exceeded the expected by chance (Fisher’s exact test p-value < 6.94e-145; permutation p-value < 10e-4; 10 expected on average), we can hypothesize that these shared genes may have an impact on the development of comorbidity between asthma and hypertension. Importantly, this observation cannot be explained simply by shared GWAS hits. Only one missense variant, rs3184504 in SH2B3 gene, was in the intersection of asthma and hypertension GWAS associations. It was independently associated with hypertension, blood eosinophil count and many other traits and controlled only 39 of 107 genes associated with asthma and hypertension through eQTL regulation. Interestingly, among these 39 genes 5 were up-regulated in blood of patients with hypertension and included in the “blood pressure signature” proposed by Huan et al.71: FOS, MYADM, PPP1R15A, S100A10, and TAGAP.
To illustrate the novelty of our findings, we compared our set of shared genes with gene sets obtained from Open Targets v18.02 and DisGeNET v5.0 using trivial disease names. In our list of 330 shared genes, 82 were novel (nodes shown in bold frames in Fig. 2), i.e. not associated with asthma and hypertension in OpenTargets and DisGeNET. These two large gene-disease association databases use distinct sets of evidence sources and interpret some associations differently. For example, they do not consider gene regulation data for establishing gene-disease associations, although it is implicated in asthma72 and hypertension71. Open Targets and DisGeNET include gene-disease associations obtained directly from GWAS Catalog, where disease-associated variants are mapped to neighboring genes and do not differentiate synonymous and non-synonymous variants. As a result, most of the newly identified shared genes were associated via eQTLs and composed modules 5 and 6.
Adverse drug reactions may mediate comorbidity of asthma and hypertension
We observed more common drugs (and therefore drug targets) relevant to asthma and hypertension than expected by chance. We composed four lists of drugs that influence asthma or hypertension (Supplementary Table S3): drugs with positive effects on asthma or hypertension (i.e. used to treat or relieve their symptoms) and drugs with negative effects on these diseases (i.e. contraindicated or worsening disease symptoms). Eight drugs used to treat hypertension were harmful for asthma patients: timolol, nadolol, sotalol, pindolol, carvedilol, labetalol, propranolol. All these drugs belong to the class of non-selective beta-blockers and may exacerbate asthma19. At the same time, seven anti-asthmatic drugs occurred in the list of drugs that may increase the risk of hypertension or elevate blood pressure: triamcinolone, prednisolone, methylprednisolone, dexamethasone, hydrocortisone (corticosteroids) and epinephrine, ephedra, ephedrine (beta-agonists). Both overlaps were statistically significant with p-value < 2.64e-06 and p-value < 4.50e-06, respectively in Fisher’s exact test.
To find target genes whose activation or inhibition exhibit positive or negative influence on asthma and hypertension, we performed target overrepresentation analysis in four drug groups. 96 genes were significantly overrepresented among targets of at least one of four drug groups (Table 2 and Supplementary Table S4), only 16 of which were in asthma-hypertension network. Interestingly, 8 of these 16 genes associated with asthma or hypertension through drug evidence appeared in module 4, enriched with genes involved in smooth muscle contraction. Figure 3 summarizes all significant drug targets, their relationships with drugs, and drug effects on asthma and hypertension. Supplementary Table S5 provides this network in tabular format. ADRB1 and ADRB2 were targeted by drugs from all the four groups, and activation and inhibition of beta-adrenoreceptors caused opposite effects on asthma and hypertension. Similarly, NR3C1, a glucocorticoid receptor, was activated by drugs indicated in asthma but potentially harmful for hypertension. ANXA1 is another target of corticosteroid drugs, such as dexamethasone and hydrocortisone, its activation mediates the anti-inflammatory effect via inhibition of phospholipase A273. ANXA1 is up-regulated in blood of hypertensive patients71. PTGS2 inhibition seemed to have a negative effect on both diseases, while PTGS1 was overrepresented only among targets of drugs contraindicated in asthma.
Our results agree with prior knowledge about asthma and hypertension medications. It is well known that \(\beta \)-blockers and agonists have opposite effects on asthma and hypertension. Although medications for treating hypertension and asthma target different receptors (ADRB1 blockers have anti-hypertensive effect2 and ADRB2 agonists have anti-asthmatic effect1), there are many components non-specifically targeting multiple types of adrenoreceptors19. Blockers of \({\alpha }_{1}\) and agonists of \({\alpha }_{2}\)-adrenergic receptors cure hypertension2, and opposite actions on these receptors may provoke blood pressure elevation. Alternative treatment options for asthma patients who poorly respond to beta-agonists are inhaled or systemic glucocorticoids74. Overdosage of glucocorticoids induces elevation of blood pressure, although inhaled glucocorticoids are relatively safe compared to systemic steroids because of their local action21. Patients with an elevated level of glucocorticoids due to their overproduction or abnormal metabolism develop Cushing syndrome75, characterized by high blood pressure among other symptoms. Other classes of anti-inflammatory drugs, such as non-steroid anti-inflammatory drugs (NSAID) may also influence asthma and hypertension. NSAID-Exacerbated Respiratory Disease (NERD), is a classic example of adverse drug reaction in response to aspirin and some other NSAIDs. NERD is also called aspirin-induced asthma because it resembles main asthma symptoms, such as bronchial obstruction and dyspnea, and is prevalent among asthma patients76. NERD is thought to result from PTGS1 inhibition, therefore the usage of PTGS2 selective inhibitors is considered to be relatively safe for asthmatics77, but it is associated with a risk of hypertension78,79. Also, NSAIDs may decrease the effect of anti-hypertensive medications80.
