Abstract
The hemp plant (Cannabis sativa L.) has a long tradition of being used for many different purposes such as industry, medicine and nutrition. In particular, because hemp seed (HS) is rich in oil protein and considerable amounts of dietary fiber, vitamins and minerals that are particularly suitable also for animal nutrition. Different studies have evaluated HS on qualitative and quantitative properties of livestock products but as of today, nobody has investigated the molecular pathway behind HS supplementation in farm animals. Thus, in this study, we will report the first RNA sequencing of the whole-blood transcriptome of ewes fed either with a controlled diet (CTR, n = 5) or with a diet supplemented with 5% of hemp seed (HSG, n = 5). Applying a false discovery rate (FDR) <0.05 and a log2FC either higher than 0.5 or lower than −0.5, we identified 314 differentially regulated genes in the HS-supplemented group compared to the CTR group. Several genes encoding for different subunits belonging to the complex I, II, III, IV and ATP-synthase were up-regulated making oxidative phosphorylation (FDR: 3.05e-19) and thermogenesis (FDR: 2.17e-16) the highest up-regulated pathways in our study. Moreover, we found up-regulation in different genes involved in lactose biosyntheses such as GALK1 and PGM1 and, as a result, we observed a statistically higher lactose percentage in the HSG group (p < 0.05). These results indicate that HS supplementation positively affects the energy production pathway in lactating ewes conferring them also more resistance to adverse climatic conditions such as low temperature. Finally, the higher milk lactose content makes the derived dairy products more profitable.
Similar content being viewed by others
Introduction
Recently in the field of animal nutrition, people have paid more attention to meal formulations that meet both the consumers’ needs and ameliorated animals’ health. Thus, it is becoming common that animal diets are often supplemented either with micronutrients1,2,3 but also with by-products of agriculture4,5. Moreover, there is also more interest in the use of plants rich in secondary compounds. Of these, Cannabis sativa L. (hemp) deserves special attention because it has been used for centuries for different purposes such as a source of fiber but also as a medicinal plant thanks to the cannabinoids contained in the leaves and flowers6. In the European Union (EU), the cultivation of hemp is permitted only for the varieties where the maximum content of tetrahydrocannabinol (THC), the main psychoactive substance, is limited to 0.2% (w/w) as dry matter7. Hemp seed (HS) is generally used for animal nutrition and is considered THC free; however, some studies have reported traces of THC present in the seed sample probably because it was contaminated with plant debris6.
HS is rich in oil and proteins and provides a considerable amount of dietary fiber, vitamins and minerals6; moreover, the by-product following oil extraction called HS cake can be used as a more economical source of protein8. Studies about the use of HS for animal nutrition have mainly focused on the improvement of the quantity and quality of production. In feedlot cattle, different amounts of HS included in the diet did not show any negative effects on either the matter intake or rate or efficiency of gaining weight; however, positive effects on beef tissue quality were recorded because the content of conjugated linoleic acids (CLAs) was relatively higher but, at the same time, saturated fatty acids (SFAs) also increased9. According to this study, also in a different trial, HS cake supplementation in comparison to soybean meal favorably affected FAs composition by boosting the ratio of polyunsaturated on monounsaturated fatty acid (PUFA/MUFA)s in both fresh and cooked meat from bovine M. longissimus dorsi10. HS cake was also evaluated as a source of crude protein in ewes compared to peas and rapeseed cake. While the crude protein amount was similar between all supplements, Karlsoon et al.11 have shown the metabolized energy produced by HS cake was lower in comparison to the other two supplements probably because the nondegradable rumen component negatively affects intestinal digestibility. Thus, while the HS cake showed controversial results as a protein food source, a HS supplemented diet favorably effects the fatty FA composition in lactating goats, increasing the quality of milk from a human consumer’s point of view12. Indeed, HS feeding also showed an increased amount of fat, shifting the composition in favor of PUFAs and in contrast to the lipids that are considered related to hypercholesterolaemic diseases13. In agreement with these studies, also using HS oil as a dietary supplement, it was possible to modify the goat milk’s fatty acid quality with a higher ratio of PUFAs on SFAs without affecting liver function14. HS supplementation has been used also in broiler and laying hens for assessing affects on meat and egg quality respectively15,16. Importantly, in broilers, HS did not affect negatively growth performance but positively influenced the serum lipid profile showing a decrease of triglyceride, low-density lipoproteins (LDL) and total cholesterol. Moreover, it also improved gut health as a consequence of reducing coliform bacteria associated with HS supplementation15. In laying hens, both raw and heat-treated HS improved the egg quality, egg FA profile and hen performance values; moreover, the heat treatment modified the flavour of HS restoring the feed consumption at the control group level16.