Discussion
In this study, we investigated the genetic overlap between a pair of comorbid disorders, asthma and hypertension. A correlation of gene perturbation with disease status points to the possible involvement of this gene in the disease mechanism. At the same time, different molecular lesions may have similar effects on the phenotype. These considerations motivated us to construct two sets of genes independently associated with each disease through various perturbation types and investigate their overlap. We found 330 genes simultaneously associated with both diseases and potentially responsible for their comorbidity. Projecting these shared genes to an interaction network revealed six functional modules comprising tightly interconnected genes. We tested these modules for the overrepresentation of GO terms and pathways, tissue-specificity and evidence types supporting gene associations.
We observed an excess of genes jointly controlled by asthma- and hypertension-associated eQTLs in modules enriched with genes involved in interferon-gamma signaling and chromatin assembly. This observation suggests that the coincidence of asthma and hypertension may be at least partially explained by concordantly altered genetic regulation of certain biological processes of functions. Our finding agrees with very recent results published by Li et al.81, who analyzed multiple disease pairs and demonstrated that comorbid diseases share significantly more eQTL-regulated transcripts than expected by chance.
We also found more drugs with opposite effects on asthma and hypertension than expected by chance. Based on this observation, we hypothesized that comorbidity might be the result of drug side effects, when drugs against one disease may predispose the patient to the development of another. This particular case of asthma and hypertension demonstrates that genes targeted by contraindicated drugs may also participate in pathophysiologic mechanisms of comorbidity.
An important limitation of this study is the absence of direct validation of the resulting associated gene set. Since the genetic basis of asthma and hypertension comorbidity is poorly understood, we have no gold standard to compare with. Another way of result evaluation would be an experimental validation of gene roles in the comorbidity, which was not in the scope of this work.
The validity of our results is partially confirmed by the presence of some known shared genes in our set. Four genes (ADRB1, ADRB2, TLR4, and CD14) listed in Table 1 appear among 330 genes associated with asthma and hypertension. This limited overlap may be explained by the fact that most of the papers included in Table 1 do not focus on asthma or hypertension and only list them among multiple phenotypes associated with certain genes. Therefore, if associations of these genes with asthma and hypertension do not appear in abstracts, they remain invisible for text mining tools.
Another indirect confirmation of the result validity is our observation that comorbid diseases share many genes controlled by eQTLs. This agrees with the results of Li et al.81, obtained independently on different datasets.
Methods
Monogenic associations via hereditary asthma and hypertension
From MalaCards82 v4.5, and literature2,83 we extracted genes that carry mutations causing monogenic syndromes with asthma or hypertension among other symptoms. In total, we found 37 genes associated with familial forms of hypertension or with Mendelian disorders characterized by hypertension. No monogenic forms of asthma were found, although several Mendelian syndromes characterized by frequent asthma attacks considered in the literature83. From human phenotype ontology84 (HPO) we selected phenotype terms related to asthma (HP:0002099 - Asthma, HP:0012042 - Aspirin-induced asthma, HP:0025428 - Bronchospasm) and hypertension (HP:0000822 - Hypertension, HP:0000875 - Episodic hypertension, HP:0004421 - Elevated systolic blood pressure, HP:0004972 - Elevated mean arterial pressure, HP:0005117 - Elevated diastolic blood pressure). We excluded genes associated with specific kinds of hypertension (e.g. ocular hypertension) that are not related to essential hypertension (HP:0007906, HP:0001409, HP:0002092, HP:0002640, HP:0008071, HP:0100817, HP:0005168). Mapping of genes to HPO terms was downloaded from http://compbio.charite.de/jenkins/job/hpo.annotations.monthly/lastStableBuild/, table ALL_SOURCES_FREQUENT_diseases_to_genes_to_phenotypes.txt” available on 26.01.2018. We used gene-phenotype pairs marked as “frequent”, which means that this phenotype was manifested in at least 50% patients with a disease.
Coding GWAS variants
GWAS Catalog85 v1.0.1 (downloaded on 22.08.2017) includes 29 studies related to asthma which report 407 associations in total (395 unique). In addition, we included 17 associations from 4 studies of asthma-related traits such as NERD, eosinophil count or Immunoglobulin E (IgE) levels. Similarly, 120 associations with hypertension and blood pressure traits were obtained from 16 studies listed in GWAS Catalog. All associations have p-value < 10e-5 (default in GWAS Catalog). Only 29 and 9 single-nucleotide polymorphisms (SNPs) associated with asthma and asthma-related traits and with hypertension and blood pressure respectively were annotated as splice_region_variant, missense_variant, synonymous_variant or non_coding_transcript_exon_variant and therefore affected transcript sequences.
Regulation
Although only a small fraction of variants discovered in GWAS affects transcripts, about a half of them overlap with eQTLs86. From three recent blood eQTL studies86,87,88 comprising thousands of individuals, we obtained lists of eQTL SNPs and overlapped them with variants associated with asthma and hypertension in GWAS. Furthermore, we included variants associated with tissue-specific expression from GTEx v689. This dataset contained eQTLs in 44 tissues including whole blood. The details of eQTL data sources used in this study are presented in Table 3. From each study, we retained only SNP-gene pairs which passed the false discovery rate (FDR) threshold of 0.05. All SNPs coordinates were lifted over to hg38 and their IDs were converted into dbSNP90 v150 to match SNPs IDs used in GWAS catalog.
Differentially expressed genes
We obtained asthma and hypertension expression signatures from the two biggest expression profiling studies of blood pressure71 and asthma91. From Huan et al. we took 34 genes with expressions associated with hypertension diagnosis or with systolic and diastolic blood pressure. Asthma expression signature included 541 genes differentially expressed in the blood of asthma patients in Bigler et al. dataset. In addition to these two signatures, we composed two lists of genes that demonstrated altered expression levels in asthma or hypertension according to ANDSystem (published on 09.10.2014) and DisGeNET v5.0. These two resources provide collections of differentially expressed genes automatically extracted from biomedical literature. To reduce false positives, we took genes whose altered expression in asthma and hypertension was supported by at least two evidence sources.
Drugs and drug targets
Drugs indicated and contraindicated in asthma and hypertension were obtained from DrugBank92 v5.0.9 and NDF-RT93 (released on 02.10.2017). Surprisingly, 5 anti-asthmatic drugs (budesonide, flunisolide, salmeterol, formoterol, and cromoglicic acid) were at the same time contraindicated in asthma according to NDF-RT. These drugs are known to cause paradoxical drug reactions, for example, when the intake of an anti-asthmatic drug provokes bronchospasm94. We excluded such drugs from contraindications list but kept them in the list of indications, because paradoxical reactions are extremely rare. Finally, the list of contraindicated drugs was extended with drugs reported to induce the rise of blood pressure and asthma, NERD or bronchospasms. From DrugBank, we obtained genes targeted by selected drugs. All target genes encoding non-human proteins were excluded. All drug action types used in DrugBank were converted to simplified effects reflecting the direction of drug action on its target, e.g. “positive” for agonists and activators, “negative” for inhibitors and antagonists, and “other” for modifiers. Supplementary Table S3 contains selected drugs, their effects on diseases and their targets.
Statistical tests
Gene set overrepresentation tests were carried out in TargetMine95 (build 20180327) with default parameters (one-tailed Fisher’s exact test followed by Benjamini-Hochberg correction for multiple testing; p-value threshold 0.05; all genes from the tested database were considered as the background set). Similarly, overrepresentation of drug targets in groups of drugs evaluated using the one-sided Fisher’s exact test implemented in scipy python library. We considered all the drugs from DrugBank with defined action on any human target as the background, and applied p-value threshold of 0.05 after Benjamini-Hochberg correction for multiple testing.
To demonstrate that asthma and hypertension share more eQTL regulated genes than expected by chance, we generated 10000 pairs of random eQTL sets and calculated the number of shared genes for each pair. Sizes and overlap of randomly generated eQTL sets were set the same as real sets of asthma and hypertension eQTLs.
Data availability
All data generated during this study are included in Supplementary Information files.
References
Global strategy for asthma management and prevention. https://ginasthma.org (2017).
Mancia, G., Grassi, G. & Redon, J. (eds) Manual of Hypertension of the European Society of Hypertension, Second Edition, https://doi.org/10.1201/b17072 (CRC Press, 2014).
Dogra, S., Ardern, C. I. & Baker, J. The relationship between age of asthma onset and cardiovascular disease in canadians. J. Asthma 44, 849–854, https://doi.org/10.1080/02770900701752391 (2007).
Christiansen, S. C. et al. Hypertension and asthma: A comorbid relationship. The J. Allergy Clin. Immunol. In Pract. 4, 76–81, https://doi.org/10.1016/j.jaip.2015.07.009 (2016).