This data encourages the use of hemp plants and its role as a supplement in animal diets. Despite the positive effects on the quality of products, transcriptomic data behind the HS supplementation is lacking. Thus, in this study using a high-throughput RNA-sequencing approach, we elucidate the molecular mechanisms, signaling pathways and metabolic processes behind the hemp seed supplementation in sheep.
Results
Influence of HS supplemented diet on whole blood cell count and milk composition
To evaluate the overall health of the lactating ewes fed with a 5% HS supplemented diet, the complete whole blood count was evaluated in the control (CTR, n = 10) and HS group (HSG, n = 10) at the end of the supplementation period (day 30). As shown in Fig. 1, no differences were detected either in total white blood cell counts or in the cell type sub-set (neutrophils, lymphocytes, eosinphils and monocytes), indicating that HS does not have a detrimental effect on either innate and adaptive immune system. We assessed also the influence of HS on milk parameters; while total amount of proteins (including caseins) and lipids were not hanged between CTR and HSG groups, lactose showed a statistically significant increase in HSG (4.69 ± 0.04 vs 4.84 ± 0.03). Finally, the somatic cell counts were measured as a common parameter used for diagnosis of infectious disease and no differences were found between the two groups indicating that HS is not relevant to the mammary gland health status (Fig. 2).
Effect of hemp seed HS-supplemented diet on complete whole blood formula. (A) white blood cell, (B) neutrophils, (C) lymphocyte, (D) eosinophil and (E) monocyte were measured in CTR and HSG groups at the end of supplementation period. Each point represents a single subject, and data are expressed as percentage. Any possible differences were analyzed by using the Student’s t-test. CTR: a control group.
Effect of hemp seed HS-supplemented diet on milk composition. (A) caseins, (B) proteins, (C) lactose, (D) lipids and (E) somatic cells were quantified in CTR and HSG groups at the end of supplementation period. Each point represents a single subject, and data are expressed as percentage for proteins, caseins, lactose and lipids while somatic cells are expressed as absolute number. Any possible differences were analyzed by using the Student’s t-test and a p-value <0.05 is considered significant. *p-value <0.05. CTR: a control group.
Influence of HS supplemented diet on blood transcriptome
In order to identify the transcriptomic signature in lactating ewes fed with HS supplemented diet, we sequenced the RNA isolated from whole blood collected from animals of both groups (CTR, n = 5; HSG, n = 5). Filtering the expression by false discovery rate (FDR) <0.05 combined with a log2 fold change (log2FC) either higher than 0.5 or lower than −0.5, we identified314 differentially expressed genes (DEGs) (Supplementary Table 1), of which 270 and 44 DEGs were up- and down-regulated, respectively (Supplementary Fig. 1). To identify molecular pathways associated with our data set, we interrogated STRING using the highest available interaction score (0.9) in order to increase stringency and avoid possible uncorrected data interpretation. Up- and down-regulated genes were analyzed separately and no enriched pathways were associated with the down-regulated genes (data not shown). Analyzing the up-regulated genes, in Table 1, we show all the statistically significant enriched pathways; in Supplementary Fig. 2, we show graphically all possible interactions between the up-regulated DEGs and it is possible to appreciate that they mainly clustered in three groups (I, II, and III). The first group includes genes that are shared between oxidative phosphorylation (FDR: 3.05e-19) and thermogenesis (FDR 2.17e-16) pathways; the second group involves gene related to sugar metabolism like Glycolysis/Gluconeogenesis (3.92e-08) and Galactose metabolism (5.72e-06) and more generally to carbon metabolism (5.72e-06); finally, the third group includes genes involved in the protein synthesis and more specifically genes related to the ribosome and protein translation (8.85e-07). Then, we looked for enriched functional GO terms represented in our data set: the significantly enriched biological processes (BP) included genes involved generation of precursor metabolites and energy production (FDR: 3.03e-05), nitrogen compound metabolic process, purine ribonucleoside monophosphate metabolic process, electron transport chain, ATP metabolic process (all significant BP are reported in Supplementary Table 2). In addition, oxidoreductase activity, catalytic activity and NADH dehydrogenase (ubiquinone) activity were the top three enriched molecular functions (MF) GO terms (Supplementary Table 3).
Co-expression network analysis
Because HS supplementation influences milk lactose probably influencing pathways related to sugar metabolism –mainly Glycolysis/Gluconeogenesis and galactose metabolism- we aimed to identify a consensus network connected with the trait of interest. Thus, using the lactose amount of each of the ten samples as trait for the weighted gene-correlation network analysis (WGCNA), 47 different modules were identified (Fig. 3A). Each module had a different number of genes (Supplementary Table 4), ranging from a minimum of 34 (module brown4) to a maximum of 9.591 (module grey). This latter contained all genes with low expression values, and it was discarded from further analysis. All modules were positively or negatively correlated with lactose, with a correlation value ranging from −0.82 (module lightcyan) to 0.64 (module lightgreen) (Fig. 3B and Supplementary Table 4). Among them, two modules (lightgreen and blue), which include 177 and 2467 genes respectively, showed a correlation coefficient higher than 0.45 and a p-value lower than 0.05 (Supplementary Tables 5 and 6). For both modules, we focused on the genes with the highest intramodular connectivity (hub genes, MM >0.7), as these genes may represent points of biological interest in defining our phenotypes. Among those belonging to the lightgreen module, we identified PGM1 (MM = 0,81; p.value = 0.004), which is strongly associated with galactose metabolism. Furthermore, applying the same criteria to the other significantly associated module (blue), we noticed that GALK1 showed a high MM value (MM = 0.99, p.value = 3.25e-08) and it is also involved in lactose metabolism. Interestingly, about 40% (125 of 316) of our DEGs are represented in the blue module confirming the strengths of our experimental analysis.
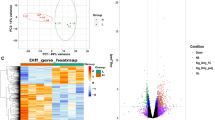
Composite visualization of co-expression analysis. (A) Gene dendrogram carried out by average linkage hierarchical clustering. The color row represents the module assignment determined by the Dynamic Tree Cut; (B) heatmap plot of the adjacencies in the eigengene network including the trait lactose. Each row and column in the heatmap corresponds to one module eigengene (labeled by color) or trait. In the heatmap, blue color represents low adjacency (negative correlation), while red represents high adjacency (positive correlation). Squares of red color along the diagonal are the meta-modules; (C) determination of soft-threshold power in WGCNA. The histogram of connectivity distribution and the scale-free topology are shown in the first two panels, respectively. The scale-free topology index and the mean connectivity for each power value between 1 and 20 are shown in scale independence and mean connectivity panels, respectively; (D) a scatterplot of gene significance for weight (GS) versus module membership (MM) in the blue and lighgreen modules. GS and MM exhibit a very significant correlation, implying that hub genes of the brown module also tend to be highly correlated with lactose. Eigene values from both modules (significantly associated with lactose amount) across the ten samples(CTR and HS) are also reported.