Johnson, M., Nriagu, J., Hammad, A., Savoie, K. & Jamil, H. Asthma, environmental risk factors, and hypertension among arab americans in metro detroit. J. Immigr. Minor. Heal. 12, 640–651, https://doi.org/10.1007/s10903-008-9205-8 (2010).
Ferguson, S. et al. Factors associated with systemic hypertension in asthma. Lung 192, 675–683, https://doi.org/10.1007/s00408-014-9600-y (2014).
Heck, S. et al. High probability of comorbidities in bronchial asthma in germany. npj Prim. Care Respir. Medicine 27, https://doi.org/10.1038/s41533-017-0026-x (2017).
Hidalgo, C. A., Blumm, N., Barabási, A.-L. & Christakis, N. A. A dynamic network approach for the study of human phenotypes. PLoS Comput. Biol. 5, e1000353, https://doi.org/10.1371/journal.pcbi.1000353 (2009).
Hu, J. X., Thomas, C. E. & Brunak, S. Network biology concepts in complex disease comorbidities. Nat. Rev. Genet. 17, 615–629, https://doi.org/10.1038/nrg.2016.87 (2016).
Blair, D. R. et al. A nondegenerate code of deleterious variants in mendelian loci contributes to complex disease risk. Cell 155, 70–80, https://doi.org/10.1016/j.cell.2013.08.030 (2013).
Melamed, R. D., Emmett, K. J., Madubata, C., Rzhetsky, A. & Rabadan, R. Genetic similarity between cancers and comorbid mendelian diseases identifies candidate driver genes. Nat. Commun. 6, https://doi.org/10.1038/ncomms8033 (2015).
Tabarés-Seisdedos, R. & Baudot, A. Editorial: Direct and inverse comorbidities between complex disorders. Front. Physiol. 7, https://doi.org/10.3389/fphys.2016.00117 (2016).
Catalá-López, F. et al. Inverse and direct cancer comorbidity in people with central nervous system disorders: A meta-analysis of cancer incidence in 577, 013 participants of 50 observational studies. Psychother. Psychosom. 83, 89–105, https://doi.org/10.1159/000356498 (2014).
Ibáñez, K., Boullosa, C., Tabarés-Seisdedos, R., Baudot, A. & Valencia, A. Molecular evidence for the inverse comorbidity between central nervous system disorders and cancers detected by transcriptomic meta-analyses. PLoS Genet. 10, e1004173, https://doi.org/10.1371/journal.pgen.1004173 (2014).
Samet, J. M. Tobacco smoking. Thorac. Surg. Clin. 23, 103–112, https://doi.org/10.1016/j.thorsurg.2013.01.009 (2013).
Messerli, F. Cardiovascular Effects of Obesity and hypertension. The Lancet 319, 1165–1168, https://doi.org/10.1016/s0140-6736(82)92234-6 (1982).
Scott, H. A. et al. Dietary restriction and exercise improve airway inflammation and clinical outcomes in overweight and obese asthma: a randomized trial. Clin. & Exp. Allergy 43, 36–49, https://doi.org/10.1111/cea.12004 (2012).
Benson, M. K., Berrill, W. T., Cruickshank, J. M. & Sterling, G. S. A comparison of four beta-adrenoceptor antagonists in patients with asthma. Br J Clin Pharmacol 5, 415–419 (1978).
Morales, D. R., Lipworth, B. J., Donnan, P. T., Jackson, C. & Guthrie, B. Respiratory effect of beta-blockers in people with asthma and cardiovascular disease: population-based nested case control study. BMC Medicine 15, https://doi.org/10.1186/s12916-017-0781-0 (2017).
Wong, C. S., Pavord, I. D., Williams, J., Britton, J. R. & Tattersfield, A. E. Bronchodilator, cardiovascular, and hypokalaemic effects of fenoterol, salbutamol, and terbutaline in asthma. Lancet 336, 1396–1399 (1990).
Grossman, E. & Messerli, F. H. Drug-induced hypertension: An unappreciated cause of secondary hypertension. The Am. J. Medicine 125, 14–22, https://doi.org/10.1016/j.amjmed.2011.05.024 (2012).
Rzhetsky, A., Wajngurt, D., Park, N. & Zheng, T. Probing genetic overlap among complex human phenotypes. Proc. Natl. Acad. Sci. 104, 11694–11699, https://doi.org/10.1073/pnas.0704820104 (2007).
Ziyab, A. H. et al. Allergic sensitization and filaggrin variants predispose to the comorbidity of eczema, asthma, and rhinitis: results from the isle of wight birth cohort. Clin. & Exp. Allergy 44, 1170–1178, https://doi.org/10.1111/cea.12321 (2014).
Bagley, S. C., Sirota, M., Chen, R., Butte, A. J. & Altman, R. B. Constraints on biological mechanism from disease comorbidity using electronic medical records and database of genetic variants. Plos Comput. Biol. 12, e1004885, https://doi.org/10.1371/journal.pcbi.1004885 (2016).