As a further confirmation that both light green and blue modules are related to lactose differences between HS and CTR groups, we calculated their eigengene values. As expected, the histogram showed an opposite pattern between the two experimental groups in both modules (Fig. 3D).
Discussion
The last years have seen an increasing interest in livestock diet supplementation with different ingredients for improving the quality of products and to ameliorate the health status of animals. In this scenario, the use of hemp seed is gaining interest because it is characterized by a high content of oil (26–37.5%), protein (25%), and fiber (28%) with a digestibility of about 20%6. Specifically, oil is constituted mostly by PUFA with an approximate ratio of 3:1 between ω-6 and ω-3 fatty acids, which is considered the best condition for human health17. Indeed, ω-3 FAs are being widely investigated and a large body of evidence indicates their prominent role as protective lipid mediators in the context of inflammatory, metabolic, neurodegenerative, and neoplastic diseases18,19. However, the knowledge around the use of HS in animal nutrition is still limited to tolerability and the effects on production traits8,9,10,11,12,13,14,15,16. Thus, using a RNA-seq approach, which represents one of the most relevant techniques for obtaining a comprehensive analysis of gene expression changes and molecular processes influenced by the diet components, we investigated the effects of an HS-supplemented diet on the whole blood transcriptome of lactating sheep. Whole blood represents a quick and accessible source of biological material. Most importantly, gene expression analysis from this tissue is being successfully used to validate metabolic status from our and different groups1,2,4,5,20. However, before assessing the effects of hempseed-supplementation on gene expression, we evaluated the complete whole blood formula in both groups—CTR and HSG—to evaluate the potential effects of HS on cellular subpopulation but did not find any difference (Fig. 1).
HS supplementation influences the expression of 314 genes. About 271 genes were up-regulated, and when we looked for the enriched biological pathways associated to them, the highest one was “oxidative phosphorylation (oas00190)” (Table 1). This upregulation was as result of the genes that are related to all four complexes and ATP-synthase of the electron transport chain (Supplementary Fig. 1). In detail, ND4, ND5, ND6, NDUFS7, NDUFS8, and NDUFV1 are subunits that belong to the core of NADH dehydrogenase, and constitute the minimal assembly required for the transfer of electrons from NADH to ubiquinone before they move to the respiratory chain. We also found up-regulation of NDUFA3, NDUFA7, NDUFA8, NDUFA11, and NDUFA13 that belong to subunit A and NDUFB2 and NDUFB7 that belong to subunit B; both subunits have accessory functions in the NADH dehydrogenase complex and are not involved in catalytic activity. SDHA, the major catalytic subunit of succinate-ubiquinone oxidoreductase, was also up-regulated, which further reinforced the effect of HS supplementation on complex I and II of the electron transport chain. With regards to complex III, we found up-regulated CYC1, CYTB, UQCR10, and UQCRC1, while COX1, COX2, and COX4I1 were found on complex IV and were also up-regulated. Finally, for the ATP-synthase complex, we found up-regulated ATP5D, ATP5I, and ATP621. However, the protonic gradient generated by electron chain transport is often associated with thermogenesis, which in our data set, is the second most up-regulated pathway. Perinatal mortality caused by hypothermia is one of major concerns within the sheep industry. Thus, the possibility to increase cold tolerance with HS supplementation is desirable. Indeed, it has been previously shown that the high intake of ω-3 fatty acids, together with a ω-6 increase in rectal temperature of newborn lambs can improve cold tolerance22. However, in a different study, ewes fed with a docosahexaenoic acid supplement diet at late gestation and early lactation do not ameliorate thermogenesis indexes in lambs but have a potential positive effect on the immune system due to a higher level of immunoglobulins in the colostrum23. Thus, because in our study we show a general upregulation of genes involved in the thermogenesis activation, we could speculate that HS supplementation confers a better adaptation to cold stress.
Glycolysis and galactose metabolism pathways are also overexpressed following HS diet Supplementation (Table 1). Specifically, we found upregulation in GALK1 that catalyze the phosphorylation of galactose, which is one of the steps involved in the production of lactose. Moreover, PGM1 that together UGP2 catalyze the conversion of glucose to UDP-galactose is upregulated. Because it has been demonstrated that expression of PGM1 is strongly correlated with the amount of lactose in milk24, in agreement, we found a higher content of lactose in animals that were fed hemp seed. Moreover, we performed a co-expression analysis that was successfully used in several studies, including those for candidate biomarkers about boar taint in non-castrated pigs25 and identification of molecular signatures related to feed effiency26. When applying the WGCNA analysis to our dataset, we found that GALK1 and PGM1 belong in the blue and lightgreen cluster, which represent the only two clusters that are statistically correlated to the lactose level. Moreover, several genes that are differentially expressed between our experimental groups belong to those two modules, further supporting our findings.
In conclusion, in this study, we have assessed the transcriptome signature induced by 5% hemp seed supplemented diet in ewes. The findings suggest that pathways related to energy production were the most affected. In addition, we found that this condition could also be potentially beneficial for adaptation to low temperatures. Moreover, we found a higher content of lactose, which makes the derived dairy products more profitable.
Materials and Methods
Experimental design and diets
In this study none of the animals was sacrificed. All the experiments were planned and conducted according to Directive 2010/63/EU of the European Parliament (European Union, 2010) and Directive 86/609/EEC (European Economic Community, 1986), which deal with the protection of animals used for scientific purposes.
Thirty two half-bred ewes, homogeneous for age (3–5 years), for days in milking (74 ± 4 days) and for weight (59 ± 4 kg) have been randomly divided into two groups of sixteen ewes each: a control group (CTR) and an experimental group (HSG) whose diet was supplemented with the 5% hemp seeds. To verify eventual variations among the selected groups before the trial, individual milk samples were collected to obtain information about milk yield, chemical-nutritional composition and fatty acid profile. The ewes were housed for the entire trial period in two separate areas of free housing with an access to an identical feeding area with individual pens (0.5 × 1.2 × 1 m) and free access to clean drinking water. The study was conducted for a period of 30 days, in which all animals received total mixed rations (TMR) whose composition, reported in Table 2, was defined taking into account the parameters reported on the Nutrient Requirements of Small Ruminants (Board on Agriculture, Division on Earth, & Life Studies; 2007). Samples of TMR were analyzed, according to AOAC methods (1990)27, for crude protein (CD; method 930.15) and ether extract (EE; method 920.39); detergent procedures reported by Van Soest et al.28 were used for the determination of neutral detergent fiber (NDF) and acid detergent fiber (ADF).
Blood and milk sampling
Milk, and whole blood (WB) samples were collected from each group at the beginning (T0) and after 30 days (T30) of the supplementation time. Although in the study we have used a larger number of animals, for the RNA-Seq analysis, 2.5 mL of jugular venous blood were collected from 10 animals at T30 (5 from the control group and 5 from HSG). The selection was performed with the aim to have all ewes with the same age (3 years) in order to avoid potential variations induced by this parameter. Duplicate WB samples from each individual animal were collected in PAXgeneTM tubes (Qiagen SpA, Milan, MI, Italy), stored at room temperature overnight and then at −20 °C until RNA isolation, as per the manufacturer’s instructions.
Blood analysis
Complete blood cell count with leukocyte formula (total white blood cells, monocyte, lymphocyte, basophils, neutrophils, and eosinophils) for both CTR (n = 10) and HSG (n = 10) groups at T0 and T30 were performed at the Veterinary and Public Health Institute (Teramo, Italy) using a laser-based hematology analyzer with software applications for animal species (ADVIA 120 hematology system, Siemens, Munich, Germany) and following the routine procedure of the institute (Veterinary and Public Health Institute “G. Caporale”, Teramo, Italy).
Chemical analysis of milk
Chemical composition of milk (fat, protein, casein, and lactose) was determined by MilkoScan FT 6000 (Foss Integrator IMT; Foss, Hillerød, Denmark), whereas the somatic cells count (SCC) was performed using the Fossomatic FC (Foss).