Zhou, H. et al. Genetic risk variants associated with comorbid alcohol dependence and major depression. JAMA Psychiatry 74, 1234, https://doi.org/10.1001/jamapsychiatry.2017.3275 (2017).
Park, J., Lee, D.-S., Christakis, N. A. & Barabási, A.-L. The impact of cellular networks on disease comorbidity. Mol. Syst. Biol. 5, https://doi.org/10.1038/msb.2009.16 (2009).
Menche, J. et al. Uncovering disease-disease relationships through the incomplete interactome. Science 347, 1257601–1257601, https://doi.org/10.1126/science.1257601 (2015).
Ko, Y., Cho, M., Lee, J.-S. & Kim, J. Identification of disease comorbidity through hidden molecular mechanisms. Sci. Reports 6, https://doi.org/10.1038/srep39433 (2016).
Rubio-Perez, C. et al. Genetic and functional characterization of disease associations explains comorbidity. Sci. Reports 7, https://doi.org/10.1038/s41598-017-04939-4 (2017).
Lee, D.-S. et al. The implications of human metabolic network topology for disease comorbidity. Proc. Natl. Acad. Sci. 105, 9880–9885, https://doi.org/10.1073/pnas.0802208105 (2008).
Ohn, J. H. The landscape of genetic susceptibility correlations among diseases and traits. J Am Med Inf. Assoc 24, 921–926 (2017).
Bulik-Sullivan, B. et al. An atlas of genetic correlations across human diseases and traits. Nat. Genet. 47, 1236–1241 (2015).
Saik, O. V. et al. Novel candidate genes important for asthma and hypertension comorbidity revealed from associative gene networks. BMC Med. Genomics 11, https://doi.org/10.1186/s12920-018-0331-4 (2018).
Ivanisenko, V. A. et al. ANDSystem: an associative network discovery system for automated literature mining in the field of biology. BMC Syst. Biol. 9, S2, https://doi.org/10.1186/1752-0509-9-s2-s2 (2015).
Ivanisenko, V. A., Demenkov, P. S., Ivanisenko, T. V., Mishchenko, E. L. & Saik, O. V. A new version of the ANDSystem tool for automatic extraction of knowledge from scientific publications with expanded functionality for reconstruction of associative gene networks by considering tissue-specific gene expression. BMC Bioinforma. 20, https://doi.org/10.1186/s12859-018-2567-6 (2019).
Chen, J., Bardes, E. E., Aronow, B. J. & Jegga, A. G. ToppGene suite for gene list enrichment analysis and candidate gene prioritization. Nucleic Acids Res. 37, W305–W311, https://doi.org/10.1093/nar/gkp427 (2009).
Tranchevent, L.-C. et al. Candidate gene prioritization with endeavour. Nucleic Acids Res. 44, W117–W121, https://doi.org/10.1093/nar/gkw365 (2016).
Drevytska, T. et al. shRNA-induced knockdown of a bioinformatically predicted target IL10 influences functional parameters in spontaneously hypertensive rats with asthma. J. Integr. Bioinforma. 15, https://doi.org/10.1515/jib-2018-0053 (2018).
Bragina, E. Y. et al. Molecular relationships between bronchial asthma and hypertension as comorbid diseases. J. Integr. Bioinforma. 15, https://doi.org/10.1515/jib-2018-0052 (2018).
Saik, O. V. et al. Search for new candidate genes involved in the comorbidity of asthma and hypertension based on automatic analysis of scientific literature. J. Integr. Bioinforma. 15, https://doi.org/10.1515/jib-2018-0054 (2018).
Butler, M. G. Genetics of hypertension. Current status. J Med Liban 58, 175–178 (2010).
Wenzel, S. E. Asthma phenotypes: the evolution from clinical to molecular approaches. Nat. Medicine 18, 716–725, https://doi.org/10.1038/nm.2678 (2012).
Carvalho-Silva, D. et al. Open targets platform: new developments and updates two years on. Nucleic Acids Res. 47, D1056–D1065, https://doi.org/10.1093/nar/gky1133 (2018).
Piñero, J. et al. DisGeNET: a comprehensive platform integrating information on human disease-associated genes and variants. Nucleic Acids Res. 45, D833–D839, https://doi.org/10.1093/nar/gkw943 (2016).
Shoshi, A. et al. GenCoNet – a graph database for the analysis of comorbidities by gene networks. J. Integr. Bioinforma. 15, https://doi.org/10.1515/jib-2018-0049 (2018).
Shannon, P. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504, https://doi.org/10.1101/gr.1239303 (2003).
Doncheva, N. T., Morris, J. H., Gorodkin, J. & Jensen, L. J. Cytoscape StringApp: Network analysis and visualization of proteomics data. J. Proteome Res. 18, 623–632, https://doi.org/10.1021/acs.jproteome.8b00702 (2018).