Library preparation and RNA-Seq analysis
Next generation sequencing experiments, comprising RNA extraction and bioinformatics analysis, were performed by an external company (Genomix4life SRL, Baronissi, Salerno, Italy). Total RNA was extracted using TRIzol and following manufacturer’s instruction (Invitrogen, Carlsbad, CA, USA). Indexed libraries were prepared from 300 ng/ea purified RNA with TruSeq Stranded mRNA Sample Prep Kit (Illumina, San Diego, CA, USA) according to the manufacturer’s instructions. Libraries were quantified using the TapeStation 4200 (Agilent Technologies, Santa Clara, CA, USA) and pooled such that each index-tagged sample was present in equimolar amounts, with final concentration of the pooled samples of 2 nM. The pooled samples were subject to cluster generation and sequencing using an Illumina NextSeq. 500 System (Illumina, San Diego, CA, USA) in a 2 × 75 paired end format at a final concentration of 1.8 pmol.
The raw sequence files generated (.fastq files) underwent quality control analysis using FastQC tool Version 0.11.8 using default settings (http://www.bioinformatics.babraham.ac.uk/projects/fastqc/). To remove the adapter sequences, cutadapt (version 1.4) was used29. The mapping of paired-end reads was performed using STAR (version 2.5)30 on reference genome Oar_v3.1 (GCA_000298735.1) from Ensembl (http://www.ensembl.org/Ovis_aries/Info/Index). The quantification of transcripts expressed for each replicate of the sequenced samples was performed using HTSeq-Count algorithm DESeq. 231 was used to perform the differential expression analysis. Genes with a false discovery rate (FDR) less than 0.05 were considered as differentially expressed genes (DEGs). Raw data associated with this project are deposited in the GenBank’s Sequence Read Archive (SRA) under the accession Bioproject number PRJNA528905.
Enriched pathway analysis
The STRING software (v 11 http://string-db.org/) was used to identify canonical pathways using the dataset of 314 DEGs identified between HSG group and the CTR one with a false discovery rate (FDR) <0.05. We set the interaction score as 0.9, the highest value permitted by the software to avoid false positive. The significance of the canonical pathway was measured with the p-value and the ratio of DEG/number of genes in the pathway
Unsigned weighted correlation networks analysis (WGCNA)
We used the unsigned WGCNA32,33 to perform the hierarchical clustering and identify co-expressed genes (“hub genes”), which may have main regulatory functions and/or major impact on our candidate genes. In detail, an adjacency matrix was first created between pairs of all expression values, before to be replaced with the weighted adjacency matrix obtained by raising the correlations to the power of 7 (Fig. 3C). The weighted adjacency matrix was then transformed into a topological overlap matrix (TOM), which allow the calculation of dissimilarity values used to minimize effects of spurious associations. The result was used as input for the linkage hierarchical clustering and the modules (clusters of highly interconnected genes) were identified in the resulting dendrogram through a dynamic hybrid tree cutting algorithm (DynamicTreeCut algorithm).
Using the module eigengene values, we finally estimated the relationships between each module and the lactose amount of each one of the ten samples used in the RNAseq experiment by calculating the Pearson’s correlation values. Modules were filtered out using a correlation value >0.45 and p.value <0.05. Interesting modules were in deep investigated.
Statistics
GraphPad Prism (GraphPad v.6 Software, La Jolla, CA, USA) was used for statistical analysis. Differences in cell subsets from peripheral blood and plasma samples were assessed using Student’s t-test.
References
Elgendy, R. et al. Transcriptomic signature of high dietary organic selenium supplementation in sheep: A nutrigenomic insight using a custom microarray platform and gene set enrichment analysis. J. Anim. Sci. 94, 3169–3184 (2016).
Elgendy, R. et al. Transcriptome profiling and functional analysis of sheep fed with high zinc-supplemented diet: A nutrigenomic approach. Anim. Feed Sci. Technol. 234, 195–204 (2017).
Ianni, A. et al. Zinc supplementation of Friesian cows: Effect on chemical-nutritional composition and aromatic profile of dairy products. J. Dairy Sci. 102, 2918–2927 (2019).
Iannaccone, M. et al. RNA Sequencing-Based Whole-Transcriptome Analysis of Friesian Cattle Fed with Grape Pomace-Supplemented Diet. Animals 8, 188, https://doi.org/10.3390/ani8110188 (2018).
Iannaccone, M. et al. Whole Blood Transcriptome Analysis Reveals Positive Effects of Dried Olive Pomace-Supplemented Diet on Inflammation and Cholesterol in Laying Hens. Animals 9, 427, https://doi.org/10.3390/ani9070427 (2019).
Callaway, J. C. Hempseed as a nutritional resource: An overview. Euphytica 140, 65–72 (2004).
European Food Safety Authority. Scientific Opinion on the safety of hemp (Cannabis genus) for use as animal. EFSA 9, 1–41 (2011).
Hessle, A., Eriksson, M., Nadeau, E., Turner, T. & Johansson, B. Cold-pressed hempseed cake as a protein feed for growing cattle. Acta Agriculturae Scand Section A 58, 136–145 (2008).
Gibb, D. J., Shah, M. A., Mir, P. S. & Mcallister, T. A. Effect of full-fat hemp seed on performance and tissue fatty acids of feedlot cattle. Can. J. Anim. Sci. 85, 223–230 (2005).
Turner, T., Hessle, A., Lundström, K. & Pickova, J. Influence of hempseed cake and soybean meal on lipid fractions in bovine M. longissimus dorsi. Acta Agriculturae Scand Section A 58, 152–160 (2008).
Karlsson, L., Finell, M. & Martinsson, K. Effects of increasing amounts of hempseed cake in the diet of dairy cows on the production and composition of milk. Animal 4, 1854–1860 (2010).
Mierlita, D. Fatty acid profile and health lipid indices in the raw milk of ewes grazing part-time and hemp seed supplementation of lactating ewes. South African Journal of Animal Sciences 46, 237–246 (2016).
Pilarczyk, R., Wójcik, J., Sablik, P. & Czerniak, P. Fatty acid profile and health lipid indices in the raw milk of Simmental and Holstein-Friesian cows from an organic farm. South African Journal of Animal Sciences 45, 30–38 (2015).
Cozma, A. et al. Effect of hemp seed oil supplementation on plasma lipid profile, liver function, milk fatty acid, cholesterol, and vitamin A concentrations in Carpathian goats. Czech Journal of Animal Science 60, 289–301 (2015).
Vispute, M. et al. Effect of dietary supplementation of hemp (Cannabis sativa) and dill seed (Anethum graveolens) on performance, serum biochemicals and gut health of broiler chickens. J. Anim. Physiol. Anim. Nutr. 103, 525–533 (2019).
Konca, Y. et al. Effects of heat-treated hempseed supplementation on performance, egg quality, sensory evaluation and antioxidant activity of laying hens. British Poultry Science 60, 39–46 (2019).
WHO 2003. Diet, Nutrition and the Prevention of Report of a Joint WHO/FAO Expert Consultation. 1–149 (2003).
Calviello, G. et al. Experimental Evidence of ω -3 Polyunsaturated Fatty Acid Modulation of Inflammatory Cytokines and Bioactive Lipid Mediators: Their Potential Role in Inflammatory, Neurodegenerative, and Neoplastic Diseases. BioMed Reserch International 743171, https://doi.org/10.1155/2013/743171 (2013).
Calder, P. C. Conference on ‘ The future of animal products in the human diet: health and environmental concerns’ Plenary Lecture 3 n -3 PUFA and health: fact, fi ction and the future Very long-chain n -3 fatty acids and human health: fact, fiction and the future. Proceedings of the Nutrition Society 77, 52–72 (2019).