Wang, J. et al. ClusterViz: A cytoscape APP for cluster analysis of biological network. IEEE/ACM Transactions on Comput. Biol. Bioinforma. 12, 815–822, https://doi.org/10.1109/tcbb.2014.2361348 (2015).
Shen, H., Cheng, X., Cai, K. & Hu, M.-B. Detect overlapping and hierarchical community structure in networks. Phys. A: Stat. Mech. its Appl. 388, 1706–1712, https://doi.org/10.1016/j.physa.2008.12.021 (2009).
Barnes, P. J. Cytokine-directed therapies for asthma. J. Allergy Clin. Immunol. 108, S72–S76, https://doi.org/10.1067/mai.2001.116435 (2001).
Abdullah. Elevated th17 and IL-23 in hypertensive patients with acutly increased blood pressure. Am. J. Immunol. 8, 27–32, https://doi.org/10.3844/ajisp.2012.27.32 (2012).
Nakajima, H. & Hirose, K. Role of IL-23 and th17 cells in airway inflammation in asthma. Immune Netw. 10, 1, https://doi.org/10.4110/in.2010.10.1.1 (2010).
Christiansen, S. C. & Zuraw, B. L. Treatment of hypertension in patients with asthma. New Engl. J. Medicine 381, 1046–1057, https://doi.org/10.1056/nejmra1800345 (2019).
Henricks, P. A. & Nijkamp, F. P. Reactive oxygen species as mediators in asthma. Pulm. Pharmacol. & Ther. 14, 409–421, https://doi.org/10.1006/pupt.2001.0319 (2001).
Rodrigo, R. et al. Relationship between oxidative stress and essential hypertension. Hypertens. Res. 30, 1159–1167, https://doi.org/10.1291/hypres.30.1159 (2007).
Taniyama, Y. & Griendling, K. K. Reactive oxygen species in the vasculature. Hypertension 42, 1075–1081, https://doi.org/10.1161/01.hyp.0000100443.09293.4f (2003).
Polonikov, A. V. et al. Antioxidant defense enzyme genes and asthma susceptibility: Gender-specific effects and heterogeneity in gene-gene interactions between pathogenetic variants of the disease. BioMed Res. Int. 2014, 1–17, https://doi.org/10.1155/2014/708903 (2014).
Minelli, C. et al. Glutathione-s-transferase genes and asthma phenotypes: a human genome epidemiology (HuGE) systematic review and meta-analysis including unpublished data. Int. J. Epidemiol. 39, 539–562, https://doi.org/10.1093/ije/dyp337 (2009).
Ghosh, S. et al. Nitrotyrosine proteome survey in asthma identifies oxidative mechanism of catalase inactivation. The J. Immunol. 176, 5587–5597, https://doi.org/10.4049/jimmunol.176.9.5587 (2006).
Bessa, S. S., Ali, E. M. & Hamdy, S. M. The role of glutathione s- transferase m1 and t1 gene polymorphisms and oxidative stress-related parameters in egyptian patients with essential hypertension. Eur. J. Intern. Medicine 20, 625–630, https://doi.org/10.1016/j.ejim.2009.06.003 (2009).
Zhou, X. F. et al. Polymorphisms in the promoter region of catalase gene and essential hypertension. Dis. Markers 21, 3–7, https://doi.org/10.1155/2005/487014 (2005).
Erne, P., Resink, T. J., Burgisser, E. & Buhler, F. R. Platelets and hypertension. J. Cardiovasc. Pharmacol. 7(Suppl 6), S103–108 (1985).
Kornerup, K. N. & Page, C. P. The role of platelets in the pathophysiology of asthma. Platelets 18, 319–328, https://doi.org/10.1080/09537100701230436 (2007).
Mitchell, C., Provost, K., Niu, N., Homer, R. & Cohn, L. IFN- acts on the airway epithelium to inhibit local and systemic pathology in allergic airway disease. The J. Immunol. 187, 3815–3820, https://doi.org/10.4049/jimmunol.1100436 (2011).
Leibowitz, A. & Schiffrin, E. L. Immune mechanisms in hypertension. Curr. Hypertens. Reports 13, 465–472, https://doi.org/10.1007/s11906-011-0224-9 (2011).
Kidd, C. D. A., Thompson, P. J., Barrett, L. & Baltic, S. Histone modifications and asthma. the interface of the epigenetic and genetic landscapes. Am. J. Respir. Cell Mol. Biol. 54, 3–12, https://doi.org/10.1165/rcmb.2015-0050tr (2016).
Cardinale, J. P. et al. HDAC inhibition attenuates inflammatory, hypertrophic, and hypertensive responses in spontaneously hypertensive rats. Hypertension 56, 437–444, https://doi.org/10.1161/hypertensionaha.110.154567 (2010).
Bhavsar, P., Ahmad, T. & Adcock, I. M. The role of histone deacetylases in asthma and allergic diseases. J. Allergy Clin. Immunol. 121, 580–584, https://doi.org/10.1016/j.jaci.2007.12.1156 (2008).
Lee, H.-A. et al. Histone deacetylase inhibition attenuates transcriptional activity of mineralocorticoid receptor through its acetylation and prevents development of hypertension. Circ. Res. 112, 1004–1012, https://doi.org/10.1161/circresaha.113.301071 (2013).
Yu, W., Gwinn, M., Clyne, M., Yesupriya, A. & Khoury, M. J. A navigator for human genome epidemiology. Nat. Genet. 40, 124–125, https://doi.org/10.1038/ng0208-124 (2008).
Huan, T. et al. A meta-analysis of gene expression signatures of blood pressure and hypertension. PLOS Genet. 11, e1005035, https://doi.org/10.1371/journal.pgen.1005035 (2015).
Li, X. et al. eQTL of bronchial epithelial cells and bronchial alveolar lavage deciphers GWAS-identified asthma genes. Allergy 70, 1309–1318, https://doi.org/10.1111/all.12683 (2015).
Parente, L. & Solito, E. Annexin 1: more than an anti-phospholipase protein. Inflamm. Res. 53, 125–132, https://doi.org/10.1007/s00011-003-1235-z (2004).
Alangari, A. Corticosteroids in the treatment of acute asthma. Annals Thorac. Medicine 9, 187, https://doi.org/10.4103/1817-1737.140120 (2014).
Bertagna, X., Guignat, L., Groussin, L. & Bertherat, J. Cushing’s disease. Best Pract. & Res. Clin. Endocrinol. & Metab. 23, 607–623, https://doi.org/10.1016/j.beem.2009.06.001 (2009).
Kowalski, M. L. et al. Classification and practical approach to the diagnosis and management of hypersensitivity to nonsteroidal anti-inflammatory drugs. Allergy 68, 1219–1232, https://doi.org/10.1111/all.12260 (2013).
Szczeklik, A. et al. Safety of a specific COX-2 inhibitor in aspirin-induced asthma. Clin. Exp. Allergy 31, 219–225, https://doi.org/10.1046/j.1365-2222.2001.01075.x (2001).
Solomon, D. H., Schneeweiss, S., Levin, R. & Avorn, J. Relationship between COX-2 specific inhibitors and hypertension. Hypertension 44, 140–145, https://doi.org/10.1161/01.hyp.0000136134.31846.83 (2004).
Chan, C. C. et al. Do COX-2 inhibitors raise blood pressure more than nonselective NSAIDs and placebo? an updated metaanalysis. J. Hypertens. 27, 2332–2341, https://doi.org/10.1097/hjh.0b013e3283310dc9 (2009).
Gualtierotti, R. et al. Effect of naproxen and acetaminophen on blood pressure lowering by ramipril, valsartan and aliskiren in hypertensive patients. Expert. Opin. on Pharmacother. 14, 1875–1884, https://doi.org/10.1517/14656566.2013.816286 (2013).
Li, H. et al. Novel disease syndromes unveiled by integrative multiscale network analysis of diseases sharing molecular effectors and comorbidities. BMC Med. Genomics 11, https://doi.org/10.1186/s12920-018-0428-9 (2018).
Rappaport, N. et al. MalaCards: an amalgamated human disease compendium with diverse clinical and genetic annotation and structured search. Nucleic Acids Res. 45, D877–D887, https://doi.org/10.1093/nar/gkw1012 (2016).
Cookson, W. O. & Moffatt, M. F. Genetics of asthma and allergic disease. Hum. Mol. Genet. 9, 2359–2364 (2000).
Köhler, S. et al. Expansion of the human phenotype ontology (HPO) knowledge base and resources. Nucleic Acids Res. 47, D1018–D1027, https://doi.org/10.1093/nar/gky1105 (2018).
Buniello, A. et al. The NHGRI-EBI GWAS catalog of published genome-wide association studies, targeted arrays and summary statistics 2019. Nucleic Acids Res. 47, D1005–D1012, https://doi.org/10.1093/nar/gky1120 (2018).
Joehanes, R. et al. Integrated genome-wide analysis of expression quantitative trait loci aids interpretation of genomic association studies. Genome Biol. 18, https://doi.org/10.1186/s13059-016-1142-6 (2017).
Westra, H.-J. et al. Systematic identification of trans eQTLs as putative drivers of known disease associations. Nat. Genet. 45, 1238–1243, https://doi.org/10.1038/ng.2756 (2013).
Jansen, R. et al. Conditional eQTL analysis reveals allelic heterogeneity of gene expression. Hum. Mol. Genet. 26, 1444–1451, https://doi.org/10.1093/hmg/ddx043 (2017).
Ardlie, K. G. et al. The genotype-tissue expression (GTEx) pilot analysis: Multitissue gene regulation in humans. Science 348, 648–660, https://doi.org/10.1126/science.1262110 (2015).