Liew, C., Ma, J., Tang, H., Zheng, R. & Dempsey, A. A. The peripheral-blood transcriptome: new insights into disease and risk assessment. J. Lab. Clin. Med. 147, 126–132 (2007).
Hatefi, Y. The mitochondrial electron transport and oxidative phosphorylation system. Ann. Rev. Biochem. 54, 1015–1069.
Chen, C. Y. et al. Dietary Supplementation of High Levels of Saturated and Monounsaturated Fatty Acids to Ewes during Late Gestation Reduces Thermogenesis in Newborn Lambs by Depressing Fatty Acid Oxidation in Perirenal Brown Adipose Tissue. J. Nutr. 137, 43–48 (2007).
Keithly, J. I., Kott, R. W., Berardinelli, J. G., Moreaux, S. & Hatfield, P. G. Thermogenesis, blood metabolites and hormones, and growth of lambs born to ewes supplemented with algae-derived docosahexaenoic acid. J. Anim. Sci. 89, 4305–4313 (2008).
Mohammad, M. A., Hadsell, D. L. & Haymond, M. W. Gene regulation of UDP-galactose synthesis and transport: potential rate-limiting processes in initiation of milk production in humans. Am. J. Physiol. Endocrinol. Metab. 303, 365–376 (2012).
Drag, M. et al. Differential expression and co-expression gene networks reveal candidate biomarkers of boar taint in non-castrated pigs. Sci. rep. 7, 12205 (2017).
Salleh, S. M. et al. Gene co-expression networks from RNA sequencing of dairy cattle identifies genes and pathways affecting feed efficiency. BMC Bioinf. 19, 513 (2018).
AOAC. Official Methods of Analysis. Vol. 1. 15th ed. Association of Official Analytical Chemists, Arlington, VA (1990).
Van Soest, P. J., Robertson, J. B. & Lewis, B. A. Methods for Dietary Fiber, Neutral Detergent Fiber, and Nonstarch Polysaccharides in Relation to Animal Nutrition. J. Dairy Sci. 74, 3583–3597 (1991).
Martin, M. Cutadapt Removes Adapter Sequences From High-Throughput Sequencing Reads. EmboNet J ournal 17, https://doi.org/10.14806/ej.17.1.200 (2011).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 29, 15–21 (2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq. 2. Genome Biol. 15, 1–21 (2014).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics 9, 1 (2008).
Langfelder, P. & Horvath, S. Fast R functions for robust correlations and hierarchical clustering. J Stat Softw 46 (2012).
Acknowledgements
The research work reported in this paper was supported by PSR Regione Abruzzo 2007–2013 Misura 124 “IprOv: Caratterizzazione e miglioramento degli indici salutistici e sicurezza alimentare delle produzioni ovine tipiche abruzzesi a marchio di origine” (Principal investigator: G.M.).
Author information
Authors and Affiliations
Contributions
G.M. conceived the study and designed the research plan; M.I., A.I., C.M., F.B., E.D.A., L.G., D.G. and F.P. performed the experiments; M.I., A.I., F.C., S.E. and F.B. analyzed the data; M.I., A.I., F.C. and S.E. draft the manuscript that was revised by G.M.; all Authors revised and approved the final version of manuscript.
Corresponding author
Ethics declarations
Competing interests
Raw data associated with this project are deposited in the GenBank’s Sequence Read Archive (SRA) under the accession Bioproject Number PRJNA528905.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Iannaccone, M., Ianni, A., Contaldi, F. et al. Whole blood transcriptome analysis in ewes fed with hemp seed supplemented diet. Sci Rep 9, 16192 (2019). https://doi.org/10.1038/s41598-019-52712-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-52712-6
This article is cited by
-
Whole-plant corn silage improves rumen fermentation and growth performance of beef cattle by altering rumen microbiota
Applied Microbiology and Biotechnology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.