Sherry, S. T. et al. dbSNP: the NCBI database of genetic variation. Nucleic Acids Res. 29, 308–311 (2001).
Bigler, J. et al. A severe asthma disease signature from gene expression profiling of peripheral blood from u-BIOPRED cohorts. Am. J. Respir. Critical Care Medicine 195, 1311–1320, https://doi.org/10.1164/rccm.201604-0866oc (2017).
Wishart, D. S. et al. DrugBank 5.0: a major update to the DrugBank database for 2018. Nucleic Acids Res. 46, D1074–D1082, https://doi.org/10.1093/nar/gkx1037 (2017).
Carter, J. S. et al. Initializing the VA medication reference terminology using UMLS metathesaurus co-occurrences. Proc AMIA Symp 116–120 (2002).
Nicklas, R. A. Paradoxical bronchospasm associated with the use of inhaled beta agonists. J. Allergy Clin. Immunol. 85, 959–964, https://doi.org/10.1016/0091-6749(90)90084-h (1990).
Chen, Y.-A., Tripathi, L. P. & Mizuguchi, K. TargetMine, an integrated data warehouse for candidate gene prioritisation and target discovery. PLoS One 6, e17844, https://doi.org/10.1371/journal.pone.0017844 (2011).
Gilmour, P. S., Schladweiler, M. C., Richards, J. H., Ledbetter, A. D. & Kodavanti, U. P. Hypertensive rats are susceptible to TLR4-mediated signaling following exposure to combustion source particulate matter. Inhalation Toxicol. 16, 5–18, https://doi.org/10.1080/08958370490442827 (2004).
Wettschureck, N. & Offermanns, S. Rho/rho-kinase mediated signaling in physiology and pathophysiology. J. Mol. Medicine 80, 629–638, https://doi.org/10.1007/s00109-002-0370-2 (2002).
Calò, L. A. & Pessina, A. C. RhoA/rho-kinase pathway: much more than just a modulation of vascular tone. evidence from studies in humans. J. Hypertens. 25, 259–264, https://doi.org/10.1097/hjh.0b013e328010d4d2 (2007).
Siehler, S. Regulation of RhoGEF proteins by g12/13-coupled receptors. Br. J. Pharmacol. 158, 41–49, https://doi.org/10.1111/j.1476-5381.2009.00121.x (2009).
Madeo, A. C., Manichaikul, A., Pryor, S. P. & Griffith, A. J. Do mutations of the pendred syndrome gene, SLC26a4, confer resistance to asthma and hypertension? J. Med. Genet. 46, 405–406, https://doi.org/10.1136/jmg.2008.063610 (2009).
Pacanowski, M. et al. β-adrenergic receptor gene polymorphisms and β-blocker treatment outcomes in hypertension. Clin. Pharmacol. & Ther. 84, 715–721, https://doi.org/10.1038/clpt.2008.139 (2008).
Leineweber, K. & Heusch, G. β1- and β2-adrenoceptor polymorphisms and cardiovascular diseases. Br. J. Pharmacol. 158, 61–69, https://doi.org/10.1111/j.1476-5381.2009.00187.x (2009).
Sharma, A. et al. A disease module in the interactome explains disease heterogeneity, drug response and captures novel pathways and genes in asthma. Hum. Mol. Genet. 24, 3005–3020 (2015).
Acknowledgements
We would like to thank Veronika Gurianova and Anna Brattseva for their kind support. This study was supported by the International DFG Research Training Group GRK 1906 and the Volkswagen Stiftung Trilateral Partnerships - Cooperation Projects between Scholars and Scientists from Ukraine, Russia, and Germany In silico screening and experimental validation of new drug targets for the treatment of comorbid multifactorial diseases (#90335). We acknowledge support for the Article Processing Charge by the Open Access Publication Fund of Bielefeld University.
Author information
Authors and Affiliations
Contributions
O.Z., O.V.S., E.Yu.B., I.G. and C. K. extracted the data, O.Z. analyzed the data and interpreted the results. V.A.I., V.E.D., M.B.F. and R.H. supervised the project. All authors provided critical feedback and contributed to the writing of the manuscript and reviewed the final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zolotareva, O., Saik, O.V., Königs, C. et al. Comorbidity of asthma and hypertension may be mediated by shared genetic dysregulation and drug side effects. Sci Rep 9, 16302 (2019). https://doi.org/10.1038/s41598-019-52762-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-52762-w
This article is cited by
-
Risk factor analysis and nomogram for predicting poor symptom control in smoking asthmatics
BMC Pulmonary Medicine (2024)
-
Efficacy observation of pushing Qiaogong point with different intensities for treating stage I essential hypertension
Journal of Acupuncture and Tuina Science (2023)
-
Case–control study exploring the short-term association of bronchiolitis with high blood pressure and hypertension in hospitalized children
Clinical Hypertension (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.